Non-Coding RNAs in the Brain-Heart Axis: The Case of Parkinson’s Disease
Abstract
1. Introduction
2. MicroRNAs
2.1. Regulation of miRNAs in Parkinson’s Disease
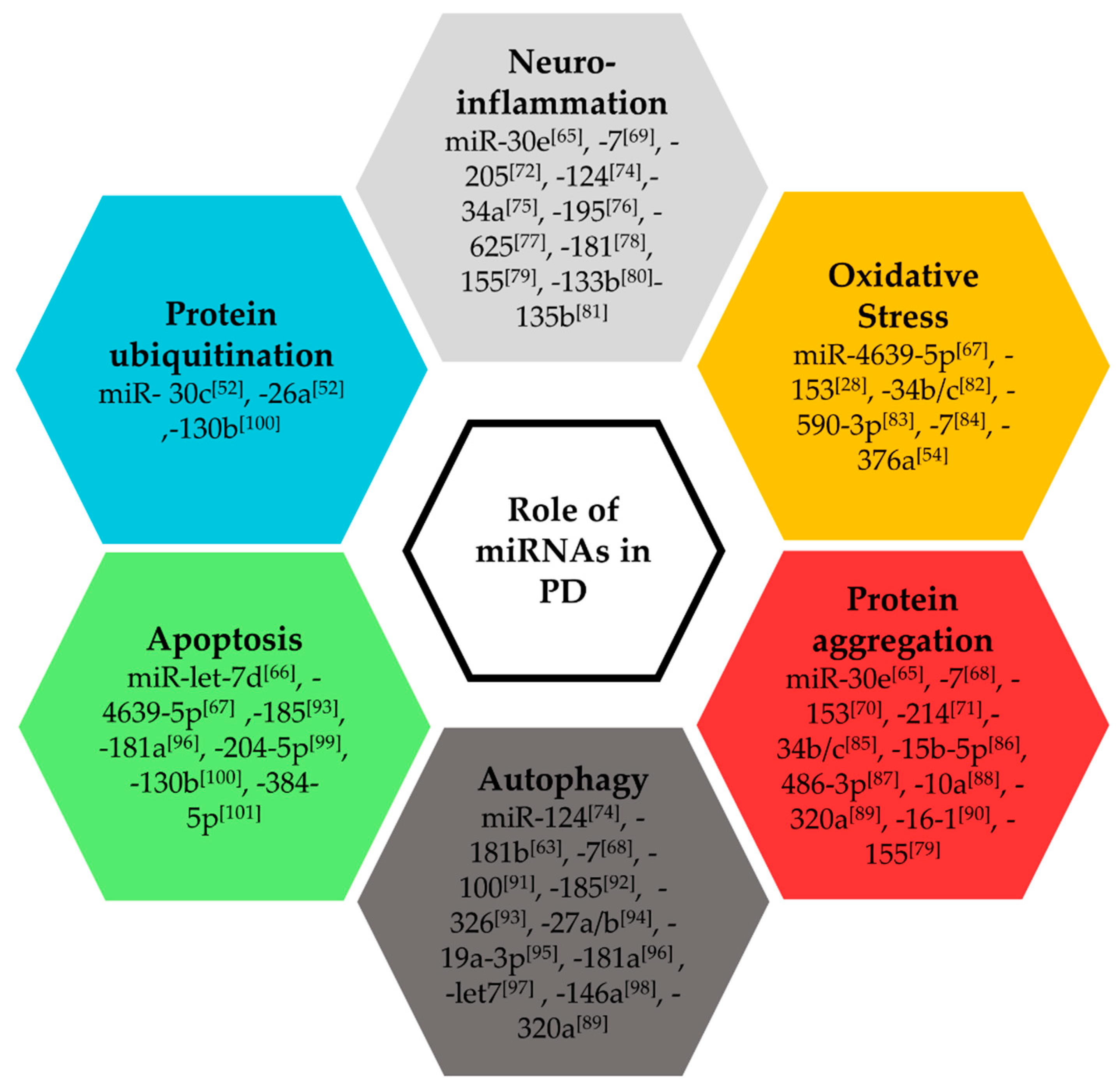
2.2. Functional Roles of miRNAs in Parkinson’s Disease
2.3. Therapeutic Potential of miRNAs in Parkinson’s Disease
3. Long Non-Coding RNAs
3.1. Regulation of lncRNAs in Parkinson’ Disease
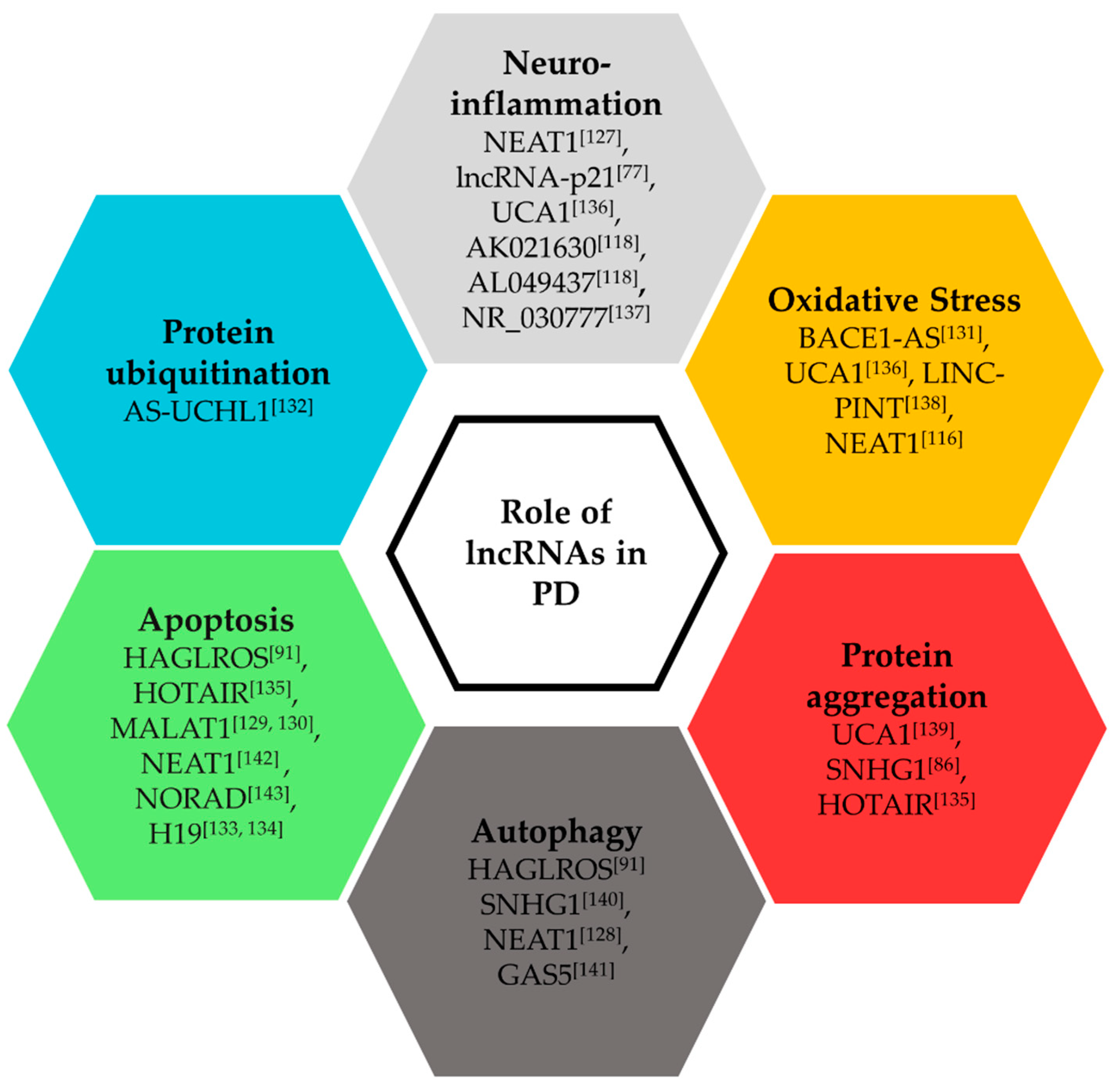
3.2. Functional Roles of lncRNAs in Parkinson’s Disease
4. Circular RNAs
5. tRNA Derived ncRNAs
6. Cardiac Comorbidities in PD
7. Non-Coding RNAs in the Brain-Heart Axis in PD
8. Translatable Techniques for Parkinson’s Disease
9. Available Gene Expression Datasets in Parkinson’s Disease
10. Harmonization of High-Throughput Datasets and Clinical Data from Parkinson’s Disease Patients
11. Systems-Level Biomedical Data Mining Applied to the Discovery of RNA Biomarkers of Parkinson’s Disease
11.1. RNA-Based Computational Biomarker Discovery for PD
11.2. Pathway- and Network-Based Biomarker Modeling for PD Research
12. Conclusions and Future Directions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| PD | Parkinson’s disease |
| miRNA | MicroRNA |
| lncRNA | Long non-coding RNA |
| circRNA | Circular RNA |
| CSF | Cerebrospinal Fluid |
| PBMC | Peripheral blood mononuclear cells |
| PPMI | Parkinson’s Progressive Markers Initiative |
| NCER-PD | National Centre of Excellence in Research on Parkinson’s Disease |
| MPTP | 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine |
| MPP | 1-methyl-4-phenylpyridinium |
| LRRK2 | Leucine-rich repeat kinase 2 |
| HOTAIR | HOX transcript antisense RNA |
| NEAT1 | Nuclear Paraspeckle Assembly Transcript 1 |
| MALAT1 | Metastasis Associated Lung Adenocarcinoma Transcript 1 |
| BACE1-AS | Beta- Secretase 1 Antisense RNA |
| HAGLROS | HAGLR Opposite Strand LncRNA |
| C. elegans | Caenorhabditis elegans |
| GEO | Gene Expression Omnibus |
References
- Rizek, P.; Kumar, N.; Jog, M. An update on the diagnosis and treatment of Parkinson disease. Can. Med. Assoc. J. 2016, 188, 1157–1165. [Google Scholar] [CrossRef]
- Bach, J.; Ziegler, U.; Deuschl, G.; Dodel, R.; Doblhammer-Reiter, G. Projected numbers of people with movement disorders in the years 2030 and 2050. Mov. Disord. 2011, 26, 2286–2290. [Google Scholar] [CrossRef]
- Antony, P.M.A.; Diederich, N.J.; Krüger, R.; Balling, R. The hallmarks of Parkinson’s disease. FEBS J. 2013, 280, 5981–5993. [Google Scholar] [CrossRef]
- Meiser, J.; Weindl, D.; Hiller, K. Complexity of dopamine metabolism. Cell Commun. Signal. 2013, 11, 34. [Google Scholar] [CrossRef] [PubMed]
- Günaydın, Z.Y.; Özer, F.F.; Karagöz, A.; Bektaş, O.; Karataş, M.B.; Vural, A.; Bayramoğlu, A.; Çelik, A.; Yaman, M. Evaluation of cardiovascular risk in patients with Parkinson disease under levodopa treatment. J. Geriatr. Cardiol. 2016, 13, 75–80. [Google Scholar]
- Scorza, F.A.; Fiorini, A.C.; Scorza, C.A.; Finsterer, J. Cardiac abnormalities in Parkinson’s disease and Parkinsonism. J. Clin. Neurosci. 2018, 53, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Troncoso-Escudero, P.; Parra, A.; Nassif, M.; Vidal, R.L. Outside in: Unraveling the Role of Neuroinflammation in the Progression of Parkinson’s Disease. Front. Neurol. 2018, 9, 860. [Google Scholar] [CrossRef] [PubMed]
- Metzger, J.M.; Moore, C.F.; Boettcher, C.A.; Brunner, K.G.; Fleddermann, R.A.; Matsoff, H.N.; Resnikoff, H.A.; Bondarenko, V.; Kamp, T.J.; Hacker, T.A.; et al. In vivo imaging of inflammation and oxidative stress in a nonhuman primate model of cardiac sympathetic neurodegeneration. NPJ Parkinson’s Dis. 2018, 4, 22. [Google Scholar] [CrossRef] [PubMed]
- Labzin, L.I.; Heneka, M.T.; Latz, E. Innate Immunity and Neurodegeneration. Annu. Rev. Med. 2018, 69, 437–449. [Google Scholar] [CrossRef]
- King, E.; Thomas, A. Systemic Inflammation in Lewy Body Diseases. Alzheimer Dis. Assoc. Disord. 2017, 31, 346–356. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, C.C.; Tarelli, R. Parkinson’s disease and systemic inflammation. Parkinsons Dis. 2011, 2011, 436813. [Google Scholar] [CrossRef] [PubMed]
- Hipp, G.; Vaillant, M.; Diederich, N.J.; Roomp, K.; Satagopam, V.P.; Banda, P.; Sandt, E.; Mommaerts, K.; Schmitz, S.K.; Longhino, L.; et al. The Luxembourg Parkinson’s Study: A Comprehensive Approach for Stratification and Early Diagnosis. Front. Aging Neurosci. 2018, 10, 326. [Google Scholar] [CrossRef] [PubMed]
- Ali, K.; Morris, H.R. Parkinson’s disease: Chameleons and mimics. Pract. Neurol. 2015, 15, 14–25. [Google Scholar] [CrossRef]
- Levin, J.; Kurz, A.; Arzberger, T.; Giese, A.; Höglinger, G.U. The Differential Diagnosis and Treatment of Atypical Parkinsonism. Dtsch. Aerzteblatt Online 2016, 113, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Le, W.; Dong, J.; Li, S.; Korczyn, A.D. Can Biomarkers Help the Early Diagnosis of Parkinson’s Disease? Neurosci. Bull. 2017, 33, 535–542. [Google Scholar] [CrossRef] [PubMed]
- Goetz, C.G.; Tilley, B.C.; Shaftman, S.R.; Stebbins, G.T.; Fahn, S.; Martinez-Martin, P.; Poewe, W.; Sampaio, C.; Stern, M.B.; Dodel, R.; et al. Movement Disorder Society-sponsored revision of the Unified Parkinson’s Disease Rating Scale (MDS-UPDRS): Scale presentation and clinimetric testing results. Mov. Disord. 2008, 23, 2129–2170. [Google Scholar] [CrossRef]
- Wang, G.; Cui, H.-L.; Liu, J.; Xiao, Q.; Wang, Y.; Ma, J.-F.; Zhou, H.-Y.; Pan, J.; Tan, Y.; Chen, S.; et al. Current approaches for the management of Parkinson’s disease in Chinese hospitals: A cross-sectional survey. BMC Neurol. 2018, 18, 122. [Google Scholar] [CrossRef]
- Michotte, A. Recent developments in the neuropathological diagnosis of Parkinson’s disease and parkinsonism. Acta Neurol. Belg. 2003, 103, 155–158. [Google Scholar]
- Goretti, E.; Wagner, D.R.; Devaux, Y. miRNAs as biomarkers of myocardial infarction: A step forward towards personalized medicine? Trends Mol. Med. 2014, 20, 716–725. [Google Scholar] [CrossRef]
- Foinquinos, A.; Batkai, S.; Genschel, C.; Viereck, J.; Rump, S.; Gyöngyösi, M.; Traxler, D.; Riesenhuber, M.; Spannbauer, A.; Lukovic, D.; et al. Preclinical development of a miR-132 inhibitor for heart failure treatment. Nat. Commun. 2020, 11, 633. [Google Scholar] [CrossRef]
- Lee, H.; Pienaar, I.S. Disruption of the blood-brain barrier in Parkinson’s disease: Curse or route to a cure? Front. Biosci. 2014, 19, 272–280. [Google Scholar] [CrossRef] [PubMed]
- Gray, M.T.; Woulfe, J.M. Striatal blood-brain barrier permeability in Parkinson’s disease. J. Cereb. Blood Flow Metab. 2015, 35, 747–750. [Google Scholar] [CrossRef] [PubMed]
- Desai, B.S.; Monahan, A.J.; Carvey, P.M.; Hendey, B. Blood-brain barrier pathology in Alzheimer’s and Parkinson’s disease: Implications for drug therapy. Cell Transplant. 2007, 16, 285–299. [Google Scholar] [CrossRef] [PubMed]
- Sweeney, M.D.; Sagare, A.P.; Zlokovic, B.V. Blood–brain barrier breakdown in Alzheimer disease and other neurodegenerative disorders. Nat. Rev. Neurol. 2018, 14, 133–150. [Google Scholar] [CrossRef]
- Network, T.C.; Devaux, Y.; Zangrando, J.; Schroen, B.; Creemers, E.E.; Pedrazzini, T.; Chang, C.-P.; Dorn, G.W.; Thum, T.; Heymans, S. Long noncoding RNAs in cardiac development and ageing. Nat. Rev. Cardiol. 2015, 12, 415–425. [Google Scholar] [CrossRef]
- Marques, T.M.; Kuiperij, H.B.; Bruinsma, I.B.; Van Rumund, A.; Aerts, M.B.; Esselink, R.A.J.; Bloem, B.R.; Verbeek, M.M. MicroRNAs in Cerebrospinal Fluid as Potential Biomarkers for Parkinson’s Disease and Multiple System Atrophy. Mol. Neurobiol. 2016, 54, 7736–7745. [Google Scholar] [CrossRef]
- Mo, M.; Xiao, Y.; Huang, S.; Cen, L.; Chen, X.; Zhang, L.; Luo, Q.; Li, S.; Yang, X.; Lin, X.; et al. MicroRNA expressing profiles in A53T mutant alpha-synuclein transgenic mice and Parkinsonian. Oncotarget 2016, 8, 15–28. [Google Scholar] [CrossRef]
- Gui, Y.-X.; Liu, H.; Zhang, L.; Lv, W.; Hu, X. Altered microRNA profiles in cerebrospinal fluid exosome in Parkinson disease and Alzheimer disease. Oncotarget 2015, 6, 37043–37053. [Google Scholar] [CrossRef]
- Jurjević, I.; Miyajima, M.; Ogino, I.; Akiba, C.; Nakajima, M.; Kondo, A.; Kikkawa, M.; Kanai, M.; Hattori, N.; Arai, H. Decreased Expression of hsa-miR-4274 in Cerebrospinal Fluid of Normal Pressure Hydrocephalus Mimics with Parkinsonian Syndromes. J. Alzheimers Dis. 2017, 56, 317–325. [Google Scholar] [CrossRef]
- Qin, L.-X.; Tan, J.-Q.; Zhang, H.-N.; Tang, J.-G.; Jiang, B.; Shen, X.-M.; Tang, B.-S.; Wang, C.-Y. Preliminary study of hsa-miR-626 change in the cerebrospinal fluid of Parkinson’s disease patients. J. Clin. Neurosci. 2019, 70, 198–201. [Google Scholar] [CrossRef]
- Cao, X.-Y.; Lu, J.-M.; Zhao, Z.-Q.; Li, M.-C.; Lu, T.; An, X.-S.; Xue, L.-J. MicroRNA biomarkers of Parkinson’s disease in serum exosome-like microvesicles. Neurosci. Lett. 2017, 644, 94–99. [Google Scholar] [CrossRef] [PubMed]
- Patil, K.S.; Basak, I.; Dalen, I.; Hoedt, E.; Lange, J.; Lunde, K.A.; Liu, Y.; Tysnes, O.-B.; Forsgren, L.; Aarsland, D.; et al. Combinatory microRNA serum signatures as classifiers of Parkinson’s disease. Park. Relat. Disord. 2019, 64, 202–210. [Google Scholar] [CrossRef]
- Li, H.; Yu, L.; Li, M.; Chen, X.; Tian, Q.; Jiang, Y.; Li, N. MicroRNA-150 serves as a diagnostic biomarker and is involved in the inflammatory pathogenesis of Parkinson’s disease. Mol. Genet. Genom. Med. 2020, 8, e1189. [Google Scholar] [CrossRef] [PubMed]
- Ozdilek, B.; Demircan, B. Serum microRNA expression levels in Turkish patients with Parkinson’s disease. Int. J. Neurosci. 2020, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, S.R.; Dionísio, P.A.; Guedes, L.C.; Gonçalves, N.; Coelho, M.M.; Rosa, M.M.; Amaral, J.D.; Ferreira, J.J.; Rodrigues, C.M.P. Circulating Inflammatory miRNAs Associated with Parkinson’s Disease Pathophysiology. Biomolecules 2020, 10, 945. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Wang, C.; Lu, S.; Yu, C.; Huang, L.; Feng, W.; Xu, H.; Chen, X.; Zen, K.; Yan, Q.; et al. A panel of four decreased serum microRNAs as a novel biomarker for early Parkinson’s disease. Biomarkers 2015, 21, 129–137. [Google Scholar] [CrossRef]
- Ma, W.; Li, Y.; Wang, C.; Xu, F.; Wang, M.; Liu, Y. Serum miR-221 serves as a biomarker for Parkinson’s disease. Cell Biochem. Funct. 2016, 34, 511–515. [Google Scholar] [CrossRef]
- Bai, X.; Tang, Y.; Yu, M.; Wu, L.; Liu, F.; Ni, J.; Wang, Z.; Wang, J.; Fei, J.; Wang, W.; et al. Downregulation of blood serum microRNA 29 family in patients with Parkinson’s disease. Sci. Rep. 2017, 7, 5411. [Google Scholar] [CrossRef]
- Khoo, S.K.; Petillo, D.; Kang, U.J.; Resau, J.H.; Berryhill, B.; Linder, J.; Forsgren, L.; Neuman, L.A.; Tan, A.C. Plasma-based circulating MicroRNA biomarkers for Parkinson’s disease. J. Parkinsons Dis. 2012, 2, 321–331. [Google Scholar] [CrossRef]
- Cardo, L.F.; Coto, E.; De Mena, L.; Ribacoba, R.; Morís, G.; Menéndez, M.; Alvarez, V. Profile of microRNAs in the plasma of Parkinson’s disease patients and healthy controls. J. Neurol. 2013, 260, 1420–1422. [Google Scholar] [CrossRef]
- Li, N.; Pan, X.; Zhang, J.; Ma, A.-J.; Xie, A.; Yang, S. Plasma levels of miR-137 and miR-124 are associated with Parkinson’s disease but not with Parkinson’s disease with depression. Neurol. Sci. 2017, 38, 761–767. [Google Scholar] [CrossRef] [PubMed]
- Schwienbacher, C.; Foco, L.; Picard, A.; Corradi, E.; Serafin, A.; Panzer, J.; Zanigni, S.; Blankenburg, H.; Facheris, M.F.; Giannini, G.; et al. Plasma and White Blood Cells Show Different miRNA Expression Profiles in Parkinson’s Disease. J. Mol. Neurosci. 2017, 62, 244–254. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Li, T.; Cui, Y.; Li, S.; Cheng, C.; Shen, B.; Le, W. Elevated Plasma microRNA-105-5p Level in Patients with Idiopathic Parkinson’s Disease: A Potential Disease Biomarker. Front. Mol. Neurosci. 2019, 13, 218. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Yang, R.; Hu, B.-L.; Lu, P.; Zhou, L.-L.; He, Z.-Y.; Wu, H.-M.; Zhu, J.-H. Reduced Circulating Levels of miR-433 and miR-133b Are Potential Biomarkers for Parkinson’s Disease. Front. Cell. Neurosci. 2017, 11, 170. [Google Scholar] [CrossRef]
- Chen, L.; Yang, J.; Lü, J.; Cao, S.; Zhao, Q.; Yu, Z. Identification of aberrant circulating miRNAs in Parkinson’s disease plasma samples. Brain Behav. 2018, 8, e00941. [Google Scholar] [CrossRef]
- Yao, Y.F.; Qu, M.W.; Li, G.C.; Zhang, F.B.; Rui, H.C. Circulating exosomal miRNAs as diagnostic biomarkers in Parkinson’s disease. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 5278–5283. [Google Scholar]
- Yang, Z.; Li, T.; Li, S.; Wei, M.; Qi, H.; Shen, B.; Chang, R.C.-C.; Le, W.; Piao, F. Altered Expression Levels of MicroRNA-132 and Nurr1 in Peripheral Blood of Parkinson’s Disease: Potential Disease Biomarkers. ACS Chem. Neurosci. 2019, 10, 2243–2249. [Google Scholar] [CrossRef]
- Ravanidis, S.; Bougea, A.; Papagiannakis, N.; Maniati, M.; Koros, C.; Simitsi, A.M.; Bozi, M.; Pachi, I.; Stamelou, M.; Paraskevas, G.P.; et al. Circulating Brain-enriched MicroRNAs for detection and discrimination of idiopathic and genetic Parkinson’s disease. Mov. Disord. 2020, 35, 457–467. [Google Scholar] [CrossRef]
- Pasinetti, G.M. Role of Personalized Medicine in the Identification and Characterization of Parkinson’s Disease in Asymptomatic Subjects. J. Alzheimers Dis. Parkinsonism 2012, 2. [Google Scholar] [CrossRef]
- Serafin, A.; Foco, L.; Zanigni, S.; Blankenburg, H.; Picard, A.; Zanon, A.; Giannini, G.; Pichler, I.; Facheris, M.F.; Cortelli, P.; et al. Overexpression of blood microRNAs 103a, 30b, and 29a in L-dopa-treated patients with PD. Neurology 2015, 84, 645–653. [Google Scholar] [CrossRef]
- Caggiu, E.; Paulus, K.; Mameli, G.; Arru, G.; Sechi, G.P.; Sechi, L.A. Differential expression of miRNA 155 and miRNA 146a in Parkinson’s disease patients. eNeurologicalSci 2018, 13, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Martins, M.; Rosa, A.; Guedes, L.C.; Fonseca, B.V.; Gotovac, K.; Violante, S.; Mestre, T.; Coelho, M.; Rosa, M.M.; Martin, E.R.; et al. Convergence of miRNA expression profiling, alpha-synuclein interacton and GWAS in Parkinson’s disease. PLoS ONE 2011, 6, e25443. [Google Scholar] [CrossRef] [PubMed]
- Behbahanipour, M.; Peymani, M.; Salari, M.; Hashemi, M.S.; Nasr-Esfahani, M.H.; Ghaedi, K. Expression Profiling of Blood microRNAs 885, 361, and 17 in the Patients with the Parkinson’s disease: Integrating Interaction Data to Uncover the Possible Triggering Age-Related Mechanisms. Sci Rep. 2019, 9, 13759. [Google Scholar] [CrossRef] [PubMed]
- Baghi, M.; Rostamian Delavar, M.; Yadegari, E.; Peymani, M.; Pozo, D.; Hossein Nasr-Esfahani, M.; Ghaedi, K. Modified level of miR-376a is associated with Parkinson’s disease. J. Cell. Mol. Med. 2020, 24, 2622–2634. [Google Scholar] [CrossRef] [PubMed]
- Alieva, A.K.; Filatova, E.V.; Karabanov, A.V.; Illarioshkin, S.N.; Limborska, S.A.; Shadrina, M.I.; Slominsky, P.A. miRNA expression is highly sensitive to a drug therapy in Parkinson’s disease. Park. Relat. Disord. 2015, 21, 72–74. [Google Scholar] [CrossRef] [PubMed]
- Cressatti, M.; Juwara, L.; Galindez, J.M.; Velly, A.M.; Nkurunziza, E.S.; Marier, S.; Canie, O.; Gornistky, M.; Schipper, H.M. Salivary microR-153 and microR-223 Levels as Potential Diagnostic Biomarkers of Idiopathic Parkinson’s Disease. Mov. Disord. 2019, 35, 468–477. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zheng, J.; Su, L.; Chen, F.; Zhu, R.; Chen, X.; Ye, Q. Increased Salivary microRNAs That Regulate DJ-1 Gene Expression as Potential Markers for Parkinson’s Disease. Front. Aging Neurosci. 2020, 12, 210. [Google Scholar] [CrossRef]
- Burgos, K.; Malenica, I.; Metpally, R.; Courtright, A.; Rakela, B.; Beach, T.; Shill, H.; Adler, C.; Sabbagh, M.; Villa, S.; et al. Profiles of extracellular miRNA in cerebrospinal fluid and serum from patients with Alzheimer’s and Parkinson’s diseases correlate with disease status and features of pathology. PLoS ONE 2014, 9, e94839. [Google Scholar] [CrossRef]
- Devaux, Y.; Dankiewicz, J.; Salgado-Somoza, A.; Stammet, P.; Collignon, O.; Gilje, P.; Gidlof, O.; Zhang, L.; Vausort, M.; Hassager, C.; et al. Association of Circulating MicroRNA-124-3p Levels with Outcomes after out-of-Hospital Cardiac Arrest: A Substudy of a Randomized Clinical Trial. JAMA Cardiol. 2016, 1, 305–313. [Google Scholar] [CrossRef]
- Devaux, Y.; Salgado-Somoza, A.; Dankiewicz, J.; Boileau, A.; Stammet, P.; Schritz, A.; Zhang, L.; Vausort, M.; Gilje, P.; Erlinge, D.; et al. Incremental Value of Circulating MiR-122-5p to Predict Outcome after out of Hospital Cardiac Arrest. Theranostics 2017, 7, 2555–2564. [Google Scholar] [CrossRef]
- Braza-Boïls, A.; Barwari, T.; Gutmann, C.; Thomas, M.R.; Judge, H.M.; Joshi, A.; Pechlaner, R.; Shankar-Hari, M.; Ajjan, R.A.; Sabroe, I.; et al. Circulating MicroRNA Levels Indicate Platelet and Leukocyte Activation in Endotoxemia Despite Platelet P2Y12 Inhibition. Int. J. Mol. Sci. 2020, 21, 2897. [Google Scholar] [CrossRef] [PubMed]
- Devaux, Y. Transcriptome of blood cells as a reservoir of cardiovascular biomarkers. Biochim. Biophys. Acta (BBA) Bioenerg. 2017, 1864, 209–216. [Google Scholar] [CrossRef] [PubMed]
- Kern, F.; Fehlmann, T.; Violich, I.; Alsop, E.; Hutchins, E.D.; Kahraman, M.; Grammes, N.; Guimaraes, P.; Backes, C.; Poston, K.L.; et al. Deep sncRNA-seq of the PPMI cohort to study Parkinson’s disease progression. BioRxiv 2020. [Google Scholar] [CrossRef]
- Briggs, C.E.; Wang, Y.; Kong, B.; Woo, T.-U.W.; Iyer, L.K.; Sonntag, K.C. Midbrain dopamine neurons in Parkinson’s disease exhibit a dysregulated miRNA and target-gene network. Brain Res. 2015, 1618, 111–121. [Google Scholar] [CrossRef]
- Li, D.; Yang, H.; Ma, J.; Luo, S.; Chen, S.; Gu, Q. MicroRNA-30e regulates neuroinflammation in MPTP model of Parkinson’s disease by targeting Nlrp3. Hum. Cell 2017, 31, 106–115. [Google Scholar] [CrossRef]
- Li, L.; Liu, H.; Song, H.; Qin, Y.; Wang, Y.; Xu, M.; Liu, C.; Gao, J.; Sun, S. Let-7d microRNA Attenuates 6-OHDA-Induced Injury by Targeting Caspase-3 in MN9D Cells. J. Mol. Neurosci. 2017, 63, 403–411. [Google Scholar] [CrossRef]
- Chen, Y.; Gao, C.; Sun, Q.; Pan, H.; Huang, P.; Ding, J.; Chen, S. MicroRNA-4639 Is a Regulator of DJ-1 Expression and a Potential Early Diagnostic Marker for Parkinson’s Disease. Front. Aging Neurosci. 2017, 9, 232. [Google Scholar] [CrossRef]
- Choi, D.C.; Yoo, M.; Kabaria, S.; Junn, E. MicroRNA-7 facilitates the degradation of alpha-synuclein and its aggregates by promoting autophagy. Neurosci. Lett. 2018, 678, 118–123. [Google Scholar] [CrossRef]
- Cao, B.; Wang, T.; Qu, Q.; Kang, T.; Yang, Q. Long Noncoding RNA SNHG1 Promotes Neuroinflammation in Parkinson’s Disease via Regulating miR-7/NLRP3 Pathway. Neuroscience 2018, 388, 118–127. [Google Scholar] [CrossRef]
- Je, G.; Kim, Y.-S. Mitochondrial ROS-mediated post-transcriptional regulation of α-synuclein through miR-7 and miR-153. Neurosci. Lett. 2017, 661, 132–136. [Google Scholar] [CrossRef]
- Wang, Z.-H.; Zhang, J.-L.; Duan, Y.-L.; Zhang, Q.-S.; Li, G.-F.; Zheng, D.-L. MicroRNA-214 participates in the neuroprotective effect of Resveratrol via inhibiting α-synuclein expression in MPTP-induced Parkinson’s disease mouse. Biomed. Pharmacother. 2015, 74, 252–256. [Google Scholar] [CrossRef] [PubMed]
- Cho, H.J.; Liu, G.; Jin, S.M.; Parisiadou, L.; Xie, C.; Yu, J.; Sun, L.; Ma, B.; Ding, J.; Vancraenenbroeck, R.; et al. MicroRNA-205 regulates the expression of Parkinson’s disease-related leucine-rich repeat kinase 2 protein. Hum. Mol. Genet. 2013, 22, 608–620. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Jiang, Y.; Wang, Y.; Yang, S.; Bi, X.; Pan, X.; Ma, A.-J. MiR-181b regulates autophagy in a model of Parkinson’s disease by targeting the PTEN/Akt/mTOR signaling pathway. Neurosci. Lett. 2018, 675, 83–88. [Google Scholar] [CrossRef] [PubMed]
- Yao, L.; Ye, Y.; Mao, H.; Lu, F.; He, X.; Lu, G.; Zhang, S. MicroRNA-124 regulates the expression of MEKK3 in the inflammatory pathogenesis of Parkinson’s disease. J. Neuroinflam. 2018, 15, 13. [Google Scholar] [CrossRef] [PubMed]
- Horst, C.H.; Titze-De-Almeida, R.; De Almeida, S.S.T. The involvement of Eag1 potassium channels and miR-34a in rotenone-induced death of dopaminergic SH-SY5Y cells. Mol. Med. Rep. 2017, 15, 1479–1488. [Google Scholar] [CrossRef]
- Ren, Y.; Li, H.; Xie, W.; Wei, N.; Liu, M. MicroRNA195 triggers neuroinflammation in Parkinson’s disease in a Rhoassociated kinase 1dependent manner. Mol. Med. Rep. 2019, 19, 5153–5161. [Google Scholar] [CrossRef]
- Ding, X.-M.; Zhao, L.-J.; Qiao, H.-Y.; Wu, S.-L.; Wang, X.-H. Long non-coding RNA-p21 regulates MPP+-induced neuronal injury by targeting miR-625 and derepressing TRPM2 in SH-SY5Y cells. Chem. Interact. 2019, 307, 73–81. [Google Scholar] [CrossRef]
- Ye, Y.; He, X.; Lu, F.; Mao, H.; Zhu, Z.; Yao, L.; Luo, W.; Sun, X.; Wang, B.; Qian, C.; et al. A lincRNA-p21/miR-181 family feedback loop regulates microglial activation during systemic LPS- and MPTP- induced neuroinflammation. Cell Death Dis. 2018, 9, 803. [Google Scholar] [CrossRef]
- Thome, A.D.; Harms, A.S.; Volpicelli-Daley, L.A.; Standaert, D.G. microRNA-155 Regulates Alpha-Synuclein-Induced Inflammatory Responses in Models of Parkinson Disease. J. Neurosci. 2016, 36, 2383–2390. [Google Scholar] [CrossRef]
- Zhang, L.-M.; Wang, M.-H.; Yang, H.-C.; Tian, T.; Sun, G.-F.; Ji, Y.-F.; Hu, W.-T.; Liu, X.; Wang, J.-P.; Lu, H. Dopaminergic neuron injury in Parkinson’s disease is mitigated by interfering lncRNA SNHG14 expression to regulate the miR-133b/ α-synuclein pathway. Aging 2019, 11, 9264–9279. [Google Scholar] [CrossRef]
- Zhang, J.; Liu, W.; Wang, Y.; Zhao, S.; Chang, N. miR-135b Plays a Neuroprotective Role by Targeting GSK3beta in MPP(+)-Intoxicated SH-SY5Y Cells. Dis. Markers 2017, 2017, 5806146. [Google Scholar] [CrossRef] [PubMed]
- Minones-Moyano, E.; Porta, S.; Escaramis, G.; Rabionet, R.; Iraola, S.; Kagerbauer, B.; Espinosa-Parrilla, Y.; Ferrer, I.; Estivill, X.; Marti, E. MicroRNA profiling of Parkinson’s disease brains identifies early downregulation of miR-34b/c which modulate mitochondrial function. Hum. Mol. Genet. 2011, 20, 3067–3078. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Le, T.; Wei, R.; Jiao, Y. Knockdown of JMJD1C, a target gene of hsa-miR-590-3p, inhibits mitochondrial dysfunction and oxidative stress in MPP+-treated MES23.5 and SH-SY5Y cells. Cell. Mol. Biol. 2016, 62, 39–45. [Google Scholar]
- Chaudhuri, A.D.; Choi, D.C.; Kabaria, S.; Tran, A.; Junn, E. MicroRNA-7 Regulates the Function of Mitochondrial Permeability Transition Pore by Targeting VDAC1 Expression. J. Biol. Chem. 2016, 291, 6483–6493. [Google Scholar] [CrossRef] [PubMed]
- Kabaria, S.; Choi, D.C.; Chaudhuri, A.D.; Mouradian, M.M.; Junn, E. Inhibition of miR-34b and miR-34c enhances α-synuclein expression in Parkinson’s disease. FEBS Lett. 2014, 589, 319–325. [Google Scholar] [CrossRef]
- Chen, Y.; Lian, Y.-J.; Ma, Y.-Q.; Wu, C.-J.; Zheng, Y.-K.; Xie, N.-C. LncRNA SNHG1 promotes α-synuclein aggregation and toxicity by targeting miR-15b-5p to activate SIAH1 in human neuroblastoma SH-SY5Y cells. NeuroToxicology 2018, 68, 212–221. [Google Scholar] [CrossRef]
- Wang, Y.; Cai, Y.; Huang, H.; Chen, X.; Chen, X.; Chen, X.; Mai, H.; Li, X.; Zhao, J.; Yang, J.; et al. miR-486-3p Influences the Neurotoxicity of a-Synuclein by Targeting the SIRT2 Gene and the Polymorphisms at Target Sites Contributing to Parkinson’s Disease. Cell. Physiol. Biochem. 2018, 51, 2732–2745. [Google Scholar] [CrossRef]
- Liang, H.; Ding, B.; Liang, J.; Shi, X.; Jiang, X.; Gao, Y. MicroRNA-10a inhibits A30P α-synuclein aggregation and toxicity by targeting proapoptotic protein BCL2L11. Int. J. Clin. Exp. Pathol. 2018, 11, 624–633. [Google Scholar]
- Li, G.; Yang, H.; Zhu, D.; Huang, H.; Liu, G.; Lun, P. Targeted Suppression of Chaperone-Mediated Autophagy by miR-320a Promotes α-Synuclein Aggregation. Int. J. Mol. Sci. 2014, 15, 15845–15857. [Google Scholar] [CrossRef]
- Zhang, Z.; Cheng, Y. miR-16-1 Promotes the Aberrant α-Synuclein Accumulation in Parkinson Disease via Targeting Heat Shock Protein 70. Sci. World J. 2014, 2014, 938348. [Google Scholar] [CrossRef]
- Peng, T.; Liu, X.; Wang, J.; Liu, Y.; Fu, Z.; Ma, X.; Li, J.; Sun, G.; Ji, Y.; Lu, J.; et al. Long noncoding RNA HAGLROS regulates apoptosis and autophagy in Parkinson’s disease via regulating miR-100/ATG10 axis and PI3K/Akt/mTOR pathway activation. Artif. Cells Nanomed. Biotechnol. 2019, 47, 2764–2774. [Google Scholar] [CrossRef]
- Wen, Z.; Zhang, J.; Tang, P.; Tu, N.; Wang, K.; Wu, G. Overexpression of miR185 inhibits autophagy and apoptosis of dopaminergic neurons by regulating the AMPK/mTOR signaling pathway in Parkinson’s disease. Mol. Med. Rep. 2018, 17, 131–137. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.H.; Wang, Y.B.; Yang, J.; Liu, H.Q.; Wang, L.L. MicroRNA-326 suppresses iNOS expression and promotes autophagy of dopaminergic neurons through the JNK signaling by targeting XBP1 in a mouse model of Parkinson’s disease. J. Cell. Biochem. 2019, 120, 14995–15006. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Fiesel, F.C.; Belmonte, K.C.; Hudec, R.; Wang, W.-X.; Kim, C.; Nelson, P.T.; Springer, W.; Kim, J. miR-27a and miR-27b regulate autophagic clearance of damaged mitochondria by targeting PTEN-induced putative kinase 1 (PINK1). Mol. Neurodegener. 2016, 11, 55. [Google Scholar] [CrossRef]
- Zhou, T.; Lin, D.; Chen, Y.; Peng, S.; Jing, X.; Lei, M.; Enxiang, T.; Liang, Y. α-synuclein accumulation in SH-SY5Y cell impairs autophagy in microglia by exosomes overloading miR-19a-3p. Epigenomics 2019, 11, 1661–1677. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Song, Y.; Zhu, X. MicroRNA-181a Regulates Apoptosis and Autophagy Process in Parkinson’s Disease by Inhibiting p38 Mitogen-Activated Protein Kinase (MAPK)/c-Jun N-Terminal Kinases (JNK) Signaling Pathways. Med. Sci. Monit. 2017, 23, 1597–1606. [Google Scholar] [CrossRef] [PubMed]
- Kumar, L.; Nazir, A. Modulation of Alpha-synuclein Expression and Associated Effects by MicroRNA Let-7 in Transgenic C. elegans. Front. Mol. Neurosci. 2017, 10, 328. [Google Scholar] [CrossRef]
- Jauhari, A.; Singh, T.; Mishra, S.; Shankar, J.; Yadav, S. Coordinated Action of miR-146a and Parkin Gene Regulate Rotenone-induced Neurodegeneration. Toxicol. Sci. 2020, 176, 433–445. [Google Scholar] [CrossRef]
- Chiu, C.-C.; Yeh, T.-H.; Chen, R.-S.; Chen, H.-C.; Huang, Y.-Z.; Weng, Y.-H.; Cheng, Y.-C.; Liu, Y.-C.; Cheng, A.-J.; Lu, Y.-C.; et al. Upregulated Expression of MicroRNA-204-5p Leads to the Death of Dopaminergic Cells by Targeting DYRK1A-Mediated Apoptotic Signaling Cascade. Front. Cell. Neurosci. 2019, 13, 399. [Google Scholar] [CrossRef]
- Xu, L.; Jia, Y.; Yang, X.-H.; Han, F.; Zheng, Y.; Ni, Y.; Chen, X.; Hong, J.; Liu, J.-Q.; Li, Q.; et al. MicroRNA-130b transcriptionally regulated by histone H3 deacetylation renders Akt ubiquitination and apoptosis resistance to 6-OHDA. Biochim. Biophys. Acta (BBA) Mol. Basis Dis. 2017, 1863, 1678–1689. [Google Scholar] [CrossRef]
- Tao, H.; Liu, Y.; Hou, Y. miRNA3845p regulates the progression of Parkinson’s disease by targeting SIRT1 in mice and SHSY5Y cell. Int. J. Mol. Med. 2020, 45, 441–450. [Google Scholar] [PubMed]
- Bauer, M.; Kinkl, N.; Meixner, A.; Kremmer, E.; Riemenschneider, M.; Forstl, H.; Gasser, T.; Ueffing, M. Prevention of interferon-stimulated gene expression using microRNA-designed hairpins. Gene. Ther. 2009, 16, 142–147. [Google Scholar] [CrossRef] [PubMed]
- Sibley, C.R.; Seow, Y.; Curtis, H.J.; Weinberg, M.S.; Wood, M. Silencing of Parkinson’s disease-associated genes with artificial mirtron mimics of miR-1224. Nucleic Acids Res. 2012, 40, 9863–9875. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Khodr, C.E.; Sapru, M.K.; Pedapati, J.; Bohn, M.C. A microRNA embedded AAV alpha-synuclein gene silencing vector for dopaminergic neurons. Brain Res. 2011, 1386, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Poole, E.; Kuan, W.L.; Barker, R.; Sinclair, J. The human cytomegalovirus non-coding Beta2.7 RNA as a novel therapeutic for Parkinson’s disease—Translational research with no translation. Virus Res. 2016, 212, 64–69. [Google Scholar] [CrossRef] [PubMed]
- Saraiva, C.; Paiva, J.; Santos, T.; Ferreira, L.; Bernardino, L. MicroRNA-124 loaded nanoparticles enhance brain repair in Parkinson’s disease. J. Control. Release 2016, 235, 291–305. [Google Scholar] [CrossRef]
- Dong, R.F.; Zhang, B.; Tai, L.W.; Liu, H.M.; Shi, F.K.; Liu, N.N. The Neuroprotective Role of MiR-124-3p in a 6-Hydroxydopamine-Induced Cell Model of Parkinson’s Disease via the Regulation of ANAX5. J. Cell. Biochem. 2018, 119, 269–277. [Google Scholar] [CrossRef]
- Andersen, R.E.; Lim, D.A. Forging our understanding of lncRNAs in the brain. Cell Tissue Res. 2017, 371, 55–71. [Google Scholar] [CrossRef]
- Shi, C.; Zhang, L.; Qin, C. Long non-coding RNAs in brain development, synaptic biology, and Alzheimer’s disease. Brain Res. Bull. 2017, 132, 160–169. [Google Scholar] [CrossRef]
- Majidinia, M.; Mihanfar, A.; Rahbarghazi, R.; Nourazarian, A.; Bagca, B.; Avci, C.B. The roles of non-coding RNAs in Parkinson’s disease. Mol. Biol. Rep. 2016, 43, 1193–1204. [Google Scholar] [CrossRef]
- Salta, E.; De Strooper, B. Noncoding RNAs in neurodegeneration. Nat. Rev. Neurosci. 2017, 18, 627–640. [Google Scholar] [CrossRef]
- Devaux, Y.; Creemers, E.E.; Boon, R.A.; Werfel, S.; Thum, T.; Engelhardt, S.; Dimmeler, S.; Squire, I.; Network, O.B.O.T.C. Circular RNAs in heart failure. Eur. J. Hear. Fail. 2017, 19, 701–709. [Google Scholar] [CrossRef] [PubMed]
- Gomes, C.P.D.C.; Schroen, B.; Kuster, G.M.; Robinson, E.L.; Ford, K.; Squire, I.B.; Heymans, S.; Martelli, F.; Emanueli, C.; Devaux, Y.; et al. Regulatory RNAs in Heart Failure. Circulation 2020, 141, 313–328. [Google Scholar] [CrossRef]
- Mortezaei, Z.; Lanjanian, H.; Masoudi-Nejad, A. Candidate novel long noncoding RNAs, MicroRNAs and putative drugs for Parkinson’s disease using a robust and efficient genome-wide association study. Genomics 2017, 109, 158–164. [Google Scholar] [CrossRef]
- Kraus, T.F.J.; Haider, M.; Spanner, J.; Steinmaurer, M.; Dietinger, V.; Kretzschmar, H.A. Altered Long Noncoding RNA Expression Precedes the Course of Parkinson’s Disease—A Preliminary Report. Mol. Neurobiol. 2016, 54, 2869–2877. [Google Scholar] [CrossRef] [PubMed]
- Simchovitz, A.; Hanan, M.; Niederhoffer, N.; Madrer, N.; Yayon, N.; Bennett, E.R.; Greenberg, D.S.; Kadener, S.; Soreq, H. NEAT1 is overexpressed in Parkinson’s disease substantia nigra and confers drug-inducible neuroprotection from oxidative stress. FASEB J. 2019, 33, 11223–11234. [Google Scholar] [CrossRef] [PubMed]
- Boros, F.A.; Maszlag-Török, R.; Vécsei, L.; Klivényi, P. Increased level of NEAT1 long non-coding RNA is detectable in peripheral blood cells of patients with Parkinson’s disease. Brain Res. 2020, 1730, 146672. [Google Scholar] [CrossRef] [PubMed]
- Ni, Y.; Huang, H.; Chen, Y.; Cao, M.; Zhou, H.; Zhang, Y. Investigation of Long Non-coding RNA Expression Profiles in the Substantia Nigra of Parkinson’s Disease. Cell. Mol. Neurobiol. 2016, 37, 329–338. [Google Scholar] [CrossRef] [PubMed]
- Soreq, L.; Guffanti, A.; Salomonis, N.; Simchovitz, A.; Israel, Z.; Bergman, H.; Soreq, H. Long non-coding RNA and alternative splicing modulations in Parkinson’s leukocytes identified by RNA sequencing. PLoS Comput. Biol. 2014, 10, e1003517. [Google Scholar] [CrossRef]
- Fan, Y.; Li, J.; Yang, Q.; Gong, C.; Gao, H.; Mao, Z.; Yuan, X.; Zhu, S.; Xue, Z. Dysregulated Long Non-coding RNAs in Parkinson’s Disease Contribute to the Apoptosis of Human Neuroblastoma Cells. Front. Mol. Neurosci. 2019, 13, 1320. [Google Scholar] [CrossRef]
- Fan, Y.; Xue, Z. Plasma-based circulating long non-coding RNA AS-Uch1 as a novel diagnostic biomarker for Parkinson’s disease. In Proceedings of the 2019 International Congress, Nice, France, 22–26 September 2019. [Google Scholar]
- Zhao, J.; Geng, L.; Chen, Y.; Wu, C. SNHG1 promotes MPP+-induced cytotoxicity by regulating PTEN/AKT/mTOR signaling pathway in SH-SY5Y cells via sponging miR-153-3p. Biol. Res. 2020, 53, 1. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Wang, J.; Liu, Q.; Xiao, Z.; Dai, Q. Knockdown of long non-coding RNA AL049437 mitigates MPP+ -induced neuronal injury in SH-SY5Y cells via the microRNA-205-5p/MAPK1 axis. NeuroToxicology 2020, 78, 29–35. [Google Scholar] [CrossRef]
- Liu, S.; Cui, B.; Dai, Z.-X.; Shi, P.-K.; Wang, Z.-H.; Guo, Y.-Y. Long Non-coding RNA HOTAIR Promotes Parkinson’s Disease Induced by MPTP through up-regulating the Expression of LRRK2. Curr. Neurovascular Res. 2016, 13, 115–120. [Google Scholar] [CrossRef] [PubMed]
- Coupland, K.; Kim, W.S.; Halliday, G.M.; Hallupp, M.; Dobson-Stone, C.; Kwok, J.B. Role of the Long Non-Coding RNA MAPT-AS1 in Regulation of Microtubule Associated Protein Tau (MAPT) Expression in Parkinson’s Disease. PLoS ONE 2016, 11, e0157924. [Google Scholar] [CrossRef] [PubMed]
- Lin, D.; Liang, Y.; Jing, X.; Chen, Y.; Lei, M.; Zeng, Z.; Zhou, T.; Wu, X.; Peng, S.; Zheng, D.; et al. Microarray analysis of an synthetic α-synuclein induced cellular model reveals the expression profile of long non-coding RNA in Parkinson’s disease. Brain Res. 2018, 1678, 384–396. [Google Scholar] [CrossRef] [PubMed]
- Xie, S.-P.; Zhou, F.; Li, J.; Duan, S.-J. NEAT1 regulates MPP+-induced neuronal injury by targeting miR-124 in neuroblastoma cells. Neurosci. Lett. 2019, 708, 134340. [Google Scholar] [CrossRef]
- Yan, W.; Chen, Z.Y.; Chen, J.Q.; Chen, H.M. LncRNA NEAT1 promotes autophagy in MPTP-induced Parkinson’s disease through stabilizing PINK1 protein. Biochem. Biophys. Res. Commun. 2018, 496, 1019–1024. [Google Scholar] [CrossRef]
- Liu, W.; Zhang, Q.; Zhang, J.; Pan, W.; Zhao, J.; Xu, Y. Long non-coding RNA MALAT1 contributes to cell apoptosis by sponging miR-124 in Parkinson disease. Cell Biosci. 2017, 7, 19. [Google Scholar] [CrossRef]
- Lu, Y.; Gong, Z.; Jin, X.; Zhao, P.; Zhang, Y.; Wang, Z. LncRNA MALAT1 targeting miR-124-3p regulates DAPK1 expression contributes to cell apoptosis in Parkinson’s Disease. J. Cell. Biochem. 2020. [Google Scholar] [CrossRef]
- Li, Y.; Fang, J.; Zhou, Z.; Zhou, Q.; Sun, S.; Jin, Z.; Xi, Z.; Wei, J. Downregulation of lncRNA BACE1-AS improves dopamine-dependent oxidative stress in rats with Parkinson’s disease by upregulating microRNA-34b-5p and downregulating BACE1. Cell Cycle 2020, 19, 1158–1171. [Google Scholar] [CrossRef]
- Carrieri, C.; Forrest, A.R.; Santoro, C.; Persichetti, F.; Carninci, P.; Zucchelli, S.; Gustincich, S. Expression analysis of the long non-coding RNA antisense to Uchl1 (AS Uchl1) during dopaminergic cells’ differentiation in vitro and in neurochemical models of Parkinson’s disease. Front. Cell. Neurosci. 2015, 9, 114. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Piao, X.; Hu, S.; Gao, J.; Bao, M. LncRNA H19 diminishes dopaminergic neuron loss by mediating microRNA-301b-3p in Parkinson’s disease via the HPRT1-mediated Wnt/beta-catenin signaling pathway. Aging (Albany NY) 2020, 12, 8820–8836. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Xia, Q.; Lin, J. LncRNA H19 Attenuates Apoptosis in MPTP-Induced Parkinson’s Disease through Regulating miR-585-3p/PIK3R3. Neurochem. Res. 2020, 45, 1700–1710. [Google Scholar] [CrossRef] [PubMed]
- Lin, Q.; Hou, S.; Dai, Y.; Jiang, N.; Lin, Y. LncRNA HOTAIR targets miR-126-5p to promote the progression of Parkinson’s disease through RAB3IP. Biol. Chem. 2019, 400, 1217–1228. [Google Scholar] [CrossRef]
- Cai, L.; Tu, L.; Li, T.; Yang, X.; Ren, Y.; Gu, R.; Zhang, Q.; Yao, H.; Qu, X.; Wang, Q.; et al. Downregulation of lncRNA UCA1 ameliorates the damage of dopaminergic neurons, reduces oxidative stress and inflammation in Parkinson’s disease through the inhibition of the PI3K/Akt signaling pathway. Int. Immunopharmacol. 2019, 75, 105734. [Google Scholar] [CrossRef]
- Yang, H.; Lin, Q.; Chen, N.; Luo, Z.; Zheng, C.; Li, J.; Zheng, F.; Guo, Z.; Cai, P.; Wu, S.; et al. LncRNA NR_030777 Alleviates Paraquat-induced Neurotoxicity by Regulating Zfp326 and Cpne5. Toxicol. Sci. 2020. [Google Scholar] [CrossRef]
- Simchovitz, A.; Hanan, M.; Yayon, N.; Lee, S.; Bennett, E.R.; Greenberg, D.S.; Kadener, S.; Soreq, H. A lncRNA survey finds increases in neuroprotective LINC-PINT in Parkinson’s disease substantia nigra. Aging Cell 2020, 19, e13115. [Google Scholar] [CrossRef]
- Lu, M.; Sun, W.-L.; Shen, J.; Wei, M.; Chen, B.; Qi, Y.-J.; Xu, C.-S. LncRNA-UCA1 promotes PD development by upregulating SNCA. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 7908–7915. [Google Scholar]
- Qian, C.; Ye, Y.; Mao, H.; Yao, L.; Sun, X.; Wang, B.; Zhang, H.; Xie, L.; Zhang, H.; Zhang, Y.; et al. Downregulated lncRNA-SNHG1 enhances autophagy and prevents cell death through the miR-221/222 /p27/mTOR pathway in Parkinson’s disease. Exp. Cell Res. 2019, 384, 111614. [Google Scholar] [CrossRef]
- Xu, W.; Zhang, L.; Geng, Y.; Liu, Y.; Zhang, N. Long noncoding RNA GAS5 promotes microglial inflammatory response in Parkinson’s disease by regulating NLRP3 pathway through sponging miR-223-3p. Int. Immunopharmacol. 2020, 85, 106614. [Google Scholar] [CrossRef]
- Liu, R.; Li, F.; Zhao, W. Long noncoding RNA NEAT1 knockdown inhibits MPP(+)-induced apoptosis, in fl ammation and cytotoxicity in SK-N-SH cells by regulating miR-212-5p/RAB3IP axis. Neurosci. Lett. 2020, 731, 135060. [Google Scholar] [CrossRef] [PubMed]
- Song, Q.; Geng, Y.; Li, Y.; Wang, L.; Qin, J. Long noncoding RNA NORAD regulates MPP+-induced Parkinson’s disease model cells. J. Chem. Neuroanat. 2019, 101, 101668. [Google Scholar] [CrossRef] [PubMed]
- D’Ambra, E.; Capauto, D.; Morlando, M. Exploring the Regulatory Role of Circular RNAs in Neurodegenerative Disorders. Int. J. Mol. Sci. 2019, 20, 5477. [Google Scholar] [CrossRef] [PubMed]
- Lukiw, W.J. Circular RNA (circRNA) in Alzheimer’s disease (AD). Front. Genet. 2013, 4, 307. [Google Scholar] [CrossRef] [PubMed]
- Choi, D.C.; Chae, Y.-J.; Kabaria, S.; Chaudhuri, A.D.; Jain, M.R.; Li, H.; Mouradian, M.M.; Junn, E. MicroRNA-7 protects against 1-methyl-4-phenylpyridinium-induced cell death by targeting RelA. J. Neurosci. 2014, 34, 12725–12737. [Google Scholar] [CrossRef] [PubMed]
- Sang, Q.; Liu, X.; Wang, L.; Qi, L.; Sun, W.; Wang, W.; Sun, Y.; Zhang, H. CircSNCA downregulation by pramipexole treatment mediates cell apoptosis and autophagy in Parkinson’s disease by targeting miR-7. Aging 2018, 10, 1281–1293. [Google Scholar] [CrossRef]
- Feng, Z.; Zhang, L.; Wang, S.; Hong, Q. Circular RNA circDLGAP4 exerts neuroprotective effects via modulating miR-134-5p/CREB pathway in Parkinson’s disease. Biochem. Biophys. Res. Commun. 2020, 522, 388–394. [Google Scholar] [CrossRef]
- Kumar, L.; Jadiya, P.; Haque, R.; Shukla, S.; Nazir, A. Functional Characterization of Novel Circular RNA Molecule, circzip-2 and Its Synthesizing Gene zip-2 in C. elegans Model of Parkinson’s Disease. Mol. Neurobiol. 2018, 55, 6914–6926. [Google Scholar] [CrossRef]
- Jia, E.; Zhou, Y.; Liu, Z.; Wang, L.; Ouyang, T.; Pan, M.; Bai, Y.; Ge, Q. Transcriptomic Profiling of Circular RNA in Different Brain Regions of Parkinson’s Disease in a Mouse Model. Int. J. Mol. Sci. 2020, 21, 3006. [Google Scholar] [CrossRef]
- Hanan, M.; Simchovitz, A.; Yayon, N.; Vaknine, S.; Cohen-Fultheim, R.; Karmon, M.; Madrer, N.; Rohrlich, T.M.; Maman, M.; Bennett, E.R.; et al. A Parkinson’s disease Circ RNA s Resource reveals a link between circ SLC 8A1 and oxidative stress. EMBO Mol. Med. 2020, 11942. [Google Scholar] [CrossRef]
- Pekarsky, Y.; Balatti, V.; Palamarchuk, A.; Rizzotto, L.; Veneziano, D.; Nigita, G.; Rassenti, L.Z.; Pass, H.; Kipps, T.J.; Liu, C.-G.; et al. Dysregulation of a family of short noncoding RNAs, tsRNAs, in human cancer. Proc. Natl. Acad. Sci. USA 2016, 113, 5071–5076. [Google Scholar] [CrossRef] [PubMed]
- Balatti, V.; Nigita, G.; Veneziano, D.; Drusco, A.; Stein, G.S.; Messier, T.L.; Farina, N.H.; Lian, J.B.; Tomasello, L.; Liu, C.-G.; et al. tsRNA signatures in cancer. Proc. Natl. Acad. Sci. USA 2017, 114, 8071–8076. [Google Scholar] [CrossRef] [PubMed]
- Blanco, S.; Dietmann, S.; Flores, J.V.; Hussain, S.; Kutter, C.; Humphreys, P.; Lukk, M.; Lombard, P.; Treps, L.; Popis, M.; et al. Aberrant methylation of t RNA s links cellular stress to neuro-developmental disorders. EMBO J. 2014, 33, 2020–2039. [Google Scholar] [CrossRef]
- Van Es, M.A.; Schelhaas, H.J.; Van Vught, P.W.J.; Ticozzi, N.; Andersen, P.M.; Groen, E.J.N.; Schulte, C.; Blauw, H.M.; Koppers, M.; Diekstra, F.P.; et al. Angiogenin variants in Parkinson disease and amyotrophic lateral sclerosis. Ann. Neurol. 2011, 70, 964–973. [Google Scholar] [CrossRef] [PubMed]
- Magee, R.; Londin, E.; Rigoutsos, I. TRNA-derived fragments as sex-dependent circulating candidate biomarkers for Parkinson’s disease. Parkinsonism Relat. Disord. 2019, 65, 203–209. [Google Scholar] [CrossRef]
- Hong, C.T.; Hu, H.H.; Chan, L.; Bai, C.-H. Prevalent cerebrovascular and cardiovascular disease in people with Parkinson’s disease: A meta-analysis. Clin. Epidemiol. 2018, 10, 1147–1154. [Google Scholar] [CrossRef]
- Muqtadar, H.; Testai, F.D.; Gorelick, P.B. The Dementia of Cardiac Disease. Curr. Cardiol. Rep. 2012, 14, 732–740. [Google Scholar] [CrossRef]
- Chen, H.; Zhang, S.M.; Schwarzschild, M.A.; Hernan, M.A.; Willett, W.C.; Ascherio, A. Obesity and the risk of Parkinson’s disease. Am. J. Epidemiol. 2004, 159, 547–555. [Google Scholar] [CrossRef]
- Simon, K.C.; Chen, H.; Schwarzschild, M.; Ascherio, A. Hypertension, hypercholesterolemia, diabetes, and risk of Parkinson disease. Neurology 2007, 69, 1688–1695. [Google Scholar] [CrossRef]
- Dölle, C.; Flønes, I.; Nido, G.S.; Miletic, H.; Osuagwu, N.; Kristoffersen, S.; Lilleng, P.K.; Larsen, J.P.; Tysnes, O.-B.; Haugarvoll, K.; et al. Defective mitochondrial DNA homeostasis in the substantia nigra in Parkinson disease. Nat. Commun. 2016, 7, 13548. [Google Scholar] [CrossRef]
- Yu, E.P.K.; Bennett, M.R. The role of mitochondrial DNA damage in the development of atherosclerosis. Free Radic. Biol. Med. 2016, 100, 223–230. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.-P.; Chen, L.-S.; Yen, M.-F.; Fann, C.-Y.; Chiu, Y.-H.; Chen, H.-H.; Pan, S.-L. Parkinson’s Disease Is Related to an Increased Risk of Ischemic Stroke—A Population-Based Propensity Score-Matched Follow-up Study. PLoS ONE 2013, 8, e68314. [Google Scholar] [CrossRef]
- Liang, H.-W.; Huang, Y.-P.; Pan, S.-L. Parkinson disease and risk of acute myocardial infarction: A population-based, propensity score–matched, longitudinal follow-up study. Am. Hear. J. 2015, 169, 508–514. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.D.; Malaty, I.; Price, C.C.; Okun, M.; Bowers, D. Health comorbidities and cognition in 1948 patients with idiopathic Parkinson’s disease. Park. Relat. Disord. 2012, 18, 1073–1078. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zeng, F.; Jin, W.S.; Zhu, C.; Wang, Q.H.; Bu, X.L.; Luo, H.B.; Zou, H.Q.; Pu, J.; Zhou, Z.H.; et al. Comorbidity burden of patients with Parkinson’s disease and Parkinsonism between 2003 and 2012: A multicentre, nationwide, retrospective study in China. Sci. Rep. 2017, 7, 1671. [Google Scholar] [CrossRef]
- Peralta, C.; Wolf, E.; Alber, H.; Seppi, K.; Muller, S.; Bosch, S.; Wenning, G.K.; Pachinger, O.; Poewe, W. Valvular heart disease in Parkinson’s disease vs. controls: An echocardiographic study. Mov. Disord. 2006, 21, 1109–1113. [Google Scholar] [CrossRef] [PubMed]
- Van Camp, G.; Flamez, A.; Cosyns, B.; Weytjens, C.; Muyldermans, L.; Van Zandijcke, M.; De Sutter, J.; Santens, P.; Decoodt, P.; Moerman, C.; et al. Treatment of Parkinson’s disease with pergolide and relation to restrictive valvular heart disease. Lancet 2004, 363, 1179–1183. [Google Scholar] [CrossRef]
- Whitsett, T.L.; Goldberg, L.I. Effects of levodopa on systolic preejection period, blood pressure, and heart rate during acute and chronic treatment of Parkinson’s disease. Circulation 1972, 45, 97–106. [Google Scholar] [CrossRef]
- Noack, C.; Schroeder, C.; Heusser, K.; Lipp, A. Cardiovascular effects of levodopa in Parkinson’s disease. Parkinsonism Relat. Disord. 2014, 20, 815–818. [Google Scholar] [CrossRef]
- Apraxine, M.; Pasquet, A.; Jeanjean, A.P. Pramipexole-Induced Reversible Heart Failure. Mov. Disord. Clin. Pract. 2014, 1, 381–382. [Google Scholar] [CrossRef]
- Galassi, G.; Georgoulopoulou, E.; Ariatti, A. Amiodarone neurotoxicity: The other side of the medal. Open Med. 2014, 9, 437–442. [Google Scholar] [CrossRef][Green Version]
- Ishida, S.; Sugino, M.; Hosokawa, T.; Sato, T.; Furutama, D.; Fukuda, A.; Kimura, F.; Kuwabara, H.; Shibayama, Y.; Hanafusa, T. Amiodarone-induced liver cirrhosis and parkinsonism: A case report. Clin. Neuropathol. 2010, 29, 84–88. [Google Scholar] [CrossRef] [PubMed]
- Fabiani, G.; Pastro, P.C.; Froehner, C. Parkinsonism and other movement disorders in outpatients in chronic use of cinnarizine and flunarizine. Arq. Neuro-Psiquiatr. 2004, 62, 784–788. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, U.A.; Ong, S.-B.; Hausenloy, D.J.; Ong, S.-G. Parkinson’s disease proteins: Novel mitochondrial targets for cardioprotection. Pharmacol. Ther. 2015, 156, 34–43. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.-X.; Zhuang, L.-L.; Liu, J.; Jing, Y.-Y.; Sun, J.; Gong, L.; Liu, X.-Y. The role of Parkin protein in cardiac function and ventricular remodeling in myocardial infarction rats. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 5004–5013. [Google Scholar]
- Kim, J.; Inoue, K.; Ishii, J.; Vanti, W.B.; Voronov, S.V.; Murchison, E.; Hannon, G.; Abeliovich, A. A MicroRNA Feedback Circuit in Midbrain Dopamine Neurons. Science 2007, 317, 1220–1224. [Google Scholar] [CrossRef]
- Boštjančič, E.; Brandner, T.; Zidar, N.; Glavač, D.; Štajer, D. Down-regulation of miR-133a/b in patients with myocardial infarction correlates with the presence of ventricular fibrillation. Biomed. Pharmacother. 2018, 99, 65–71. [Google Scholar] [CrossRef]
- Yu, Y.; Liu, H.; Yang, D.; He, F.; Yuan, Y.; Guo, J.; Hu, J.; Yu, J.; Yan, X.; Wang, S.; et al. Aloe-emodin attenuates myocardial infarction and apoptosis via up-regulating miR-133 expression. Pharmacol. Res. 2019, 146, 104315. [Google Scholar] [CrossRef]
- Liu, Y.; Li, Y.; Ni, J.; Shu, Y.; Wang, H.; Hu, T. MiR-124 attenuates doxorubicin-induced cardiac injury via inhibiting p66Shc-mediated oxidative stress. Biochem. Biophys. Res. Commun. 2019, 521, 420–426. [Google Scholar] [CrossRef]
- Zhao, Y.; Yan, M.; Chen, C.; Gong, W.; Yin, Z.; Li, H.; Fan, J.; Zhang, X.A.; Wang, D.W.; Zuo, H. MiR-124 aggravates failing hearts by suppressing CD151-facilitated angiogenesis in heart. Oncotarget 2018, 9, 14382–14396. [Google Scholar] [CrossRef]
- Geng, L.; Liu, W.; Chen, Y. miR-124-3p attenuates MPP+-induced neuronal injury by targeting STAT3 in SH-SY5Y cells. Exp. Biol. Med. 2017, 242, 1757–1764. [Google Scholar] [CrossRef] [PubMed]
- Yao, L.; Zhu, Z.; Wu, J.; Zhang, Y.; Zhang, H.; Sun, X.; Qian, C.; Wang, B.; Xie, L.; Zhang, S.; et al. MicroRNA-124 regulates the expression of p62/p38 and promotes autophagy in the inflammatory pathogenesis of Parkinson’s disease. FASEB J. 2019, 33, 8648–8665. [Google Scholar] [CrossRef]
- Gong, X.; Zhu, Y.; Chang, H.; Li, Y.; Ma, F. Long noncoding RNA MALAT1 promotes cardiomyocyte apoptosis after myocardial infarction via targeting miR-144-3p. Biosci. Rep. 2019, 39. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Zhang, L.; Song, J.; Wang, Z.; Huang, X.; Guo, Z.; Chen, F.; Zhao, X.-X. Long noncoding RNA MALAT1 mediates cardiac fibrosis in experimental postinfarct myocardium mice model. J. Cell. Physiol. 2018, 234, 2997–3006. [Google Scholar] [CrossRef]
- Zhang, Q.-S.; Wang, Z.-H.; Zhang, J.-L.; Duan, Y.-L.; Li, G.-F.; Zheng, D.-L. Beta-asarone protects against MPTP-induced Parkinson’s disease via regulating long non-coding RNA MALAT1 and inhibiting α-synuclein protein expression. Biomed. Pharmacother. 2016, 83, 153–159. [Google Scholar] [CrossRef]
- Chen, Q.; Huang, X.; Li, R. lncRNA MALAT1/miR-205-5p axis regulates MPP(+)-induced cell apoptosis in MN9D cells by directly targeting LRRK2. Am. J. Transl. Res. 2018, 10, 563–572. [Google Scholar] [PubMed]
- Gao, L.; Liu, Y.; Guo, S.; Yao, R.; Wu, L.; Xiao, L.; Wang, Z.; Liu, Y.; Zhang, Y. Circulating Long Noncoding RNA HOTAIR is an Essential Mediator of Acute Myocardial Infarction. Cell. Physiol. Biochem. 2017, 44, 1497–1508. [Google Scholar]
- Jiang, Y.; Mo, H.; Luo, J.; Zhao, S.; Liang, S.; Zhang, M.; Yuan, J. HOTAIR Is a Potential Novel Biomarker in Patients with Congenital Heart Diseases. BioMed Res. Int. 2018, 2018, 2850657. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Zhang, X.; Guo, Y.; Rong, H.; Liu, T. The long noncoding RNA HOTAIR promotes Parkinson’s disease by upregulating LRRK2 expression. Oncotarget 2017, 8, 24449–24456. [Google Scholar] [CrossRef]
- Zhou, F.; Xie, S.; Li, J.; Duan, S. Long noncoding RNA HOTAIR promotes cell apoptosis by sponging miR-221 in Parkinson’s disease. RSC Adv. 2019, 9, 29502–29510. [Google Scholar] [CrossRef]
- Caudle, W.M.; Bammler, T.K.; Lin, Y.; Pan, S.; Zhang, J. Using ‘omics’ to define pathogenesis and biomarkers of Parkinson’s disease. Expert. Rev. Neurother. 2010, 10, 925–942. [Google Scholar] [CrossRef] [PubMed]
- Schapira, A. Mitochondrial dysfunction in neurodegenerative disorders. Biochim. Biophys. Acta (BBA) Bioenerg. 1998, 1366, 225–233. [Google Scholar] [CrossRef]
- Chang, D.; Nalls, M.A.; Hallgrimsdottir, I.B.; Hunkapiller, J.; van der Brug, M.; Cai, F.; International Parkinson’s Disease Genomics Consortium; 23andMe Research Team; Kerchner, G.A.; Ayalon, G.; et al. A meta-analysis of genome-wide association studies identifies 17 new Parkinson’s disease risk loci. Nat. Genet. 2017, 49, 1511–1516. [Google Scholar] [CrossRef] [PubMed]
- Kikuchi, T.; Morizane, A.; Doi, D.; Magotani, H.; Onoe, H.; Hayashi, T.; Mizuma, H.; Takara, S.; Takahashi, R.; Inoue, H.; et al. Human iPS cell-derived dopaminergic neurons function in a primate Parkinson’s disease model. Nature 2017, 548, 592–596. [Google Scholar] [CrossRef] [PubMed]
- Barrett, T.; Wilhite, S.E.; Ledoux, P.; Evangelista, C.; Kim, I.F.; Tomashevsky, M.; Marshall, K.A.; Phillippy, K.H.; Sherman, P.M.; Holko, M.; et al. NCBI GEO: Archive for functional genomics data sets—Update. Nucleic Acids Res. 2013, 41, D991–D995. [Google Scholar] [CrossRef]
- Hoss, A.G.; Labadorf, A.; Beach, T.G.; Latourelle, J.C.; Myers, R.H. microRNA Profiles in Parkinson’s Disease Prefrontal Cortex. Front. Aging Neurosci. 2016, 8, 36. [Google Scholar] [CrossRef]
- Schulze, M.; Sommer, A.; Plötz, S.; Farrell, M.; Winner, B.; Grosch, J.; Winkler, J.; Riemenschneider, M.J. Sporadic Parkinson’s disease derived neuronal cells show disease-specific mRNA and small RNA signatures with abundant deregulation of piRNAs. Acta Neuropathol. Commun. 2018, 6, 58. [Google Scholar] [CrossRef]
- Pantano, L.; Friedländer, M.R.; Escaramís, G.; Lizano, E.; Pallarès-Albanell, J.; Ferrer, I.; Estivill, X.; Martí, E. Specific small-RNA signatures in the amygdala at premotor and motor stages of Parkinson’s disease revealed by deep sequencing analysis. Bioinformatics 2015, 32, 673–681. [Google Scholar] [CrossRef]
- Soreq, L.; Salomonis, N.; Bronstein, M.; Greenberg, D.S.; Israel, Z.; Bergman, H.; Soreq, H. Small RNA sequencing-microarray analyses in Parkinson leukocytes reveal deep brain stimulation-induced splicing changes that classify brain region transcriptomes. Front. Mol. Neurosci. 2013, 6, 10. [Google Scholar] [CrossRef]
- Zhang, L.; Hamad, A.; Vausort, M.; Funakoshi, H.; Feldman, A.M.; Wagner, D.R.; Devaux, Y. Identification of Candidate Long Noncoding RNAs Associated with Left Ventricular Hypertrophy. Clin. Transl. Sci. 2014, 8, 100–106. [Google Scholar] [CrossRef]
- Zhang, L.; Salgado-Somoza, A.; Vausort, M.; Leszek, P.; Devaux, Y. A heart-enriched antisense long non-coding RNA regulates the balance between cardiac and skeletal muscle triadin. Biochim. Biophys. Acta (BBA) Bioenerg. 2018, 1865, 247–258. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.; Metge, F.; Dieterich, C. Specific identification and quantification of circular RNAs from sequencing data. Bioinformatics 2015, 32, 1094–1096. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Mollenhauer, B.; Zimmermann, J.; Sixel-Döring, F.; Focke, N.K.; Wicke, T.; Ebentheuer, J.; Schaumburg, M.; Lang, E.; Trautmann, E.; Zetterberg, H.; et al. Monitoring of 30 marker candidates in early Parkinson disease as progression markers. Neurology 2016, 87, 168–177. [Google Scholar] [CrossRef]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef]
- Kim, S.-Y.; Volsky, D.J. PAGE: Parametric Analysis of Gene Set Enrichment. BMC Bioinform. 2005, 6, 144. [Google Scholar] [CrossRef]
- Glaab, E.; Schneider, R. PathVar: Analysis of gene and protein expression variance in cellular pathways using microarray data. Bioinformatics 2011, 28, 446–447. [Google Scholar] [CrossRef]
- Lee, E.; Chuang, H.-Y.; Kim, J.-W.; Ideker, T.; Lee, D. Inferring Pathway Activity toward Precise Disease Classification. PLoS Comput. Biol. 2008, 4, e1000217. [Google Scholar] [CrossRef]
- Dittrich, M.T.; Klau, G.W.; Rosenwald, A.; Dandekar, T.; Müller, T. Identifying functional modules in protein-protein interaction networks: An integrated exact approach. Bioinformatics 2008, 24, i223–i231. [Google Scholar] [CrossRef]
- Kittas, A.; Delobelle, A.; Schmitt, S.; Breuhahn, K.; Guziolowski, C.; Grabe, N. Directed random walks and constraint programming reveal active pathways in hepatocyte growth factor signaling. FEBS J. 2016, 283, 350–360. [Google Scholar] [CrossRef]
- Catlett, N.L.; Bargnesi, A.J.; Ungerer, S.; Seagaran, T.; Ladd, W.; Elliston, K.O.; Pratt, D. Reverse causal reasoning: Applying qualitative causal knowledge to the interpretation of high-throughput data. BMC Bioinform. 2013, 14, 340. [Google Scholar] [CrossRef] [PubMed]
- Chindelevitch, L.; Ziemek, D.; EnayetAllah, A.; Randhawa, R.; Sidders, B.; Brockel, C.; Huang, E.S. Causal reasoning on biological networks: Interpreting transcriptional changes. Bioinformatics 2012, 28, 1114–1121. [Google Scholar] [CrossRef] [PubMed]
- Fakhry, C.T.; Choudhary, P.; Gutteridge, A.; Sidders, B.; Chen, P.; Ziemek, D.; Zarringhalam, K. Interpreting transcriptional changes using causal graphs: New methods and their practical utility on public networks. BMC Bioinform. 2016, 17, 318. [Google Scholar] [CrossRef] [PubMed]
- Jaeger, S.; Min, J.; Nigsch, F.; Camargo, M.; Hutz, J.E.; Cornett, A.; Cleaver, S.; Buckler, A.; Jenkins, J.L. Causal Network Models for Predicting Compound Targets and Driving Pathways in Cancer. J. Biomol. Screen. 2014, 19, 791–802. [Google Scholar] [CrossRef]
- Edwards, Y.J.K.; Beecham, G.W.; Scott, W.K.; Khuri, S.; Bademci, G.; Tekin, D.; Martin, E.R.; Jiang, Z.; Mash, D.C.; Ffrench-Mullen, J.; et al. Identifying Consensus Disease Pathways in Parkinson’s Disease Using an Integrative Systems Biology Approach. PLoS ONE 2011, 6, e16917. [Google Scholar] [CrossRef]
- Chandrasekaran, S.; Bonchev, D. A network view on Parkinson’s disease. Comput. Struct. Biotechnol. J. 2013, 7, e201304004. [Google Scholar] [CrossRef]
- Fujita, K.A.; Ostaszewski, M.; Matsuoka, Y.; Ghosh, S.; Glaab, E.; Trefois, C.; Crespo, I.; Perumal, T.M.; Jurkowski, W.; Antony, P.M.A.; et al. Integrating Pathways of Parkinson’s Disease in a Molecular Interaction Map. Mol. Neurobiol. 2013, 49, 88–102. [Google Scholar] [CrossRef]
- Santiago, J.A.; Potashkin, J.A. Integrative network analysis unveils convergent molecular pathways in Parkinson’s disease and diabetes. PLoS ONE 2013, 8, e83940. [Google Scholar] [CrossRef]
- Fu, L.M.; Fu, K.A. Analysis of Parkinson’s disease pathophysiology using an integrated genomics-bioinformatics approach. Pathophysiology 2015, 22, 15–29. [Google Scholar] [CrossRef]
- Wang, Q.; Zhang, Y.; Wang, M.; Song, W.M.; Shen, Q.; McKenzie, A.; Choi, I.; Zhou, X.; Pan, P.-Y.; Yue, Z.; et al. The landscape of multiscale transcriptomic networks and key regulators in Parkinson’s disease. Nat. Commun. 2019, 10, 5234. [Google Scholar] [CrossRef]
- Riley, B.E.; Gardai, S.J.; Emig-Agius, D.; Bessarabova, M.; Ivliev, A.E.; Schule, B.; Alexander, J.; Wallace, W.; Halliday, G.M.; Langston, J.W.; et al. Systems-based analyses of brain regions functionally impacted in Parkinson’s disease reveals underlying causal mechanisms. PLoS ONE 2014, 9, e102909. [Google Scholar] [CrossRef] [PubMed]
| miRNA | Regulation | No. of Samples | Sample Type | Reference |
|---|---|---|---|---|
| miR-205 miR-24 * | ↑ ↓ | 28 PD vs. 28 controls | CSF | [26] |
| miR-144-5p miR-200a-3p miR-542-5p | ↑ ↑ ↑ | 44 PD vs. 42 controls | CSF | [27] |
| miR-153* miR-409-3p * miR-10a-5p let-7g3p miR-136-3p miR-433 * miR-30b miR-1 miR-19b-3p | ↑ ↑ ↑ ↑ ↑ ↑ ↑ ↓ ↓ | 47 PD vs. 27 controls | CSF Exosomes | [28] |
| miR-4274 | ↓ | 28 PD vs. 6 controls | CSF | [29] |
| miR-626 * | ↓ | 20 PD vs. 27 controls | CSF | [30] |
| miR-24 * miR-195 miR-19b | ↑ ↑ ↓ | 109 PD vs. 40 controls | Serum Exosomes | [31] |
| miR-335-5p * miR-3613-3p miR-6865-3p | ↑ ↑ ↑ | 16 PD vs. 8 controls | Serum | [32] |
| miR-150 | ↓ | 80 PD vs. 60 controls | Serum | [33] |
| miR-29c * | ↑ | 51 PD vs. 20 controls | Serum | [34] |
| miR-146a miR-335-3 miR-335-5p * | ↓ ↓ ↓ | 85 PD vs. 40 controls | Serum | [35] |
| miR-141 miR-214 miR-146b-5p miR-193a-3p | ↓ ↓ ↓ ↓ | 169 PD vs. 180 Controls | Serum | [36] |
| miR-221 | ↓ | 138 PD vs. 112 controls | Serum | [37] |
| miR-29 | ↓ | 80 PD vs. 80 controls | Serum | [38] |
| miR-222 * miR-505 * miR-626 * | ↓ ↓ ↓ | 42 PD vs. 30 controls | Plasma | [39] |
| miR-331-5p * | ↑ | 31 PD vs. 25 controls | Plasma | [40] |
| miR-124 miR-137 | ↓ ↑ | 60 PD vs. 60 controls | Plasma | [41] |
| miR-30a-5p | ↑ | 60 PD vs. 60 controls | Plasma | [42] |
| miR-105-5p | ↑ | 319 PD vs. 273 controls | Plasma | [43] |
| miR-433 * miR-133b | ↓ ↓ | 46 PD vs. 49 controls | Plasma | [44] |
| miR-27a Let-7a Let-7f miR-142-3p miR-222 * | ↑ ↓ ↓ ↓ ↓ | 25 PD vs. 25 controls | Plasma | [45] |
| miR-331-5p miR-505 * | ↑ ↑ | 52 PD vs. 48 controls | Plasma/exosomes | [46] |
| miR-132 * | ↑ | 269 PD vs. 222 controls | Plasma | [47] |
| miR-7-5p miR-22-3p* miR-124-3p miR-136-3p * miR-139-5p miR-330-5p miR-433-3p miR-495-3p miR-132-3p miR-431-3p miR-128-3p miR-136-3p * miR-154-5p miR-323a-3p miR-338-3p miR-382-5p miR-409-3p * miR-410-3p miR-485-5p miR-22-3p * | ↑ ↑ ↑ ↑ ↑ ↑ ↑ ↑ ↑ ↑ ↓ ↓ ↓ ↓ ↓ ↓ ↓ ↓ ↓ ↓ | 99 Idiopathic PD vs. 101 controls, 27 patients with GBA mutations vs. 101 controls and 26 SNCA-A53T mutation carriers vs. 101 controls | Plasma | [48] |
| miR-29c * miR-424 miR-30e-5p | ↑ ↑ ↑ | 13 PD vs 10 non-PD | PBMC | [49] |
| miR-103a-3p miR-30b-5p miR-29a-3p | ↑ ↑ ↑ | 46 PD vs. 46 controls | PBMC | [50] |
| miR-155-5p miR-146-5p | ↑ ↓ | 37 PD vs. 43 controls | PBMC | [51] |
| miR-30c miR-26a | ↓ ↓ | 19 PD vs. 13 controls | PBMC | [52] |
| miR-885 miR-17 miR-361 | ↑ ↑ ↓ | 36 PD vs. 16 controls | PBMC | [53] |
| miR-376a | ↓ | 33 PD vs. 25 controls | PBMC | [54] |
| miR-7 miR-9-5p miR-9-3p miR-129 miR-132 * | ↑ ↑ ↑ ↑ ↑ | 20 non-medicated PD vs. 18 medicated PD | Peripheral blood lymphocytes | [55] |
| miR-223 miR-153 * | ↓ ↓ | 83 PD vs. 77 controls | Saliva | [56] |
| miR-874 miR-145-3p | ↑ ↑ | 30 PD vs. 30 controls | Saliva | [57] |
| lncRNA | Regulation | No. of Samples | Sample Type | Reference |
|---|---|---|---|---|
| lincRNA-p21 MALAT1 SNHG1 NEAT1 * H19 | ↑ ↑ ↑ ↑ ↓ | 20 PD vs. 10 controls | Human brain specimens | [115] |
| AL049437 AK021630 | ↑ ↓ | 11 PD vs. 14 controls | Tissue samples | [118] |
| U1 RP11-462G22.1 | ↓ ↓ | 3 PD vs. 3 controls | Blood Leukocytes | [119] |
| AC131056.3-001 HOTAIRM1 lnc-MOK-6:1 RF01976.1-201 | ↑ ↑ ↑ ↑ | 72 PD vs. 22 controls | Blood Leukocytes | [120] |
| NEAT1 * | ↑ | 61 PD vs. 42 controls | PBMC | [117] |
| AS-Uchl1 | ↓ | 68 PD vs. 65 controls | Plasma | [121] |
| GEO Reference | Profiling Technique | Main Observations | Sample Type | Reference |
|---|---|---|---|---|
| GSE72962 | RNA-seq | 125 miRNAs are regulated in PD A set of 29 miRNAs classifies PD | Frozen brain tissue from pre-frontal cortex of 29 PD patients and 33 controls. | [197] |
| GSE16658 | Microarray | 18 miRNAs are regulated in PD 11 miRNAs are over-represented in pathways linked to PD | PBMCs of 19 PD patients and 13 controls | [53] |
| GSE110719 | RNA-seq | 99 miRNAs are regulated in the substantia nigra of PD patients | Fibroblasts and iPSCs from 6 controls and nine PD Dopaminergic neurons from five controls and six PD. | [198] |
| GSE97285 | RNA-seq | Several groups of miRNAs are regulated in PD at different disease progression stages | Brain samples from the amygdala—14 PD and 14 controls | [199] |
| GSE40915 | RNA-seq | 16 miRNAs are regulated in blood leukocytes of PD patient’s pre-treatment 11 miRNAs are regulated after brain stimulation, among which five vary inversely to disease evolution. | Blood leukocytes from nine PD and three controls. | [200] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Acharya, S.; Salgado-Somoza, A.; Stefanizzi, F.M.; Lumley, A.I.; Zhang, L.; Glaab, E.; May, P.; Devaux, Y. Non-Coding RNAs in the Brain-Heart Axis: The Case of Parkinson’s Disease. Int. J. Mol. Sci. 2020, 21, 6513. https://doi.org/10.3390/ijms21186513
Acharya S, Salgado-Somoza A, Stefanizzi FM, Lumley AI, Zhang L, Glaab E, May P, Devaux Y. Non-Coding RNAs in the Brain-Heart Axis: The Case of Parkinson’s Disease. International Journal of Molecular Sciences. 2020; 21(18):6513. https://doi.org/10.3390/ijms21186513
Chicago/Turabian StyleAcharya, Shubhra, Antonio Salgado-Somoza, Francesca Maria Stefanizzi, Andrew I. Lumley, Lu Zhang, Enrico Glaab, Patrick May, and Yvan Devaux. 2020. "Non-Coding RNAs in the Brain-Heart Axis: The Case of Parkinson’s Disease" International Journal of Molecular Sciences 21, no. 18: 6513. https://doi.org/10.3390/ijms21186513
APA StyleAcharya, S., Salgado-Somoza, A., Stefanizzi, F. M., Lumley, A. I., Zhang, L., Glaab, E., May, P., & Devaux, Y. (2020). Non-Coding RNAs in the Brain-Heart Axis: The Case of Parkinson’s Disease. International Journal of Molecular Sciences, 21(18), 6513. https://doi.org/10.3390/ijms21186513







