Rolling Circle Amplification (RCA)-Mediated Genome-Wide ihpRNAi Mutant Library Construction in Brassica napus
Abstract
1. Introduction
2. Results
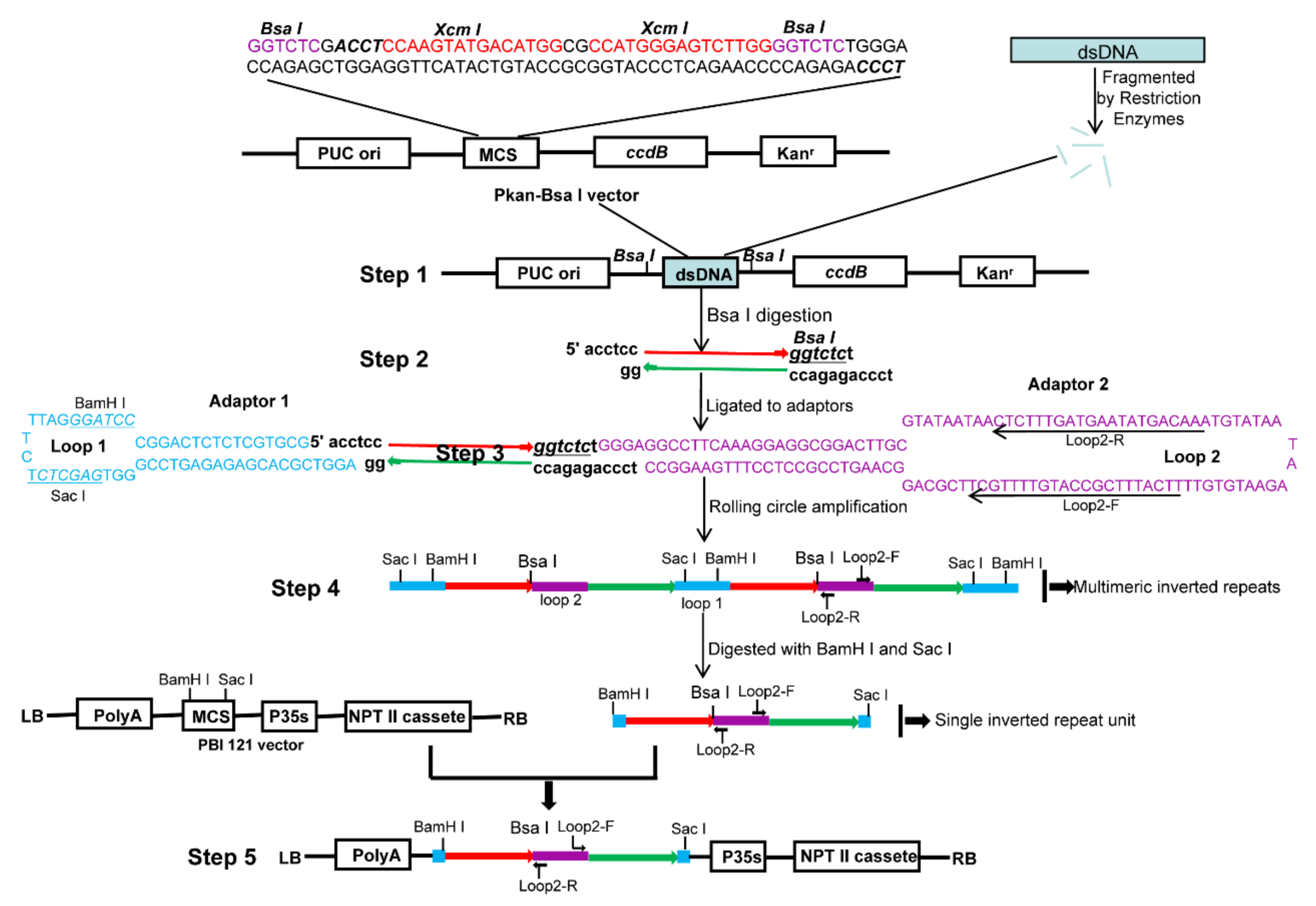
2.1. RCA-Mediated ihpRNA Library Construction Procedure
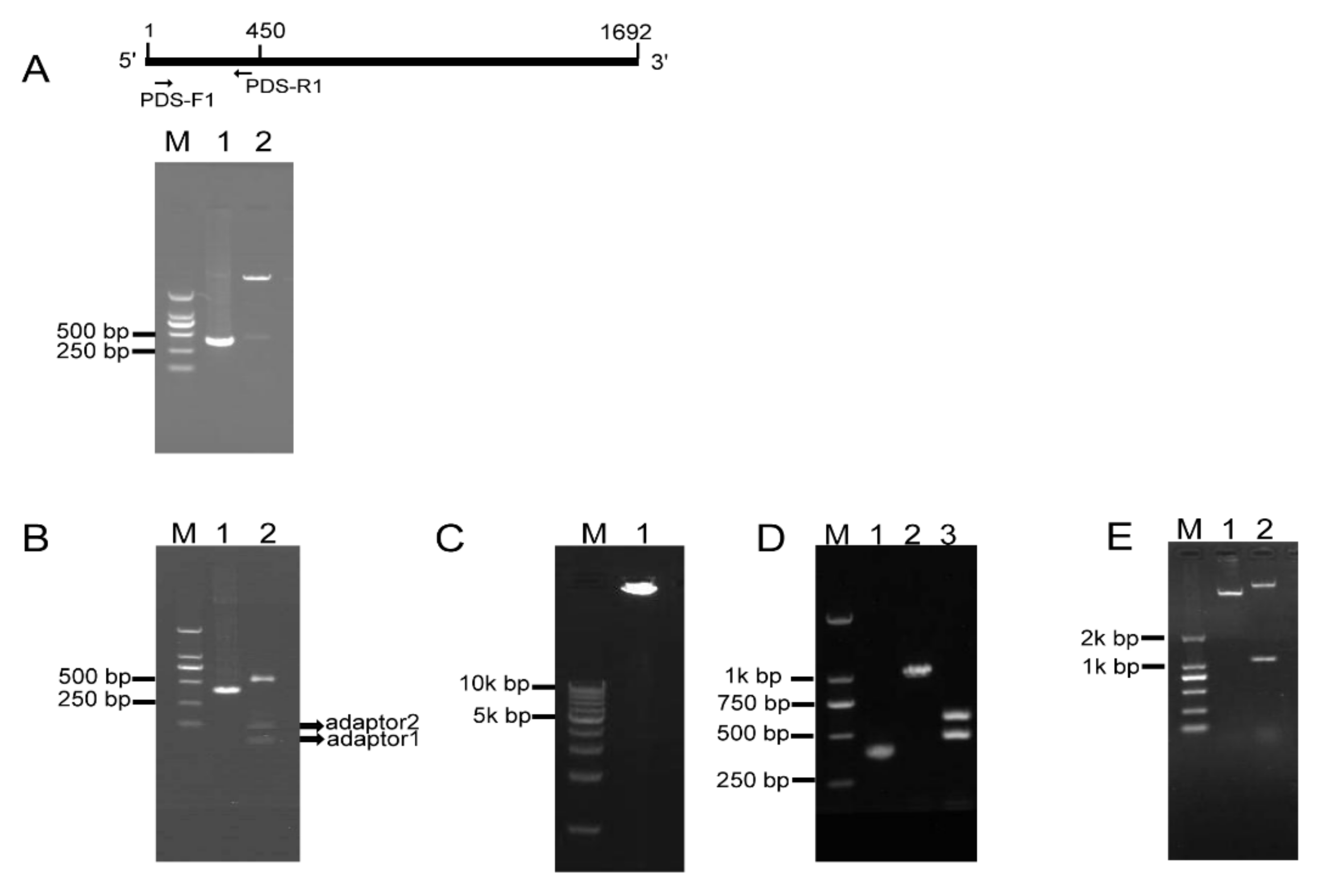
2.2. RCA-Mediated Construction of PDS ihpRNA
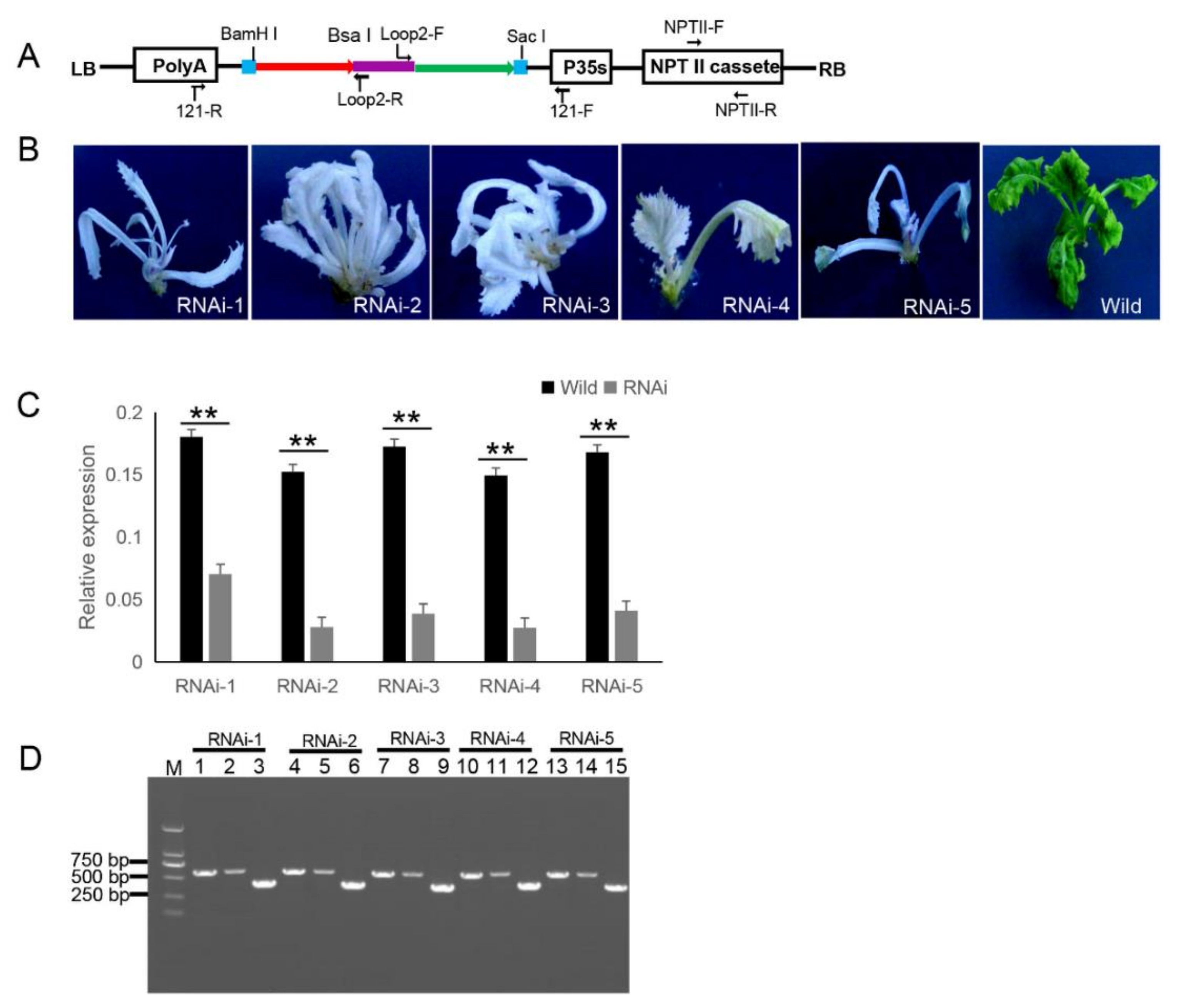
2.3. Testing of Eukaryotic Expression 121-PDS-ihpRNA Construct
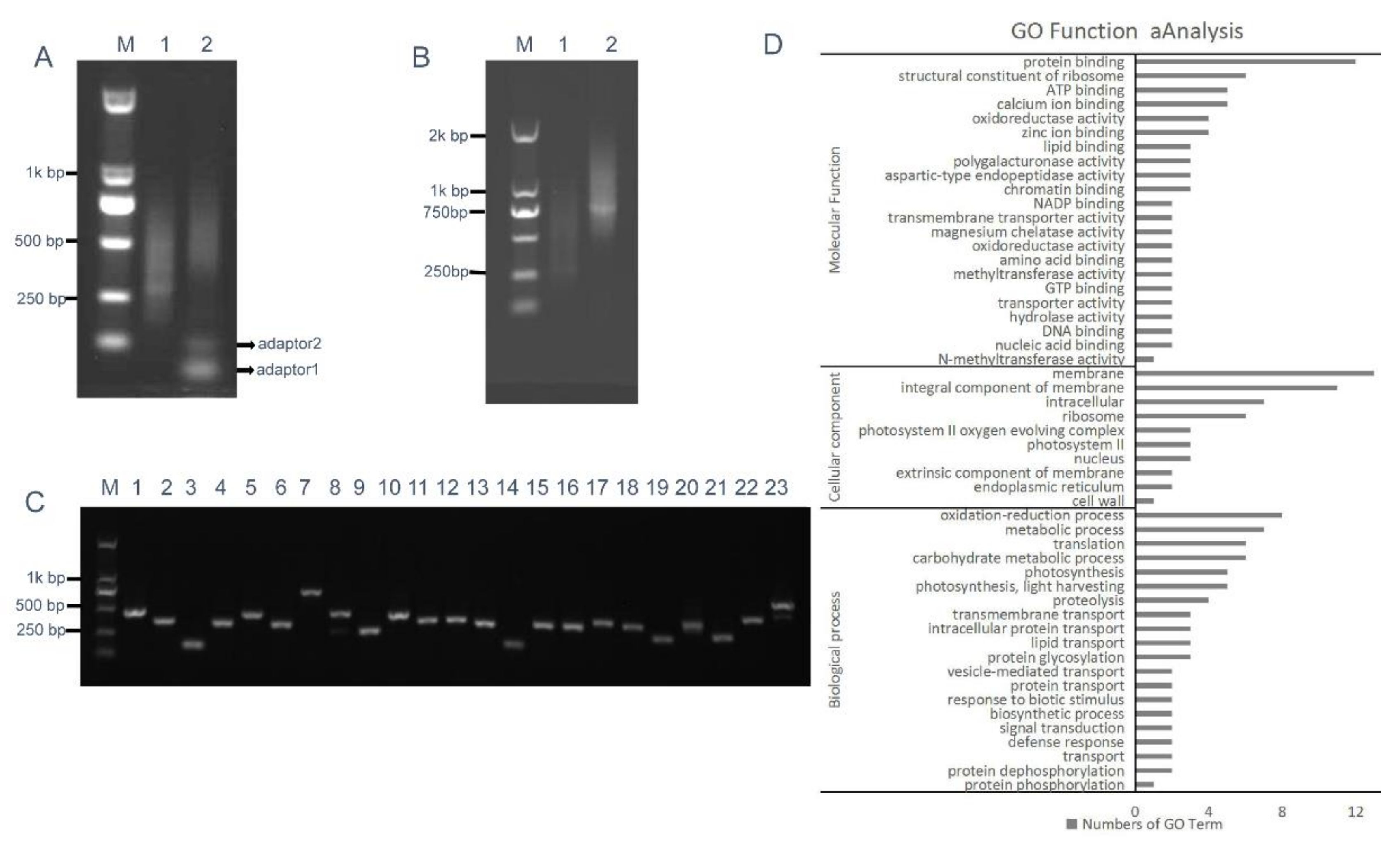
2.4. Construction of the Intermediate B. napus cDNA Library
2.5. RCA-Induced ihpRNA Library Construction of B. napus
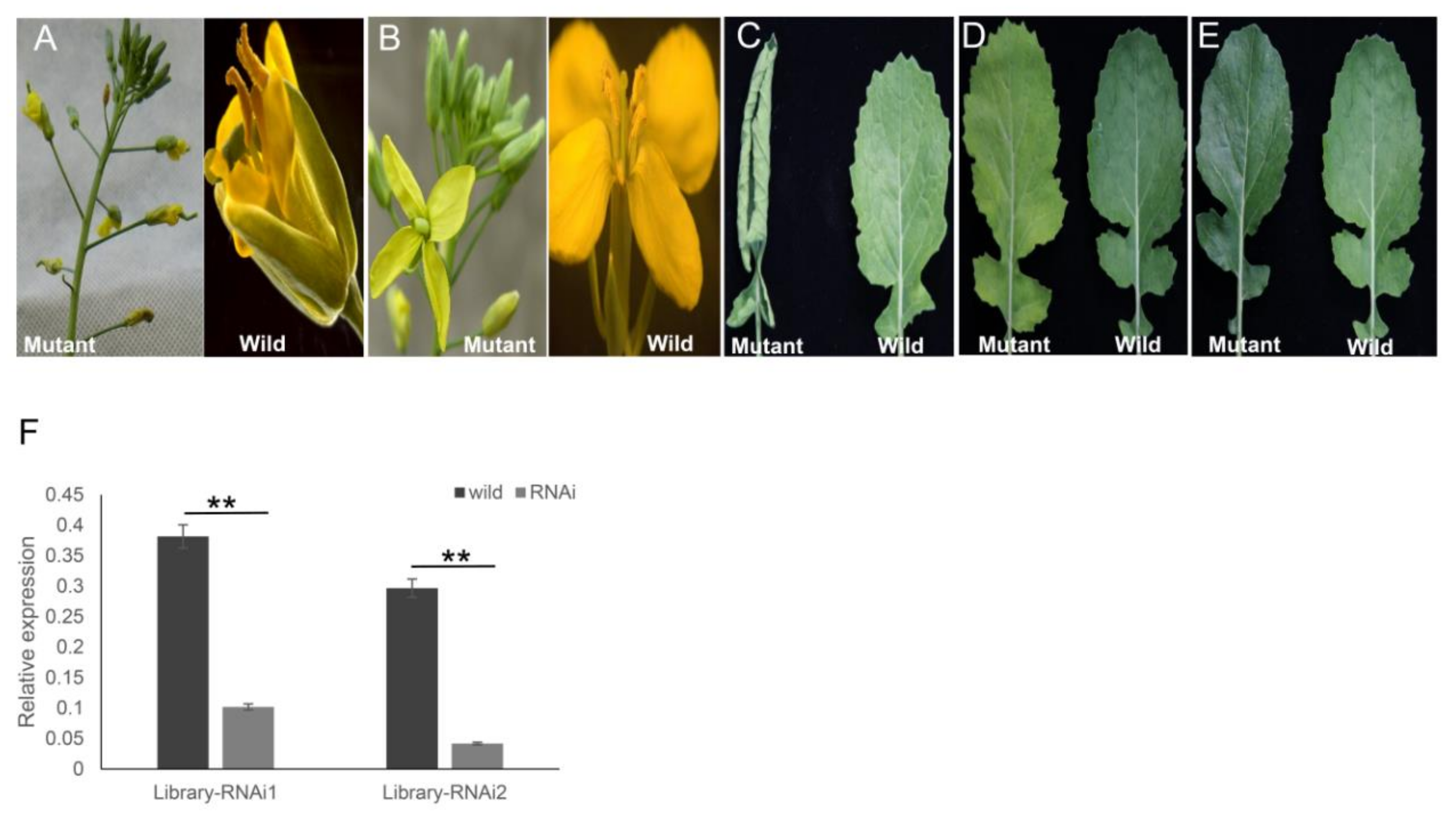
2.6. Generation of the Transgenic ihpRNA Mutant Population of B. napus
3. Discussion
4. Materials and Methods
4.1. Construction of the pKAN-BsaI Vector and Cloning of the PDS Gene Fragment
4.2. B. napus Intermediate cDNA Library Construction
4.3. The Rolling Circle Amplification (RCA)
4.4. ihpRNA Library Construction
4.5. B. napus Transformation
4.6. Quantitative Real-Time PCR (qRT-PCR)
4.7. Gene Ontology (GO) Function Analysis of Randomly Selected Genes
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ihpRNA | Long intron-spliced hairpin RNA |
| RCA | Rolling circle amplifica |
| RNAi | RNA interference |
| PTGS | Post-transcriptional gene silencing |
| dsRNA | Double-stranded RNA |
| siRNA | Small interfering RNA |
| RISCs | RNA-induced silencing complexes |
| PDS | Phytoene desaturase |
References
- Wang, X.; Wang, H.; Wang, J.; Sun, R.; Wu, J.; Liu, S.; Bai, Y.; Mun, J.H.; Bancroft, I.; Cheng, F.; et al. The genome of the mesopolyploid crop species Brassica rapa. Nat. Genet. 2011, 43, 1035–1039. [Google Scholar] [CrossRef]
- Ahn, J.H.; Kim, J.; Yoo, S.J.; Yoo, S.Y.; Roh, H.; Choi, J.H.; Choi, M.S.; Chung, K.S.; Han, E.J.; Hong, S.M.; et al. Isolation of 151 mutants that have developmental defects from T-DNA tagging. Plant Cell Physiol. 2007, 48, 169–178. [Google Scholar] [CrossRef]
- Bolle, C.; Schneider, A.; Leister, D. Perspectives on systematic analyses of gene function in Arabidopsis thaliana: New tools, topics and trends. Curr. Genom. 2011, 12, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.; Long, T.; Wu, C. Effort and contribution of T-DNA insertion mutant library for rice functional genomics research in China: Review and perspective. J. Integr. Plant Biol. 2012, 54, 953–966. [Google Scholar] [CrossRef]
- Kolesnik, T.; Szeverenyi, I.; Bachmann, D.; Kumar, C.S.; Jiang, S.; Ramamoorthy, R.; Cai, M.; Ma, Z.G.; Sundaresan, V.; Ramachandran, S. Establishing an efficient Ac/Ds tagging system in rice: Large-scale analysis of Ds flanking sequences. Plant J. 2004, 37, 301–314. [Google Scholar] [CrossRef] [PubMed]
- Piffanelli, P.; Droc, G.; Mieulet, D.; Lanau, N.; Bes, M.; Bourgeois, E.; Rouviere, C.; Gavory, F.; Cruaud, C.; Ghesquiere, A.; et al. Large-scale characterization of Tos17 insertion sites in a rice T-DNA mutant library. Plant Mol. Biol. 2007, 65, 587–601. [Google Scholar] [CrossRef] [PubMed]
- Till, B.J.; Cooper, J.; Tai, T.H.; Colowit, P.; Greene, E.A.; Henikoff, S.; Comai, L. Discovery of chemically induced mutations in rice by TILLING. BMC Plant Biol. 2007, 7, 19. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Long, T.; Yao, W.; Xiong, L.; Zhang, Q.; Wu, C. Mutant resources for the functional analysis of the rice genome. Mol. Plant 2013, 6, 596–604. [Google Scholar] [CrossRef]
- Matthew, L. RNAi for plant functional genomics. Comp. Funct. Genom. 2004, 5, 240–244. [Google Scholar] [CrossRef]
- Kim, S.K. http://C.Elegans: Mining the functional genomic landscape. Nature Rev. Genet. 2001, 2, 681–689. [Google Scholar] [CrossRef]
- Stracke, R.; Werber, M.; Weisshaar, B. The R2R3-MYB gene family in Arabidopsis thaliana. Curr. Opin. Plant Biol. 2001, 4, 447–456. [Google Scholar] [CrossRef]
- Johnston, J.S.; Pepper, A.E.; Hall, A.E.; Chen, Z.J.; Hodnett, G.; Drabek, J.; Lopez, R.; Price, H.J. Evolution of genome size in Brassicaceae. Ann. Bot. 2005, 95, 229–235. [Google Scholar] [CrossRef] [PubMed]
- Yogeeswaran, K.; Frary, A.; York, T.L.; Amenta, A.; Lesser, A.H.; Nasrallah, J.B.; Tanksley, S.D.; Nasrallah, M.E. Comparative genome analyses of Arabidopsis spp.: Inferring chromosomal rearrangement events in the evolutionary history of A. thaliana. Genome Res. 2005, 15, 505–515. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Liu, Y.; Yang, X.; Tong, C.; Edwards, D.; Parkin, I.A.; Zhao, M.; Ma, J.; Yu, J.; Huang, S.; et al. The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat. Commun. 2014, 5, 3930. [Google Scholar] [CrossRef] [PubMed]
- Chalhoub, B.; Denoeud, F.; Liu, S.; Parkin, I.A.; Tang, H.; Wang, X.; Chiquet, J.; Belcram, H.; Tong, C.; Samans, B.; et al. Plant genetics. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 2014, 345, 950–953. [Google Scholar] [CrossRef]
- Axtell, M.J. Classification and comparison of small RNAs from plants. Annu. Rev. Plant Biol. 2013, 64, 137–159. [Google Scholar] [CrossRef]
- Wang, M.B.; Masuta, C.; Smith, N.A.; Shimura, H. RNA silencing and plant viral diseases. Mol. Plant Microbe Interact. 2012, 25, 1275–1285. [Google Scholar] [CrossRef]
- Jones-Rhoades, M.W.; Bartel, D.P.; Bartel, B. MicroRNAs and their regulatory roles in plants. Annu. Rev. Plant Biol. 2006, 57, 19–53. [Google Scholar] [CrossRef]
- Chen, S.; Hofius, D.; Sonnewald, U.; Börnke, F. Temporal and spatial control of gene silencing in transgenic plants by inducible expression of double-stranded RNA. Plant J. 2003, 36, 731–740. [Google Scholar] [CrossRef]
- Guo, H.S.; Fei, J.F.; Xie, Q.; Chua, N.H. A chemical-regulated inducible RNAi system in plants. Plant J. 2003, 34, 383–392. [Google Scholar] [CrossRef]
- Wesley, S.V.; Helliwell, C.A.; Smith, N.A.; Wang, M.B.; Rouse, D.T.; Liu, Q.; Gooding, P.S.; Singh, S.P.; Abbott, D.; Stoutjesdijk, P.A.; et al. Construct design for efficient, effective and high-throughput gene silencing in plants. Plant J. 2001, 27, 581–590. [Google Scholar] [CrossRef] [PubMed]
- Helliwell, C.A.; Waterhouse, P.M. Constructs and methods for hairpin RNA-mediated gene silencing in plants. Methods Enzymol. 2005, 392, 24–35. [Google Scholar] [PubMed]
- Mansoor, S.; Amin, I.; Hussain, M.; Zafar, Y.; Briddon, R.W. Engineering novel traits in plants through RNA interference. Trends Plant Sci. 2006, 11, 559–565. [Google Scholar] [CrossRef]
- Smith, N.A.; Singh, S.P.; Wang, M.B.; Stoutjesdijk, P.A.; Green, A.G.; Waterhouse, P.M. Total silencing by intron-spliced hairpin RNAs. Nature 2000, 407, 319–320. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, P.M.; Helliwell, C.A. Exploring plant genomes by RNA-induced gene silencing. Nat. Rev. Genet. 2003, 4, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Fan, Y.L. Rolling circle amplification-mediated long hairpin RNA library construction in plants. Methods Mol Biol. 2012, 894, 309–321. [Google Scholar]
- Wang, L.; Zheng, J.; Luo, Y.; Xu, T.; Zhang, Q.; Zhang, L.; Xu, M.; Wan, J.; Wang, M.B.; Zhang, C.; et al. Construction of a genomewide RNAi mutant library in rice. Plant Biotechnol. J. 2013, 11, 997–1005. [Google Scholar] [CrossRef]
- Dean, F.B.; Nelson, J.R.; Giesler, T.L.; Lasken, R.S. Rapid amplification of plasmid and phage DNA using Phi29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res. 2001, 11, 1095–1099. [Google Scholar] [CrossRef]
- Hutchison, C.A.; Smith, H.O.; Pfannkoch, C.; Venter, J.C. Cell-free cloning using phi29 DNA polymerase. Proc. Natl. Acad. Sci. USA 2005, 102, 17332–17336. [Google Scholar] [CrossRef]
- Zeng, X.; Yan, X.; Yuan, R.; Li, K.; Wu, Y.; Liu, F.; Luo, J.; Li, J.; Wu, G. Identification and analysis of MS5d: A gene that affects double-strand break (DSB) repair during meiosis I in Brassica napus microsporocytes. Front. Plant Sci. 2016, 7, 1966. [Google Scholar] [CrossRef]
- Zou, C.; Li, G.; Qu, Z.; Chen, D.; Cheng, Y.; Zheng, P. Breeding of Brassica napus cultivar Zhongshuang No. 6 with double-low, higher-yield and resistance to Sclerotinia sclerotiorum. Chin. J. Oil Crop Sci. 2003, 25, 115–116. [Google Scholar]
- Yan, X.; Zhang, L.; Chen, B.; Xiong, Z.; Chen, C.; Wang, L.; Yu, J.; Lu, C.; Wei, W. Functional identification and characterization of the Brassica napus transcription factor gene BnAP2, the ortholog of Arabidopsis thaliana APETALA2. PLoS ONE 2012, 7, e33890. [Google Scholar]
- Liu, F.; Xiong, X.; Wang, P.; Lei, L.; Zeng, X.; Zhu, L.; Li, Y.; Luo, J.; Fu, D.; Fu, P. Effects of non-procedural factors in Brassica napus genetic transformation. Chin. J. Oil Crop Sci. 2017, 2, 106–121. [Google Scholar]
- Yang, L.; Kawakatsu, T.; Wakasa, Y.; Yoine, M.; Takaiwa, F. RNA silencing is induced by the expression of foreign recombinant products in transgenic rice. Plant Sci. 2014, 225, 138–146. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.B.; Helliwell, C.A.; Wu, L.M.; Waterhouse, P.M.; Peacock, W.J.; Dennis, E.S. Hairpin RNAs derived from RNA polymerase II and polymerase III promoter-directed transgenes are processed differently in plants. RNA 2008, 14, 903–913. [Google Scholar] [CrossRef]
- Dunoyer, P.; Himber, C.; Voinnet, O. DICER-LIKE 4 is required for RNA interference and produces the 21-nucleotide small interfering RNA component of the plant cell-to-cell silencing signal. Nat. Genet. 2005, 37, 1356–1360. [Google Scholar] [CrossRef]
- Gasciolli, V.; Mallory, A.C.; Bartel, D.P.; Vaucheret, H. Partially Redundant Functions of Arabidopsis DICER-like Enzymes and a Role for DCL4 in Producing trans-Acting siRNAs. Curr. Biol. 2005, 15, 1494–1500. [Google Scholar] [CrossRef]
- Fusaro, A.F.; Matthew, L.; Smith, N.A.; Curtin, S.J.; Dedic-Hagan, J.; Ellacott, G.A.; Watson, J.M.; Wang, M.-B.; Brosnan, C.; Carroll, B.J.; et al. RNA interference-inducing hairpin RNAs in plants act through the viral defence pathway. EMBO Rep. 2006, 7, 1168–1175. [Google Scholar] [CrossRef]
- Barnes, W.M. PCR amplification of up to 35-kb DNA with high fidelity and high yield from lambda bacteriophage templates. Proc. Natl. Acad. Sci. USA 1994, 91, 2216–2220. [Google Scholar] [CrossRef]
- Jofuku, K.D.; den Boer, B.G.; van Montagu, M.; Okamuro, J.K. Control of Arabidopsis flower and seed development by the homeotic gene APETALA2. Plant Cell 1994, 6, 1211–1225. [Google Scholar]
- Lu, W.; Liu, J.; Xin, Q.; Wan, L.; Hong, D.; Yang, G. A triallelic genetic male sterility locus in Brassica napus: An integrative strategy for its physical mapping and possible local chromosome evolution around it. Ann. Bot. 2013, 111, 305–315. [Google Scholar] [CrossRef] [PubMed]
- Xin, Q.; Shen, Y.; Li, X.; Lu, W.; Wang, X.; Han, X.; Dong, F.; Wan, L.; Yang, G.; Hong, D. MS5 mediates early meiotic progression and its natural variants may have applications for hybridproduction in Brassica napus. Plant Cell 2016, 28, 1263–1278. [Google Scholar] [CrossRef] [PubMed]
- Lizardi, P.M.; Huang, X.; Zhu, Z.; Bray-Ward, P.; Thomas, D.C.; Ward, D.C. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat. Genet. 1998, 19, 225–232. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Luo, Y.Z.; Zhang, L.; Jiao, X.M.; Wang, M.B.; Fan, Y.L. Rolling circle amplification-mediated hairpin RNA (RMHR) library construction in plants. Nucleic Acids Res. 2008, 36, e149. [Google Scholar] [CrossRef] [PubMed]
- Hirochika, H.; Guiderdoni, E.; An, G.; Hsing, Y.I.; Eun, M.Y.; Han, C.D.; Upadhyaya, N.; Ramachandran, S.; Zhang, Q.; Pereira, A. Rice mutant resources for gene discovery. Plant Mol. Biol. 2004, 54, 325–334. [Google Scholar] [CrossRef]
- Fraser, A.G.; Kamath, R.S.; Zipperlen, P.; Martinez-Campos, M.; Sohrmann, M.; Ahringer, J. Functional genomic analysis of C. elegans chromosome I by systematic RNA interference. Nature 2000, 408, 325–330. [Google Scholar] [CrossRef]
- Kamath, R.S.; Fraser, A.G.; Dong, Y.; Poulin, G.; Durbin, R.; Gotta, M.; Kanapin, A.; Le Bot, N.; Moreno, S.; Sohrmann, M.; et al. Systematic functional analysis of the Caenorhabditis elegans genome using RNAi. Nature 2003, 421, 231–237. [Google Scholar] [CrossRef]
- Wang, L.; Zhao, J.; Fan, Y. Gene cloning and function analysis of ABP9 protein which specifically binds to ABRE2 motif of maize Cat1 gene. Chin. Sci. Bull. 2002, 47, 1871–1875. [Google Scholar] [CrossRef]
- De Block, M.; de Brouwer, D.; Tenning, P. Transformation of Brassica napus and Brassica oleracea using Agrobacterium tumefaciens and the expression of the bar and neo genes in the transgenic Plants. Plant Physiol. 1989, 91, 694–701. [Google Scholar] [CrossRef]
- Xu, M.Y.; Dong, Y.; Zhang, Q.X.; Zhang, L.; Luo, Y.Z.; Sun, J.; Fan, Y.L.; Wang, L. Identification of miRNAs and their targets from Brassica napus by high-throughput sequencing and degradome analysis. BMC Genom. 2012, 13, 421. [Google Scholar] [CrossRef]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, S.; Luo, J.; Zeng, X.; Li, K.; Yuan, R.; Zhu, L.; Li, X.; Wu, G.; Yan, X. Rolling Circle Amplification (RCA)-Mediated Genome-Wide ihpRNAi Mutant Library Construction in Brassica napus. Int. J. Mol. Sci. 2020, 21, 7243. https://doi.org/10.3390/ijms21197243
Zhao S, Luo J, Zeng X, Li K, Yuan R, Zhu L, Li X, Wu G, Yan X. Rolling Circle Amplification (RCA)-Mediated Genome-Wide ihpRNAi Mutant Library Construction in Brassica napus. International Journal of Molecular Sciences. 2020; 21(19):7243. https://doi.org/10.3390/ijms21197243
Chicago/Turabian StyleZhao, Shengbo, Junling Luo, Xinhua Zeng, Keqi Li, Rong Yuan, Li Zhu, Xiaofei Li, Gang Wu, and Xiaohong Yan. 2020. "Rolling Circle Amplification (RCA)-Mediated Genome-Wide ihpRNAi Mutant Library Construction in Brassica napus" International Journal of Molecular Sciences 21, no. 19: 7243. https://doi.org/10.3390/ijms21197243
APA StyleZhao, S., Luo, J., Zeng, X., Li, K., Yuan, R., Zhu, L., Li, X., Wu, G., & Yan, X. (2020). Rolling Circle Amplification (RCA)-Mediated Genome-Wide ihpRNAi Mutant Library Construction in Brassica napus. International Journal of Molecular Sciences, 21(19), 7243. https://doi.org/10.3390/ijms21197243



