The Role of Micro RNA and Long-Non-Coding RNA in Osteoporosis
Abstract
1. Overview
2. The Role of Micro RNA (miRNA) and Long-Non-Coding RNA (lncRNA) in Osteoporosis
2.1. Micro RNA (miRNA)
2.1.1. miR-9-5p
2.1.2. miR-21
2.1.3. miR-29
2.1.4. miR-30b-5p
2.1.5. miR-31
2.1.6. miR-100
2.1.7. miR-103-3p
2.1.8. miR-122-5p
2.1.9. miR-124
2.1.10. miR-133 Family
2.1.11. miR-135a-5p
2.1.12. miR-146a
2.1.13. miR-148a
2.1.14. miR-155
2.1.15. miR-182-5p
2.1.16. miR-194-5p
2.1.17. miR-200a-3p
2.1.18. miR-203a
2.1.19. miR-214-5p
2.1.20. miR-221
2.1.21. miR-223
2.1.22. miR-338 Cluster
2.1.23. miR-365
2.1.24. miR-410
2.1.25. miR-422a
2.1.26. miR-449b-5p
2.1.27. miR-503
2.1.28. miR-543
2.1.29. miR-579-3p
2.1.30. miR-874
2.1.31. miR-1297
2.1.32. miR-2861
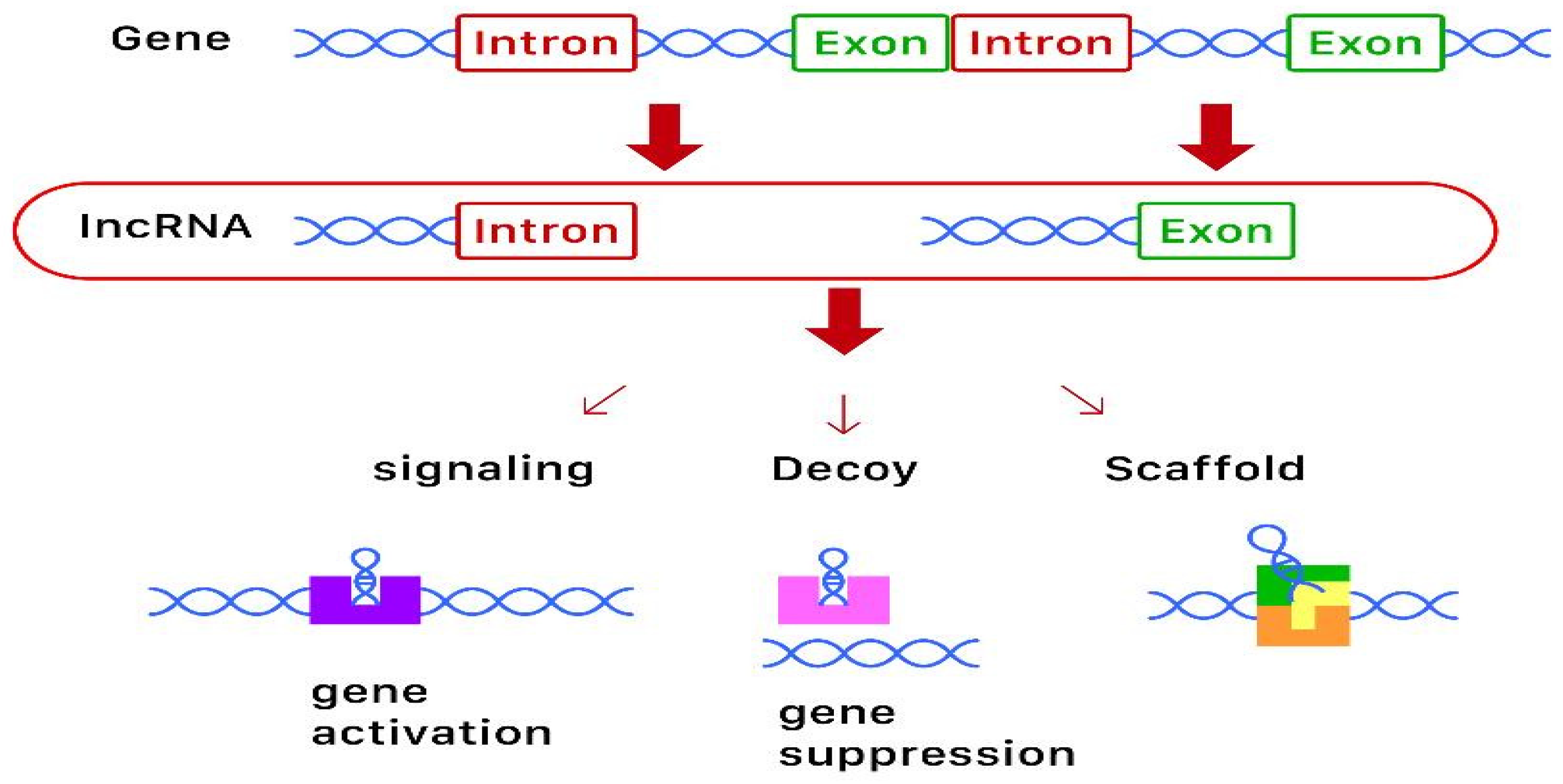
2.2. Long-Non-Coding RNA (lncRNA)
2.2.1. lncRNA-ANCR
2.2.2. lncRNA BMNCR
2.2.3. lncRNA CASC11
2.2.4. lncRNA CRNDE
2.2.5. lncRNA GAS5
2.2.6. lncRNA MALAT1
2.2.7. lncRNA MEG3
2.2.8. lncRNA MSC-AS1
2.2.9. lncRNA NEF
2.2.10. lncRNA SNHG1
2.2.11. lncRNA TUG1
2.2.12. lncRNA XIXT
3. Discussion
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| 3-UTR | 3-untranslated regions |
| ADCY6 | adenylate cyclase 6 |
| AKT | phosphorylated-protein kinase B |
| ALP | alkaline phosphatase |
| BMD | bone mineral density |
| BMSC | bone mesenchymal stem cell |
| BMP-2 | bone morphogenetic protein 2 |
| COL1A1 | collagen α-1 (I) chain |
| COL4A1 | collagen type IV α1 chain |
| CXCL12 | C-X-C motif chemokine ligand 12 |
| DElncRNAs | differentially expressed lncRNAs |
| DEmRNAs | differentially expressed mRNAs |
| ERα | estrogen receptor alpha |
| GIOP | glucocorticoid-induced osteoporosis |
| HDAC5 | histone deacetylase 5 |
| lncRNA | long non-coding RNA |
| MAPK | mitogen-activated protein kinase |
| mRNA | messenger RNA |
| miRNA | micro RNA |
| MITF | microphthalmia-associated transcription factor |
| MMP-9 | matrix metalloproteinase-9 |
| PI3K | phosphoinositide-3-kinase regulatory subunit 1 |
| PMOP | postmenopausal osteoporosis |
| RANKL | receptor activator of nuclear factor-κB ligand |
| RNA | ribonucleic acid |
| RT-qPCR | real-time quantitative polymerase chain reaction |
| RUNX2 | runt-related transcription factor 2 |
| SATB2 | special AT-rich sequence-binding protein 2 |
| SOX4 | SRY-box transcription factor 4 |
| SUFU | suppressor of fused gene |
| TGF | transforming growth factor |
| YAF-2 | YY1-associated factor 2 |
References
- Tang, P.; Xiong, Q.; Ge, W.; Zhang, L. The Role of MicroRNAs in Osteoclasts and Osteoporosis. RNA Biol. 2014, 11, 1355–1363. [Google Scholar] [CrossRef] [PubMed]
- Feng, Q.; Zheng, S.; Zheng, J. The Emerging Role of MicroRNAs in Bone Remodeling and Its Therapeutic Implications for Osteoporosis. Biosci. Rep. 2018, 38. [Google Scholar] [CrossRef]
- van Wijnen, A.; van de Peppel, J.; van Leeuwen, J.; Lian, J.; Stein, G.; Westendorf, J.; Oursler, M.; Im, H.J.; Taipaleenmäki, H.; Hesse, E.; et al. MicroRNA Functions in Osteogenesis and Dysfunctions in Osteoporosis. Cur. Osteoporos. Rep. 2013, 2, 72–82. [Google Scholar] [CrossRef] [PubMed]
- Ge, D.; Wang, W.; Chen, H.; Yang, L.; Cao, X. Functions of MicroRNAs in Osteoporosis. Eur. Rev. Med. and Pharmacol. Sci. 2017, 21, 4784–4789. [Google Scholar]
- Foessl, I.; Kotzbeck, P.; Obermayer-Pietsch, B. MiRNAs as Novel Biomarkers for Bone Related Diseases. J. Lab. Precis. Med. 2019, 4. Available online: http://jlpm.amegroups.com/article/view/4655 (accessed on 2 July 2020). [CrossRef]
- Yoon, J.H.; Kim, J.; Gorospe, M. Long noncoding RNA turnover. Biochimie 2015, 117, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Bellavia, D.; De Luca, A.; Carina, V.; Costa, V.; Raimondi, L.; Salamanna, F.; Alessandro, R.; Fini, M.; Giavaresi, G. Deregulated MiRNAs in Bone Health: Epigenetic Roles in Osteoporosis. Bone 2019, 122, 52–75. [Google Scholar] [CrossRef]
- Silva, A.; Moura, S.; Teixeira, J.; Barbosa, M.; Santos, S.; Almeida, M. Long Noncoding RNAs: A Missing Link in Osteoporosis. Bone Res. 2019, 7. [Google Scholar] [CrossRef]
- Wu, Q.; Li, X.; Miao, Z.; Ye, J.; Wang, B.; Zhang, F.; Xu, R.; Jiang, D.; Zhao, M.; Yuan, F. Long Non-Coding RNAs: A New Regulatory Code for Osteoporosis. Front. Endocrinol. 2018, 9. [Google Scholar] [CrossRef]
- Li, W.; Zhu, H.; Xu, H.; Zhang, B.; Huang, S. CRNDE Impacts the Proliferation of Osteoclast by Estrogen Deficiency in Postmenopausal Osteoporosis. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 5815–5821. [Google Scholar] [CrossRef]
- Hao, L.; Fu, J.; Tian, Y.; Wu, J. Systematic Analysis of LncRNAs, MiRNAs and MRNAs for the Identification of Biomarkers for Osteoporosis in the Mandible of Ovariectomized Mice. Int. J. Mol. Med. 2017, 40, 689–702. [Google Scholar] [CrossRef] [PubMed]
- Fei, Q.; Bai, S.; Lin, J.; Meng, H.; Yang, Y.; Guo, A. Identification of Aberrantly Expressed Long Non-Coding RNAs in Postmenopausal Osteoporosis. Int. J. Mol. Med. 2018, 41, 3537–3550. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Xu, C.; Zhu, W.; He, H.; Zhang, L.; Tang, B.; Zeng, Y.; Tian, Q.; Deng, H. Long Noncoding RNA Analyses for Osteoporosis Risk in Caucasian Women. Calcif. Tissue Int. 2019, 105, 183–192. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Wang, X.; Zhao, H.; Zhou, C. MicroRNA-9-5p Promotes Osteoporosis Development through Inhibiting Osteogenesis and Promoting Adipogenesis via Targeting Wnt3a. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 456–463. [Google Scholar] [CrossRef]
- Wang, S.; Tang, C.; Zhang, Q.; Chen, W. Reduced miR-9 and miR-181a expression down-regulates Bim concentration and promote osteoclasts survival. Int. J. Clin. Exp. Pathol. 2014, 7, 2209–2218. [Google Scholar]
- Jiang, L.; Tian, L.; Zhang, C. Bone Marrow Stem Cells-Derived Exosomes Extracted from Osteoporosis Patients Inhibit Osteogenesis via MicroRNA-21/SMAD7. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 6221–6229. [Google Scholar] [CrossRef]
- Cheng, V.; Au, P.; Tan, K.; Cheung, C. MicroRNA and Human Bone Health. JBMR Plus 2019, 3, 2–13. [Google Scholar] [CrossRef]
- Seeliger, C.; Karpinski, K.; Haug, A.; Vester, H.; Schmitt, A.; Bauer, J.; van Griensven, M. Five Freely Circulating MiRNAs and Bone Tissue MiRNAs Are Associated with Osteoporotic Fractures. J. Bone Miner. Res. 2014, 29, 1718–1728. [Google Scholar] [CrossRef]
- Yavropoulou, M.; Anastasilakis, A.; Makras, P.; Tsalikakis, D.; Grammatiki, M.; Yovos, J. Expression of MicroRNAs That Regulate Bone Turnover in the Serum of Postmenopausal Women with Low Bone Mass and Vertebral Fractures. Eur. J. Endocrinol. 2017, 176, 169–176. [Google Scholar] [CrossRef]
- Kelch, S.; Balmayor, E.R.; Seeliger, C.; Vester, H.; Kirschke, J.S.; van Griensven, M. miRNAs in bone tissue correlate to bone mineral density and circulating miRNAs are gender independent in osteoporotic patients. Sci. Rep. 2017, 7, 15861. [Google Scholar] [CrossRef]
- Li, H.; Wang, Z.; Fu, Q.; Zhang, J. Plasma miRNA levels correlate with sensitivity to bone mineral density in postmenopausal osteoporosis patients. Biomarkers. 2014, 19, 553–556. [Google Scholar] [CrossRef] [PubMed]
- Weilner, S.; Skalicky, S.; Salzer, B.; Keider, V.; Wagner, M.; Hildner, F.; Gabriel, C.; Dovjak, P.; Pietschmann, P.; Grillari-Voglauer, R.; et al. Differentially circulating miRNAs after recent osteoporotic fractures can influence osteogenic differentiation. Bone 2015, 79, 43–51. [Google Scholar] [CrossRef] [PubMed]
- Yang, N.; Wang, G.; Hu, C.; Shi, Y.; Liao, L.; Shi, S.; Cai, Y.; Cheng, S.; Wang, X.; Liu, Y.; et al. Tumor necrosis factor α suppresses the mesenchymal stem cell osteogenesis promoter miR-21 in estrogen deficiency-induced osteoporosis. J. Bone Miner. Res. 2013, 28, 559–573. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.H.; Sui, B.D.; Du, F.Y.; Shuai, Y.; Zheng, C.X.; Zhao, P.; Yu, X.R.; Jin, Y. miR-21 deficiency inhibits osteoclast function and prevents bone loss in mice. Sci. Rep. 2017, 7, 43191. [Google Scholar] [CrossRef]
- Lian, W.; Ko, J.; Chen, Y.; Ke, H.; Hsieh, C.; Kuo, C.; Wang, S.; Huang, B.; Tseng, J.; Wang, F. MicroRNA-29a Represses Osteoclast Formation and Protects against Osteoporosis by Regulating PCAF-Mediated RANKL and CXCL12. Cell Death Dis. 2019, 10, 705. [Google Scholar] [CrossRef]
- Kocijan, R.; Muschitz, C.; Geiger, E.; Skalicky, S.; Baierl, A.; Dormann, R.; Plachel, F.; Feichtinger, X.; Heimel, P.; Fahrleitner-Pammer, A.; et al. Circulating MicroRNA Signatures in Patients With Idiopathic and Postmenopausal Osteoporosis and Fragility Fractures. J. Clin. Endocrinol. Metab. 2016, 101, 4125–4134. [Google Scholar] [CrossRef]
- Li, Z.; Hassan, M.Q.; Jafferji, M.; Aqeilan, R.I.; Garzon, R.; Croce, C.M.; Van Wijnen, A.J.; Stein, J.L.; Stein, G.S.; Lian, J.B. Biological functions of miR-29b contribute to positive regulation of osteoblast differentiation. J. Biol. Chem. 2009, 284, 15676–15684. [Google Scholar] [CrossRef]
- Kapinas, K.; Kessler, C.; Ricks, T.; Gronowicz, G.; Delany, A.M. miR-29 modulates Wnt signaling in human osteoblasts through a positive feedback loop. J. Biol. Chem. 2010, 285, 25221–25231. [Google Scholar] [CrossRef]
- Rossi, M.; Pitari, M.R.; Amodio, N.; Di Martino, M.T.; Conforti, F.; Leone, E.; Botta, C.; Paolino, F.M.; Del Giudice, T.; Iuliano, E.; et al. miR-29b negatively regulates human osteoclastic cell differentiation and function: Implications for the treatment of multiple myeloma-related bone disease. J. Cell. Physiol. 2013, 228, 1506–1515. [Google Scholar] [CrossRef]
- Feichtinger, X.; Muschitz, C.; Heimel, P.; Baierl, A.; Fahrleitner-Pammer, A.; Redl, H.; Resch, H.; Geiger, E.; Skalicky, S.; Dormann, R.; et al. Bone-related Circulating MicroRNAs miR-29b-3p, miR-550a-3p, and miR-324-3p and their Association to Bone Microstructure and Histomorphometry. Sci. Rep. 2018, 8, 4867. [Google Scholar] [CrossRef]
- Bottani, M.; Banfi, G.; Lombardi, G. Perspectives on MiRAs as Epigenetic Markers in Osteoporosis and Bone Fracture Risk: A Step Forward in Personalized Diagnosis. Front. Genet. 2019, 10. [Google Scholar] [CrossRef]
- Mäkitie, R.; Hackl, M.; Niinimäki, R.; Kakko, S.; Grillari, J.; Mäkitie, O. Altered MicroRNA Profile in Osteoporosis Caused by Impaired WNT Signaling. J. Clin. Endocrinol. Metab. 2018, 103, 1985–1996. [Google Scholar] [CrossRef] [PubMed]
- Mandourah, A.; Ranganath, L.; Barraclough, R.; Vinjamuri, S.; Van’T Hof, R.; Hamill, S.; Czanner, G.; Dera, A.; Wang, D.; Barraclough, D. Circulating MicroRNAs as Potential Diagnostic Biomarkers for Osteoporosis. Sci. Rep. 2018, 8, 8421. [Google Scholar] [CrossRef]
- Qadir, A.S.; Um, S.; Lee, H.; Baek, K.; Seo, B.M.; Lee, G.; Kim, G.S.; Woo, K.M.; Ryoo, H.M.; Baek, J.H. miR-124 Negatively Regulates Osteogenic Differentiation and In vivo Bone Formation of Mesenchymal Stem Cells. J. Cell. Biochem. 2015, 116, 730–742. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, L.; Moore, B.T.; Peng, X.H.; Fang, X.; Lappe, J.M.; Recker, R.R.; Xiao, P. MiR-133a in human circulating monocytes: A potential biomarker associated with postmenopausal osteoporosis. PLoS ONE 2012, 7, e34641. [Google Scholar] [CrossRef] [PubMed]
- Kocijan, R.; Weigl, M.; Skalicky, S.; Geiger, E.; Ferguson, J.; Leinfellner, G.; Heimel, P.; Pietschmann, P.; Grillari, J.; Redl, H.; et al. MicroRNA Levels in Bone and Blood Change during Bisphosphonate and Teriparatide Therapy in an Animal Model of Postmenopausal Osteoporosis. Bone 2020, 131, 115104. [Google Scholar] [CrossRef]
- Shi, X.; Zhang, Z. MicroRNA-135a-5p Is Involved in Osteoporosis Progression through Regulation of Osteogenic Differentiation by Targeting RUNX2. Exp. Ther. Med. 2019, 18, 2393–2400. [Google Scholar] [CrossRef]
- Xiao, Y.; Li, B.; Liu, J. MicroRNA-148a Inhibition Protects against Ovariectomy-induced Osteoporosis through PI3K/AKT Signaling by Estrogen Receptor α. Mol. Med. Rep. 2018, 17, 7789–7796. [Google Scholar] [CrossRef]
- Pan, B.; Tong, Z.; Li, S.; Wu, L.; Liao, J.; Yang, Y.; Li, H.; Dai, Y.; Li, J.; Pan, L. Decreased MicroRNA-182-5p Helps Alendronate Promote Osteoblast Proliferation and Differentiation in Osteoporosis via the Rap1/MAPK Pathway. Biosci. Rep. 2018, 38. [Google Scholar] [CrossRef]
- Lv, R.; Pan, X.; Song, L.; Sun, Q.; Guo, C.; Zou, S.; Zhou, Q. MicroRNA-200a-3p Accelerates the Progression of Osteoporosis by Targeting Glutaminase to Inhibit Osteogenic Differentiation of Bone Marrow Mesenchymal Stem Cells. Biomed. Pharmacother. 2019, 116, 108960. [Google Scholar] [CrossRef]
- Qiu, J.; Huang, G.; Na, N.; Chen, L. MicroRNA-214-5p/TGF-β/Smad2 Signaling Alters Adipogenic Differentiation of Bone Marrow Stem Cells in Postmenopausal Osteoporosis. Mol. Med. Rep. 2018, 17, 6301–6310. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Gao, Y.; Cai, L.; Li, F.; Lou, Y.; Xu, N.; Kang, Y.; Yang, H. MicroRNA-221 Is Involved in the Regulation of Osteoporosis through Regulates RUNX2 Protein Expression and Osteoblast Differentiation. Am. J. Transl. Res. 2017, 9, 126–135. [Google Scholar] [PubMed]
- Pickering, M.; Millet, M.; Rousseau, J.; Croset, M.; Szulc, P.; Borel, O.; Rendu, E.; Chapurlat, R. Selected Serum MicroRNA, Abdominal Aortic Calcification and Risk of Osteoporotic Fracture. PLoS ONE 2019, 14, e0216947. [Google Scholar] [CrossRef] [PubMed]
- Guo, D.; Han, Y.; Cong, L.; Liang, D.; Tu, G. Resveratrol Prevents Osteoporosis in Ovariectomized Rats by Regulating MicroRNA-338-3p. Mol. Med. Rep. 2015, 12, 2098–2106. [Google Scholar] [CrossRef]
- Lin, C.; Yu, S.; Jin, R.; Xiao, Y.; Pan, M.; Pei, F.; Zhu, X.; Huang, H.; Zhang, Z.; Chen, S.; et al. Circulating MiR-338 Cluster Activities on Osteoblast Differentiation: Potential Diagnostic and Therapeutic Targets for Postmenopausal Osteoporosis. Theranostics 2019, 9, 3780–3797. [Google Scholar] [CrossRef]
- Li, G.; Bu, J.; Zhu, Y.; Xiao, X.; Liang, X.; Zhang, R. Curcumin Improves Bone Microarchitecture in Glucocorticoid-Induced Secondary Osteoporosis Mice through the Activation of MicroRNA-365 via Regulating MMP-9. Int. J. Clin. Exp. Pathol. 2015, 8, 15684–15695. [Google Scholar]
- Zhang, H.; Ding, W.; Ji, F.; Wu, D. MicroRNA-410 Participates in the Pathological Process of Postmenopausal Osteoporosis by Downregulating Bone Morphogenetic Protein-2. Exp. Ther. Med. 2019, 18, 3659–3666. [Google Scholar] [CrossRef]
- Cao, Z.; Moore, B.; Wang, Y.; Peng, X.; Lappe, J.; Recker, R.; Xiao, P. MiR-422a as a Potential Cellular MicroRNA Biomarker for Postmenopausal Osteoporosis. PLoS ONE 2014, 9, e97098. [Google Scholar] [CrossRef]
- Li, J.; Wei, X.; Sun, Q.; Zhao, X.; Zheng, C.; Bai, C.; Du, J.; Zhang, Z.; Zhu, L.; Jia, Y. MicroRNA-449b-5p Promotes the Progression of Osteoporosis by Inhibiting Osteogenic Differentiation of BMSCs via Targeting Satb2. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 6394–6403. [Google Scholar] [CrossRef]
- Li, X.; Ning, L.; Zhao, X.; Wan, S. MicroRNA-543 Promotes Ovariectomy-Induced Osteoporosis through Inhibition of AKT/P38 MAPK Signaling Pathway by Targeting YAF2. J. Cell. Biochem. 2018. [Google Scholar] [CrossRef]
- Luo, B.; Yang, J.; Wang, Y.; Qu, G.; Hao, P.; Zeng, Z.; Yuan, J.; Yang, R.; Yuan, Y. MicroRNA-579-3p Promotes the Progression of Osteoporosis by Inhibiting Osteogenic Differentiation of Mesenchymal Stem Cells through Regulating Sirt1. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 6791–6799. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Liu, Z.; Yu, B.; Zhang, X. MicroRNA-874 Targeting SUFU Involves in Osteoblast Proliferation and Differentiation in Osteoporosis Rats through the Hedgehog Signaling Pathway. Biochem. Biophys. Res. Comm. 2018, 506, 194–203. [Google Scholar] [CrossRef]
- Wang, Q.; Wang, C.; Meng, Y. MicroRNA-1297 Promotes the Progression of Osteoporosis through Regulation of Osteogenesis of Bone Marrow Mesenchymal Stem Cells by Targeting WNT5A. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 4541–4550. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Xie, H.; Liu, W.; Hu, R.; Huang, B.; Tan, Y.F.; Liao, E.Y.; Xu, K.; Sheng, Z.F.; Zhou, H.D.; et al. A Novel MicroRNA Targeting HDAC5 Regulates Osteoblast Differentiation in Mice and Contributes to Primary Osteoporosis in Humans. J. Clin. Invest. 2009, 119, 3666–3677. [Google Scholar] [CrossRef]
- Cai, N.; Li, C.; Wang, F. Silencing of LncRNA-ANCR Promotes the Osteogenesis of Osteoblast Cells in Postmenopausal Osteoporosis via Targeting EZH2 and RUNX2. Yonsei Med. J. 2019, 60, 751–759. [Google Scholar] [CrossRef] [PubMed]
- Tong, X.; Gu, P.; Xu, S.; Lin, X. Long Non-Coding RNA-DANCR in Human Circulating Monocytes: A Potential Biomarker Associated with Postmenopausal Osteoporosis. Biosci. Biotechnol. Biochem. 2015, 79, 732–737. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Zhang, X.; Zhu, X.; Wang, C. LncRNA Bmncr Alleviates the Progression of Osteoporosis by Inhibiting RANML-Induced Osteoclast Differentiation. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 9199–9206. [Google Scholar] [CrossRef]
- Yu, H.; Zhou, W.; Yan, W.; Xu, Z.; Xie, Y.; Zhang, P. LncRNA CASC11 Is Upregulated in Postmenopausal Osteoporosis and Is Correlated with TNF-α. Clin. Interv. Aging 2019, 14, 1663–1669. [Google Scholar] [CrossRef]
- Feng, J.; Wang, J.; Li, C. LncRNA GAS5 Overexpression Alleviates the Development of Osteoporosis through Promoting Osteogenic Differentiation of MSCs via Targeting MicroRNA-498 to Regulate RUNX2. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 7757–7765. [Google Scholar] [CrossRef]
- Yang, X.; Yang, J.; Lei, P.; Wen, T. LncRNA MALAT1 Shuttled by Bone Marrow-Derived Mesenchymal Stem Cells-Secreted Exosomes Alleviates Osteoporosis through Mediating MicroRNA-34c/SATB2 Axis. Aging 2019, 11, 8777–8791. [Google Scholar] [CrossRef]
- Zheng, S.; Wang, Y.; Yang, Y.; Chen, B.; Wang, C.; Li, R.; Huang, D. LncRNA MALAT1 Inhibits Osteogenic Differentiation of Mesenchymal Stem Cells in Osteoporosis Rats through MAPK Signaling Pathway. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 4609–46017. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Li, Y.; Zhang, Y.; Ma, L.; Lin, L.; Meng, J.; Jiang, L.; Wang, L.; Zhou, P.; Zhang, Y. LncRNA MEG3 Inhibited Osteogenic Differentiation of Bone Marrow Mesenchymal Stem Cells from Postmenopausal Osteoporosis by Targeting MiR-133a-3p. Biomed. Pharmacother. 2017, 89, 1178–1186. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Peng, G.; Wu, H.; Liu, M.; Mao, G.; Ning, X.; Yang, H.; Deng, J. Long non-coding RNA MEG3 is involved in osteogenic differentiation and bone diseases (Review). Biomed. Rep. 2020, 13, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Hu, X.; He, S.; Ding, W.; Wang, F.; Zhao, Y.; Huang, Z. LncRNA MSC-AS1 Promotes Osteogenic Differentiation and Alleviates Osteoporosis through Sponging MicroRNA-140-5p to Upregulate BMP2. Biochem/ Biophys. Res. Comm. 2019, 519, 790–796. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Guo, Z.; Gao, W.; Wang, J.; Liu, Y.; Gao, F.; Sun, S.; Zhou, X.; Yang, Z.; Zheng, W. LncRNA-NEF Is Downregulated in Postmenopausal Osteoporosis and Is Related to Course of Treatment and Recurrence. J. Int. Med. Res. 2019, 47, 3299–3306. [Google Scholar] [CrossRef]
- Huang, S.; Zhu, X.; Xiao, D.; Zhuang, J.; Liang, G.; Liang, C.; Zheng, X.; Ke, Y.; Chang, Y. LncRNA SNHG1 Was Down-Regulated after Menopause and Participates in Postmenopausal Osteoporosis. Biosci. Rep. 2019, 39. [Google Scholar] [CrossRef]
- Han, Y.; Liu, C.; Lei, M.; Sun, S.; Zheng, W.; Niu, Y.; Xia, X. LncRNA TUG1 Was Upregulated in Osteoporosis and Regulates the Proliferation and Apoptosis of Osteoclasts. J. Orthop. Surg. Res. 2019, 14. [Google Scholar] [CrossRef]
- Zhang, H.; Du, X.; Dong, Q. LncRNA XIXT Promotes Osteogenic Differentiation of Bone Mesenchymal Stem Cells and Alleviates Osteoporosis Progression by Targeting MiRNA-30a-5p. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 8721–8729. [Google Scholar] [CrossRef]
- Feurer, E.; Kan, C.; Croset, M.; Sornay-Rendu, E.; Chapurlat, R. Lack of Association Between Select Circulating miRNAs and Bone Mass, Turnover, and Fractures: Data from the OFELY Cohort. J. Bone Miner. Res. 2019, 34, 1074–1085. [Google Scholar] [CrossRef]
- Ladang, A.; Beaudart, C.; Locquet, M.; Reginster, J.-Y.; Bruyère, O.; Cavalier, E. Evaluation of a Panel of MicroRNAs That Predicts Fragility Fracture Risk: A Pilot Study. Calcif. Tissue Int. 2020, 106, 239–247. [Google Scholar] [CrossRef]
- Sun, X.; Guo, Q.; Wei, W.; Robertson, S.; Yuan, Y.; Luo, Y. Current Progress on MicroRNA-Based Gene Delivery in the Treatment of Osteoporosis and Osteoporotic Fracture. Int. J. Endocrinol. 2019, 3, 1–17. [Google Scholar] [CrossRef] [PubMed]
| miRNA | Action | Expression in PMOP | Sources | References |
|---|---|---|---|---|
| miR-9-5p | inhibit osteogenesis, promote adipogenesis promote osteoclastogenesis | high | serum | Zhang et al. [14] Wang et al. [15] |
| miR-21 | promote osteoclastogenesis promote osteogenesis and inhibit osteoclastogenesis | unclear | Jiang et al. [16] Cheng at al. [17] Seeliger et al. [18] Yavropoulou et al. [19] Yang et al. [23] Hu et al. [24] | |
| miR-29 | unclear | low | serum | Tang et al. [1] Lian et al. [25] Kocijan et al. [26] Li et al. [27] Kapinas et al. [28] Rossi et al. [29] Bottani et al. [31] |
| miR-30b-5p | negatively regulate osteoblast differentiation | low | serum | Bottani et al. [31] |
| miR-31 | unclear | low | bone | Tang et a. [1] Foessl et al. [5] Mäkitie et al. [32] |
| miR-100 | inhibit osteogenic differentiation | high | bone and serum | Cheng at al. [17] Seeliger et al. [18] |
| miR-103-3p | inhibit osteoblast differentiation and proliferation | low | serum | Bottani et al. [31] |
| miR-122-5p | inhibit osteoblast differentiation | low | serum | Mandourah et al. [33] |
| miR-124 | inhibit osteoclast formation inhibit osteogenesis | high | serum | Tang et al. [1] Yavropoulou et al. [19] Qadiret et al. [34] |
| miR-133 | inhibit osteoblast differentiation increase osteoclastogenesis | high | bone and serum (mouse) | Tang et al. [1] Cheng at al. [17] Wang et al. [35] Kocijan et al. [36] |
| miR-135a-5p | inhibit osteogenic differentiation | high | bone | Shi et al. [37] |
| miR-146a | inhibit osteoclastogenesis | high | bone (mouse) | Tang et al. [1] |
| miR-148a | induce osteoclast formation | high | Serum (mouse) | Tang et al. [1] Xiao et al. [38] |
| miR-155 | regulate osteoclastogenesis | high | unclear (mouse) | Tang et al. [1] |
| miR-182-5p | inhibited ADCY6 expression and Rap1/MAPK signaling pathway activation | high | bone and serum (mouse) | Pan et al. [39] |
| miR-194-5p | unclear | high | whole blood lysate | Foessl et al. [5] |
| miR-200a-3p | inhibit osteogenic differentiation | high | serum | Lv et al. [40] |
| miR-203a | slow osteoblast differentiation | high | bone | Kocijan et al. [36] |
| miR-214-5p | promote adipogenic differentiation | high | (in vitro) | Qiu et al. [41] |
| miR-221 | inhibit osteogenic inhibition | low | bone | Zhang et al. [42] |
| miR-223 | inhibit osteoclast differentiation | unclear | serum | Tang et al. [1] Seeliger et al. [18] Pickering et al. [43] |
| miR-338 | regulate osteoblast differentiation | high | serum | Guo et al. [44] Lin et al. [35] |
| miR-365 | suppresses MMP-9 | low | bone (mouse) | Li et al. [46] |
| miR-410 | regulate BMP-2 expression | high | serum | van Wijnen et al. [3] Zhang et al. [47] |
| miR-422a | may stimulate osteoclastogenesis | high | human circulating monocytes | Bottani et al. [31] Cao et al. [48] |
| miR-449b-5p | inhibit osteogenic differentiation | unclear | (in vivo) | Li et al. [49] |
| miR-503 | inhibit osteoclast differentiation | low | human circulating monocytes | Tang et al. [1] |
| miR-543 | promote osteoblast apoptosis | high | bone (mouse) | Li et al. [50] |
| miR-579-3p | inhibit osteogenic differentiation | high | serum | Luo et al. [51] |
| miR-874 | promote osteoblast proliferation | low | bone (mouse) | Lin et al. [52] |
| miR-1297 | inhibit osteogenic differentiation | high | bone | Wang et al. [53] |
| miR-2861 | promote osteoblast differentiation | high | serum | Yavropoulou et al. [19] Li et al. [54] |
| lncRNA | Action | Expression in PMOP | Sources | References |
|---|---|---|---|---|
| ANCR | inhibit osteoblasts, increase osteoclastogenesis | high | blood mononuclear cells | Wu et al. [9] Cai et al. [55] Tong et al. [56] |
| BMNCR | inhibit osteoporosis | low | bone (mouse) | Chen et al. [57] |
| CASC11 | lead to TNF-α upregulation in osteoclasts | high | plasma | Yu et al. [58] |
| CRNDE | regulate cell apoptosis | high | bone | Li et al. [10] |
| GAS5 | regulate osteogenic differentiation | low | bone | Feng et al. [59] |
| MALAT1 | unclear | low | bone (mouse) | Yang et al. [60] Zheng et al. [61] |
| MEG3 | unclear | high | bone | Wu et al. [9] Wang et al. [62] Sun et al. [63] |
| MSC-AS1 | induce osteogenic differentiation | unclear | bone (mouse) | Zhang et al. [64] |
| NEF | interact with IL-6 | low | plasma | Ma et al. [65] |
| SNHG1 | unclear | low | plasma | Huang et al. [66] |
| TUG1 | may promote osteoclast differentiation | high | plasma | Han et al. [67] |
| XIXT | promote osteogenic differentiation of BMSCs | low | Serum | Zhang et al. [68] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ko, N.-Y.; Chen, L.-R.; Chen, K.-H. The Role of Micro RNA and Long-Non-Coding RNA in Osteoporosis. Int. J. Mol. Sci. 2020, 21, 4886. https://doi.org/10.3390/ijms21144886
Ko N-Y, Chen L-R, Chen K-H. The Role of Micro RNA and Long-Non-Coding RNA in Osteoporosis. International Journal of Molecular Sciences. 2020; 21(14):4886. https://doi.org/10.3390/ijms21144886
Chicago/Turabian StyleKo, Nai-Yu, Li-Ru Chen, and Kuo-Hu Chen. 2020. "The Role of Micro RNA and Long-Non-Coding RNA in Osteoporosis" International Journal of Molecular Sciences 21, no. 14: 4886. https://doi.org/10.3390/ijms21144886
APA StyleKo, N.-Y., Chen, L.-R., & Chen, K.-H. (2020). The Role of Micro RNA and Long-Non-Coding RNA in Osteoporosis. International Journal of Molecular Sciences, 21(14), 4886. https://doi.org/10.3390/ijms21144886






