Analysis of Results of Specific IgE in 100 Atopic Dermatitis Patients with the Use of Multiplex Examination ALEX2—Allergy Explorer
Abstract
1. Introduction
2. Results
- Blo t 5—Storage mites, Blomia tropicalis
- Aca s—Storage mites, Acarus siro
- Gly d 2, Lep d 2—Storage mites, NPC2 family
- Can f 1, Can f 2, Can f 4 and Can f 6—Lipocalins
- Fel d 1, Ory c 3—Uteroglobin
- Mala s 11—Mn superoxide dismutase
- Asp f 6, Bla g 9, Der p 20, Pen m 2—Arginine kinase
- Fag s 1, Mal d 1, Cor a 1.0401, Cor a 1.0103—PR-10 proteins
- Cyn d 1, Lol p 1, Phl p 1—Beta-expansins
- Phl p 2—Expansin
- Der p 21, Der p 23—Peritrophin-like domain.
- Secc pollen—Cultivated rye
- Cra c 6—Troponin C (North Sea shrimp)
- Hom g—Lobster
- Ara h 6-2S Albumin, Peanut, Ara h 1-Cupin (Vicillin-type, 7S globulin), Peanut
- Alt a 6—Enolase, Alternaria alternata
- Lep d 2, Der p 2, Der f 2—NPC2 family
- Der p 1—Cystein protease
- Der p 21, Der p 23—Peritrophin-like protein domain
- Der p 5, Der p 7—European house dust mite
- Loc m—Migratory locust
- Phl p 1—Beta expansin
- Phl p 2—Expansin
- Secc pollen—Cultivated rye
- Tyr p—Storage mite
- Api g 1, Bet v 1, Cor a 1.0103, rCor a 1.0401, Dau c 1, Fag s 1—PR-10 proteins
- Cyn d 1—Beta expansin
- Can f—1 Lipocalin
- Fel d 1—Uteroglobin
- Der f 1—Cystein proteasa
- Phl p 5, Phl p 6—Grass group 5, 6 Timothy.
- Can f 1, Can f 2, Can f 4, Cav p 1, Fel d 4, Fel d 7—Lipocalins
- Phl p 1—Beta expansin
- Phl p 2—Expansin
- Fel d 1—Uteroglobin
- Der p 21, Der p 5—House dust mites and
- Asp f 6—Mn superoxide dismutase, Aspergillus fumigatus
- Der f 2, Der p 2—NPC2 family, house dust mite
- Fel d 1—Uteroglobin
- Fel d 4, Mus m 1—Lipocalins
- Mala s 5—Malassezia sympodialis
- Rat—extract specific IgE of Rat
- Aca s, Blo t 5, Tyr p—Storage mites
- Pen m 2, Der p 20, Bla g 9—Arginine kinase
- Asp f 6—Mn superoxide dismutase, Aspergillus fumigatus
- Der f 2, Der p 2—NPC2 family, house dust mite
- Gly d 2 and Lep d 2—NPC2 family, storage mite
- Der p 5, Der p 7, Der p21—Molecular components from house dust mites
- Der p 23—Peritrophin-like domain
- Can f Fel d 1-like—Uteroglobin
- Cor a 1.0103, Fag s1—PR-10 proteins
- Phl p 1—Beta expansin
- Phl p 2—Expansin
2.1. Comparison of Mild and Severe Form of AD
2.2. Comparison of Mild and Moderate Forms of AD
2.3. Comparison of Moderate and Severe Forms of AD
3. Discussion
4. Materials and Methods
4.1. Statistical Analysis
4.2. Commentary on Statistical Evaluation
4.3. Patients and Methods
4.4. Dermatological Examination
4.5. Examination of SpecificIgE to Molecular Components
4.6. Bronchial Asthma
4.7. Allergic Rhinitis
4.8. The Evaluation of Duration of Atopic Dermatitis
4.9. The Onset of Atopic Dermatitis
4.10. The Family History of Atopy
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Czarnowicki, T.; Krueger, J.; Guttman-Yassky, E. Novel concepts of prevention and treatment of atopic dermatitis through barrier and immune manipulations with implications for the atopic march. J. Allergy Clin. Immunol. 2017, 139, 1723–1734. [Google Scholar] [CrossRef]
- Paller, A.; Spergel, J.; Mina-Osorio, P.; Irvine, A. The atopic march and atopic multimorbidity: Many trajectories, many pathways. J. Allergy Clin. Immunol. 2019, 143, 46–55. [Google Scholar] [CrossRef] [PubMed]
- Illi, S.; von Mutius, E.; Lau, S.; Nickel, R.; Grüber, C.; Niggemann, B.; Wahn, U. The Multicenter Allergy Study Gro The natural course of atopic dermatitis from birth to age 7 years and the association with asthma. J. Allergy Clin. Immunol. 2004, 113, 925–931. [Google Scholar] [CrossRef] [PubMed]
- Tokura, Y. Evolution of Atopic Dermatitis in the 21st Century; Extrinsic and Intrinsic Atopic Dermatitis; Springer: Singapore, 2018; pp. 181–199. [Google Scholar]
- Burbank, A.J.; Sood, A.K.; Kesic, M.J.; Peden, D.B.; Hernandez, M.L. Environmental determinants of allergy and asthma in early life. J. Allergy Clin. Immunol. 2017, 140, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Saglani, S.; Gregory, L.G.; Manghera, A.K. Inception of early-life allergen-induced airway hyperresponsiveness is reliant on IL-13(+)CD4(+) T cells. Sci. Immunol. 2018, 3, eaan4128. [Google Scholar] [CrossRef] [PubMed]
- Tovey, E.R.; Almqvist, C.; Li, Q.; Crisafulli, D.; Marks, G.B. Nonlinear relationship of mite allergen exposure to mite sensitization and asthma in a birth cohort. J. Allergy Clin. Immunol. 2008, 122, 114–118. [Google Scholar] [CrossRef] [PubMed]
- Zakzuk, J.; Mercado, D.; Bornacelly, A. Hygienic conditions influence sensitization to Blomia tropicalis allergenic components: Results from the FRAAT birth cohort. Pediatr. Allergy Immunol. 2019, 30, 172–178. [Google Scholar] [CrossRef]
- Matricardi, P.M.; Kleine-Tebbe, J.; Hoffmann, H.J.; Valenta, R.; Hilger, C.; Hofmaier, S.; Aalberse, R.C.; Agache, I.; Asero, R.; Ballmer-Weber, B.; et al. EAACI molecular Allergology User’s guide. Pediatr. Allergy Immunol. 2016, 27 (Suppl. S23), 1–250. [Google Scholar] [CrossRef]
- Pulsawat, P.; Soongrung, T.; Satitsuksanoa, P. The house dust mite allergen Der p 5 binds lipid ligands and stimulates airway epithelial cells through a TLR2-dependent pathway. Clin. Exp. Allergy 2019, 49, 378–390. [Google Scholar] [CrossRef]
- Soongrung, T.; Mongkorntanyatip, K.; Peepim, T. The Blomia tropicalis allergen Blot 7 stimulates innate immune signalling pathways through TLR2. Clin. Exp. Allergy 2018, 48, 464–474. [Google Scholar] [CrossRef]
- Satitsuksanoa, P.; Kennedy, M.; Gilis, D. The minor house dust mite allergen Der p 13 is a fatty acid-binding protein and an activator of a TLR2-mediated innate immune response. Allergy 2016, 71, 1425–1434. [Google Scholar] [CrossRef]
- Resch, Y.; Blatt, K.; Malkus, U. Molecular, Structural and Immunological Characterization of Der p 18, a Chitinase-Like House Dust Mite Allergen. PLoS ONE 2016, 11, e0160641. [Google Scholar]
- Zakzuk, J.; Benedetti, I.; Fernandez-Caldas, E.; Caraballo, L. The influence of chitin on the immune response to the house dust mite allergen Blo t 12. Int. Arch. Allergy Immunol. 2014, 163, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Trompette, A.; Divanovic, S.; Visintin, A. Allergenicity resulting from functional mimicry of a Toll-like receptor complex protein. Nature 2009, 457, 585–588. [Google Scholar] [CrossRef]
- Hewitt, C.R.; Horton, H.; Jones, R.M.; Pritchard, D.I. Heterogeneous proteolytic specificity and activity of the house dust mite proteinase allergen Der p, I. Clin. Exp. Allergy 1997, 27, 201–207. [Google Scholar] [CrossRef] [PubMed]
- Anto, J.M.; Bousquet, J.; Akdis, M.; Auffray, C.; Keil, T.; Momas, I.; Postma, D.S.; Valenta, R.; Wickman, M.; Cambon-Thomsen, A.; et al. Mechanisms of the development of allergy (MeDALL): Introducing novel concepts in allergy phenotypes. J. Allergy Clin. Immunol. 2017, 139, 388–399. [Google Scholar] [CrossRef]
- van Hage, M.; Hamsten, C.; Valenta, R. ImmunoCAP assays: Pros and cons in allergology. J. Allergy Clin. Immunol. 2017, 140, 974–977. [Google Scholar] [CrossRef]
- Heffler, E.; Puggioni, F.; Peveri, S.; Montagni, M.; Canonica, G.W.; Melioli, G. Extended IgE profile based on an allergen macroarray: A novel tool for precision medicine in allergy diagnosis. World Allergy Organ. J. 2018, 11, 7. [Google Scholar] [CrossRef]
- Steering Committee Authors; Review Panel Members. A WAO—ARIA—GA2LEN consensus document on molecular-based allergy diagnosis (PAMD@): Update 2020. World Allergy Organ. J. 2020, 13, 100091. [Google Scholar]
- Čelakovská, J.; Bukač, J.; Vaňková, R.; Krcmova, I.; Krejsek, J.; Andrýs, C. ISAC multiplex testing—results of examination in 100 patients suffering from atopic dermatitis. Food Agric. Immunol. 2020, 31, 1014–1035. [Google Scholar] [CrossRef]
- Čelakovská, J.; Bukač, J.; Vaňková, R.; Krcmova, I.; Krejsek, J.; Andrýs, C. Cluster analysis of molecular components in 100 patients suffering from atopic dermatitis according to the ISAC Multiplex testing. Food Agric. Immunol. 2020, 31, 827–848. [Google Scholar] [CrossRef]
- Di Fraia, M.; Arasi, S.; Castelli, S. A new molecular multiplex IgE assay for the diagnosis of pollen allergy in Mediterranean countries: A validation study. Clin. Exp. Allergy 2019, 49, 341–349. [Google Scholar] [CrossRef] [PubMed]
- Melioli, G.; Bonifazi, F.; Bonini, S. The ImmunoCAP ISAC molecular allergology approach in adult multi-sensitized Italian patients with respiratory symptoms. Clin. Biochem. 2011, 44, 1005–1011. [Google Scholar] [CrossRef] [PubMed]
- Weghofer, M.; Grote, M.; Resch, Y.; Casset, A.; Kneidinger, M.; Kopec, J.; Thomas, W.R.; Fernández-Caldas, E.; Kabesch, M.; Ferrara, R.; et al. Identification of Der p 23, a peritrophin-like protein, as a new major Dermatophagoides pteronyssinus allergen associated with the peritrophic matrix of mite fecal pellets. J. Immunol. 2013, 190, 3059–3067. [Google Scholar] [CrossRef]
- Flöistrup, H.; Swartz, J.; Bergström, A.; Alm, J.S.; Scheynius, A.; van Hage, M.; Waser, M.; Braun-Fahrländer, C.; Schram-Bijkerk, D.; Huber, M.; et al. Allergic disease and sensitization in Steiner school children. J. Allergy Clin. Immunol. 2006, 117, 59–66. [Google Scholar] [CrossRef]
- Bousquet, J.; Lockey, R.; Malling, H.J.; Alvarez-Cuesta, E.; Canonica, G.W.; Chapman, M.D.; Creticos, P.J.; Dayer, J.M.; Durham, S.R.; Demoly, P.; et al. Allergen immunotherapy: Therapeutic vaccines for allergic diseases—World Health Organization. American academy of Allergy, Asthma and Immunology. Ann. Allergy Asthma Immunol. 1998, 81, 401–405. [Google Scholar] [CrossRef]
- Carvalho Kdos, A.; de Melo-Neto, O.P.; Magalhães, F.B.; Ponte, J.C.; Felipe, F.A.; dos Santos, M.C.; dos Santos Lima, G.; Cruz, Á.A.; Pinheiro, C.S.; Pontes-de-Carvalho, L.C.; et al. Blomia tropicalis Blo t 5 and Blo t 21 recombinant allergens might confer higher specificity to serodiagnostic assays than whole mite extract. BMC Immunol. 2013, 27, 11. [Google Scholar] [CrossRef]
- Pytelková, J.; Lepšík, M.; Sanda, M.; Talacko, P.; Marešová, L.; Mareš, M. Enzymatic activity and immunoreactivity of Aca s 4, an alpha-amylase allergen from the storage mite Acarus siro. BMC Biochem. 2012, 13, 3. [Google Scholar] [CrossRef]
- Banerjee, S.; Resch, Y.; Chen, K.; Swoboda, I.; Focke-Tejkl, M.; Blatt, K.; Novak, N.; Wickman, M.; van Hage, M.; Ferrara, R.; et al. Der p 11 is a major allergen for house dust miteallergic patients suffering from atopic dermatitis. J. Investig. Dermatol. 2015, 135, 102–109. [Google Scholar] [CrossRef]
- Holzweber, F.; Svehla, E.; Fellner, W. Inhibition of IgE binding to cross-reactive carbohydrate determinants enhances diagnostic selectivity. Allergy 2013, 68, 1269–1277. [Google Scholar] [CrossRef]
- Biedermann, T.; Winther, L.; Till, S.J.; Panzner, P.; Knulst, A.; Valovirta, E. Birch pollen allergy in Europe. Allergy 2019, 74, 1237–1248. [Google Scholar] [CrossRef]
- Panzner, P.; Vachová, M.; Vítovcová, P.; Brodská, P.; Vlas, T. A comprehensive analysis of middle-european molecular sensitization profiles to pollen allergens. Int. Arch. Allergy Immunol. 2014, 164, 74–82. [Google Scholar] [CrossRef]
- Panzner, P.; Vachová, M.; Vlas, T.; Vítovcová, P.; Brodská, P.; Malý, M. Cross-sectional study on sensitization to mite and cockroach allergen components in allergy patients in the Central European region. Clin. Transl. Allergy 2018, 8, 1–9. [Google Scholar] [CrossRef]
- Vachová, M.; Panzner, P.; Vlas, T.; Vítovcová, P. Analysis of Sensitization Profiles in Central European Allergy Patients Focused on Animal Allergen Molecules. Int. Arch. Allergy Immunol. 2020, 181, 278–284. [Google Scholar] [CrossRef]
- Cabanas, R.; Lopez-Serrano, M.C.; Carreira, J. Importance of albumin in cross-reactivity among cat, dog and horse allergens. J. Investig. Allergol. Clin. Immunol. 2000, 10, 71–77. [Google Scholar]
- Basagana, M.; Bartolome, B.; Pastor, C. Allergy to human seminal fluid: Cross-reactivity with dog dander. J. Allergy Clin. Immunol. 2008, 121, 233–239. [Google Scholar] [CrossRef]
- Khurana, T.; Newman-Lindsay, S.; Young, P.R.; Slater, J.E. The NPC2 protein: A novel dog allergen. Ann. Allergy Asthma Immunol. 2016, 116, 440–446.e442. [Google Scholar] [CrossRef]
- Konradsen, J.R.; Fujisawa, T.; van Hage, M. Allergy to furry animals: New insights, diagnostic approaches, and challenges. J. Allergy Clin. Immunol. 2015, 135, 616–625. [Google Scholar] [CrossRef]
- Asarnoj, A.; Hamsten, C.; Waden, K. Sensitization to cat and dog allergen molecules in childhood and prediction of symptoms of cat and dog allergy in adolescence: A BAMSE/MeDALL study. J. Allergy Clin. Immunol. 2016, 137, 813–821.e817. [Google Scholar] [CrossRef]
- Bjerg, A.; Winberg, A.; Berthold, M.; Mattsson, L.; Borres, M.P.; Ronmark, E. A population-based study of animal component sensitization, asthma, and rhinitis in schoolchildren. Pediatr. Allergy Immunol. 2015, 26, 557–563. [Google Scholar] [CrossRef]
- Mittermann, I.; Wikberg, G.; Johansson, C.; Lupinek, C.; Lundeberg, L.; Crameri, R.; Valenta, R.; Scheynius, A. IgE sensitization profiles differ between adult patients with severe and moderate atopic dermatitis. PLoS ONE 2016, 11, e0156077. [Google Scholar] [CrossRef] [PubMed]
- Röckmann, H.; van Geel, M.; Knulst, A.C.; Huiskes, J.; Bruijnzeel-Koomen, C.; de Bruin-Weller, M. Food allergen sensitization pattern in adults in relation to severity of atopic dermatitis. Clin. Transl. Allergy 2014, 4, 9. [Google Scholar] [CrossRef]
- Hanifin, J.; Rajka, G. Diagnostic features of atopic dermatitis. Acta Derm. Venereol. 1980, 92, 44–47. [Google Scholar]
- European Task Force on Atopic Dermatitis. Severity scoring of atopic dermatitis: The SCORAD Index (consensus report of the European Task Force on Atopic Dermatitis). Dermatology 1993, 186, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Jakob, T.; Forstenlechner, P.; Matricardi, P.; Kleine-Tebbe, J. Molecular allergy diagnostics using multiplex assays: Methodological and practical considerations for use in research and clinical routine. Allergol. J. Int. 2015, 24, 320–332. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.; Roh, J.; Lee, J. Clinical Availability of Component-Resolved Diagnosis Using Microarray Technology in Atopic Dermatitis. Ann. Dermatol. 2014, 26, 437–446. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Du, K.; She, W.; Ouyang, Y.; Sima, Y.; Liu, C.; Zhang, L. Recent advances in the diagnosis of allergic rhinitis. Expert Rev. Clin. Immunol. 2018, 14, 957–964. [Google Scholar] [CrossRef]
| Characteristic of Patients | Number of Patients |
|---|---|
| Patients suffering from atopic dermatitis | 100 patients |
| Sex | 48 men, 52 women |
| The average age | 40.9 years (min. age 14 years, max. age 67 years) |
| The average SCORAD | 39 points, s.d. 13.1 points |
| Severity of AD | Mild form 14 (14%) |
| Moderate form 61 (61%) | |
| Severe form 25 (25%) | |
| Family history about atopy | Positive family history 48 patients (48%) |
| Negative family history 52 patients (52%) | |
| Onset of AD | Under 5 years of age 61 patients (61%) |
| After 5 year of age 39 patients (%) | |
| Eczematic lesions | Persistent 57 patients (57%) |
| Occasional 43 patients (43%) | |
| Bronchial asthma | 55 (55%) |
| Allergic rhinitis | 74 (74%) |
| Allergen Reagents (Allergenic Extracts and Molecular Components) | Sensitization Confirmed in Patients (%) |
|---|---|
| Phl p 1 (Beta expansin, Timothy grass) | 57.0 |
| Bet v 1 (PR-10 protein, birch), Lol p 1 (Beta expansin, rye grass), Sec c_pollen (cultivated rye, pollen) | 53.0 |
| Fag s1 (PR-10 protein, European beech) | 49.0 |
| Cor a 1.0103 (PR-10 protein, hazel pollen) Cyn d 1 (Beta expansin, Bermuda grass) | 48.0 |
| Cor a_pollen (hazel pollen) | 47.0 |
| Der f 2 (NPC2 family, house dust mite) Fra a 1 + 3 (PR-10 protein + non-specific lipid transfer protein, type 1, strawberry) Phl p 2 (expansin, Timothy grass) | 45.0 |
| Cor a 1.0401 (PR-10 protein, hazel pollen) Cyn d (Bermuda grass) Der p 2 (NPC2 family, house dust mite) Fel d 1 (uteroglobin, cat) | 44.0 |
| Aln g 1 (PR-10 protein, alder) | 43.0 |
| Pas n (bahia grass) Phl p 5.0101 (grass group 5/6, Timothy grass), Phl p 6 (grass group 5/6, Timothy grass) | 42.0 |
| Lep d 2 (NPC2 family, storage mite) Mal d 1 (PR-10 protein, apple) | 41.0 |
| Can f 1 (lipocalin, dog) Der p 23 (peritrophin-like protein domain, house dust mite) | 36.0 |
| Gly m 4 (PR-10 protein, soybean) | 35.0 |
| Der p 1 (cysteine protease, house dust mite) | 34.0 |
| Ara h 8 (PR-10 protein, peanut) Gly d 2 (NPC2 family, storage mite) | 33.0 |
| Art v (mugwort), Can f 6 (lipocalin, dog) | 32.0 |
| Der f 1 (cysteine protease, house dust mite) Fel d 7 (lipocalin, cat) | 31.0 |
| Can f_male urine (male dog urine) | 30.0 |
| Can f 4 (lipocalin, dog) Equ c 1 (lipocalin, horse) | 29.0 |
| Pla l (English plantain) | 28.0 |
| Ory c 3 (uteroglobin, rabbit) Phr c (common reed) | 27.0 |
| Aca s (storage mite) Alt a 1 (unknown, Alternaria alternata) Der p 5 (unknown, house dust mite) Fel d 4 (lipocalin, cat) | 26.0 |
| Amb a (ragweed) Tyr p (storage mite) | 25.0 |
| Ach d (house cricket) Der p 7 (mites group 7, bactericidal permeability-increasing-like protein, house dust mite) Loc m (migratory locust) Mala s 11 (Mn superoxide dismutase, Malassezia sympodialis) | 24.0 |
| Amb a 4 (plant defensin, ragweed) Der p 20 (arginine kinase, house dust mite), Ten m (mealworm) | 23.0 23.0 |
| Api g 1 (PR-10 protein, celery), Mus m 1 (lipocalin and urinary prealbumin, house mouse) | 22.0 |
| Bla g 9 (arginine kinase, German cockroach) Can f_Fd1 (uteroglobin, dog), Cav p 1 (lipocalin, guinea pig) | 21.0 |
| Asp f 6 (Mn superoxide dismutase, Aspergillus fumigatus) Bla g 4 (calycin, lipocalin, German cockroach) Fra e 1 (Ole e 1-like protein family, European ash), Jug r_pollen (walnut pollen) | 20.0 |
| Art v 1 (plant defensin, mugwort) Dau c (carrot) Dau c 1 (PR-10 protein, carrot) | 19.0 |
| Par j 2 (non-specific lipid transfer protein, type 1, pellitory of the wall) Pen m 2 (arginine kinase, shrimp) Ves v 5 (antigen 5, yellow jacket venom) | 18.0 |
| Ama r (redroot pigweed) Can f 2 (lipocalin, dog) Rat n (rat) | 17.0 |
| Api m (honey bee venom) Cup a 1 (pectate lyase, cypress) Fra e (European ash), Gal d_white (egg white) Per a (American cockroach) | 16.0 |
| Aca m (acacia) Api m 1 (phospholipase A2, honey bee venom) Asp f 3 (peroxysomal protein, Aspergillus fumigatus) Cla h 8 (mannitol dehydrogenase, cladosporium herbarum) Cry j 1 (pectate lyase, Japanese cedar) Der p 21 (unknown, house dust mite) Hel a (sunflower seed) Hom g (lobster) Pla l 1 (Ole e 1-like protein family, English plantain) Sal k (Russian thistle, saltwort) | 15.0 |
| Act d 2 (thaumatin-like protein, kiwi fruit) Mala s 6 (cyclophilin, malassezia sympodialis) Sac c (Saccharomyces cerevisiae) Sol t (potato) | 14.0 |
| Api m 10 (icarapin variant 2, honey bee venom) Pha v (green bean, French bean) | 13.0 |
| Alt a 6 (enolase, Alternaria alternata) Amb a 1 (pectate lyase, ragweed), Api g 2 (non-specific lipid transfer protein, type 1, celery) Blo t 5 (mites group 5, storage mite) Can s (hemp), Hor v (barley) Lol spp. (squid) Pan b (Northern shrimp), Phl p 7 (polcalcin, Timothy grass) Phod s 1 (lipocalin, Siberian hamster) Sal k 1 (pectin methylesterase, Russian thistle, saltwort) Urt d (nettle) | 12.0 |
| Bet v 2 (profilin, birch) Cuc m 2 (profilin, muskmelon) Pec spp. (scallop) Sec c_flour (cultivated rye) Ses i 1 (2S albumin, sesame) Tyr p 2 (NPC2 family, storage mite) Ves v (yellow jacket venom), Ves v 1 (phospholipase A1, yellow jacket venom) Zea m 14 (non-specific lipid transfer protein, type 1, maize) | 11.0 |
| Che q (quinoa) Mala s 5 (unknown, Malassezia sympodialis) Ole e 1 (Ole e 1-family, olive) Pers a (avocado) Phl p 12 (profilin, Timothy grass) Pla a 2 (polygalacturonase, London plane tree) Pol d (paper wasp venom) Pyr c (pear) Sola l (tomato) Ulm c (elm) Zea m (maize) | 10.0 |
| The Number of Patients According to the Level of Specific IgE in Classes 0–4 in 75 Selected Allergen Reagents, (100 Patients Included in the Study = 100%) | ||||||
|---|---|---|---|---|---|---|
| Allergen Reagents (Allergen Extract, Molecular Components) | Class 0 | Class 1 | Class 2 | Class 3 | Class 4 | Class 3 + Class 4 |
| Acas | 74 | 5 | 7 | 8 | 6 | 14 |
| Achd | 76 | 6 | 3 | 7 | 8 | 15 |
| Alng1 | 57 | 0 | 12 | 14 | 17 | 31 |
| Alta1 | 74 | 0 | 0 | 4 | 22 | 26 |
| Alta6 | 88 | 2 | 4 | 4 | 2 | 6 |
| Amba | 75 | 5 | 8 | 8 | 4 | 12 |
| Amba4 | 77 | 7 | 5 | 8 | 3 | 11 |
| Apig1 | 78 | 5 | 3 | 8 | 6 | 14 |
| Apim10 | 87 | 6 | 1 | 6 | 0 | 6 |
| Arah8 | 67 | 8 | 10 | 8 | 7 | 15 |
| Artv | 68 | 5 | 7 | 10 | 10 | 20 |
| Artv1 | 81 | 2 | 7 | 9 | 1 | 10 |
| Aspf6 | 80 | 2 | 4 | 9 | 5 | 14 |
| Betv1 | 47 | 2 | 6 | 5 | 40 | 45 |
| Blag9 | 79 | 0 | 1 | 2 | 18 | 20 |
| Blot5 | 88 | 3 | 3 | 3 | 3 | 6 |
| Canf_Fd1 | 79 | 6 | 9 | 6 | 0 | 6 |
| Canf_maleurine | 70 | 2 | 8 | 7 | 13 | 20 |
| Canf1 | 64 | 1 | 8 | 10 | 17 | 27 |
| Canf2 | 83 | 1 | 2 | 4 | 10 | 14 |
| Canf4 | 71 | 1 | 6 | 5 | 17 | 22 |
| Canf6 | 68 | 0 | 6 | 5 | 21 | 26 |
| Cavp1 | 79 | 1 | 4 | 7 | 9 | 16 |
| Cora_pollen | 53 | 4 | 7 | 19 | 17 | 36 |
| Cora1.0103 | 52 | 2 | 7 | 7 | 32 | 39 |
| Cora1.0401 | 56 | 0 | 14 | 10 | 20 | 30 |
| Cynd | 56 | 12 | 16 | 14 | 2 | 16 |
| Cynd1 | 52 | 8 | 16 | 16 | 8 | 24 |
| Dauc | 81 | 6 | 3 | 7 | 3 | 10 |
| Dauc1 | 81 | 5 | 2 | 6 | 6 | 12 |
| Derf1 | 69 | 3 | 5 | 12 | 11 | 23 |
| Derf2 | 55 | 1 | 2 | 2 | 40 | 42 |
| Derp1 | 66 | 2 | 7 | 7 | 18 | 25 |
| Derp2 | 56 | 0 | 2 | 2 | 40 | 42 |
| Derp20 | 77 | 2 | 2 | 1 | 18 | 19 |
| Derp21 | 85 | 0 | 1 | 2 | 12 | 14 |
| Derp23 | 64 | 1 | 7 | 5 | 23 | 28 |
| Derp5 | 74 | 8 | 3 | 1 | 14 | 15 |
| Derp7 | 76 | 6 | 12 | 3 | 3 | 6 |
| Equc1 | 71 | 2 | 8 | 1 | 18 | 19 |
| Fags1 | 51 | 3 | 6 | 15 | 25 | 40 |
| Feld1 | 56 | 4 | 6 | 2 | 32 | 34 |
| Feld4 | 74 | 3 | 8 | 6 | 9 | 15 |
| Feld7 | 69 | 5 | 10 | 3 | 13 | 16 |
| Fraa13 | 55 | 6 | 13 | 12 | 14 | 26 |
| Frae | 84 | 2 | 4 | 2 | 8 | 10 |
| Frae1 | 80 | 4 | 5 | 2 | 9 | 11 |
| Glyd2 | 67 | 9 | 8 | 4 | 12 | 16 |
| Glym4 | 65 | 7 | 12 | 8 | 8 | 16 |
| Homg | 85 | 2 | 7 | 2 | 4 | 6 |
| Chispp_ | 91 | 1 | 2 | 3 | 3 | 6 |
| Lepd2 | 59 | 4 | 9 | 18 | 10 | 28 |
| Locm | 76 | 8 | 6 | 4 | 6 | 10 |
| Lolp1 | 47 | 7 | 6 | 10 | 30 | 40 |
| Malas11 | 76 | 0 | 7 | 1 | 16 | 17 |
| Malas5 | 90 | 1 | 3 | 1 | 5 | 6 |
| Mald1 | 59 | 1 | 17 | 10 | 13 | 23 |
| Musm1 | 78 | 1 | 6 | 7 | 8 | 15 |
| Olee1 | 90 | 0 | 3 | 2 | 5 | 7 |
| Oryc3 | 73 | 1 | 2 | 10 | 14 | 24 |
| Panb | 88 | 0 | 5 | 5 | 2 | 7 |
| Pasn | 58 | 9 | 19 | 12 | 2 | 14 |
| Penm2 | 82 | 0 | 6 | 5 | 7 | 12 |
| Phlp1 | 43 | 5 | 9 | 4 | 39 | 43 |
| Phlp2 | 55 | 4 | 9 | 7 | 25 | 32 |
| Phlp5_0101 | 58 | 0 | 5 | 5 | 32 | 37 |
| Phlp6 | 58 | 7 | 6 | 8 | 21 | 29 |
| Phods1 | 88 | 3 | 2 | 2 | 5 | 7 |
| Plal | 72 | 14 | 2 | 2 | 10 | 12 |
| Plal1 | 85 | 3 | 0 | 2 | 10 | 12 |
| Ratn | 83 | 3 | 6 | 6 | 2 | 8 |
| Secc_pollen | 47 | 5 | 17 | 17 | 14 | 31 |
| Tenm | 77 | 7 | 6 | 6 | 4 | 10 |
| tIgE | 18 | 0 | 0 | 0 | 82 | 82 |
| Tyrp | 75 | 5 | 7 | 8 | 5 | 13 |
| Allergen Reagents, Allergenic Extracts and Molecular Components (Biochemical Name). r = Recombinant, n= Natural | Number of Patients | Severity of AD | Bronchial Asthma | Allergic Rhinitis | Onset of AD | Family History | Duration of Lesion | |
|---|---|---|---|---|---|---|---|---|
| Aca s | Storage mite | 14 | 0.0437 | - | - | - | - | 0.00316 |
| Ach d | House cricket | 15 | - | - | - | - | - | 0.00164 |
| rAln g 1 | PR-10 protein, alder | 31 | - | - | - | - | - | - |
| rAlt a 1 | Unknown, Alternaria alternata | 26 | - | - | - | - | - | - |
| rAlt a 6 | Enolase, Alternaria alternata | 6 | - | 0.0312 | - | - | - | - |
| Amb a | Ragweed | 12 | - | - | - | - | - | - |
| rAmb a 4 | Plant defensin, ragweed | 11 | - | - | - | - | - | - |
| rApi g 1 | PR-10 protein, celery | 14 | - | - | 0.0182 | - | - | - |
| rApi m 10 | Icarapin variant 2, honey bee venom | 6 | - | - | - | - | - | |
| nAra h 1 | Cupin (vicillin-type, 7S globulin), peanut | 5 | 0.0349 | - | - | - | - | - |
| nAra h 6 | 2S albumin, peanut | 5 | 0.0349 | - | - | - | - | - |
| rAra h 8 | PR-10 protein, peanut | 15 | - | - | - | - | - | - |
| Art v | Mugwort | 20 | - | - | - | - | - | - |
| rArt v 1 | Plant defensin, mugwort | 10 | - | - | - | - | - | - |
| rAsp f 6 | Mn superoxide dismutase, Aspergillus fumigatus | 14 | 0.00145 | - | - | 0.00784 | - | 0.0214 |
| rBet v 1 | PR-10 protein, birch | 45 | - | - | 0.009 | - | - | - |
| rBla g 9 | Arginine kinase, German cockroach | 20 | 0.00439 | - | - | - | - | 0.0008 |
| rBlo t 5 | Blomia tropicalis, storage mite | 6 | 0.023 | - | - | - | - | 0.0356 |
| rCan f 1 | Lipocalin, dog | 27 | 0.0235 | - | 0.01 | 0.00262 | - | - |
| rCan f 2 | Lipocalin, dog | 14 | 0.0437 | - | - | 0.00784 | - | - |
| rCan f 4 | Lipocalin, dog | 22 | - | - | - | 0.00623 | - | - |
| rCan f 6 | Lipocalin, dog | 26 | 0.00907 | - | - | - | - | - |
| rCanf_Fel d 1-like | Uteroglobin, dog | 6 | - | - | - | - | - | 0.0356 |
| Can f_male urine | Male dog urine | 20 | - | - | - | - | - | - |
| rCav p 1 | Lipocalin, guinea pig | 16 | - | - | 0.0238 | |||
| rCor a 1.0103 | PR-10 protein, hazel pollen | 39 | 0.0163 | - | 0.0163 | - | - | 0.0482 |
| rCor a 1.0401 | PR-10 protein, hazelnut | 30 | 0.00819 | - | - | - | - | - |
| Cor a_pollen | Hazel pollen | 36 | 0.00728 | - | 0.0164 | - | - | 0.0211 |
| rCra c 6 | Troponin C, North sea shrimp | 5 | 0.0349 | - | - | - | - | - |
| Cyn d | Bermuda grass | 16 | 0.0288 | - | - | - | - | - |
| rCyn d 1 | Beta expansin, Bermuda grass | 24 | 0.00168 | - | 0.0312 | - | - | |
| Dau c | Carrot | 10 | - | - | - | - | - | 0.0402 |
| rDau c 1 | PR-10 protein, carrot | 12 | - | - | 0.0329 | - | - | - |
| rDer f 1 | Cysteine protease, house dust mite | 23 | - | - | 0.0327 | - | - | - |
| rDer f 2 | NPC2 family, house dust mite | 42 | 0.046 | 0.0497 | 0.0384 | |||
| rDer p 1 | Cysteine protease, house dust mite | 25 | - | 0.0485 | - | - | - | - |
| rDer p 2 | NPC2 family, house dust mite | 42 | 0.046 | - | - | 0.0497 | 0.0384 | |
| rDer p 20 | Arginine kinase, house dust mite | 19 | 0.00224 | - | - | - | - | 0.00017 |
| rDer p 21 | Unknown, house dust mite | 14 | 0.0437 | 0.0185 | - | 0.00784 | - | 0.00316 |
| rDer p 23 | Peritrophin-like protein domain, house dust mite | 28 | 0.00785 | 0.0122 | - | - | - | 0.00658 |
| rDer p 5 | Unknown, house dust mite | 15 | - | 0.0482 | - | 0.042 | - | 0.00164 |
| rDer p 7 | Bactericidal permeability-increasing-like protein, house dust mite | 6 | - | 0.0312 | - | - | - | 0.0356 |
| rEqu c 1 | Lipocalin, horse | 19 | - | - | - | - | - | - |
| rFag s 1 | PR-10 protein, European beech | 40 | 0.017 | - | 0.0189 | - | - | 0.032 |
| rFel d 1 | Uteroglobin, cat | 34 | 0.0319 | 0.0291 | 0.00168 | 0.00117 | 0.00476 | |
| rFel d 4 | Lipocalin, cat | 15 | - | - | - | 0.042 | 0.00713 | - |
| rFel d 7 | Lipocalin, cat | 16 | - | - | - | 0.00017 | - | - |
| rFra a 1 + 3 | PR-10 protein+ non-specific lipid transfer protein type 1, strawberry | 26 | - | - | - | - | - | - |
| Fra e | European ash | 10 | - | - | - | - | - | - |
| rFra e 1 | Ole e 1-like protein family, European ash | 11 | - | - | - | - | - | - |
| rGly d 2 | NPC2 family, storage mite | 16 | 0.0455 | - | - | - | - | 0.00085 |
| rGly m 4 | PR-10 protein, soybean | 16 | - | - | - | - | - | - |
| Hom g | Lobster | 6 | 0.0337 | - | - | - | - | - |
| Chi spp. | Crab | 6 | - | - | - | - | - | - |
| rLep d 2 | NPC2 family, storage mite | 28 | 0.0168 | 0.0462 | - | - | - | 0.00154 |
| Loc m | Migratory locust | 10 | - | 0.0402 | - | - | - | 0.0402 |
| nLol p 1 | Beta-espansin, Rye grass | 40 | 0.0132 | 0.0406 | 0.0106 | |||
| Lol spp. | Squid | 5 | - | - | - | - | - | - |
| rMal d 1 | PR-10 protein, apple | 23 | 0.0249 | - | - | - | - | - |
| rMala s 11 | Mn superoxide dismutase, Malassezia sympodialis | 17 | 0.00855 | - | - | - | - | - |
| rMala s 5 | Unknown, Malassezia sympodialis | 6 | - | - | - | - | 0.0103 | - |
| nMus m 1 | Lipocalin and urinary prealbumin, house mouse | 15 | - | - | - | - | 0.00147 | - |
| nOle e 1 | Ole e 1-family, olive | 7 | - | - | - | - | - | 0.0185 |
| rOry c 1 | Lipocalin, rabbit | 5 | - | - | - | - | - | - |
| rOry c 3 | Uteroglobin, rabbit | 24 | 0.0431 | - | - | - | - | 0.041 |
| Pan b | Northern shrimp | 7 | - | - | - | - | - | - |
| Pas n | Bahia grass | 14 | 0.0437 | - | - | - | - | - |
| rPen m 2 | Arginine kinase, shrimp | 12 | 0.0264 | - | - | - | - | 0.001 |
| rPer a 7 | Tropomyosin, American cockroach | 5 | - | - | - | - | - | - |
| rPhl p 1 | Beta expansin, Timothy | 43 | 0.0273 | 0.0298 | - | 0.0482 | - | 0.0251 |
| rPhl p 2 | Expansin, Timothy | 32 | 0.0164 | 0.02 | - | 0.049 | - | 0.0126 |
| rPhl p 5.0101 | Grass group 5/6, Timothy | 37 | - | - | 0.0291 | - | - | 0.04 |
| rPhl p 6 | Grass group 5/6, Timothy | 29 | - | - | 0.0248 | - | - | - |
| rPhod s 1 | Lipocalin, Siberian hamster | 7 | - | - | - | - | - | - |
| Pla l | English plantain | 12 | - | - | - | - | - | - |
| rPla l 1 | Ole e 1-like protein family, English plantain | 12 | - | - | - | - | - | - |
| Rat n | Rat | 8 | - | - | - | - | 0.0267 | - |
| Sec c_pollen | Cultivated rye, pollen | 31 | 0.0118 | 0.00971 | - | - | - | - |
| Ten m | Mealworm | 10 | - | - | - | - | - | 0.0402 |
| Tyr p | Storage mite | 13 | - | 0.0338 | - | - | - | 0.00604 |
| rXip g 1 | Beta-parvalbumin, swordfish | 5 | - | - | - | - | - | - |
| Comparison of Column Proportions | ||||
|---|---|---|---|---|
| Allegen Reagents | The Level of Specific IgE in Classes 0–4 | Mild Form of AD (A) | Moderate Form of AD (B) | Severe Form of AD (C) |
| Acam | 0 | .a | C(0.023) | |
| 1 | .a | B(0.022) | ||
| Acas | 0 | C(0.030) | C(0.004) | |
| Allc | 0 | .a | C(0.035) | |
| Alta6 | 0 | .a | ||
| 3 | .a | B(0.038) | ||
| Amar | 0 | .a | C(0.016) | |
| 1 | .a | |||
| 2 | .a | B(0.010) | ||
| Amba | 0 | |||
| 4 | .a | B(0.038) | ||
| Apig1 | 0 | C(0.047) | C(0.047) | |
| Arah1 | 0 | .a | C(0.009) | |
| 4 | .a | B(0.038) | ||
| Arah2 | 0 | .a | C(0.038) | |
| Arah3 | 0 | .a | C(0.038) | |
| Aspf1 | 0 | .a | C(0.038) | |
| Aspf6 | 0 | .a | C(0.019) | |
| 3 | .a | B(0.009) | ||
| Betv1 | 0 | |||
| 2 | B(0.040) | .a | ||
| 4 | A(0.017) | |||
| Betv2 | 0 | .a | C(0.046) | |
| Blag4 | 0 | .a | C(0.019) | |
| 1 | .a | |||
| 2 | .a | B(0.009) | ||
| Blag9 | 0 | C(0.025) | C(0.011) | |
| 2 | .a | .a | ||
| 3 | .a | .a | ||
| 4 | A(0.025) B(0.001) | |||
| Blot21 | 0 | .a | C(0.010) | |
| Blot5 | 0 | C(0.021) | ||
| Canf1 | 0 | B(0.032) C(0.032) | ||
| Canf2 | 0 | .a | ||
| 3 | .a | B(0.038) | ||
| Canf6 | 0 | C(0.015) | ||
| 4 | A(0.043) B(0.043) | |||
| Capa | 0 | .a | C(0.029) | |
| Carp | 0 | .a | C(0.038) | |
| Clah | 0 | .a | C(0.002) | |
| 1 | .a | B(0.010) | ||
| Clah8 | 0 | .a | C(0.023) | |
| 4 | A(0.015) | |||
| Crac6 | 0 | C(0.007) | ||
| Cucm2 | 0 | .a | C(0.046) | |
| Cupa1 | 0 | .a | C(0.041) | |
| 1 | .a | B(0.035) | ||
| Cynd | 0 | C(0.008) | ||
| Cynd1 | 0 | C(0.016) | ||
| 4 | .a | B(0.029) | ||
| 1 | .a | B(0.035) | ||
| Derf2 | 0 | C(0.032) | ||
| Derp20 | 0 | C(0.014) | C(0.007) | |
| 4 | A(0.025) B(0.001) | |||
| Derp21 | 0 | |||
| 4 | .a | B(0.016) | ||
| Derp23 | 0 | |||
| 4 | .a | B(0.021) | ||
| Dolspp | 0 | .a | C(0.038) | |
| Equc1 | 0 | C(0.015) | C(0.021) | |
| Equc3 | 0 | .a | C(0.038) | |
| Fags1 | 0 | |||
| 2 | B(0.040) | .a | ||
| Feld4 | 0 | C(0.008) | C(0.008) | |
| 4 | B(0.008) | |||
| Fraa1 + 3 | 0 | C(0.032) | ||
| Frae | 0 | C(0.011) | ||
| Frae1 | 0 | C(0.016) | ||
| Gadm | 0 | .a | C(0.038) | |
| Gald1 | 0 | .a | C(0.002) | |
| Gald2 | 0 | .a | C(0.001) | |
| 2 | .a | B(0.038) | ||
| Gald3 | 0 | .a | C(0.010) | |
| 2 | .a | B(0.038) | ||
| Gald5 | 0 | .a | C(0.010) | |
| Gald_white | 0 | .a | C(0.008) | |
| Gald_yolk | 0 | .a | C(0.002) | |
| 1 | .a | B(0.038) | ||
| Glyd2 | 0 | |||
| 4 | B(.021) | |||
| Glym4 | 0 | |||
| 4 | .a | B(.029) | ||
| Hela | 0 | .a | C(.004) | |
| 1 | .a | B(.001) | ||
| Hevb8 | 0 | .a | C(.003) | |
| 4 | .a | B(.038) | ||
| Horv | 0 | .a | C(.016) | |
| Chea | 0 | .a | C(.002) | |
| Cheq | 0 | .a | ||
| 2 | .a | B(0.038) | ||
| Jugr_pollen | 0 | .a | C(0.004) | |
| 1 | .a | |||
| 2 | .a | B(.002) | ||
| Lepd2 | 0 | C(0.017) | ||
| Lolp1 | 0 | |||
| 4 | A(0.015) B(0.033) | |||
| Lupa | 0 | .a | C(0.002) | |
| 2 | .a | B(.038) | ||
| 3 | .a | .a | ||
| Mald1 | 0 | C(0.017) | ||
| Malas11 | 0 | .a | C(0.001) | |
| 4 | .a | B(0.008) | ||
| Malas6 | 0 | .a | C(0.011) | |
| 1 | .a | B(0.010) | ||
| Mani | 0 | .a | C(0.038) | |
| Mera1 | 0 | .a | C(0.035) | |
| Musa | 0 | .a | C(0.010) | |
| 1 | .a | B(0.035) | ||
| Oryc2 | 0 | .a | C(0.035) | |
| Parj2 | 0 | .a | ||
| 1 | .a | |||
| 2 | .a | B(0.029) | ||
| Pecspp_ | 0 | |||
| 1 | ||||
| 2 | .a | B(0.038) | ||
| Penm2 | 0 | C(0.043) | C(0.008) | |
| Pera | 0 | C(0.034) | ||
| Pera7 | 0 | C(0.030) | ||
| Persa | 0 | .a | C(0.002) | |
| 2 | .a | B(0.010) | ||
| Phav | 0 | .a | ||
| 2 | .a | B(0.010) | ||
| Phlp1 | 0 | |||
| 4 | A(0.044) B(0.044) | |||
| Phlp12 | 0 | .a | C(0.022) | |
| Phlp2 | 0 | |||
| 4 | B(0.037) | |||
| Phlp5_0101 | 0 | |||
| 4 | A(0.028) | |||
| Phod2 | 0 | .a | C(0.003) | |
| 3 | .a | B(0.038) | ||
| Pima | 0 | .a | C(0.010) | |
| 1 | .a | B(0.035) | ||
| Pyrc | 0 | .a | ||
| 2 | .a | B(0.038) | ||
| Rajc | 0 | .a | .a | .a |
| Rudspp_ | 0 | C(0.030) | ||
| Sacc | 0 | .a | C(0.011) | |
| 1 | .a | |||
| Salk1 | 0 | .a | C(0.016) | |
| Scos | 0 | .a | C(0.035) | |
| Secc_flour | 0 | .a | C(0.001) | |
| 1 | .a | B(0.010) | ||
| Sesi | 0 | .a | C(0.035) | |
| Sesi1 | 0 | .a | C(0.046) | |
| Solt | 0 | .a | C(0.011) | |
| Trifo | 0 | .a | C(0.035) | |
| Tyrp | 0 | C(0.030) | C(0.002) | |
| Tyrp2 | 0 | |||
| 2 | B(0.029) | |||
| Urtd | 0 | .a | C(0.002) | |
| 1 | .a | B(0.001) | ||
| Vesv | 0 | .a | C(0.046) | |
| 1 | .a | B(0.029) | ||
| 3 | .a | |||
| tIgE | 0 | B(0.001) | .a | |
| 4 | A(0.001) | .a | ||
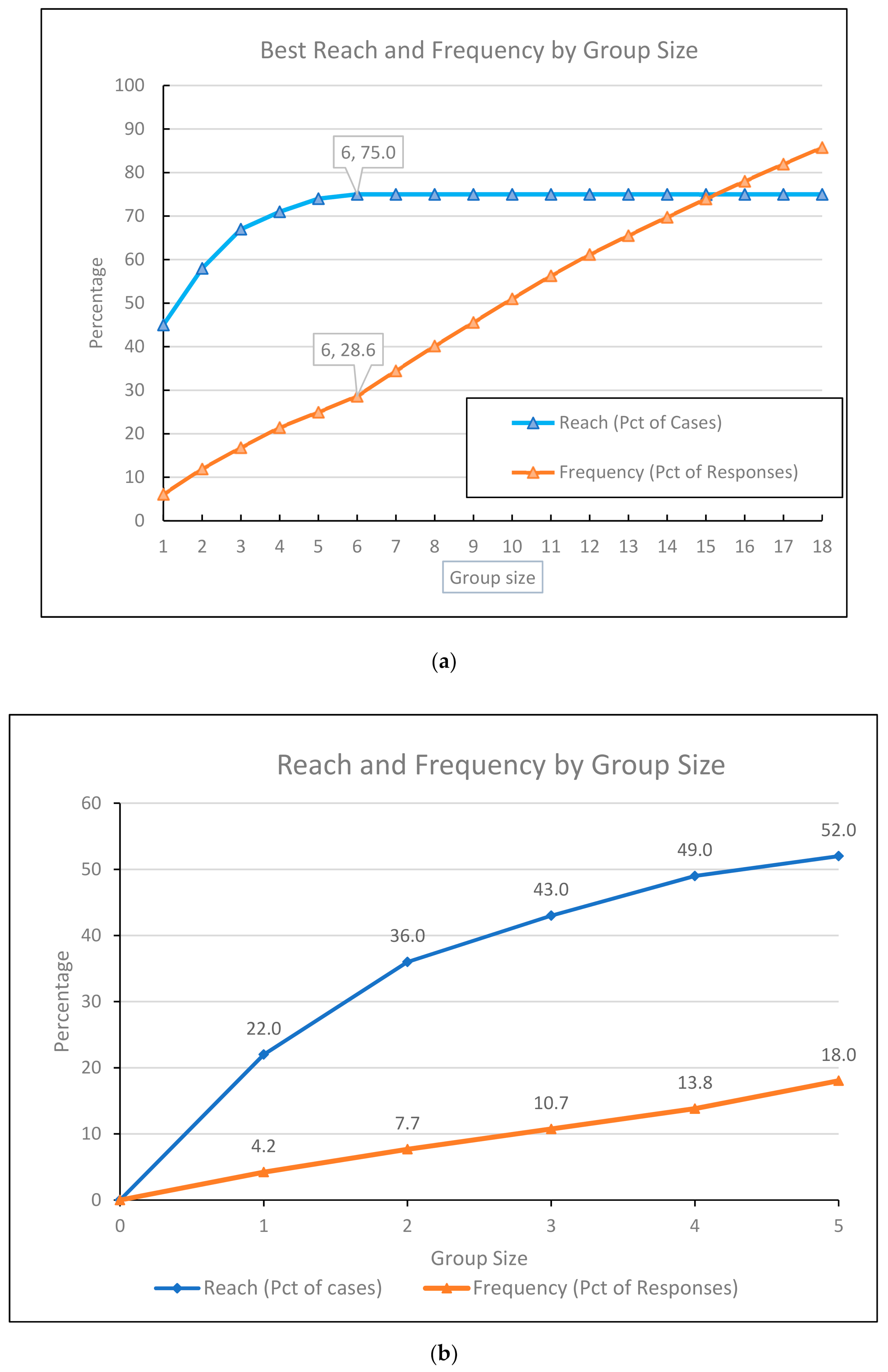
| Allergen Reagent, Molecular Components | Group Size | Reach | Pct of Cases | Frequency | Pct of Responses |
|---|---|---|---|---|---|
| a | |||||
| Alng1, Alta1, Betv1, Canf1, Cora1.0103, Cora1.0401, Cora_pollen, Derf2, Derp2, Fags1, Feld1, Lolp1, Phlp1, Phlp2, Phlp5_0101, Phlp6, Secc_pollen, Der p 23 | 18 | 75 | 75.0 | 632 | 85.8 |
| b | |||||
| Canf4, Artv, Blag9, Cynd, Feld7 | 5 | 52 | 52.0 | 94 | 18 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Čelakovská, J.; Bukač, J.; Cermákova, E.; Vaňková, R.; Skalská, H.; Krejsek, J.; Andrýs, C. Analysis of Results of Specific IgE in 100 Atopic Dermatitis Patients with the Use of Multiplex Examination ALEX2—Allergy Explorer. Int. J. Mol. Sci. 2021, 22, 5286. https://doi.org/10.3390/ijms22105286
Čelakovská J, Bukač J, Cermákova E, Vaňková R, Skalská H, Krejsek J, Andrýs C. Analysis of Results of Specific IgE in 100 Atopic Dermatitis Patients with the Use of Multiplex Examination ALEX2—Allergy Explorer. International Journal of Molecular Sciences. 2021; 22(10):5286. https://doi.org/10.3390/ijms22105286
Chicago/Turabian StyleČelakovská, Jarmila, Josef Bukač, Eva Cermákova, Radka Vaňková, Hana Skalská, Jan Krejsek, and Ctirad Andrýs. 2021. "Analysis of Results of Specific IgE in 100 Atopic Dermatitis Patients with the Use of Multiplex Examination ALEX2—Allergy Explorer" International Journal of Molecular Sciences 22, no. 10: 5286. https://doi.org/10.3390/ijms22105286
APA StyleČelakovská, J., Bukač, J., Cermákova, E., Vaňková, R., Skalská, H., Krejsek, J., & Andrýs, C. (2021). Analysis of Results of Specific IgE in 100 Atopic Dermatitis Patients with the Use of Multiplex Examination ALEX2—Allergy Explorer. International Journal of Molecular Sciences, 22(10), 5286. https://doi.org/10.3390/ijms22105286






