MoMCP1, a Cytochrome P450 Gene, Is Required for Alleviating Manganese Toxin Revealed by Transcriptomics Analysis in Magnaporthe oryzae
Abstract
1. Introduction
2. Results
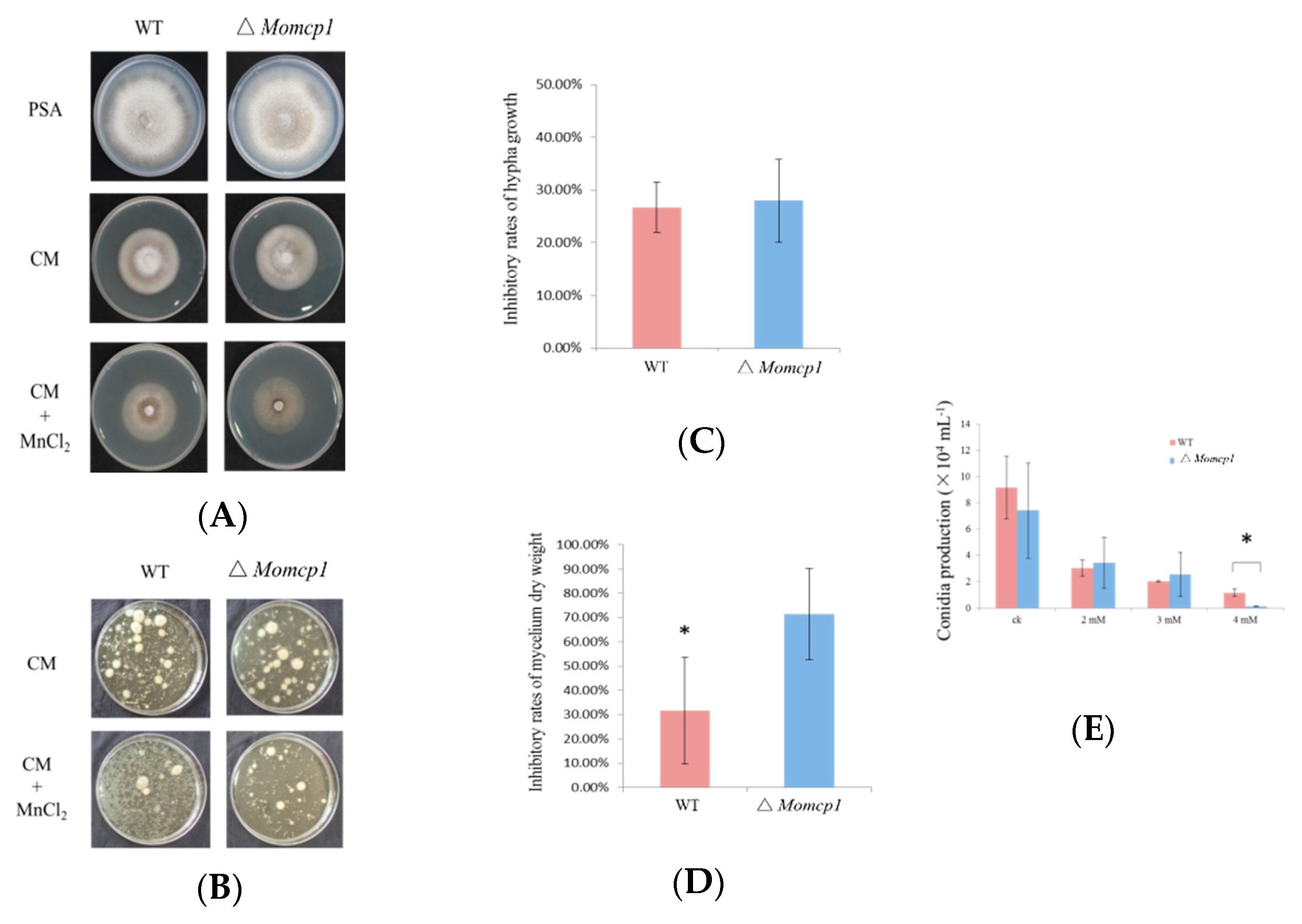
2.1. ∆Momcp1 Mutant Is More Sensitive to Mn2+
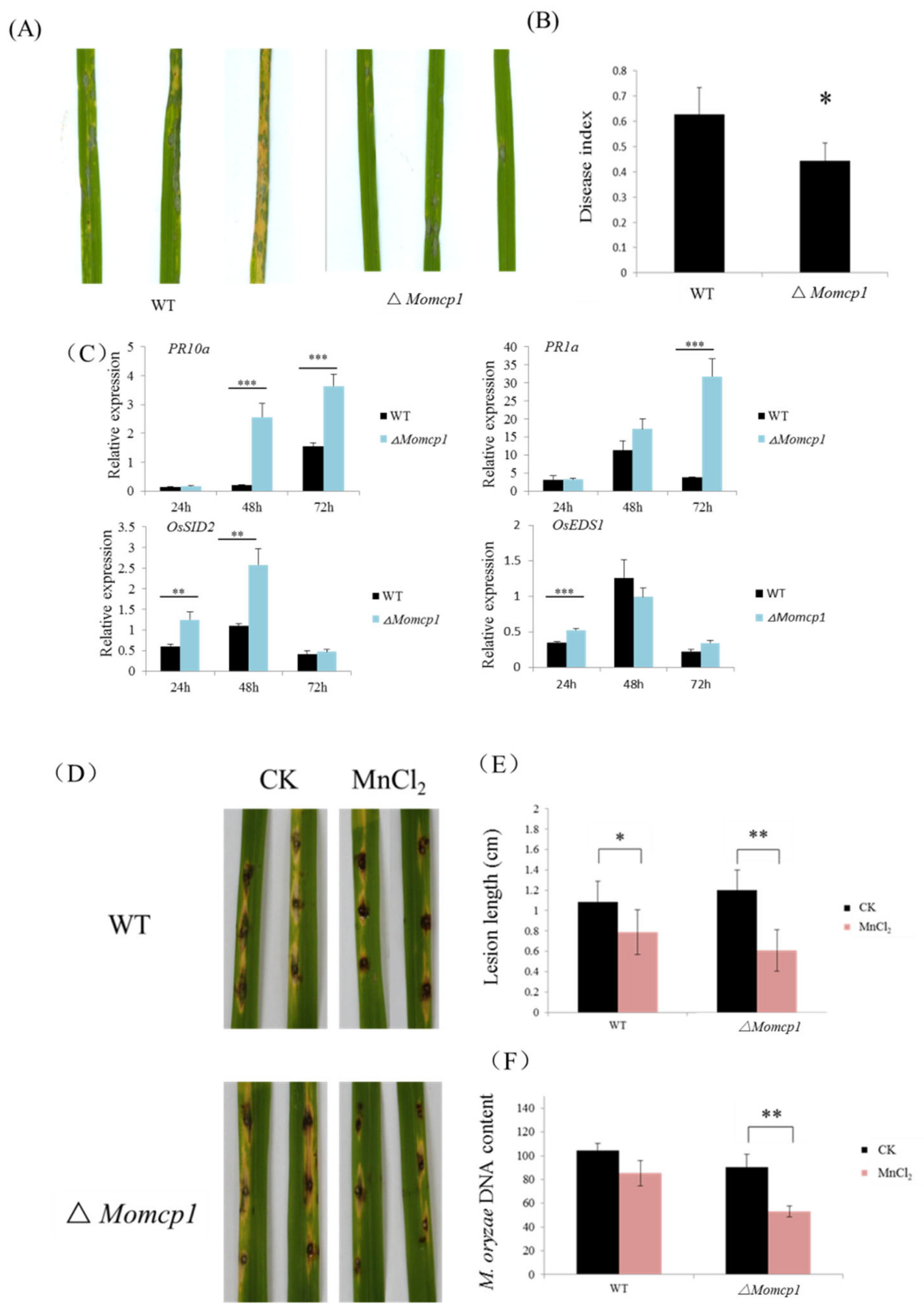
2.2. MoMCP1 Is Involved in the Pathogenicity of Rice
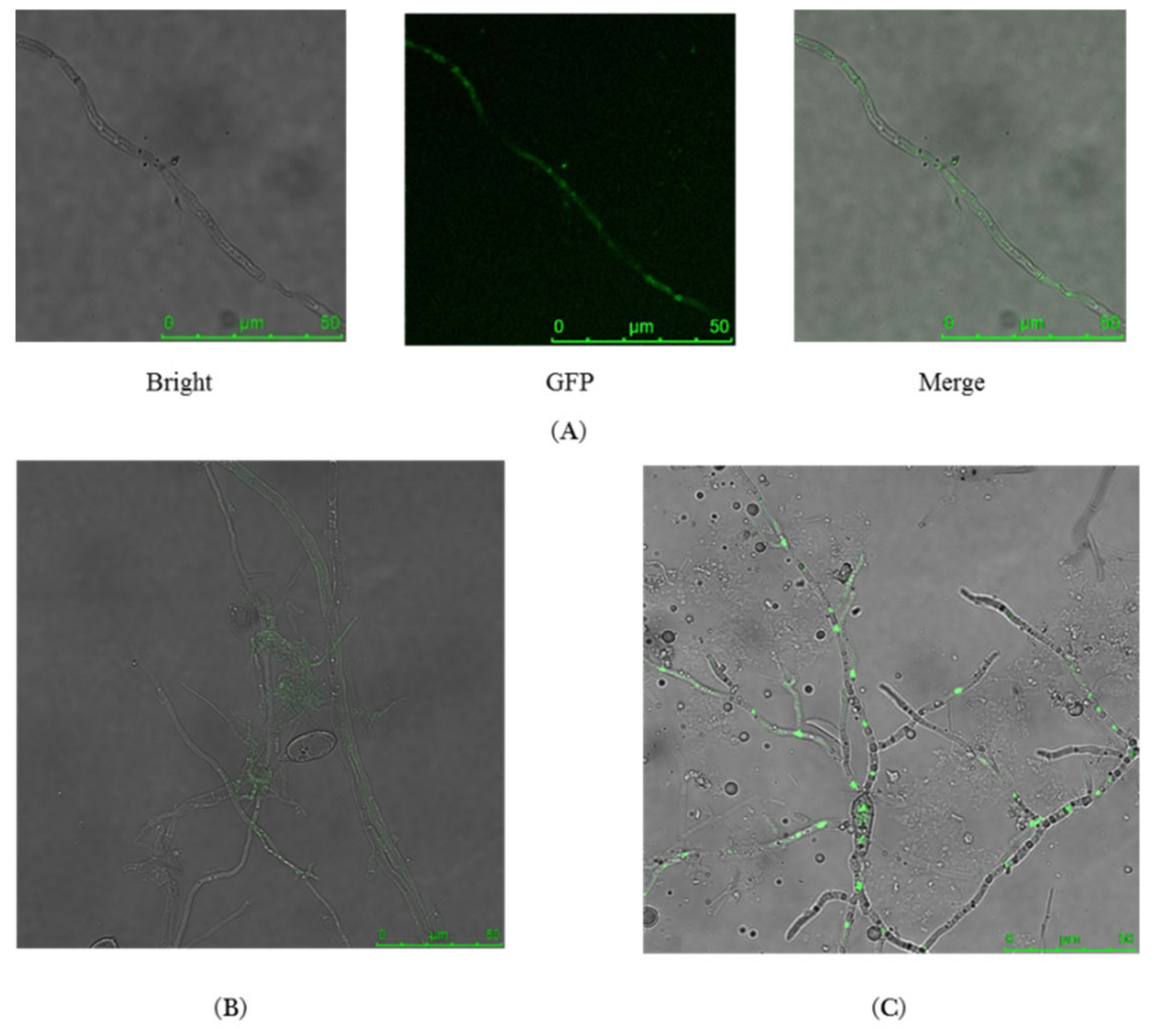
2.3. MoMCP1-GFP Fusion Protein Is Only Observed in Hypha
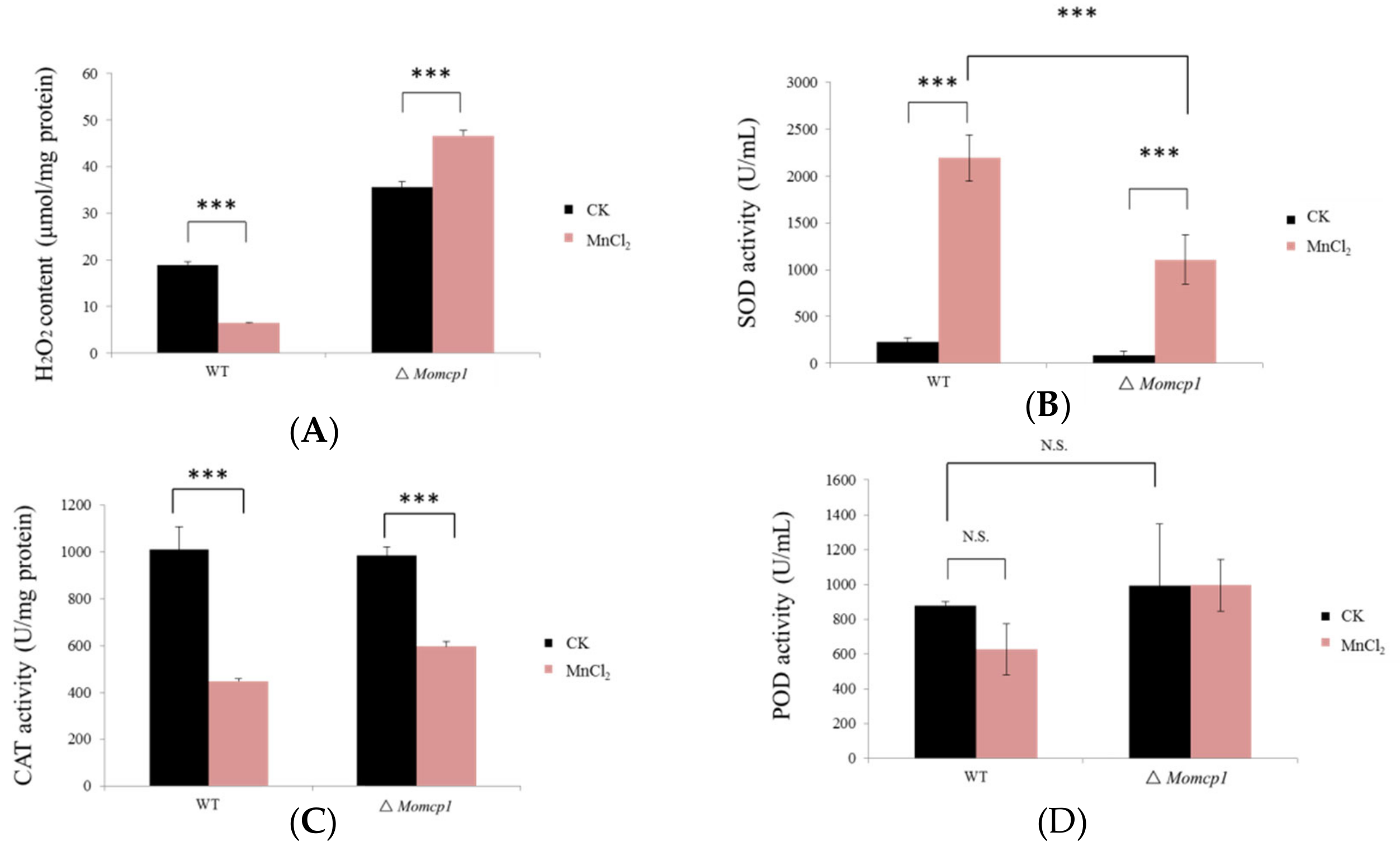
2.4. Excessive Mn2+ Increased Intracellular Content of H2O2 and Decreased ROS Related Enzymes in ∆Momcp1 Mutant
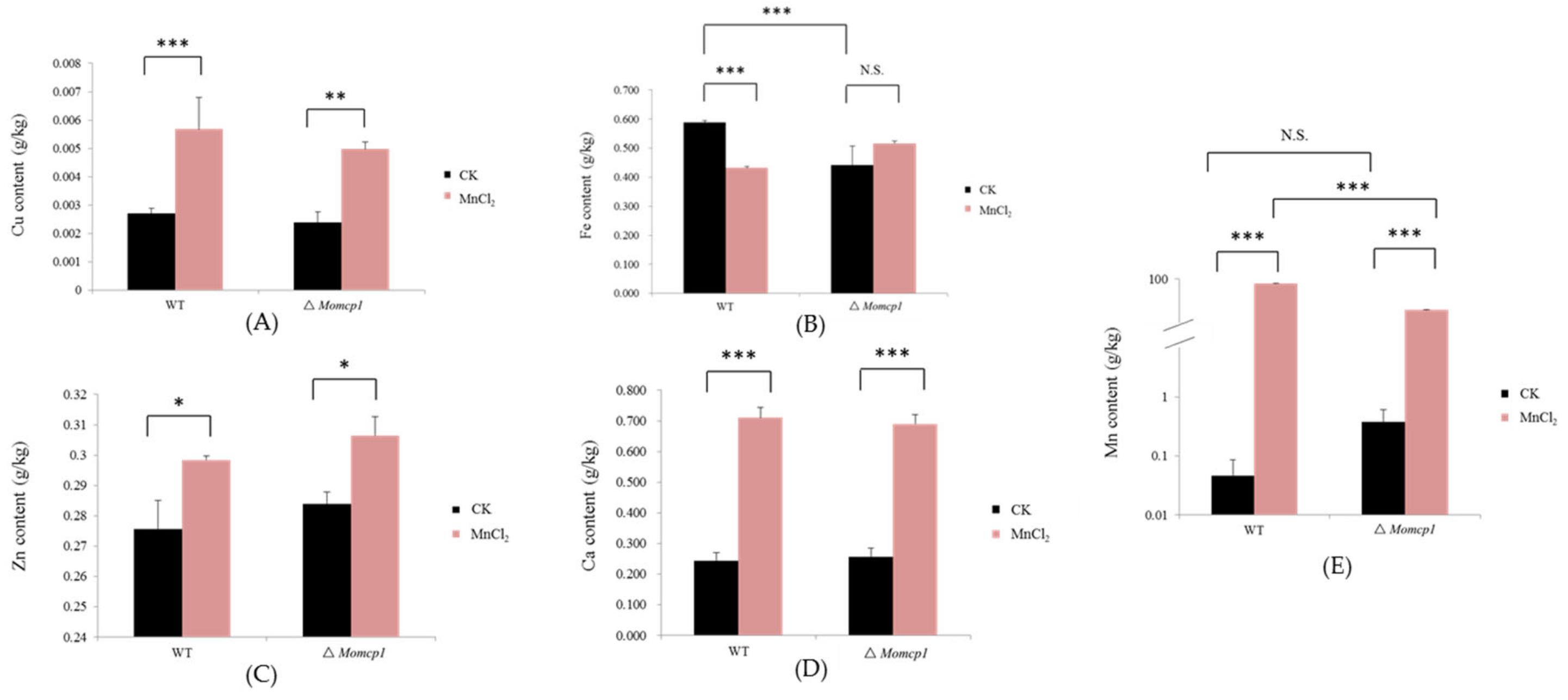
2.5. Excessive Mn Affected the Contents of Cu, Zn, Fe, and Ca in M. oryzae
2.6. Differential Gene Expression Analysis between ∆Momcp1 Mutant and WT Strains Using Transcriptomes
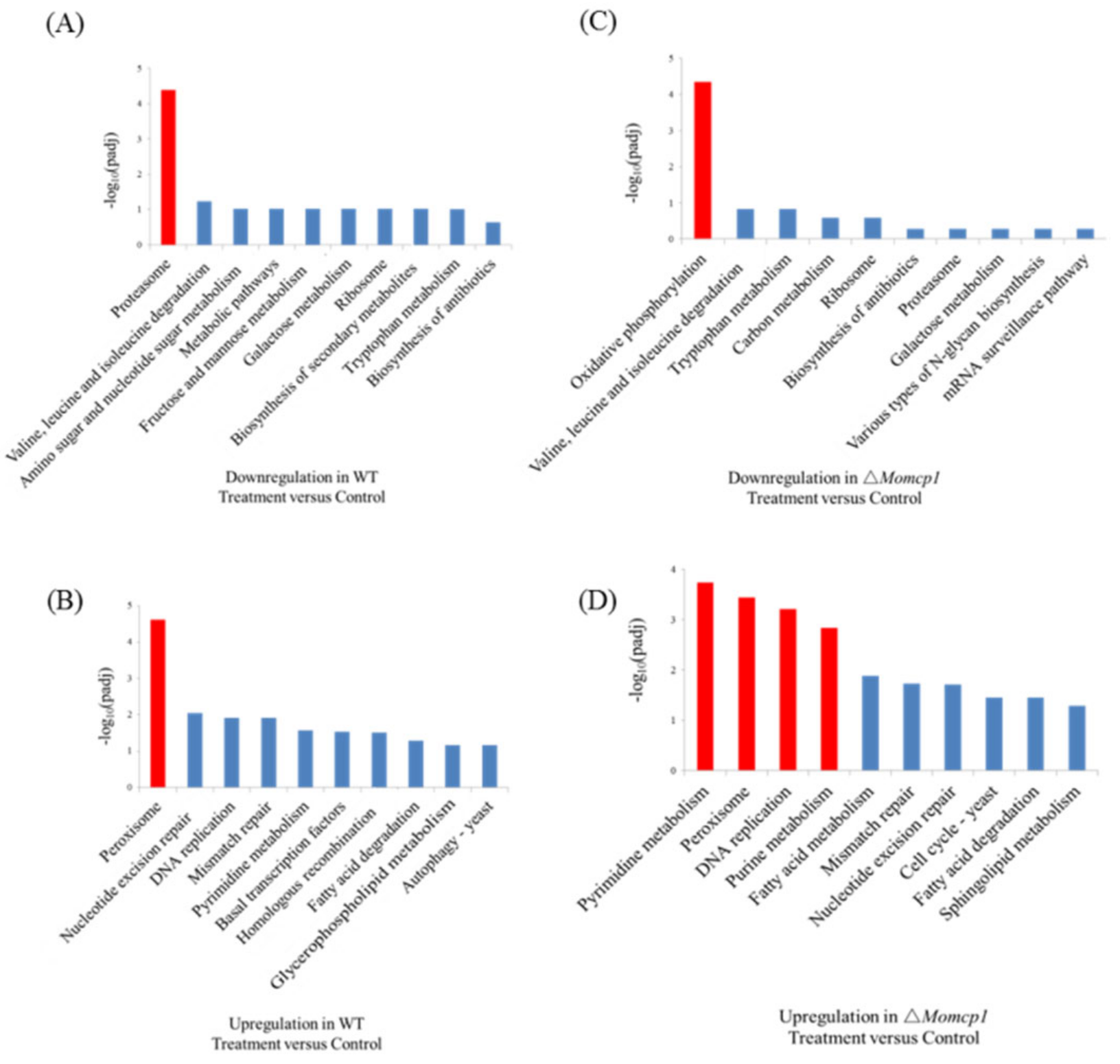
2.7. Transcriptome Analysis of the Sensitivity of Mn2+ in △Momcp1 Mutant
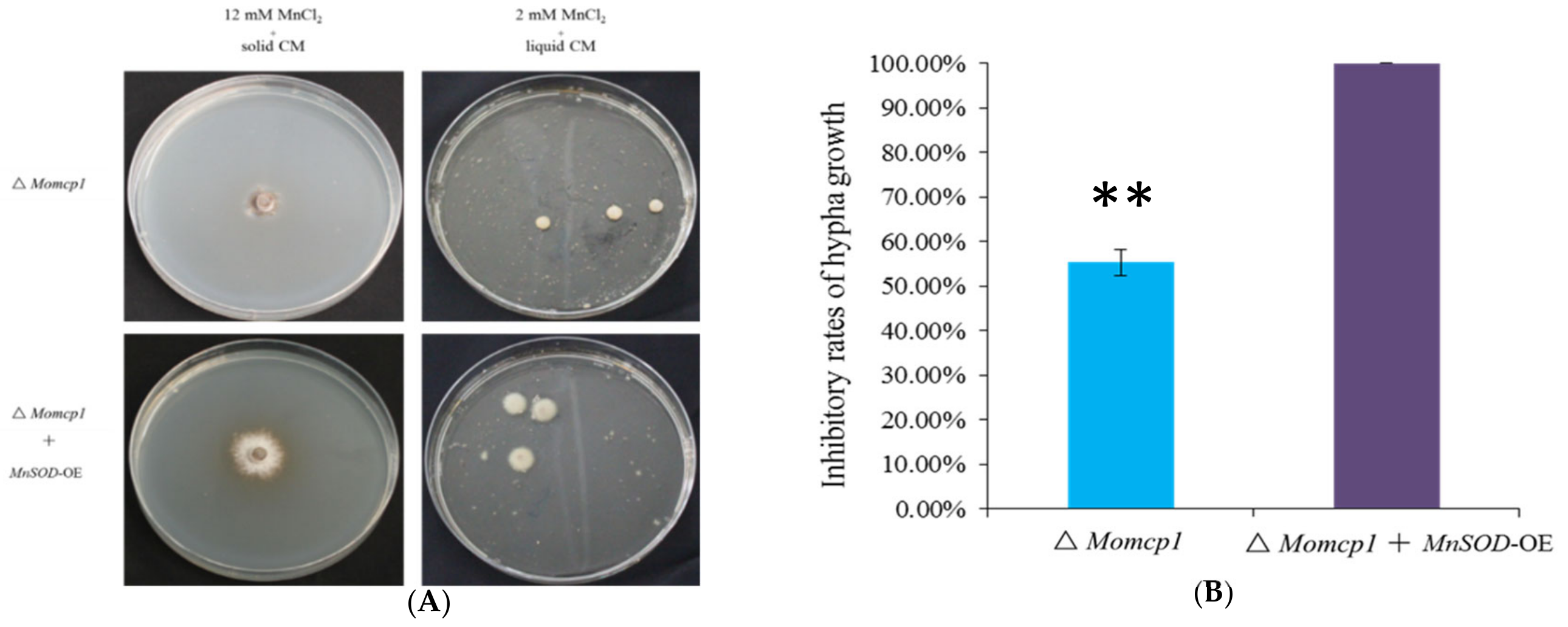
2.8. Overexpression of MnSOD Restored the Mn2+ Tolerance in ∆Momcp1 Mutant
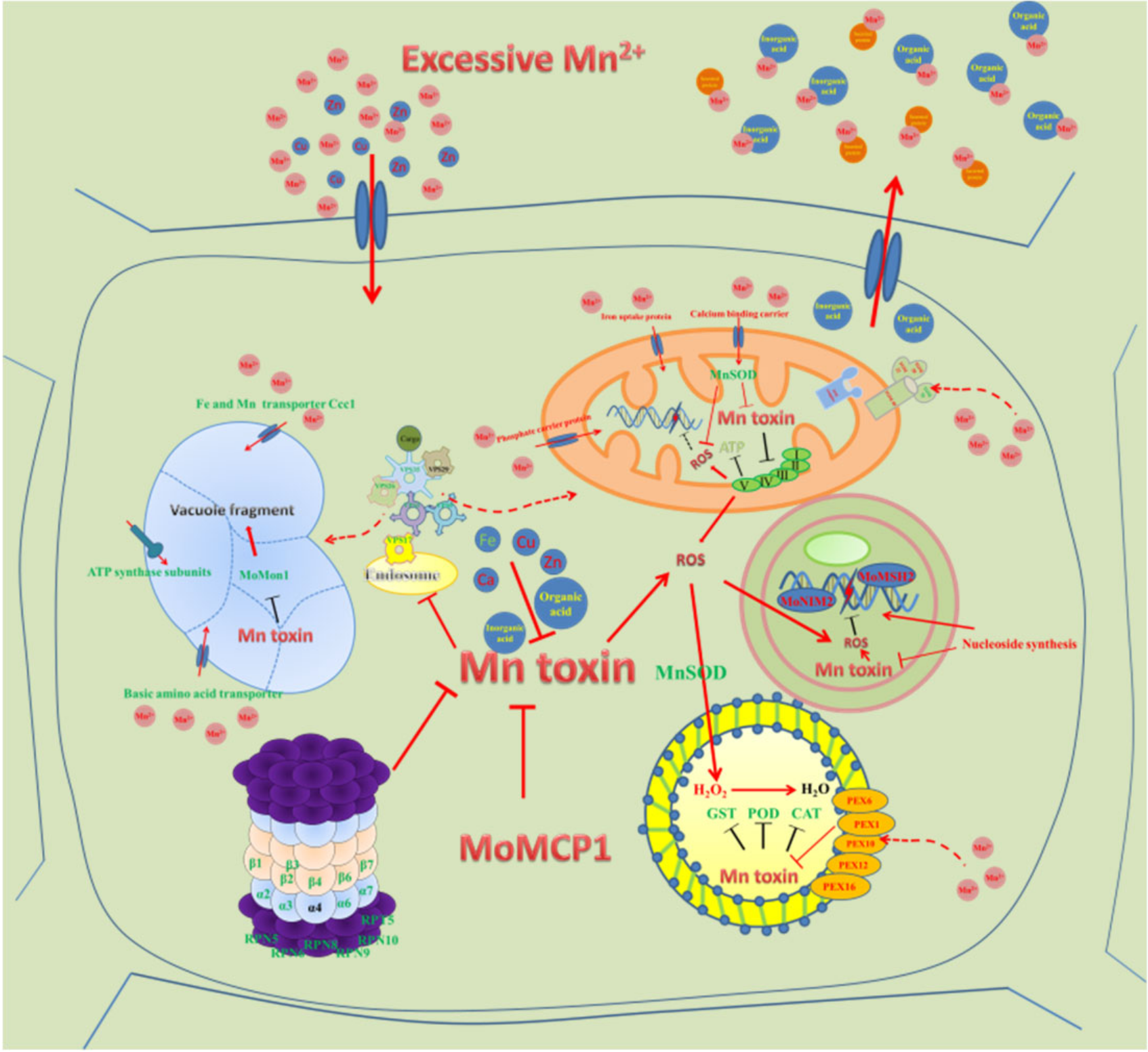
3. Discussion
3.1. Pathogenicity Related Genes
3.2. Mitochondria Related Genes
3.3. Vacuole Related Genes
3.4. Peroxisome Related Genes, Especially MnSOD, Detoxify Mn2+ Induced Oxidative Stress
3.5. Proteasome Related Genes
3.6. Nucleotide Synthesis Related Genes
3.7. Secreted Compound and Transport Related Genes
3.8. Changes of Other Metals by Excessive Mn2+
3.9. Cytochrome P450 Genes and Mn Stress
4. Materials and Methods
4.1. Strains and Growth Conditions
4.2. Mycelial Growth, Mycelial Dry Weight, Conidial Production, Germination, Appressorium Formation and Plant Infection Assays
4.3. Gene Deletion
4.4. RNA Isolation and RT-qPCR
4.5. RNAseq Analysis
4.6. Generation of Transformants Expressing the MoMCP1–eGFP
4.7. Overexpression of MnSOD in ∆Momcp1
4.8. Determination of H2O2 Content, Activities of SOD, CAT and POD
4.9. Determination of Different Metal Contents by ICP-OES
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| WT | Wild type |
| CYP450 | Cytochrome P450 |
| OsNramp5 | Natural resistance associated macrophage protein 5 |
| OsMTP9 | Metal tolerance protein 9 |
| OsYSL6 | Yellow stripe-like protein 6 |
| ROS | Reactive oxygen species |
| CAT | Catalase |
| SOD | Superoxide dismutase |
| POD | Peroxidase |
| dpi | Day post inoculation |
| hpi | Hour post inoculation |
| DEG | Differently expressed gene |
| TOM | Translocase of the outer mitochondrial membrane |
| TIM | Translocase of the inner mitochondrial membrane |
| VPS | Vacuolar protein sorting-associated genes |
| PEX | Peroxin |
References
- Aguirre, J.D.; Culotta, V.C. Battles with iron: Manganese in oxidative stress protection. J. Biol. Chem. 2012, 287, 13541–13548. [Google Scholar] [CrossRef] [PubMed]
- Dambach, M.; Sandoval, M.; Updegrove, T.B.; Anantharaman, V.; Aravind, L.; Waters, L.S.; Storz, G. The ubiquitous yybP-ykoY riboswitch is a manganese-responsive regulatory element. Mol. Cell 2015, 57, 1099–1109. [Google Scholar] [CrossRef] [PubMed]
- Kwakye, G.F.; Paoliello, M.M.; Mukhopadhyay, S.; Bowman, A.B.; Aschner, M. Manganese-induced Parkinsonism and Parkinson’s disease: Shared and distinguishable features. Int. J. Environ. Res. Public Health 2015, 12, 7519–7540. [Google Scholar] [CrossRef] [PubMed]
- Fuhrs, H.; Gotze, S.; Specht, A.; Erban, A.; Gallien, S.; Heintz, D.; Van Dorsselaer, A.; Kopka, J.; Braun, H.P.; Horst, W.J. Characterization of leaf apoplastic peroxidases and metabolites in Vigna unguiculata in response to toxic manganese supply and silicon. J. Exp. Bot. 2009, 60, 1663–1678. [Google Scholar] [CrossRef]
- Sasaki, A.; Yamaji, N.; Yokosho, K.; Ma, J.F. Nramp5 is a major transporter responsible for manganese and cadmium uptake in rice. Plant Cell 2012, 24, 2155–2167. [Google Scholar] [CrossRef] [PubMed]
- Ueno, D.; Sasaki, A.; Yamaji, N.; Miyaji, T.; Fujii, Y.; Takemoto, Y.; Moriyama, S.; Che, J.; Moriyama, Y.; Iwasaki, K.; et al. A polarly localized transporter for efficient manganese uptake in rice. Nat. Plants 2015, 1, 15170. [Google Scholar] [CrossRef]
- Sasaki, A.; Yamaji, N.; Xia, J.; Ma, J.F. OsYSL6 is involved in the detoxification of excess manganese in rice. Plant Physiol. 2011, 157, 1832–1840. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Fujii, Y.; Yamaji, N.; Masuda, S.; Takemoto, Y.; Kamiya, T.; Yusuyin, Y.; Iwasaki, K.; Kato, S.; Maeshima, M.; et al. Mn tolerance in rice is mediated by MTP8.1, a member of the cation diffusion facilitator family. J. Exp. Bot. 2013, 64, 4375–4387. [Google Scholar] [CrossRef]
- Ma, G.; Li, J.; Li, J.; Li, Y.; Gu, D.; Chen, C.; Cui, J.; Chen, X.; Zhang, W. OsMTP11, a trans-Golgi network localized transporter, is involved in manganese tolerance in rice. Plant Sci. 2018, 274, 59–69. [Google Scholar] [CrossRef]
- Reddi, A.R.; Jensen, L.T.; Naranuntarat, A.; Rosenfeld, L.; Leung, E.; Shah, R.; Culotta, V.C. The overlapping roles of manganese and Cu/Zn SOD in oxidative stress protection. Free Radic. Biol. Med. 2009, 46, 154–162. [Google Scholar] [CrossRef]
- Luk, E.E.-C.; Culotta, V.C. Manganese superoxide dismutase in Saccharomyces cerevisiae acquires its metal co-factor through a pathway involving the nramp metal transporter, Smf2p. J. Biol. Chem. 2001, 276, 47556–47562. [Google Scholar] [CrossRef]
- Jensen, L.T.; Carroll, M.C.; Hall, M.D.; Harvey, C.J.; Beese, S.E.; Culotta, V.C. Down-regulation of a manganese transporter in the face of metal toxin. Mol. Biol. Cell 2009, 20, 2810–2819. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Chen, O.S.; McVey Ward, D.; Kaplan, J. CCC1 is a transporter that mediates vacuolar iron storage in yeast. J. Biol. Chem. 2001, 276, 29515–29519. [Google Scholar] [CrossRef] [PubMed]
- Lisher, J.P.; Giedroc, D.P. Manganese acquisition and homeostasis at the host-pathogen interface. Front. Cell Infect. Microbiol. 2013, 3, 91. [Google Scholar] [CrossRef] [PubMed]
- Corbin, B.D.; Seeley, E.H.; Raab, A.; Feldmann, J.; Miller, M.R.; Torres, V.J.; Anderson, K.L.; Dattilo, B.M.; Dunman, P.M.; Gerads, R.; et al. Metal Chelation and Inhibition of Bacterial Growth in Tissue Abscesses. Science 2008, 319, 962–965. [Google Scholar] [CrossRef] [PubMed]
- Jabado, N.; Jankowski, A.; Dougaparsad, S.; Picard, V.; Grinstein, S.; Gros, P. Natural resistance to intracellular infections: Natural resistance–associated macrophage protein 1 (Nramp1) functions as a pH-dependent manganese transporter at the phagosomal membrane. J. Exp. Med. 2000, 192, 1237–1248. [Google Scholar] [CrossRef]
- Juttukonda, L.J.; Skaar, E.P. Manganese homeostasis and utilization in pathogenic bacteria. Mol. Microbiol. 2015, 97, 216–228. [Google Scholar] [CrossRef]
- Radin, J.N.; Kelliher, J.L.; Parraga Solorzano, P.K.; Kehl-Fie, T.E. The two-component system ArlRS and alterations in metabolism enable Staphylococcus aureus to resist calprotectin-Induced Manganese Starvation. PLoS Pathog. 2016, 12, e1006040. [Google Scholar] [CrossRef]
- Si, M.; Zhao, C.; Burkinshaw, B.; Zhang, B.; Wei, D.; Wang, Y.; Dong, T.G.; Shen, X. Manganese scavenging and oxidative stress response mediated by type VI secretion system in Burkholderia thailandensis. Proc. Natl. Acad. Sci. USA 2017, 114, E2233–E2242. [Google Scholar] [CrossRef]
- Wang, C.; Guan, Y.; Lv, M.; Zhang, R.; Guo, Z.; Wei, X.; Du, X.; Yang, J.; Li, T.; Wan, Y.; et al. Manganese increases the sensitivity of the cGAS-STING pathway for double-Stranded DNA and is required for the host defense against DNA viruses. Immunity 2018, 48, 675–687.e7. [Google Scholar] [CrossRef]
- Li, W.; Zhu, Z.; Chern, M.; Yin, J.; Yang, C.; Ran, L.; Cheng, M.; He, M.; Wang, K.; Wang, J.; et al. A natural allele of a transcription factor in rice confers broad-spectrum blast resistance. Cell 2017, 170, 114–126. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Cao, X.L.; Zhu, Y.; Yang, X.M.; Zhang, K.N.; Xiao, Z.Y.; Wang, H.; Zhao, J.H.; Zhang, L.L.; Li, G.B.; et al. Osa-miR398b boosts H2O2 production and rice blast disease-resistance via multiple superoxide dismutases. New Phytol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Talbot, N.J.; Kershaw, M.J.; Wakley, G.E.; De Vries, O.M.; Wessels, J.G.; Hamer, J.E. MPG1 encodes a fungal hydrophobin involved in surface interactions during infection-related development of Magnaporthe grisea. Plant Cell 1996, 8, 985–999. [Google Scholar] [CrossRef] [PubMed]
- DeZwaan, T.M.; Carroll, A.M.; Valent, B.; Sweigard, J.A. Magnaporthe grisea pth11p is a novel plasma membrane protein that mediates appressorium differentiation in response to inductive substrate cues. Plant Cell 1999, 11, 2013–2030. [Google Scholar] [CrossRef] [PubMed]
- Dagdas, Y.F.; Yoshino, K.; Dagdas, G.; Ryder, L.S.; Bielska, E.; Steinberg, G.; Talbot, N.J. Septin-mediated plant cell invasion by the rice blast fungus, Magnaporthe oryzae. Science 2012, 336, 1590–1595. [Google Scholar] [CrossRef]
- Guo, M.; Tan, L.; Nie, X.; Zhang, Z. A class-II myosin is required for growth, conidiation, cell wall integrity and pathogenicity of Magnaporthe oryzae. Virulence 2017, 8, 1335–1354. [Google Scholar] [CrossRef] [PubMed]
- Egan, M.J.; Wang, Z.Y.; Jones, M.A.; Smirnoff, N.; Talbot, N.J. Generation of reactive oxygen species by fungal NADPH oxidases is required for rice blast disease. Proc. Natl. Acad. Sci. USA 2007, 104, 11772–11777. [Google Scholar] [CrossRef]
- Spence, C.A.; Lakshmanan, V.; Donofrio, N.; Bais, H.P. Crucial Roles of Abscisic Acid Biogenesis in Virulence of Rice Blast Fungus Magnaporthe oryzae. Front. Plant Sci. 2015, 6, 1082. [Google Scholar] [CrossRef]
- Kong, S.; Park, S.Y.; Lee, Y.H. Systematic characterization of the bZIP transcription factor gene family in the rice blast fungus, Magnaporthe oryzae. Environ. Microbiol. 2015, 17, 1425–1443. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Ma, W.B.; Li, Y.; Wang, H.; Que, Y.W.; Ma, Z.H.; Talbot, N.J.; Wang, Z.Y. A sterol 14alpha-demethylase is required for conidiation, virulence and for mediating sensitivity to sterol demethylation inhibitors by the rice blast fungus Magnaporthe oryzae. Fungal Genet. Biol. 2011, 48, 144–153. [Google Scholar] [CrossRef]
- Kim, S.; Park, S.Y.; Kim, K.S.; Rho, H.S.; Chi, M.H.; Choi, J.; Park, J.; Kong, S.; Park, J.; Goh, J.; et al. Homeobox transcription factors are required for conidiation and appressorium development in the rice blast fungus Magnaporthe oryzae. PLoS Genet. 2009, 5, e1000757. [Google Scholar] [CrossRef]
- Sun, D.; Cao, H.; Shi, Y.; Huang, P.; Dong, B.; Liu, X.; Lin, F.; Lu, J. The regulatory factor X protein MoRfx1 is required for development and pathogenicity in the rice blast fungus Magnaporthe oryzae. Mol. Plant Pathol. 2017, 18, 1075–1088. [Google Scholar] [CrossRef]
- Kong, L.A.; Yang, J.; Li, G.T.; Qi, L.L.; Zhang, Y.J.; Wang, C.F.; Zhao, W.S.; Xu, J.R.; Peng, Y.L. Different chitin synthase genes are required for various developmental and plant infection processes in the rice blast fungus Magnaporthe oryzae. PLoS Pathog. 2012, 8, e1002526. [Google Scholar] [CrossRef]
- Guo, M.; Chen, Y.; Du, Y.; Dong, Y.; Guo, W.; Zhai, S.; Zhang, H.; Dong, S.; Zhang, Z.; Wang, Y.; et al. The bZIP transcription factor MoAP1 mediates the oxidative stress response and is critical for pathogenicity of the rice blast fungus Magnaporthe oryzae. PLoS Pathog. 2011, 7, e1001302. [Google Scholar] [CrossRef]
- Gao, H.M.; Liu, X.G.; Shi, H.B.; Lu, J.P.; Yang, J.; Lin, F.C.; Liu, X.H. MoMon1 is required for vacuolar assembly, conidiogenesis and pathogenicity in the rice blast fungus Magnaporthe oryzae. Res. Microbiol. 2013, 164, 300–309. [Google Scholar] [CrossRef]
- Samalova, M.; Melida, H.; Vilaplana, F.; Bulone, V.; Soanes, D.M.; Talbot, N.J.; Gurr, S.J. The beta-1,3-glucanosyltransferases (Gels) affect the structure of the rice blast fungal cell wall during appressorium-mediated plant infection. Cell. Microbiol. 2017, 19, e12659. [Google Scholar] [CrossRef]
- Qi, Y.; Marlin, M.C.; Liang, Z.; Berry, W.L.; Janknecht, R.; Zhou, J.; Wang, Z.; Lu, G.; Li, G. Distinct biochemical and functional properties of two Rab5 homologs from the rice blast fungus Magnaporthe oryzae. J. Biol. Chem. 2014, 289, 28299–28309. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Guo, Z.; Xi, Y.; Yuan, M.; Lin, Y.; Wu, C.; Abubakar, Y.S.; Dou, X.; Li, G.; Wang, Z.; et al. Sorting nexin (MoVps17) is required for fungal development and plant infection by regulating endosome dynamics in the rice blast fungus. Environ. Microbiol. 2017, 19, 4301–4317. [Google Scholar] [PubMed]
- Goh, J.; Jeon, J.; Lee, Y.H. ER retention receptor, MoERR1 is required for fungal development and pathogenicity in the rice blast fungus, Magnaporthe oryzae. Sci. Rep. 2017, 7, 1259. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Liu, W.; Wang, C.; Xu, Q.; Wang, Y.; Ding, S.; Xu, J.R. A MADS-box transcription factor MoMcm1 is required for male fertility, microconidium production and virulence in Magnaporthe oryzae. Mol. Microbiol. 2011, 80, 33–53. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Liu, M.; Liu, X.; Yin, Z.; Sun, Y.; Zhang, H.; Zheng, X.; Wang, P.; Zhang, Z. Heat-Shock Proteins MoSsb1, MoSsz1, and MoZuo1 Attenuate MoMkk1-Mediated Cell-Wall Integrity Signaling and Are Important for Growth and Pathogenicity of Magnaporthe oryzae. Mol. Plant Microbe Interact. 2018, 31, 1211–1221. [Google Scholar] [CrossRef]
- Galhano, R.; Illana, A.; Ryder, L.S.; Rodriguez-Romero, J.; Demuez, M.; Badaruddin, M.; Martinez-Rocha, A.L.; Soanes, D.M.; Studholme, D.J.; Talbot, N.J.; et al. Tpc1 is an important Zn(II)2Cys6 transcriptional regulator required for polarized growth and virulence in the rice blast fungus. PLoS Pathog. 2017, 13, e1006516. [Google Scholar] [CrossRef]
- Yang, L.; Ru, Y.; Cai, X.; Yin, Z.; Liu, X.; Xiao, Y.; Zhang, H.; Zheng, X.; Wang, P.; Zhang, Z. MoImd4 mediates crosstalk between MoPdeH-cAMP signalling and purine metabolism to govern growth and pathogenicity in Magnaporthe oryzae. Mol. Plant Pathol. 2019, 20, 500–518. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yang, J.; Qian, B.; Cai, Y.; Zou, X.; Zhang, H.; Zheng, X.; Wang, P.; Zhang, Z. MoYvh1 subverts rice defense through functions of ribosomal protein MoMrt4 in Magnaporthe oryzae. PLoS Pathog. 2018, 14, e1007016. [Google Scholar] [CrossRef]
- Sabnam, N.; Roy Barman, S. WISH, a novel CFEM GPCR is indispensable for surface sensing, asexual and pathogenic differentiation in rice blast fungus. Fungal Genet. Biol. 2017, 105, 37–51. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Ahn, I.P.; Rho, H.S.; Lee, Y.H. MHP1, a Magnaporthe grisea hydrophobin gene, is required for fungal development and plant colonization. Mol. Microbiol. 2005, 57, 1224–1237. [Google Scholar] [CrossRef] [PubMed]
- Kulkarni, R.D.; Thon, M.R.; Pan, H.; Dean, R.A. Novel G-protein-coupled receptor-like proteins in the plant pathogenic fungus Magnaporthe grisea. Genome Biol. 2005, 6, R24. [Google Scholar] [CrossRef] [PubMed]
- Chung, H.; Choi, J.; Park, S.Y.; Jeon, J.; Lee, Y.H. Two conidiation-related Zn(II)2Cys6 transcription factor genes in the rice blast fungus. Fungal Genet. Biol. 2013, 61, 133–141. [Google Scholar] [CrossRef]
- Kwon, S.; Lee, J.; Jeon, J.; Kim, S.; Park, S.Y.; Jeon, J.; Lee, Y.H. Role of the Histone Acetyltransferase Rtt109 in Development and Pathogenicity of the Rice Blast Fungus. Mol. Plant Microbe Interact. 2018, 31, 1200–1210. [Google Scholar] [CrossRef] [PubMed]
- Jeon, J.; Rho, H.; Kim, S.; Kim, K.S.; Lee, Y.H. Role of MoAND1-mediated nuclear positioning in morphogenesis and pathogenicity in the rice blast fungus, Magnaporthe oryzae. Fungal Genet. Biol. 2014, 69, 43–51. [Google Scholar] [CrossRef]
- Motoyama, T.; Ochiai, N.; Morita, M.; Iida, Y.; Usami, R.; Kudo, T. Involvement of putative response regulator genes of the rice blast fungus Magnaporthe oryzae in osmotic stress response, fungicide action, and pathogenicity. Curr. Genet. 2008, 54, 185–195. [Google Scholar] [CrossRef]
- Bruno, K.S.; Tenjo, F.; Li, L.; Hamer, J.E.; Xu, J.R. Cellular localization and role of kinase activity of PMK1 in Magnaporthe grisea. Eukaryot. Cell 2004, 3, 1525–1532. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, J.; Wright, J.D.; Hartline, D.; Quispe, C.F.; Madayiputhiya, N.; Wilson, R.A. Principles of carbon catabolite repression in the rice blast fungus: Tps1, Nmr1-3, and a MATE–Family Pump regulate glucose metabolism during Infection. PLoS Genet. 2012, 8, e1002673. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.C.; Ham, K.S.; Darvill, A.G.; Albersheim, P. Deletion of two endo-β-1, 4-xylanase genes reveals additional isozymes secreted by the rice blast fungus. Mol. Plant Microbe Interact. 1997, 10, 700–708. [Google Scholar] [CrossRef]
- Yi, M.; Chi, M.H.; Khang, C.H.; Park, S.Y.; Kang, S.; Valent, B.; Lee, Y.H. The ER chaperone LHS1 is involved in asexual development and rice infection by the blast fungus Magnaporthe oryzae. Plant Cell 2009, 21, 681–695. [Google Scholar] [CrossRef] [PubMed]
- Yi, M.; Park, J.H.; Ahn, J.H.; Lee, Y.H. MoSNF1 regulates sporulation and pathogenicity in the rice blast fungus Magnaporthe oryzae. Fungal Genet. Biol. 2008, 45, 1172–1181. [Google Scholar] [CrossRef]
- Oses-Ruiz, M.; Sakulkoo, W.; Littlejohn, G.R.; Martin-Urdiroz, M.; Talbot, N.J. Two independent S-phase checkpoints regulate appressorium-mediated plant infection by the rice blast fungus Magnaporthe oryzae. Proc. Natl. Acad. Sci. USA 2017, 114, E237–E244. [Google Scholar] [CrossRef] [PubMed]
- Gavin, C.E.; Gunter, K.K.; Gunter, T.E. Mn2+ sequestration by mitochondria and inhibition of oxidative phosphorylation. Toxicol. Appl. Pharmacol. 1992, 115, 1–5. [Google Scholar] [CrossRef]
- Luk, E.; Carroll, M.; Baker, M.; Culotta, V.C. Manganese activation of superoxide dismutase 2 in Saccharomyces cerevisiae requires MTM1, a member of the mitochondrial carrier family. Proc. Natl. Acad. Sci. USA 2003, 100, 10353–10357. [Google Scholar] [CrossRef] [PubMed]
- Harbauer, A.B.; Zahedi, R.P.; Sickmann, A.; Pfanner, N.; Meisinger, C. The protein import machinery of mitochondria-a regulatory hub in metabolism, stress, and disease. Cell Metab. 2014, 19, 357–372. [Google Scholar] [CrossRef] [PubMed]
- Becker, T.; Bottinger, L.; Pfanner, N. Mitochondrial protein import: From transport pathways to an integrated network. Trends Biochem. Sci. 2012, 37, 85–91. [Google Scholar] [CrossRef]
- Stewart, J.B.; Chinnery, P.F. The dynamics of mitochondrial DNA heteroplasmy: Implications for human health and disease. Nat. Rev. Genet. 2015, 16, 530–542. [Google Scholar] [CrossRef]
- Kaur, G.; Kumar, V.; Arora, A.; Tomar, A.; Ashish; Sur, R.; Dutta, D. Affected energy metabolism under manganese stress governs cellular toxin. Sci. Rep. 2017, 7, 11645. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.S.; Dixon, E.W.; Vincelli, P.; Farman, M.L. Field resistance to strobilurin (QoI) fungicides in Pyricularia grisea caused by mutations in the mitochondrial cytochrome b gene. Phytopathology 2003, 93, 891–900. [Google Scholar] [CrossRef]
- Seaman, M.N.; McCaffery, J.M.; Emr, S.D. A membrane coat complex essential for endosome-to-Golgi retrograde transport in yeast. J. Cell Biol. 1998, 142, 665–681. [Google Scholar] [CrossRef] [PubMed]
- Thorsen, M.; Perrone, G.G.; Kristiansson, E.; Traini, M.; Ye, T.; Dawes, I.W.; Nerman, O.; Tamas, M.J. Genetic basis of arsenite and cadmium tolerance in Saccharomyces cerevisiae. BMC Genom. 2009, 10, 105. [Google Scholar] [CrossRef]
- Caza, M.; Hu, G.; Nielson, E.D.; Cho, M.; Jung, W.H.; Kronstad, J.W. The Sec1/Munc18 (SM) protein Vps45 is involved in iron uptake, mitochondrial function and virulence in the pathogenic fungus Cryptococcus neoformans. PLoS Pathog. 2018, 14, e1007220. [Google Scholar] [CrossRef]
- Zheng, W.; Zhou, J.; He, Y.; Xie, Q.; Chen, A.; Zheng, H.; Shi, L.; Zhao, X.; Zhang, C.; Huang, Q.; et al. Retromer is essential for autophagy-dependent plant infection by the rice blast fungus. PLoS Genet. 2015, 11, e1005704. [Google Scholar] [CrossRef] [PubMed]
- Zheng, W.; Zheng, H.; Zhao, X.; Zhang, Y.; Xie, Q.; Lin, X.; Chen, A.; Yu, W.; Lu, G.; Shim, W.B.; et al. Retrograde trafficking from the endosome to the trans-Golgi network mediated by the retromer is required for fungal development and pathogenicity in Fusarium graminearum. New Phytol. 2016, 210, 1327–1343. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, S.; Rokad, D.; Malovic, E.; Luo, J.; Harischandra, D.S.; Jin, H.; Kanthasamy, A.G. Manganese activates NLRP3 inflammasome signaling and propagates exosomal release of ASC in microglial cells. Sci. Signal 2019, 12, eaat9900. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.M.; Liang, S.; Shi, H.B.; Lu, J.P.; Dong, B.; Liao, Q.S.; Lin, F.C.; Liu, X.H. VPS9 domain-containing proteins are essential for autophagy and endocytosis in Pyricularia oryzae. Environ. Microbiol. 2018, 20, 1516–1530. [Google Scholar] [CrossRef]
- Li, Y.; Li, B.; Liu, L.; Chen, H.; Zhang, H.; Zheng, X.; Zhang, Z. FgMon1, a guanine nucleotide exchange factor of FgRab7, is important for vacuole fusion, autophagy and plant infection in Fusarium graminearum. Sci. Rep. 2015, 5, 18101. [Google Scholar] [CrossRef]
- Schrader, M.; Fahimi, H.D. Peroxisomes and oxidative stress. Biochim. Biophys. Acta 2006, 1763, 1755–1766. [Google Scholar] [CrossRef]
- Chen, X.L.; Wang, Z.; Liu, C. Roles of peroxisomes in the rice blast fungus. Biomed. Res. Int. 2016, 2016, 9343417. [Google Scholar] [CrossRef] [PubMed]
- Idnurm, A.; Giles, S.S.; Perfect, J.R.; Heitman, J. Peroxisome function regulates growth on glucose in the basidiomycete fungus Cryptococcus neoformans. Eukaryot. Cell 2007, 6, 60–72. [Google Scholar] [CrossRef] [PubMed]
- Kimoto, D.; Kadooka, C.; Saenrungrot, P.; Okutsu, K.; Yoshizaki, Y.; Takamine, K.; Goto, M.; Tamaki, H.; Futagami, T. Pex16 is involved in peroxisome and Woronin body formation in the white koji fungus, Aspergillus luchuensis mut. kawachii. J. Biosci. Bioeng. 2019, 127, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Finley, E.J.; Gavin, C.E.; Aschner, M.; Gunter, T.E. Manganese neuro toxin and the role of reactive oxygen species. Free Radic. Biol. Med. 2013, 62, 65–75. [Google Scholar] [CrossRef]
- Singh, A.K.; Dobashi, K.; Gupta, M.P.; Asayama, K.; Singh, I.; Orak, J.K. Manganese superoxide dismutase in rat liver peroxisomes: biochemical and immunochemical evidence. Mol. Cell. Biochem. 1999, 197, 7–12. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, J.; Hao, L.; Bijli, K.M.; Chandler, J.D.; Orr, M.; Hu, X.; Jones, D.P.; Go, Y.M. From the Cover: Manganese stimulates mitochondrial H2O2 production in SH-SY5Y human neuroblastoma cells over physiologic as well as toxicologic range. Toxicol. Sci. 2017, 155, 213–223. [Google Scholar] [CrossRef] [PubMed]
- Hasan, M.K.; Cheng, Y.; Kanwar, M.K.; Chu, X.Y.; Ahammed, G.J.; Qi, Z.Y. Responses of plant proteins to heavy metal stress—A review. Front. Plant Sci. 2017, 8, 1492. [Google Scholar] [CrossRef]
- Sung, D.Y.; Kim, T.H.; Komives, E.A.; Mendoza-Cozatl, D.G.; Schroeder, J.I. ARS5 is a component of the 26S proteasome complex, and negatively regulates thiol biosynthesis and arsenic tolerance in Arabidopsis. Plant J. 2009, 59, 802–813. [Google Scholar] [CrossRef] [PubMed]
- Kusmierczyk, A.R.; Kunjappu, M.J.; Funakoshi, M.; Hochstrasser, M. A multimeric assembly factor controls the formation of alternative 20S proteasomes. Nat. Struct. Mol. Biol. 2008, 15, 237–244. [Google Scholar] [CrossRef] [PubMed]
- Padmanabhan, A.; Vuong, S.A.; Hochstrasser, M. Assembly of an evolutionarily conserved alternative proteasome Isoform in human cells. Cell Rep. 2016, 14, 2962–2974. [Google Scholar] [CrossRef] [PubMed]
- Sachadyn, P. Conservation and diversity of MutS proteins. Mutat. Res. 2010, 694, 20–30. [Google Scholar] [CrossRef] [PubMed]
- Saunders, D.G.; Dagdas, Y.F.; Talbot, N.J. Spatial uncoupling of mitosis and cytokinesis during appressorium-mediated plant infection by the rice blast fungus Magnaporthe oryzae. Plant Cell 2010, 22, 2417–2428. [Google Scholar] [CrossRef] [PubMed]
- Gube, M. Fungal Molecular Response to Heavy Metal Stress. In The Mycota-Biochemistry and Molecular Biology, 3rd ed.; Esser, K., Ed.; Springer: Cham, Switzerland, 2016; Volume 4, pp. 47–68. [Google Scholar]
- Soanes, D.M.; Chakrabarti, A.; Paszkiewicz, K.H.; Dawe, A.L.; Talbot, N.J. Genome-wide transcriptional profiling of appressorium development by the rice blast fungus Magnaporthe oryzae. PLoS Pathog. 2012, 8, e1002514. [Google Scholar] [CrossRef] [PubMed]
- Shao, J.F.; Yamaji, N.; Shen, R.F.; Ma, J.F. The key to Mn homeostasis in plants: Regulation of Mn transporters. Trends Plant Sci. 2017, 22, 215–224. [Google Scholar] [CrossRef]
- Cherrad, S.; Girard, V.; Dieryckx, C.; Goncalves, I.R.; Dupuy, J.W.; Bonneu, M.; Rascle, C.; Job, C.; Job, D.; Vacher, S.; et al. Proteomic analysis of proteins secreted by Botrytis cinerea in response to heavy metal toxin. Metallomics 2012, 4, 835–846. [Google Scholar] [CrossRef]
- Wang, L.; Li, H.; Wei, H.; Wu, X.; Ke, L. Identification of cadmium-induced Agaricus blazei genes through suppression subtractive hybridization. Food Chem. Toxicol. 2014, 63, 84–90. [Google Scholar] [CrossRef]
- Irving, H.M.N.H.; Williams, R.J.P. Order of stability of metal complexes. Nature 1948, 162, 746. [Google Scholar] [CrossRef]
- Kim, S.; Hu, J.; Oh, Y.; Park, J.; Choi, J.; Lee, Y.H.; Dean, R.A.; Mitchell, T.K. Combining ChIP-chip and expression profiling to model the MoCRZ1 mediated circuit for Ca/calcineurin signaling in the rice blast fungus. PLoS Pathog. 2010, 6, e1000909. [Google Scholar] [CrossRef] [PubMed]
- Wettmarshausen, J.; Goh, V.; Huang, K.T.; Arduino, D.M.; Tripathi, U.; Leimpek, A.; Cheng, Y.; Pittis, A.A.; Gabaldon, T.; Mokranjac, D.; et al. MICU1 confers protection from MCU-dependent manganese toxin. Cell Rep. 2018, 25, 1425–1435. [Google Scholar] [CrossRef] [PubMed]
- Fitsanakis, V.A.; Zhang, N.; Garcia, S.; Aschner, M. Manganese (Mn) and iron (Fe): Interdependency of transport and regulation. Neurotox. Res. 2010, 18, 124–131. [Google Scholar] [CrossRef] [PubMed]
- Halliwell, B.; Gutteridge, J.M. Biologically relevant metal ion-dependent hydroxyl radical generation an update. FEBS Lett. 1992, 307, 108–112. [Google Scholar] [CrossRef]
- Haas, H. Molecular genetics of fungal siderophore biosynthesis and uptake: The role of siderophores in iron uptake and storage. Appl. Microbiol. Biotechnol. 2003, 62, 316–330. [Google Scholar] [CrossRef]
- Chen, L.H.; Lin, C.H.; Chung, K.R. A nonribosomal peptide synthetase mediates siderophore production and virulence in the citrus fungal pathogen Alternaria alternata. Mol. Plant Pathol. 2013, 14, 497–505. [Google Scholar] [CrossRef] [PubMed]
- Shin, J.Y.; Bui, D.C.; Lee, Y.; Nam, H.; Jung, S.; Fang, M.; Kim, J.C.; Lee, T.; Kim, H.; Choi, G.J.; et al. Functional characterization of cytochrome P450 monooxygenases in the cereal head blight fungus Fusarium graminearum. Environ. Microbiol. 2017, 19, 2053–2067. [Google Scholar] [CrossRef]
- Rai, A.; Singh, R.; Shirke, P.A.; Tripathi, R.D.; Trivedi, P.K.; Chakrabarty, D. Expression of rice CYP450-like gene (Os08g01480) in Arabidopsis modulates regulatory network leading to heavy metal and other abiotic stress tolerance. PLoS ONE 2015, 10, e0138574. [Google Scholar] [CrossRef]
- Korashy, H.M.; El-Kadi, A.O. Regulatory mechanisms modulating the expression of cytochrome P450 1A1 gene by heavy metals. Toxicol. Sci. 2005, 88, 39–51. [Google Scholar] [CrossRef]
- Ceyhun, S.B.; Aksakal, E.; Ekinci, D.; Erdogan, O.; Beydemir, S. Influence of cobalt and zinc exposure on mRNA expression profiles of metallothionein and cytocrome P450 in rainbow trout. Biol. Trace Elem. Res. 2011, 144, 781–789. [Google Scholar] [CrossRef]
- Zheng, Y.X.; Chan, P.; Pan, Z.F.; Shi, N.N.; Wang, Z.X.; Pan, J.; He, F.S. Polymorphism of metabolic genes and susceptibility to occupational chronic manganism. Biomarkers 2002, 7, 337–346. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Lu, L.; Jia, Y.; Li, C. Effectiveness and durability of the rice pi-ta gene in Yunnan province of China. Phytopathology 2014, 104, 762–768. [Google Scholar] [CrossRef] [PubMed]
- Kawano, Y.; Akamatsu, A.; Hayashi, K.; Housen, Y.; Okuda, J.; Yao, A.; Nakashima, A.; Takahashi, H.; Yoshida, H.; Wong, H.L.; et al. Activation of a Rac GTPase by the NLR family disease resistance protein Pit plays a critical role in rice innate immunity. Cell Host Microbe 2010, 7, 362–375. [Google Scholar] [CrossRef]
- Sweigard, J.A.; Carroll, A.M.; Farrall, L.; Chumley, F.G.; Valent, B. Magnaporthe grisea pathogenicity genes obtained through insertional mutagenesis. Mol. Plant Microbe Interact. 1998, 11, 404–412. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−△△Ct Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Wang, Y.; Li, Y.; Wang, H.; Liu, L.; Liu, Y.; Yang, J.; Liu, L.; Li, C. Proteomic analyses of Magnaporthe oryzae development disrupted by salicylic acid. Physiol. Mol. Plant Pathol. 2018, 102, 55–66. [Google Scholar] [CrossRef]

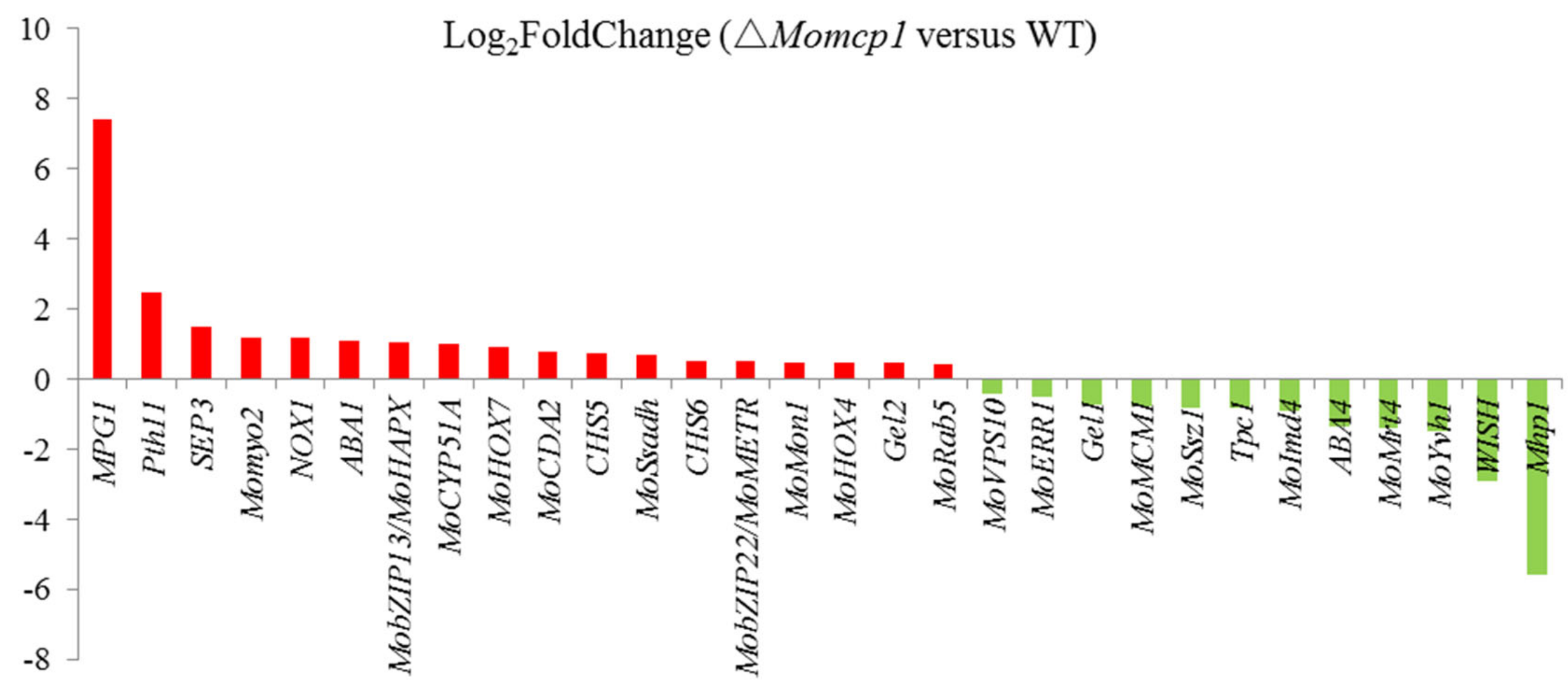
| Gene ID | Description | Log2FoldChange (△Momcp1 versus Wild Type) | Name | Reference |
|---|---|---|---|---|
| MGG_09134 | hypothetical protein | 7.42 | MPG1 | [23] |
| MGG_05871 | hypothetical protein | 2.45 | Pth11 | [24] |
| MGG_01521 | cell division control protein 3 | 1.47 | SEP3 | [25] |
| MGG_03060 | myosin type II heavy chain | 1.16 | Momyo2 | [26] |
| MGG_00750 | cytochrome b-245 heavychain subunit beta | 1.16 | NOX1 | [27] |
| MGG_07626 | cytochrome P450 monooxygenase | 1.07 | ABA1 | [28] |
| MGG_05959 | bZIP transcription factor | 1.05 | MobZIP13/MoHAPX | [29] |
| MGG_04628 | cytochrome P450 51 | 1.01 | MoCYP51A | [30] |
| MGG_12865 | hypothetical protein | 0.91 | MoHOX7 | [31] |
| MGG_08774 | chitin deacetylase | 0.78 | MoCDA2 | [32] |
| MGG_13014 | class V chitin synthase | 0.75 | CHS5 | [33] |
| MGG_01230 | succinate-semialdehyde dehydrogenase | 0.69 | MoSsadh | [34] |
| MGG_13013 | chitin synthase 8 | 0.51 | CHS6 | [33] |
| MGG_14561 | regulatory protein Cys-3 | 0.49 | MobZIP22/MoMETR | [29] |
| MGG_05755 | vacuolar fusion protein | 0.48 | MoMon1 | [35] |
| MGG_06285 | hypothetical protein | 0.46 | MoHOX4 | [31] |
| MGG_06722 | 1%2C3-beta-glucanosyltransferase | 0.46 | Gel2 | [36] |
| MGG_01185 | GTP-binding protein ypt5 | 0.40 | MoRab5 | [37] |
| MGG_00506 | hypothetical protein | −0.42 | MoVPS10 | [38] |
| MGG_16126 | hypothetical protein | −0.51 | MoERR1 | [39] |
| MGG_07331 | 1%2C3-beta-glucanosyltransferase | −0.72 | Gel1 | [36] |
| MGG_02773 | MADS box protein | −0.77 | MoMCM1 | [40] |
| MGG_02842 | hsp70-like protein | −0.81 | MoSsz1 | [41] |
| MGG_01285 | C6 finger domain-containing protein | −0.82 | Tpc1 | [42] |
| MGG_03699 | inosine-5’-monophosphate dehydrogenase | −0.91 | MoImd4 | [43] |
| MGG_07514 | 3-oxoacyl-[acyl-carrier-protein] reductase | −1.34 | ABA4 | [28] |
| MGG_08908 | mRNA turnover protein 4 | −1.39 | MoMrt4 | [44] |
| MGG_09700 | tyrosine-protein phosphatase | −1.49 | MoYvh1 | [44] |
| MGG_09022 | hypothetical protein | −2.91 | WISH | [45] |
| MGG_10105 | hypothetical protein | −5.58 | Mhp1 | [46] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Y.; Wu, Q.; Liu, L.; Li, X.; Lin, A.; Li, C. MoMCP1, a Cytochrome P450 Gene, Is Required for Alleviating Manganese Toxin Revealed by Transcriptomics Analysis in Magnaporthe oryzae. Int. J. Mol. Sci. 2019, 20, 1590. https://doi.org/10.3390/ijms20071590
Wang Y, Wu Q, Liu L, Li X, Lin A, Li C. MoMCP1, a Cytochrome P450 Gene, Is Required for Alleviating Manganese Toxin Revealed by Transcriptomics Analysis in Magnaporthe oryzae. International Journal of Molecular Sciences. 2019; 20(7):1590. https://doi.org/10.3390/ijms20071590
Chicago/Turabian StyleWang, Yi, Qi Wu, Lina Liu, Xiaoling Li, Aijia Lin, and Chengyun Li. 2019. "MoMCP1, a Cytochrome P450 Gene, Is Required for Alleviating Manganese Toxin Revealed by Transcriptomics Analysis in Magnaporthe oryzae" International Journal of Molecular Sciences 20, no. 7: 1590. https://doi.org/10.3390/ijms20071590
APA StyleWang, Y., Wu, Q., Liu, L., Li, X., Lin, A., & Li, C. (2019). MoMCP1, a Cytochrome P450 Gene, Is Required for Alleviating Manganese Toxin Revealed by Transcriptomics Analysis in Magnaporthe oryzae. International Journal of Molecular Sciences, 20(7), 1590. https://doi.org/10.3390/ijms20071590




