The Two-Step Clustering Approach for Metastable States Learning
Abstract
1. Introduction
2. Learning Metastable States from MD Data
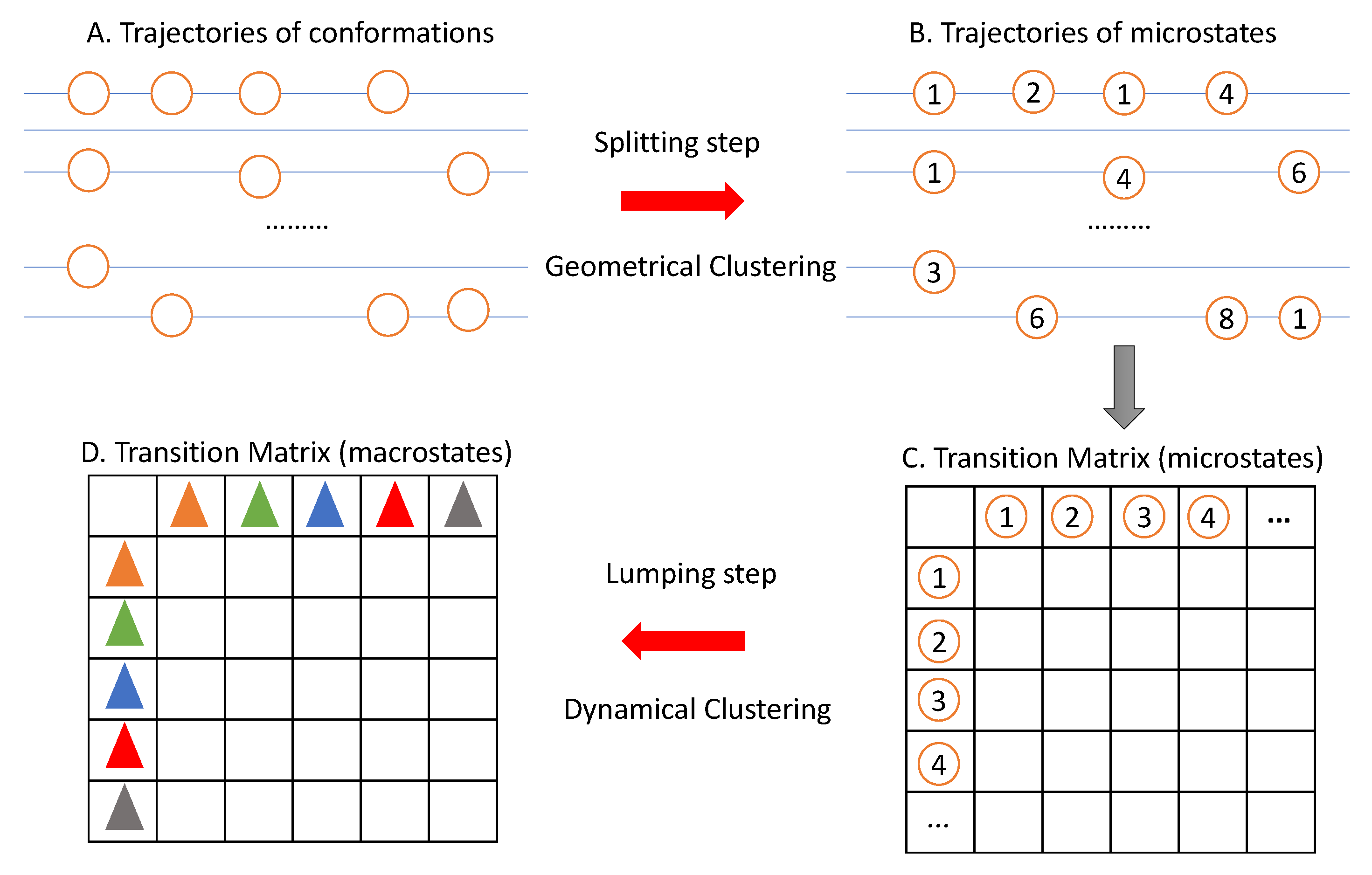
3. The Two-Step Clustering Framework
3.1. The Splitting Step: Geometrical Clustering
3.2. The Lumping Step: Dynamical Clustering
3.3. Refinements to The Framework
4. Some Extensions
5. Discussion and Outlook
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Finkelstein, A.V.; Ptitsyn, O. Protein Physics: A Course of Lectures; Academic Press: Cambridge, MA, USA, 2002. [Google Scholar]
- Schor, M.; Mey, A.S.; MacPhee, C.E. Analytical methods for structural ensembles and dynamics of intrinsically disordered proteins. Biophys. Rev. 2016, 8, 429–439. [Google Scholar] [CrossRef] [PubMed]
- Sponer, J.; Bussi, G.; Krepl, M.; Banas, P.; Bottaro, S.; Cunha, R.A.; Gil-Ley, A.; Pinamonti, G.; Poblete, S.; Jurecka, P.; et al. RNA structural dynamics as captured by molecular simulations: A comprehensive overview. Chem. Rev. 2018, 118, 4177–4338. [Google Scholar] [CrossRef] [PubMed]
- Selkoe, D.J. Folding proteins in fatal ways. Nature 2003, 426, 900. [Google Scholar] [CrossRef] [PubMed]
- Chapman, H.N.; Fromme, P.; Barty, A.; White, T.A.; Kirian, R.A.; Aquila, A.; Hunter, M.S.; Schulz, J.; DePonte, D.P.; Weierstall, U.; et al. Femtosecond X-ray protein nanocrystallography. Nature 2011, 470, 73. [Google Scholar] [CrossRef] [PubMed]
- Kabsch, W.; Rösch, P. Nuclear magnetic resonance: Protein structure determination. Nature 1986, 321, 469. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ha, T. Single-molecule fluorescence resonance energy transfer. Methods 2001, 25, 78–86. [Google Scholar] [CrossRef]
- Carroni, M.; Saibil, H.R. Cryo electron microscopy to determine the structure of macromolecular complexes. Methods 2016, 95, 78–85. [Google Scholar] [CrossRef]
- Boomsma, W.; Mardia, K.V.; Taylor, C.C.; Ferkinghoff-Borg, J.; Krogh, A.; Hamelryck, T. A generative, probabilistic model of local protein structure. Proc. Natl. Acad. Sci. USA 2008, 105, 8932–8937. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.W.; Liu, J.S.; Kou, S. Exploring the conformational space for protein folding with sequential Monte Carlo. Ann. Appl. Stat. 2018, 12, 1628–1654. [Google Scholar] [CrossRef]
- Moult, J.; Fidelis, K.; Kryshtafovych, A.; Rost, B.; Hubbard, T.; Tramontano, A. Critical assessment of methods of protein structure prediction—Round VII. Proteins Struct. Funct. Bioinform. 2007, 69, 3–9. [Google Scholar] [CrossRef]
- Moult, J.; Fidelis, K.; Kryshtafovych, A.; Rost, B.; Tramontano, A. Critical assessment of methods of protein structure prediction—Round VIII. Proteins Struct. Funct. Bioinform. 2009, 77, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Kryshtafovych, A.; Schwede, T.; Topf, M.; Fidelis, K.; Moult, J. Critical assessment of methods of protein structure prediction (CASP)—Round XIII. Proteins Struct. Funct. Bioinform. 2019, 87, 1011–1020. [Google Scholar] [CrossRef]
- Lena, P.D.; Nagata, K.; Baldi, P.F. Deep spatio-temporal architectures and learning for protein structure prediction. In Advances in Neural Information Processing Systems; Curran Associates Inc.: Red Hook, NY, USA, 2012; pp. 512–520. [Google Scholar]
- Wang, S.; Sun, S.; Li, Z.; Zhang, R.; Xu, J. Accurate de novo prediction of protein contact map by ultra-deep learning model. PLoS Comput. Biol. 2017, 13, e1005324. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; Adhikari, B.; Cheng, J. DeepSF: Deep convolutional neural network for mapping protein sequences to folds. Bioinformatics 2017, 34, 1295–1303. [Google Scholar] [CrossRef]
- Mardt, A.; Pasquali, L.; Wu, H.; Noé, F. VAMPnets for deep learning of molecular kinetics. Nat. Commun. 2018, 9, 5. [Google Scholar] [CrossRef] [PubMed]
- AlQuraishi, M. AlphaFold at CASP13. Bioinformatics 2019. [Google Scholar] [CrossRef] [PubMed]
- Dill, K.A.; MacCallum, J.L. The protein-folding problem, 50 years on. Science 2012, 338, 1042–1046. [Google Scholar] [CrossRef] [PubMed]
- Karplus, M.; McCammon, J.A. Molecular dynamics simulations of biomolecules. Nat. Struct. Mol. Biol. 2002, 9, 646. [Google Scholar] [CrossRef] [PubMed]
- Berg, B.A.; Neuhaus, T. Multicanonical algorithms for first order phase transitions. Phys. Lett. B 1991, 267, 249–253. [Google Scholar] [CrossRef]
- Sugita, Y.; Okamoto, Y. Replica-exchange molecular dynamics method for protein folding. Chem. Phys. Lett. 1999, 314, 141–151. [Google Scholar] [CrossRef]
- Mitsutake, A.; Sugita, Y.; Okamoto, Y. Generalized-ensemble algorithms for molecular simulations of biopolymers. Pept. Sci. Orig. Res. Biomol. 2001, 60, 96–123. [Google Scholar] [CrossRef]
- Bowman, G.R.; Huang, X.; Pande, V.S. Using generalized ensemble simulations and Markov state models to identify conformational states. Methods 2009, 49, 197–201. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Yao, Y.; Bowman, G.R.; Sun, J.; Guibas, L.J.; Carlsson, G.; Pande, V.S. Constructing multi-resolution Markov state models (MSMs) to elucidate RNA hairpin folding mechanisms. In Biocomputing 2010; World Scientific: Singapore, 2010; pp. 228–239. [Google Scholar]
- Lane, T.J.; Bowman, G.R.; Beauchamp, K.; Voelz, V.A.; Pande, V.S. Markov state model reveals folding and functional dynamics in ultra-long MD trajectories. J. Am. Chem. Soc. 2011, 133, 18413–18419. [Google Scholar] [CrossRef]
- McGibbon, R.T.; Pande, V.S. Learning kinetic distance metrics for Markov state models of protein conformational dynamics. J. Chem. Theory Comput. 2013, 9, 2900–2906. [Google Scholar] [CrossRef]
- Schwantes, C.R.; McGibbon, R.T.; Pande, V.S. Perspective: Markov models for long-timescale biomolecular dynamics. J. Chem. Phys. 2014, 141, 090901. [Google Scholar] [CrossRef] [PubMed]
- Nüske, F.; Wu, H.; Prinz, J.H.; Wehmeyer, C.; Clementi, C.; Noé, F. Markov state models from short non-equilibrium simulations—Analysis and correction of estimation bias. J. Chem. Phys. 2017, 146, 094104. [Google Scholar] [CrossRef]
- Husic, B.E.; Pande, V.S. Markov state models: From an art to a science. J. Am. Chem. Soc. 2018, 140, 2386–2396. [Google Scholar] [CrossRef]
- Chodera, J.D.; Noé, F. Markov state models of biomolecular conformational dynamics. Curr. Opin. Struct. Biol. 2014, 25, 135–144. [Google Scholar] [CrossRef]
- Wang, W.; Cao, S.; Zhu, L.; Huang, X. Constructing Markov State Models to elucidate the functional conformational changes of complex biomolecules. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2018, 8, e1343. [Google Scholar] [CrossRef]
- Lu, L.; Jiang, H.; Wong, W.H. Multivariate density estimation by Bayesian sequential partitioning. J. Am. Stat. Assoc. 2013, 108, 1402–1410. [Google Scholar] [CrossRef]
- Breiman, L.; Friedman, J.; Stone, C.J.; Olshen, R.A. Classification and Regression Trees; Routledge: London, UK, 1984. [Google Scholar]
- Vassilvitskii, S.; Arthur, D. k-means++: The advantages of careful seeding. In Proceedings of the Eighteenth Annual ACM-SIAM Symposium on Discrete Algorithms, New Orleans, LA, USA, 7–9 January 2007; pp. 1027–1035. [Google Scholar]
- Reynolds, A.P.; Richards, G.; Rayward-Smith, V.J. The application of k-medoids and pam to the clustering of rules. In Proceedings of the International Conference on Intelligent Data Engineering and Automated Learning, Exeter, UK, 25–27 August 2004; Springer: Berlin/Heidelberg, Germany, 2004; pp. 173–178. [Google Scholar]
- Mu, Y.; Nguyen, P.H.; Stock, G. Energy landscape of a small peptide revealed by dihedral angle principal component analysis. Proteins Struct. Funct. Bioinform. 2005, 58, 45–52. [Google Scholar] [CrossRef]
- Altis, A.; Nguyen, P.H.; Hegger, R.; Stock, G. Dihedral angle principal component analysis of molecular dynamics simulations. J. Chem. Phys. 2007, 126, 244111. [Google Scholar] [CrossRef]
- Sittel, F.; Jain, A.; Stock, G. Principal component analysis of molecular dynamics: On the use of Cartesian vs. internal coordinates. J. Chem. Phys. 2014, 141, 07B605_1. [Google Scholar] [CrossRef]
- Jain, A.K. Data clustering: 50 years beyond K-means. Pattern Recognit. Lett. 2010, 31, 651–666. [Google Scholar] [CrossRef]
- Chodera, J.D.; Swope, W.C.; Pitera, J.W.; Dill, K.A. Long-time protein folding dynamics from short-time molecular dynamics simulations. Multiscale Model. Simul. 2006, 5, 1214–1226. [Google Scholar] [CrossRef]
- Deuflhard, P.; Huisinga, W.; Fischer, A.; Schütte, C. Identification of almost invant aggregates in reversible nearly uncoupled Markov chains. Linear Algebra Its Appl. 2000, 315, 39–59. [Google Scholar] [CrossRef]
- Deuflhard, P.; Weber, M. Robust Perron cluster analysis in conformation dynamics. Linear Algebra Its Appl. 2005, 398, 161–184. [Google Scholar] [CrossRef]
- Beauchamp, K.A.; McGibbon, R.; Lin, Y.S.; Pande, V.S. Simple few-state models reveal hidden complexity in protein folding. Proc. Natl. Acad. Sci. USA 2012, 109, 17807–17813. [Google Scholar] [CrossRef]
- Wang, W.; Liang, T.; Sheong, F.K.; Fan, X.; Huang, X. An efficient Bayesian kinetic lumping algorithm to identify metastable conformational states via Gibbs sampling. J. Chem. Phys. 2018, 149, 072337. [Google Scholar] [CrossRef]
- Jain, A.; Stock, G. Identifying metastable states of folding proteins. J. Chem. Theory Comput. 2012, 8, 3810–3819. [Google Scholar] [CrossRef] [PubMed]
- Husic, B.E.; McKiernan, K.A.; Wayment-Steele, H.K.; Sultan, M.M.; Pande, V.S. A minimum variance clustering approach produces robust and interpretable coarse-grained models. J. Chem. Theory Comput. 2018, 14, 1071–1082. [Google Scholar] [CrossRef]
- Chodera, J.D.; Singhal, N.; Pande, V.S.; Dill, K.A.; Swope, W.C. Automatic discovery of metastable states for the construction of Markov models of macromolecular conformational dynamics. J. Chem. Phys. 2007, 126, 155101. [Google Scholar] [CrossRef]
- Sheong, F.K.; Silva, D.A.; Meng, L.; Zhao, Y.; Huang, X. Automatic state partitioning for multibody systems (APM): An efficient algorithm for constructing Markov state models to elucidate conformational dynamics of multibody systems. J. Chem. Theory Comput. 2014, 11, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Sittel, F.; Stock, G. Robust density-based clustering to identify metastable conformational states of proteins. J. Chem. Theory Comput. 2016, 12, 2426–2435. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Zhu, L.; Sheong, F.K.; Wang, W.; Huang, X. Adaptive partitioning by local density-peaks: An efficient density-based clustering algorithm for analyzing molecular dynamics trajectories. J. Comput. Chem. 2017, 38, 152–160. [Google Scholar] [CrossRef]
- Ester, M.; Kriegel, H.P.; Sander, J.; Xu, X. A Density-Based Algorithm for Discovering Clusters in Large Spatial Databases with Noise; KDD: Portland, OR, USA, 1996; Volume 96, pp. 226–231. [Google Scholar]
- Rodriguez, A.; Laio, A. Clustering by fast search and find of density peaks. Science 2014, 344, 1492–1496. [Google Scholar] [CrossRef] [PubMed]
- Sittel, F.; Stock, G. Perspective: Identification of collective variables and metastable states of protein dynamics. J. Chem. Phys. 2018, 149, 150901. [Google Scholar] [CrossRef]
- Bowman, G.R. Improved coarse-graining of Markov state models via explicit consideration of statistical uncertainty. J. Chem. Phys. 2012, 137, 134111. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Cui, R.Z.; Bowman, G.R.; Silva, D.A.; Sun, J.; Huang, X. Hierarchical Nyström methods for constructing Markov state models for conformational dynamics. J. Chem. Phys. 2013, 138, 174106. [Google Scholar] [CrossRef] [PubMed]
- Bowman, G.R.; Meng, L.; Huang, X. Quantitative comparison of alternative methods for coarse-graining biological networks. J. Chem. Phys. 2013, 139, 121905. [Google Scholar] [CrossRef]
- Krivov, S.V. Protein Folding Free Energy Landscape along the Committor-the Optimal Folding Coordinate. J. Chem. Theory Comput. 2018, 14, 3418–3427. [Google Scholar] [CrossRef]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Mardt, A.; Pasquali, L.; Noe, F. Deep generative Markov state models. In Advances in Neural Information Processing Systems; Curran Associates Inc.: Red Hook, NY, USA, 2018; pp. 3975–3984. [Google Scholar]
- Noé, F. Machine Learning for Molecular Dynamics on Long Timescales. arXiv 2018, arXiv:1812.07669. [Google Scholar]
- Noé, F.; Wu, H.; Prinz, J.H.; Plattner, N. Projected and hidden Markov models for calculating kinetics and metastable states of complex molecules. J. Chem. Phys. 2013, 139, 11B609_1. [Google Scholar] [CrossRef] [PubMed]
- Olsson, S.; Noé, F. Dynamic graphical models of molecular kinetics. Proc. Natl. Acad. Sci. USA 2019, 116, 15001–15006. [Google Scholar] [CrossRef] [PubMed]

| S1 | S2 | S3 | S4 | S5 | S6 | |
|---|---|---|---|---|---|---|
| S1 | 0.9457 | 0.0477 | 0.0062 | 0.0004 | 0.0000 | 0.0000 |
| S2 | 0.0609 | 0.9365 | 0.0004 | 0.0021 | 0.0000 | 0.0002 |
| S3 | 0.0403 | 0.0021 | 0.8939 | 0.0636 | 0.0000 | 0.0000 |
| S4 | 0.0020 | 0.0090 | 0.0526 | 0.9356 | 0.0008 | 0.0000 |
| S5 | 0.0013 | 0.0013 | 0.0000 | 0.0098 | 0.9718 | 0.0158 |
| S6 | 0.0000 | 0.0401 | 0.0000 | 0.0000 | 0.0519 | 0.9080 |
| Sum of diagonals: 5.591479 | ||||||
| Mean of diagonals: 0.9319131 | ||||||
| Minimal of diagonals: 0.8939 | ||||||
| S1 | S2 | S3 | S4 | S5 | S6 | |
|---|---|---|---|---|---|---|
| S1 | 0.9352 | 0.0003 | 0.0018 | 0.0000 | 0.0000 | 0.0626 |
| S2 | 0.0477 | 0.9131 | 0.0000 | 0.0068 | 0.0324 | 0.0000 |
| S3 | 0.0042 | 0.0000 | 0.9752 | 0.0000 | 0.0004 | 0.0202 |
| S4 | 0.0000 | 0.0032 | 0.0000 | 0.9104 | 0.0816 | 0.0048 |
| S5 | 0.0000 | 0.0269 | 0.0175 | 0.0672 | 0.8884 | 0.0000 |
| S6 | 0.0508 | 0.0000 | 0.0068 | 0.0000 | 0.0000 | 0.9424 |
| Sum of diagonals: 5.564797 | ||||||
| Mean of diagonals: 0.9274662 | ||||||
| Minimal of diagonals: 0.8884 | ||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, H.; Fan, X. The Two-Step Clustering Approach for Metastable States Learning. Int. J. Mol. Sci. 2021, 22, 6576. https://doi.org/10.3390/ijms22126576
Jiang H, Fan X. The Two-Step Clustering Approach for Metastable States Learning. International Journal of Molecular Sciences. 2021; 22(12):6576. https://doi.org/10.3390/ijms22126576
Chicago/Turabian StyleJiang, Hangjin, and Xiaodan Fan. 2021. "The Two-Step Clustering Approach for Metastable States Learning" International Journal of Molecular Sciences 22, no. 12: 6576. https://doi.org/10.3390/ijms22126576
APA StyleJiang, H., & Fan, X. (2021). The Two-Step Clustering Approach for Metastable States Learning. International Journal of Molecular Sciences, 22(12), 6576. https://doi.org/10.3390/ijms22126576






