Post-Embryonic Phase Transitions Mediated by Polycomb Repressive Complexes in Plants
Abstract
:1. Introduction
2. PRC1 and PRC2 in Plants
2.1. PRC2 in Plants
2.2. PRC1 in Plants
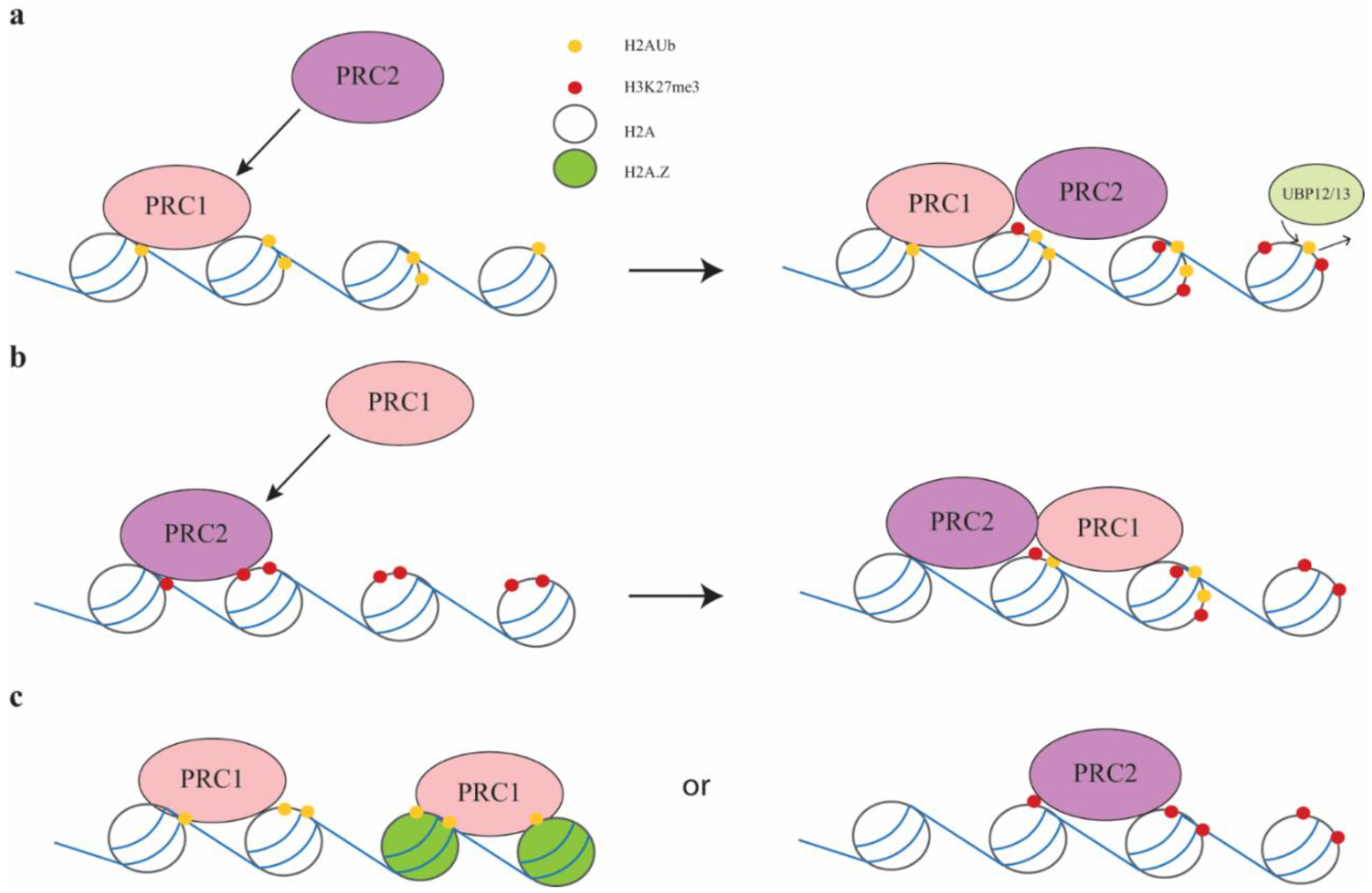
2.3. PRC1 and PRC2 Interactions in Plants
3. Post-Embryonic Developmental Transitions Mediated by PRC2 and PRC1
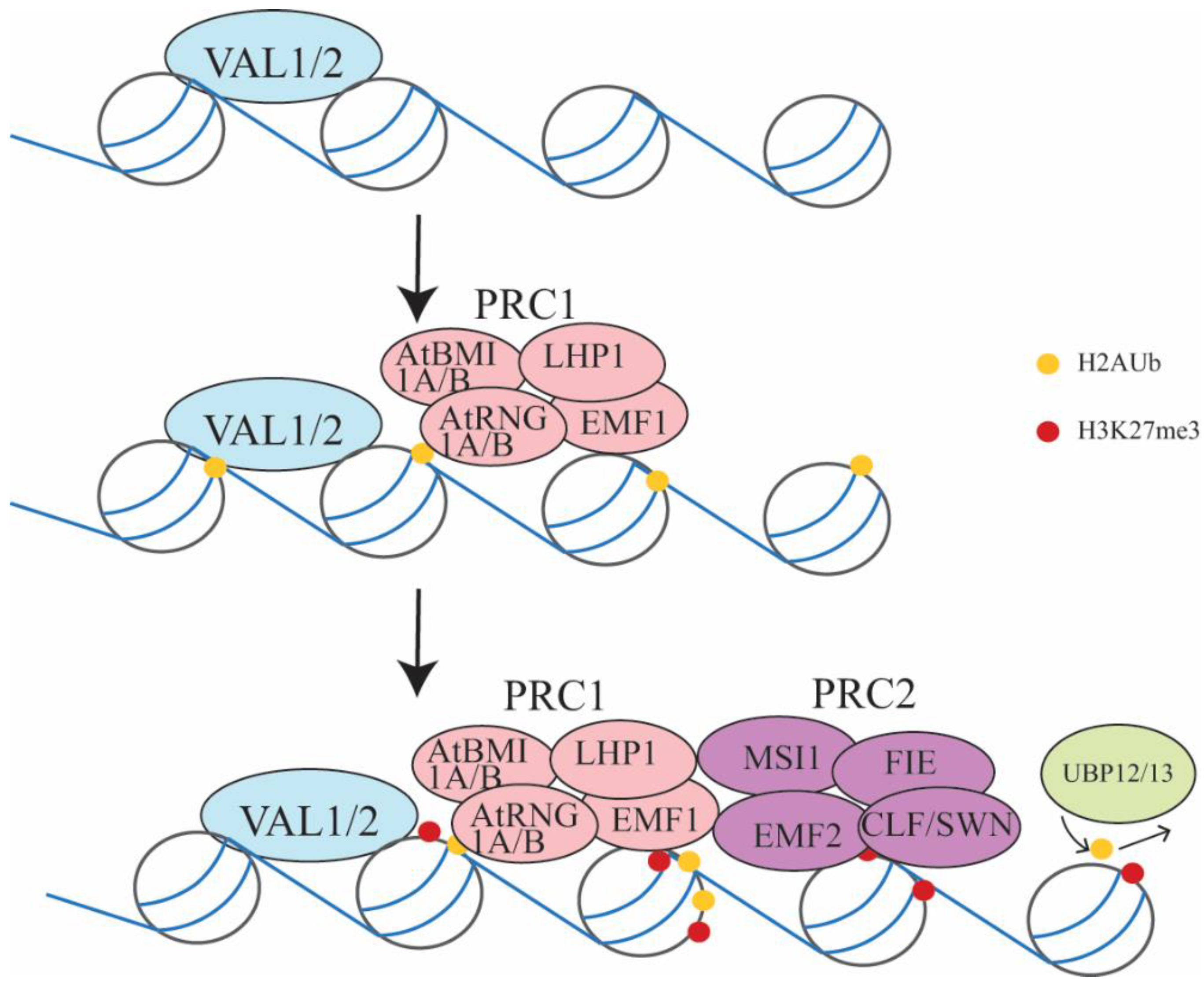
3.1. Embryo-to-Seedling Transition Mediated by PRC2 and PRC1
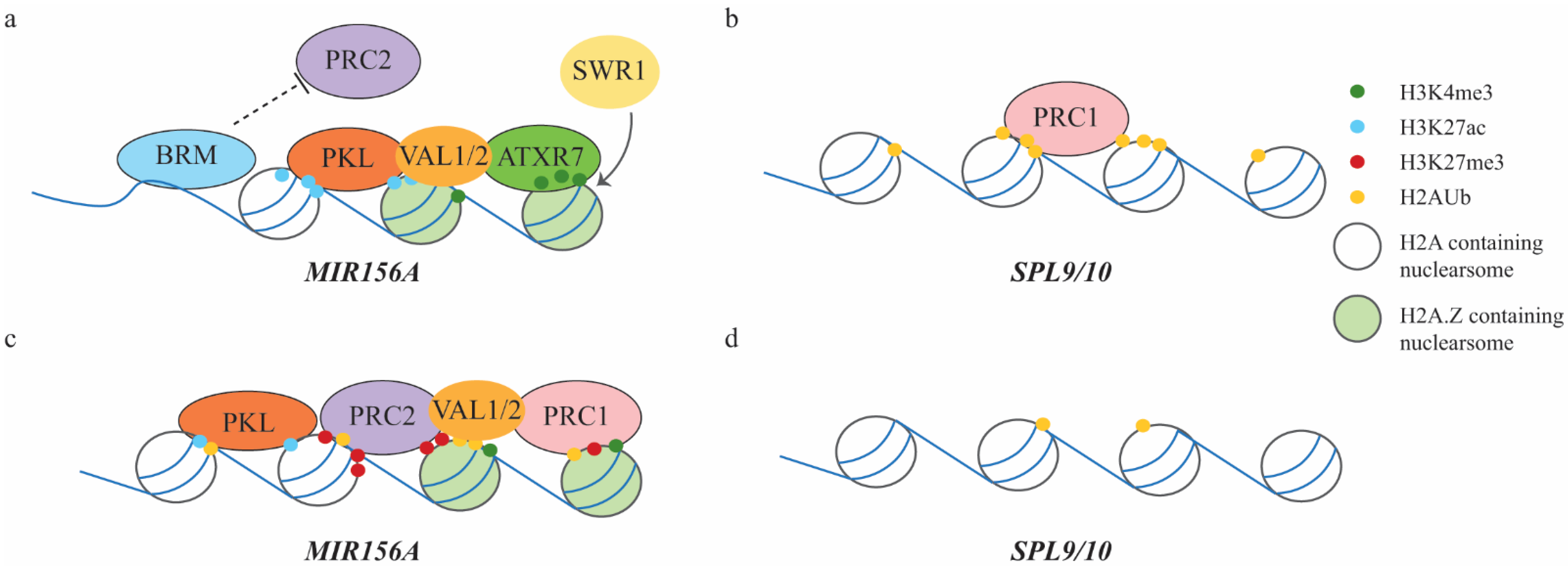
3.2. Juvenile-to-Adult Vegetative Phase Transition Mediated by PRC2 and PRC1
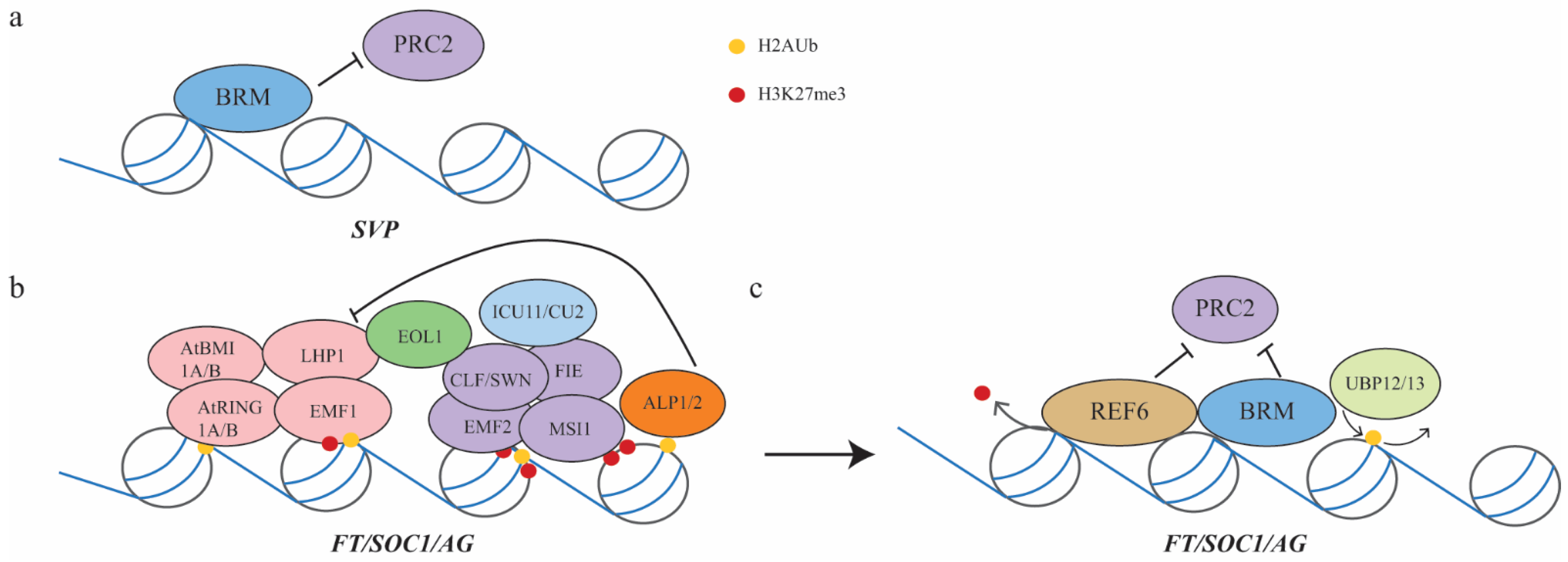
3.3. Vegetative-to-Reproductive Phase Transition Mediated by PRC2 and PRC1
3.4. How Is PRC2 Recruited and How Is H3K27me3 Spread?
4. PRC1 and PRC2 in Plants Other Than Arabidopsis
5. Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Baurle, I.; Dean, C. The timing of developmental transitions in plants. Cell 2006, 125, 655–664. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mariño-Ramírez, L.; Kann, M.G.; Shoemaker, B.A.; Landsman, D. Histone structure and nucleosome stability. Expert Rev. Proteom. 2005, 2, 719–729. [Google Scholar] [CrossRef]
- Gan, E.S.; Xu, Y.; Ito, T. Dynamics of H3K27me3 methylation and demethylation in plant development. Plant Signal. Behav. 2015, 10, e1027851. [Google Scholar] [CrossRef] [Green Version]
- Strahl, B.D.; Allis, C.D. The language of covalent histone modifications. Nature 2000, 403, 41–45. [Google Scholar] [CrossRef]
- Ahringer, J.; Gasser, S.M. Repressive Chromatin in Caenorhabditis elegans: Establishment, Composition, and Function. Genetics 2018, 208, 491–511. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.; Clarenz, O.; Cokus, S.; Bernatavichute, Y.V.; Pellegrini, M.; Goodrich, J.; Jacobsen, S.E. Whole-genome analysis of histone H3 lysine 27 trimethylation in Arabidopsis. PLoS Biol. 2007, 5, e129. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lafos, M.; Kroll, P.; Hohenstatt, M.L.; Thorpe, F.L.; Clarenz, O.; Schubert, D. Dynamic regulation of H3K27 trimethylation during Arabidopsis differentiation. PLoS Genet. 2011, 7, e1002040. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lewis, E.B. A gene complex controlling segmentation in Drosophila. Nature 1978, 276, 565–570. [Google Scholar] [CrossRef] [PubMed]
- Shao, Z.; Raible, F.; Mollaaghababa, R.; Guyon, J.R.; Wu, C.T.; Bender, W.; Kingston, R.E. Stabilization of chromatin structure by PRC1, a Polycomb complex. Cell 1999, 98, 37–46. [Google Scholar] [CrossRef] [Green Version]
- Kuzmichev, A.; Nishioka, K.; Erdjument-Bromage, H.; Tempst, P.; Reinberg, D. Histone methyltransferase activity associated with a human multiprotein complex containing the Enhancer of Zeste protein. Genes Dev. 2002, 16, 2893–2905. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, C.; Bratzel, F.; Hohmann, N.; Koch, M.; Turck, F.; Calonje, M. VAL- and AtBMI1-mediated H2Aub initiate the switch from embryonic to postgerminative growth in Arabidopsis. Curr. Biol. CB 2013, 23, 1324–1329. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mozgova, I.; Köhler, C.; Hennig, L. Keeping the gate closed: Functions of the polycomb repressive complex PRC2 in development. Plant J. Cell Mol. Biol. 2015, 83, 121–132. [Google Scholar] [CrossRef]
- Kleinmanns, J.A.; Schubert, D. Polycomb and Trithorax group protein-mediated control of stress responses in plants. Biol. Chem. 2014, 395, 1291–1300. [Google Scholar] [CrossRef]
- Bratzel, F.; López-Torrejón, G.; Koch, M.; Del Pozo, J.C.; Calonje, M. Keeping cell identity in Arabidopsis requires PRC1 Ring-finger homologs that catalyze H2A monoubiquitination. Curr. Biol. CB 2010, 20, 1853–1859. [Google Scholar] [CrossRef] [Green Version]
- Calonje, M.; Sanchez, R.; Chen, L.; Sung, Z.R. Embryonic Flower1 participates in polycomb group-mediated AG gene silencing in Arabidopsis. Plant Cell 2008, 20, 277–291. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chanvivattana, Y.; Bishopp, A.; Schubert, D.; Stock, C.; Moon, Y.H.; Sung, Z.R.; Goodrich, J. Interaction of Polycomb-group proteins controlling flowering in Arabidopsis. Development 2004, 131, 5263–5276. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oliva, M.; Butenko, Y.; Hsieh, T.F.; Hakim, O.; Katz, A.; Smorodinsky, N.I.; Michaeli, D.; Fischer, R.L.; Ohad, N. FIE, a nuclear PRC2 protein, forms cytoplasmic complexes in Arabidopsis thaliana. J. Exp. Bot. 2016, 67, 6111–6123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deng, W.; Buzas, D.M.; Ying, H.; Robertson, M.; Taylor, J.; Peacock, W.J.; Dennis, E.S.; Helliwell, C. Arabidopsis Polycomb Repressive Complex 2 binding sites contain putative GAGA factor binding motifs within coding regions of genes. BMC Genom. 2013, 14, 593. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xiao, J.; Jin, R.; Yu, X.; Shen, M.; Wagner, J.D.; Pai, A.; Song, C.; Zhuang, M.; Klasfeld, S.; He, C.; et al. Cis and trans determinants of epigenetic silencing by Polycomb repressive complex 2 in Arabidopsis. Nat. Genet. 2017, 49, 1546–1552. [Google Scholar] [CrossRef] [Green Version]
- Yoshida, N.; Yanai, Y.; Chen, L.; Kato, Y.; Hiratsuka, J.; Miwa, T.; Sung, Z.R.; Takahashi, S. Embryonic Flower2, a novel polycomb group protein homolog, mediates shoot development and flowering in Arabidopsis. Plant Cell 2001, 13, 2471–2481. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Helliwell, C.A.; Wood, C.C.; Robertson, M.; James Peacock, W.; Dennis, E.S. The Arabidopsis FLC protein interacts directly in vivo with SOC1 and FT chromatin and is part of a high-molecular-weight protein complex. Plant J. Cell Mol. Biol. 2006, 46, 183–192. [Google Scholar] [CrossRef] [PubMed]
- Hennig, L.; Bouveret, R.; Gruissem, W. MSI1-like proteins: An escort service for chromatin assembly and remodeling complexes. Trends Cell Biol. 2005, 15, 295–302. [Google Scholar] [CrossRef]
- de Lucas, M.; Pu, L.; Turco, G.; Gaudinier, A.; Morao, A.K.; Harashima, H.; Kim, D.; Ron, M.; Sugimoto, K.; Roudier, F.; et al. Transcriptional Regulation of Arabidopsis Polycomb Repressive Complex 2 Coordinates Cell-Type Proliferation and Differentiation. Plant Cell 2016, 28, 2616–2631. [Google Scholar] [CrossRef] [Green Version]
- Simon, J.A.; Kingston, R.E. Mechanisms of polycomb gene silencing: Knowns and unknowns. Nat. Rev. Mol. Cell Biol. 2009, 10, 697–708. [Google Scholar] [CrossRef]
- Zhang, X.; Germann, S.; Blus, B.J.; Khorasanizadeh, S.; Gaudin, V.; Jacobsen, S.E. The Arabidopsis LHP1 protein colocalizes with histone H3 Lys27 trimethylation. Nat. Struct. Mol. Biol. 2007, 14, 869–871. [Google Scholar] [CrossRef]
- Turck, F.; Roudier, F.; Farrona, S.; Martin-Magniette, M.L.; Guillaume, E.; Buisine, N.; Gagnot, S.; Martienssen, R.A.; Coupland, G.; Colot, V. Arabidopsis TFL2/LHP1 specifically associates with genes marked by trimethylation of histone H3 lysine 27. PLoS Genet. 2007, 3, e86. [Google Scholar] [CrossRef]
- Chen, D.; Molitor, A.; Liu, C.; Shen, W.H. The Arabidopsis PRC1-like ring-finger proteins are necessary for repression of embryonic traits during vegetative growth. Cell Res. 2010, 20, 1332–1344. [Google Scholar] [CrossRef] [Green Version]
- Chen, D.; Molitor, A.M.; Xu, L.; Shen, W.H. Arabidopsis PRC1 core component AtRING1 regulates stem cell-determining carpel development mainly through repression of class I KNOX genes. BMC Biol. 2016, 14, 112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Calonje, M. PRC1 marks the difference in plant PcG repression. Mol. Plant 2014, 7, 459–471. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bratzel, F.; Yang, C.; Angelova, A.; López-Torrejón, G.; Koch, M.; del Pozo, J.C.; Calonje, M. Regulation of the new Arabidopsis imprinted gene AtBMI1C requires the interplay of different epigenetic mechanisms. Mol. Plant 2012, 5, 260–269. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Exner, V.; Aichinger, E.; Shu, H.; Wildhaber, T.; Alfarano, P.; Caflisch, A.; Gruissem, W.; Köhler, C.; Hennig, L. The chromodomain of LIKE HETEROCHROMATIN PROTEIN 1 is essential for H3K27me3 binding and function during Arabidopsis development. PLoS ONE 2009, 4, e5335. [Google Scholar] [CrossRef] [Green Version]
- Aubert, D.; Chen, L.; Moon, Y.H.; Martin, D.; Castle, L.A.; Yang, C.H.; Sung, Z.R. EMF1, a novel protein involved in the control of shoot architecture and flowering in Arabidopsis. Plant Cell 2001, 13, 1865–1875. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gómez-Zambrano, Á.; Merini, W.; Calonje, M. The repressive role of Arabidopsis H2A.Z in transcriptional regulation depends on AtBMI1 activity. Nat. Commun. 2019, 10, 2828. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Berke, L.; Snel, B. The plant Polycomb repressive complex 1 (PRC1) existed in the ancestor of seed plants and has a complex duplication history. BMC Evol. Biol. 2015, 15, 44. [Google Scholar] [CrossRef] [Green Version]
- Merini, W.; Romero-Campero, F.J.; Gomez-Zambrano, A.; Zhou, Y.; Turck, F.; Calonje, M. The Arabidopsis Polycomb Repressive Complex 1 (PRC1) Components AtBMI1A, B, and C Impact Gene Networks throughout All Stages of Plant Development. Plant Physiol. 2017, 173, 627–641. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Derkacheva, M.; Steinbach, Y.; Wildhaber, T.; Mozgová, I.; Mahrez, W.; Nanni, P.; Bischof, S.; Gruissem, W.; Hennig, L. Arabidopsis MSI1 connects LHP1 to PRC2 complexes. EMBO J. 2013, 32, 2073–2085. [Google Scholar] [CrossRef] [Green Version]
- Cao, R.; Wang, L.; Wang, H.; Xia, L.; Erdjument-Bromage, H.; Tempst, P.; Jones, R.S.; Zhang, Y. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 2002, 298, 1039–1043. [Google Scholar] [CrossRef] [Green Version]
- Blackledge, N.P.; Farcas, A.M.; Kondo, T.; King, H.W.; McGouran, J.F.; Hanssen, L.L.P.; Ito, S.; Cooper, S.; Kondo, K.; Koseki, Y.; et al. Variant PRC1 complex-dependent H2A ubiquitylation drives PRC2 recruitment and polycomb domain formation. Cell 2014, 157, 1445–1459. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, Y.; Romero-Campero, F.J.; Gómez-Zambrano, Á.; Turck, F.; Calonje, M. H2A monoubiquitination in Arabidopsis thaliana is generally independent of LHP1 and PRC2 activity. Genome Biol. 2017, 18, 69. [Google Scholar] [CrossRef] [Green Version]
- Kralemann, L.E.M.; Liu, S.; Trejo-Arellano, M.S.; Muñoz-Viana, R.; Köhler, C.; Hennig, L. Removal of H2Aub1 by ubiquitin-specific proteases 12 and 13 is required for stable Polycomb-mediated gene repression in Arabidopsis. Genome Biol. 2020, 21, 144. [Google Scholar] [CrossRef] [PubMed]
- Derkacheva, M.; Liu, S.; Figueiredo, D.D.; Gentry, M.; Mozgova, I.; Nanni, P.; Tang, M.; Mannervik, M.; Köhler, C.; Hennig, L. H2A deubiquitinases UBP12/13 are part of the Arabidopsis polycomb group protein system. Nat. Plants 2016, 2, 16126. [Google Scholar] [CrossRef] [PubMed]
- Iovino, N.; Ciabrelli, F.; Cavalli, G. PRC2 controls Drosophila oocyte cell fate by repressing cell cycle genes. Dev. Cell 2013, 26, 431–439. [Google Scholar] [CrossRef] [Green Version]
- Birve, A.; Sengupta, A.K.; Beuchle, D.; Larsson, J.; Kennison, J.A.; Rasmuson-Lestander, A.; Müller, J. Su(z)12, a novel Drosophila Polycomb group gene that is conserved in vertebrates and plants. Development 2001, 128, 3371–3379. [Google Scholar] [CrossRef]
- Bouyer, D.; Roudier, F.; Heese, M.; Andersen, E.D.; Gey, D.; Nowack, M.K.; Goodrich, J.; Renou, J.P.; Grini, P.E.; Colot, V.; et al. Polycomb repressive complex 2 controls the embryo-to-seedling phase transition. PLoS Genet. 2011, 7, e1002014. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, X.; Lim, M.H.; Pelletier, J.; Tang, M.; Nguyen, V.; Keller, W.A.; Tsang, E.W.; Wang, A.; Rothstein, S.J.; Harada, J.J.; et al. Synergistic repression of the embryonic programme by Set Domain Group 8 and Embryonic Flower 2 in Arabidopsis seedlings. J. Exp. Bot. 2012, 63, 1391–1404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Köhler, C.; Hennig, L.; Bouveret, R.; Gheyselinck, J.; Grossniklaus, U.; Gruissem, W. Arabidopsis MSI1 is a component of the MEA/FIE Polycomb group complex and required for seed development. EMBO J. 2003, 22, 4804–4814. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hennig, L.; Taranto, P.; Walser, M.; Schönrock, N.; Gruissem, W. Arabidopsis MSI1 is required for epigenetic maintenance of reproductive development. Development 2003, 130, 2555–2565. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, L.; Shen, W.H. Polycomb silencing of KNOX genes confines shoot stem cell niches in Arabidopsis. Curr. Biol. CB 2008, 18, 1966–1971. [Google Scholar] [CrossRef] [Green Version]
- Molitor, A.M.; Bu, Z.; Yu, Y.; Shen, W.H. Arabidopsis AL PHD-PRC1 complexes promote seed germination through H3K4me3-to-H3K27me3 chromatin state switch in repression of seed developmental genes. PLoS Genet. 2014, 10, e1004091. [Google Scholar] [CrossRef] [Green Version]
- Peng, L.; Wang, L.; Zhang, Y.; Dong, A.; Shen, W.H.; Huang, Y. Structural Analysis of the Arabidopsis AL2-PAL and PRC1 Complex Provides Mechanistic Insight into Active-to-Repressive Chromatin State Switch. J. Mol. Biol. 2018, 430, 4245–4259. [Google Scholar] [CrossRef]
- Feng, J.; Chen, D.; Berr, A.; Shen, W.H. ZRF1 Chromatin Regulators Have Polycomb Silencing and Independent Roles in Development. Plant Physiol. 2016, 172, 1746–1759. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Bishop, B.; Ringenberg, W.; Muir, W.M.; Ogas, J. The CHD3 remodeler PICKLE associates with genes enriched for trimethylation of histone H3 lysine 27. Plant Physiol. 2012, 159, 418–432. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Rider, S.D., Jr.; Henderson, J.T.; Fountain, M.; Chuang, K.; Kandachar, V.; Simons, A.; Edenberg, H.J.; Romero-Severson, J.; Muir, W.M.; et al. The CHD3 remodeler PICKLE promotes trimethylation of histone H3 lysine 27. J. Biol. Chem. 2008, 283, 22637–22648. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, H.C.; Chuang, K.; Henderson, J.T.; Rider, S.D., Jr.; Bai, Y.; Zhang, H.; Fountain, M.; Gerber, J.; Ogas, J. PICKLE acts during germination to repress expression of embryonic traits. Plant J. Cell Mol. Biol. 2005, 44, 1010–1022. [Google Scholar] [CrossRef] [PubMed]
- Aichinger, E.; Villar, C.B.; Di Mambro, R.; Sabatini, S.; Köhler, C. The CHD3 chromatin remodeler PICKLE and polycomb group proteins antagonistically regulate meristem activity in the Arabidopsis root. Plant Cell 2011, 23, 1047–1060. [Google Scholar] [CrossRef] [Green Version]
- Aichinger, E.; Villar, C.B.; Farrona, S.; Reyes, J.C.; Hennig, L.; Köhler, C. CHD3 proteins and polycomb group proteins antagonistically determine cell identity in Arabidopsis. PLoS Genet. 2009, 5, e1000605. [Google Scholar] [CrossRef] [Green Version]
- Poethig, R.S. The past, present, and future of vegetative phase change. Plant Physiol. 2010, 154, 541–544. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Poethig, R.S. Phase change and the regulation of developmental timing in plants. Science 2003, 301, 334–336. [Google Scholar] [CrossRef]
- Poethig, R.S. Vegetative phase change and shoot maturation in plants. Curr. Top. Dev. Biol. 2013, 105, 125–152. [Google Scholar] [CrossRef] [Green Version]
- Wu, G.; Park, M.Y.; Conway, S.R.; Wang, J.W.; Weigel, D.; Poethig, R.S. The sequential action of miR156 and miR172 regulates developmental timing in Arabidopsis. Cell 2009, 138, 750–759. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, M.; Hu, T.; Zhao, J.; Park, M.Y.; Earley, K.W.; Wu, G.; Yang, L.; Poethig, R.S. Developmental Functions of miR156-Regulated Squamosa Promoter Binding Protein-Like (SPL) Genes in Arabidopsis thaliana. PLoS Genet. 2016, 12, e1006263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, J.; Xu, M.; Willmann, M.R.; McCormick, K.; Hu, T.; Yang, L.; Starker, C.G.; Voytas, D.F.; Meyers, B.C.; Poethig, R.S. Threshold-dependent repression of SPL gene expression by miR156/miR157 controls vegetative phase change in Arabidopsis thaliana. PLoS Genet. 2018, 14, e1007337. [Google Scholar] [CrossRef] [Green Version]
- Mao, Y.B.; Liu, Y.Q.; Chen, D.Y.; Chen, F.Y.; Fang, X.; Hong, G.J.; Wang, L.J.; Wang, J.W.; Chen, X.Y. Jasmonate response decay and defense metabolite accumulation contributes to age-regulated dynamics of plant insect resistance. Nat. Commun. 2017, 8, 13925. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Zhou, L.; Shi, H.; Chern, M.; Yu, H.; Yi, H.; He, M.; Yin, J.; Zhu, X.; Li, Y.; et al. A single transcription factor promotes both yield and immunity in rice. Science 2018, 361, 1026–1028. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, N.; Niu, Q.W.; Ng, K.H.; Chua, N.H. The role of miR156/SPLs modules in Arabidopsis lateral root development. Plant J. Cell Mol. Biol. 2015, 83, 673–685. [Google Scholar] [CrossRef]
- Hyun, Y.; Richter, R.; Coupland, G. Competence to Flower: Age-Controlled Sensitivity to Environmental Cues. Plant Physiol. 2017, 173, 36–46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hyun, Y.; Vincent, C.; Tilmes, V.; Bergonzi, S.; Kiefer, C.; Richter, R.; Martinez-Gallegos, R.; Severing, E.; Coupland, G. A regulatory circuit conferring varied flowering response to cold in annual and perennial plants. Science 2019, 363, 409–412. [Google Scholar] [CrossRef]
- Stief, A.; Altmann, S.; Hoffmann, K.; Pant, B.D.; Scheible, W.R.; Bäurle, I. Arabidopsis miR156 Regulates Tolerance to Recurring Environmental Stress through SPL Transcription Factors. Plant Cell 2014, 26, 1792–1807. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.W.; Park, M.Y.; Wang, L.J.; Koo, Y.; Chen, X.Y.; Weigel, D.; Poethig, R.S. miRNA control of vegetative phase change in trees. PLoS Genet. 2011, 7, e1002012. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leichty, A.R.; Poethig, R.S. Development and evolution of age-dependent defenses in ant-acacias. Proc. Natl. Acad. Sci. USA 2019, 116, 15596–15601. [Google Scholar] [CrossRef] [Green Version]
- Yu, S.; Cao, L.; Zhou, C.M.; Zhang, T.Q.; Lian, H.; Sun, Y.; Wu, J.; Huang, J.; Wang, G.; Wang, J.W. Sugar is an endogenous cue for juvenile-to-adult phase transition in plants. eLife 2013, 2, e00269. [Google Scholar] [CrossRef]
- Yang, L.; Xu, M.; Koo, Y.; He, J.; Poethig, R.S. Sugar promotes vegetative phase change in Arabidopsis thaliana by repressing the expression of MIR156A and MIR156C. eLife 2013, 2, e00260. [Google Scholar] [CrossRef]
- Fouracre, J.P.; Poethig, R.S. Role for the shoot apical meristem in the specification of juvenile leaf identity in Arabidopsis. Proc. Natl. Acad. Sci. USA 2019, 116, 10168–10177. [Google Scholar] [CrossRef] [Green Version]
- Xie, Y.; Liu, Y.; Wang, H.; Ma, X.; Wang, B.; Wu, G.; Wang, H. Phytochrome-interacting factors directly suppress MIR156 expression to enhance shade-avoidance syndrome in Arabidopsis. Nat. Commun. 2017, 8, 348. [Google Scholar] [CrossRef] [Green Version]
- Xu, M.; Hu, T.; Smith, M.R.; Poethig, R.S. Epigenetic Regulation of Vegetative Phase Change in Arabidopsis. Plant Cell 2016, 28, 28–41. [Google Scholar] [CrossRef] [Green Version]
- Xu, M.; Leichty, A.R.; Hu, T.; Poethig, R.S. H2A.Z promotes the transcription of MIR156A and MIR156C in Arabidopsis by facilitating the deposition of H3K4me3. Development 2018, 145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, Y.; Guo, C.; Zhou, B.; Li, C.; Wang, H.; Zheng, B.; Ding, H.; Zhu, Z.; Peragine, A.; Cui, Y.; et al. Regulation of Vegetative Phase Change by SWI2/SNF2 Chromatin Remodeling ATPase BRAHMA. Plant Physiol. 2016, 172, 2416–2428. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Picó, S.; Ortiz-Marchena, M.I.; Merini, W.; Calonje, M. Deciphering the Role of Polycomb Repressive Complex1 Variants in Regulating the Acquisition of Flowering Competence in Arabidopsis. Plant Physiol. 2015, 168, 1286–1297. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Wang, Z.; Hu, Y.; Cao, Y.; Ma, L. Polycomb Group Proteins RING1A and RING1B Regulate the Vegetative Phase Transition in Arabidopsis. Front. Plant Sci. 2017, 8, 867. [Google Scholar] [CrossRef] [Green Version]
- Petesch, S.J.; Lis, J.T. Overcoming the nucleosome barrier during transcript elongation. Trends Genet. 2012, 28, 285–294. [Google Scholar] [CrossRef] [Green Version]
- Li, C.; Chen, C.; Gao, L.; Yang, S.; Nguyen, V.; Shi, X.; Siminovitch, K.; Kohalmi, S.E.; Huang, S.; Wu, K.; et al. The Arabidopsis SWI2/SNF2 chromatin Remodeler BRAHMA regulates polycomb function during vegetative development and directly activates the flowering repressor gene SVP. PLoS Genet. 2015, 11, e1004944. [Google Scholar] [CrossRef] [Green Version]
- Tamada, Y.; Yun, J.Y.; Woo, S.C.; Amasino, R.M. Arabidopsis Trithorax-Related7 is required for methylation of lysine 4 of histone H3 and for transcriptional activation of FLOWERING LOCUS C. Plant Cell 2009, 21, 3257–3269. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fouracre, J.P.; He, J.; Chen, V.J.; Sidoli, S.; Poethig, R.S. VAL genes regulate vegetative phase change via miR156-dependent and independent mechanisms. PLoS Genet. 2021, 17, e1009626. [Google Scholar] [CrossRef]
- Qüesta, J.I.; Song, J.; Geraldo, N.; An, H.; Dean, C. Arabidopsis transcriptional repressor VAL1 triggers Polycomb silencing at FLC during vernalization. Science 2016, 353, 485–488. [Google Scholar] [CrossRef]
- He, Y. Chromatin regulation of flowering. Trends Plant Sci. 2012, 17, 556–562. [Google Scholar] [CrossRef] [PubMed]
- Costa, S.; Dean, C. Storing memories: The distinct phases of Polycomb-mediated silencing of Arabidopsis FLC. Biochem. Soc. Trans. 2019, 47, 1187–1196. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Whittaker, C.; Dean, C. The FLC Locus: A Platform for Discoveries in Epigenetics and Adaptation. Annu. Rev. Cell Dev. Biol. 2017, 33, 555–575. [Google Scholar] [CrossRef]
- Kim, S.Y.; Lee, J.; Eshed-Williams, L.; Zilberman, D.; Sung, Z.R. EMF1 and PRC2 cooperate to repress key regulators of Arabidopsis development. PLoS Genet. 2012, 8, e1002512. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, S.Y.; Zhu, T.; Sung, Z.R. Epigenetic regulation of gene programs by EMF1 and EMF2 in Arabidopsis. Plant Physiol. 2010, 152, 516–528. [Google Scholar] [CrossRef] [Green Version]
- Schönrock, N.; Bouveret, R.; Leroy, O.; Borghi, L.; Köhler, C.; Gruissem, W.; Hennig, L. Polycomb-group proteins repress the floral activator AGL19 in the FLC-independent vernalization pathway. Genes Dev. 2006, 20, 1667–1678. [Google Scholar] [CrossRef] [Green Version]
- Goodrich, J.; Puangsomlee, P.; Martin, M.; Long, D.; Meyerowitz, E.M.; Coupland, G. A Polycomb-group gene regulates homeotic gene expression in Arabidopsis. Nature 1997, 386, 44–51. [Google Scholar] [CrossRef]
- Jiang, D.; Wang, Y.; Wang, Y.; He, Y. Repression of FLOWERING LOCUS C and FLOWERING LOCUS T by the Arabidopsis Polycomb repressive complex 2 components. PLoS ONE 2008, 3, e3404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shu, J.; Chen, C.; Thapa, R.K.; Bian, S.; Nguyen, V.; Yu, K.; Yuan, Z.C.; Liu, J.; Kohalmi, S.E.; Li, C.; et al. Genome-wide occupancy of histone H3K27 methyltransferases CURLY LEAF and SWINGER in Arabidopsis seedlings. Plant Direct. 2019, 3, e00100. [Google Scholar] [CrossRef] [Green Version]
- Lu, F.; Cui, X.; Zhang, S.; Jenuwein, T.; Cao, X. Arabidopsis REF6 is a histone H3 lysine 27 demethylase. Nat. Genet. 2011, 43, 715–719. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Gu, L.; Gao, L.; Chen, C.; Wei, C.Q.; Qiu, Q.; Chien, C.W.; Wang, S.; Jiang, L.; Ai, L.F.; et al. Concerted genomic targeting of H3K27 demethylase REF6 and chromatin-remodeling ATPase BRM in Arabidopsis. Nat. Genet. 2016, 48, 687–693. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.; Lu, F.; Qiu, Q.; Zhou, B.; Gu, L.; Zhang, S.; Kang, Y.; Cui, X.; Ma, X.; Yao, Q.; et al. REF6 recognizes a specific DNA sequence to demethylate H3K27me3 and regulate organ boundary formation in Arabidopsis. Nat. Genet. 2016, 48, 694–699. [Google Scholar] [CrossRef] [PubMed]
- Liang, S.C.; Hartwig, B.; Perera, P.; Mora-García, S.; de Leau, E.; Thornton, H.; de Lima Alves, F.; Rappsilber, J.; Yang, S.; James, G.V.; et al. Kicking against the PRCs—A Domesticated Transposase Antagonises Silencing Mediated by Polycomb Group Proteins and is an Accessory Component of Polycomb Repressive Complex 2. PLoS Genet. 2015, 11, e1005660. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Tergemina, E.; Cui, H.; Förderer, A.; Hartwig, B.; Velikkakam James, G.; Schneeberger, K.; Turck, F. Ctf4-related protein recruits LHP1-PRC2 to maintain H3K27me3 levels in dividing cells in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2017, 114, 4833–4838. [Google Scholar] [CrossRef] [Green Version]
- Bloomer, R.H.; Hutchison, C.E.; Bäurle, I.; Walker, J.; Fang, X.; Perera, P.; Velanis, C.N.; Gümüs, S.; Spanos, C.; Rappsilber, J.; et al. The Arabidopsis epigenetic regulator ICU11 as an accessory protein of Polycomb Repressive Complex 2. Proc. Natl. Acad. Sci. USA 2020, 117, 16660–16666. [Google Scholar] [CrossRef] [PubMed]
- Mateo-Bonmatí, E.; Esteve-Bruna, D.; Juan-Vicente, L.; Nadi, R.; Candela, H.; Lozano, F.M.; Ponce, M.R.; Pérez-Pérez, J.M.; Micol, J.L. INCURVATA11 and CUPULIFORMIS2 Are Redundant Genes That Encode Epigenetic Machinery Components in Arabidopsis. Plant Cell 2018, 30, 1596–1616. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Y.; Wang, Y.; Krause, K.; Yang, T.; Dongus, J.A.; Zhang, Y.; Turck, F. Telobox motifs recruit CLF/SWN-PRC2 for H3K27me3 deposition via TRB factors in Arabidopsis. Nat. Genet. 2018, 50, 638–644. [Google Scholar] [CrossRef] [PubMed]
- Angel, A.; Song, J.; Dean, C.; Howard, M. A Polycomb-based switch underlying quantitative epigenetic memory. Nature 2011, 476, 105–108. [Google Scholar] [CrossRef]
- Coustham, V.; Li, P.; Strange, A.; Lister, C.; Song, J.; Dean, C. Quantitative modulation of polycomb silencing underlies natural variation in vernalization. Science 2012, 337, 584–587. [Google Scholar] [CrossRef] [PubMed]
- De Lucia, F.; Crevillen, P.; Jones, A.M.; Greb, T.; Dean, C. A PHD-polycomb repressive complex 2 triggers the epigenetic silencing of FLC during vernalization. Proc. Natl. Acad. Sci. USA 2008, 105, 16831–16836. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, H.; Berry, S.; Olsson, T.S.G.; Hartley, M.; Howard, M.; Dean, C. Distinct phases of Polycomb silencing to hold epigenetic memory of cold in Arabidopsis. Science 2017, 357, 1142–1145. [Google Scholar] [CrossRef]
- Strejčková, B.; Čegan, R.; Pecinka, A.; Milec, Z.; Šafář, J. Identification of polycomb repressive complex 1 and 2 core components in hexaploid bread wheat. BMC Plant Biol. 2020, 20, 175. [Google Scholar] [CrossRef]
- Chen, D.H.; Huang, Y.; Ruan, Y.; Shen, W.H. The evolutionary landscape of PRC1 core components in green lineage. Planta 2016, 243, 825–846. [Google Scholar] [CrossRef]
- Parihar, V.; Arya, D.; Walia, A.; Tyagi, V.; Dangwal, M.; Verma, V.; Khurana, R.; Boora, N.; Kapoor, S.; Kapoor, M. Functional characterization of LIKE HETEROCHROMATIN PROTEIN 1 in the moss Physcomitrella patens: Its conserved protein interactions in land plants. Plant J. Cell Mol. Biol. 2019, 97, 221–239. [Google Scholar] [CrossRef]
- Shaver, S.; Casas-Mollano, J.A.; Cerny, R.L.; Cerutti, H. Origin of the polycomb repressive complex 2 and gene silencing by an E(z) homolog in the unicellular alga Chlamydomonas. Epigenetics 2010, 5, 301–312. [Google Scholar] [CrossRef] [Green Version]
- Mosquna, A.; Katz, A.; Decker, E.L.; Rensing, S.A.; Reski, R.; Ohad, N. Regulation of stem cell maintenance by the Polycomb protein FIE has been conserved during land plant evolution. Development 2009, 136, 2433–2444. [Google Scholar] [CrossRef] [Green Version]
- Okano, Y.; Aono, N.; Hiwatashi, Y.; Murata, T.; Nishiyama, T.; Ishikawa, T.; Kubo, M.; Hasebe, M. A polycomb repressive complex 2 gene regulates apogamy and gives evolutionary insights into early land plant evolution. Proc. Natl. Acad. Sci. USA 2009, 106, 16321–16326. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luo, M.; Platten, D.; Chaudhury, A.; Peacock, W.J.; Dennis, E.S. Expression, imprinting, and evolution of rice homologs of the polycomb group genes. Mol. Plant 2009, 2, 711–723. [Google Scholar] [CrossRef]
- Nallamilli, B.R.; Zhang, J.; Mujahid, H.; Malone, B.M.; Bridges, S.M.; Peng, Z. Polycomb group gene OsFIE2 regulates rice (Oryza sativa) seed development and grain filling via a mechanism distinct from Arabidopsis. PLoS Genet. 2013, 9, e1003322. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.; Lee, S.; Hang, R.; Kim, S.R.; Lee, Y.S.; Cao, X.; Amasino, R.; An, G. OsVIL2 functions with PRC2 to induce flowering by repressing OsLFL1 in rice. Plant J. Cell Mol. Biol. 2013, 73, 566–578. [Google Scholar] [CrossRef] [PubMed]
- Ni, J.; Ma, X.; Feng, Y.; Tian, Q.; Wang, Y.; Xu, N.; Tang, J.; Wang, G. Updating and interaction of polycomb repressive complex 2 components in maize (Zea mays). Planta 2019, 250, 573–588. [Google Scholar] [CrossRef] [PubMed]
- Kapazoglou, A.; Tondelli, A.; Papaefthimiou, D.; Ampatzidou, H.; Francia, E.; Stanca, M.A.; Bladenopoulos, K.; Tsaftaris, A.S. Epigenetic chromatin modifiers in barley: IV. The study of barley polycomb group (PcG) genes during seed development and in response to external ABA. BMC Plant Biol. 2010, 10, 73. [Google Scholar] [CrossRef] [Green Version]
- Baile, F.; Merini, W.; Hidalgo, I.; Calonje, M. EAR domain-containing transcription factors trigger PRC2-mediated chromatin marking in Arabidopsis. Plant Cell 2021. [Google Scholar] [CrossRef]
- Zeng, X.; Gao, Z.; Jiang, C.; Yang, Y.; Liu, R.; He, Y. HISTONE DEACETYLASE 9 Functions with Polycomb Silencing to Repress FLOWERING LOCUS C Expression. Plant Physiol. 2020, 182, 555–565. [Google Scholar] [CrossRef]
- Liu, X.; Yang, S.; Zhao, M.; Luo, M.; Yu, C.W.; Chen, C.Y.; Tai, R.; Wu, K. Transcriptional repression by histone deacetylases in plants. Mol. Plant 2014, 7, 764–772. [Google Scholar] [CrossRef] [Green Version]
- Shu, K.; Liu, X.D.; Xie, Q.; He, Z.H. Two Faces of One Seed: Hormonal Regulation of Dormancy and Germination. Mol. Plant 2016, 9, 34–45. [Google Scholar] [CrossRef] [Green Version]
- Conti, L. Hormonal control of the floral transition: Can one catch them all? Dev. Biol. 2017, 430, 288–301. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Ichihashi, Y.; Suzuki, T.; Shibata, A.; Shirasu, K.; Yamaguchi, N.; Ito, T. Abscisic acid-dependent histone demethylation during postgermination growth arrest in Arabidopsis. Plant Cell Environ. 2019, 42, 2198–2214. [Google Scholar] [CrossRef] [PubMed]
- Veluchamy, A.; Jégu, T.; Ariel, F.; Latrasse, D.; Mariappan, K.G.; Kim, S.K.; Crespi, M.; Hirt, H.; Bergounioux, C.; Raynaud, C.; et al. LHP1 Regulates H3K27me3 Spreading and Shapes the Three-Dimensional Conformation of the Arabidopsis Genome. PLoS ONE 2016, 11, e0158936. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Mammals | Drosophila | Arabidopsis | Characteristic Domain | Activities | |
|---|---|---|---|---|---|
| PRC1 | RING1A/ RING1B | dRing/ Sce | AtRING1A/ 1B | RING finger domain and Ring-finger and WD-40 Associated Ubiquitin-Like (RAWUL) domain | E3 ubiquitin ligase activity for H2A |
| PCGF1-6 | Psc | AtBMI1A/ 1B/1C | RING finger domain and RAWUL domain | Co-factors for E3 ubiquitin ligase and compact nucleosomes | |
| CBX2/4/6/7/8 | Pc | LHP1(TFL2)/ VRN1 | Chromodomain and chromo shadow domain | Recognizes and binds to H3K27me3 | |
| PHC1/ PHC2/ PHC3 | Ph | UNKNOWN | Sterile Alpha Motif (SAM) domain and Zinc Finger domain | Mediates monoubiquitination of histone H2A | |
| PRC2 | EZH1/2 | E(z) | CLF/SWN/ MEA | SET domain | H3K27 methyltransferase |
| SUZ12 | Su(z)12 | EMF2/VRN2 FIS2 | Zinc Finger | Assists with E(z) catalytic activity | |
| EED | Esc | FIE | WD-40 repeat domain | Assists with E(z) catalytic activity | |
| RBAP48/46 | P55/Nurf55 | MSI1-5 | WD-40 repeat domain | Histone binding |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hinsch, V.; Adkins, S.; Manuela, D.; Xu, M. Post-Embryonic Phase Transitions Mediated by Polycomb Repressive Complexes in Plants. Int. J. Mol. Sci. 2021, 22, 7533. https://doi.org/10.3390/ijms22147533
Hinsch V, Adkins S, Manuela D, Xu M. Post-Embryonic Phase Transitions Mediated by Polycomb Repressive Complexes in Plants. International Journal of Molecular Sciences. 2021; 22(14):7533. https://doi.org/10.3390/ijms22147533
Chicago/Turabian StyleHinsch, Valerie, Samuel Adkins, Darren Manuela, and Mingli Xu. 2021. "Post-Embryonic Phase Transitions Mediated by Polycomb Repressive Complexes in Plants" International Journal of Molecular Sciences 22, no. 14: 7533. https://doi.org/10.3390/ijms22147533
APA StyleHinsch, V., Adkins, S., Manuela, D., & Xu, M. (2021). Post-Embryonic Phase Transitions Mediated by Polycomb Repressive Complexes in Plants. International Journal of Molecular Sciences, 22(14), 7533. https://doi.org/10.3390/ijms22147533






