Ectopic Expression of Jatropha curcas TREHALOSE-6-PHOSPHATE PHOSPHATASE J Causes Late-Flowering and Heterostylous Phenotypes in Arabidopsis but not in Jatropha
Abstract
:1. Introduction
2. Results
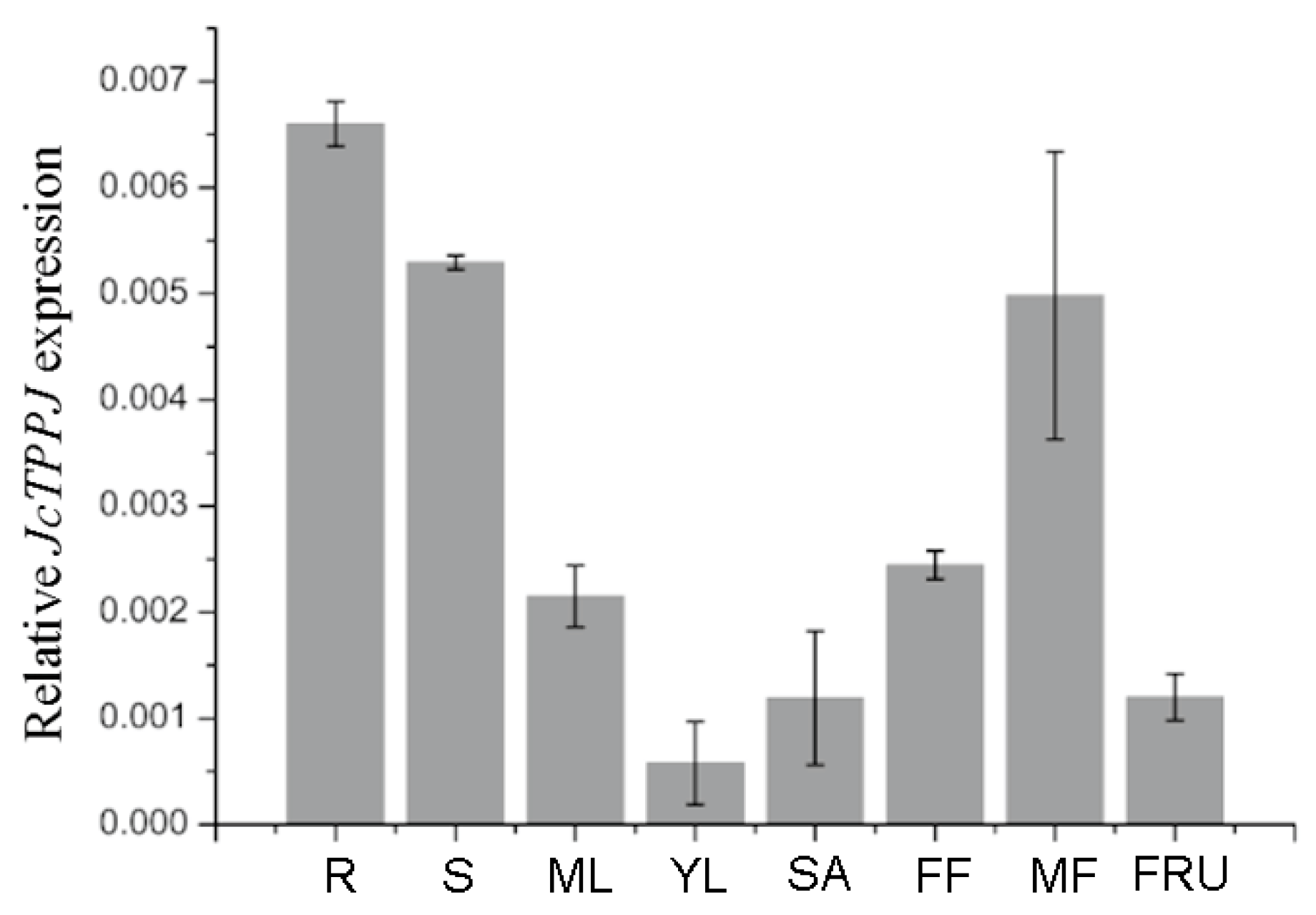
2.1. JcTPPJ is Highly Expressed in Male Flowers
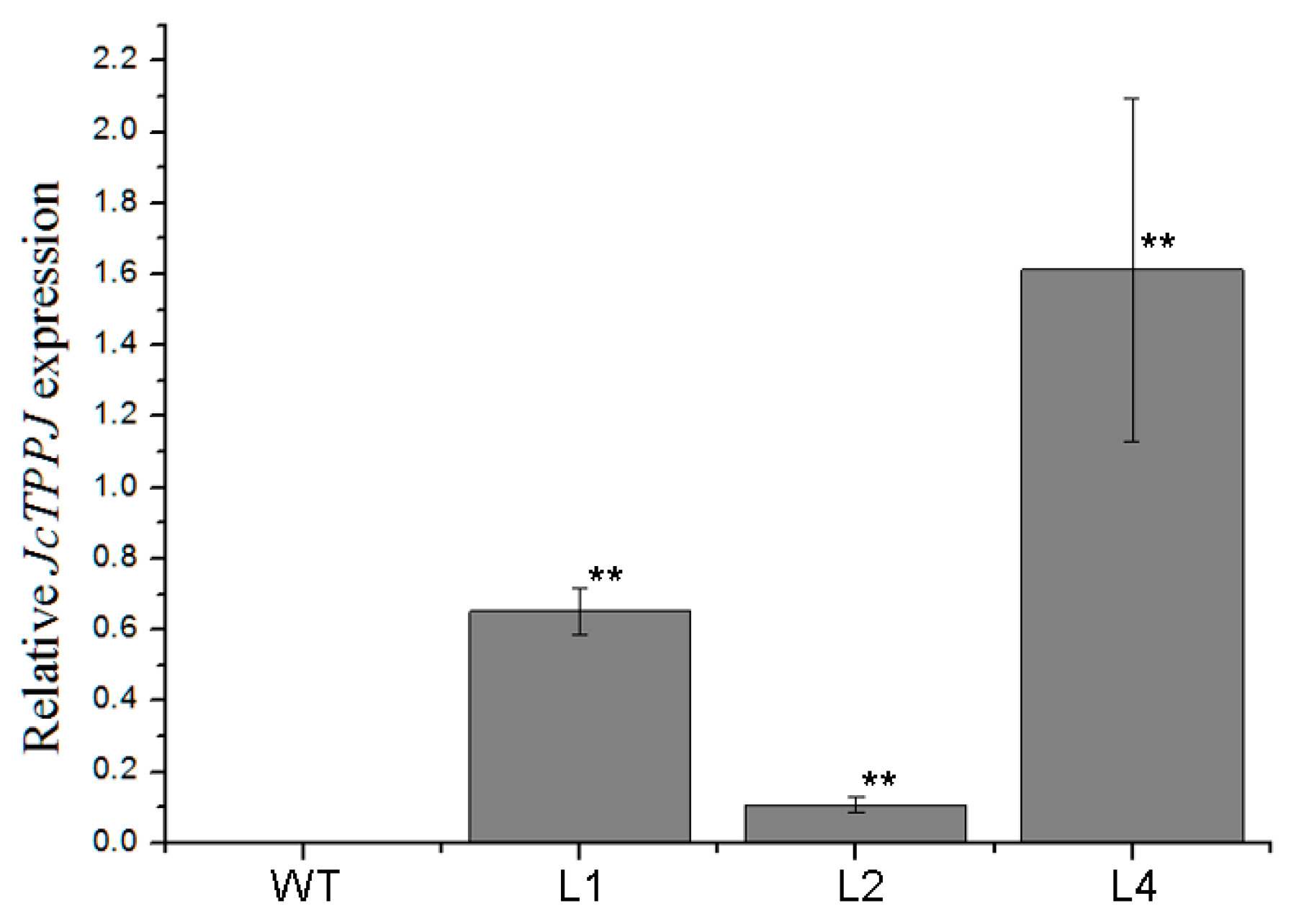
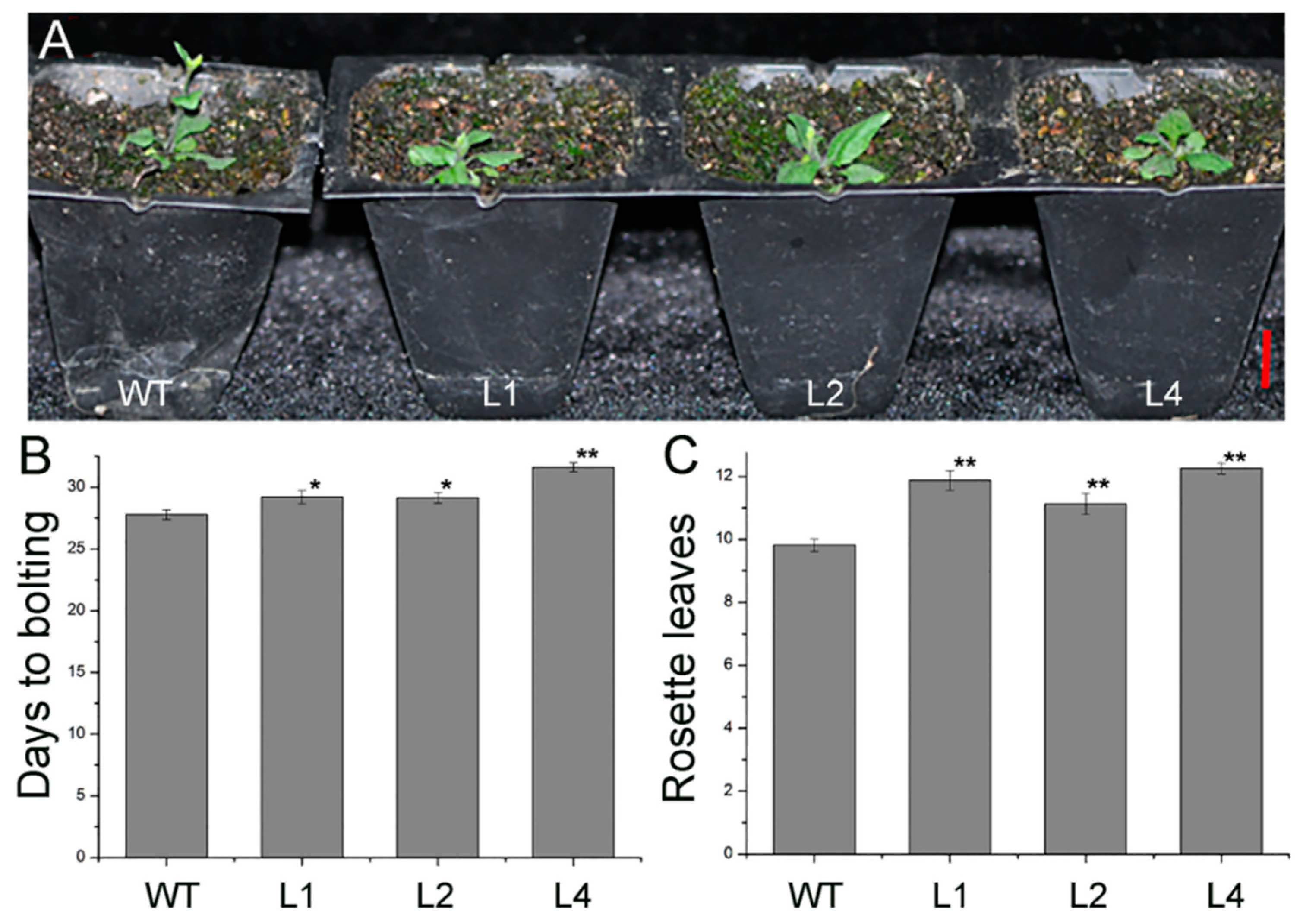
2.2. Overexpression of JcTPPJ in Arabidopsis Delayed Flowering
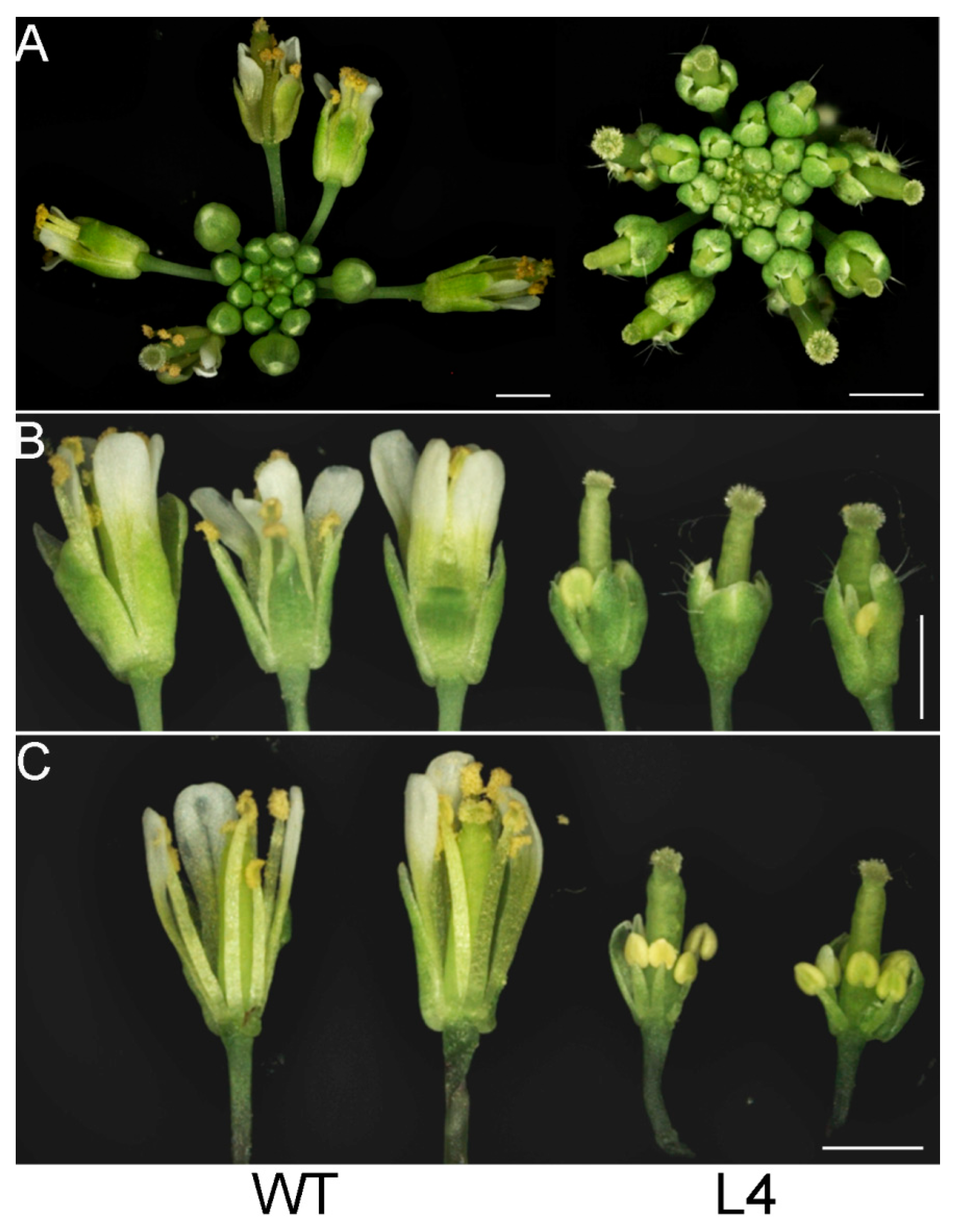
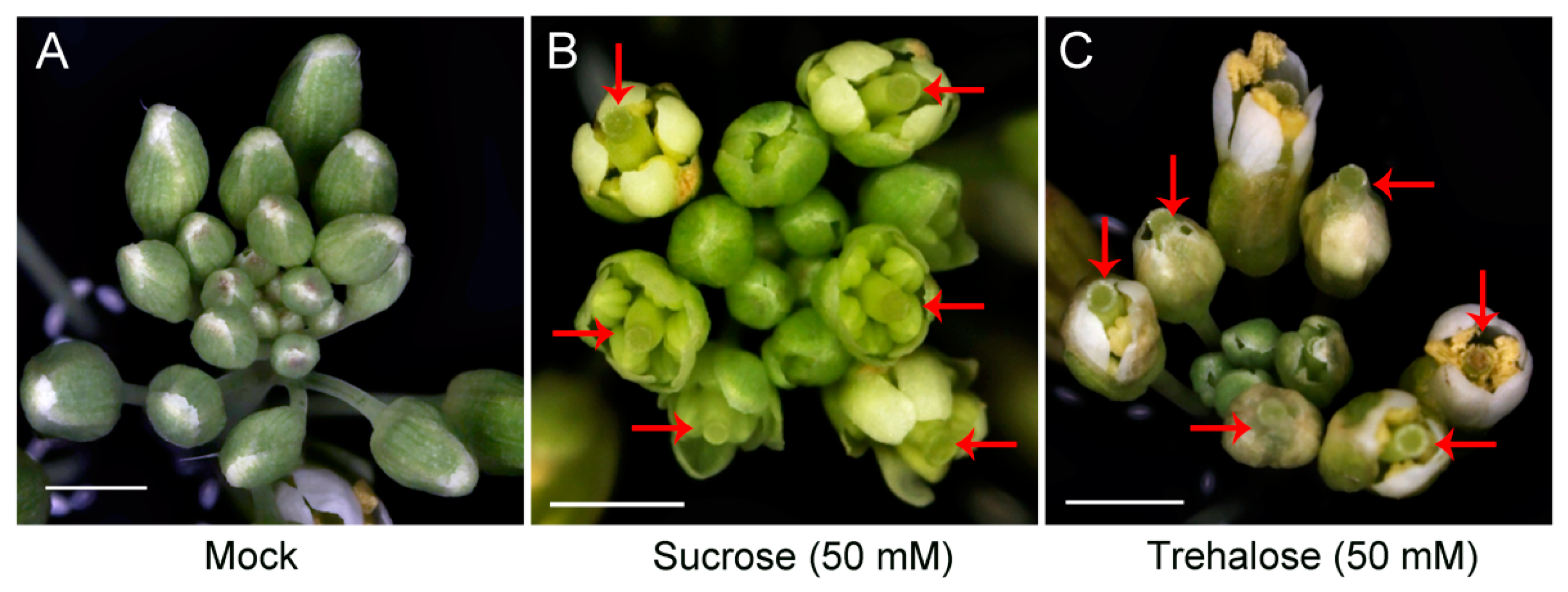
2.3. Overexpression of JcTPPJ in Arabidopsis Resulted in Delayed Perianth and Stamen Filaments Development
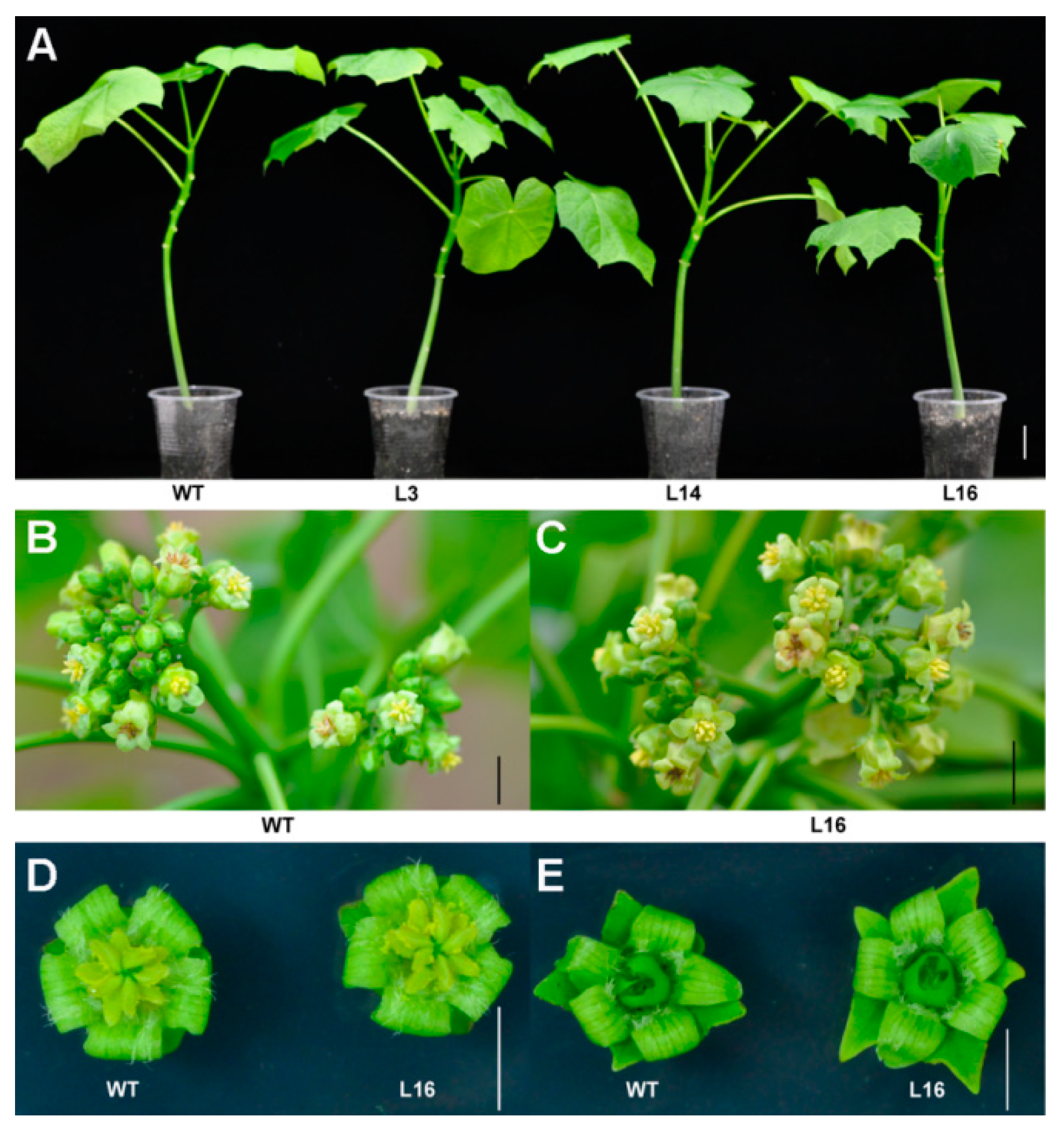
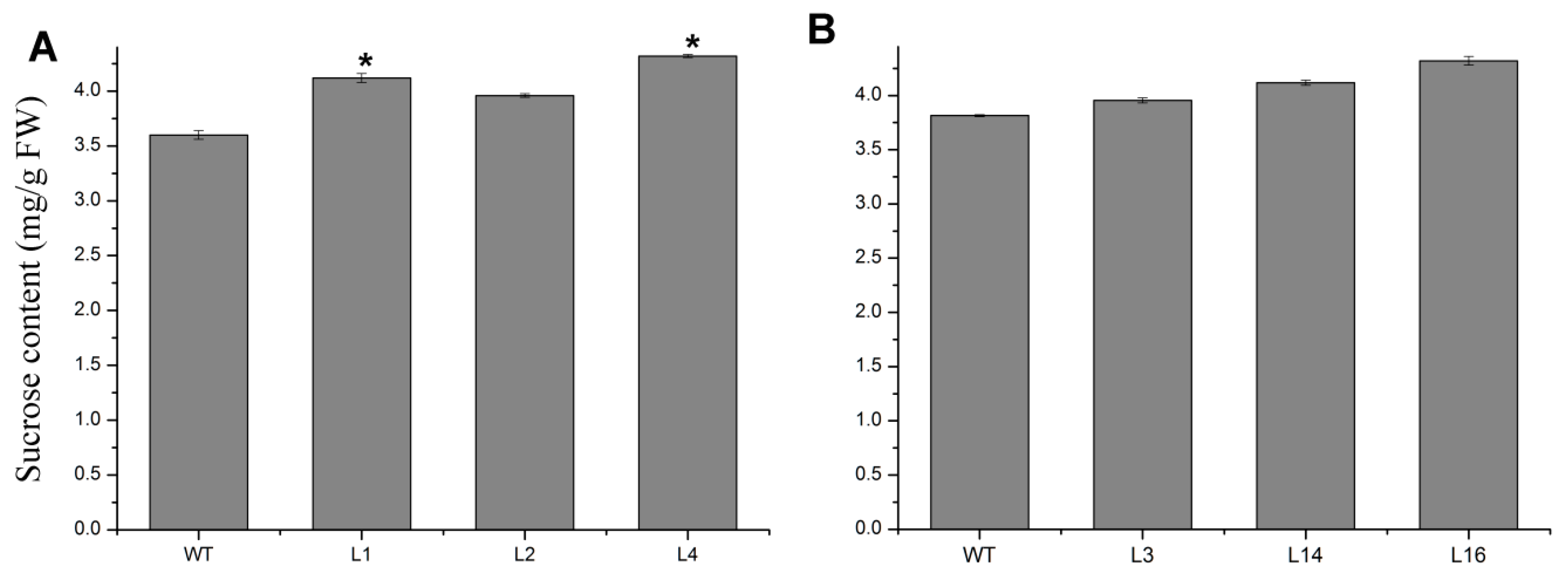
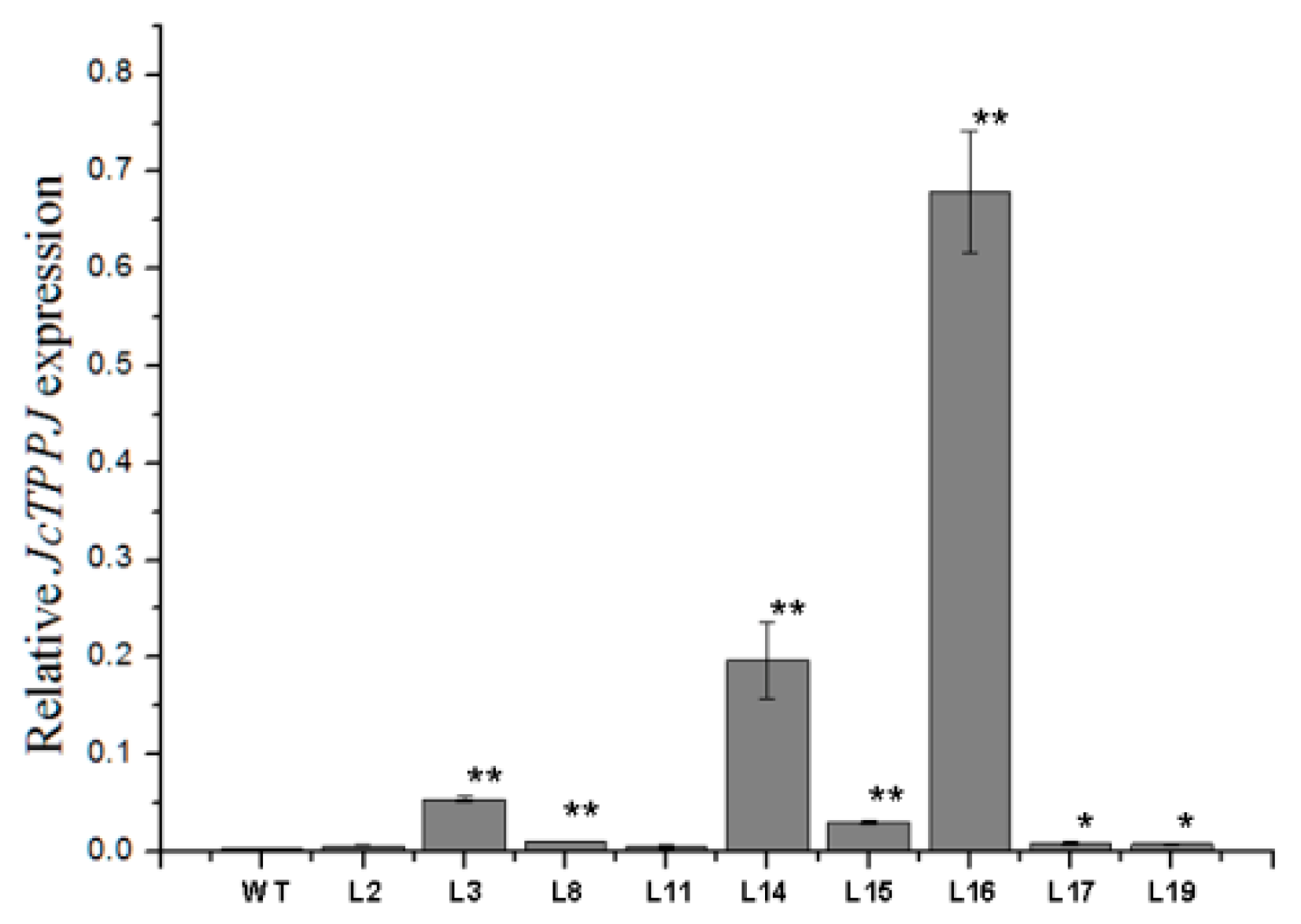
2.4. Overexpression of JcTPPJ Did Not Affect Flower Development in Jatropha
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Phenotypic Analysis
4.3. Isolation of JcTPPJ cDNA
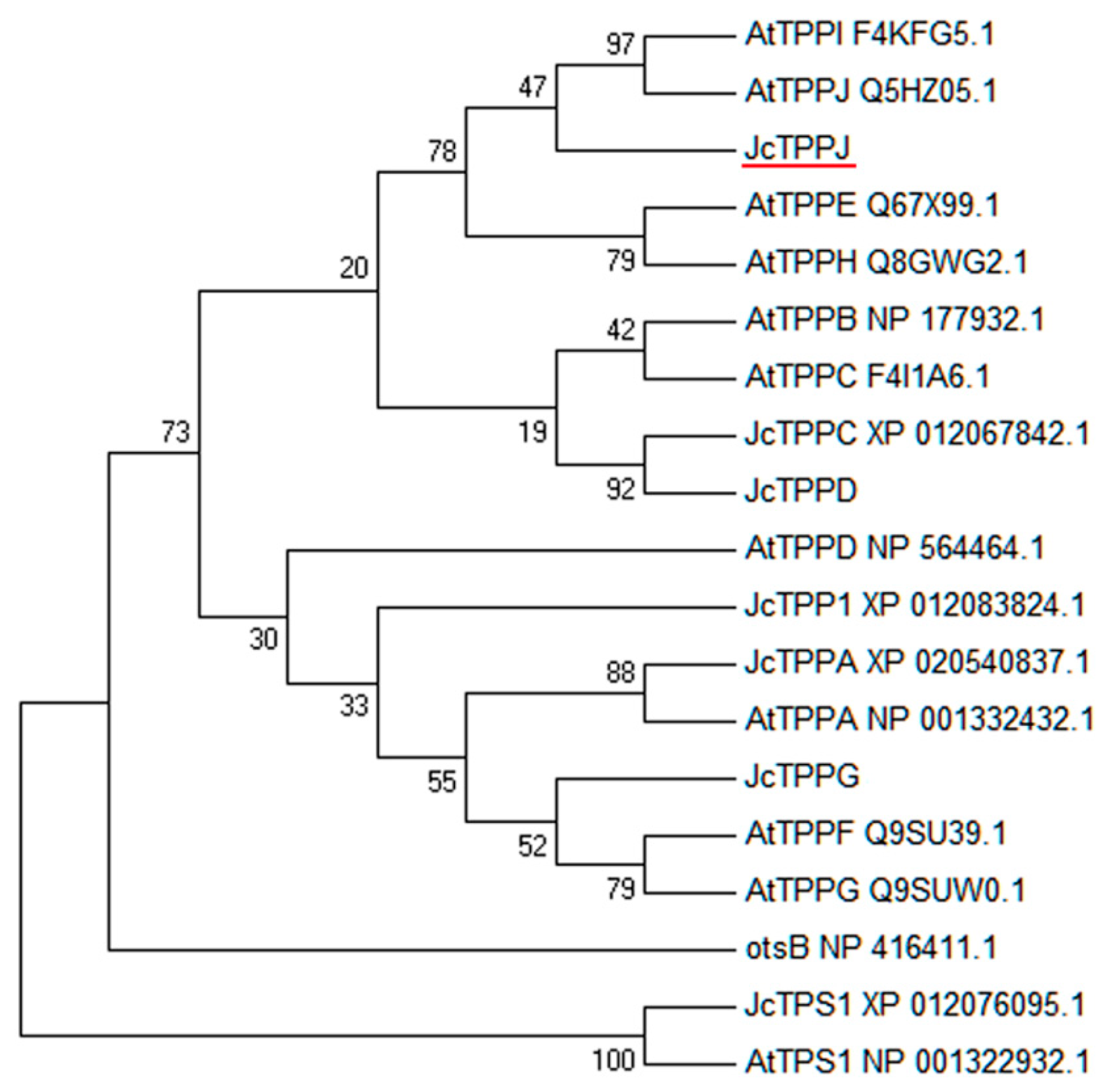
4.4. Phylogenetic Analysis and Sequence Alignment
4.5. Vector Construction and Transformation
4.6. Quantitative RT-PCR Analysis
4.7. Sucrose Treatment and Determination
4.8. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| CaMV | cauliflower mosaic virus |
| qRT-PCR | quantitative reverse transcriptase-polymerase chain reaction |
| T6P | trehalose-6-phosphate |
| TPP | trehalose-6-phosphate phosphatase |
| TPS | trehalose-6-phosphate synthase |
| TRE | trehalase |
| UDP-Glu | uridine diphosphate glucose |
| WT | wild-type |
References
- Paul, M.J.; Primavesi, L.F.; Jhurreea, D.; Zhang, Y. Trehalose metabolism and signaling. Annu. Rev. Plant Biol. 2008, 59, 417–441. [Google Scholar] [CrossRef] [PubMed]
- Walmagh, M.; Zhao, R.; Desmet, T. Trehalose Analogues: Latest Insights in Properties and Biocatalytic Production. Int. J. Mol. Sci. 2015, 16, 13729–13745. [Google Scholar] [CrossRef]
- Wingler, A. The function of trehalose biosynthesis in plants. Phytochemistry 2002, 60, 437–440. [Google Scholar] [CrossRef]
- Iturriaga, G.; Suarez, R.; Nova-Franco, B. Trehalose metabolism: from osmoprotection to signaling. Int. J. Mol. Sci. 2009, 10, 3793–3810. [Google Scholar] [CrossRef] [PubMed]
- Han, B.; Fu, L.; Zhang, D.; He, X.; Chen, Q.; Peng, M.; Zhang, J. Interspecies and Intraspecies Analysis of Trehalose Contents and the Biosynthesis Pathway Gene Family Reveals Crucial Roles of Trehalose in Osmotic-Stress Tolerance in Cassava. Int. J. Mol. Sci. 2016, 17, 1077. [Google Scholar] [CrossRef]
- Müller, J.; Aeschbacher, R.A.; Wingler, A.; Boller, T.; Wiemken, A. Trehalose and trehalase in Arabidopsis. Plant Physiol. 2001, 125, 1086–1093. [Google Scholar] [CrossRef]
- Leyman, B.; van Dijck, P.; Thevelein, J.M. An unexpected plethora of trehalose biosynthesis genes in Arabidopsis thaliana. Trends Plant Sci. 2001, 6, 510–513. [Google Scholar] [CrossRef]
- Krasensky, J.; Broyart, C.; Rabanal, F.A.; Jonak, C. The redox-sensitive chloroplast trehalose-6-phosphate phosphatase AtTPPD regulates salt stress tolerance. Antioxid. Redox Signal. 2014, 21, 1289–1304. [Google Scholar] [CrossRef]
- Lunn, J.E. Gene families and evolution of trehalose metabolism in plants. Funct. Plant Biol. 2007, 34, 550–563. [Google Scholar] [CrossRef]
- Ramon, M.; De Smet, I.; Vandesteene, L.; Naudts, M.; Leyman, B.; van Dijck, P.; Rolland, F.; Beeckman, T.; Thevelein, J.M. Extensive expression regulation and lack of heterologous enzymatic activity of the Class II trehalose metabolism proteins from Arabidopsis thaliana. Plant Cell Environ. 2009, 32, 1015–1032. [Google Scholar] [CrossRef]
- Vandesteene, L.; Ramon, M.; Le Roy, K.; van Dijck, P.; Rolland, F. A single active trehalose-6-P synthase (TPS) and a family of putative regulatory TPS-like proteins in Arabidopsis. Mol. Plant 2010, 3, 406–419. [Google Scholar] [CrossRef]
- Vandesteene, L.; Lopez-Galvis, L.; Vanneste, K.; Feil, R.; Maere, S.; Lammens, W.; Rolland, F.; Lunn, J.E.; Avonce, N.; Beeckman, T.; et al. Expansive evolution of the trehalose-6-phosphate phosphatase gene family in Arabidopsis. Plant Physiol. 2012, 160, 884–896. [Google Scholar] [CrossRef]
- Blázquez, M.A.; Santos, E.; Flores, C.-L.; Martínez-Zapater, J.M.; Salinas, J.; Gancedo, C. Isolation and molecular characterization of the Arabidopsis TPS1 gene, encoding trehalose-6-phosphate synthase. Plant J. 1998, 13, 685–689. [Google Scholar] [CrossRef]
- Schluepmann, H.; Pellny, T.K.; Van Dijken, A.; Smeekens, S.; Paul, M.J. Trehalose 6-phosphate is indispensable for carbohydrate utilization and growth in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2003, 100, 6849–6854. [Google Scholar] [CrossRef] [PubMed]
- Figueroa, C.M.; Feil, R.; Ishihara, H.; Watanabe, M.; Kolling, K.; Krause, U.; Hohne, M.; Encke, B.; Plaxton, W.C.; Zeeman, S.C.; et al. Trehalose 6-phosphate coordinates organic and amino acid metabolism with carbon availability. Plant J. 2016, 85, 410–423. [Google Scholar] [CrossRef]
- Gomez, L.D.; Baud, S.; Gilday, A.; Li, Y.; Graham, I.A. Delayed embryo development in the ARABIDOPSIS TREHALOSE-6-PHOSPHATE SYNTHASE 1 mutant is associated with altered cell wall structure, decreased cell division and starch accumulation. Plant J. 2006, 46, 69–84. [Google Scholar] [CrossRef]
- Eastmond, P.J.; van Dijken, A.J.H.; Spielman, M.; Kerr, A.; Tissier, A.F.; Dickinson, H.G.; Jones, J.D.G.; Smeekens, S.C.; Graham, I.A. Trehalose-6-phosphate synthase 1, which catalyses the first step in trehalose synthesis, is essential for Arabidopsis embryo maturation. Plant J. 2002, 29, 225–235. [Google Scholar] [CrossRef]
- Schluepmann, H.; van Dijken, A.; Aghdasi, M.; Wobbes, B.; Paul, M.; Smeekens, S. Trehalose mediated growth inhibition of Arabidopsis seedlings is due to trehalose-6-phosphate accumulation. Plant Physiol. 2004, 135, 879–890. [Google Scholar] [CrossRef]
- Satoh-Nagasawa, N.; Nagasawa, N.; Malcomber, S.; Sakai, H.; Jackson, D. A trehalose metabolic enzyme controls inflorescence architecture in maize. Nature 2006, 441, 227–230. [Google Scholar] [CrossRef]
- Wingler, A.; Delatte, T.L.; O’Hara, L.E.; Primavesi, L.F.; Jhurreea, D.; Paul, M.J.; Schluepmann, H. Trehalose 6-phosphate is required for the onset of leaf senescence associated with high carbon availability. Plant Physiol. 2012, 158, 1241–1251. [Google Scholar] [CrossRef]
- Wahl, V.; Ponnu, J.; Schlereth, A.; Arrivault, S.; Langenecker, T.; Franke, A.; Feil, R.; Lunn, J.E.; Stitt, M.; Schmid, M. Regulation of flowering by trehalose-6-phosphate signaling in Arabidopsis thaliana. Science 2013, 339, 704–707. [Google Scholar] [CrossRef] [PubMed]
- Kolbe, A.; Tiessen, A.; Schluepmann, H.; Paul, M.J.; Ulrich, S.; Geigenberger, P. Trehalose 6-phosphate regulates starch synthesis via posttranslational redox activation of ADP-glucose pyrophosphorylase. Proc. Natl. Acad. Sci. USA 2005, 102, 11118–11123. [Google Scholar] [CrossRef] [PubMed]
- Pilon-Smits, E.A.H.; Terry, N.; Sears, T.; Kim, H.; Zayed, A.; Hwang, S.; van Dun, K.; Voogd, E.; Verwoerd, T.C.; Krutwagen, R.W.H.H.; et al. Trehalose-producing transgenic tobacco plants show improved growth performance under drought stress. J. Plant Physiol. 1998, 152, 525–532. [Google Scholar] [CrossRef]
- Almeida, A.M.; Villalobos, E.; Araújo, S.S.; Leyman, B.; van Dijck, P.; Alfaro-Cardoso, L.; Fevereiro, P.S.; Torné, J.M.; Santos, D.M. Transformation of tobacco with an Arabidopsis thaliana gene involved in trehalose biosynthesis increases tolerance to several abiotic stresses. Euphytica 2005, 146, 165–176. [Google Scholar] [CrossRef]
- Karim, S.; Aronsson, H.; Ericson, H.; Pirhonen, M.; Leyman, B.; Welin, B.; Mantyla, E.; Palva, E.T.; Van Dijck, P.; Holmstrom, K.O. Improved drought tolerance without undesired side effects in transgenic plants producing trehalose. Plant Mol. Biol. 2007, 64, 371–386. [Google Scholar] [CrossRef] [PubMed]
- Miranda, J.A.; Avonce, N.; Suarez, R.; Thevelein, J.M.; van Dijck, P.; Iturriaga, G. A bifunctional TPS-TPP enzyme from yeast confers tolerance to multiple and extreme abiotic-stress conditions in transgenic Arabidopsis. Planta 2007, 226, 1411–1421. [Google Scholar] [CrossRef] [PubMed]
- Nuccio, M.L.; Wu, J.; Mowers, R.; Zhou, H.-P.; Meghji, M.; Primavesi, L.F.; Paul, M.J.; Chen, X.; Gao, Y.; Haque, E.; et al. Expression of trehalose-6-phosphate phosphatase in maize ears improves yield in well-watered and drought conditions. Nat. Biotechnol. 2015, 33, 862–869. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Zhang, H.; Li, J.; Cui, J.; Zhang, X.; Ren, X. Enhancement of biocontrol efficacy of Pichia carribbica to postharvest diseases of strawberries by addition of trehalose to the growth medium. Int. J. Mol. Sci. 2012, 13, 3916–3932. [Google Scholar] [CrossRef]
- Fernandez, O.; Bethencourt, L.; Quero, A.; Sangwan, R.S.; Clement, C. Trehalose and plant stress responses: friend or foe? Trends Plant Sci. 2010, 15, 409–417. [Google Scholar] [CrossRef]
- Kretzschmar, T.; Pelayo, M.A.; Trijatmiko, K.R.; Gabunada, L.F.; Alam, R.; Jimenez, R.; Mendioro, M.S.; Slamet-Loedin, I.H.; Sreenivasulu, N.; Bailey-Serres, J.; et al. A trehalose-6-phosphate phosphatase enhances anaerobic germination tolerance in rice. Nat. Plants 2015, 1, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Ohto, M.A.; Onai, K.; Furukawa, Y.; Aoki, E.; Nakamura, A.K. Effects of sugar on vegetative development and floral transition in Arabidopsis. Plant Physiol. 2001, 127, 252–261. [Google Scholar] [CrossRef]
- Rolland, F.; Moore, B.; Sheen, J. Sugar Sensing and Signaling in Plants. Plant Cell 2002, 14, S185–S205. [Google Scholar] [CrossRef]
- Wang, J.; Guan, H.; Dong, R.; Liu, C.; Liu, Q.; Liu, T.; Wang, L.; He, C. Overexpression of maize sucrose non-fermenting-1-related protein kinase 1 genes, ZmSnRK1s, causes alteration in carbon metabolism and leaf senescence in Arabidopsis thaliana. Gene 2019, 691, 34–44. [Google Scholar] [CrossRef] [PubMed]
- Gazzarrini, S.; Tsuchiya, Y.; Lumba, S.; Okamoto, M.; McCourt, P. The transcription factor FUSCA3 controls developmental timing in Arabidopsis through the hormones gibberellin and abscisic acid. Dev. Cell 2004, 7, 373–385. [Google Scholar] [CrossRef] [PubMed]
- Seo, P.J.; Ryu, J.; Kang, S.K.; Park, C.M. Modulation of sugar metabolism by an INDETERMINATE DOMAIN transcription factor contributes to photoperiodic flowering in Arabidopsis. Plant J. Cell Mol. Biol. 2011, 65, 418–429. [Google Scholar] [CrossRef]
- Tsai, A.Y.; Gazzarrini, S. AKIN10 and FUSCA3 interact to control lateral organ development and phase transitions in Arabidopsis. Plant J. Cell Mol. Biol. 2012, 69, 809–821. [Google Scholar] [CrossRef] [PubMed]
- Jeong, E.Y.; Seo, P.J.; Woo, J.C.; Park, C.M. AKIN10 delays flowering by inactivating IDD8 transcription factor through protein phosphorylation in Arabidopsis. BMC Plant Biol. 2015, 15, 110. [Google Scholar] [CrossRef]
- Sun, L.L.; Sui, X.L.; Lucas, W.J.; Li, Y.X.; Feng, S.; Ma, S.; Fan, J.; Gao, L.; Zhang, Z.X. Down-regulation of the Sucrose Transporter CsSUT1 Causes Male Sterility by Altering Carbohydrate Supply. Plant Physiol. 2019. [Google Scholar] [CrossRef]
- Alam, N.C.; Abdullah, T.L.; Abdullah, N.A. Flowering and Fruit Set Under Malaysian Climate of Jatropha curcas L. Am. J. Agric. Biol. Sci. 2011, 6, 142–147. [Google Scholar]
- Hu, Y.X.; Tao, Y.B.; Xu, Z.F. Overexpression of Jatropha Gibberellin 2-oxidase 6 (JcGA2ox6) Induces Dwarfism and Smaller Leaves, Flowers and Fruits in Arabidopsis and Jatropha. Front. Plant Sci. 2017, 8, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Li, C.Q.; Fu, Q.T.; Niu, L.J.; Luo, L.; Chen, J.H.; Xu, Z.F. Three TFL1 homologues regulate floral initiation in the biofuel plant Jatropha curcas. Sci. Rep. 2017, 7, 1–9. [Google Scholar] [CrossRef]
- Cai, L.; Zhang, L.; Fu, Q.T.; Xu, Z.F. Identification and expression analysis of cytokinin metabolic genes IPTs, CYP735A and CKXs in the biofuel plant Jatropha curcas. PeerJ 2018, 6, 1–20. [Google Scholar] [CrossRef]
- Li, J.L.; Pan, B.Z.; Niu, L.J.; Chen, M.S.; Tang, M.Y.; Xu, Z.F. Gibberellin Inhibits Floral Initiation in the Perennial Woody Plant Jatropha curcas. J. Plant Growth Regul. 2018, 37, 999–1006. [Google Scholar] [CrossRef]
- Hui, W.K.; Wang, Y.; Chen, X.Y.; Zayed, M.Z.; Wu, G.J. Analysis of Transcriptional Responses of the Inflorescence Meristems in Jatropha curcas Following Gibberellin Treatment. Int. J. Mol. Sci. 2018, 19, 432. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Li, J.; Gao, X.; Zhang, D.; Li, A.; Liu, C. Genome-Wide Screening and Characterization of the Dof Gene Family in Physic Nut (Jatropha curcas L.). Int. J. Mol. Sci. 2018, 19, 1598. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wang, F.F.; Peng, X.J.; Huang, J.H.; Shen, S.H. Global Phosphoproteomic Analysis Reveals the Defense and Response Mechanisms of Jatropha Curcas Seedling under Chilling Stress. Int. J. Mol. Sci. 2019, 20, 208. [Google Scholar] [CrossRef]
- Avonce, N.; Mendoza-Vargas, A.; Morett, E.; Iturriaga, G. Insights on the evolution of trehalose biosynthesis. BMC Evol. Biol. 2006, 6, 1–15. [Google Scholar] [CrossRef]
- Lunn, J.E.; Delorge, I.; Figueroa, C.M.; van Dijck, P.; Stitt, M. Trehalose metabolism in plants. Plant J. 2014, 79, 544–567. [Google Scholar] [CrossRef]
- Habibur Rahman Pramanik, M.; Imai, R. Functional identification of a trehalose 6-phosphate phosphatase gene that is involved in transient induction of trehalose biosynthesis during chilling stress in rice. Plant Mol. Biol. 2005, 58, 751–762. [Google Scholar] [CrossRef]
- Li, P.; Ma, S.; Bohnert, H.J. Coexpression characteristics of trehalose-6-phosphate phosphatase subfamily genes reveal different functions in a network context. Physiol. Plant. 2008, 133, 544–556. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, O.; Vandesteene, L.; Feil, R.; Baillieul, F.; Lunn, J.E.; Clement, C. Trehalose metabolism is activated upon chilling in grapevine and might participate in Burkholderia phytofirmans induced chilling tolerance. Planta 2012, 236, 355–369. [Google Scholar] [CrossRef] [PubMed]
- Barrett, S.C.H. The evolution and adaptive significance of heterostyly. Trends Ecol. Evol. 1990, 5, 144–148. [Google Scholar] [CrossRef]
- Weller, S.G. The different forms of flowers—What have we learned since Darwin? Bot. J. Linn. Soc. 2009, 160, 249–261. [Google Scholar] [CrossRef]
- Cohen, J.I. “A case to which no parallel exists”: The influence of Darwin’s Different Forms of Flowers. Am. J. Bot. 2010, 97, 701–716. [Google Scholar] [CrossRef] [PubMed]
- Barrett, S.C.H.; Arroyo, M.T.K. Variation in floral morph ratios in tristylous Oxalis squamata (Oxalidaceae): an Andean alpine endemic. Botany 2012, 90, 1180–1185. [Google Scholar] [CrossRef]
- De Vos, J.M.; Hughes, C.E.; Schneeweiss, G.M.; Moore, B.R.; Conti, E. Heterostyly accelerates diversification via reduced extinction in primroses. Proc. R. Soc. B Biol. Sci. 2014, 281, 20140075. [Google Scholar] [CrossRef]
- Ganders, F.R. The biology of heterostyly. N. Z. J. Bot. 1979, 17, 607–635. [Google Scholar] [CrossRef]
- Tang, M.; Tao, Y.B.; Fu, Q.T.; Song, Y.L.; Niu, L.J.; Xu, Z.F. An ortholog of LEAFY in Jatropha curcas regulates flowering time and floral organ development. Sci. Rep. 2016, 6, 1–15. [Google Scholar] [CrossRef]
- Ni, J.; Gao, C.C.; Chen, M.S.; Pan, B.Z.; Ye, K.Q.; Xu, Z.F. Gibberellin promotes shoot branching in the perennial woody plant Jatropha curcas. Plant Cell Physiol. 2015, 56, 1655–1666. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]
- Fu, Q.T.; Li, C.Q.; Tang, M.Y.; Tao, Y.B.; Pan, B.Z.; Zhang, L.; Niu, L.J.; He, H.Y.; Wang, X.L.; Xu, Z.F. An efficient protocol for Agrobacterium-mediated transformation of the biofuel plant Jatropha curcas by optimizing kanamycin concentration and duration of delayed selection. Plant Biotechnol. Rep. 2015, 9, 405–416. [Google Scholar] [CrossRef]
- Zhang, L.; He, L.L.; Fu, Q.T.; Xu, Z.F. Selection of reliable reference genes for gene expression studies in the biofuel plant Jatropha curcas using real-time quantitative PCR. Int. J. Mol. Sci. 2013, 14, 24338–24354. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Fils-Lycaon, B.; Julianus, P.; Chillet, M.; Galas, C.; Hubert, O.; Rinaldo, D.; Mbéguié-A-Mbéguié, D. Acid invertase as a serious candidate to control the balance sucrose versus (glucose+fructose) of banana fruit during ripening. Sci. Hortic. 2011, 129, 197–206. [Google Scholar] [CrossRef]



© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, M.-L.; Ni, J.; Chen, M.-S.; Xu, Z.-F. Ectopic Expression of Jatropha curcas TREHALOSE-6-PHOSPHATE PHOSPHATASE J Causes Late-Flowering and Heterostylous Phenotypes in Arabidopsis but not in Jatropha. Int. J. Mol. Sci. 2019, 20, 2165. https://doi.org/10.3390/ijms20092165
Zhao M-L, Ni J, Chen M-S, Xu Z-F. Ectopic Expression of Jatropha curcas TREHALOSE-6-PHOSPHATE PHOSPHATASE J Causes Late-Flowering and Heterostylous Phenotypes in Arabidopsis but not in Jatropha. International Journal of Molecular Sciences. 2019; 20(9):2165. https://doi.org/10.3390/ijms20092165
Chicago/Turabian StyleZhao, Mei-Li, Jun Ni, Mao-Sheng Chen, and Zeng-Fu Xu. 2019. "Ectopic Expression of Jatropha curcas TREHALOSE-6-PHOSPHATE PHOSPHATASE J Causes Late-Flowering and Heterostylous Phenotypes in Arabidopsis but not in Jatropha" International Journal of Molecular Sciences 20, no. 9: 2165. https://doi.org/10.3390/ijms20092165
APA StyleZhao, M.-L., Ni, J., Chen, M.-S., & Xu, Z.-F. (2019). Ectopic Expression of Jatropha curcas TREHALOSE-6-PHOSPHATE PHOSPHATASE J Causes Late-Flowering and Heterostylous Phenotypes in Arabidopsis but not in Jatropha. International Journal of Molecular Sciences, 20(9), 2165. https://doi.org/10.3390/ijms20092165





