1. Introduction
Radiotherapy is one of the major modalities for the efficient treatment of cancer. Tumor therapy with heavy ions is increasingly implemented, as dose distributions that are increasingly confined to the neoplasm and spare healthy tissues can be achieved [
1]. Moreover, this dosimetric advantage is combined with the enhanced relative biological effectiveness (RBE) of this densely ionizing radiation (IR) [
2]. Ionizing particles deposit concentrated ionization events along their trajectories, thereby presumably causing clustered DNA lesions [
3,
4]. Compared to conventional photon-based radiotherapy this unique energy deposition pattern of charged ions substantially improves therapeutic efficacy [
5].
Repair of radiation-induced DSBs occurs within the higher-order chromatin structure [
6]. The nuclear localization of DSBs can be related to the condensation status of chromatin, with the two extremes euchromatin versus heterochromatin [
7]. Within the nucleus, chromatin is a highly dynamic structure and nucleosome modifications regulate DNA accessibility, flexibility and mobility for the recruitment of molecular machines ensuring transcription, replication and repair [
8]. The chromatin state may influence the forms of radiation-induced DNA damage and may also affect repair processing [
9]. Non-homologous end-joining (NHEJ) is widely considered as the prevalent DSB repair pathway, particularly in non-proliferating cells [
10]. NHEJ mediates the fast ligation of broken DNA ends and is initiated by the binding of the Ku70/Ku80 heterodimer to DSB termini to generate the binding scaffold for other NHEJ factors [
11].
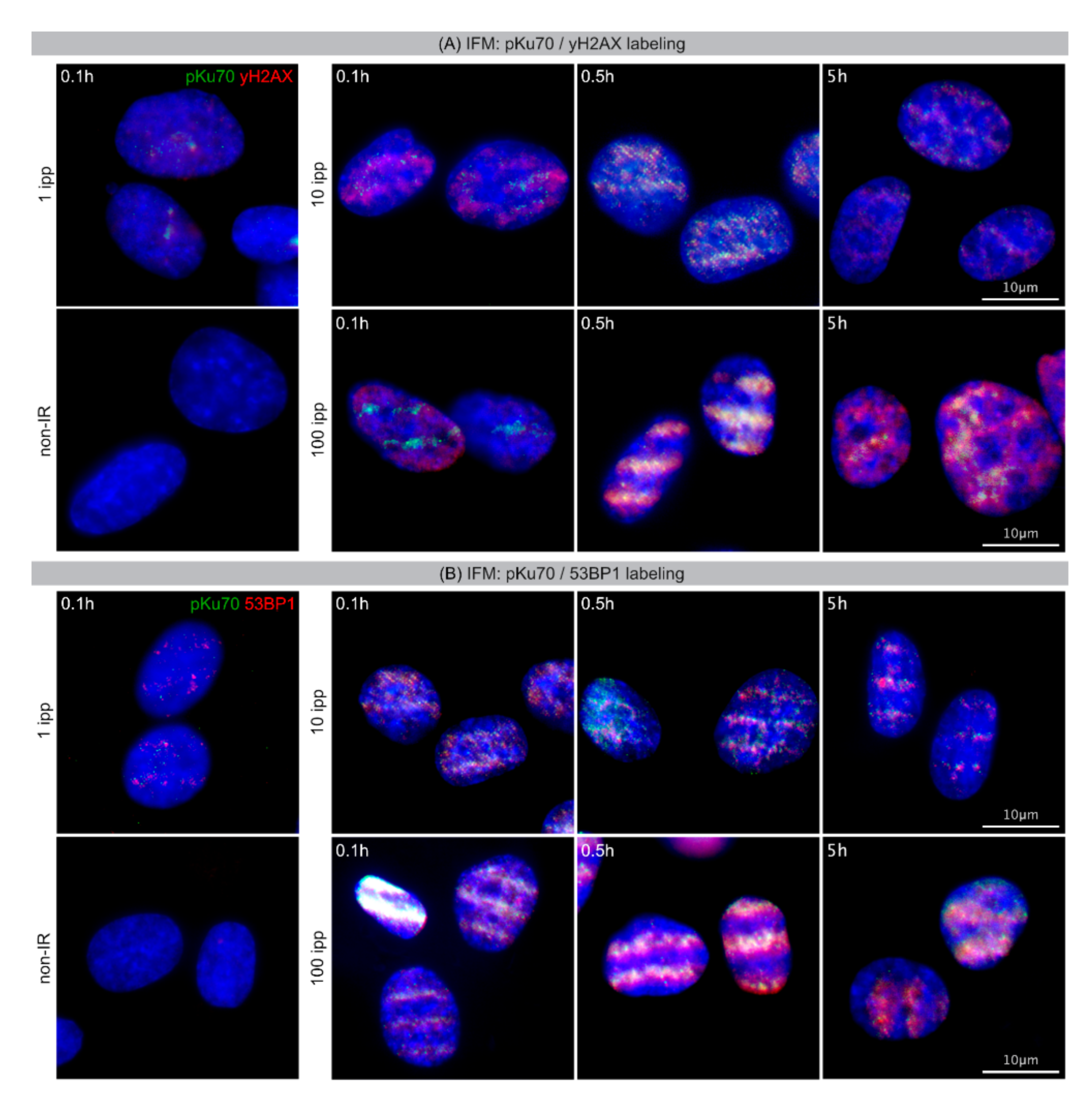
Currently, different experimental techniques exist to visualize DSBs. Phosphorylated H2AX (γH2AX) and p53 binding protein 1 (53BP1) appear in foci of DNA damage under immunofluorescence microscopy (IFM) and are considered surrogate markers of DSBs [
12]. However, the limited resolution of conventional light microscopy has restricted further insights into the detailed structure of repair foci. In recent years we established immunogold-labelling techniques to detect DNA-repair proteins within the chromatin ultrastructure using transmission electron microscopy (TEM) [
13,
14,
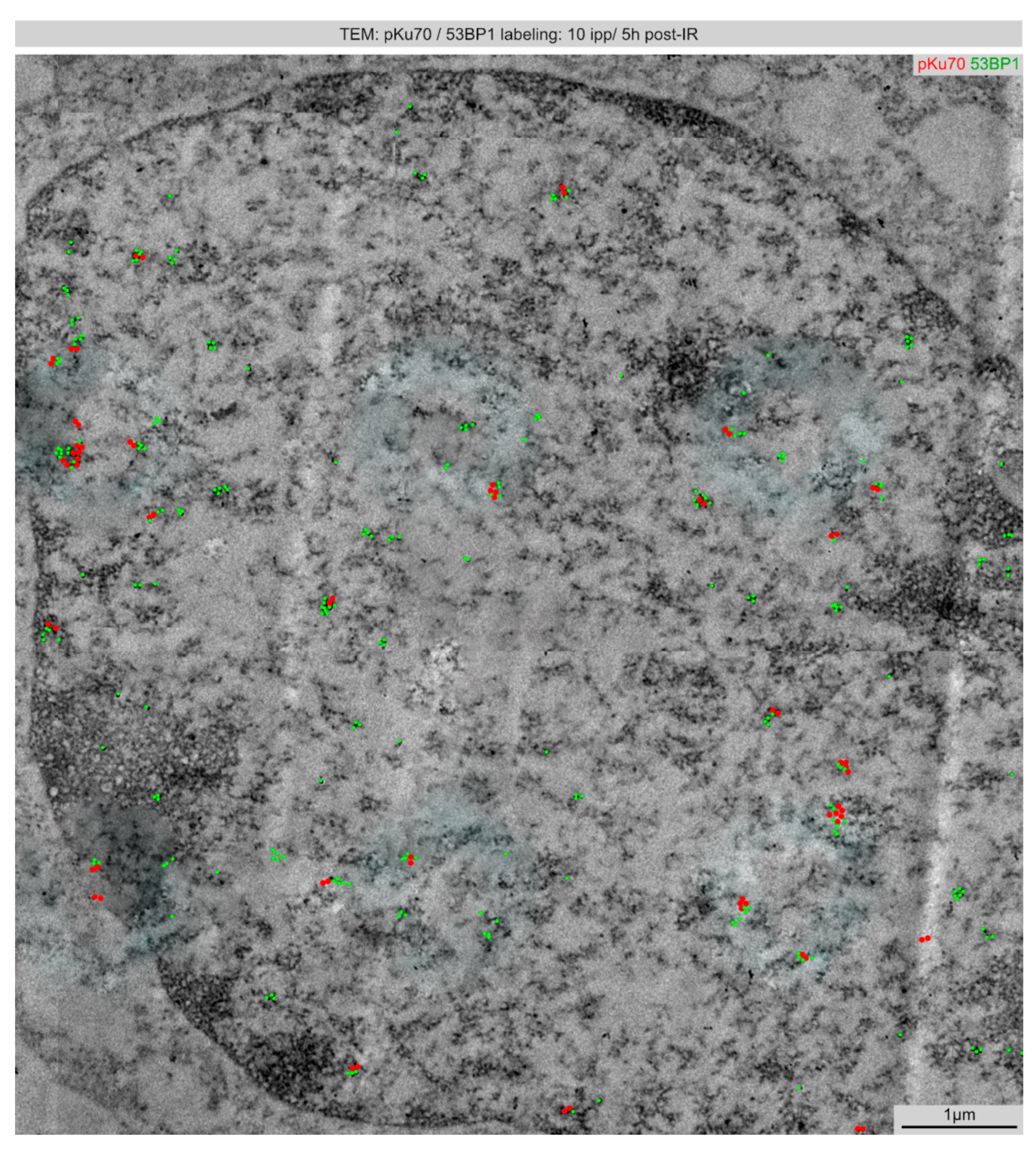
15]. The nanometer resolution of TEM permits the visualization of repair proteins at single-molecule level in different chromatin compartments. By labeling phosphorylated Ku70 (pKu70, phosphorylated at serine 6) which binds directly to broken DNA ends in preparation for re-joining, this TEM approach permits the reliable detection of unrepaired DSBs in electron-lucent euchromatin and electron-dense heterochromatin [
16]. In this study, we analyzed human fibroblasts, because the (epi)genome of these normal tissue cells is highly organized into chromosome territories, as well as into heterochromatin and euchromatin domains. The disruption of (epi)genomic integrity with the accumulation of various kinds of DNA damage due to deficient DNA repair capacities and epigenetic abnormalities constitute hallmarks of malignant tumors. Another important factor influencing radiation-induced DNA damage in tumor cells is the variable production of reactive oxygen species (ROS) as a consequence of hypoxic or metabolic stress. Due to these confounding factors in heterogeneous cancer cell populations we used human fibroblasts to analyze the fundamental chromatin-associated DNA repair mechanism.
The heavy-ion microbeam SNAKE (Superconducting Nanoprobe for Applied nuclear physics Experiments) enables the targeted irradiation of cell layers with defined numbers of carbon ions (1, 10 or 100 ions per point, ipp) focused to micrometer-sized stripes or spots [
17]. The sub-micrometer beam size allows the targeted irradiation of cellular substructures with high targeting accuracy [
18]. By scanning one beam stripe/spot after another, the heavy-ion microbeam can irradiate several cm
2 of cell layers with pre-defined patterns. Using the focusing system of SNAKE, single and multiple carbon ions were delivered to micrometer-sized beam stripes/spots in targeted cell nuclei and at defined time-points after exposure DNA lesions were visualized by different high-resolution microscopy approaches. The objective of this investigation was to study the influence of high local damage density on the clustering of DSBs in different chromatin regions.
3. Discussion
Particle radiotherapy is an important treatment modality for malignant tumors due to the favorable depth-dose profile and high relative biological effectiveness within the Bragg peak [
1]. Treatment planning of particle radiotherapy relies on biophysical modelling to predict the biological effects induced by diverse types of ion beams [
20]. These track-structure simulations enable the assessment of physical and chemical interactions of primary particles and their secondary electrons with the traversed medium. For a better understanding of the complex biological effects, however, these theoretical simulations need to be proven by experimental investigations in organic cells with physiological chromatin organization.
Previous reports using super-resolution fluorescence microscopy have shown the formation of clustered DNA repair foci along the particle tracks in heavy ion-irradiated cells [
21,
22]. Super-resolution fluorescence microscopy has managed to overcome the diffraction limit, thereby enabling the imaging of cellular structures with an improved resolution for many applications. At the same time, these super-resolution techniques retain the advantages of optical microscopy with regard to sample preservation, imaging flexibility and target specificity [
23]. Although these advanced optical technologies do not reach the resolution values of electron microscopy (EM), they allow the extraction of quantitative information on spatial distributions of proteins or other macromolecules within subcellular compartments. An early and powerful approach to obtain detailed information at the nanoscale level is using electrons instead of photons. Following the same physical principal, but with much smaller wavelength, electron microscopy is able to achieve clearly higher resolving power. However, transmission and scanning EM techniques are technically demanding, relatively costly and time-consuming. Importantly, due to principles of signal detection, the possibilities to specifically label and visualize multiple cellular structures or components in one specimen is still limited. Moreover, chemical fixation and contrasting procedures and/or physical sectioning render specimens vulnerable to artifacts and exclude the option to observe living cells or organisms at high resolution in their unperturbed state. However, due the highest resolution power the TEM approach permits nanoscale visualization of clustered DSBs and characterization of lesion complexity in the local chromatin ultrastructure [
13,
14,
15].
Accumulating experimental evidence indicates that the induction of radiation-induced damage to cellular DNA also depends on the local chromatin density [
24]. Ion beams crossing different chromatin regions may influence the radiation-induced DNA damage pattern in human cell nuclei and may cause damage to variable degrees with different functional importance, from affecting single nucleosomes to entire chromatin fiber loops [
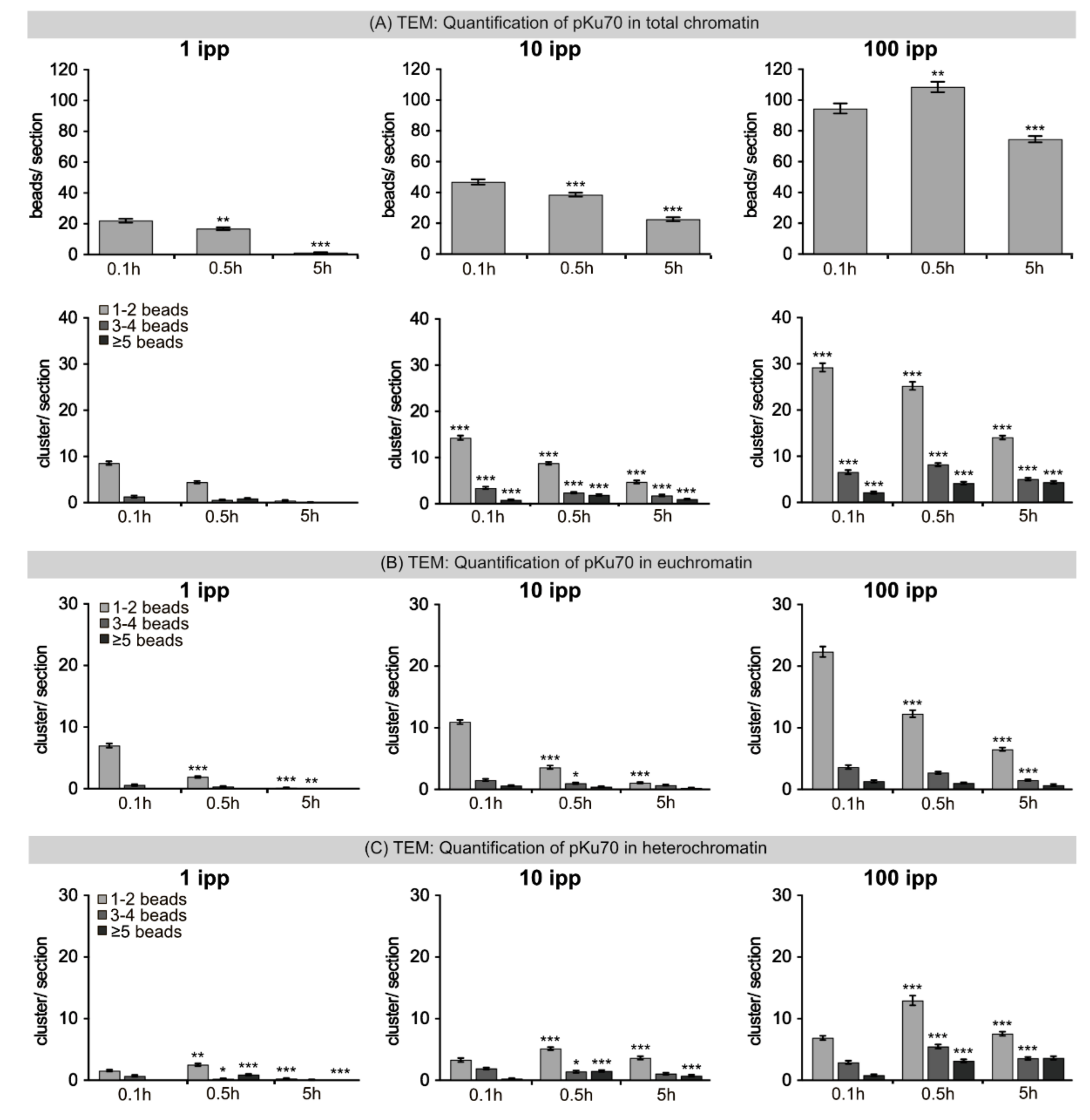
25]. This work focused on the induction of DSBs and their clustering, as DSBs are commonly considered the most relevant DNA lesions for the biological effects of IR. The defining feature of DSBs is the complete disruption of the molecular continuity. By affecting both DNA strands, DSBs lack the possibility to use the complementary, undamaged strand as template to restore the sequence in the damaged strand. In this experimental study with human fibroblasts, radiation-induced DNA damage after microbeam IR was analyzed by fluorescence and electron microscopy to obtain information at different scales. While fluorescence microscopic analysis highlights the nuclear distribution of DNA damage, electron microscopic study provides high-resolution information on the spatial clustering of DNA lesions in different chromatin domains at the micro- to nanoscale level. Defined numbers of carbon ions were delivered in micrometer-sized beam spots, so that the quantitative effects of single and multiple ions regarding the induction of damage to cellular DNA and its repair by the cell can be studied in small sub-regions of the nucleus. During DSB formation, each broken terminus binds a Ku heterodimer and the two heterodimers then associate to form a bridging complex that recruits additional NHEJ proteins. By labeling the activated Ku heterodimer, we were able to capture some structural aspects of DSB lesions and to define different levels of DSB complexity (pKu70 clusters with 1–2, 3–4 or ≥5 beads). Multiple DSBs in close vicinity (≤10 nm distance) were scored as clustered lesions and the number of DSBs in clusters is referred to as DSB complexity. The quantification of pKu70 clusters in different chromatin compartments indicates that more severe forms of damage complexity (clusters with 3–4 and ≥5 beads) were generated in heterochromatic regions. Moreover, the number of clustered DSBs in heterochromatin increased with time, suggesting that the repair of these clustered DSBs was not only delayed, but actually inherently inefficient. Our findings suggest that the increased DSB complexity in heterochromatic domains may interfere with damage processing and likely increase the risk for errors.
Labeling the activated Ku heterodimer permits visualization of unrepaired DSBs in close proximity within their chromatin environment at the micro- to nanoscale level by high-resolution electron microscopy. In previous studies we established TEM-based techniques to investigate specific DNA repair factors in the chromatin ultrastructure by post-embedding immunogold-labeling procedures for fixed cell and tissue specimens [
13,
14,
15]. The exploration of co-localization events in ultrathin sections is achieved by double-labeling with gold-conjugated secondary antibodies, characterized by differentially sized colloidal gold particles (6–10 nm in diameter). Scoring these gold-particles in electron-lucent and electron-dense regions of ultrathin nuclear sections permits quantification of specific repair factors in euchromatic and heterochromatic domains. Restriction endonucleases generate the simplest possible form of DSB, as they disrupt the continuity of the DNA molecule without chemically altering any of its constituent moieties. To verify our TEM-based detection technique for clustered DSBs, restriction endonuclease I-SceI was used to generate defined numbers of DSBs at specific sites of the genome [
26]. Following fixation and embedding of these I-
SceI transfected cells, ultra-thin sections were prepared and genomically-integrated restriction sites were subsequently labeled for pKu70. Screening of these nuclear sections by TEM revealed clusters of pKu70 dimers, which bind proportionally to the restriction cuts located at specific nucleotide sequences within 440 base pairs (bp) in length (
Supplementary Figure S2).
Due to varying electron absorption in euchromatin versus heterochromatin TEM permits to capture chromatin density distribution in the nucleus by different grey scales. In our TEM study gold-beads and clusters were counted and assigned to euchromatic or heterochromatic compartments based on their different chromatin densities; however, this approach do not reveal possible variations regarding structural organization. In previous experiments, we established gold-labeling of p53BP1 in combination with euchromatic or heterochromatic histone modifications (e.g., H3K9ac, H3K9me3) to analyze the impact of differently compacted chromatin on DNA repair [
13,
14,
15]. The nuclear distribution of these histone modifications correlated clearly with chromatin density defined by different grey levels. Further characterization of these radiation-induced lesions by gold-labeling other histone modifications or repair factors would be beneficial, but these examinations would require much time and effort, especially in this experimental setting.
Collectively, our findings support the idea that the biological significance of clustered DSBs is attributed to the cells inability to process them efficiently. Among DNA lesions, clustered DSBs represent the highest level of overall damage complexity, likely adding substantial difficulty to processing attempts. Additionally, DSBs have the greatest probability of causing adverse biological effects, including cell death, mutation as well as malignant transformations [
26]. The clustering of DSBs may, depending on overall organization, destabilize chromatin and jeopardize its potential to orchestrate efficient processing by NHEJ. Our experimental results improve the mechanistic knowledge of DSB clustering that underpin the biological effectiveness of particle irradiation and may have implications for radiotherapy with heavy ions as well as for radiation protection in space research.
4. Materials and Methods
Cell Culture: Human TERT fibroblasts (normal foreskin fibroblasts immortalized by ectopic expression of the catalytic subunit of the telomerase enzyme) were grown on cell carrier foils in Fibroblast Growth Medium (PromoCell) at 37 °C and 5% CO
2. Confluent cell layers on carrier foils were used for experiments. Cells transfected with I-
SceI plasmid were kindly provided by G. Iliakis [
26].
Ion irradiation: Carbon ion IR was performed at the Scanning Ion Microbeam SNAKE installed at the Munich 14 MV tandem accelerator. IR was performed by stripe/spot application of counted, individual carbon ions (1, 10, 100 ions per pulse, ipp) with initial beam energy in vacuum of 55 MeV. After leaving the beam transport vacuum through 7.5 µm Kapton foil, traversing an air gap (~30 µm) and cell carrier foil (~6 µm) the ions hit the cells. Due to energy loss in material the ions have an energy of 49.5 MeV and corresponding LET of 333 keV/µm. For matrix IR, beam spots (diameter ≈1 µm) were arranged in uniform 2D grids with 3µm and 4µm spacing for the x- and y-coordinates, respectively. Electrostatically scanning fields of 500 µm × 500 µm (for 1 and 10 ipp) or 300 µm × 300 µm (for 100 ipp) were irradiated with pre-defined matrices. In order to apply the right number of ions, PMT detectors were placed behind the samples enabling the counting of single ions. Once sufficient amounts of ions were applied the beam was stopped using fast electrostatic beam switch. The chosen IR geometry with defined ion numbers gives fluences F of 0.083 ions/µm2 for 1 ipp, 0.83 ions/µm2 for 10 ipp and 8.3 ions/µm2 for 100 ipp. By using the formula the dose D can be calculated using the density of water as approximation for cell density. This results in corresponding doses of 4.4 Gy, 44.2 Gy and 442.2 Gy, respectively.
For STED analysis non-focused small angle IR using SNAKE was used as described previously [
2]. Cells were cultured on 22 × 22 mm
2 coverslips and irradiated in air. During IR cells were covered with medium layers of ≈7.5 µm thickness. Energy loss led to final energy of 27 MeV and corresponding LET of 500 keV/µm. The center of the coverslips was irradiated with field sizes of 3.5 × 22 mm
2 under an angle of 9° and lasted only few seconds. IR was performed with 0.03 ions/µm
2 fluence and resulted in 2.4 Gy.
Immunofluorescence Microscopy: Cells on carrier foils were fixed in 4% paraformaldehyde and permeabilized in 0.2% Triton X-100. After blocking with 1% BSA in PBS overnight, cells were incubated with primary antibodies (γH2AX; 53BP1, Novus Biologicals, Wiesbaden Nordenstadt, Germany; pKu70, pSer5, Abcam, Cambridge, UK) followed by AlexaFluor-488 or AlexaFluor-568 secondary antibodies (Invitrogen, Karlsruhe, Germany). Finally, cells were mounted in VECTAshieldTM mounting medium with 4′,6-diamidino-2-phenylindole (DAPI, Vector Laboratories, Burlingame, CA, USA). Fluorescence images were captured using Nikon-Eclipse Ni fluorescence microscope equipped with charge-coupled-device camera and acquisition software (Nikon, Düsseldorf, Germany).
For STED microscopy cells were processed similarly and labeled with the same primary antibodies, but goat-anti-rabbit Abberior STAR 440SXP (Abberior Instruments, Göttingen, Germany) and goat-anti-mouse Chromeo 505 (Active Motif, La Hulpe, Belgium) were used as secondary antibodies. After embedding using ProlongGold antifade, cells were imaged using super-resolution optical CW STED microscope (Leica TCS SP 8 3X, Leica Microsystems, Wetzlar, Germany). Excitation laser wavelength of 470 nm was used for Abberior STAR 440SX and for Chromeo505 wavelength of 514 nm, both with power of ≈1 mW each. Detection ranges were 473–504 nm and 518–580 nm, respectively, with depletion laser at 592 nm (≈70 mW). STED laser was subdivided in lateral STED beam (40%) and axial STED beam (60% of the power).
Transmission Electron Microscopy: Cells were fixed with 2% paraformaldehyde and 0.05% glutaraldehyde in PBS. Fixed samples were dehydrated using increasing concentrations of ethanol and infiltrated with LR White resin overnight (Plano, Wetzlar, Germany). Subsequently, samples were embedded in fresh resin with accelerator at 37 °C until the resin polymerized. Ultrathin sections (≈70 nm) were prepared on Microtome Ultracut UCT (Leica, Wetzlar, Germany) with diamond knives (Diatome, Biel, Switzerland), gathered up on pioloform-coated nickel grids and processed for immunogold-labeling. Sections were placed on drops of blocking solution (Aurion, Wageningen, The Netherlands) to block non-specific staining. Afterwards sections were rinsed and incubated with primary antibodies (anti-53BP1, Novus Biologicals, Wiesbaden Nordenstadt, Germany; anti-pKu70, pSer5, Abcam, Cambridge, UK) overnight at 4 °C. After rinsing, secondary antibodies conjugated with 6 nm or 10 nm gold particles (Aurion, Wageningen, The Netherlands) were applied to the grids for 1 h. Sections were rinsed and fixed with 2% glutaraldehyde in PBS. All sections were stained with uranyl acetate and examined using Tecnai Biotwin™ transmission electron microscope (FEI, Eindhoven, The Netherlands). For quantification, single beads and bead clusters were counted in ≥25 randomly chosen nuclear sections. Detection, localization and quantification of gold beads and clusters were performed at the electron microscope by eye. Absolute numbers of gold particles were mapped over the entire nuclear compartment of 25 cross sections. Gold beads and clusters were counted and assigned to euchromatic or heterochromatic compartments based on chromatin density (defined by different gray scale levels). To facilitate the detection of beads and clusters in compact heterochromatin, regions of interest were overexposed by the electron beam to visualize electron-dense gold particles with their strong scattering. In TEM micrographs gold-particles were overlayed with colored dots.
Statistical analysis: Statistical analysis was performed using GraphPad Prism (San Diego, CA, USA) software. Data sets were analyzed for normal distribution using Shapiro–Wilk. Normally distributed data sets were subsequently analyzed using unpaired two-tailed Student’s t-test (equal variances under F-test) or Welch’s t-test (unequal variances under F-test). Non-parametric data were analyzed for statistical significance using two-tailed Mann–Whitney U test. A p < 0.05 was considered statistically significant, p < 0.01 as highly significant and p < 0.001 as extremely statistically significant.