Genetic Predisposition to the Mortality in Septic Shock Patients: From GWAS to the Identification of a Regulatory Variant Modulating the Activity of a CISH Enhancer
Abstract
1. Introduction
2. Results
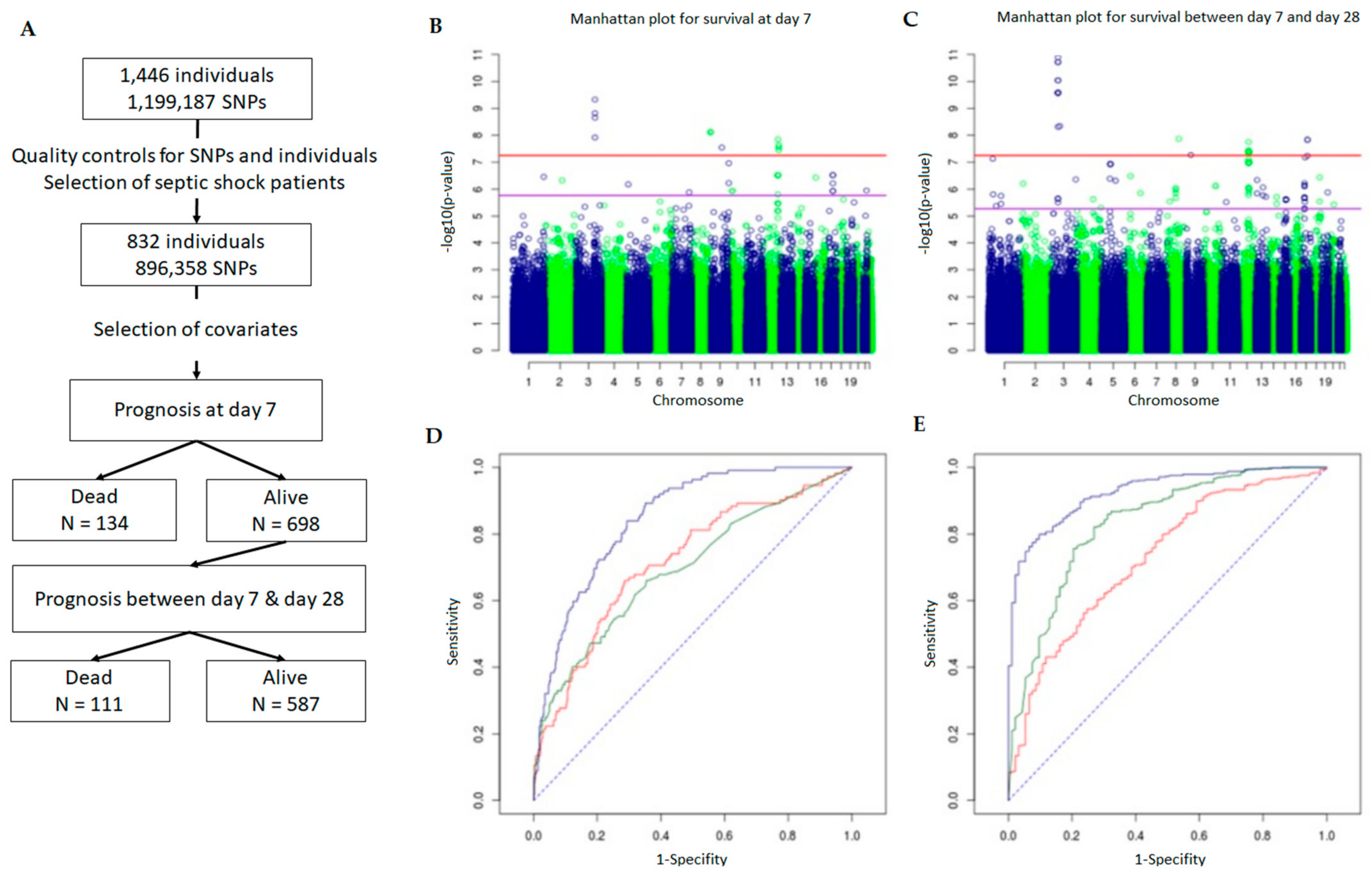
2.1. Patients and Covariates
2.2. Genome-Wide Association Analysis
2.3. Usefulness of GWAS to Predict Septic Shock Outcome
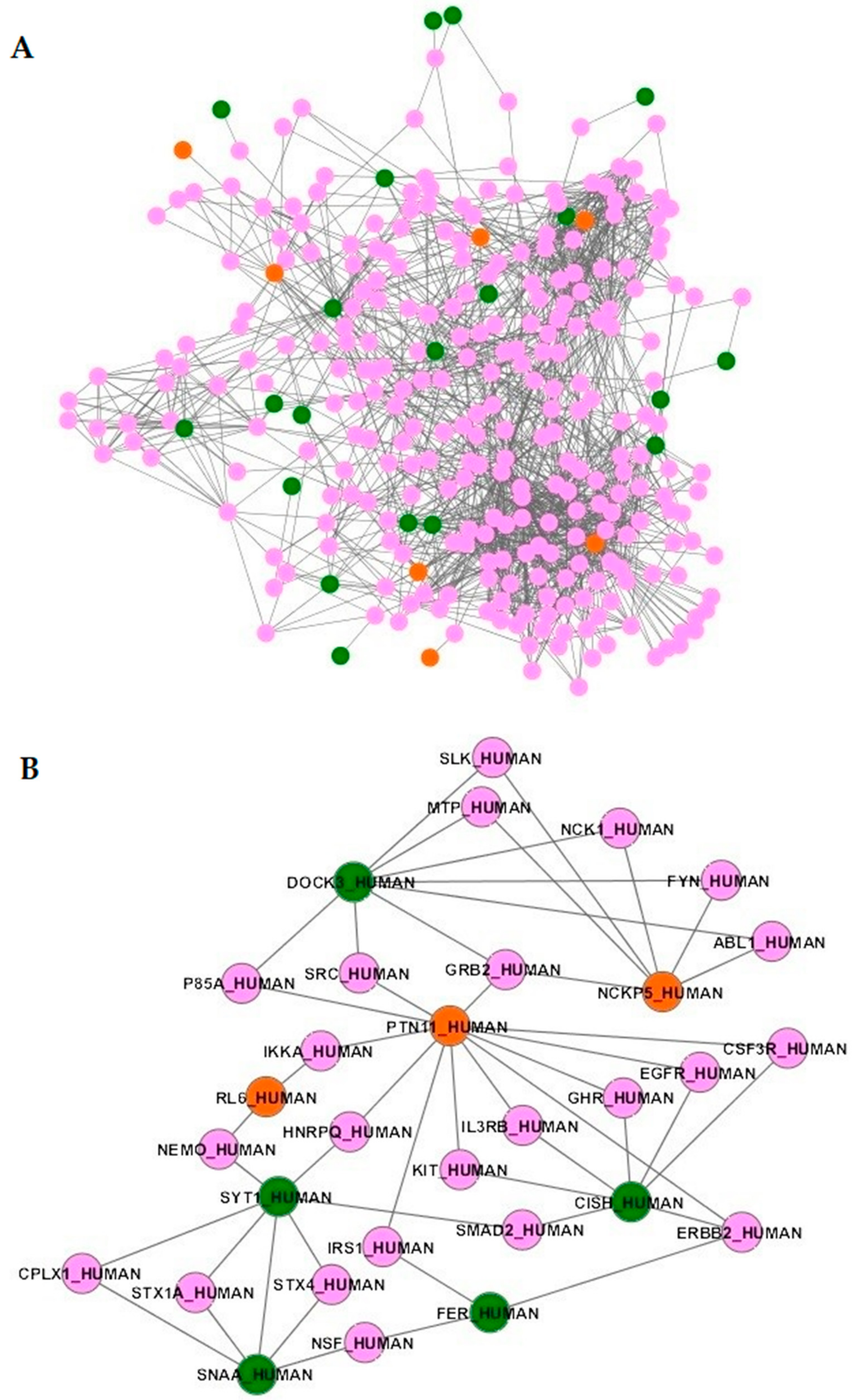
2.4. Protein–Protein Networks and Functional Enrichments
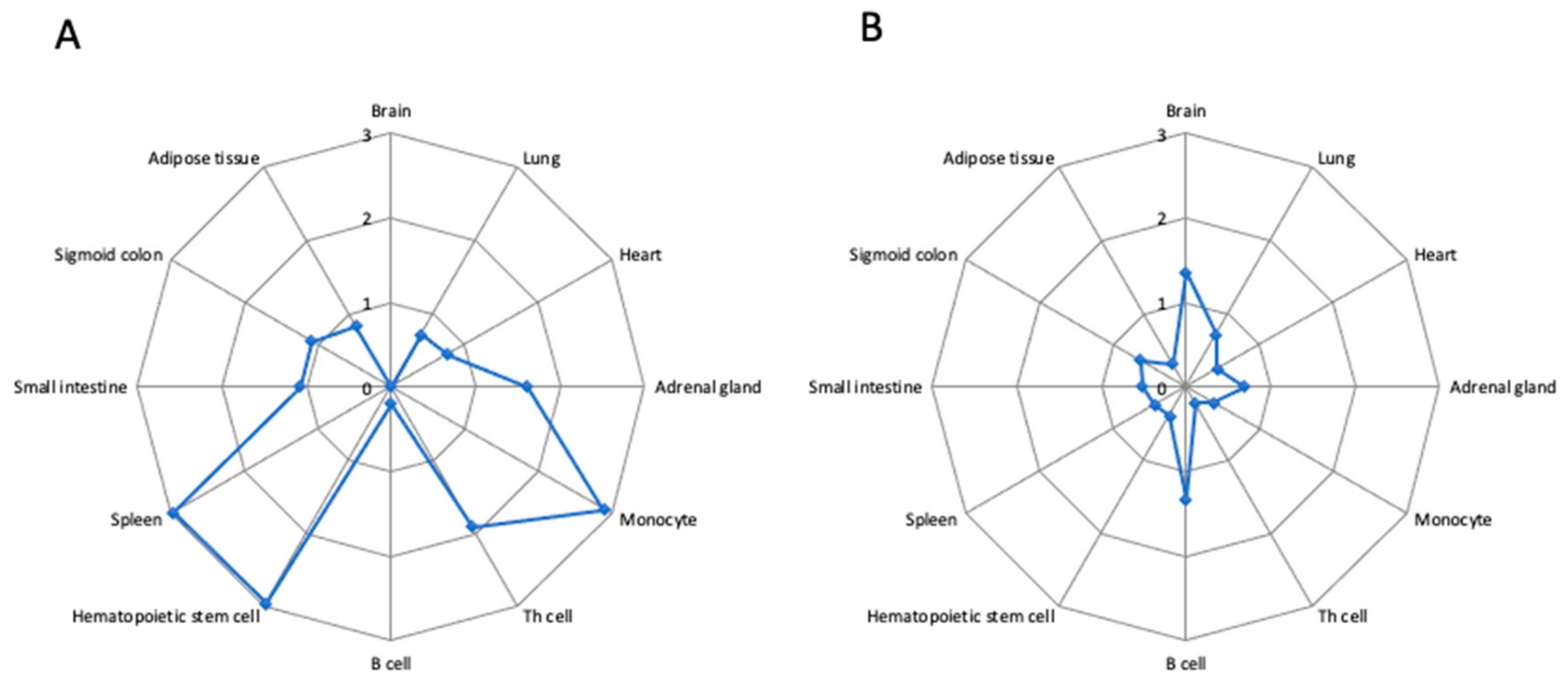
2.5. Sepsis-Associated SNPs in Super-Enhancers
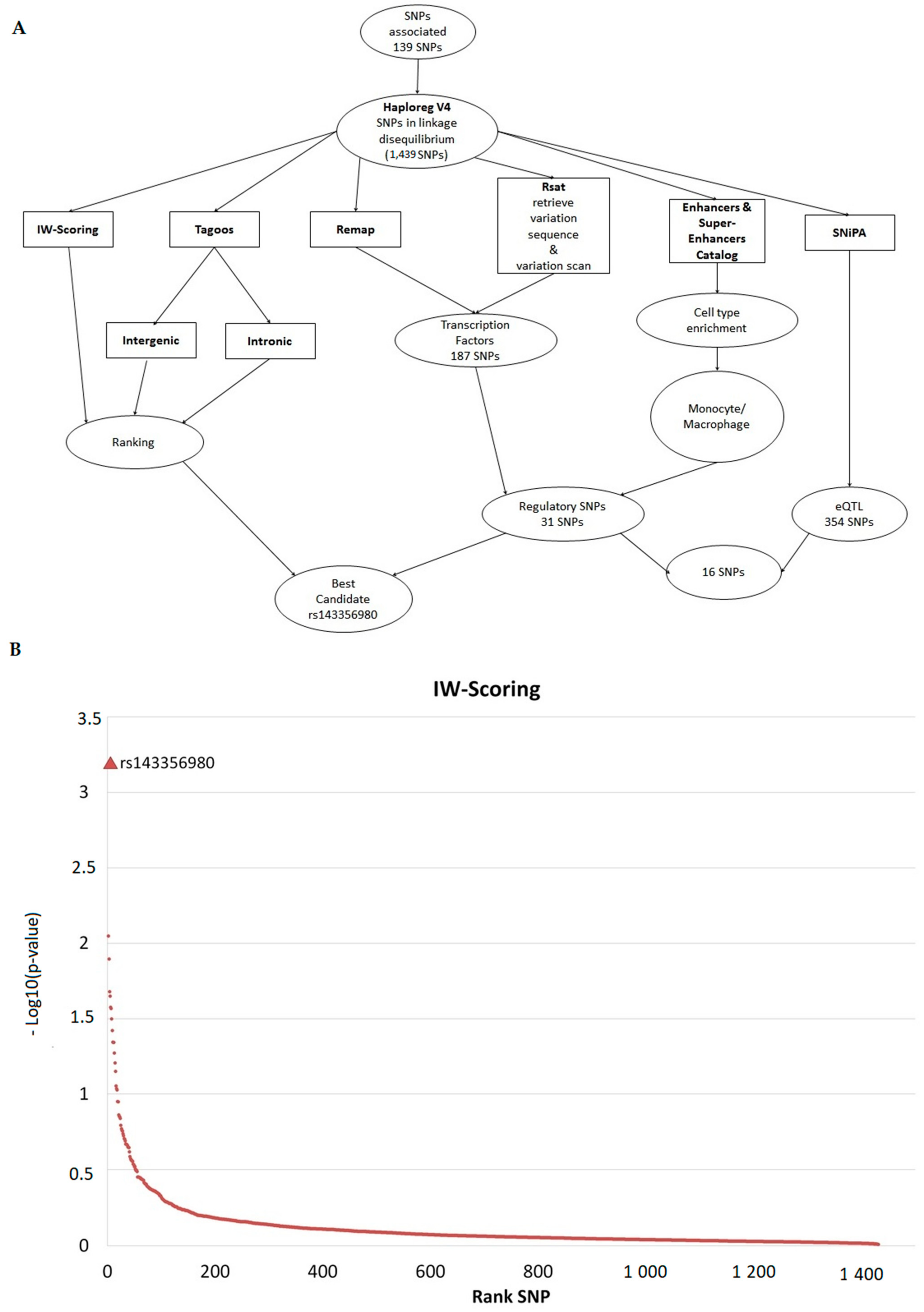
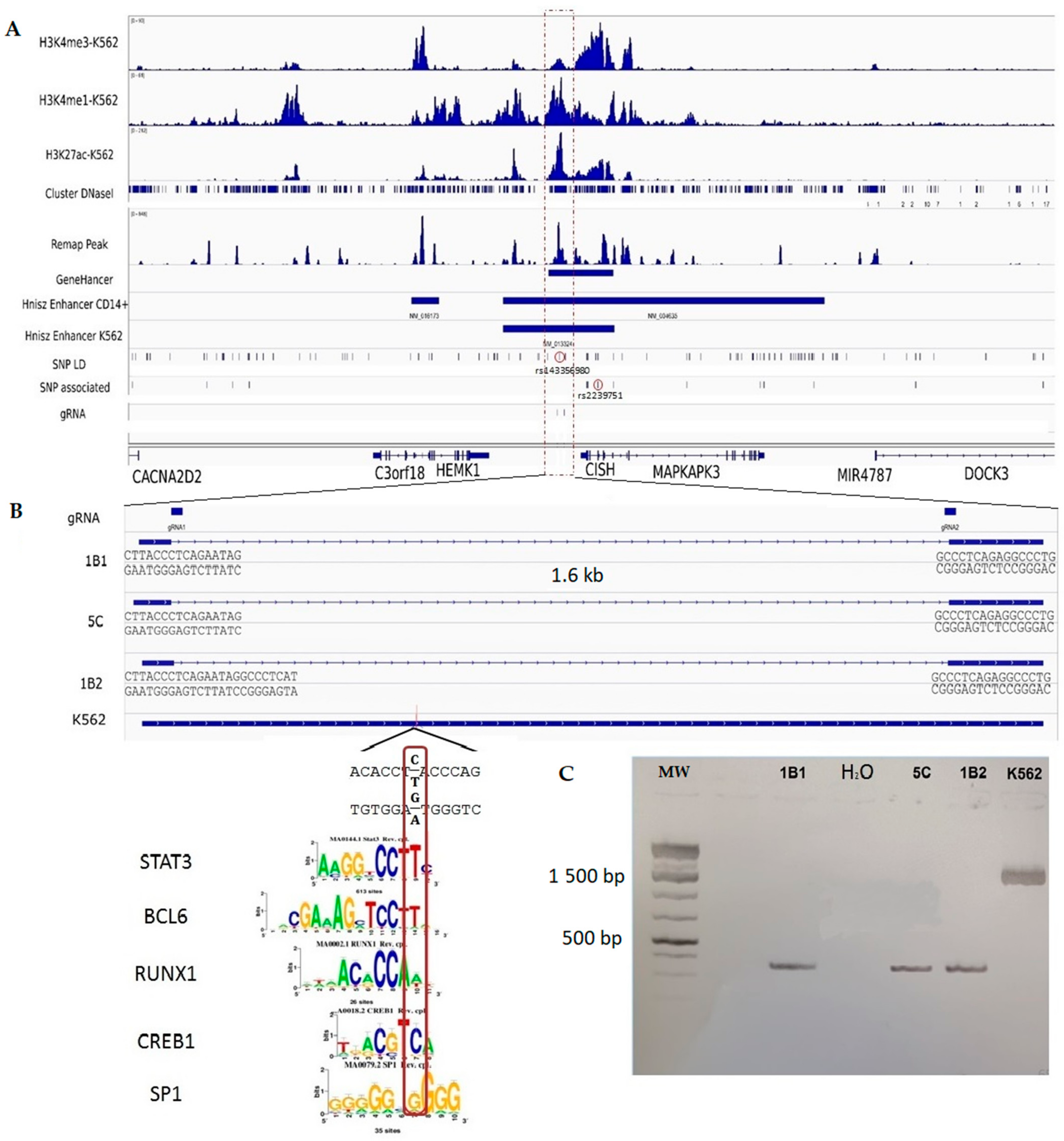
2.6. Prioritization and Annotation of Non-Coding Functional SNPs
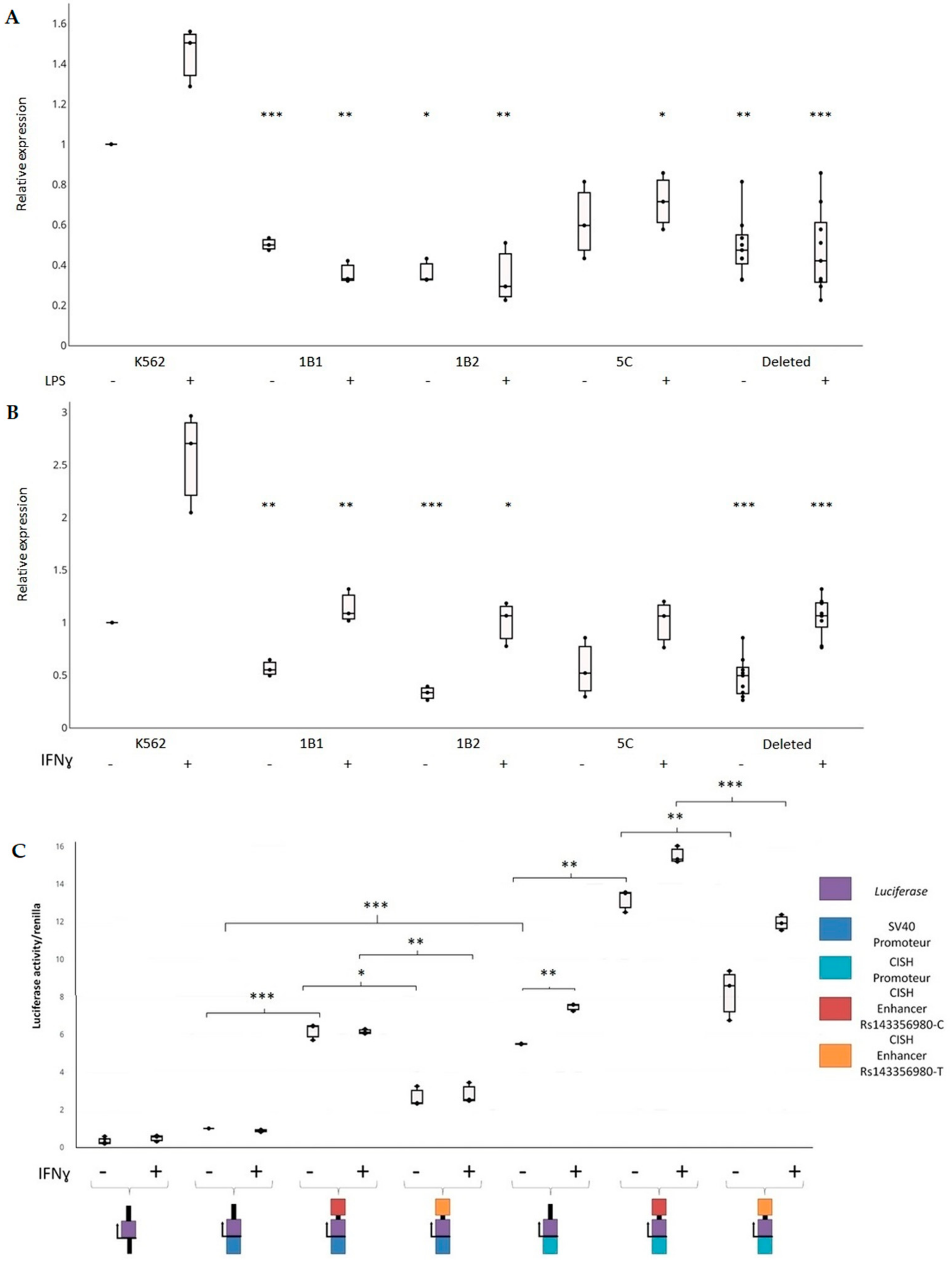
2.7. Experimental Validation of the Regulatory Effect of rs143356980
3. Discussion
4. Materials and Methods
4.1. Patients, Database, and Study Design
4.2. Cell Line and Culture Conditions
4.3. Single Nucleotide Polymorphism (SNP) Selection
4.4. Association Analyses and Statistical Methods
4.5. Protein–Protein Network and Functional Enrichment
4.6. SNP Annotation and Prioritization
4.7. Genome Editing Using the CRISPR-Cas9 Method
4.8. cDNA Synthesis and qRT-PCR
4.9. Gene Reporter Assays
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CISH | cytokine inducible SH2 containing protein |
| eQTL | expression quantitative trait loci |
| FDR | false discovery rate |
| GWAS | genome-wide association studies |
| KEGG | Kyoto encyclopedia of genes and genomes |
| LPS | lipopolysaccharide |
| PTP | protein tyrosine phosphatase |
| RCTs | randomized clinical trials |
| ROC | receiving operating characteristic |
| RSAT | regulatory sequence analysis tools |
| SNP | single nucleotide polymorphism |
| SOFA | sequential organ failure assessment |
| SOCS | suppressor of cytokine signaling |
References
- Stevenson, E.K.; Rubenstein, A.R.; Radin, G.T.; Wiener, R.S.; Walkey, A.J. Two decades of mortality trends among patients with severe sepsis: A comparative meta-analysis. Crit. Care Med. 2014, 42, 625–631. [Google Scholar] [CrossRef]
- Caironi, P.; Tognoni, G.; Masson, S.; Fumagalli, R.; Pesenti, A.; Romero, M.; Fanizza, C.; Caspani, L.; Faenza, S.; Grasselli, G.; et al. Albumin Replacement in Patients with Severe Sepsis or Septic Shock. N. Engl. J. Med. 2014, 370, 1412–1421. [Google Scholar] [CrossRef] [PubMed]
- Ranieri, V.M.; Thompson, B.T.; Barie, P.S.; Dhainaut, J.-F.; Douglas, I.S.; Finfer, S.; Gårdlund, B.; Marshall, J.C.; Rhodes, A.; Artigas, A.; et al. Drotrecogin Alfa (Activated) in Adults with Septic Shock. N. Engl. J. Med. 2012, 366, 2055–2064. [Google Scholar] [CrossRef] [PubMed]
- Sprung, C.L.; Annane, D.; Keh, D.; Moreno, R.; Singer, M.; Freivogel, K.; Weiss, Y.G.; Benbenishty, J.; Kalenka, A.; Forst, H.; et al. Hydrocortisone Therapy for Patients with Septic Shock. N. Engl. J. Med. 2008, 358, 111–124. [Google Scholar] [CrossRef] [PubMed]
- Valenta, J.; Brodska, H.; Drabek, T.; Hendl, J.; Kazda, A. High-dose selenium substitution in sepsis: A prospective randomized clinical trial. Intensiv. Care Med. 2011, 37, 808–815. [Google Scholar] [CrossRef]
- Boomer, J.S.; To, K.; Chang, K.C.; Takasu, O.; Osborne, D.F.; Walton, A.H.; Bricker, T.L.; Jarman, S.D., 2nd; Kreisel, D.; Krupnick, A.S.; et al. Immunosuppression in patients who die of sepsis and multiple organ failure. JAMA 2011, 306, 2594–2605. [Google Scholar] [CrossRef] [PubMed]
- Esper, A.M.; Moss, M.; Lewis, C.A.; Nisbet, R.; Mannino, D.M.; Martin, G.S. The role of infection and comorbidity: Factors that influence disparities in sepsis. Crit. Care Med. 2006, 34, 2576–2582. [Google Scholar] [CrossRef]
- Cohen, J.; Opal, S.; Calandra, T. Sepsis studies need new direction. Lancet Infect. Dis. 2012, 12, 503–505. [Google Scholar] [CrossRef]
- Mira, J.P.; Cariou, A.; Grall, F.; Delclaux, C.; Losser, M.R.; Heshmati, F.; Cheval, C.; Monchi, M.; Teboul, J.L.; Riche, F.; et al. Association of TNF2, a TNF-alpha promoter polymorphism, with septic shock susceptibility and mortality: A multicenter study. JAMA 1999, 282, 561–568. [Google Scholar] [CrossRef]
- Lowe, P.R.; Galley, H.F.; Abdel-Fattah, A.; Webster, N.R. Influence of interleukin-10 polymorphisms on interleukin-10 expression and survival in critically ill patients. Crit. Care Med. 2003, 31, 34–38. [Google Scholar] [CrossRef]
- Clark, M.F.; Baudouin, S.V. A systematic review of the quality of genetic association studies in human sepsis. Intensive Care Med. 2006, 32, 1706–1712. [Google Scholar] [CrossRef] [PubMed]
- Manolio, T.A. Genomewide Association Studies and Assessment of the Risk of Disease. N. Engl. J. Med. 2010, 363, 166–176. [Google Scholar] [CrossRef]
- The International Meningococcal Genetics Consortium; Davila, S.; Wright, V.J.; Khor, C.C.; Sim, K.S.; Binder, A.; Breunis, W.B.; Inwald, D.; Nadel, S.; Betts, H.; et al. Genome-wide association study identifies variants in the CFH region associated with host susceptibility to meningococcal disease. Nat. Genet. 2010, 42, 772–776. [Google Scholar] [CrossRef]
- Timmann, C.; Thye, T.; Vens, M.; Evans, J.; May, J.; Ehmen, C.; Sievertsen, J.; Muntau, B.; Ruge, G.; Loag, W.; et al. Genome-wide association study indicates two novel resistance loci for severe malaria. Nat. Cell Biol. 2012, 489, 443–446. [Google Scholar] [CrossRef] [PubMed]
- Rautanen, A.; Mills, T.C.; Gordon, A.C.; Hutton, P.; Steffens, M.; Nuamah, R.; Chiche, J.-D.; Parks, T.; Chapman, S.J.; Davenport, E.E.; et al. Genome-wide association study of survival from sepsis due to pneumonia: An observational cohort study. Lancet Respir. Med. 2015, 3, 53–60. [Google Scholar] [CrossRef]
- Scherag, A.; Schöneweck, F.; Kesselmeier, M.; Taudien, S.; Platzer, M.; Felder, M.; Sponholz, C.; Rautanen, A.; Hill, A.V.; Hinds, C.J.; et al. Genetic Factors of the Disease Course after Sepsis: A Genome-Wide Study for 28 Day Mortality. EBioMedicine 2016, 12, 239–246. [Google Scholar] [CrossRef]
- Flores, C. Host genetics shapes adult sepsis survival. Lancet Respir. Med. 2015, 3, 7–8. [Google Scholar] [CrossRef]
- Man, M.; Close, S.L.; Shaw, A.D.; Bernard, G.R.; Douglas, I.S.; Kaner, R.J.; Payen, D.; Vincent, J.-L.; Fossceco, S.; Janes, J.M.; et al. Beyond single-marker analyses: Mining whole genome scans for insights into treatment responses in severe sepsis. Pharm. J. 2013, 13, 218–226. [Google Scholar] [CrossRef]
- Hotchkiss, R.S.; Monneret, G.; Payen, D. Immunosuppression in sepsis: A novel understanding of the disorder and a new therapeutic approach. Lancet Infect. Dis. 2013, 13, 260–268. [Google Scholar] [CrossRef]
- Riché, F.; Gayat, E.; Barthélémy, R.; Le Dorze, M.; Matéo, J.; Payen, D. Reversal of neutrophil-to-lymphocyte count ratio in early versus late death from septic shock. Crit. Care 2015, 19, 1–10. [Google Scholar] [CrossRef]
- Carter, T.C.; Ye, Z.; Ivacic, L.C.; Budi, N.; Rose, W.E.; Shukla, S.K. Association of variants in selected genes mediating host immune response with duration of Staphylococcus aureus bacteremia. Genes Immun. 2020, 21, 240–248. [Google Scholar] [CrossRef]
- Ba, M.; Rawat, S.; Lao, R.; Grous, M.; Salmon, M.; Halayko, A.J.; Gerthoffer, W.T.; Singer, C.A. Differential regulation of cytokine and chemokine expression by MK2 and MK3 in airway smooth muscle cells. Pulm. Pharmacol. Ther. 2018, 53, 12–19. [Google Scholar] [CrossRef]
- Carow, B.; Gao, Y.; Terán, G.; Yang, X.O.; Dong, C.; Yoshimura, A.; Rottenberg, M.E. CISH controls bacterial burden early after infection with Mycobacterium tuberculosis in mice. Tuberculosis 2017, 107, 175–180. [Google Scholar] [CrossRef]
- Gaestel, M. What goes up must come down: Molecular basis of MAPKAP kinase 2/3-dependent regulation of the inflammatory response and its inhibition. Biol. Chem. 2013, 394, 1301–1315. [Google Scholar] [CrossRef]
- Khor, C.C.; Vannberg, F.O.; Chapman, S.J.; Guo, H.; Wong, S.H.; Walley, A.J.; Vukcevic, D.; Rautanen, A.; Mills, T.C.; Chang, K.C.; et al. CISH and susceptibility to infectious diseases. N. Engl. J. Med. 2010, 362, 2092–2101. [Google Scholar] [CrossRef]
- Tsukada, H.; Ochi, H.; Maekawa, T.; Abe, H.; Fujimoto, Y.; Tsuge, M.; Takahashi, H.; Kumada, H.; Kamatani, N.; Nakamura, Y.; et al. A Polymorphism in MAPKAPK3 Affects Response to Interferon Therapy for Chronic Hepatitis C. Gastroenterology 2009, 136, 1796–1805.e6. [Google Scholar] [CrossRef]
- Reid, A.L.; Wang, Y.; Samani, A.; Hightower, R.M.; Lopez, M.A.; Gilbert, S.R.; Ianov, L.; Crossman, D.K.; Dell’Italia, L.J.; Millay, D.P.; et al. DOCK3 is a dosage-sensitive regulator of skeletal muscle and Duchenne muscular dystrophy-associated pathologies. Hum. Mol. Genet. 2020, 29, 2855–2871. [Google Scholar] [CrossRef]
- Namekata, K.; Watanabe, H.; Guo, X.; Kittaka, D.; Kawamura, K.; Kimura, A.; Harada, C.; Harada, T. Dock3 regulates BDNF-TrkB signaling for neurite outgrowth by forming a ternary complex with Elmo and RhoG. Genes Cells 2012, 17, 688–697. [Google Scholar] [CrossRef]
- Wiltrout, K.; Ferrer, A.; Van De Laar, I.; Namekata, K.; Harada, T.; Klee, E.W.; Zimmerman, M.T.; Cousin, M.A.; Kempainen, J.L.; Babovic-Vuksanovic, D.; et al. Variants in DOCK3 cause developmental delay and hypotonia. Eur. J. Hum. Genet. 2019, 27, 1225–1234. [Google Scholar] [CrossRef]
- Chapple, C.E.; Robisson, B.; Spinelli, L.; Guien, C.; Becker, E.; Brun, C. Extreme multifunctional proteins identified from a human protein interaction network. Nat. Commun. 2015, 6, 7412. [Google Scholar] [CrossRef]
- Liu, X.; Ye, L.; Bai, Y.; Mojidi, H.; Simister, N.E.; Zhu, X. Activation of the JAK/STAT-1 signaling pathway by IFN-gamma can down-regulate functional expression of the MHC class I-related neonatal Fc receptor for IgG. J. Immunol. 2008, 181, 449–463. [Google Scholar] [CrossRef]
- Pellegrino, M.J.; Habecker, B.A. STAT3 integrates cytokine and neurotrophin signals to promote sympathetic axon regeneration. Mol. Cell. Neurosci. 2013, 56, 272–282. [Google Scholar] [CrossRef] [PubMed]
- Hnisz, D.; Abraham, B.; Lee, T.I.; Lau, A.; Saint-André, V.; Sigova, A.A.; Hoke, H.A.; Young, R.A. Super-Enhancers in the Control of Cell Identity and Disease. Cell 2013, 155, 934–947. [Google Scholar] [CrossRef] [PubMed]
- Cheneby, J.; Gheorghe, M.; Artufel, M.; Mathelier, A.; Ballester, B. ReMap 2018: An updated atlas of regulatory regions from an integrative analysis of DNA-binding ChIP-seq experiments. Nucleic Acids Res. 2018, 46, D267–D275. [Google Scholar] [CrossRef]
- Nguyen, N.T.T.; Contreras-Moreira, B.; Castro-Mondragon, J.A.; Santana-Garcia, W.; Ossio, R.; Robles-Espinoza, C.D.; Bahin, M.; Collombet, S.; Vincens, P.; Thieffry, D.; et al. RSAT 2018: Regulatory sequence analysis tools 20th anniversary. Nucleic Acids Res. 2018, 46, W209–W214. [Google Scholar] [CrossRef]
- González, A.; Artufel, M.; Rihet, P. TAGOOS: Genome-wide supervised learning of non-coding loci associated to complex phenotypes. Nucleic Acids Res. 2019, 47, e79. [Google Scholar] [CrossRef]
- Wang, J.; Ullah, A.Z.D.; Chelala, C. IW-Scoring: An Integrative Weighted Scoring framework for annotating and prioritizing genetic variations in the noncoding genome. Nucleic Acids Res. 2018, 46, e47. [Google Scholar] [CrossRef]
- Fishilevich, S.; Nudel, R.; Rappaport, N.; Hadar, R.; Plaschkes, I.; Stein, T.I.; Rosen, N.; Kohn, A.; Twik, M.; Safran, M.; et al. GeneHancer: Genome-wide integration of enhancers and target genes in GeneCards. Database 2017, 2017. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.-M.; Luo, X.-G.; Liang, J.-H.; Zhang, C.; Meng, X.-P.; Guo, D.-W. Critical selection of internal control genes for quantitative real-time RT-PCR studies in lipopolysaccharide-stimulated human THP-1 and K562 cells. Biochem. Biophys. Res. Commun. 2012, 427, 366–372. [Google Scholar] [CrossRef]
- Sutherland, J.A.; Turner, A.R.; Mannoni, P.; McGann, L.E.; Turc, J.M. Differentiation of K562 leukemia cells along erythroid, macrophage, and megakaryocyte lineages. J. Boil. Response Modif. 1986, 5, 250–262. [Google Scholar]
- De Chanville, C.B.; Chousterman, B.G.; Hamon, P.; Laviron, M.; Guillou, N.; Loyher, P.L.; Meghraoui-Kheddar, A.; Barthelemy, S.; Deterre, P.; Boissonnas, A.; et al. Sepsis Triggers a Late Expansion of Functionally Impaired Tissue-Vascular Inflammatory Monocytes During Clinical Recovery. Front. Immunol. 2020, 11, 675. [Google Scholar] [CrossRef] [PubMed]
- Lukaszewicz, A.C.; Grienay, M.; Resche-Rigon, M.; Pirracchio, R.; Faivre, V.; Boval, B.; Payen, D. Monocytic HLA-DR expression in intensive care patients: Interest for prognosis and secondary infection prediction. Crit. Care Med. 2009, 37, 2746–2752. [Google Scholar]
- Bress, A.; Han, J.; Patel, S.R.; Desai, A.A.; Mansour, I.; Groo, V.; Progar, K.; Shah, E.; Stamos, T.D.; Wing, C.; et al. Association of Aldosterone Synthase Polymorphism (CYP11B2 −344T > C) and Genetic Ancestry with Atrial Fibrillation and Serum Aldosterone in African Americans with Heart Failure. PLoS ONE 2013, 8, e71268. [Google Scholar]
- Baron, M.; Davignon, J.L. Inhibition of IFN-gamma-induced STAT1 tyrosine phosphorylation by human CMV is mediated by SHP2. J. Immunol. 2008, 181, 5530–5536. [Google Scholar] [CrossRef]
- Chemnitz, J.M.; Parry, R.V.; Nichols, K.E.; June, C.H.; Riley, J.L. SHP-1 and SHP-2 Associate with Immunoreceptor Tyrosine-Based Switch Motif of Programmed Death 1 upon Primary Human T Cell Stimulation, but Only Receptor Ligation Prevents T Cell Activation. J. Immunol. 2004, 173, 945–954. [Google Scholar] [CrossRef]
- McCafferty, D.-M.; Craig, A.W.B.; Senis, Y.A.; Greer, P.A. Absence of Fer Protein-Tyrosine Kinase Exacerbates Leukocyte Recruitment in Response to Endotoxin. J. Immunol. 2002, 168, 4930–4935. [Google Scholar] [CrossRef]
- Khajah, M.; Andonegui, G.; Chan, R.; Craig, A.W.; Greer, P.A.; McCafferty, D.M. Fer Kinase Limits Neutrophil Chemotaxis toward End Target Chemoattractants. J. Immunol. 2013, 190, 2208–2216. [Google Scholar] [CrossRef] [PubMed]
- Le, K.T.T.; Matzaraki, V.; Netea, M.G.; Wijmenga, C.; Moser, J.; Kumar, V. Functional Annotation of Genetic Loci Associated With Sepsis Prioritizes Immune and Endothelial Cell Pathways. Front. Immunol. 2019, 10, 1949. [Google Scholar] [CrossRef] [PubMed]
- Zhang, A.; Gu, W.; Lu, H.; Zeng, L.; Zhang, L.; Du, D.; Hao, J.; Wen, D.; Wang, X.; Jiang, J. Genetic contribution of suppressor of cytokine signalling polymorphisms to the susceptibility to infection after traumatic injury. Clin. Exp. Immunol. 2018, 194, 93–102. [Google Scholar] [CrossRef]
- Hindorff, L.A.; Sethupathy, P.; Junkins, H.A.; Ramos, E.M.; Mehta, J.P.; Collins, F.S.; Manolio, T.A. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl. Acad. Sci. USA 2009, 106, 9362–9367. [Google Scholar] [CrossRef]
- Nicolae, D.L.; Gamazon, E.; Zhang, W.; Duan, S.; Dolan, M.E.; Cox, N.J. Trait-Associated SNPs Are More Likely to Be eQTLs: Annotation to Enhance Discovery from GWAS. PLoS Genet. 2010, 6, e1000888. [Google Scholar] [CrossRef]
- Ji, L.-D.; Xu, W.-N.; Chai, P.-F.; Zheng, W.; Qian, H.-X.; Xu, J. Polymorphisms in the CISH gene are associated with susceptibility to tuberculosis in the Chinese Han population. Infect. Genet. Evol. 2014, 28, 240–244. [Google Scholar] [CrossRef]
- Zhao, L.; Chu, H.; Xu, X.; Yue, J.; Li, H.; Wang, M. Association Between Single-Nucleotide Polymorphism in CISH Gene and Susceptibility to Tuberculosis in Chinese Han Population. Cell Biophys. 2014, 68, 529–534. [Google Scholar] [CrossRef]
- Hu, Z.; Yang, J.; Wu, Y.; Xiong, G.; Wang, Y.; Yang, J.; Deng, L. Polymorphisms in CISH Gene Are Associated with Persistent Hepatitis B Virus Infection in Han Chinese Population. PLoS ONE 2014, 9, e100826. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, A.; Masuhara, M.; Mitsui, K.; Yokouchi, M.; Ohtsubo, M.; Misawa, H.; Miyajima, A.; Yoshimura, A. CIS, a Cytokine Inducible SH2 Protein, Is a Target of the JAK-STAT5 Pathway and Modulates STAT5 Activation. Blood 1997, 89, 3148–3154. [Google Scholar] [CrossRef] [PubMed]
- Yoshimura, A.; Ohkubo, T.; Kiguchi, T.; Jenkins, N.; Gilbert, D.; Copeland, N.; Hara, T.; Miyajima, A. A novel cytokine-inducible gene CIS encodes an SH2-containing protein that binds to tyrosine-phosphorylated interleukin 3 and erythropoietin receptors. EMBO J. 1995, 14, 2816–2826. [Google Scholar] [CrossRef]
- Delconte, R.B.; Kolesnik, T.B.; Dagley, L.F.; Rautela, J.; Shi, W.; Putz, E.M.; Stannard, K.; Zhang, J.-G.; Teh, C.; Firth, M.; et al. CIS is a potent checkpoint in NK cell–mediated tumor immunity. Nat. Immunol. 2016, 17, 816–824. [Google Scholar] [CrossRef]
- Feng, T.; Liao, X.; Yang, X.; Yang, C.; Lin, F.; Guo, Y.; Kang, Y.; Li, H. A shift toward inhibitory receptors and impaired effector functions on NK cells contribute to immunosuppression during sepsis. J. Leukoc. Biol. 2020, 107, 57–67. [Google Scholar] [CrossRef]
- Queval, C.J.; Song, O.-R.; Carralot, J.-P.; Saliou, J.-M.; Bongiovanni, A.; Deloison, G.; Deboosère, N.; Jouny, S.; Iantomasi, R.; Delorme, V.; et al. Mycobacterium tuberculosis Controls Phagosomal Acidification by Targeting CISH-Mediated Signaling. Cell Rep. 2017, 20, 3188–3198. [Google Scholar] [CrossRef]
- Fairfax, B.P.; Humburg, P.; Makino, S.; Naranbhai, V.; Wong, D.; Lau, E.; Jostins, L.; Plant, K.; Andrews, R.; McGee, C.; et al. Innate Immune Activity Conditions the Effect of Regulatory Variants upon Monocyte Gene Expression. Science 2014, 343, 1246949. [Google Scholar] [CrossRef]
- Davenport, E.E.; Burnham, K.L.; Radhakrishnan, J.; Humburg, P.; Hutton, P.; Mills, T.C.; Rautanen, A.; Gordon, A.C.; Garrard, C.; Hill, A.V.S.; et al. Genomic landscape of the individual host response and outcomes in sepsis: A prospective cohort study. Lancet Respir. Med. 2016, 4, 259–271. [Google Scholar] [CrossRef]
- Van der Poll, T.; van de Veerdonk, F.L.; Scicluna, B.P.; Netea, M.G. The immunopathology of sepsis and potential therapeutic targets. Nat Rev Immunol. 2017, 17, 407–420. [Google Scholar] [CrossRef]
- Bernard, G.R.; Vincent, J.-L.; Laterre, P.-F.; LaRosa, S.P.; Dhainaut, J.-F.; Lopez-Rodriguez, A.; Steingrub, J.S.; Garber, G.E.; Helterbrand, J.D.; Ely, E.W.; et al. Efficacy and Safety of Recombinant Human Activated Protein C for Severe Sepsis. N. Engl. J. Med. 2001, 344, 699–709. [Google Scholar] [CrossRef]
- Singer, M.; Deutschman, C.S.; Seymour, C.W.; Shankar-Hari, M.; Annane, D.; Bauer, M.; Bellomo, R.; Bernard, G.R.; Chiche, J.-D.; Coopersmith, C.M.; et al. The Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3). JAMA 2016, 315, 801–810. [Google Scholar] [CrossRef] [PubMed]
- Bozza, F.A.; Salluh, J.I.; Japiassu, A.M.; Soares, M.; Assis, E.F.; Gomes, R.N.; Bozza, M.T.; Castro-Faria-Neto, H.C.; Bozza, P.T. Cytokine profiles as markers of disease severity in sepsis: A multiplex analysis. Crit. Care 2007, 11, R49. [Google Scholar] [CrossRef]
- Hotchkiss, R.S.; Monneret, G.; Payen, D. Sepsis-induced immunosuppression: From cellular dysfunctions to immunotherapy. Nat. Rev. Immunol. 2013, 13, 862–874. [Google Scholar] [CrossRef] [PubMed]
- Aulchenko, Y.S.; Ripke, S.; Isaacs, A.; Van Duijn, C.M. GenABEL: An R library for genome-wide association analysis. Bioinformatics 2007, 23, 1294–1296. [Google Scholar] [CrossRef] [PubMed]
- R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020.
- Venables, W.N.; Ripley, B.D. Modern Applied Statistics with S, 4th ed.; Springer: New York, NY, USA, 2002. [Google Scholar]
- Ward, L.D.; Kellis, M. HaploReg v4: Systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and disease. Nucleic Acids Res. 2016, 44, D877–D881. [Google Scholar] [CrossRef]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009, 37, 1–13. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, D. BioCarta. Biotech Software & Internet Report. Comput. Softw. J. Scient 2001, 2, 117–120. [Google Scholar]
- Ferré, Q.; Charbonnier, G.; Sadouni, N.; Lopez, F.; Kermezli, Y.; Spicuglia, S.; Capponi, C.; Ghattas, B.; Puthier, D. OLOGRAM: Determining significance of total overlap length between genomic regions sets. Bioinformatics 2019. [Google Scholar] [CrossRef]
- Arnold, M.; Raffler, J.; Pfeufer, A.; Suhre, K.; Kastenmüller, G. SNiPA: An interactive, genetic variant-centered annotation browser. Bioinformatics 2015, 31, 1334–1336. [Google Scholar] [CrossRef] [PubMed]
- Naito, Y.; Hino, K.; Bono, H.; Ui-Tei, K. CRISPRdirect: Software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 2015, 31, 1120–1123. [Google Scholar] [CrossRef]
- Mali, P.; Yang, L.; Esvelt, K.M.; Aach, J.; Guell, M.; DiCarlo, J.E.; Norville, J.E.; Church, G.M. RNA-Guided Human Genome Engineering via Cas9. Science 2013, 339, 823–826. [Google Scholar] [CrossRef]
| SNP | CHR: Position | Alleles (MAF) | Risk Allele | p Value (q Value) | Unadjusted OR (Adjusted OR) | LD Region r2 > 0.8 | Genes Containing SNP | Genes in LD | Phenotype Associated |
|---|---|---|---|---|---|---|---|---|---|
| rs16857698 | 3: 145685067 | A > G (0.014) | G | 1.75 × 10−9 (7.53 × 10−4) | 3.52 (4.51) | 3:145665563-145685067 | D7 | ||
| rs5029231 | 3: 145701146 | C > T (0.019) | T | 1.37 × 10−8 (1.73 × 10−3) | 3.54 (3.99) | 3:145686379-145759412 | D7 | ||
| rs6763296 | 3: 145709314 | T > C (0.018) | C | 2.55 × 10−9 (7.53 × 10−4) | 3.93 (4.67) | 3:145686379-145759412 | D7 | ||
| rs16857836 | 3: 145752473 | G > T (0.014) | T | 5.51 × 10−10 (4.89 × 10−4) | 3.79 (5.20) | 3:145686379-145759412 | D7 | ||
| rs4544 | 8: 143994806 | T > C (0.010) | C | 8.86 × 10−9 (1.31 × 10−3) | 3.24 (6.87) | 8:143983592-144018027 | CYP11B2 | GML | D7 |
| rs11991278 | 8: 144001245 | C > T (0.010) | T | 8.48 × 10−9 (1.31 × 10−3) | 3.23 (6.87) | 8:143983592-144018027 | CYP11B2 | GML | D7 |
| rs6981918 | 8: 144007939 | C > A (0.010) | A | 8.74 × 10−9 (1.31 × 10−3) | 3.22 (6.85) | 8:143983592-144018027 | CYP11B2 | GML | D7 |
| rs956727 | 9: 86846933 | A > G (0.009) | G | 3.22 × 10−8 (2.60 × 10−3) | 4.85 (17.43) | 9:86814655-86862104 | SLC28A3 | D7 | |
| rs7974468 | 12: 112927208 | G > A (0.013) | A | 1.60 × 10−8 (1.78 × 10−3) | 3.10 (3.34) | 12:112819245-112985734 | PTPNN11 | RPH3A | D7 |
| rs10849640 | 12: 119712137 | G > A (0.116) | A | 3.22 × 10−8 (2.60 × 10−3) | 1.65 (2.25) | 12:119712137-119725314 | D7 | ||
| rs10849641 | 12: 119721354 | C > T (0.115) | T | 2.65 × 10−8 (2.60 × 10−3) | 1.67 (2.27) | 12:119712137-119725314 | D7 | ||
| rs10849642 | 12: 119725314 | C > T (0.117) | T | 4.04 × 10−8 (2.99 × 10−3) | 1.62 (2.25) | 12:119712137-119725314 | D7 | ||
| rs12491812 | 3: 50556581 | C > T (0.011) | T | 4.18 × 10−11 (1.25 × 10−5) | 5.62 (7.32) | 3:50534635-50645413 | CACNA2D2 | C3orf18, HEMK1, CISH | D28 |
| rs2239753 | 3: 50645158 | T > C (0.011) | C | 2.80 × 10−11 (1.25 × 10−5) | 4.97 (7.02) | 3:50555933-50645413 | CISH | C3orf18, HEMK1, CACNA2D2 | D28 |
| rs2239752 | 3: 50645413 | C > T (0.011) | T | 5.43 × 10−10 (4.86 × 10−5) | 4.32 (5.62) | 3:50555933-50645413 | CISH | C3orf18, HEMK1, CACNA2D2 | D28 |
| rs2239751 | 3: 50647888 | A > C (0.011) | C | 5.21 × 10−10 (4.86 × 10−5) | 4.32 (5.62) | 3:50531386-50875635 | CISH | C3orf18, HEMK1, CACNA2D2, MAPKAPK3, DOCK3 | D28 |
| rs743753 | 3: 50651395 | C > T (0.011) | T | 5.21 × 10−10 (4.86 × 10−5) | 4.32 (5.62) | 3:50531386-50875635 | MAPKAPK3 | C3orf18, HEMK1, CACNA2D2, CISH, DOCK3 | D28 |
| rs616689 | 3: 50668532 | G > A (0.014) | A | 1.87 × 10−10 (3.35 × 10−5) | 5.09 (5.79) | 3:50647343-50751643 | MAPKAPK3 | CISH, DOCK3 | D28 |
| rs9879397 | 3: 50685642 | G > A (0.012) | A | 8.79 × 10−9 (6.57 × 10−4) | 4.00 (4.86) | 3:50647343-50751643 | MAPKAPK3 | CISH, DOCK3 | D28 |
| rs2170840 | 3: 50686517 | A > C (0.014) | C | 1.87 × 10−10 (3.35 × 10−5) | 5.09 (5.78) | 3:50647343-50751643 | MAPKAPK3 | CISH, DOCK3 | D28 |
| rs12492982 | 3: 50698155 | C > T (0.011) | T | 4.18 × 10−11 (1.25 × 10−5) | 4.85 (7.32) | 3:50531386-50875635 | MAPKAPK3 | C3orf18, HEMK1, CACNA2D2, CISH, DOCK3 | D28 |
| rs2035484 | 3: 50721892 | A > G (0.011) | G | 5.21 × 10−10 (4.86 × 10−5) | 4.32 (5.62) | NA | DOCK3 | D28 | |
| rs17051403 | 3: 50751643 | C > A (0.011) | A | 5.21 × 10−10 (4.86 × 10−5) | 4.32 (5.62) | 3:50531386-50875635 | DOCK3 | C3orf18, HEMK1, CACNA2D2, CISH, MAPKAPK3 | D28 |
| rs17072628 | 3: 65229760 | G > A (0.012) | A | 8.25 × 10−9 (6.57 × 10−4) | 3.88 (4.96) | 3:65214495-65241577 | D28 | ||
| rs7840669 | 8: 89929277 | A > G (0.015) | G | 2.38 × 10−8 (1.53 × 10−3) | 4.23 (3.77) | 8:89901960-90133835 | D28 | ||
| rs7953683 | 12: 79993704 | C > T (0.024) | T | 3.07 × 10−8 (1.72 × 10−3) | 2.06 (2.07) | 12:79919466-80080618 | PAWR | D28 | |
| rs1502522 | 17: 51544776 | A > G (0.029) | G | 2.57 × 10−8 (1.53 × 10−3) | 3.24 (2.64) | 17:51519876-51590268 | D28 | ||
| rs1393467 | 17: 51560869 | T > C (0.029) | C | 2.57 × 10−8 (1.53 × 10−3) | 3.24 (2.64) | 17:51519876-51590268 | D28 |
| Early Death | Late Death | |||
|---|---|---|---|---|
| Nb of Risk Alleles | Non-Adjusted OR | Adjusted OR | Non Adjusted OR | Adjusted OR |
| 0 | 1 | 1 | 1 | 1 |
| 1 | 1.56 | 1.44 | 4.75 | 3.53 |
| 2 | 1.71 | 1.58 | 7.95 | 7.82 |
| 3 | 3.38 | 3.48 | 17.96 | 17.86 |
| ≥4 | 9.40 | 12.33 | 70.75 | 123.35 |
| HGNC Symbol | UniProt Symbol | Presence in the Interactome | Presence in the Sub-Network |
|---|---|---|---|
| ADAP2 | ADAP2_HUMAN | yes | yes |
| ANKFN1 | ANKF1_HUMAN | no | no |
| ANKH | ANKH_HUMAN | yes | yes |
| ARIH1 | ARI1_HUMAN | yes | yes |
| ASIC2 | ASIC2_HUMAN | yes | yes |
| ATAD5 | ATAD5_HUMAN | yes | yes |
| C3orf18 | CC018_HUMAN | no | no |
| C6orf170 | BROMI_HUMAN | no | no |
| CACNA2D2 | CA2D2_HUMAN | no | no |
| CISH | CISH_HUMAN | yes | yes |
| CRLF3 | CRLF3_HUMAN | yes | yes |
| CYP11B2 | C11B2_HUMAN | no | no |
| DOCK3 | DOCK3_HUMAN | yes | yes |
| DPYD | DPYD_HUMAN | yes | yes |
| EHMT1 | EHMT1_HUMAN | yes | yes |
| FER | FER_HUMAN | yes | yes |
| GML | GML_HUMAN | no | no |
| GPR158 | GP158_HUMAN | yes | yes |
| GREM2 | GREM2_HUMAN | yes | no |
| HECTD4 | HECD4_HUMAN | no | no |
| HEMK1 | HEMK1_HUMAN | yes | yes |
| IFIT1B | IFT1B_HUMAN | no | no |
| KPTN | KPTN_HUMAN | yes | yes |
| LBP | LBP_HUMAN | yes | yes |
| LIPA | LICH_HUMAN | no | no |
| MAPKAPK3 | MAPK3_HUMAN | yes | yes |
| NAPA | SNAA_HUMAN | yes | yes |
| NCKAP5 | NCKP5_HUMAN | yes | yes |
| NLN | NEUL_HUMAN | no | no |
| OSCP1 | OSCP1_HUMAN | no | no |
| PAWR | PAWR_HUMAN | yes | yes |
| PPFIA2 | LIPA2_HUMAN | yes | yes |
| PTPN11 | PTN11_HUMAN | yes | yes |
| RBFOX1 | RFOX1_HUMAN | yes | yes |
| RPL6 | RL6_HUMAN | yes | yes |
| RNF135 | RN135_HUMAN | yes | yes |
| SLC15A1 | S15A1_HUMAN | yes | no |
| SLC28A3 | S28A3_HUMAN | no | no |
| SLFN13 | SLN13_HUMAN | no | no |
| SLFN12L | SN12L_HUMAN | no | no |
| SYNC | SYNCI_HUMAN | yes | yes |
| SYT1 | SYT1_HUMAN | yes | yes |
| TRAFD1 | TRAD1_HUMAN | yes | no |
| U6 | SNR27_HUMAN | yes | yes |
| WDR85 | DPH7_HUMAN | no | no |
| Initial Cohort (n = 832) | Survivors after 7 Days (n = 698) | ||||
|---|---|---|---|---|---|
| All Cohort | Dead before Day 7 | Alive at Day 7 | Dead before Day 28 | Alive at Day 28 | |
| n = 832 | n = 134 | n = 698 | n = 111 | n = 587 | |
| Age (year) | 67.3 ± 22.7 a | 70.8 ± 13.1 | 65.8 ± 24.0 | 71.4 ± 14.8 | 63.4 ± 24.5 |
| Male sex (%) | 467 (56.1) | 78 (58.2) | 389 (55.7) | 66 (59.5) | 323 (55) |
| Drotrecogin alpha (%) | 407 (48.9) | 59 (44) | 348 (49.9) | 44 (39.6) | 304 (51.8) |
| Prior and preexisting conditions (%) | |||||
| Hypertension | 297 (35.7) | 53 (39.6) | 244 (35.0) | 43 (38.7) | 201 (34.2) |
| Myocardial infarction | 129 (15.5) | 30 (22.4) | 99 (14.2) | 28 (25.2) | 71 (12.1) |
| Congestive cardiomyopathy | 67 (8.1) | 7 (5.2) | 60 (8.6) | 20 (18) | 40 (6.8) |
| Diabetes | 169 (20.3) | 30 (22.4) | 139 (19.9) | 27 (24.3) | 112 (19.1) |
| Pancreatitis | 30 (3.6) | 5 (3.7) | 25 (3.6) | 4 (3.6) | 21 (3.6) |
| Liver disease | 14 (1.7) | 3 (2.2) | 11 (1.6) | 4 (3.6) | 7 (1.2) |
| COPD b | 226 (27.2) | 39 (29.1) | 187 (26.8) | 36 (32.4) | 151 (25.7) |
| Cancer | 169 (20.3) | 33 (24.6) | 136 (19.5) | 29 (26.1) | 107 (18.2) |
| Apache II score | 25 ± 10 | 28 ± 11.8 | 25 ± 10 | 28 ± 9 | 24 ± 10 |
| SOFA score c | 8 ± 3 | 9.5 ± 3 | 8 ± 3 | 8 ± 3 | 8 ± 3 |
| log(IL-6) d | 6.4 ± 3.1 | 7.4 ± 4.0 | 6.1 ± 2.9 | 6.3 ± 2.6 | 6.1 ± 2.9 |
| Survival at Day 7 | Survival between Day 7 and Day 28 | |||
|---|---|---|---|---|
| p-Value | OR (CI) a | p-Value | OR (CI) | |
| Age (years) | 4.98 × 10−4 | 1.03 (1.01; 1.05) | 1.62 × 10−3 | 1.03 (1.01; 1.05) |
| Gender (M/F) | NS | NS | NS | NS |
| Hypertension | NS | NS | NS | NS |
| Myocardial infarct | NS | NS | NS | NS |
| Cardiomyopathy | NS | NS | 1.53 × 10−3 | 3.11 (1.52; 6.24) |
| Chronic obstructive pulmonary disease (COPD) | NS | NS | NS | NS |
| Diabetes | NS | NS | NS | NS |
| Liver disease | NS | NS | NS | NS |
| Malignancy | NS | NS | NS | NS |
| Pre-infusion APACHE score | NS | NS | NS | NS |
| Log of baseline IL-6 concentration | 2.64 × 10−6 | 1.29 (1.16; 1.44) | NS | NS |
| Treatment by Activated Prot C or not | NS | NS | 2.22 × 10−2 | 0.56 (0.33; 0.91) |
| baseline SOFA score (without neuro component) | 4.77 × 10−2 | 1.10 (1.00; 1.22) | 1.74 × 10−2 | 1.14 (1.02; 1.27) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rosier, F.; Brisebarre, A.; Dupuis, C.; Baaklini, S.; Puthier, D.; Brun, C.; Pradel, L.C.; Rihet, P.; Payen, D. Genetic Predisposition to the Mortality in Septic Shock Patients: From GWAS to the Identification of a Regulatory Variant Modulating the Activity of a CISH Enhancer. Int. J. Mol. Sci. 2021, 22, 5852. https://doi.org/10.3390/ijms22115852
Rosier F, Brisebarre A, Dupuis C, Baaklini S, Puthier D, Brun C, Pradel LC, Rihet P, Payen D. Genetic Predisposition to the Mortality in Septic Shock Patients: From GWAS to the Identification of a Regulatory Variant Modulating the Activity of a CISH Enhancer. International Journal of Molecular Sciences. 2021; 22(11):5852. https://doi.org/10.3390/ijms22115852
Chicago/Turabian StyleRosier, Florian, Audrey Brisebarre, Claire Dupuis, Sabrina Baaklini, Denis Puthier, Christine Brun, Lydie C. Pradel, Pascal Rihet, and Didier Payen. 2021. "Genetic Predisposition to the Mortality in Septic Shock Patients: From GWAS to the Identification of a Regulatory Variant Modulating the Activity of a CISH Enhancer" International Journal of Molecular Sciences 22, no. 11: 5852. https://doi.org/10.3390/ijms22115852
APA StyleRosier, F., Brisebarre, A., Dupuis, C., Baaklini, S., Puthier, D., Brun, C., Pradel, L. C., Rihet, P., & Payen, D. (2021). Genetic Predisposition to the Mortality in Septic Shock Patients: From GWAS to the Identification of a Regulatory Variant Modulating the Activity of a CISH Enhancer. International Journal of Molecular Sciences, 22(11), 5852. https://doi.org/10.3390/ijms22115852








