Nationwide Study of Drug Resistance Mutations in HIV-1 Infected Individuals under Antiretroviral Therapy in Brazil
Abstract
1. Introduction
2. Results
2.1. Characterization of the Study Population
2.2. Prevalence of Surveillance Drug-Resistance Mutations
2.3. A Gradual Alteration on the Prevalence of Surveillance Drug-Resistance Mutations
2.4. A Shift on Treatment Scheme during the Years
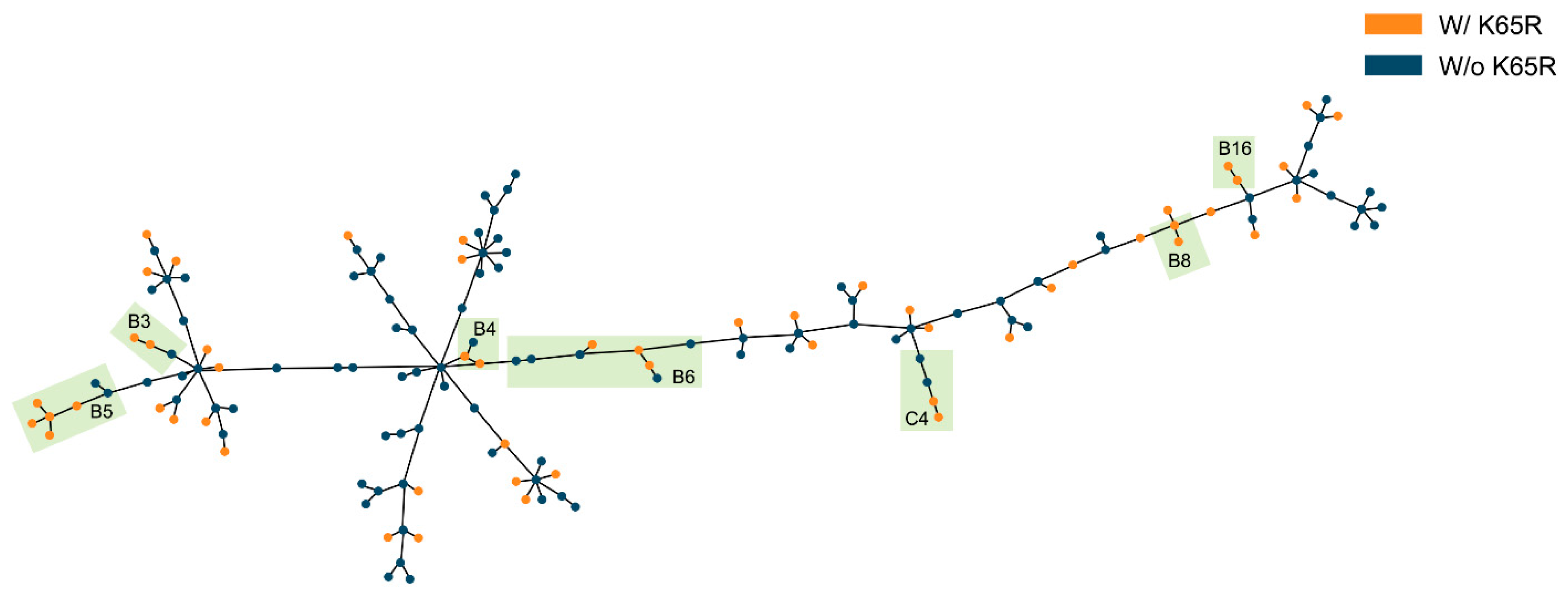
2.5. Evidence for Transmission of K65R
3. Discussion
4. Materials and Methods
4.1. Study Population
4.2. HIV-1 Subtypes and Drug Resistance Mutations
4.3. Phylogenetic Analysis of K65R Sequences and Transmission Clusters
4.4. HLA Binding Affinity Predictions
4.5. HLA Prevalence in the Brazilian Population
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Joint United Nations Programme on HIV/AIDS. UNAIDS Data 2020; UNAIDS: Geneva, Switzerland, 2020.
- Souza, P.R.B.D., Jr.; Szwarcwald, C.L.; Castilho, E.A.D. Self-rated health by HIV-infected individuals undergoing antiretroviral therapy in Brazil. Cad. Saúde Pública/Ministério Saúde, Fundação Oswaldo Cruz. Esc. Nac. Saúde Pública 2011. [Google Scholar] [CrossRef]
- Teeraananchai, S.; Kerr, S.J.; Amin, J.; Ruxrungtham, K.; Law, M.G. Life expectancy of HIV-positive people after starting combination antiretroviral therapy: A meta-analysis. HIV Med. 2017, 18, 256–266. [Google Scholar] [CrossRef]
- Trickey, A.; May, M.T.; Vehreschild, J.J.; Obel, N.; Gill, M.J.; Crane, H.M.; Boesecke, C.; Patterson, S.; Grabar, S.; Cazanave, C.; et al. Survival of HIV-positive patients starting antiretroviral therapy between 1996 and 2013: A collaborative analysis of cohort studies. Lancet HIV 2017, 4, e349–e356. [Google Scholar] [CrossRef]
- Saag, M.S.; Gandhi, R.T.; Hoy, J.F.; Landovitz, R.J.; Thompson, M.A.; Sax, P.E.; Smith, D.M.; Benson, C.A.; Buchbinder, S.P.; Del Rio, C.; et al. Antiretroviral Drugs for Treatment and Prevention of HIV Infection in Adults: 2020 Recommendations of the International Antiviral Society-USA Panel. JAMA 2020, 324, 1651–1669. [Google Scholar] [CrossRef]
- Lundgren, J.; Phillips, A. Prevention of HIV transmission by antiretroviral therapy. Lancet HIV 2018, 5, e108–e109. [Google Scholar] [CrossRef]
- Eisinger, R.W.; Dieffenbach, C.W.; Fauci, A.S. HIV viral load and transmissibility of HIV infection undetectable equals untransmittable. JAMA 2019, 321, 451–452. [Google Scholar] [CrossRef]
- Cohen, M.S.; Chen, Y.Q.; McCauley, M.; Gamble, T.; Hosseinipour, M.C.; Kumarasamy, N.; Hakim, J.G.; Kumwenda, J.; Grinsztejn, B.; Pilotto, J.H.S.; et al. Antiretroviral Therapy for the Prevention of HIV-1 Transmission. N. Engl. J. Med. 2016, 375, 830–839. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T.S.O.; Leite, T.F.; Freitas, S.Z.; Cesar, G.A.; De Rezende, G.R.; Lindenberg, A.D.S.C.; Guimarães, M.L.; Motta-Castro, A.R.C. HIV-1 molecular epidemiology, transmission clusters and transmitted drug resistance mutations in central Brazil. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef]
- Booth, C.L.; Geretti, A.M. Prevalence and determinants of transmitted antiretroviral drug resistance in HIV-1 infection. J. Antimicrob. Chemother. 2007, 59, 1047–1056. [Google Scholar] [CrossRef]
- Arruda, M.B.; Boullosa, L.T.; Cardoso, C.C.; da Costa, C.M.; Brites, C.; de Lima, S.T.S.; Kaminski, H.T.; Aleixo, A.W.; Esposito, A.O.P.; Cavalcanti, A.M.S.; et al. Brazilian network for HIV drug resistance surveillance (HIV-BresNet): A survey of treatment-naive individuals. J. Int. AIDS Soc. 2018, 21. [Google Scholar] [CrossRef]
- Geretti, A.M. Epidemiology of antiretroviral drug resistance in drug-naïve persons. Curr. Opin. Infect. Dis. 2007, 20, 22–32. [Google Scholar] [CrossRef] [PubMed]
- Frentz, D.; Boucher, C.A.B.; Van De Vijver, D.A.M.C. Temporal changes in the epidemiology of transmission of drug-resistant HIV-1 across the world. AIDS Rev. 2012, 14, 17–27. [Google Scholar]
- Sui, H.; Gui, T.; Jia, L.; Guo, W.; Han, J.; Liu, Y.; Bao, Z.; Li, H.; Li, J.; Li, L. Different Frequencies of Drug Resistance Mutations among HIV-1 Subtypes Circulating in China: A Comprehensive Study. PLoS ONE 2014, 9, e91803. [Google Scholar] [CrossRef]
- Chaplin, B.; Eisen, G.; Idoko, J.; Onwujekwe, D.; Idigbe, E.; Adewole, I.; Gashau, W.; Meloni, S.; Sarr, A.D.; Sankalé, J.L.; et al. Impact of HIV type 1 subtype on drug resistance mutations in Nigerian patients failing first-line therapy. AIDS Res. Hum. Retrovir. 2011, 27, 71–80. [Google Scholar] [CrossRef] [PubMed]
- Bahls, L.D.; Canezin, P.H.; Reiche, E.M.V.; Fernandez, J.C.C.; Dias, J.R.C.; Meneguetti, V.A.F.; Ueda, L.T.; Bertolini, D.A. Moderate prevalence of HIV-1 transmitted drug resistance mutations in southern Brazil. AIDS Res. Ther. 2019. [Google Scholar] [CrossRef] [PubMed]
- Duani, H.; Aleixo, A.W.; Tupinambás, U. Trends and predictors of HIV-1 acquired drug resistance in Minas Gerais, Brazil: 2002–2012. Braz. J. Infect. Dis. 2017, 21, 148–154. [Google Scholar] [CrossRef] [PubMed]
- Cavalcanti, A.M.S.; Lacerda, H.R.; De Brito, A.M.; Pereira, S.; Medeiros, D.; Oliveira, S. Antiretroviral resistance in individuals presenting therapeutic failure and subtypes of the human immunodeficiency virus type 1 in the Northeast Region of Brazil. Mem. Inst. Oswaldo Cruz 2007, 102, 785–792. [Google Scholar] [CrossRef][Green Version]
- Toledo, P.V.M.; de Carvalho, D.S.; Romagnoli, L.; Marcinko, G.; da Cunha, C.A.; de Souza, M.N.; Brindeiro, R.; de Queiroz-Telles, F. HIV-1 genotypic resistance profile of patients failing antiretroviral therapy in Paraná, Brazil. Braz. J. Infect. Dis. 2010, 14, 360–371. [Google Scholar] [CrossRef][Green Version]
- Coelho, L.P.O.; Matsuda, E.M.; Nogueira, R.S.; de Moraes, M.J.; Jamal, L.F.; Madruga, J.V.R.; Tancredi, M.V.; de Leão, A.C.Q.; de Faria Romero Soldi, G.; de Macedo Brígido, L.F.; et al. Prevalence of HIV-1 transmitted drug resistance and viral suppression among recently diagnosed adults in São Paulo, Brazil. Arch. Virol. 2019, 164, 699–706. [Google Scholar] [CrossRef]
- Ministério da Saúde. Clinical Protocol and Therapeutic Guidelines for Handling HIV Infection in Adults, 1st ed.; Ministério da Saúde, Secretaria de Vigilância em Saúde, Departamento de Vigilância, Prevenção e Controle das Infecções Sexualmente Transmissíveis, do HIV/Aids e das Hepatites Virais: Brasilia, Brazil, 2018; ISBN 9788533426436.
- Benzaken, A.S.; Pereira, G.F.M.; Costa, L.; Tanuri, A.; Santos, A.F.; Soares, M.A. Antiretroviral treatment, government policy and economy of HIV/AIDS in Brazil: Is it time for HIV cure in the country? AIDS Res. Ther. 2019, 16. [Google Scholar] [CrossRef]
- European AIDS Clinical Society. European AIDS clinical society guidelines. Version 10.0. November 2019. IEEE Trans. Sonics Ultrason. 2019, 6–11. [Google Scholar]
- Panel on Antiretroviral Guidelines for Adults and Adolescents Guidelines for the Use of Antiretroviral Agents in Adults and Adolescents with HIV. Department of Health and Human Services. Available online: https://clinicalinfo.hiv.gov/sites/default/files/guidelines/documents/AdultandAdolescentGL.pdf (accessed on 18 May 2021).
- Günthard, H.F.; Calvez, V.; Paredes, R.; Pillay, D.; Shafer, R.W.; Wensing, A.M.; Jacobsen, D.M.; Richman, D.D. Human Immunodeficiency Virus Drug Resistance: 2018 Recommendations of the International Antiviral Society-USA Panel. Clin. Infect. Dis. 2019, 68, 177–187. [Google Scholar] [CrossRef] [PubMed]
- Bennett, D.E.; Camacho, R.J.; Otelea, D.; Kuritzkes, D.R.; Fleury, H.; Kiuchi, M.; Heneine, W.; Kantor, R.; Jordan, M.R.; Schapiro, J.M.; et al. Drug Resistance Mutations for Surveillance of Transmitted HIV-1 Drug-Resistance: 2009 Update. PLoS ONE 2009, 4, e4724. [Google Scholar] [CrossRef] [PubMed]
- Brites, C.; Pinto-Neto, L.; Medeiros, M.; Nunes, E.; Sprinz, E.; Carvalho, M. Extensive variation in drug-resistance mutational profile of Brazilian patients failing antiretroviral therapy in five large Brazilian cities. Braz. J. Infect. Dis. 2016, 20, 323–329. [Google Scholar] [CrossRef]
- Touloumi, G.; Pantazis, N.; Pillay, D.; Paraskevis, D.; Chaix, M.-L.; Bucher, H.C.; Kucherer, C.; Zangerle, R.; Kran, A.-M.B.; Porter, K.; et al. Impact of HIV-1 Subtype on CD4 Count at HIV Seroconversion, Rate of Decline, and Viral Load Set Point in European Seroconverter Cohorts. Clin. Infect. Dis. 2013, 56, 888–897. [Google Scholar] [CrossRef]
- Wensing, A.M.; Calvez, V.; Ceccherini-Silberstein, F.; Charpentier, C.; Günthard, H.F.; Paredes, R.; Shafer, R.W.; Richman, D.D. 2019 update of the drug resistance mutations in HIV-1. Top. Antivir. Med. 2019, 27, 111–121. [Google Scholar]
- White, K.L.; Chen, J.M.; Feng, J.Y.; Margot, N.A.; Ly, J.K.; Ray, A.S.; Macarthur, H.L.; McDermott, M.J.; Swaminathan, S.; Miller, M.D. The K65R reverse transcriptase mutation in HIV-1 reverses the excision phenotype of zidovudine resistance mutations. Antivir. Ther. 2006, 11, 155–163. [Google Scholar]
- Brenner, B.G.; Coutsinos, D. The K65R mutation in HIV-1 reverse transcriptase: Genetic barriers, resistance profile and clinical implications. HIV Ther. 2009, 3, 583–594. [Google Scholar] [CrossRef]
- Smit, E.; White, E.; Clark, D.; Churchill, D.; Zhang, H.; Collins, S.; Pillay, D.; Sabin, C.; Nelson, M.; Winston, A.; et al. An association between K65R and HIV-1 subtype C viruses in patients treated with multiple NRTIs. J. Antimicrob. Chemother. 2017, 72, 2075–2082. [Google Scholar] [CrossRef]
- Inzaule, S.C.; Weidle, P.J.; Yang, C.; Ndiege, K.; Hamers, R.L.; Rinke de Wit, T.F.; Thomas, T.; Zeh, C. Prevalence and dynamics of the K65R drug resistance mutation in HIV-1-infected infants exposed to maternal therapy with lamivudine, zidovudine and either nevirapine or nelfinavir in breast milk. J. Antimicrob. Chemother. 2016, 71, 1619–1626. [Google Scholar] [CrossRef]
- Theys, K.; Vercauteren, J.; Snoeck, J.; Zazzi, M.; Camacho, R.J.; Torti, C.; Schülter, E.; Clotet, B.; Sönnerborg, A.; De Luca, A.; et al. HIV-1 subtype is an independent predictor of reverse transcriptase mutation k65r in HIV-1 patients treated with combination antiretroviral therapy including tenofovir. Antimicrob. Agents Chemother. 2013, 57, 1053–1056. [Google Scholar] [CrossRef] [PubMed]
- Coutsinos, D.; Invernizzi, C.F.; Moisi, D.; Oliveira, M.; Martinez-Cajas, J.L.; Brenner, B.G.; Wainberg, M.A. A Template-Dependent Dislocation Mechanism Potentiates K65R Reverse Transcriptase Mutation Development in Subtype C Variants of HIV-1. PLoS ONE 2011, 6, e20208. [Google Scholar] [CrossRef] [PubMed]
- Ministério da Saúde. Boletim Epidemiológico de HIV e Aids; Ministério da Saúde Secretaria de Vigilância em Saúde Departamento de Vigilância, Prevenção e Controle das Infecções Sexualmente Transmissíveis, do HIV/Aids e das Hepatites Virais: Brasilia, Brazil, 2018; ISBN 1517-1159.
- Guindon, S.; Dufayard, J.-F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New Algorithms and Methods to Estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Sprinz, E.; Netto, E.M.; Lima, M.P.J.S.; Furtado, J.J.D.; Da Eira, M.; Zajdenverg, R.; Madruga, J.V.; Lewi, D.S.; Pedro, R.J.; Soares, M.A. Primary antiretroviral drug resistance among HIV Type 1-infected individuals in Brazil. AIDS Res. Hum. Retrovir. 2009, 25, 861–867. [Google Scholar] [CrossRef] [PubMed]
- Lopes, C.A.F.; Soares, M.A.; Falci, D.R.; Sprinz, E. The Evolving Genotypic Profile of HIV-1 Mutations Related to Antiretroviral Treatment in the North Region of Brazil. Biomed. Res. Int. 2015, 2015. [Google Scholar] [CrossRef]
- Diaz, R.S.; Inocêncio, L.A.; Sucupira, M.C.A.; Pereira, A.A.; Hunter, J.; Ferreira, J.E.; Araújo, L.V.; Souza, D.F.C.; Sabino, E.C. The Virological and Immunological Characteristics of the HIV-1-Infected Population in Brazil: From Initial Diagnosis to Impact of Antiretroviral Use. PLoS ONE 2015, 10, e0139677. [Google Scholar] [CrossRef]
- Wolf, K.; Walter, H.; Beerenwinkel, N.; Keulen, W.; Kaiser, R.; Hoffmann, D.; Lengauer, T.; Selbig, J.; Vandamme, A.M.; Korn, K.; et al. Tenofovir Resistance and Resensitization. Antimicrob. Agents Chemother. 2003, 47, 3478–3484. [Google Scholar] [CrossRef]
- Götte, M.; Arion, D.; Parniak, M.A.; Wainberg, M.A. The M184V Mutation in the Reverse Transcriptase of Human Immunodeficiency Virus Type 1 Impairs Rescue of Chain-Terminated DNA Synthesis. J. Virol. 2000, 74, 3579–3585. [Google Scholar] [CrossRef]
- Turner, D.; Brenner, B.G.; Routy, J.P.; Petrella, M.; Wainberg, M.A. Rationale for maintenance of the M184v resistance mutation in human immunodeficiency virus type 1 reverse transcriptase in treatment experienced patients. New Microbiol. 2004, 27, 31–39. [Google Scholar]
- Pingarilho, M.; Pimentel, V.; Diogo, I.; Fernandes, S.; Miranda, M.; Pineda-Pena, A.; Libin, P.; Theys, K.; Martins, M.R.O.; Vandamme, A.M.; et al. Increasing Prevalence of HIV-1 Transmitted Drug Resistance in Portugal: Implications for First Line Treatment Recommendations. Viruses 2020, 12, 1238. [Google Scholar] [CrossRef] [PubMed]
- Franzetti, M.; Violin, M.; Antinori, A.; De Luca, A.; Ceccherini-Silberstein, F.; Gianotti, N.; Torti, C.; Bonora, S.; Zazzi, M.; Balotta, C. Trends and correlates of HIV-1 resistance among subjects failing an antiretroviral treatment over the 2003–2012 decade in Italy. BMC Infect. Dis. 2014, 14, 398. [Google Scholar] [CrossRef]
- Rocheleau, G.; Brumme, C.J.; Shoveller, J.; Lima, V.D.; Harrigan, P.R. Longitudinal trends of HIV drug resistance in a large Canadian cohort, 1996–2016. Clin. Microbiol. Infect. 2018, 24, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Ministério da Saúde. Protocolo Clínico E Diretrizes Terapêtuicas Para O Manejo Da Infecção Pelo HIV; Ministério da Saúde Secretaria de Vigilância em Saúde Departamento de Vigilância, Prevenção e Controle das Infecções Sexualmente Transmissíveis, do HIV/Aids e das Hepatites Virais: Brasilia, Brazil, 2013.
- Bacheler, L.T.; Anton, E.D.; Kudish, P.; Baker, D.; Bunville, J.; Krakowski, K.; Bolling, L.; Aujay, M.; Wang, X.V.; Ellis, D.; et al. Human immunodeficiency virus type 1 mutations selected in patients failing efavirenz combination therapy. Antimicrob. Agents Chemother. 2000, 44, 2475–2484. [Google Scholar] [CrossRef] [PubMed]
- Alcaro, S.; Alteri, C.; Artese, A.; Ceccherini-Silberstein, F.; Costa, G.; Ortuso, F.; Bertoli, A.; Forbici, F.; Santoro, M.M.; Parrotta, L.; et al. Docking Analysis and Resistance Evaluation of Clinically Relevant Mutations Associated with the HIV-1 Non-nucleoside Reverse Transcriptase Inhibitors Nevirapine, Efavirenz and Etravirine. Chem. Med. Chem. 2011, 6, 2203–2213. [Google Scholar] [CrossRef] [PubMed]
- Bacheler, L.; Jeffrey, S.; Hanna, G.; D’Aquila, R.; Wallace, L.; Logue, K.; Cordova, B.; Hertogs, K.; Larder, B.; Buckery, R.; et al. Genotypic Correlates of Phenotypic Resistance to Efavirenz in Virus Isolates from Patients Failing Nonnucleoside Reverse Transcriptase Inhibitor Therapy. J. Virol. 2001, 75, 4999–5008. [Google Scholar] [CrossRef]
- Garforth, S.J.; Lwatula, C.; Prasad, V.R. The lysine 65 residue in HIV-1 reverse transcriptase function and in nucleoside analog drug resistance. Viruses 2014, 6, 4080–4094. [Google Scholar] [CrossRef]
- Ayitewala, A.; Kyeyune, F.; Ainembabazi, P.; Nabulime, E.; Kato, C.D.; Nankya, I. Comparison of HIV drug resistance profiles across HIV-1 subtypes A and D for patients receiving a tenofovir-based and zidovudine-based first line regimens in Uganda. AIDS Res. Ther. 2020, 17. [Google Scholar] [CrossRef]
- Luo, X.L.; Mo, L.D.; Su, G.S.; Huang, J.P.; Wu, J.Y.; Su, H.Z.; Huang, W.H.; Luo, S.D.; Ni, Z.Y. Incidence and types of HIV-1 drug resistance mutation among patients failing first-line antiretroviral therapy. J. Pharmacol. Sci. 2019, 139, 275–279. [Google Scholar] [CrossRef]
- Parker Hudson, F.; Mulenga, L.; Westfall, A.O.; Warrier, R.; Mweemba, A.; Saag, M.S.; Stringer, J.S.A.; Eron, J.J.; Chi, B.H. Evolution of HIV-1 drug resistance after virological failure of first-line antiretroviral therapy in Lusaka, Zambia. Antivir. Ther. 2019, 24, 291–300. [Google Scholar] [CrossRef]
- Van Zyl, G.U.; Liu, T.F.; Claassen, M.; Engelbrecht, S.; de Oliveira, T.; Preiser, W.; Wood, N.T.; Travers, S.; Shafer, R.W. Trends in Genotypic HIV-1 Antiretroviral Resistance between 2006 and 2012 in South African Patients Receiving First-and Second-Line Antiretroviral Treatment Regimens. PLoS ONE 2013, 8, e67188. [Google Scholar] [CrossRef]
- Karade, S.; Chaturbhuj, D.N.; Sen, S.; Joshi, R.K.; Kulkarni, S.S.; Shankar, S.; Gangakhedkar, R.R. HIV drug resistance following a decade of the free antiretroviral therapy programme in India: A review. Int. J. Infect. Dis. 2018, 66, 33–41. [Google Scholar] [CrossRef] [PubMed]
- Tittle, V.; Boffito, M.; McOwan, A.; Whitlock, G. Antiretroviral resistance and management after pre-exposure to prophylaxis. Lancet HIV 2020, 7, e84. [Google Scholar] [CrossRef]
- Charpentier, C.; Lambert-Niclot, S.; Visseaux, B.; Morand-Joubert, L.; Storto, A.; Larrouy, L.; Landman, R.; Calvez, V.; Marcelin, A.-G.; Descamps, D. Evolution of the K65R, K103N and M184V/I reverse transcriptase mutations in HIV-1-infected patients experiencing virological failure between 2005 and 2010. J. Antimicrob. Chemother. 2013, 68, 2197–2198. [Google Scholar] [CrossRef] [PubMed]
- Theys, K.; Snoeck, J.; Vercauteren, J.; Abecasis, A.B.; Vandamme, A.M.; Camacho, R.J.; Mansinho, K.; Miranda, A.C.; Aldir, I.; Ventura, F.; et al. Decreasing population selection rates of resistance mutation K65R over time in HIV-1 patients receiving combination therapy including tenofovir. J. Antimicrob. Chemother. 2013, 68, 419–423. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Dolling, D.; Sabin, C.; Delpech, V.; Smit, E.; Pozniak, A.; Asboe, D.; Brown, A.L.; Churchill, D.; Williams, I.; Geretti, A.M.; et al. Time trends in drug resistant HIV-1 infections in the United Kingdom up to 2009: Multicentre observational study. BMJ 2012, 345. [Google Scholar] [CrossRef]
- Reinheimer, C.; Wesner, A.; Keppler, O.T.; Doerr, H.W.; Herrmann, E.; Stürmer, M.; Stephan, C. Prevalence of K65R in patients treated with tenofovir disoproxil fumarate: Recommendations based on the Frankfurt HIV Cohort Study Resistance Database (FHCS-RD). Med. Microbiol. Immunol. 2016, 205, 315–320. [Google Scholar] [CrossRef]
- Ministério da Saúde. Protocolo Clínico E Diretrizes Terapêuticas Para Profilaxia Pré-Exposição (Prep) De Risco À Infecção Pelo HIV, 1st ed.; Ministério da Saúde Secretaria de Vigilância em Saúde Departamento de Vigilância, Prevenção e Controle das Infecções Sexualmente Transmissíveis, do HIV/Aids e das Hepatites Virais: Brasilia, Brazil, 2018; Volume 29, ISBN 978-85-334-2582-8.
- Gibas, K.M.; van den Berg, P.; Powell, V.E.; Krakower, D.S. Drug Resistance During HIV Pre-Exposure Prophylaxis. Drugs 2019, 79, 609–619. [Google Scholar] [CrossRef]
- Parikh, U.M.; Mellors, J.W. Should we fear resistance from tenofovir/emtricitabine preexposure prophylaxis? Curr. Opin. HIV AIDS 2016, 11, 49–55. [Google Scholar] [CrossRef]
- Margot, N.A.; Waters, J.M.; Miller, M.D. In vitro human immunodeficiency virus type 1 resistance selections with combinations of tenofovir and emtricitabine or abacavir and lamivudine. Antimicrob. Agents Chemother. 2006, 50, 4087–4095. [Google Scholar] [CrossRef]
- Diallo, K.; Götte, M.; Wainberg, M.A. Molecular impact of the M184V mutation in human immunodeficiency virus type 1 reverse transcriptase. Antimicrob. Agents Chemother. 2003, 47, 3377–3383. [Google Scholar] [CrossRef]
- Wagner, B.G.; Garcia-Lerma, J.G.; Blower, S. Factors limiting the transmission of HIV mutations conferring drug resistance: Fitness costs and genetic bottlenecks. Sci. Rep. 2012, 2. [Google Scholar] [CrossRef]
- Weber, J.; Chakraborty, B.; Weberova, J.; Miller, M.D.; Quiñones-Mateu, M.E. Diminished replicative fitness of primary human immunodeficiency virus type 1 isolates harboring the K65R mutation. J. Clin. Microbiol. 2005, 43, 1395–1400. [Google Scholar] [CrossRef] [PubMed]
- Chan, P.A.; Huang, A.; Kantor, R. Low prevalence of transmitted K65R and other tenofovir resistance mutations across different HIV-1 subtypes: Implications for pre-exposure prophylaxis. J. Int. AIDS Soc. 2012, 15. [Google Scholar] [CrossRef] [PubMed]
- Margot, N.A.; Wong, P.; Kulkarni, R.; White, K.; Porter, D.; Abram, M.E.; Callebaut, C.; Miller, M.D. Commonly transmitted HIV-1 drug resistance mutations in reverse-transcriptase and protease in antiretroviral treatment-naive patients and response to regimens containing tenofovir disoproxil fumarate or tenofovir alafenamide. J. Infect. Dis. 2017, 215, 920–927. [Google Scholar] [CrossRef] [PubMed]
- Rhee, S.Y.; Jordan, M.R.; Raizes, E.; Chua, A.; Parkin, N.; Kantor, R.; Van Zy, G.U.; Mukui, I.; Hosseinipour, M.C.; Frenkel, L.M.; et al. HIV-1 drug resistance mutations: Potential applications for point-of-care Genotypic resistance testing. PLoS ONE 2015, 10, e145772. [Google Scholar] [CrossRef]
- Gregson, J.; Tang, M.; Ndembi, N.; Hamers, R.L.; Marconi, V.C.; Brooks, K.; Theys, K.; Arruda, M.; Garcia, F.; Monge, S.; et al. Global epidemiology of drug resistance after failure of WHO recommended first-line regimens for adult HIV-1 infection: A multicentre retrospective cohort study. Lancet Infect. Dis. 2016, 16, 565–575. [Google Scholar] [CrossRef]
- Kløverpris, H.N.; Leslie, A.; Goulder, P. Role of HLA adaptation in HIV evolution. Front. Immunol. 2016, 6, 665. [Google Scholar] [CrossRef]
- Neumann-Haefelin, C. HLA-B27-mediated protection in HIV and hepatitis C virus infection and pathogenesis in spondyloarthritis: Two sides of the same coin? Curr. Opin. Rheumatol. 2013, 25, 426–433. [Google Scholar] [CrossRef]
- Roberts, R.L.; Wallace, M.C.; Jones, G.T.; van Rij, A.M.; Merriman, T.R.; Harrison, A.; White, D.; Stamp, L.K.; Ching, D.; Highton, J.; et al. Prevalence of HLA-B27 in the New Zealand population: Effect of age and ethnicity. Arthritis Res. Ther. 2013, 15, R158. [Google Scholar] [CrossRef]
- Reveille, J.D.; Hirsch, R.; Dillon, C.F.; Carroll, M.D.; Weisman, M.H. The prevalence of HLA-B27 in the US: Data from the US National Health and Nutrition Examination Survey, 2009. Arthritis Rheum. 2012, 64, 1407–1411. [Google Scholar] [CrossRef]
- Khan, M.A. HLA-B27 and its subtypes in world populations: Editorial review. Curr. Opin. Rheumatol. 1995, 7, 263–269. [Google Scholar] [CrossRef]
- Araújo, P.M.M.; Martins, J.S.; Osório, N.S. SNAPPy: A snakemake pipeline for scalable HIV-1 subtyping by phylogenetic pairing. Virus Evol. 2019, 5. [Google Scholar] [CrossRef] [PubMed]
- Schultz, A.K.; Zhang, M.; Bulla, I.; Leitner, T.; Korber, B.; Morgenstern, B.; Stanke, M. jpHMM: Improving the reliability of recombination prediction in HIV-1. Nucleic Acids Res. 2009, 37, 647–651. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013. [Google Scholar] [CrossRef]
- Lefort, V.; Longueville, J.E.; Gascuel, O. SMS: Smart Model Selection in PhyML. Mol. Biol. Evol. 2017, 34, 2422–2424. [Google Scholar] [CrossRef]
- Drummond, A.J.; Suchard, M.A.; Xie, D.; Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 2012, 29, 1969–1973. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, M.A.R.; Suchard, M.A. Bayesian analysis of elapsed times in continuous-time Markov chains. Can. J. Stat. 2008, 36, 355–368. [Google Scholar] [CrossRef]
- Araújo, P.M.M.; Carvalho, A.; Pingarilho, M.; Abecasis, A.B.; Osório, N.S. Characterization of a large cluster of HIV-1 A1 infections detected in Portugal and connected to several Western European countries. Sci. Rep. 2019, 9, 7223. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro-Gonçalves, B.; Francisco, A.P.; Vaz, C.; Ramirez, M.; Carriço, J.A. PHYLOViZ Online: Web-based tool for visualization, phylogenetic inference, analysis and sharing of minimum spanning trees. Nucleic Acids Res. 2016, 44, W246–W251. [Google Scholar] [CrossRef]
- Reynisson, B.; Alvarez, B.; Paul, S.; Peters, B.; Nielsen, M. NetMHCpan-4.1 and NetMHCIIpan-4.0: Improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data. Nucleic Acids Res. 2020, 48, W449–W454. [Google Scholar] [CrossRef]
- Gonzalez-Galarza, F.F.; McCabe, A.; dos Santos, E.J.M.; Jones, J.; Takeshita, L.; Ortega-Rivera, N.D.; Cid-Pavon, G.M.D.; Ramsbottom, K.; Ghattaoraya, G.; Alfirevic, A.; et al. Allele frequency net database (AFND) 2020 update: Gold-standard data classification, open access genotype data and new query tools. Nucleic Acids Res. 2020, 48, D783–D788. [Google Scholar] [CrossRef] [PubMed]

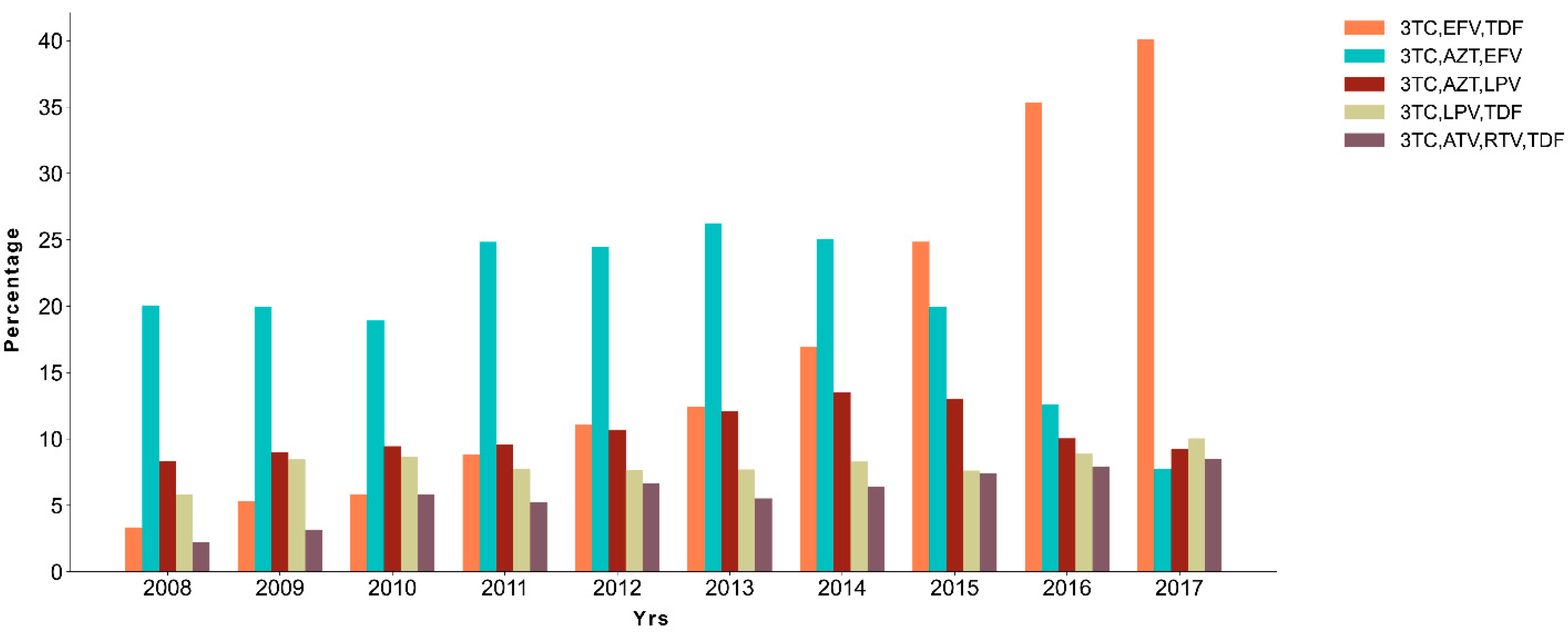

| HIV-1+ Individuals | Female | Male | |
|---|---|---|---|
| All ages | 20,226 (100.00%) | 8962 (44.3%) | 11,263 (55.7%) |
| <1 yrs old | 64 (0.32%) | 30 (0.15%) | 34 (0.17%) |
| 2–9 yrs old | 364 (1.80%) | 200 (0.99%) | 164 (0.81%) |
| 10–17 yrs old | 840 (4.15%) | 426 (2.11%) | 414 (2.05%) |
| 18–30 yrs old | 2443 (12.08%) | 1201 (5.94%) | 1242 (6.14%) |
| 30–49 yrs old | 12,426 (61.44%) | 5390 (26.65%) | 7036 (34.79%) |
| 50–79 yrs old | 4074 (20.14%) | 1709 (8.45%) | 2364 (11.69%) |
| >80 yrs old | 15 (0.07%) | 6 (0.03%) | 9 (0.04%) |
| Age (av. yrs ± std) | 39.55 ± 12.71 | 38.67 ± 13.15 | 40.25 ± 12.31 |
| Treatment (av. yrs ± std) | 2.98 ± 2.96 | 3.06 ± 2.96 | 2.92 ± 2.95 |
| Birth Federative Unit | HIV-1+ individuals | Treatment Scheme | HIV-1+ individuals |
| São Paulo | 4445 (21.98%) | 3TC,EFV,TDF | 4324 (21.38%) |
| Rio Grande do Sul | 2419 (11.96%) | 3TC,AZT,EFV | 4003 (19.79%) |
| Minas Gerais | 1878 (9.29%) | 3TC,AZT,LPV | 2258 (11.16%) |
| Rio de Janeiro | 1861 (9.20%) | 3TC,LPV,TDF | 1666 (8.24%) |
| Paraná | 1606 (7.94%) | 3TC,ATV,RTV,TDF | 1355 (6.70%) |
| Others | 8017 (39.64%) | Others | 6620 (32.73%) |
| 2008 | 2009 | 2010 | 2011 | 2012 | 2013 | 2014 | 2015 | 2016 | 2017 ** | |
|---|---|---|---|---|---|---|---|---|---|---|
| M184V | 276 (76.88) | 416 (73.37) | 375 (75.45) | 1076 (75.3) | 1781 (72.46) | 2082 (71.94) | 2004 (69.58) | 1874 (68.29) | 2684 * (53.21) | 697 (51.48) |
| K103N | 149 (41.5) | 252 (44.44) | 217 (43.66) | 638 (44.65) | 1083 (44.06) | 1215 (41.98) | 1180 (40.97) | 1293 * (47.12) | 2145 * (42.53) | 566 (41.8) |
| M41L | 125 (34.82) | 182 (32.1) | 155 (31.19) | 383 (26.8) | 561 * (22.28) | 585 * (20.21) | 425 * (14.76) | 374 (13.63) | 545 * (10.8) | 145 (10.71) |
| D67N | 114 (31.75) | 163 (28.75) | 149 (29.98) | 337 * (23.58) | 504 * (20.5) | 487 * (16.83) | 435 (15.1) | 398 (14.5) | 526 * (10.43) | 140 (10.34) |
| T215Y | 109 (30.36) | 144 (25.4) | 141 (28.37) | 299 * (20.92) | 403 * (16.4) | 434 (15) | 323 * (11.22) | 277 (10.09) | 336 * (6.66) | 72 (5.32) |
| K70R | 96 (26.74) | 133 (23.46) | 131 (26.36) | 308 * (21.55) | 451 * (18.35) | 459 * (15.86) | 368 * (12.78) | 344 (12.54) | 460 * (9.12) | 116 (8.57) |
| V82A | 82 (22.84) | 111 (19.58) | 91 (18.31) | 206 * (14.42) | 335 (13.63) | 313 * (10.82) | 251 * (8.72) | 242 (8.82) | 303 * (6.01) | 87 (6.43) |
| M46I | 81 (22.56) | 122 (21.52) | 89 (17.91) | 207 (14.49) | 302 (12.29) | 302 * (10.44) | 222 * (7.71) | 207 (7.54) | 324 (6.42) | 82 (6.06) |
| L210W | 75 (20.89) | 109 (19.22) | 96 (19.32) | 204 * (14.28) | 294 * (11.96) | 301 (10.4) | 199 * (6.91) | 172 (6.27) | 249 * (4.94) | 66 (4.87) |
| I54V | 73 (20.33) | 115 (20.28) | 79 (15.9) | 191 (13.37) | 269 * (10.94) | 273 (9.43) | 224 * (7.78) | 196 (7.14) | 241 * (4.78) | 58 (4.28) |
| G190A | 46 (12.81) | 80 (14.11) | 47 * (9.46) | 146 (10.22) | 239 (9.72) | 266 (9.19) | 271 (9.41) | 259 (9.44) | 392 * (7.77) | 102 (7.53) |
| P225H | 36 (10.03) | 56 (9.88) | 62 (12.47) | 179 (12.53) | 320 (13.02) | 403 (13.93) | 370 (12.85) | 390 (14.21) | 586 * (11.62) | 158 (11.67) |
| K65R | 8 (2.23) | 21 (3.7) | 23 (4.63) | 74 (5.18) | 161 (6.55) | 194 (6.7) | 246 * (8.74) | 374 * (13.63) | 589 * (11.68) | 164 (12.11) |
| WT SB Binding | K65R SB Binding | Allele Freq. in BR Pop. | |
|---|---|---|---|
| HLA-A01 | no | no | 9.21 |
| HLA-A02 | no | no | 25.94 |
| HLA-A03 | yes | yes | 9.26 |
| HLA-A24 | no | no | 10.00 |
| HLA-A26 | no | no | 3.35 |
| HLA-B07 | no | no | 6.92 |
| HLA-B08 | no | no | 5.12 |
| HLA-B15 | no | no | 9.08 |
| HLA-B27 | no | yes | 2.23 |
| HLA-B39 | no | no | 3.46 |
| HLA-B40 | no | no | 4.70 |
| HLA-B58 | yes | yes | 2.65 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Santos-Pereira, A.; Triunfante, V.; Araújo, P.M.M.; Martins, J.; Soares, H.; Poveda, E.; Souto, B.; Osório, N.S. Nationwide Study of Drug Resistance Mutations in HIV-1 Infected Individuals under Antiretroviral Therapy in Brazil. Int. J. Mol. Sci. 2021, 22, 5304. https://doi.org/10.3390/ijms22105304
Santos-Pereira A, Triunfante V, Araújo PMM, Martins J, Soares H, Poveda E, Souto B, Osório NS. Nationwide Study of Drug Resistance Mutations in HIV-1 Infected Individuals under Antiretroviral Therapy in Brazil. International Journal of Molecular Sciences. 2021; 22(10):5304. https://doi.org/10.3390/ijms22105304
Chicago/Turabian StyleSantos-Pereira, Ana, Vera Triunfante, Pedro M. M. Araújo, Joana Martins, Helena Soares, Eva Poveda, Bernardino Souto, and Nuno S. Osório. 2021. "Nationwide Study of Drug Resistance Mutations in HIV-1 Infected Individuals under Antiretroviral Therapy in Brazil" International Journal of Molecular Sciences 22, no. 10: 5304. https://doi.org/10.3390/ijms22105304
APA StyleSantos-Pereira, A., Triunfante, V., Araújo, P. M. M., Martins, J., Soares, H., Poveda, E., Souto, B., & Osório, N. S. (2021). Nationwide Study of Drug Resistance Mutations in HIV-1 Infected Individuals under Antiretroviral Therapy in Brazil. International Journal of Molecular Sciences, 22(10), 5304. https://doi.org/10.3390/ijms22105304









