Boon and Bane of DNA Double-Strand Breaks
Abstract
1. Background
2. Unrepaired DSBs Can Be Lethal for Dividing Cells
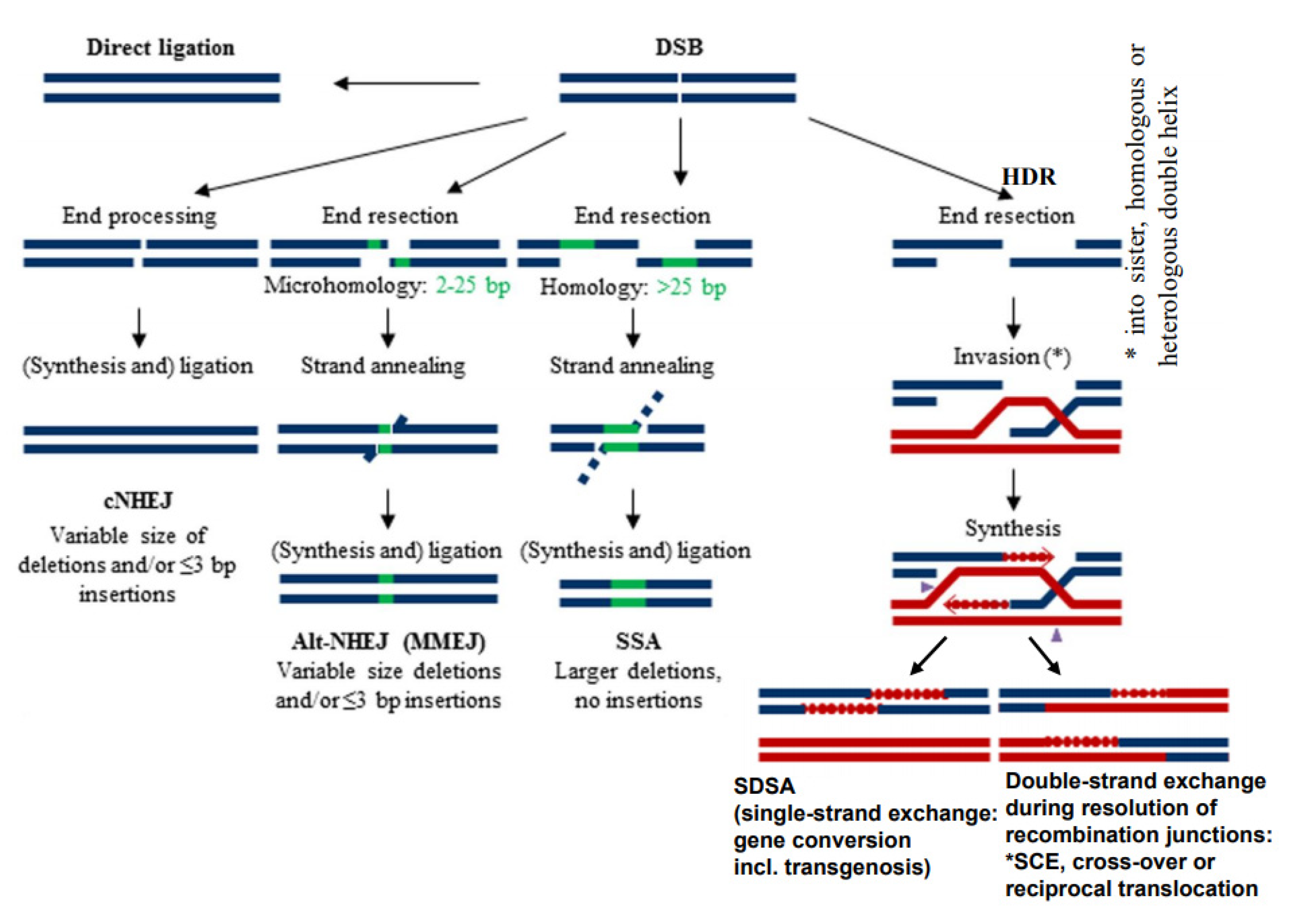
3. DSBs Can Be Repaired by Diverse Mechanisms
4. DSB Repair Generates Diverse Phenomena
5. Deleterious and Beneficial Consequences of DSB Repair
- DSBs which are programmed during meiotic prophase I are repaired in their majority without genetic consequences; a minority, via cross-over with the homologous allele, result in a new combination of maternal and paternal alleles. Cross-overs keep maternal and paternal homologous chromosomes together until reductional anaphase I, and thus enable correct segregation of parental chromosomes into gametes. The new combination of parental alleles, if beneficial for survival (and propagation) of its carriers, will be positively selected in the next generations.
- Chromosome translocations in the heterozygous state potentially reduce the fertility of carriers (due to the risk of lethality after unbalanced segregation) (Figure 4). Heterozygous inversions, if the corresponding regions engage in crossing over, will yield duplications and deletions, which are mostly lethal. Chromosome rearrangements will be eliminated if carriers bear negative features. If, however, their effect is superior to the ancestral genotype/karyotype, after passing the bottle neck towards homozygosity, the progeny will be positively selected. Such positive effects may be differential gene expression or advantageous linkage of distinct alleles, for instance. In the homozygous condition, positively selected chromosome rearrangements, and even selectively neutral ones, may contribute as initial events towards speciation (for review: [14]), because usually they act as fertility barriers.
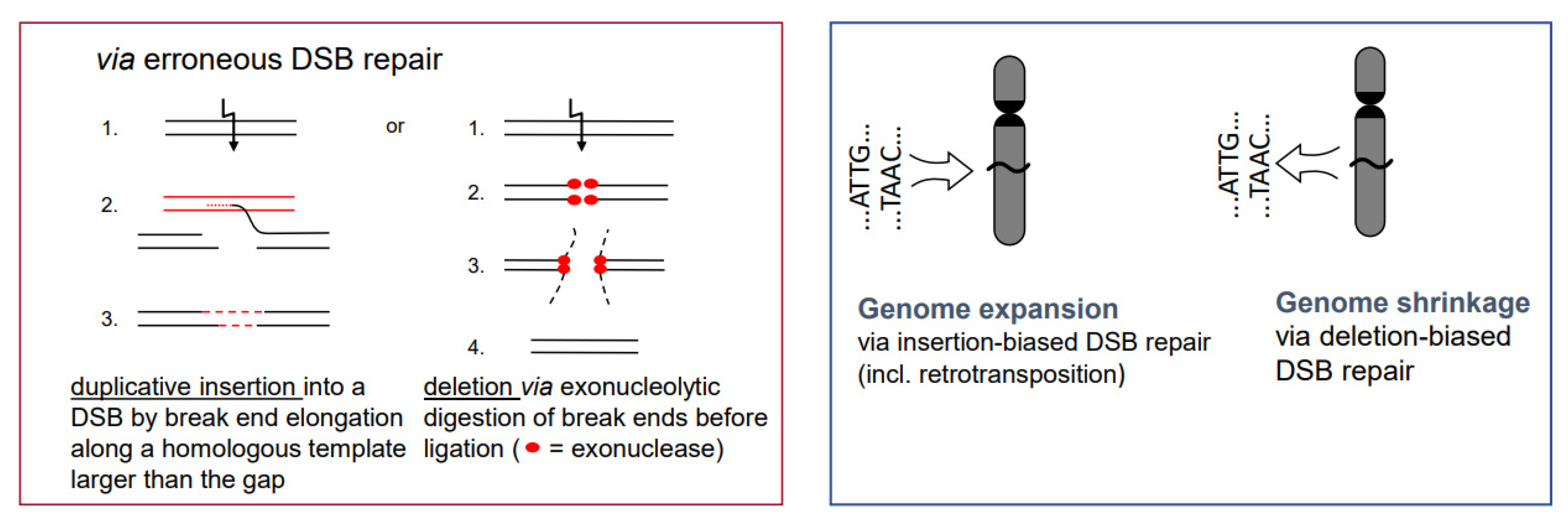
- While the correct DSB repair during meiotic prophase I results in cross-overs and leads to new combinations of pre-existing alleles, mis-repair of DSBs at any developmental stage can lead to deletion (via end-digestion), or to sequence insertion (e.g., via conversion of more than the missing sequence, via transposon invasion or via alien chromatin introgression in interspecific hybrids used in crop breeding; see Figure 1) into the break. Deletions and/or insertions create a genetic novelty which is either positive, negative or selectively neutral. Positive or neutral mutations increase genetic diversity; the latter as a playground for future mutation and selection processes.
- If there is a (genetically fixed) bias of DSB repair towards either deletions or insertions, shrinking or expansion of the genome would be the corresponding long-term consequence in a population, as long as the bias is maintained (Figure 5; for review: [11]). This might explain the C-value paradox [15], which means that the genome size is not correlated with the genetic complexity of organisms.
- 5.
- In particular, genome expansion via active retroelement amplification and dispersion is eventually the result of DSB repair [16] biased towards insertion mediated by a retroelement-encoded integrase.
- 6.
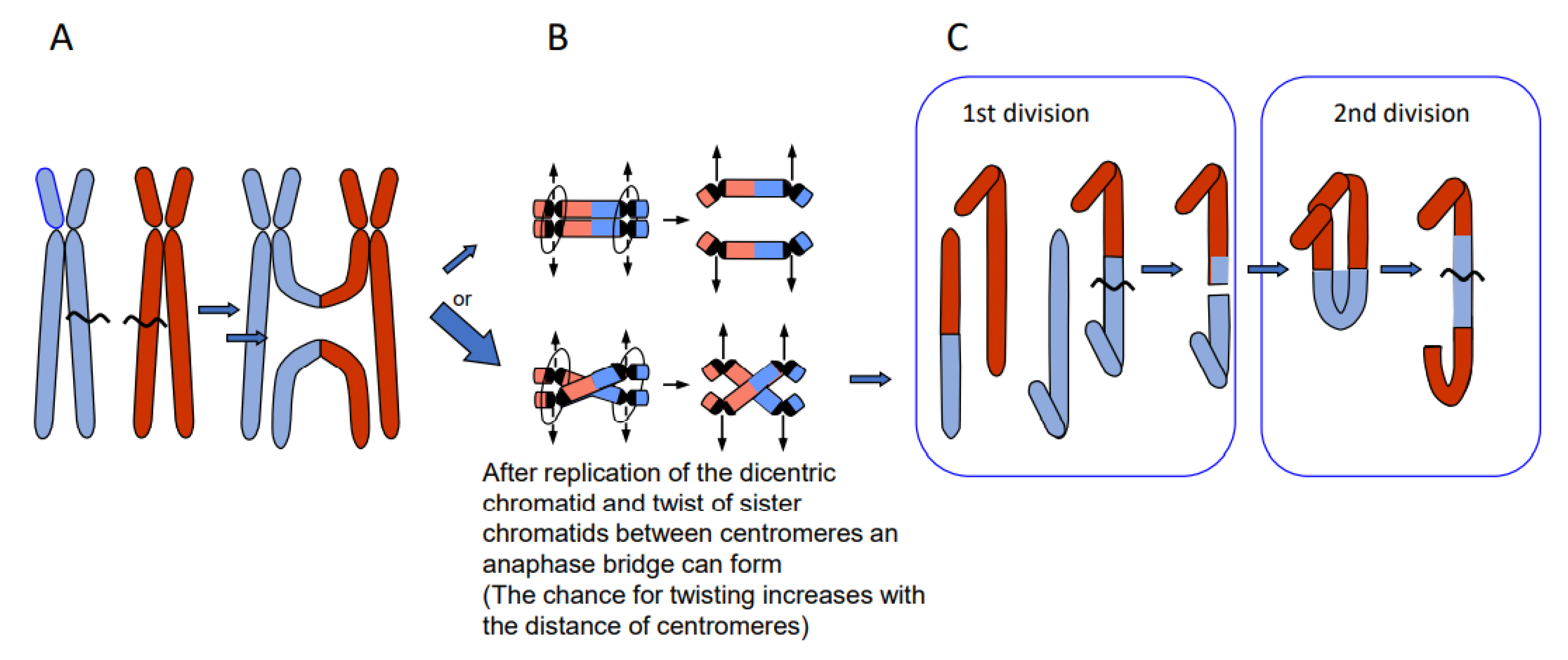
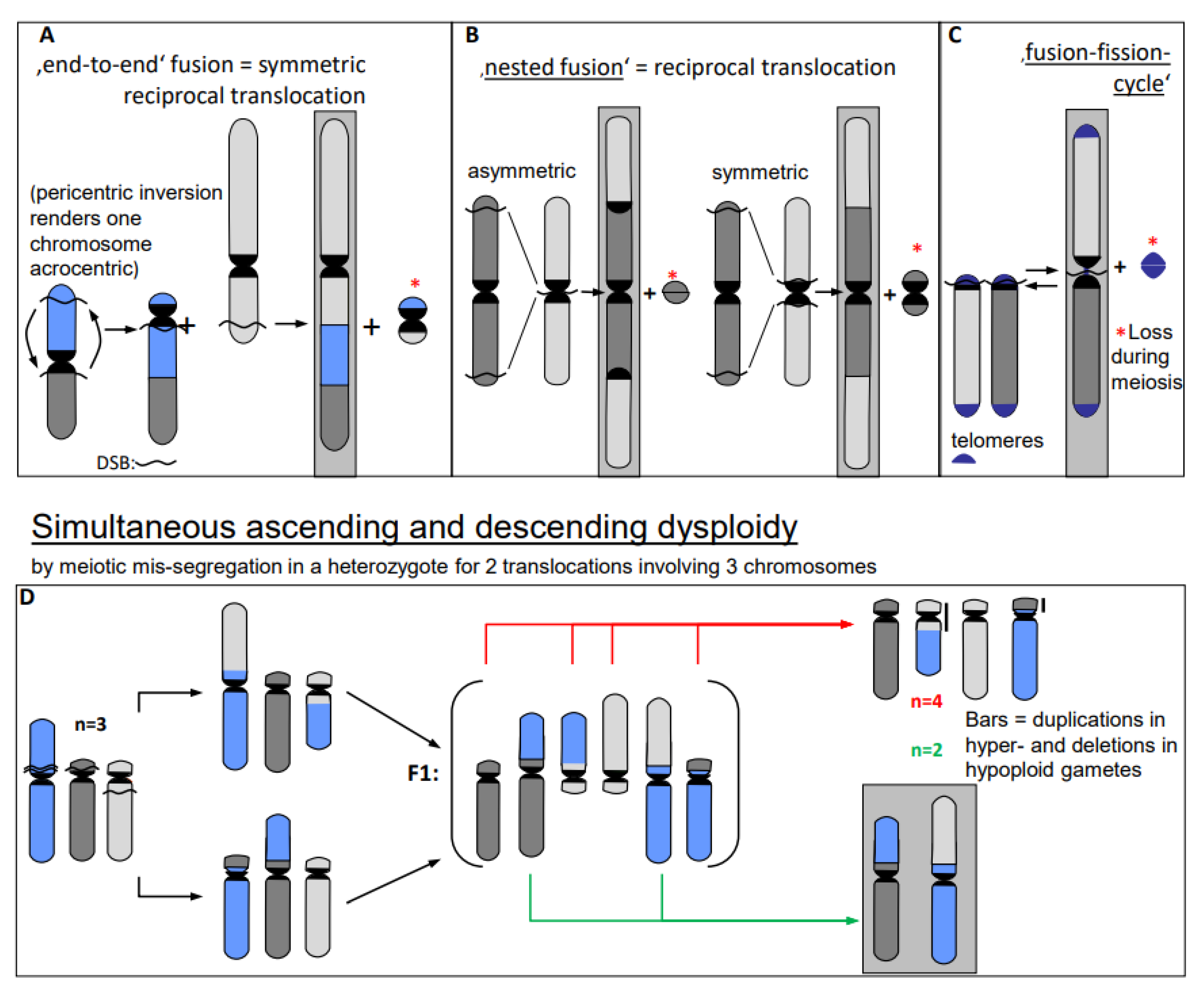
- Erroneous repair of DSBs can generate such primary chromosome rearrangements which can in turn be linked directly and/or via meiotic segregation errors with dysploid chromosome number alteration in both directions (Figure 6) (for review: [13]). Reciprocal translocation with breakpoints close to the centric ends of two acro- or telocentric chromosomes, which yield a large metacentric product and a small centric (or acentric) one, can reduce the chromosome number, if subsequent meiotic loss of the small product is tolerated [17] (Figure 6A). Similarly, insertion of a chromosome with breakpoints at both termini into a break within the centromere of a recipient chromosome reduces the chromosome number, if two telomeres and one centromere get lost or the recipient centromere becomes inactive (Figure 6B; [13]). If a metacentric gets broken in the centromere region in a way in which both fragments maintain centromere function and get stabilized by telomeric sequences, the chromosome number increases (Figure 6C, arrow to the left). This process can be reversible by translocation between these novel centric ends (Figure 6B, arrow to the right).
- 7.
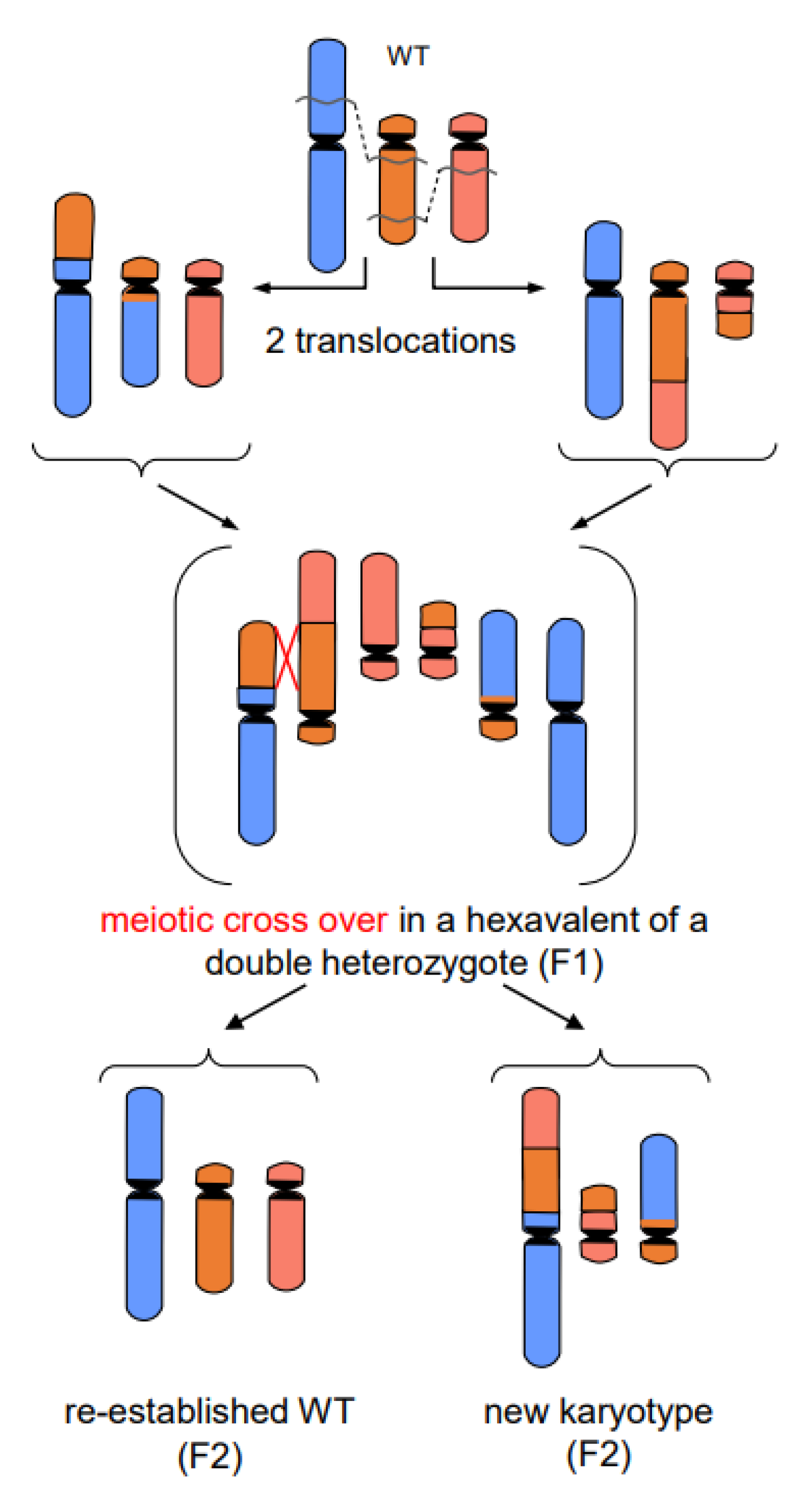
- In addition to primary rearrangements (deletion, inversion, translocation), secondary rearrangements (Figure 7) also depend on DSBs. Secondary rearrangements may occur in individuals which are heterozygous for two rearrangements with one chromosome involved in both rearrangements. If meiotic cross-over takes place between partially homozygous regions of rearranged chromosomes (flanked by different regions on either side of the cross-over), a newly rearranged chromosome segregates to one daughter nucleus and the re-established wild-type chromosome to the other. This pathway was also experimentally proven for plants, and might occur in other eukaryotes as well (for review: [13]).
- 8.
- Programmed DSBs take place during V(D)J-recombination of immunoglobulin genes in the adaptive immune system of vertebrates (for review: [18,19]). Immunoglobulins are the antibodies which recognize and neutralize antigenic proteins of invading pathogens, thus mediating disease resistance. In case of pathologic overreaction of the immune system, antibodies can cause allergies, when directed against harmless environmental antigens, or autoimmune diseases when directed against the body’s own proteins.
- 9.
- DSBs, mediated by ‘domesticated’ transposases, play an essential role in chromatin elimination. Chromatin elimination (or diminution) occurs, e.g., in protozoans, where the chromosomes of generative micronucleus are fragmented into many, much smaller (sometimes gene-sized) chromosomes of the vegetative macronucleus, removing the interspersed repetitive sequences (for review: [20]), or in somatic stem cells of roundworms (e.g., [21]). Exceptionally, B chromosomes can be eliminated from plant organs [22].
- 10.
- Programmed cell death (apoptosis) is another phenomenon accompanied by endonuclease-mediated DSBs, degrading nuclear DNA into small pieces (for review: [23]). Apoptosis represents a developmentally or extrinsically triggered suicidal cell destruction.
- 11.
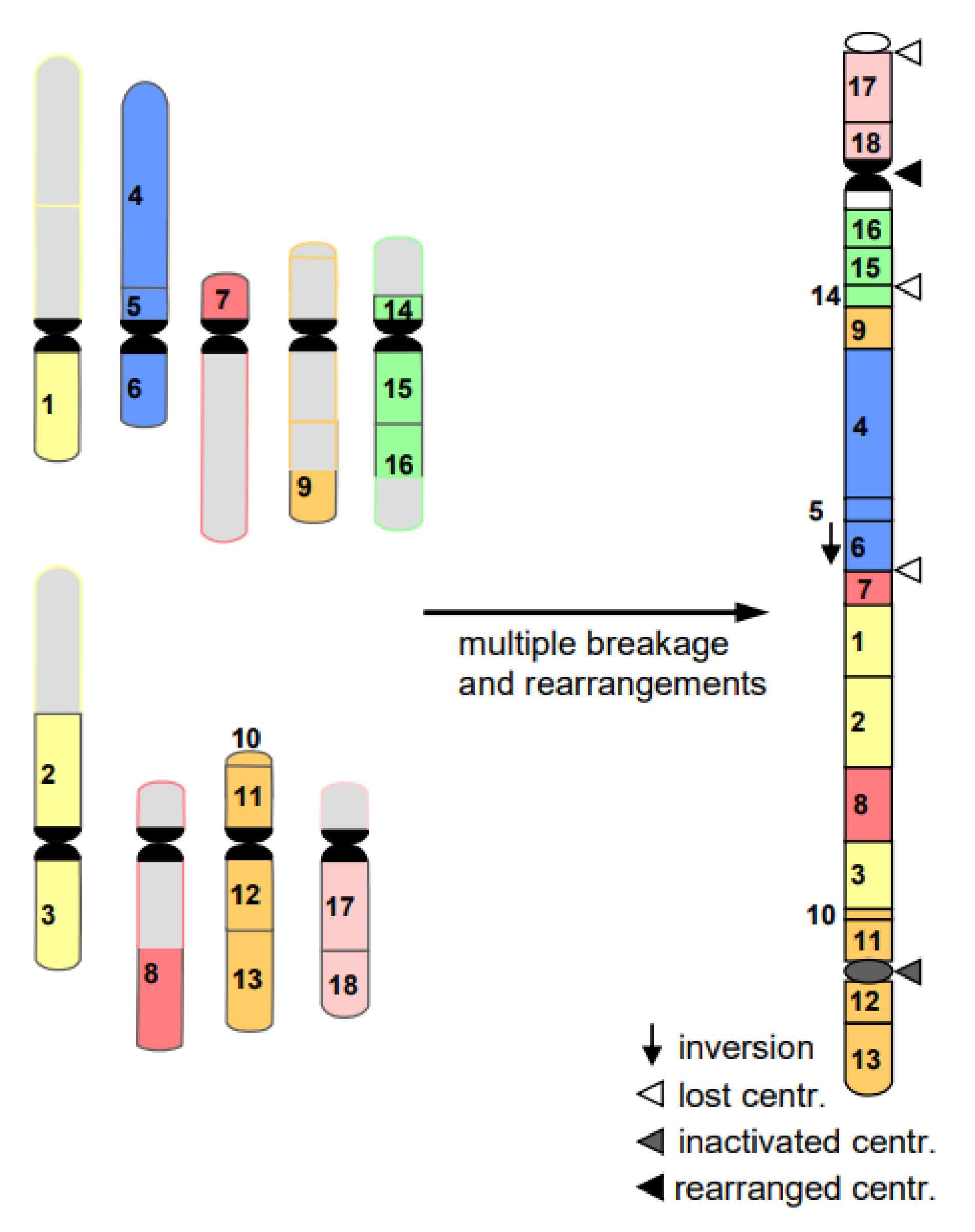
- Cancerogenesis of several tissues is also considered to start with multiple chromosome breaks as a consequence of a sudden genotoxic stress in a single cell. Such an event of catastrophic accumulation of DSBs (chromosome pulverization) and subsequent mis-repair leads simultaneously to dozens to hundreds of chromosomes rearrangements (Figure 8). The phenomenon is called chromothripsis [24]. The derivatives of the affected cell will mostly die (bottle neck) until viable versions (the malign cells) with the ability of rapid propagation prevail. This process is called evolution by several researchers (for review: [14]). True cancerogenesis is not known for plants. Nevertheless, multiple breakages and rearrangements occur during plant evolution (e.g., [25]). However, we cannot be sure whether evolutionarily fixed events appeared in a single cell, or subsequently over generations.
6. Targeted DSBs Can Modify Genetic Information for Research, Breeding and Gene Therapy
7. Concluding Remarks
Funding
Acknowledgments
Conflicts of Interest
References
- Harper, J.V.; Anderson, J.A.; O’Neill, P. Radiation induced DNA DSBs: Contribution from stalled replication forks? DNA Repair 2010, 9, 907–913. [Google Scholar] [CrossRef]
- Schiml, S.; Fauser, F.; Puchta, H. Repair of adjacent single-strand breaks is often accompanied by the formation of tandem sequence duplications in plant genomes. Proc. Natl. Acad. Sci. USA 2016, 113, 7266–7271. [Google Scholar] [CrossRef]
- Symington, L.S.; Gautier, J. Double-strand break end resection and repair pathway choice. Annu. Rev. Genet. 2011, 45, 247–271. [Google Scholar] [CrossRef]
- Schubert, V.; Neumann, P.; Marques, A.; Heckmann, S.; Macas, J.; Pedrosa-Harand, A.; Schubert, I.; Jang, T.S.; Houben, A. Super-resolution microscopy reveals diversity of plant centromere architecture. Int. J. Mol. Sci. 2020, 21, 3488. [Google Scholar] [CrossRef]
- Warecki, B.; Sullivan, W. Mechanisms driving acentric chromosome transmission. Chromosome Res. 2020, 28, 229–246. [Google Scholar] [CrossRef] [PubMed]
- Saini, N.; Ramakrishnan, S.; Elango, R.; Ayyar, S.; Zhang, Y.; Deem, A.; Ira, G.; Haber, J.E.; Lobachev, K.S.; Malkova, A. Migrating bubble during break-induced replication drives conservative DNA synthesis. Nature 2013, 502, 389–392. [Google Scholar] [CrossRef] [PubMed]
- Lieber, M.R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 2010, 79, 181–211. [Google Scholar] [CrossRef] [PubMed]
- Hanscom, T.; McVey, M. Regulation of error-prone DNA double-strand break repair and its impact on genome evolution. Cells 2020, 9, 1657. [Google Scholar] [CrossRef]
- Puchta, H. The repair of double-strand breaks in plants: Mechanisms and consequences for genome evolution. J. Exp. Bot. 2005, 56, 1–14. [Google Scholar] [CrossRef]
- Jasin, M.; Rothstein, R. Repair of strand breaks by homologous recombination. Cold Spring Harb. Perspect. Biol 2013, 5, a012740. [Google Scholar] [CrossRef]
- Schubert, I.; Vu, G.T.H. Genome stability and evolution: Attempting a holistic view. Trends Plant. Sci. 2016, 21, 749–757. [Google Scholar] [CrossRef]
- McClintock, B. The stability of broken ends of chromosomes in Zea mays. Genetics 1941, 26, 234–282. [Google Scholar] [CrossRef]
- Schubert, I.; Lysak, M.A. Interpretation of karyotype evolution should consider chromosome structural constraints. Trends Genet. 2011, 27, 207–216. [Google Scholar] [CrossRef]
- Heng, H.H. Genome Chaos: Rethinking Genetics, Evolution, and Molecular Medicine; Academic Press: Cambridge, MA, USA, 2019; p. 548. [Google Scholar]
- Thomas, C.A., Jr. The genetic organization of chromosomes. Annu. Rev. Genet. 1971, 5, 237–256. [Google Scholar] [CrossRef]
- Moore, J.K.; Haber, J.E. Capture of retrotransposon DNA at the sites of chromosomal double-strand breaks. Nature 1996, 383, 644–646. [Google Scholar] [CrossRef]
- Schubert, I. Alteration of chromosome numbers by generation of minichromosomes is there a lower limit of chromosome size for stable segregation? Cytogenet. Cell Genet. 2001, 93, 175–181. [Google Scholar] [CrossRef] [PubMed]
- Nicolas, L.; Cols, M.; Choi, J.E.; Chaudhuri, J.; Vuong, B. Generating and repairing genetically programmed DNA breaks during immunoglobulin class switch recombination. F1000 Res. 2018, 7, 458. [Google Scholar] [CrossRef] [PubMed]
- Saha, T.; Sundaravinayagam, D.; di Virgilio, M. Charting a DNA repair roadmap for immunoglobulin class switch recombination. Trends Biochem. Sci. 2021, 46, 184–199. [Google Scholar] [CrossRef]
- Rzeszutek, I.; Maurer-Alcalá, X.X.; Nowacki, M. Programmed genome rearrangements in ciliates. Cell Mol. Life Sci. 2020, 77, 4615–4629. [Google Scholar] [CrossRef] [PubMed]
- Kloc, M.; Zagrodzinska, B. Chromatin elimination–an oddity or a common mechanism in differentiation and development? Differentiation 2001, 68, 84–91. [Google Scholar] [CrossRef]
- Ruban, A.; Schmutzer, T.; Wu, D.D.; Fuchs, J.; Boudichevskaia, A.; Rubtsova, M.; Pistrick, K.; Melzer, M.; Himmelbach, A.; Schubert, V.; et al. Supernumerary B chromosomes of Aegilops speltoides undergo precise elimination in roots early in embryo development. Nat Commun 2020, 11, 2764. [Google Scholar] [CrossRef]
- D’Arcy, M.S. Cell death: A review of the major forms of apoptosis, necrosis and autophagy. Cell Biol. Int. 2019, 43, 582–592. [Google Scholar] [CrossRef] [PubMed]
- Stephens, P.J.; Greenman, C.D.; Fu, B.; Yang, F.; Bignell, G.R.; Mudie, L.J.; Pleasance, E.D.; Lau, K.W.; Beare, D.; Stebbings, L.A.; et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 2011, 144, 27–40. [Google Scholar] [CrossRef]
- Mandáková, T.; Pouch, M.; Brock, J.R.; Al-Shehbaz, I.A.; Lysak, M.A. Origin and evolution of diploid and allopolyploid Camelina genomes were accompanied by chromosome shattering. Plant. Cell 2019, 31, 2596–2612. [Google Scholar]
- Koblan, L.W.; Erdos, M.R.; Wilson, C.; Cabral, W.A.; Levy, J.M.; Xiong, Z.M.; Tavarez, U.L.; Davison, L.M.; Gete, Y.G.; Mao, X.; et al. In vivo base editing rescues Hutchinson-Gilford progeria syndrome in mice. Nature 2021, 589, 608–614. [Google Scholar] [CrossRef]
- Puchta, H. Applying CRISPR/Cas for genome engineering in plants: The best is yet to come. Curr. Opin. Plant. Biol. 2017, 36, 1–8. [Google Scholar] [CrossRef]
- Zhang, Y.; Massel, K.; Godwin, I.D.; Gao, C. Applications and potential of genome editing in crop improvement. Genome Biol. 2018, 19, 210. [Google Scholar] [CrossRef]
- Huang, T.K.; Puchta, H. CRISPR/Cas-mediated gene targeting in plants: Finally a turn for the better for homologous recombination. Plant Cell Rep. 2019, 38, 443–453. [Google Scholar] [CrossRef] [PubMed]
- Koeppel, I.; Hertig, C.; Hoffie, R.; Kumlehn, J. Cas endonuclease technology a quantum leap in the advancement of barley and wheat genetic engineering. Int. J. Mol. Sci. 2019, 20, 2647. [Google Scholar] [CrossRef]
- Schindele, A.; Dorn, A.; Puchta, H. CRISPR/Cas brings plant biology and breeding into the fast lane. Curr. Opin. Biotechnol. 2020, 61, 7–14. [Google Scholar] [CrossRef] [PubMed]
- Filler Hayut, S.; Melamed Bessudo, C.; Levy, A.A. Targeted recombination between homologous chromosomes for precise breeding in tomato. Nat. Commun. 2017, 8, 15605. [Google Scholar] [CrossRef] [PubMed]
- Beying, N.; Schmidt, C.; Pacher, M.; Houben, A.; Puchta, H. CRISPR-Cas9-mediated induction of heritable chromosomal translocations in Arabidopsis. Nat. Plants 2020, 6, 638–645. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, C.; Fransz, P.; Ronspies, M.; Dreissig, S.; Fuchs, J.; Heckmann, S.; Houben, A.; Puchta, H. Changing local recombination patterns in Arabidopsis by CRISPR/Cas mediated chromosome engineering. Nat. Commun. 2020, 11, 4418. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Sun, X.; Cormack, B.P.; Boeke, J.D. Karyotype engineering by chromosome fusion leads to reproductive isolation in yeast. Nature 2018, 560, 392–3396. [Google Scholar] [CrossRef]
- Gao, C. Genome engineering for crop improvement and future agriculture. Cell 2021, 184, 1621–31635. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schubert, I. Boon and Bane of DNA Double-Strand Breaks. Int. J. Mol. Sci. 2021, 22, 5171. https://doi.org/10.3390/ijms22105171
Schubert I. Boon and Bane of DNA Double-Strand Breaks. International Journal of Molecular Sciences. 2021; 22(10):5171. https://doi.org/10.3390/ijms22105171
Chicago/Turabian StyleSchubert, Ingo. 2021. "Boon and Bane of DNA Double-Strand Breaks" International Journal of Molecular Sciences 22, no. 10: 5171. https://doi.org/10.3390/ijms22105171
APA StyleSchubert, I. (2021). Boon and Bane of DNA Double-Strand Breaks. International Journal of Molecular Sciences, 22(10), 5171. https://doi.org/10.3390/ijms22105171




