1. Introduction
The energy transfer induced by ionizing radiation in a water medium occurs rapidly (on a scale of femtoseconds (fs)) during the physical stage of water radiolysis and is followed by the formation of radiolytic species. These species are created in a very short time (from femtoseconds (fs) to picoseconds (ps)), mainly through electronic events during the physico-chemical stage [
1,
2,
3,
4]. These events—such as thermalization, solvation of sub-excitation electrons, electronic hole migration, and fast electronic recombination—can lead to chemical bond breaks and produce species. After this stage, the chemical stage starts with heterogeneous distributions of radiation-induced spurs along the axis of the tracks from a few picoseconds (ps) to greater than ~100 nanoseconds (ns) in water at room temperature [
5]. The spur evolution largely depends on the spatial distribution of these high concentration regions. Methods using Brownian dynamics and Smoluchowski theory have been applied to describe the spur evolution, in which detailed individual trajectories of spherical particles are simulated in chemical reaction-diffusion systems (called "systems" in the following text) [
5,
6,
7,
8,
9]. In this particle-based representation, only the molecules of interest are explicitly simulated, and the solvent (water) is considered a continuum. Under these conditions, the diffusion of the species occurs until two individual reactants encounter each other (chemical interactions happen when the distance between reactants is less than a pre-defined distance). Over time, the species and their products disperse and are distributed more homogeneously. Their reactions continuously occur until they reach steady-states, where the species concentrations do not change anymore (from 100 microseconds (µs) to several seconds (s)).
Although the detail level is limited when using simplifications (spherical particles, continuum representation), the calculation time remains the main drawback of the particle-based representation when the simulations deal with a large number of species. This makes it difficult to simulate large, absorbed dose radiation systems, where the concentration of produced species may significantly impact the irradiated medium composition. Moreover, the majority of radiation chemistry experiments extend up to the homogeneous regime where radiation-induced species evolve and reach steady-states over long-term scales [
5]. Because of its computational cost, the particle-based representation cannot simulate these types of conditions. Instead, compartment-based models can be used because the level of detail is more limited [
10].
The compartment-based representation describes species evolution by gathering molecules of the same species as one group placed in a confined volume (or “compartment”). Only molecules in the compartment can react with each other. A basic requirement of compartment-based models is that the distribution of the species is a homogeneous mixture or “well-mixed” in their compartment. Therefore, no information about species positions is described during the evolution. Under this condition, the chemical master equation (CME) [
11] is used to describe the time–evolution probability of all the different possible states of the system. In general, the CME is mathematically troublesome. To address this, Gillespie developed a stochastic simulation algorithm (SSA) that equivalently reproduces the CME using a derived Monte Carlo technique [
11].
In practice, this “well-mixed” condition is usually not fulfilled since most systems of interest do not begin in homogeneous states. Many of them are first heterogeneously distributed and then tend toward homogeneity as the systems evolve over time, before reaching the fully well-mixed state. For example, in water radiolysis, the transition process between the heterogeneous and homogeneous states may occur continuously from ns to µs or longer, depending on the volume and initial species distribution. Therefore, to be able to apply this well-mixed condition to heterogeneous systems, the compartment-based model proposes dividing the simulation volume into smaller regions (“sub-volume” or “voxels”). In these voxels, the species can be considered as well-mixed (or locally homogeneous). The species can react with each other in the same voxel, and diffusion is modeled by jumps between adjacent voxels. An extension of the CME is the reaction-diffusion master equation (RDME) [
12], which is used to account for spatial distributions of species into the states of the system.
In this work, we implement, in Geant4-DNA, the compartment-based (“on-lattice”) model for simulating water radiolysis. In the result and discussion sections, we present several comparisons between the compartment-based model and the SBS method of Geant4-DNA in order to validate our compartment-based model implementation. In the materials and methods section, we detail our implementation of the compartment-based model using the reaction-diffusion master equation (RDME). We also present the next sub-volume method (NSM) [
13], the choice of voxels size, and the merging technique using “the hierarchical algorithm for the RDME” (hRDME) [
14] approach in our implementation.
3. Discussion
A challenging aspect of our method is the choice of the time
t1 when the microscopic sub-stage ends, and the mesoscopic sub-stage starts (see
Section 4). The distribution of energy deposits by ionizing particles produces regions of microscopic spurs along the tracks. Since spur sizes are comparable with their reaction radius within a few ns after exposure up to the time
t1, the microscopic sub-stage cannot be described by using the “well-mixed” model. After this period, local homogeneity eventually appears, and the compartment-based model can be applied. A small
t1 could lead to a small initial voxel size, which may cause unphysical results, while a long
t1 could make the simulation less efficient, especially for systems with a large number of species. Moreover,
t 1should also depend on the initial species distribution in which the linear energy transfer (LET) of the irradiation source is involved. However, the method to determine
t1 for different LETs is beyond the scope of this work.
In the compartment-based model, the splitting of the simulation volume into smaller volumes helps to improve the computation accuracy. Indeed, in a simple system of unimolecular reactions, Isaacson [
16] showed that a deterministic partial differential equation model can be exactly reproduced from the RDME when the voxel size
h tends to 0. However, the voxel size should not be too small. The first reason is that the voxel size must be much larger than the reaction radii to maintain the physical validity of the RDME. Since reactive events involve reactants in the same voxel, a voxel size
h comparable to the reaction radius may lead to missing reactions. This value should not be less than a critical size of about
, where
is the reaction radius [
17]. The second reason concerns the computational efficiency of the model. When the voxel size is too small, most events are diffusions with a small step length (see Equation (4)). These repetitive diffusion events slow down the simulations. Therefore, the key question is how to adapt the appropriate voxel size
h to the evolution of the system. The answer depends on each particular case. In water radiolysis, the evolution of the radiolytic species is complex. The simulation configuration such as the beam shape, the irradiated medium, temperature, etc., may significantly change the output of the system. However, for a simplified system, for example, a system in which radiation is instantaneously absorbed and the evolution of produced species starts simultaneously, we observed that the voxel size
h could be smaller at the beginning when the species were close to each other and then became increasingly coarser over time when the species distributed more homogeneously by diffusion. The coarser meshes helped in reaching longer time steps and, therefore, reduced the computational time.
To change the mesh size during the simulations, we needed to know the spatial distribution of the species over time. The first step of the NSM algorithm (see
Section 4.2.1) implies that the compartment-based representation should start from the particle-based representation by filling particle position information at the end of the microscopic sub-stage. After that, no more information on species positions is kept, and we cannot use particle position to change the voxel size. A good strategy would be to track particle position with compartment-based representation in a hybrid method [
18]. However, this may increase the computational cost when dealing with a large number of species. The solution used in this work was to merge eight adjacent octant voxels into one and, thus, obtain larger voxels (“hierarchical algorithm for the RDME—hRDME”) [
14]. Therefore, all species concentrations were initially distributed in a fine mesh and then transferred to coarser meshes over time until we reached the coarsest mesh of one single voxel. Thus, the transfer time
should be large enough for the random migration by diffusion of species between the constitutive voxels to reach a “well-mixed” state in their coarser mesh.
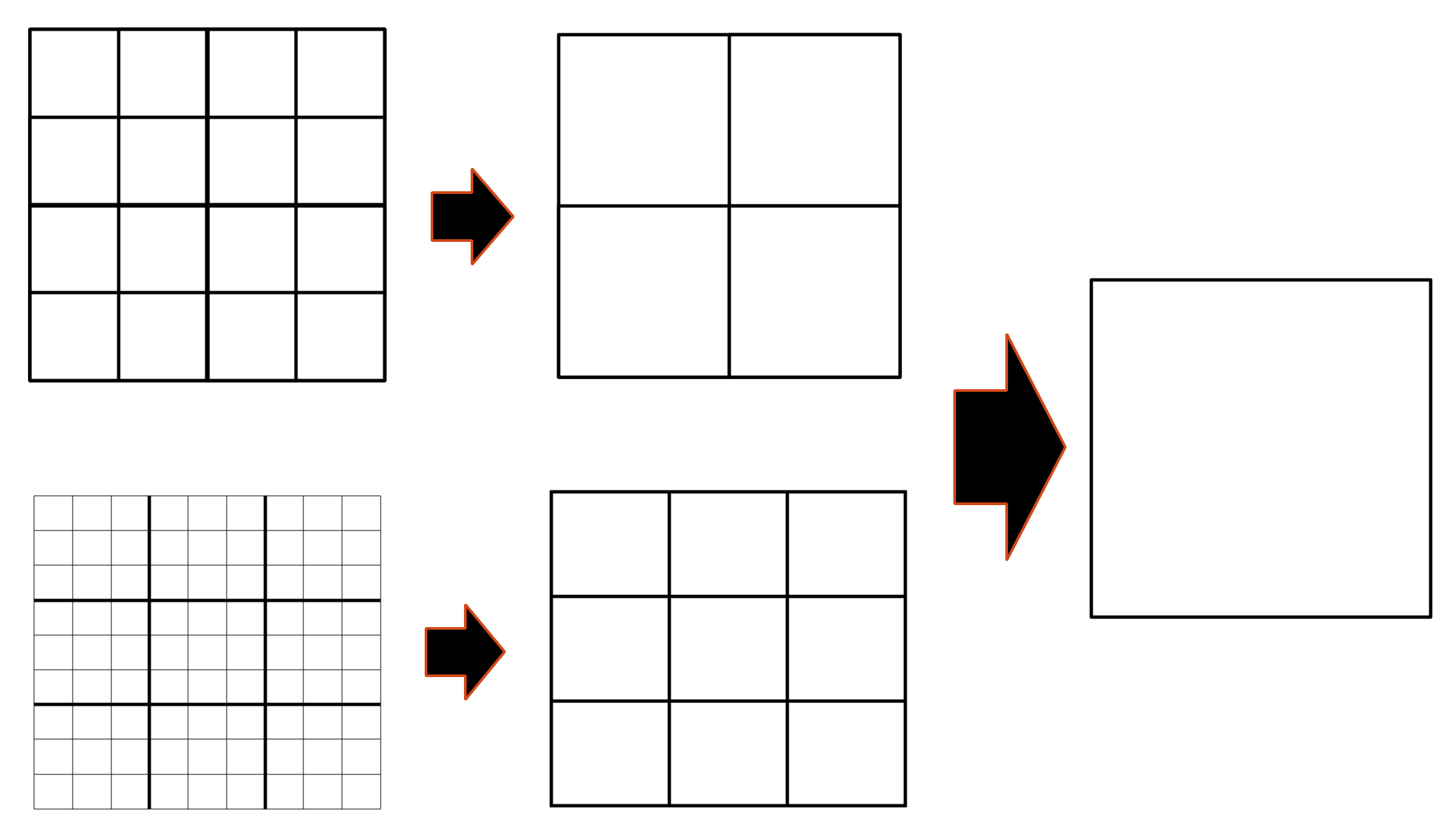
An example of the eight-octant merging operation used in the hRDME method is shown in 2D at the top of
Figure 4, where the mesh of 4
3 voxels moves to 2
3 voxels and then to one single voxel. This algorithm can also be extended to the twenty-seven-voxels merging operation, where 27 voxels were merged into one larger voxel. For example, an initial mesh of 9
3 voxels was moved to 3
3 and then one voxel (see bottom of
Figure 4). In this case, the changes of size between the finer and coarser meshes were larger than with the eight-octant merging operation, while the well-mixed requirement was still maintained since the transfer-time
for the random migration is longer in Equation (5). These larger changes of size not only speed up the simulation but also relax the choice of initial meshes, which are limited by the physical validity and the hierarchical requirement. This choice is particularly important for systems having a large geometrical volume because it helps to split the volume into a large number of voxels to start with a sufficiently fine mesh. For illustrative purposes, we considered the simple reaction-diffusion system tested in
Section 2.2, for which we extended the simulation volume to 1 × 1 × 1 µm
3 dimension. In order to keep a good start resolution close to the mean inter-particle distance, the initial mesh in the eight-octant merging operation was constrained by a fixed number of voxels equal to 512
3 or 1024
3. While an initial mesh of 1024
3 voxels involved a resolution of 0.976 nm, which is shorter than the critical voxel size (
= 1.13 nm), an initial mesh of 512
3 voxels gave a resolution of 1.9531 nm, which is larger than the mean inter-particle distance (1.63 nm) (see
Section 2.2). In this case, the twenty-seven-voxels merging operation is a better choice as an initial mesh of 729
3 voxels and provides a resolution of 1.37 nm.
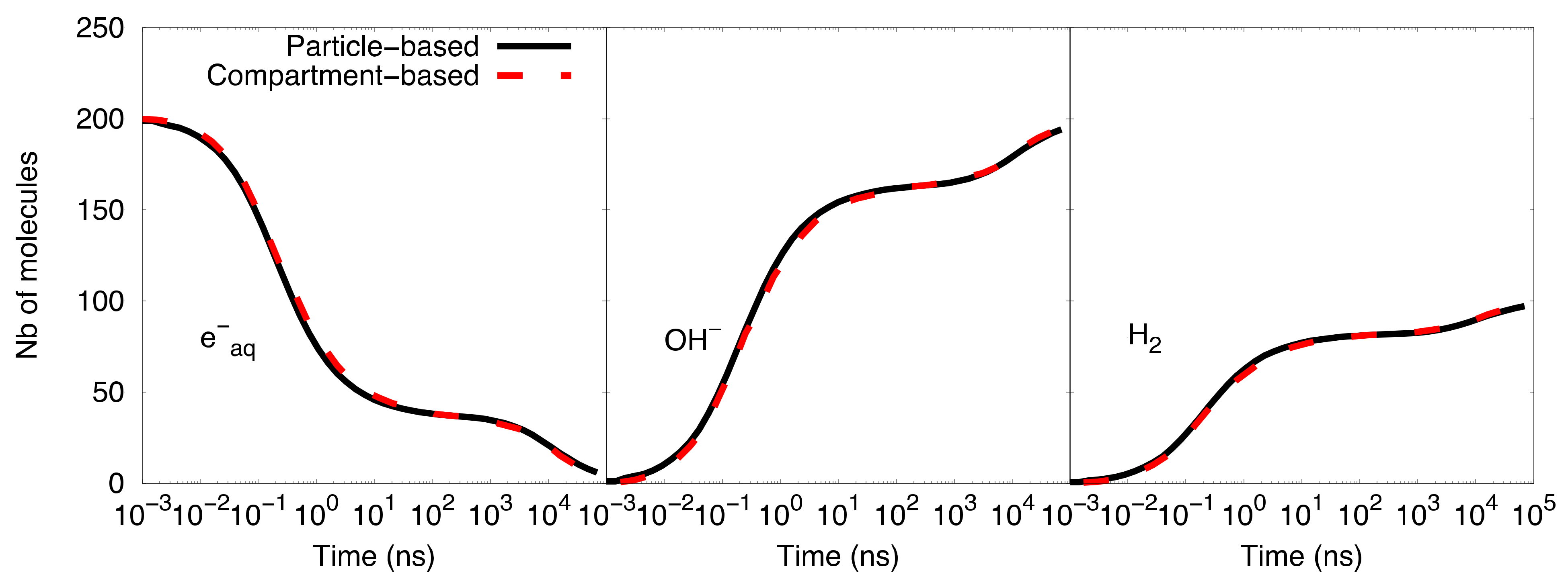
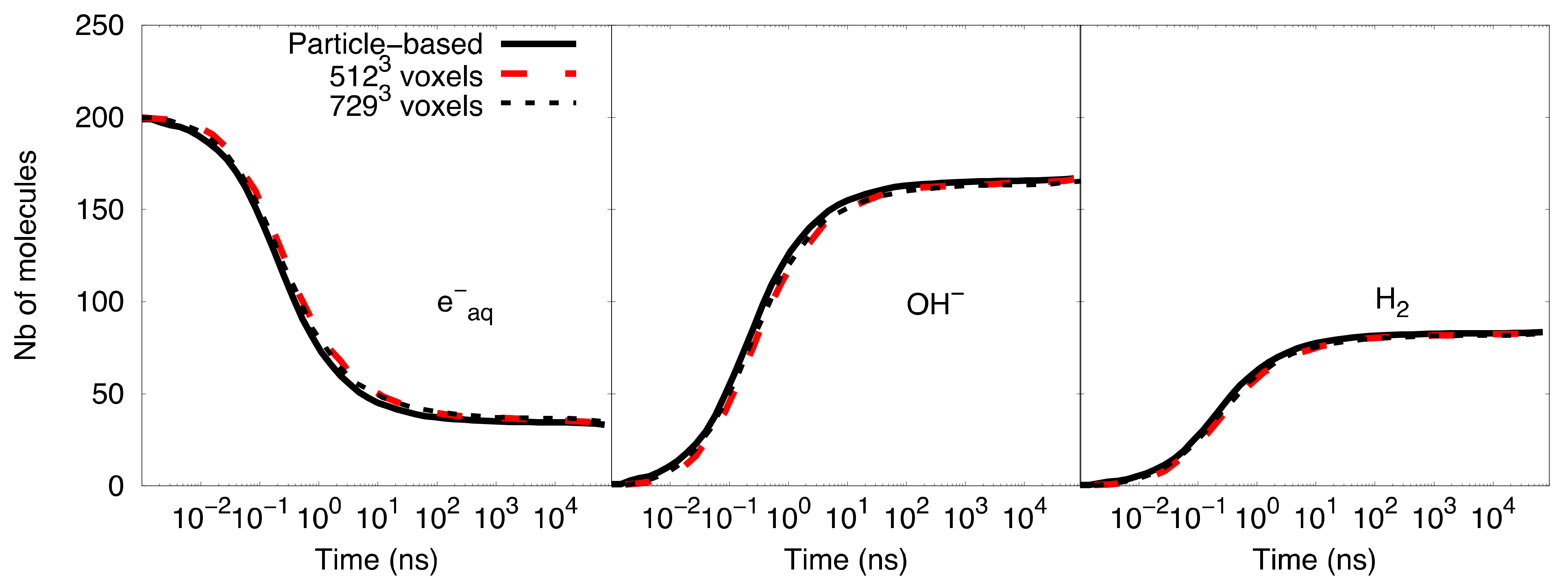
Figure 5 shows the comparison of the time-dependent yield of species (number of molecules) as computed with the particle-based SBS model and the compartment-based model starting with 729
3 voxels and 512
3 voxels. A better agreement of the species evolution was found between the compartment-based model, starting with an initial mesh of 729
3 voxels, and the particle-based SBS simulation, while the initial mesh of 512
3 voxels shows a slight deviation. This comparison illustrates the fact that the choice of initial meshes is not straightforward in the compartment-based model, as it depends on the type of simulated system.
Finally, another constraint of the compartment-based model presented in this work is the cubic volume of the simulation. This volume shape makes it possible to create uniform Cartesian meshes, which are particularly simple and efficient to handle in compartment-based simulations. A technical challenge of the compartment-based model would be to extend it to treat all forms of volumes with, for example, curved boundaries.
4. Materials and Methods
The main characteristic of our model is the combination of the SBS Brownian dynamics model already available in Geant4-DNA with the compartment-based model using the RDME (so-called “SBS-RDME model”) in water radiolysis simulations.
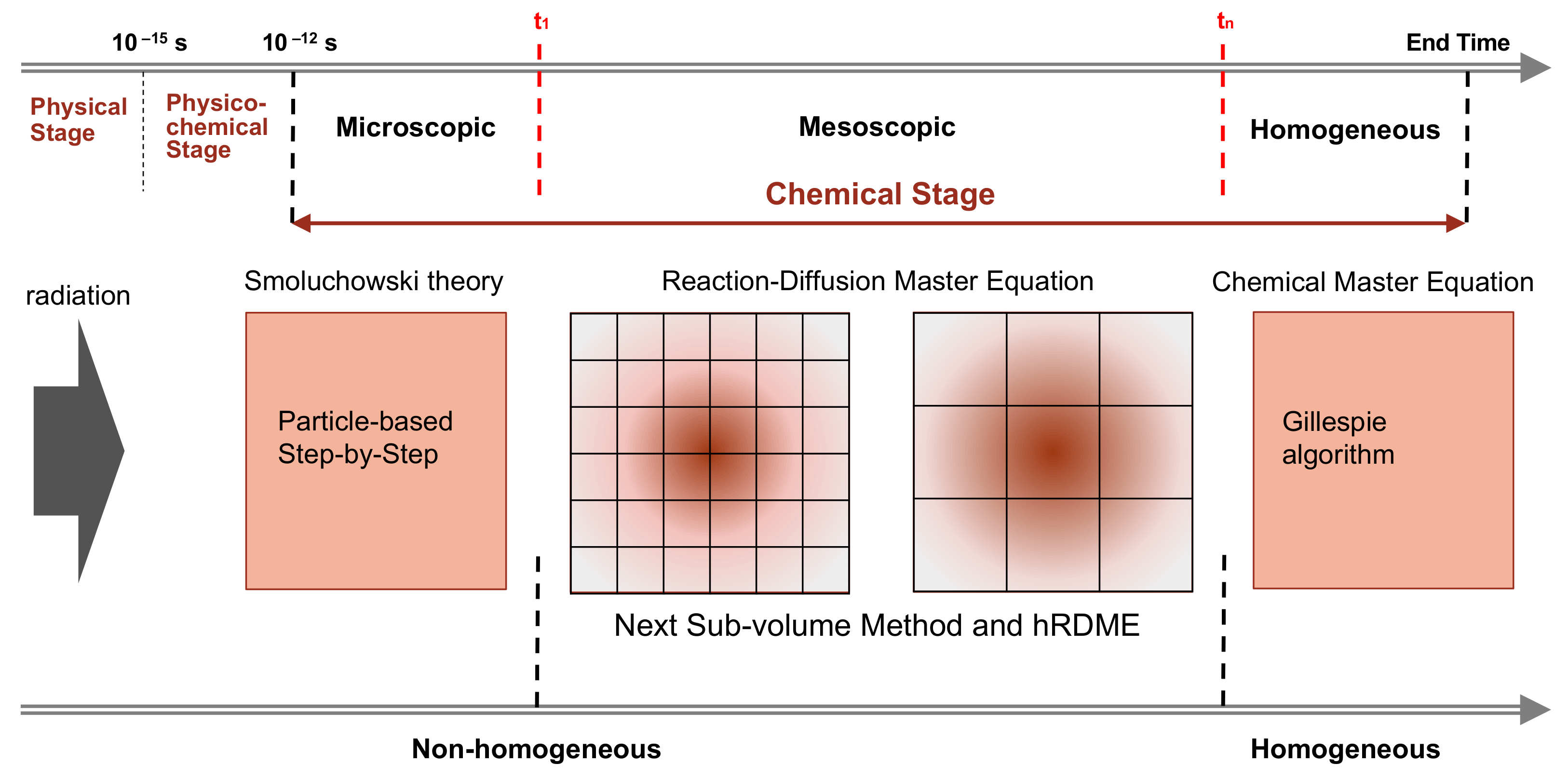
Figure 6 illustrates the simulation scheme of this combination.
We began by using Geant4-DNA [
19,
20,
21,
22,
23,
24,
25] to model the physical and physico-chemical stage up to 1 ps. Then, the chemical stage was divided into three sub-stages: the microscopic, mesoscopic, and homogeneous sub-stages. The microscopic sub-stage started from the formation of species at the beginning of the chemical stage. As the spatial distribution of the early species (e
−aq, H
•,
•OH, H
3O
+,…) was concentrated in very small volumes around the tracks, their evolutions largely depended on the species position. Therefore, at this sub-stage, we used the particle-based SBS method of Geant4-DNA to simulate the detailed trajectories of the individual species. Once the simulation reached a time
t1, we stopped the particle-based SBS simulation and initiated a uniform 3D Cartesian mesh for the compartment-based model. This was the beginning of the mesoscopic sub-stage. The initial mesh resolution should be small and selected according to the spatial distribution of species at the end of the microscopic sub-stage. We then used the hRDME approach (see details in
Section 4.2.2) to adapt the size of the voxels during the evolution of the system. The system used increasingly coarser meshes over time, to finally merge into one single voxel that covered the full simulation volume at time
tn, when the homogeneous sub-stage started. During the homogeneous sub-stage, we used the CME stochastic process to sample only reactive events.
4.1. The Microscopic Chemical Stage
During the microscopic sub-stage, the species diffused, and the reactions happened as soon as the distance between two species was shorter than their reaction radius [
6]. The diffusion and reaction processes were simulated using the particle-based step-by-step (SBS) method, which is available in Geant4-DNA [
6]. Briefly, this method proposes a time-step model that allows the choice of time steps, during which the reaction cannot occur with at least 95% (by default) confidence (named “dynamic time step”). Therefore, this model requires a minimum time step to avoid the repetition of many small time steps. For this work, we set the minimum time step at 1 ps. Details of the step-by-step (SBS) method can be found elsewhere [
6,
8].
4.2. The Mesoscopic Chemical Stage
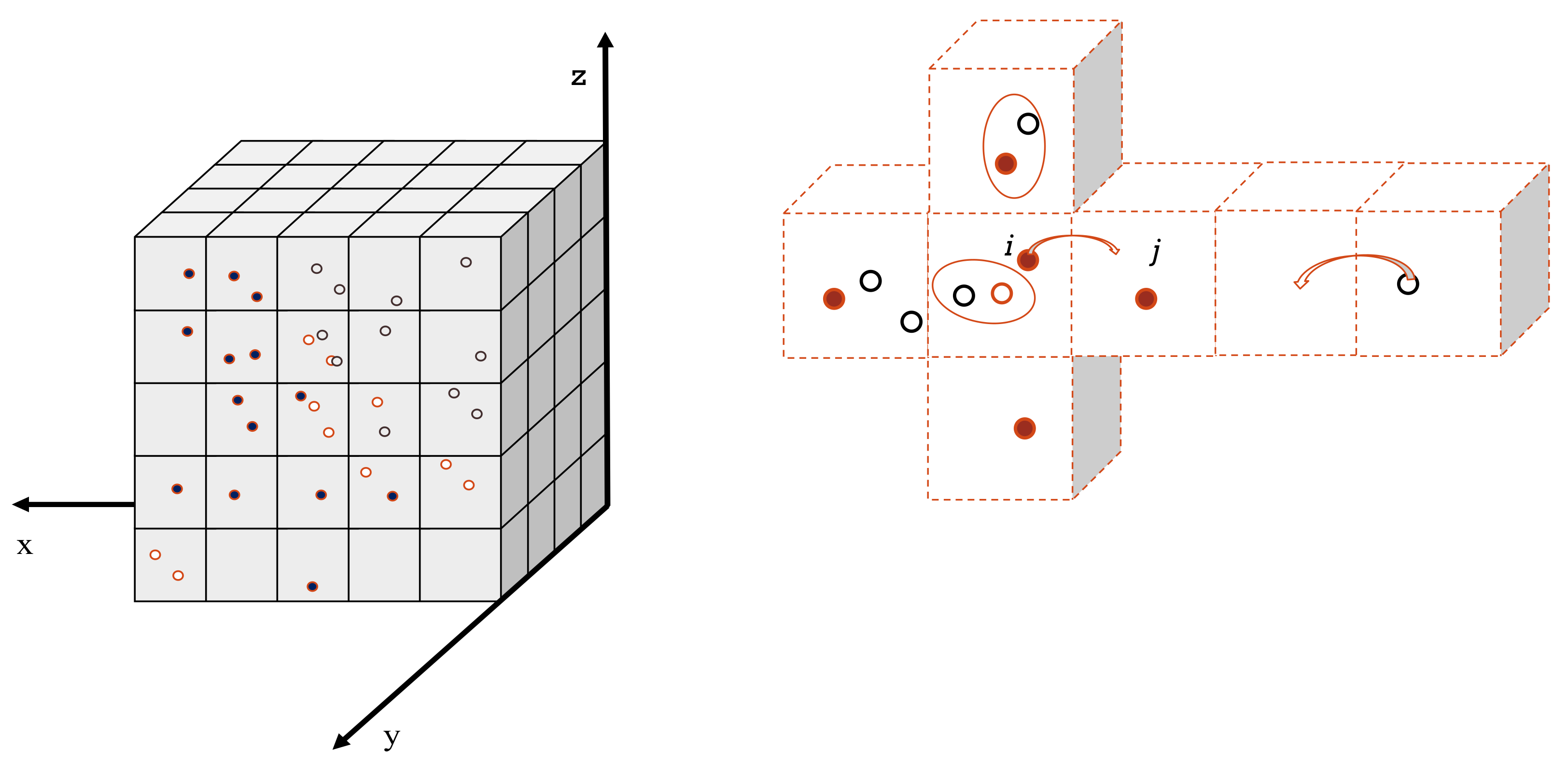
During the mesoscopic stage, we employed the compartment-based model, using the reaction-diffusion master equation (RDME) to simulate the heterogeneous chemical phase. The principal characteristic of this model is that it divides the simulation space into voxels, inside which the medium is considered homogeneous (
Figure 7—left). Reactions between molecules take place within the voxels, while species can move from one voxel to an adjacent one by diffusion (
Figure 7—right). We defined the propensity function (Equation (1)) of the RDME so that its product with a given infinitesimal time step provides the probability that the given event (reaction or diffusion) occurs in this time step. For a given voxel
i, it is given by the following equation.
The first term
is the propensity function of reaction
r among R possible bimolecular reactions (or “second-order” reactions) that is written as:
where
,
are the number of the reactants
and
in the voxel
i, respectively,
is the reaction rate, and
is the volume of the voxel
i.
The second term
in Equation (1) is the propensity function of the diffusion when a molecule of species
k among
L existing species is transferred from voxel
to voxel
among the
M possible adjacent voxels. It is written as:
where
is the number of the chemical species
in the voxel
i and
is the diffusion rate. The diffusion rate is given by:
where
D is the diffusion constant ([length]
2[s]
−1) and
h is the cubic voxel size ([length]).
The diffusion is also called the “diffusive transfer reaction” [
12]. Indeed, diffusion can be considered a sequence of a degradation reaction in voxel
and a production reaction in voxel
. Both reactions are first-order reactions that are handled by the Gillespie algorithm, as are second-order reactions between species (see
Section 4.2.1).
Figure 7.
(Left)/Voxelization of the considered volume of simulation into smaller sub-volumes. Species are represented by different types of circles. (Right)/The reactions and diffusion are triggered independently in a random process.
Figure 7.
(Left)/Voxelization of the considered volume of simulation into smaller sub-volumes. Species are represented by different types of circles. (Right)/The reactions and diffusion are triggered independently in a random process.
4.2.1. Sampling of Next Reaction/Diffusion Event
To establish the list of reactive and diffusive events over all voxels, we used the next sub-volume method (NSM), introduced by Elf and Ehrenberg [
13]. Since these events are scheduled independently in each voxel state, the NSM has an event-driven nature where, at every time step, an event occurs. This event-driven nature enables the NSM to efficiently organize events in a queue and process them one by one. The sampling of these events in each voxel is computed by using the direct method of the Gillespie algorithm [
11,
12]. In order to choose the voxel in which the next event occurs, the NSM applies the next reaction method [
26].
The numerical implementation of the NSM is summarized here [
13]:
- (1)
Computation of the species concentrations over voxels of the initial mesh at time t1.
- (2)
Calculation of the propensity functions for all voxels (see Equation (1)).
- (3)
Sampling of the time when the next event occurs for all the voxels,
where
is a uniform random number in [0,1].
- (4)
Sampling which types of reactions or diffusion will take place at a time according to their relative contribution (, ) in the propensity function .
- (5)
Sorting these events according to their occurrence times .
- (6)
Processing the first event in the queue and changing the concentrations in the voxels involved in the event.
If the event is a reaction, we eliminate reactants and create products.
If the event is a diffusion, we remove the particle in the voxel where it was located and add the particle in the voxel where it goes.
- (7)
Sampling of and the type of events in the updated voxels according to their new concentrations following points 3 and 4 and sorting the new list of events, as in point 5.
- (8)
The queue of events is processed successively following points 3 to 7 until no more events are present, or the times are longer than the end time of the simulation.
4.2.2. Adaptation of Voxel Sizes during the Evolution Time
To adapt the voxel size, we applied the hierarchical algorithm for the RDME (hRDME) [
14]. This method consists of merging the eight octant adjacent voxels (so-called “eight-octant merging operation”) into a single larger voxel. For example, an initial mesh of 32
3 voxels is gradually merged into a mesh of 16
3, 8
3, 4
3, 2
3, and finally into one voxel. All species of finer voxels are moved to a larger voxel of a coarser mesh after each “transfer time”
. In our implementation, we took the transfer time as proposed in the study of Hellander and Hellander [
14]. It is written:
where
is the voxel size corresponding to the transfer-time
, and
is the lowest diffusion constant among all species.
4.3. The Homogeneous Chemical Stage
The hRDME algorithm changes the voxel size until there is only one voxel covering the whole simulation volume. The species in the volume are considered to be well-mixed, and the Gillespie algorithm using CME is applied to sample only reactive events. The homogenous sub-stage ends when the system enters the steady-state.
4.4. Bounded Volume
In our simulation, we considered that molecules diffuse and react in a bounded volume (that is, limited by geometrical boundaries), which is also the irradiated water box volume of the physical stage. For the first time, we implemented the reflective boundary conditions [
27] in the SBS method of Geant4-DNA. These conditions enabled us to confine the chemical molecules in the considered volume by bouncing them off the walls of the volume. We also applied the bouncing off the volume border for the mesoscopic and homogeneous sub-stages.
4.5. G-Values
We calculated the yield of a chemical species induced by water radiolysis in terms of G-value. This represents the mean number of molecules of the species produced per 100 eV of energy imparted.