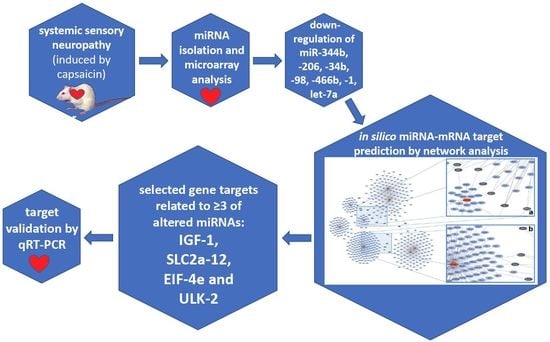
Sensory Neuropathy Affects Cardiac miRNA Expression Network Targeting IGF-1, SLC2a-12, EIF-4e, and ULK-2 mRNAs
Abstract
1. Introduction
2. Results
2.1. Basic Characteristics
2.2. Myocardial Function
2.3. miRNA Microarray Analysis
2.4. Validation of miRNA Microarray Results by qRT-PCR
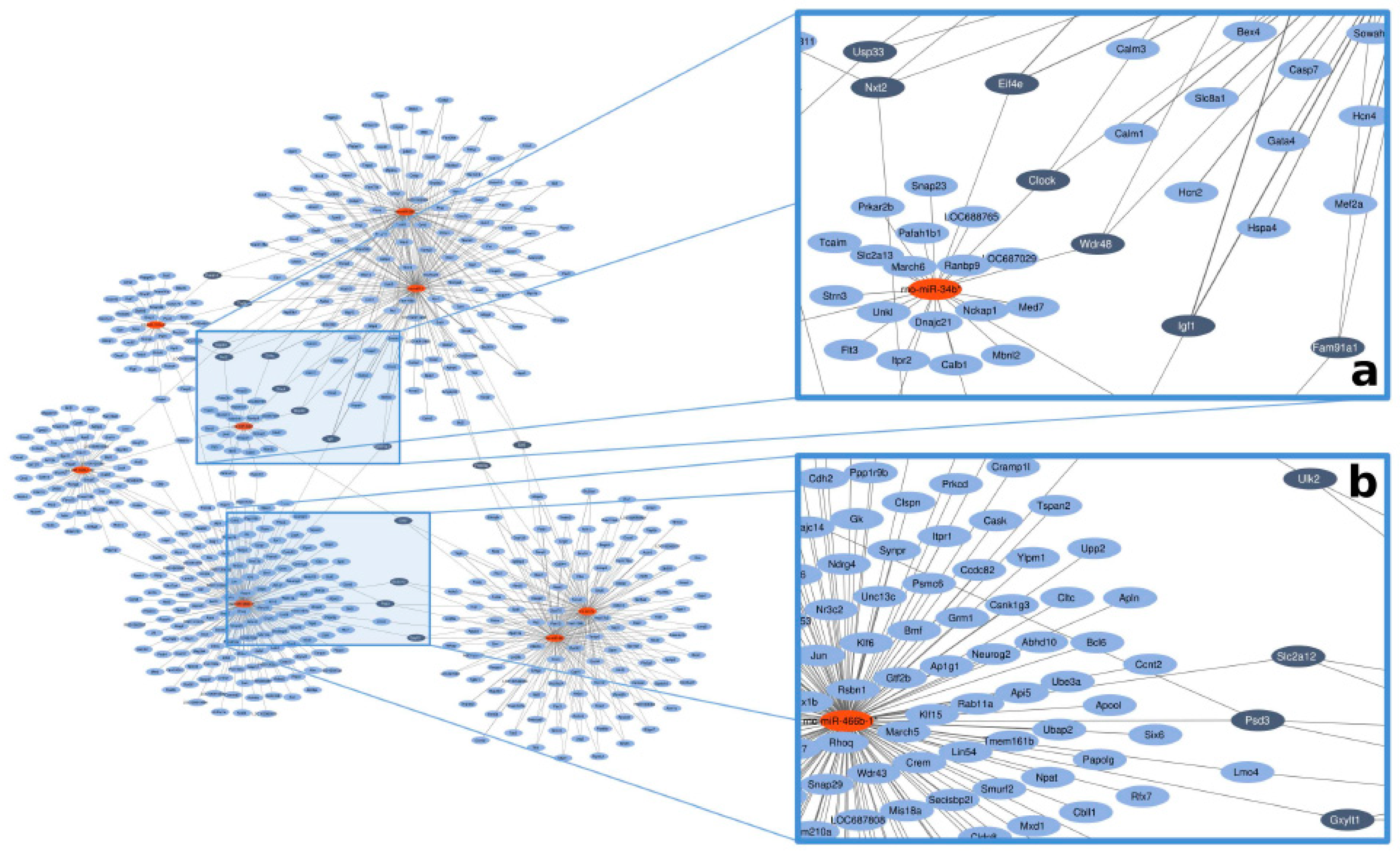
2.5. In Silico Network Analysis
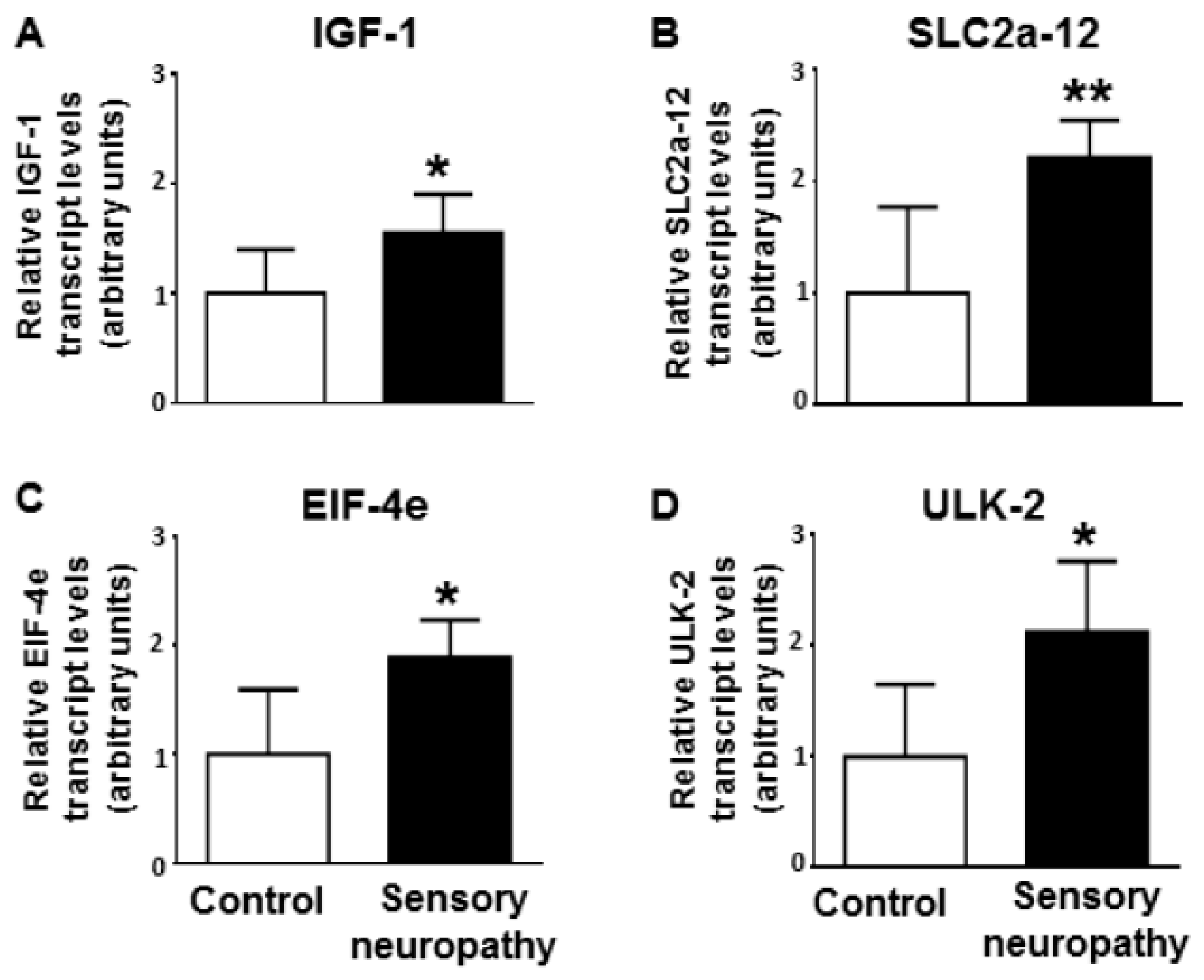
2.6. Validation of Gene Targets at mRNA Level
3. Discussion
3.1. Altered Cardiac Function Due to Sensory Neuropathy
3.2. Altered Cardiac miRNAs Expression Profile due to Sensory Neuropathy
3.3. In Silico miRNA–mRNA Target Prediction
3.4. Altered Cardiac Target mRNAs due to Sensory Neuropathy
3.5. Study Limitations
4. Materials and Methods
4.1. Induction of Selective Sensory Neuropathy by Systemic Capsaicin Treatment
4.2. Transthoracic Echocardiography
4.3. Isolation of miRNAs
4.4. miRNA Microarray
4.5. Validation of miRNA Array Results by qRT-PCR
4.6. Prediction of miRNA Targets
4.7. Selecting miRNA–Target Hubs
4.8. Validation of Predicted miRNA Target Genes at mRNA Levels
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AMI | acute myocardial infarction |
| EIF-4e | eukaryotic translation initiation factor 4e |
| eNOS | endothelial nitrogen monoxide synthase |
| GAPDH | glycerinaldehyde-3-phosphate dehydrogenase |
| GLUT-4 | glucose transporter type 4 |
| HF | heart failure |
| IGF-1 | insulin-like growth factor-1 |
| miRNA | MicroRNA |
| MMLV | Moloney Murine Leukemia Virus Reverse Transcriptase |
| mRNA | messenger RNAs |
| MTI | microRNA-target interaction |
| mTOR | mammalian target of rapamycin |
| NO | nitrogen monoxide |
| qRT-PCR | quantitative real-time PCR |
| SLC2a-12 (GLUT-12) | solute carrier family 2 facilitated glucose transporter member 12 |
| SOD1 | superoxide dismutase-1 |
| TRPV1 | transient receptor potential vanilloid 1 |
| ULK-2 | Unc-51 like autophagy activating kinase 2 |
References
- Marangoni, M.N.; Brady, S.T.; Chowdhury, S.A.; Piano, M.R. The co-occurrence of myocardial dysfunction and peripheral insensate neuropathy in a streptozotocin-induced rat model of diabetes. Cardiovasc. Diabetol. 2014, 13, 11. [Google Scholar] [CrossRef] [PubMed]
- Kostev, K.; Jockwig, A.; Hallwachs, A.; Rathmann, W. Prevalence and risk factors of neuropathy in newly diagnosed type 2 diabetes in primary care practices: A retrospective database analysis in Germany and U.K. Prim. Care Diabetes 2014, 8, 250–255. [Google Scholar] [CrossRef] [PubMed]
- Moura, J.; Borsheim, E.; Carvalho, E. The Role of MicroRNAs in Diabetic Complications-Special Emphasis on Wound Healing. Genes (Basel) 2014, 5, 926–956. [Google Scholar] [CrossRef] [PubMed]
- Tesfaye, S.; Boulton, A.J.; Dyck, P.J.; Freeman, R.; Horowitz, M.; Kempler, P.; Lauria, G.; Malik, R.A.; Spallone, V.; Vinik, A.; et al. Diabetic neuropathies: Update on definitions, diagnostic criteria, estimation of severity, and treatments. Diabetes Care 2010, 33, 2285–2293. [Google Scholar] [CrossRef] [PubMed]
- Mochizuki, Y.; Tanaka, H.; Matsumoto, K.; Sano, H.; Toki, H.; Shimoura, H.; Ooka, J.; Sawa, T.; Motoji, Y.; Ryo, K.; et al. Association of peripheral nerve conduction in diabetic neuropathy with subclinical left ventricular systolic dysfunction. Cardiovasc. Diabetol. 2015, 14, 47. [Google Scholar] [CrossRef] [PubMed]
- Jancso, G.; Kiraly, E.; Jancso-Gabor, A. Pharmacologically induced selective degeneration of chemosensitive primary sensory neurones. Nature 1977, 270, 741–743. [Google Scholar] [CrossRef] [PubMed]
- Jancso, N.; Jancso-Gabor, A.; Szolcsanyi, J. The role of sensory nerve endings in neurogenic inflammation induced in human skin and in the eye and paw of the rat. Br. J. Pharmacol. Chemother. 1968, 33, 32–41. [Google Scholar] [CrossRef] [PubMed]
- Jancso, G.; Oszlacs, O.; Santha, P. The Capsaicin Paradox: Pain Relief by an Algesic Agent. Anti-Inflamm. Anti-Allergy Agent Med. Chem. 2011, 10, 52–65. [Google Scholar] [CrossRef]
- Nagy, I.; Santha, P.; Jancso, G.; Urban, L. The role of the vanilloid (capsaicin) receptor (TRPV1) in physiology and pathology. Eur. J. Pharmacol. 2004, 500, 351–369. [Google Scholar] [CrossRef] [PubMed]
- Szallasi, A.; Cortright, D.N.; Blum, C.A.; Eid, S.R. The vanilloid receptor TRPV1: 10 years from channel cloning to antagonist proof-of-concept. Nat. Rev. Drug Discov. 2007, 6, 357–372. [Google Scholar] [CrossRef] [PubMed]
- Ferdinandy, P.; Csont, T.; Csonka, C.; Torok, M.; Dux, M.; Nemeth, J.; Horvath, L.I.; Dux, L.; Szilvassy, Z.; Jancso, G. Capsaicin-sensitive local sensory innervation is involved in pacing-induced preconditioning in rat hearts: Role of nitric oxide and CGRP? Naunyn Schmiedebergs Arch. Pharmacol. 1997, 356, 356–363. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Song, J.; Chen, H.; Cao, C.; Lee, C. TRPV1 activation is involved in the cardioprotection of remote limb ischemic postconditioning in ischemia-reperfusion injury rats. Biochem. Biophys. Res. Commun. 2015, 463, 1034–1039. [Google Scholar] [CrossRef] [PubMed]
- Franco-Cereceda, A. Calcitonin gene-related peptide and tachykinins in relation to local sensory control of cardiac contractility and coronary vascular tone. Acta Physiol. Scand. Suppl. 1988, 569, 1–63. [Google Scholar] [PubMed]
- Bencsik, P.; Kupai, K.; Giricz, Z.; Gorbe, A.; Huliak, I.; Furst, S.; Dux, L.; Csont, T.; Jancso, G.; Ferdinandy, P. Cardiac capsaicin-sensitive sensory nerves regulate myocardial relaxation via S-nitrosylation of SERCA: Role of peroxynitrite. Br. J. Pharmacol. 2008, 153, 488–496. [Google Scholar] [CrossRef] [PubMed]
- Csont, T.; Csonka, C.; Kovacs, P.; Jancso, G.; Ferdinandy, P. Capsaicin-sensitive sensory neurons regulate myocardial nitric oxide and cGMP signaling. Eur. J. Pharmacol. 2003, 476, 107–113. [Google Scholar] [CrossRef]
- Zvara, A.; Bencsik, P.; Fodor, G.; Csont, T.; Hackler, L., Jr.; Dux, M.; Furst, S.; Jancso, G.; Puskas, L.G.; Ferdinandy, P. Capsaicin-sensitive sensory neurons regulate myocardial function and gene expression pattern of rat hearts: A DNA microarray study. FASEB J. 2006, 20, 160–162. [Google Scholar] [CrossRef] [PubMed]
- Notari, M.; Pulecio, J.; Raya, A. Update on the Pathogenic Implications and Clinical Potential of microRNAs in Cardiac Disease. Biomed. Res. Int. 2015, 2015, 105620. [Google Scholar] [CrossRef] [PubMed]
- Thum, T.; Catalucci, D.; Bauersachs, J. MicroRNAs: Novel regulators in cardiac development and disease. Cardiovasc. Res. 2008, 79, 562–570. [Google Scholar] [CrossRef] [PubMed]
- Wronska, A.; Kurkowska-Jastrzebska, I.; Santulli, G. Application of microRNAs in diagnosis and treatment of cardiovascular disease. Acta Physiol. (Oxf.) 2015, 213, 60–83. [Google Scholar] [CrossRef] [PubMed]
- Leon, L.E.; Rani, S.; Fernandez, M.; Larico, M.; Calligaris, S.D. Subclinical Detection of Diabetic Cardiomyopathy with MicroRNAs: Challenges and Perspectives. J. Diabetes Res. 2016, 2016, 6143129. [Google Scholar] [CrossRef] [PubMed]
- Figueira, M.F.; Monnerat-Cahli, G.; Medei, E.; Carvalho, A.B.; Morales, M.M.; Lamas, M.E.; da Fonseca, R.N.; Souza-Menezes, J. MicroRNAs: Potential therapeutic targets in diabetic complications of the cardiovascular and renal systems. Acta Physiol. (Oxf.) 2014, 211, 491–500. [Google Scholar] [CrossRef] [PubMed]
- Perrino, C.; Barabasi, A.L.; Condorelli, G.; Davidson, S.M.; De Windt, L.; Dimmeler, S.; Engel, F.B.; Hausenloy, D.J.; Hill, J.A.; Van Laake, L.W.; et al. Epigenomic and transcriptomic approaches in the post-genomic era: Path to novel targets for diagnosis and therapy of the ischaemic heart? Position Paper of the European Society of Cardiology Working Group on Cellular Biology of the Heart. Cardiovasc. Res. 2017, 113, 725–736. [Google Scholar] [CrossRef] [PubMed]
- Agg, B.; Baranyai, T.; Makkos, A.; Veto, B.; Farago, N.; Zvara, A.; Giricz, Z.; Veres, D.V.; Csermely, P.; Aranyi, T.; et al. MicroRNA interactome analysis predicts post-transcriptional regulation of ADRB2 and PPP3R1 in the hypercholesterolemic myocardium. Sci. Rep. 2018, 8, 10134. [Google Scholar] [CrossRef] [PubMed]
- Schulz, R.; Agg, B.; Ferdinandy, P. Survival pathways in cardiac conditioning: Individual data vs. meta-analyses. What do we learn? Basic Res. Cardiol. 2017, 113, 4. [Google Scholar] [CrossRef] [PubMed]
- Varga, Z.V.; Kupai, K.; Szucs, G.; Gaspar, R.; Paloczi, J.; Farago, N.; Zvara, A.; Puskas, L.G.; Razga, Z.; Tiszlavicz, L.; et al. MicroRNA-25-dependent up-regulation of NADPH oxidase 4 (NOX4) mediates hypercholesterolemia-induced oxidative/nitrative stress and subsequent dysfunction in the heart. J. Mol. Cell. Cardiol. 2013, 62, 111–121. [Google Scholar] [CrossRef] [PubMed]
- Varga, Z.V.; Zvara, A.; Farago, N.; Kocsis, G.F.; Pipicz, M.; Gaspar, R.; Bencsik, P.; Gorbe, A.; Csonka, C.; Puskas, L.G.; et al. MicroRNAs associated with ischemia-reperfusion injury and cardioprotection by ischemic pre- and postconditioning: protectomiRs. Am. J. Physiol. Heart Circ. Physiol. 2014, 307, H216–H227. [Google Scholar] [CrossRef] [PubMed]
- Yong, F.L.; Law, C.W.; Wang, C.W. Potentiality of a triple microRNA classifier: miR-193a-3p, miR-23a and miR-338-5p for early detection of colorectal cancer. BMC Cancer 2013, 13, 280. [Google Scholar] [CrossRef] [PubMed]
- Ntoumou, E.; Tzetis, M.; Braoudaki, M.; Lambrou, G.; Poulou, M.; Malizos, K.; Stefanou, N.; Anastasopoulou, L.; Tsezou, A. Serum microRNA array analysis identifies miR-140-3p, miR-33b-3p and miR-671-3p as potential osteoarthritis biomarkers involved in metabolic processes. Clin. Epigenet. 2017, 9, 127. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Ho, D.; Vatner, D.E.; Vatner, S.F. Echocardiography in Mice. Curr. Protoc. Mouse Biol. 2011, 1, 71–83. [Google Scholar] [CrossRef] [PubMed]
- Cinca, J.; Rodriguez-Sinovas, A. Cardiovascular reflex responses induced by epicardial chemoreceptor stimulation. Cardiovasc. Res. 2000, 45, 163–171. [Google Scholar] [CrossRef]
- Garry, M.G. Abnormalities of the exercise pressor reflex in heart failure. Exerc. Sport Sci. Rev. 2011, 39, 167–176. [Google Scholar] [CrossRef] [PubMed]
- Robbins, N.; Koch, S.E.; Rubinstein, J. Targeting TRPV1 and TRPV2 for potential therapeutic interventions in cardiovascular disease. Transl. Res. 2013, 161, 469–476. [Google Scholar] [CrossRef] [PubMed]
- Dalsgaard, C.J.; Franco-Cereceda, A.; Saria, A.; Lundberg, J.M.; Theodorsson-Norheim, E.; Hokfelt, T. Distribution and origin of substance P- and neuropeptide Y-immunoreactive nerves in the guinea-pig heart. Cell Tissue Res. 1986, 243, 477–485. [Google Scholar] [CrossRef] [PubMed]
- Franco-Cereceda, A.; Lundberg, J.M. Calcitonin gene-related peptide (CGRP) and capsaicin-induced stimulation of heart contractile rate and force. Naunyn Schmiedebergs Arch. Pharmacol. 1985, 331, 146–151. [Google Scholar] [CrossRef] [PubMed]
- Franco-Cereceda, A.; Liska, J. Potential of calcitonin gene-related peptide in coronary heart disease. Pharmacology 2000, 60, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Jeng, S.F.; Rau, C.S.; Liliang, P.C.; Wu, C.J.; Lu, T.H.; Chen, Y.C.; Lin, C.J.; Hsieh, C.H. Profiling muscle-specific microRNA expression after peripheral denervation and reinnervation in a rat model. J. Neurotrauma 2009, 26, 2345–2353. [Google Scholar] [CrossRef] [PubMed]
- Rau, C.S.; Jeng, J.C.; Jeng, S.F.; Lu, T.H.; Chen, Y.C.; Liliang, P.C.; Wu, C.J.; Lin, C.J.; Hsieh, C.H. Entrapment neuropathy results in different microRNA expression patterns from denervation injury in rats. BMC Musculoskelet. Disord. 2010, 11, 181. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Nath, S.; Prasad, H.K.; Sharma, G.D.; Li, Y. MicroRNAs: A new ray of hope for diabetes mellitus. Protein Cell 2012, 3, 726–738. [Google Scholar] [CrossRef] [PubMed]
- Mitchelson, K.R.; Qin, W.Y. Roles of the canonical myomiRs miR-1, -133 and -206 in cell development and disease. World J. Biol. Chem. 2015, 6, 162–208. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zheng, S.; Geng, Y.; Xue, J.; Wang, Z.; Xie, X.; Wang, J.; Zhang, S.; Hou, Y. MicroRNA profiling of atrial fibrillation in canines: miR-206 modulates intrinsic cardiac autonomic nerve remodeling by regulating SOD1. PLoS ONE 2015, 10, e0122674. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Yuan, Y.; Li, J.; Ren, H.; Cai, Q.; Chen, X.; Liang, H.; Shan, H.; Fu, Z.D.; Gao, X.; et al. MicroRNA-1 aggravates cardiac oxidative stress by post-transcriptional modification of the antioxidant network. Cell Stress Chaperones 2015, 20, 411–420. [Google Scholar] [CrossRef] [PubMed]
- Kumarswamy, R.; Lyon, A.R.; Volkmann, I.; Mills, A.M.; Bretthauer, J.; Pahuja, A.; Geers-Knorr, C.; Kraft, T.; Hajjar, R.J.; Macleod, K.T.; et al. SERCA2a gene therapy restores microRNA-1 expression in heart failure via an Akt/FoxO3A-dependent pathway. Eur. Heart J. 2012, 33, 1067–1075. [Google Scholar] [CrossRef] [PubMed]
- Mishra, P.K.; Metreveli, N.; Tyagi, S.C. MMP-9 gene ablation and TIMP-4 mitigate PAR-1-mediated cardiomyocyte dysfunction: A plausible role of dicer and miRNA. Cell Biochem. Biophys. 2010, 57, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Feng, B.; Cao, Y.; Chen, S.; Ruiz, M.; Chakrabarti, S. Reprint of: miRNA-1 regulates endothelin-1 in diabetes. Life Sci. 2014, 118, 275–280. [Google Scholar] [CrossRef] [PubMed]
- Yildirim, S.S.; Akman, D.; Catalucci, D.; Turan, B. Relationship between downregulation of miRNAs and increase of oxidative stress in the development of diabetic cardiac dysfunction: Junctin as a target protein of miR-1. Cell Biochem. Biophys. 2013, 67, 1397–1408. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Yang, G.M.; Zhu, Y.; Wu, Y.; Chen, X.Y.; Lan, D.; Tian, K.L.; Liu, L.M. Diabetes and hyperlipidemia induce dysfunction of VSMCs: Contribution of the metabolic inflammation/miRNA pathway. Am. J. Physiol. Endocrinol. Metab. 2015, 308, E257–E269. [Google Scholar] [CrossRef] [PubMed]
- Shan, Z.X.; Lin, Q.X.; Deng, C.Y.; Zhu, J.N.; Mai, L.P.; Liu, J.L.; Fu, Y.H.; Liu, X.Y.; Li, Y.X.; Zhang, Y.Y.; et al. miR-1/miR-206 regulate Hsp60 expression contributing to glucose-mediated apoptosis in cardiomyocytes. FEBS Lett. 2010, 584, 3592–3600. [Google Scholar] [CrossRef] [PubMed]
- Matkovich, S.J.; Van Booven, D.J.; Youker, K.A.; Torre-Amione, G.; Diwan, A.; Eschenbacher, W.H.; Dorn, L.E.; Watson, M.A.; Margulies, K.B.; Dorn, G.W., 2nd. Reciprocal regulation of myocardial microRNAs and messenger RNA in human cardiomyopathy and reversal of the microRNA signature by biomechanical support. Circulation 2009, 119, 1263–1271. [Google Scholar] [CrossRef] [PubMed]
- Ai, J.; Zhang, R.; Li, Y.; Pu, J.; Lu, Y.; Jiao, J.; Li, K.; Yu, B.; Li, Z.; Wang, R.; et al. Circulating microRNA-1 as a potential novel biomarker for acute myocardial infarction. Biochem. Biophys. Res. Commun. 2010, 391, 73–77. [Google Scholar] [CrossRef] [PubMed]
- Baars, T.; Skyschally, A.; Klein-Hitpass, L.; Cario, E.; Erbel, R.; Heusch, G.; Kleinbongard, P. microRNA expression and its potential role in cardioprotection by ischemic postconditioning in pigs. Pflugers Arch. 2014, 466, 1953–1961. [Google Scholar] [CrossRef] [PubMed]
- Yin, C.; Salloum, F.N.; Kukreja, R.C. A novel role of microRNA in late preconditioning: Upregulation of endothelial nitric oxide synthase and heat shock protein 70. Circ. Res. 2009, 104, 572–575. [Google Scholar] [CrossRef] [PubMed]
- Su, J.L.; Chen, P.S.; Johansson, G.; Kuo, M.L. Function and regulation of let-7 family microRNAs. Microrna 2012, 1, 34–39. [Google Scholar] [CrossRef] [PubMed]
- Bao, M.H.; Zhang, Y.W.; Lou, X.Y.; Cheng, Y.; Zhou, H.H. Protective effects of let-7a and let-7b on oxidized low-density lipoprotein induced endothelial cell injuries. PLoS ONE 2014, 9, e106540. [Google Scholar] [CrossRef] [PubMed]
- Chavali, V.; Tyagi, S.C.; Mishra, P.K. Differential expression of dicer, miRNAs, and inflammatory markers in diabetic Ins2+/- Akita hearts. Cell Biochem. Biophys. 2014, 68, 25–35. [Google Scholar] [CrossRef] [PubMed]
- Santovito, D.; De Nardis, V.; Marcantonio, P.; Mandolini, C.; Paganelli, C.; Vitale, E.; Buttitta, F.; Bucci, M.; Mezzetti, A.; Consoli, A.; et al. Plasma exosome microRNA profiling unravels a new potential modulator of adiponectin pathway in diabetes: Effect of glycemic control. J. Clin. Endocrinol. Metab. 2014, 99, E1681–E1685. [Google Scholar] [CrossRef] [PubMed]
- Joshi, S.; Wei, J.; Bishopric, N.H. A cardiac myocyte-restricted Lin28/let-7 regulatory axis promotes hypoxia-mediated apoptosis by inducing the AKT signaling suppressor PIK3IP1. Biochim. Biophys. Acta 2016, 1862, 240–251. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wang, B.; Cui, H.; Du, Y.; Song, Y.; Yang, L.; Zhang, Q.; Sun, F.; Luo, D.; Xu, C.; et al. Let-7e replacement yields potent anti-arrhythmic efficacy via targeting beta 1-adrenergic receptor in rat heart. J. Cell. Mol. Med. 2014, 18, 1334–1343. [Google Scholar] [CrossRef] [PubMed]
- Li, X.X.; Liu, Y.M.; Li, Y.J.; Xie, N.; Yan, Y.F.; Chi, Y.L.; Zhou, L.; Xie, S.Y.; Wang, P.Y. High glucose concentration induces endothelial cell proliferation by regulating cyclin-D2-related miR-98. J. Cell. Mol. Med. 2016, 20, 1159–1169. [Google Scholar] [CrossRef] [PubMed]
- Xie, S.; Xie, N.; Li, Y.; Wang, P.; Zhang, C.; Li, Q.; Liu, X.; Deng, J.; Zhang, C.; Lv, C. Upregulation of TRB2 induced by miR-98 in the early lesions of large artery of type-2 diabetic rat. Mol. Cell. Biochem. 2012, 361, 305–314. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Wang, H.; Kornblau, S.M.; Graber, D.A.; Zhang, N.; Matthews, J.A.; Wang, M.; Weber, D.M.; Thomas, S.K.; Shah, J.J.; et al. Evidence of a role for the novel zinc-finger transcription factor ZKSCAN3 in modulating Cyclin D2 expression in multiple myeloma. Oncogene 2011, 30, 1329–1340. [Google Scholar] [CrossRef] [PubMed]
- Greco, S.; Fasanaro, P.; Castelvecchio, S.; D’Alessandra, Y.; Arcelli, D.; Di Donato, M.; Malavazos, A.; Capogrossi, M.C.; Menicanti, L.; Martelli, F. MicroRNA dysregulation in diabetic ischemic heart failure patients. Diabetes 2012, 61, 1633–1641. [Google Scholar] [CrossRef] [PubMed]
- Thum, T.; Galuppo, P.; Wolf, C.; Fiedler, J.; Kneitz, S.; van Laake, L.W.; Doevendans, P.A.; Mummery, C.L.; Borlak, J.; Haverich, A.; et al. MicroRNAs in the human heart: A clue to fetal gene reprogramming in heart failure. Circulation 2007, 116, 258–267. [Google Scholar] [CrossRef] [PubMed]
- Bernardo, B.C.; Gao, X.M.; Winbanks, C.E.; Boey, E.J.; Tham, Y.K.; Kiriazis, H.; Gregorevic, P.; Obad, S.; Kauppinen, S.; Du, X.J.; et al. Therapeutic inhibition of the miR-34 family attenuates pathological cardiac remodeling and improves heart function. Proc. Natl. Acad. Sci. USA 2012, 109, 17615–17620. [Google Scholar] [CrossRef] [PubMed]
- Shyamasundar, S.; Jadhav, S.P.; Bay, B.H.; Tay, S.S.; Kumar, S.D.; Rangasamy, D.; Dheen, S.T. Analysis of epigenetic factors in mouse embryonic neural stem cells exposed to hyperglycemia. PLoS ONE 2013, 8, e65945. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; He, H.; Zeng, T.; Huang, Z.; Fan, T.; Wu, Q. Neural-specific expression of miR-344-3p during mouse embryonic development. J. Mol. Histol. 2014, 45, 363–372. [Google Scholar] [CrossRef] [PubMed]
- Ramasamy, S.; Velmurugan, G.; Shanmugha Rajan, K.; Ramprasath, T.; Kalpana, K. MiRNAs with apoptosis regulating potential are differentially expressed in chronic exercise-induced physiologically hypertrophied hearts. PLoS ONE 2015, 10, e0121401. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, L.B.; Wang, C.; Sorensen, K.; Bang-Berthelsen, C.H.; Hansen, L.; Andersen, M.L.; Hougaard, P.; Juul, A.; Zhang, C.Y.; Pociot, F.; et al. Circulating levels of microRNA from children with newly diagnosed type 1 diabetes and healthy controls: Evidence that miR-25 associates to residual beta-cell function and glycaemic control during disease progression. Exp. Diabetes Res. 2012, 2012, 896362. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.; Li, C.; Qi, W.; Zhang, Y.; Zhang, F.; Wu, J.X.; Hu, Y.N.; Wu, D.M.; Liu, Y.; Yan, T.T.; et al. Downregulation of miR-181a upregulates sirtuin-1 (SIRT1) and improves hepatic insulin sensitivity. Diabetologia 2012, 55, 2032–2043. [Google Scholar] [CrossRef] [PubMed]
- Vlachos, I.S.; Paraskevopoulou, M.D.; Karagkouni, D.; Georgakilas, G.; Vergoulis, T.; Kanellos, I.; Anastasopoulos, I.L.; Maniou, S.; Karathanou, K.; Kalfakakou, D.; et al. DIANA-TarBase v7.0: Indexing more than half a million experimentally supported miRNA:mRNA interactions. Nucleic Acids Res. 2015, 43, D153–D159. [Google Scholar] [CrossRef] [PubMed]
- Hsu, S.D.; Tseng, Y.T.; Shrestha, S.; Lin, Y.L.; Khaleel, A.; Chou, C.H.; Chu, C.F.; Huang, H.Y.; Lin, C.M.; Ho, S.Y.; et al. miRTarBase update 2014: An information resource for experimentally validated miRNA-target interactions. Nucleic Acids Res. 2014, 42, D78–D85. [Google Scholar] [CrossRef] [PubMed]
- Betel, D.; Koppal, A.; Agius, P.; Sander, C.; Leslie, C. Comprehensive modeling of microRNA targets predicts functional non-conserved and non-canonical sites. Genome Biol. 2010, 11, R90. [Google Scholar] [CrossRef] [PubMed]
- Enright, A.J.; John, B.; Gaul, U.; Tuschl, T.; Sander, C.; Marks, D.S. MicroRNA targets in Drosophila. Genome Biol. 2003, 5, R1. [Google Scholar] [CrossRef] [PubMed]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Kertesz, M.; Iovino, N.; Unnerstall, U.; Gaul, U.; Segal, E. The role of site accessibility in microRNA target recognition. Nat. Genet. 2007, 39, 1278–1284. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, U.; Lai, X.; Winter, F.; Wolkenhauer, O.; Vera, J.; Gupta, S.K. Cooperative gene regulation by microRNA pairs and their identification using a computational workflow. Nucleic Acids Res. 2014, 42, 7539–7552. [Google Scholar] [CrossRef] [PubMed]
- HafezQorani, S.; Lafzi, A.; de Bruin, R.G.; van Zonneveld, A.J.; van der Veer, E.P.; Son, Y.A.; Kazan, H. Modeling the combined effect of RNA-binding proteins and microRNAs in post-transcriptional regulation. Nucleic Acids Res. 2016, 44, e83. [Google Scholar] [CrossRef] [PubMed]
- Wong, N.; Wang, X. miRDB: An online resource for microRNA target prediction and functional annotations. Nucleic Acids Res. 2015, 43, D146–D152. [Google Scholar] [CrossRef] [PubMed]
- Chou, C.H.; Shrestha, S.; Yang, C.D.; Chang, N.W.; Lin, Y.L.; Liao, K.W.; Huang, W.C.; Sun, T.H.; Tu, S.J.; Lee, W.H.; et al. miRTarBase update 2018: A resource for experimentally validated microRNA-target interactions. Nucleic Acids Res. 2018, 46, D296–D302. [Google Scholar] [CrossRef] [PubMed]
- Sathianathan, V.; Avelino, A.; Charrua, A.; Santha, P.; Matesz, K.; Cruz, F.; Nagy, I. Insulin induces cobalt uptake in a subpopulation of rat cultured primary sensory neurons. Eur. J. Neurosci. 2003, 18, 2477–2486. [Google Scholar] [CrossRef] [PubMed]
- Van Buren, J.J.; Bhat, S.; Rotello, R.; Pauza, M.E.; Premkumar, L.S. Sensitization and translocation of TRPV1 by insulin and IGF-I. Mol. Pain 2005, 1, 17. [Google Scholar] [CrossRef] [PubMed]
- Harada, N.; Okajima, K. Effect of capsaicin on plasma and tissue levels of insulin-like growth factor-I in spontaneously hypertensive rats. Growth Horm. IGF Res. 2008, 18, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Wang, C.; Chen, J.; Li, X.; Fang, Z.; Ma, H. The role of IGF-1 gene expression abnormality in pathogenesis of diabetic peripheral neuropathy. Chin. Med. Sci. J. 2002, 17, 204–209. [Google Scholar]
- Ishii, D.N. Implication of insulin-like growth factors in the pathogenesis of diabetic neuropathy. Brain Res. Brain Res. Rev. 1995, 20, 47–67. [Google Scholar] [CrossRef]
- Ren, J.; Samson, W.K.; Sowers, J.R. Insulin-like growth factor I as a cardiac hormone: Physiological and pathophysiological implications in heart disease. J. Mol. Cell. Cardiol. 1999, 31, 2049–2061. [Google Scholar] [CrossRef] [PubMed]
- Reiss, K.; Kajstura, J.; Zhang, X.; Li, P.; Szoke, E.; Olivetti, G.; Anversa, P. Acute myocardial infarction leads to upregulation of the IGF-1 autocrine system, DNA replication, and nuclear mitotic division in the remaining viable cardiac myocytes. Exp. Cell Res. 1994, 213, 463–472. [Google Scholar] [CrossRef] [PubMed]
- Suleiman, M.S.; Singh, R.J.; Stewart, C.E. Apoptosis and the cardiac action of insulin-like growth factor I. Pharmacol. Ther. 2007, 114, 278–294. [Google Scholar] [CrossRef] [PubMed]
- Hua, Y.; Zhang, Y.; Ren, J. IGF-1 deficiency resists cardiac hypertrophy and myocardial contractile dysfunction: Role of microRNA-1 and microRNA-133a. J. Cell. Mol. Med. 2012, 16, 83–95. [Google Scholar] [CrossRef] [PubMed]
- Elia, L.; Contu, R.; Quintavalle, M.; Varrone, F.; Chimenti, C.; Russo, M.A.; Cimino, V.; De Marinis, L.; Frustaci, A.; Catalucci, D.; et al. Reciprocal regulation of microRNA-1 and insulin-like growth factor-1 signal transduction cascade in cardiac and skeletal muscle in physiological and pathological conditions. Circulation 2009, 120, 2377–2385. [Google Scholar] [CrossRef] [PubMed]
- Purcell, S.H.; Aerni-Flessner, L.B.; Willcockson, A.R.; Diggs-Andrews, K.A.; Fisher, S.J.; Moley, K.H. Improved insulin sensitivity by GLUT12 overexpression in mice. Diabetes 2011, 60, 1478–1482. [Google Scholar] [CrossRef] [PubMed]
- Waller, A.P.; George, M.; Kalyanasundaram, A.; Kang, C.; Periasamy, M.; Hu, K.; Lacombe, V.A. GLUT12 functions as a basal and insulin-independent glucose transporter in the heart. Biochim. Biophys. Acta 2013, 1832, 121–127. [Google Scholar] [CrossRef] [PubMed]
- Fingar, D.C.; Blenis, J. Target of rapamycin (TOR): An integrator of nutrient and growth factor signals and coordinator of cell growth and cell cycle progression. Oncogene 2004, 23, 3151–3171. [Google Scholar] [CrossRef] [PubMed]
- Martins, T.; Vitorino, R.; Moreira-Goncalves, D.; Amado, F.; Duarte, J.A.; Ferreira, R. Recent insights on the molecular mechanisms and therapeutic approaches for cardiac cachexia. Clin. Biochem. 2014, 47, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Choi, C.H.; Jung, Y.K.; Oh, S.H. Autophagy induction by capsaicin in malignant human breast cells is modulated by p38 and extracellular signal-regulated mitogen-activated protein kinases and retards cell death by suppressing endoplasmic reticulum stress-mediated apoptosis. Mol. Pharmacol. 2010, 78, 114–125. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.Y.; Hopkinson, N.S.; Kemp, P.R. Myostatin induces autophagy in skeletal muscle in vitro. Biochem. Biophys. Res. Commun. 2011, 415, 632–636. [Google Scholar] [CrossRef] [PubMed]
- Varga, Z.V.; Giricz, Z.; Liaudet, L.; Hasko, G.; Ferdinandy, P.; Pacher, P. Interplay of oxidative, nitrosative/nitrative stress, inflammation, cell death and autophagy in diabetic cardiomyopathy. Biochim. Biophys. Acta 2015, 1852, 232–242. [Google Scholar] [CrossRef] [PubMed]
- Ferdinandy, P.; Hausenloy, D.J.; Heusch, G.; Baxter, G.F.; Schulz, R. Interaction of risk factors, comorbidities, and comedications with ischemia/reperfusion injury and cardioprotection by preconditioning, postconditioning, and remote conditioning. Pharmacol. Rev. 2014, 66, 1142–1174. [Google Scholar] [CrossRef] [PubMed]
- Lazar, B.A.; Jancso, G.; Oszlacs, O.; Nagy, I.; Santha, P. The Insulin Receptor Is Colocalized With the TRPV1 Nociceptive Ion Channel and Neuropeptides in Pancreatic Spinal and Vagal Primary Sensory Neurons. Pancreas 2018, 47, 110–115. [Google Scholar] [CrossRef] [PubMed]
- Fabian, M.R.; Sonenberg, N.; Filipowicz, W. Regulation of mRNA translation and stability by microRNAs. Annu. Rev. Biochem. 2010, 79, 351–379. [Google Scholar] [CrossRef] [PubMed]
- Mozsik, G.; Vincze, A.; Szolcsanyi, J. Four response stages of capsaicin-sensitive primary afferent neurons to capsaicin and its analog: Gastric acid secretion, gastric mucosal damage and protection. J. Gastroenterol. Hepatol. 2001, 16, 1093–1097. [Google Scholar] [CrossRef] [PubMed]
- Jancso, G.; Kiraly, E.; Joo, F.; Such, G.; Nagy, A. Selective degeneration by capsaicin of a subpopulation of primary sensory neurons in the adult rat. Neurosci. Lett. 1985, 59, 209–214. [Google Scholar] [CrossRef]
- Aghazadeh Tabrizi, M.; Baraldi, P.G.; Baraldi, S.; Gessi, S.; Merighi, S.; Borea, P.A. Medicinal Chemistry, Pharmacology, and Clinical Implications of TRPV1 Receptor Antagonists. Med. Res. Rev. 2017, 37, 936–983. [Google Scholar] [CrossRef] [PubMed]
- Gamse, R. Capsaicin and nociception in the rat and mouse. Possible role of substance P. Naunyn Schmiedebergs Arch. Pharmacol. 1982, 320, 205–216. [Google Scholar] [CrossRef] [PubMed]
- Borbely, E.; Botz, B.; Bolcskei, K.; Kenyer, T.; Kereskai, L.; Kiss, T.; Szolcsanyi, J.; Pinter, E.; Csepregi, J.Z.; Mocsai, A.; et al. Capsaicin-sensitive sensory nerves exert complex regulatory functions in the serum-transfer mouse model of autoimmune arthritis. Brain Behav. Immun. 2015, 45, 50–59. [Google Scholar] [CrossRef] [PubMed]
- Scheich, B.; Vincze, P.; Szoke, E.; Borbely, E.; Hunyady, A.; Szolcsanyi, J.; Denes, A.; Kornyei, Z.; Gaszner, B.; Helyes, Z. Chronic stress-induced mechanical hyperalgesia is controlled by capsaicin-sensitive neurones in the mouse. Eur. J. Pain 2017, 21, 1417–1431. [Google Scholar] [CrossRef] [PubMed]
- Csonka, C.; Szilvassy, Z.; Fulop, F.; Pali, T.; Blasig, I.E.; Tosaki, A.; Schulz, R.; Ferdinandy, P. Classic preconditioning decreases the harmful accumulation of nitric oxide during ischemia and reperfusion in rat hearts. Circulation 1999, 100, 2260–2266. [Google Scholar] [CrossRef] [PubMed]
- Farago, N.; Zvara, A.; Varga, Z.; Ferdinandy, P.; Puskas, L.G. Purification of high-quality micro RNA from the heart tissue. Acta Biol. Hung. 2011, 62, 413–425. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.E.; Silver, J.; Oshlack, A.; Holmes, M.; Diyagama, D.; Holloway, A.; Smyth, G.K. A comparison of background correction methods for two-colour microarrays. Bioinformatics 2007, 23, 2700–2707. [Google Scholar] [CrossRef] [PubMed]
- Antal, O.; Hackler, L., Jr.; Shen, J.; Man, I.; Hideghety, K.; Kitajka, K.; Puskas, L.G. Combination of unsaturated fatty acids and ionizing radiation on human glioma cells: Cellular, biochemical and gene expression analysis. Lipids Health Dis. 2014, 13, 142. [Google Scholar] [CrossRef] [PubMed]
- Lai, X.; Schmitz, U.; Gupta, S.K.; Bhattacharya, A.; Kunz, M.; Wolkenhauer, O.; Vera, J. Computational analysis of target hub gene repression regulated by multiple and cooperative miRNAs. Nucleic Acids Res. 2012, 40, 8818–8834. [Google Scholar] [CrossRef] [PubMed]
- Kovacs, I.A.; Mizsei, R.; Csermely, P. A unified data representation theory for network visualization, ordering and coarse-graining. Sci. Rep. 2015, 5, 13786. [Google Scholar] [CrossRef] [PubMed]
| BW 0 | BW 10 | ΔBW (10–0) | HW | HW/BW | |
|---|---|---|---|---|---|
| (g) | (g) | (g) | (mg) | (‰) | |
| Control | 339 ± 25 | 380 ± 40 | 41 ± 20 | 1167 ± 116 | 3.1 ± 0.2 |
| Sensory Neuropathy | 328 ± 16 | 335 ± 17 * | 7 ± 5 ** | 972 ± 98 * | 2.9 ± 0.2 |
| Parameter | Control | Sensory Neuropathy | p-Value |
|---|---|---|---|
| Heart rate | 412 ± 32 | 441 ± 35 | 0.30 |
| EDD (cm) | 0.69 ± 0.05 | 0.62 ± 0.03 | 0.03 * |
| ESD (cm) | 0.40 ± 0.10 | 0.36 ± 0.05 | 0.46 |
| IVS (cm) | 0.15 ± 0.01 | 0.13 ± 0.01 | 0.04 * |
| PW (cm) | 0.16 ± 0.01 | 0.15 ± 0.01 | 0.12 |
| EDV (ml) | 0.70 ± 0.18 | 0.54 ± 0.07 | 0.14 |
| ESV (ml) | 0.18 ± 0.10 | 0.12 ± 0.05 | 0.30 |
| SV (EDV–ESV) | 0.54 ± 0.07 | 0.42 ± 0.07 | 0.03 * |
| LVEF (%) | 75.83 ± 9.89 | 77.57 ± 8.46 | 0.79 |
| LVFS (%) | 41.14 ± 9.77 | 41.89 ± 7.35 | 0.90 |
| MVVTI (cm) | 4.14 ± 0.36 | 3.23 ± 0.75 | 0.04 * |
| E (cm/sec) | 80.67 ± 14.40 | 65.2 ± 11.39 | 0.11 |
| A (cm/sec) | 59.50 ± 27.87 | 53.2 ± 31.22 | 0.76 |
| Ea (cm/sec) | 6.33 ± 1.25 | 6.20 ± 3.19 | 0.93 |
| Aa (cm/sec) | 5.50 ± 1.71 | 6.40 ± 1.36 | 0.41 |
| Sa (cm/sec) | 5.00 ± 1.15 | 5.60 ± 1.85 | 0.57 |
| E/Ea | 13.07 ± 3.02 | 13.03 ± 5.44 | 0.99 |
| Ea/Aa | 1.26 ± 0.44 | 0.98 ± 0.48 | 0.23 |
| miRNA | Average log2 Change Sensory Neuropathy/Control | Log2 SD | Regulation |
|---|---|---|---|
| rno-miR-344b-1-3p | −1.95 * | 0.47 | down |
| rno-miR-466b-1-3p | −1.10 * | 0.49 | down |
| rno-miR-98-5p | −1.07 | 1.26 | down |
| rno-let-7a-5p | −1.03 | 1.21 | down |
| rno-miR-1-3p | −0.88 | 1.19 | down |
| rno-miR-206-3p | −0.86 | 1.23 | down |
| rno-miR-34b-3p | 0.63 | 0.46 | up |
| rno-miR-181a-2-3p | 0.75 * | 0.27 | up |
| miRNA | Average log2 Change Sensory Neuropathy/Control | log2 SD | Fold Change | Regulation | Confirmation |
|---|---|---|---|---|---|
| rno-miR-344b-1-3p | n.d. | n.d. | n.d. | n.d. | n.d. |
| rno-miR-466b-1-3p | −3.19 * | 1.59 | −9.13 * | down | yes |
| rno-miR-98-5p | −2.45 ** | 1.19 | −5.48 ** | down | yes |
| rno-let-7a-5p | −2.03 ** | 0.54 | −4.07 ** | down | yes |
| rno-miR-1-3p | −2.85 ** | 1.03 | −7.21 ** | down | yes |
| rno-miR-206-3p | −5.19 ** | 1.58 | −36.42 ** | down | yes |
| rno-miR-34b-3p | −2.93 ** | 1.76 | −7.60 ** | down | no |
| rno-miR-181a-2-3p | 3.81 ** | 1.55 | 14.03 ** | up | yes |
| Target | Regulated by | ||||||
|---|---|---|---|---|---|---|---|
| Abbreviation | Name | miR-466b | miR-98 | let-7a | miR-1 | miR-206 | miR-34b |
| IGF-1 | Insulin-like growth factor-1 | + | + | + | |||
| SLC2a-12 | Solute carrier family 2 facilitated glucose transporter member 12 | + | + | + | |||
| EIF-4e | Eukaryotic translation initiation factor 4e | + | + | + | |||
| ULK-2 | Unc-51 like autophagy activating kinase 2 | + | + | + | |||
| Target | Accession Number | Forward Primer | Reverse Primer | Efficiency | Product Size (bp) |
|---|---|---|---|---|---|
| IGF-1 | M15481.1 | CAGTTCGTGTGTGGACCAAG | GAGTCTTGGGCATGTCAGTG | 1.966 | 211 |
| SLC2a-12 | NM_001107451.1 | CCGAGACAAAGGGATGCTCT | CCTTGTAAGTCCTGCCACCA | 1.704 | 251 |
| EIF-4e | NM_053974.2 | CAAGCAAACCTTCGGTTGAT | CTCCCCGTTTGTTTTTCTCA | 1.743 | 163 |
| ULK-2 | NM_001191645.1 | CACAGAACGACCAATGGATG | TTGTTCGAAGGACCATGTGA | 2.00 | 161 |
| GAPDH | NM_017008.4 | GCATCTTCTTGTGCAGTGCC | GAGAAGGCAGCCCTGGTAAC | 1.991 | 105 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bencsik, P.; Kiss, K.; Ágg, B.; Baán, J.A.; Ágoston, G.; Varga, A.; Gömöri, K.; Mendler, L.; Faragó, N.; Zvara, Á.; et al. Sensory Neuropathy Affects Cardiac miRNA Expression Network Targeting IGF-1, SLC2a-12, EIF-4e, and ULK-2 mRNAs. Int. J. Mol. Sci. 2019, 20, 991. https://doi.org/10.3390/ijms20040991
Bencsik P, Kiss K, Ágg B, Baán JA, Ágoston G, Varga A, Gömöri K, Mendler L, Faragó N, Zvara Á, et al. Sensory Neuropathy Affects Cardiac miRNA Expression Network Targeting IGF-1, SLC2a-12, EIF-4e, and ULK-2 mRNAs. International Journal of Molecular Sciences. 2019; 20(4):991. https://doi.org/10.3390/ijms20040991
Chicago/Turabian StyleBencsik, Péter, Krisztina Kiss, Bence Ágg, Júlia A. Baán, Gergely Ágoston, Albert Varga, Kamilla Gömöri, Luca Mendler, Nóra Faragó, Ágnes Zvara, and et al. 2019. "Sensory Neuropathy Affects Cardiac miRNA Expression Network Targeting IGF-1, SLC2a-12, EIF-4e, and ULK-2 mRNAs" International Journal of Molecular Sciences 20, no. 4: 991. https://doi.org/10.3390/ijms20040991
APA StyleBencsik, P., Kiss, K., Ágg, B., Baán, J. A., Ágoston, G., Varga, A., Gömöri, K., Mendler, L., Faragó, N., Zvara, Á., Sántha, P., Puskás, L. G., Jancsó, G., & Ferdinandy, P. (2019). Sensory Neuropathy Affects Cardiac miRNA Expression Network Targeting IGF-1, SLC2a-12, EIF-4e, and ULK-2 mRNAs. International Journal of Molecular Sciences, 20(4), 991. https://doi.org/10.3390/ijms20040991