Knockout of the S-acyltransferase Gene, PbPAT14, Confers the Dwarf Yellowing Phenotype in First Generation Pear by ABA Accumulation
Abstract
1. Introduction
2. Results
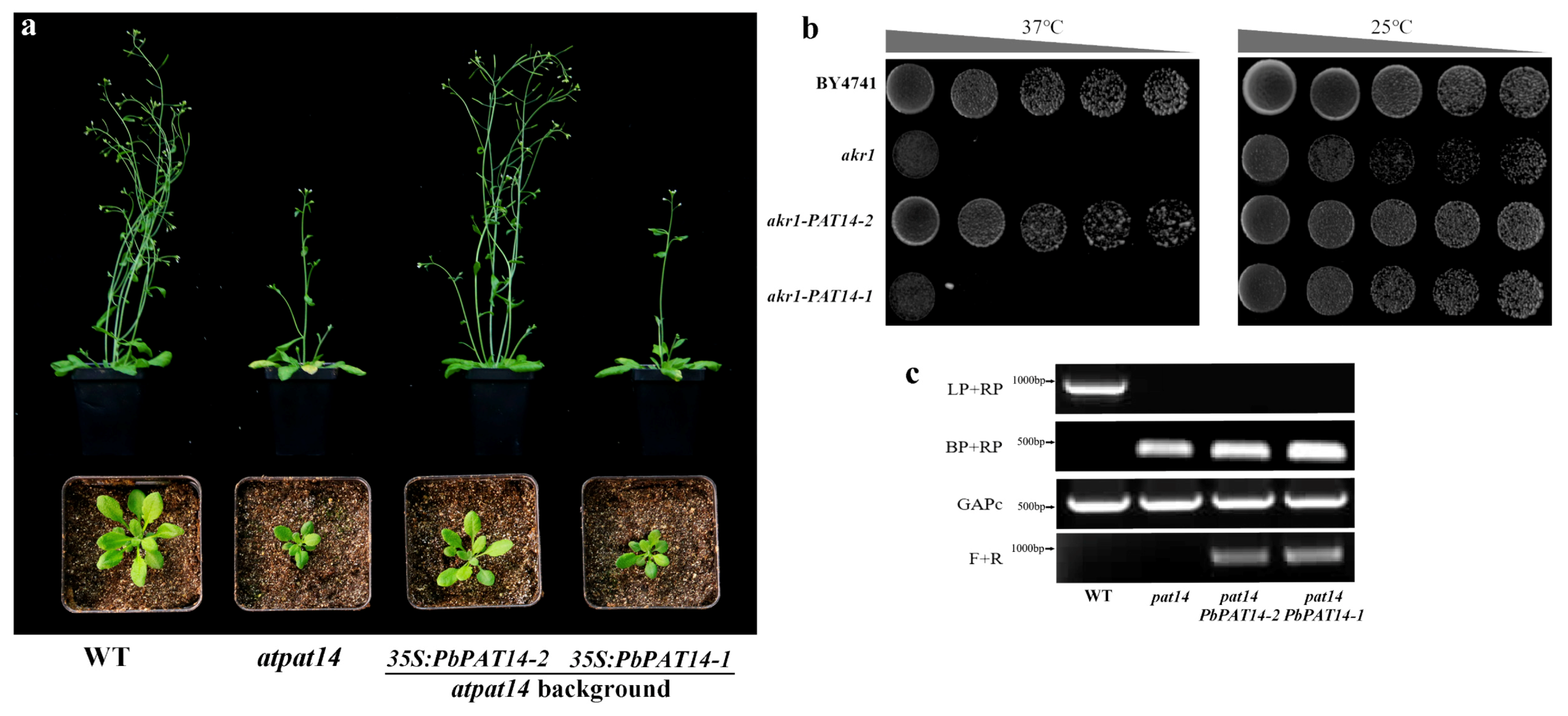
2.1. Identification and Molecular Characterization of the PbPAT14 Gene in Pear
2.2. Target Selection and Vector Construction
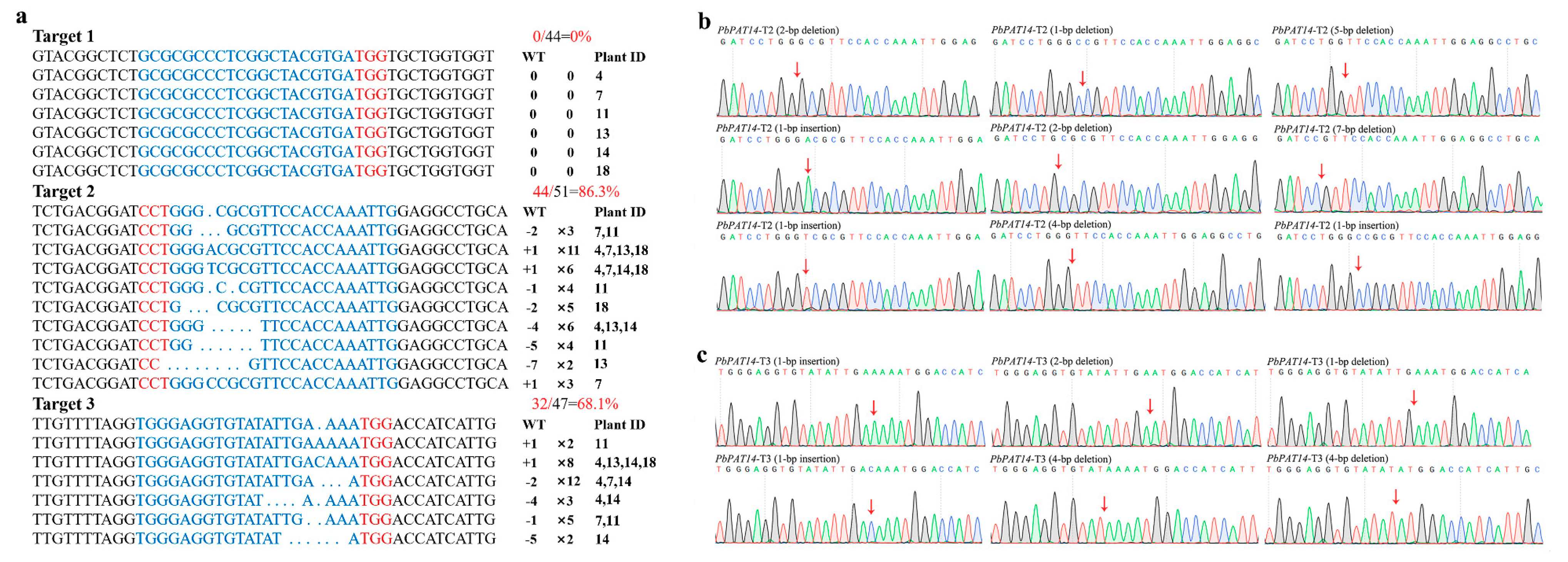
2.3. Pear Transformation and Identification of Transgenic Mutant Lines
2.4. Analysis of Potential Off-Target Mutations
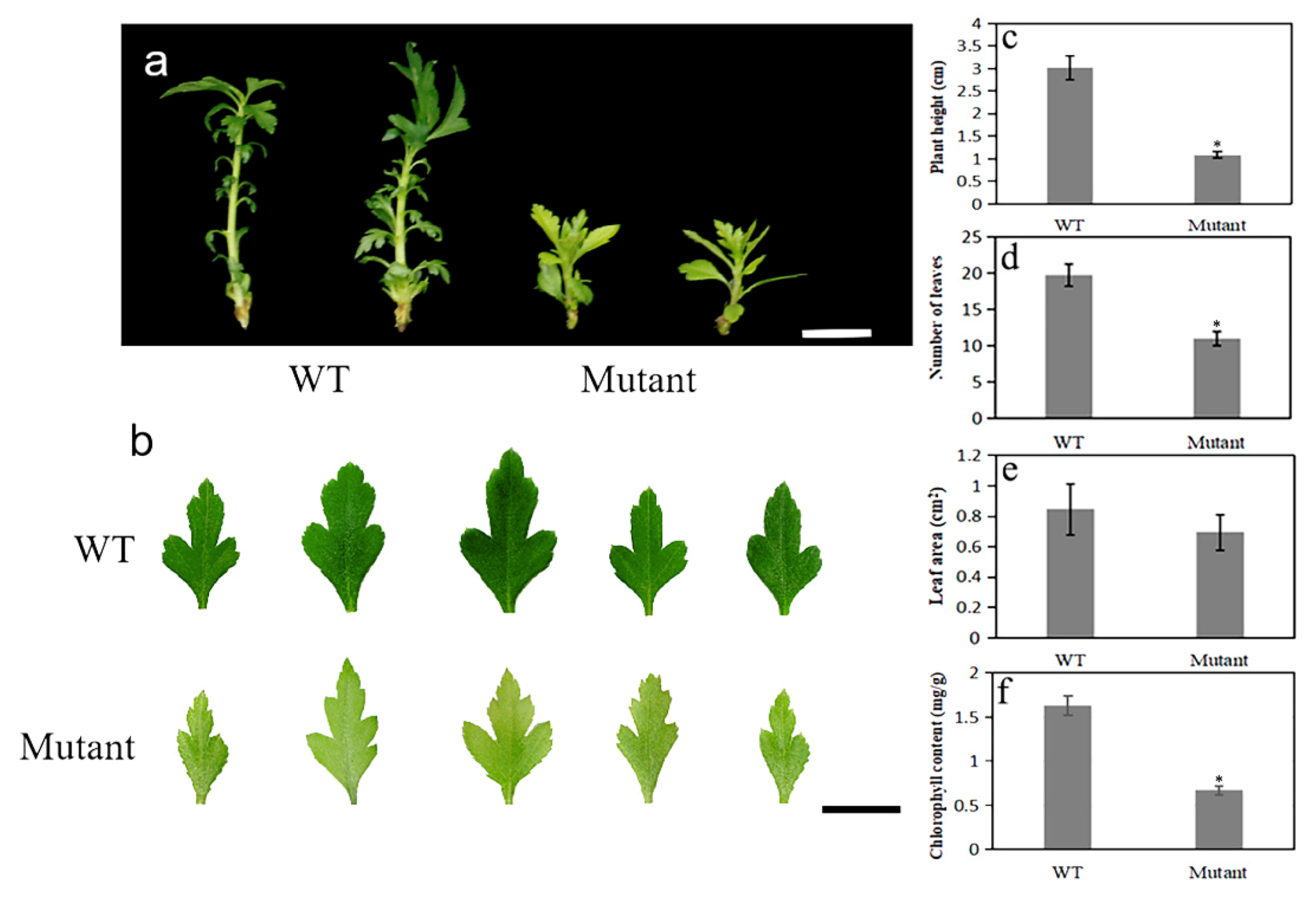
2.5. Dwarf Yellowing Phenotype of the pbpat14 Mutant Lines
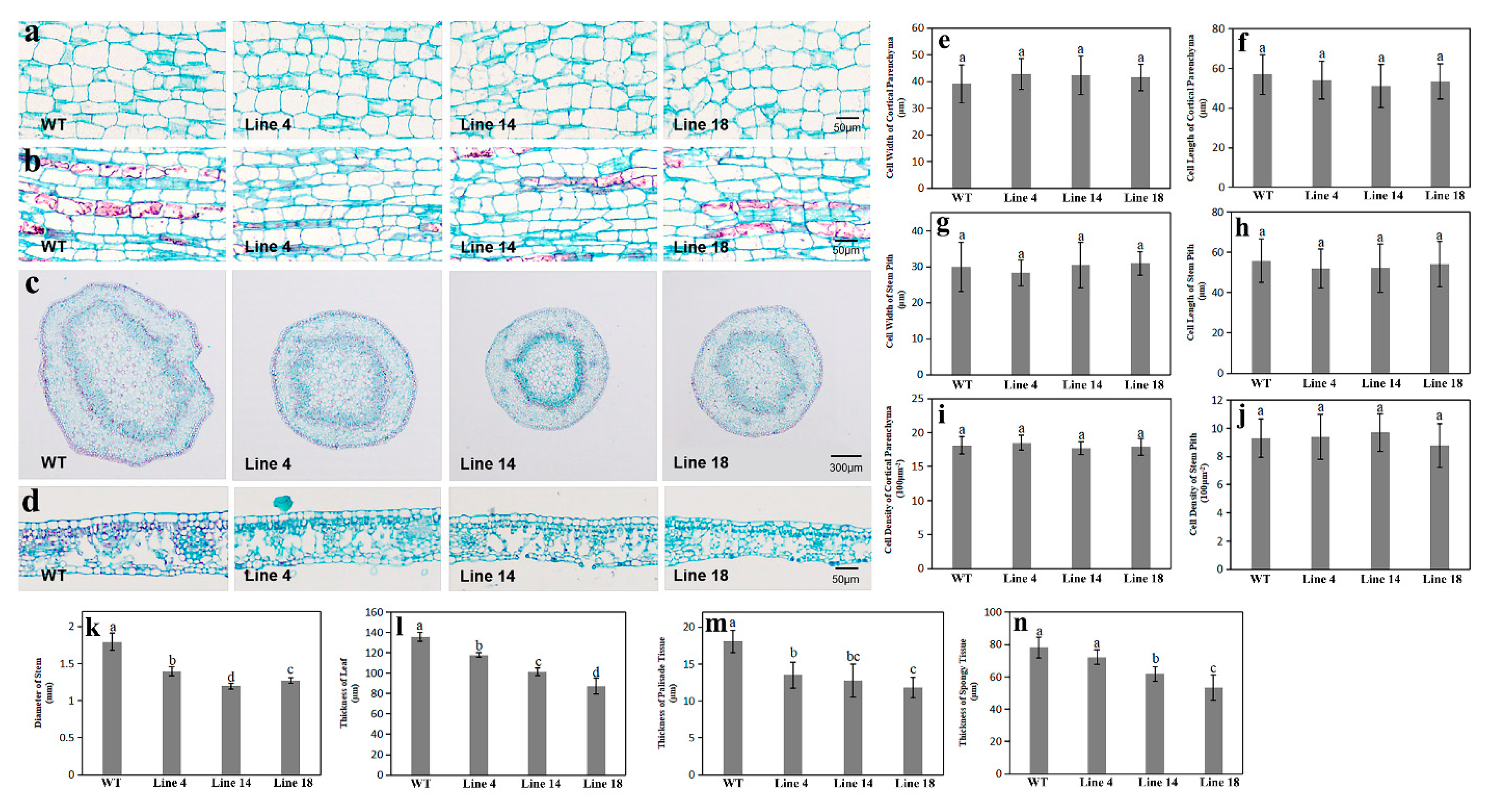
2.6. Ultrastructure Analysis of Stems and Leaves
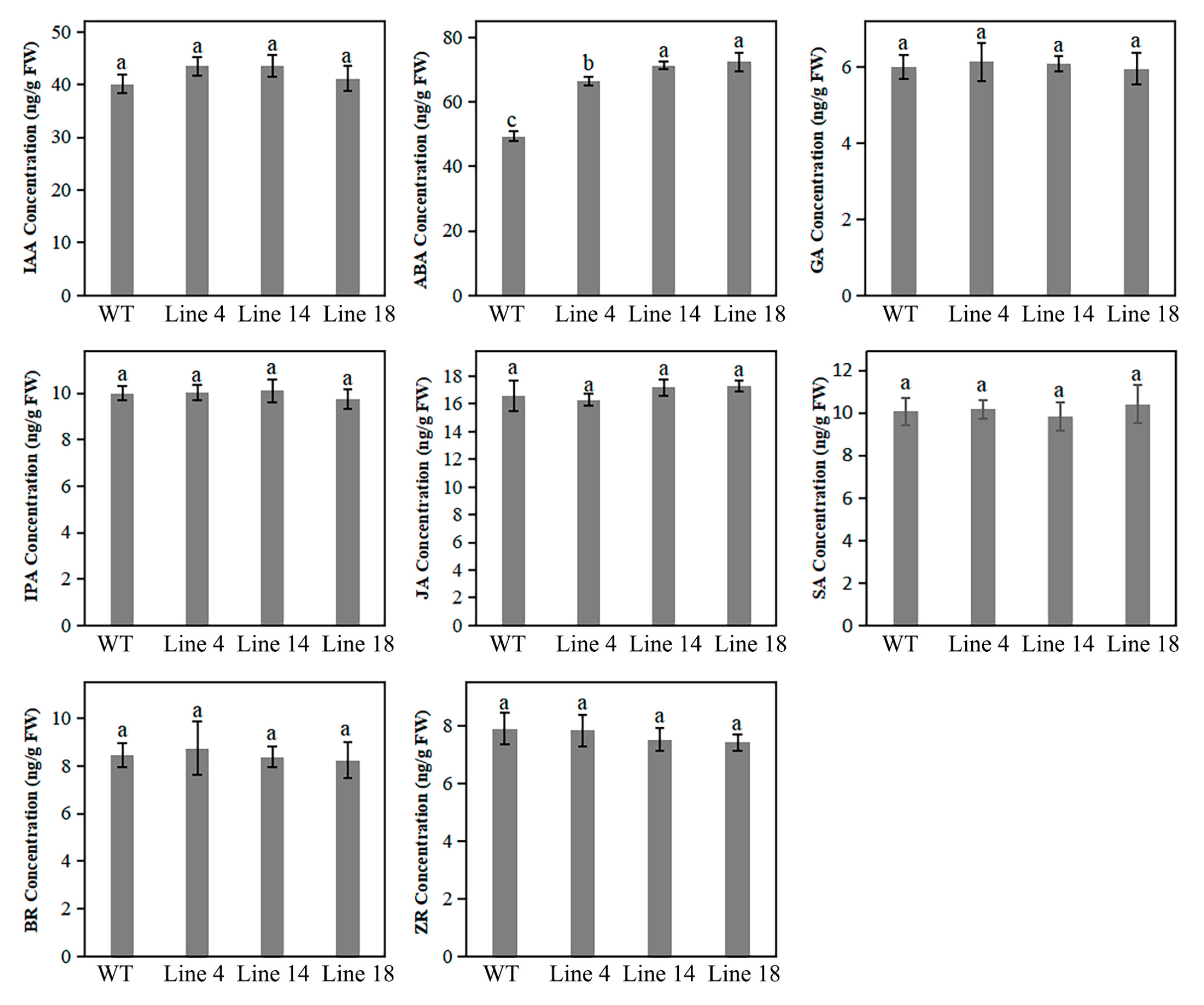
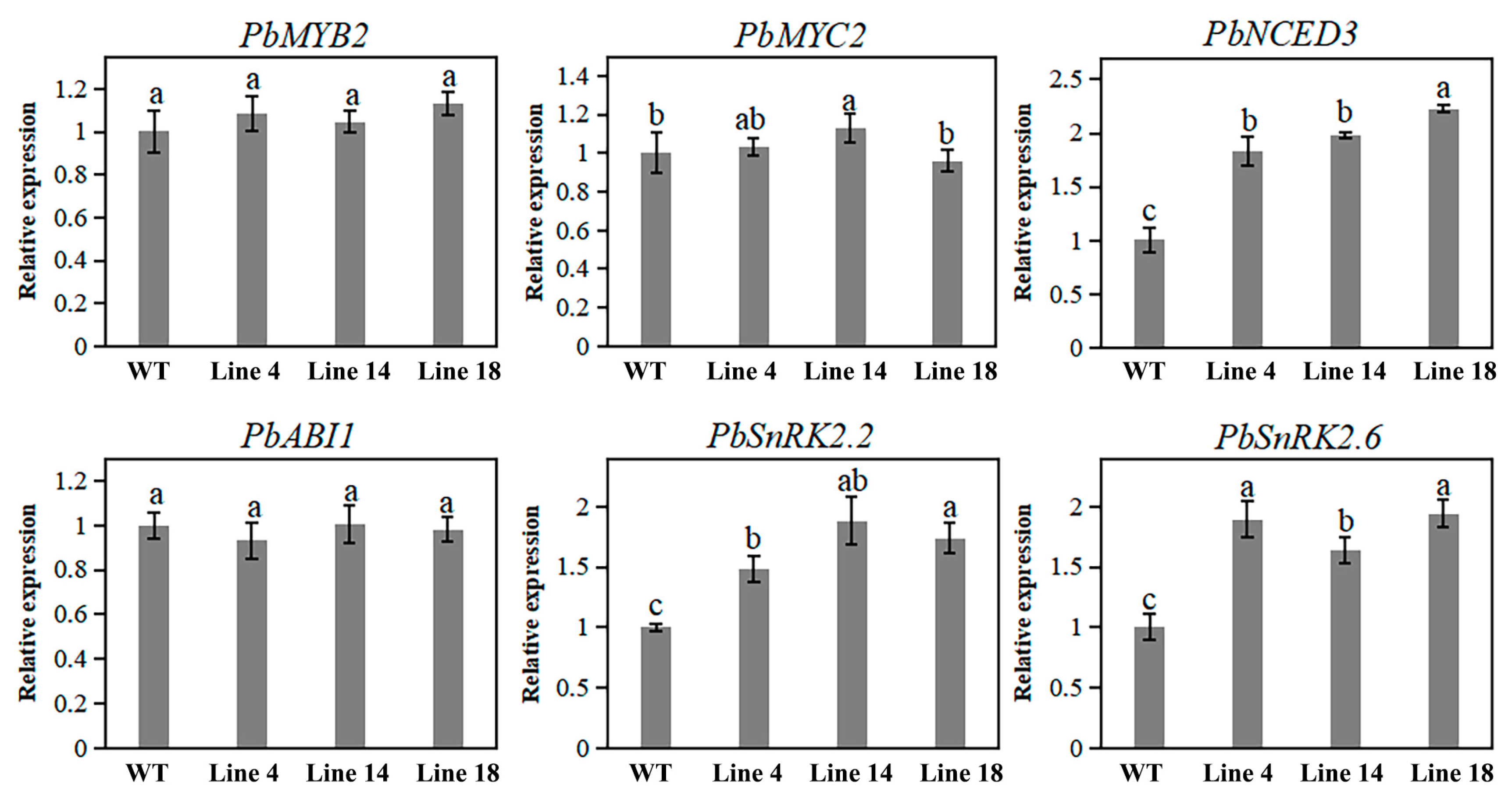
2.7. Knockout of PbPAT14 Induced ABA Accumulation
3. Discussion
4. Materials and Methods
4.1. PbPAT14 Identification and Phylogenetic Analysis
4.2. S-acylation Activity Analysis in Yeast and ARABIDOPSIS
4.3. CRISPR/Cas9 Target Site Selection and CRISPR/Cas9 Vector Construction
4.4. Transformation and Growth of Pear
4.5. Detection of Mutations
4.6. Off-Target Analysis
4.7. Growth Parameters
4.8. Measurement of Chlorophyll Content
4.9. Anatomical Structure Analysis
4.10. Concentrations of Plant Hormones
4.11. qRT-PCR Analysis
4.12. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Webster, A. Breeding and selection of apple and pear rootstocks. Proceedings of XXVI International Horticultural Congress: Genetics and Breeding of Tree Fruits and Nuts. Acta Hortic. 2003, 622, 499–512. [Google Scholar] [CrossRef]
- Xiao, Y.; Wang, C.; Tian, Y.; Yang, S.; Shen, J.; Liu, Q.; Zhang, H. Candidates responsible for dwarf pear phenotype as revealed by comparative transcriptome analysis. Mol. Breed. 2019, 39, 1. [Google Scholar] [CrossRef]
- Lauri, P.é.; Maguylo, K.; Trottier, C.J. Architecture and size relations: An essay on the apple (Malus× domestica, Rosaceae) tree. Am. J. Bot. 2006, 93, 357–368. [Google Scholar] [CrossRef] [PubMed]
- Prassinos, C.; Ko, J.H.; Lang, G.; Iezzoni, A.F.; Han, K.H. Rootstock-induced dwarfing in cherries is caused by differential cessation of terminal meristem growth and is triggered by rootstock-specific gene regulation. Tree Physiol. 2009, 29, 927–936. [Google Scholar] [CrossRef]
- Ou, C.; Jiang, S.; Wang, F.; Tang, C.; Hao, N. An RNA-Seq analysis of the pear (Pyrus communis L.) transcriptome, with a focus on genes associated with dwarf. Plant Gene 2015, 4, 69–77. [Google Scholar] [CrossRef]
- Maas, F. Proceedings of X International Pear Symposium: Evaluation of Pyrus and quince rootstocks for high density pear orchards. Acta Hortic. 2008, 800, 599–610. [Google Scholar] [CrossRef]
- Jacob, H. Proceedings of VIII International Symposium on Pear: New pear rootstocks from Geisenheim, Germany. Acta Hortic. 2002, 596, 337–344. [Google Scholar] [CrossRef]
- Webster, A.D. Dwarfing rootstocks: Past, present and future. Compact. Fruit Tree 2002, 35, 67–72. [Google Scholar]
- Elkins, R.; Bell, R.; Einhorn, T. Needs assessment for future US pear rootstock research directions based on the current state of pear production and rootstock research. J. Am. Pomol. Soc. 2012, 66, 153–163. [Google Scholar]
- Postman, J. Pear Germplasm Needs and Conservation. Pear Genome 2019, 35–50. [Google Scholar] [CrossRef]
- Ephritikhine, G.; Pagant, S.; Fujioka, S.; Takatsuto, S.; Lapous, D.; Caboche, M.; Kendrick, R.E.; Barbier-Brygoo, H. The sax1 mutation defines a new locus involved in the brassinosteroid biosynthesis pathway in Arabidopsis thaliana. Plant J. 1999, 18, 315–320. [Google Scholar] [CrossRef] [PubMed]
- Imai, A.; Komura, M.; Kawano, E.; Kuwashiro, Y.; Takahashi, T. A semi-dominant mutation in the ribosomal protein L10 gene suppresses the dwarf phenotype of the acl5 mutant in Arabidopsis thaliana. Plant J. 2008, 56, 881–890. [Google Scholar] [CrossRef] [PubMed]
- Foster, T.M.; Celton, J.M.; Chagné, D.; Tustin, D.S.; Gardiner, S.E. Two quantitative trait loci, Dw1 and Dw2, are primarily responsible for rootstock-induced dwarfing in apple. Hortic Res. 2015, 2, 15001. [Google Scholar] [CrossRef] [PubMed]
- Harrison, N.; Harrison, R.J.; Barber-Perez, N.; Cascant-Lopez, E.; Cobo-Medina, M.; Lipska, M.; Conde-Ruíz, R.; Brain, P.; Gregory, P.J.; Fernández-Fernández, F. A new three-locus model for rootstock-induced dwarfing in apple revealed by genetic mapping of root bark percentage. J. Exp. Bot. 2016, 67, 1871–1881. [Google Scholar] [CrossRef]
- Ma, Y.; Xue, H.; Zhang, L.; Zhang, F.; Ou, C.; Wang, F.; Zhang, Z. Involvement of auxin and brassinosteroid in dwarfism of autotetraploid apple (Malus × domestica). Sci Rep. 2016, 6, 26719. [Google Scholar] [CrossRef]
- Zheng, X.; Zhao, Y.; Shan, D.; Shi, K.; Wang, L.; Li, Q.; Wang, N.; Zhou, J.; Yao, J.; Xue, Y.; et al. MdWRKY 9 overexpression confers intensive dwarfing in the M26 rootstock of apple by directly inhibiting brassinosteroid synthetase MdDWF4 expression. New Phytol. 2018, 217, 1086–1098. [Google Scholar] [CrossRef]
- Sharp, R.E.; LeNoble, M.E.; Else, M.A.; Thorne, E.T.; Gherardi, F. Endogenous ABA maintains shoot growth in tomato independently of effects on plant water balance: Evidence for an interaction with ethylene. J. Exp. Bot. 2000, 51, 1575–1584. [Google Scholar] [CrossRef]
- Jia, D.; Gong, X.; Li, M.; Li, C.; Sun, T.; Ma, F. Overexpression of a novel apple NAC transcription factor gene, MdNAC1, confers the dwarf phenotype in transgenic apple (Malus domestica). Genes 2018, 9, 229. [Google Scholar] [CrossRef]
- Goto, N.; Pharis, R. Role of gibberellins in the development of floral organs of the gibberellin-deficient mutant, ga1-1, of Arabidopsis thaliana. Can. J. Bot. 1999, 77, 944–954. [Google Scholar] [CrossRef]
- Talon, M.; Koornneef, M.; Zeevaart, J. Endogenous gibberellins in Arabidopsis thaliana and possible steps blocked in the biosynthetic pathways of the semidwarf ga4 and ga5 mutants. Proc. Natl. Acad. Sci. USA 1990, 87, 7983–7987. [Google Scholar] [CrossRef]
- Noda, K.; Okuda, H.; Iwagaki, I. Indole acetic acid and abscisic acid levels in new shoots and fibrous roots of citrus scion-rootstock combinations. Sci. Hortic. 2000, 84, 245–254. [Google Scholar] [CrossRef]
- Tworkoski, T.; Fazio, G. Proceedings of IX International Symposium on Integrating Canopy: Physiological and morphological effects of size-controlling rootstocks on ’Fuji’ apple scions. Acta Hortic. 2011, 903, 865–872. [Google Scholar] [CrossRef]
- Zheng, L.; Liu, P.; Liu, Q.; Wang, T.; Dong, J. Dynamic Protein S-Acylation in Plants. Int J. Mol. Sci. 2019, 20, 560. [Google Scholar] [CrossRef] [PubMed]
- Batistic, O. Genomics and localization of the Arabidopsis DHHC-cysteine-rich domain S-acyltransferase protein family. Plant Physiol. 2012, 160, 1597–1612. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Qi, B. Progress toward Understanding Protein S-acylation: Prospective in Plants. Front. Plant Sci. 2017, 8, 346. [Google Scholar] [CrossRef]
- Putilina, T.; Wong, P.; Gentleman, S. The DHHC domain: A new highly conserved cysteine-rich motif. Mol. Cell Biochem. 1999, 195, 219–226. [Google Scholar] [CrossRef]
- Yuan, X.; Zhang, S.; Sun, M.; Liu, S.; Qi, B.; Li, X. Putative DHHC-cysteine-rich domain S-acyltransferase in plants. PLoS ONE 2013, 8, e75985. [Google Scholar] [CrossRef]
- Zhou, L.Z.; Li, S.; Feng, Q.N.; Zhang, Y.L.; Zhao, X.; Zeng, Y.L.; Wang, H.; Jiang, L.; Zhang, Y. Protein S-ACYL Transferase10 is critical for development and salt tolerance in Arabidopsis. Plant Cell 2013, 25, 1093–1107. [Google Scholar] [CrossRef]
- Qi, B.; Doughty, J.; Hooley, R. A Golgi and tonoplast localized S-acyl transferase is involved in cell expansion, cell division, vascular patterning and fertility in Arabidopsis. New Phytol. 2013, 200, 444–456. [Google Scholar] [CrossRef]
- Hemsley, P.A.; Kemp, A.C.; Grierson, C.S. The TIP GROWTH DEFECTIVE1 S-acyl transferase regulates plant cell growth in Arabidopsis. Plant Cell 2005, 17, 2554–2563. [Google Scholar] [CrossRef]
- Zhao, X.Y.; Wang, J.G.; Song, S.J.; Wang, Q.; Kang, H.; Zhang, Y.; Li, S. Precocious leaf senescence by functional loss of PROTEIN S-ACYL TRANSFERASE14 involves the NPR1-dependent salicylic acid signaling. Sci. Rep. 2016, 6, 20309. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Xu, J.; Li, G.; Wan, S.; Batistic, O.; Sun, M.; Zhang, Y.; Scott, R.; Qi, B. Protein S-acyl transferase 15 is involved in seed triacylglycerol catabolism during early seedling growth in Arabidopsis. J. Exp. Bot. 2019, 70, 5205–5216. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, H.J.; Morgan, C.; Bomblies, K.; Yang, W.; Qi, B. Both male and female gametogenesis require a fully functional protein S-acyl transferase 21 in Arabidopsis thaliana. Plant J. 2019. [Google Scholar] [CrossRef] [PubMed]
- Peng, D.; Tan, X.; Zhang, L.; Yuan, D.; Lin, J.; Liu, X.; Jiang, Y.; Zhou, B. Increasing branch and seed yield through heterologous expression of the novel rice S-acyl transferase gene OsPAT15 in Brassica napus L. Breed. Sci. 2018, 68, 326–335. [Google Scholar] [CrossRef] [PubMed]
- Zsögön, A.; Čermák, T.; Naves, E.R.; Notini, M.M.; Edel, K.H.; Weinl, S.; Freschi, L.; Voytas, D.F.; Kudla, J.; Peres, L.E.P. De novo domestication of wild tomato using genome editing. Nat. Biotechnol. 2018, 36, 1211–1216. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekaran, J.; Brumin, M.; Wolf, D.; Leibman, D.; Klap, C.; Pearlsman, M.; Sherman, A.; Arazi, T.; Gal-On, A. Development of broad virus resistance in non-transgenic cucumber using CRISPR/Cas9 technology. Mol. Plant Pathol. 2016, 17, 1140–1153. [Google Scholar] [CrossRef]
- Wang, X.; Tu, M.; Wang, D.; Liu, J.; Li, Y.; Li, Z.; Wang, Y.; Wang, X. CRISPR/Cas9-mediated efficient targeted mutagenesis in grape in the first generation. Plant Biotechnol. J. 2018, 16, 844–855. [Google Scholar] [CrossRef]
- Charrier, A.; Vergne, E.; Dousset, N.J.-P.; Richer, A.; Petiteau, A.; Chevreau, E. Efficient targeted mutagenesis in apple and first time edition of pear using the CRISPR-Cas9 system. Front. Plant. Sci. 2019, 10, 40. [Google Scholar] [CrossRef]
- Kihara, A.; Kurotsu, F.; Sano, T.; Iwaki, S.; Igarashi, Y. Long-chain base kinase Lcb4 Is anchored to the membrane through its palmitoylation by Akr1. Mol. Cell Biol. 2005, 25, 9189–9197. [Google Scholar] [CrossRef][Green Version]
- Linder, M.E.; Deschenes, R.J. Palmitoylation: Policing protein stability and traffic. Nat. Rev. Mol. Cell Biol. 2007, 8, 74–84. [Google Scholar] [CrossRef]
- Ohno, Y.; Kashio, A.; Ogata, R.; Ishitomi, A.; Yamazaki, Y.; Kihara, A. Analysis of substrate specificity of human DHHC protein acyltransferases using a yeast expression system. Mol. Biol. Cell. 2012, 23, 4543–4551. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Zhu, Q.; Chen, Y.; Liu, Y.G. CRISPR/Cas9 platforms for genome editing in plants: Developments and applications. Mol. Plant. 2016, 9, 961–974. [Google Scholar] [CrossRef] [PubMed]
- Jaganathan, D.; Ramasamy, K.; Sellamuthu, G.; Jayabalan, S.; Venkataraman, G. CRISPR for crop improvement: An update review. Front. Plant. Sci. 2018, 9, 985. [Google Scholar] [CrossRef] [PubMed]
- Soda, N.; Verma, L.; Giri, J. CRISPR-Cas9 based plant genome editing: Significance, opportunities and recent advances. Plant. Physiol. Biochem. 2018, 131, 2–11. [Google Scholar] [CrossRef] [PubMed]
- Fan, D.; Liu, T.; Li, C.; Jiao, B.; Li, S.; Hou, Y.; Luo, K. Efficient CRISPR/Cas9-mediated targeted mutagenesis in Populus in the first generation. Sci. Rep. 2015, 5, 12217. [Google Scholar] [CrossRef] [PubMed]
- Nishitani, C.; Hirai, N.; Komori, S.; Wada, M.; Okada, K.; Osakabe, K.; Yamamoto, T.; Osakabe, Y. Efficient genome editing in apple using a CRISPR/Cas9 system. Sci. Rep. 2016, 6, 31481. [Google Scholar] [CrossRef] [PubMed]
- Lincoln, C.; Britton, J.H.; Estelle, M. Growth and development of the axr1 mutants of Arabidopsis. Plant Cell 1990, 2, 1071–1080. [Google Scholar] [CrossRef]
- Depuydt, S.; Van Praet, S.; Nelissen, H.; Vanholme, B.; Vereecke, D. How plant hormones and their interactions affect cell growth. Mol. Cell Biol. Growth Differ. Plant Cells 2016, 174–195. [Google Scholar] [CrossRef]
- Finkelstein, R.R.; Rock, C.D. Abscisic acid biosynthesis and response. Arab. Book 2002, 1, e0058. [Google Scholar] [CrossRef]
- Finkelstein, R. Abscisic acid synthesis and response. Arabidopsis Book 2013, 11, e0166. [Google Scholar] [CrossRef]
- Yang, J.; Worley, E.; Udvardi, M. A NAP-AAO3 regulatory module promotes chlorophyll degradation via ABA biosynthesis in Arabidopsis leaves. Plant Cell 2014, 26, 4862–4874. [Google Scholar] [CrossRef] [PubMed]
- Rong, W.; Wang, X.; Wang, X.; Massart, S.; Zhang, Z. Molecular and ultrastructural mechanisms underlying yellow dwarf symptom formation in wheat after infection of Barley Yellow Dwarf Virus. Int. J. Mol. Sci. 2018, 19, 1187. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, M.; Ishikawa, A.; Jikumaru, Y.; Seki, M.; Umezawa, T.; Asami, T.; Maruyama-Nakashita, A.; Kudo, T.; Shinozaki, K.; Yoshida, S.; et al. Antagonistic interaction between systemic acquired resistance and the abscisic acid–mediated abiotic stress response in Arabidopsis. Plant Cell 2008, 20, 1678–1692. [Google Scholar] [CrossRef] [PubMed]
- Fan, J.; Hill, L.; Crooks, C.; Doerner, P.; Lamb, C. Abscisic acid has a key role in modulating diverse plant-pathogen interactions. Plant Physiol. 2009, 150, 1750–1761. [Google Scholar] [CrossRef] [PubMed]
- de Torres Zabala, M.; Bennett, M.H.; Truman, W.H.; Grant, M.R. Antagonism between salicylic and abscisic acid reflects early host–pathogen conflict and moulds plant defence responses. Plant J. 2009, 59, 375–386. [Google Scholar] [CrossRef] [PubMed]
- Denancé, N.; Sánchez-Vallet, A.; Goffner, D.; Molina, A. Disease resistance or growth: The role of plant hormones in balancing immune responses and fitness costs. Front. Plant Sci. 2013, 4, 155. [Google Scholar] [CrossRef]
- Prodhan, M.Y.; Munemasa, S.; Nahar, M.N.; Nakamura, Y.; Murata, Y. Guard cell salicylic acid signaling is integrated into abscisic acid signaling via the Ca2+/CPK-dependent pathway. Plant. Physiol. 2018, 178, 441–450. [Google Scholar] [CrossRef]
- Wu, J.; Wang, Z.; Shi, Z.; Zhang, S.; Ming, R.; Zhu, S.; Khan, M.A.; Tao, S.; Korban, S.S.; Wang, H.; et al. The genome of the pear (Pyrus bretschneideri Rehd.). Genome Res. 2013, 23, 396–408. [Google Scholar] [CrossRef]
- El-Gebali, S.; Mistry, J.; Bateman, A.; Eddy, S.R.; Luciani, A.; Potter, S.C.; Qureshi, M.; Richardson, L.J.; Salazar, G.A.; Smart, A.; et al. The Pfam protein families database in 2019. Nucleic Acids Res. 2019, 47, D427–D432. [Google Scholar] [CrossRef]
- Sander, J.D.; Maeder, M.L.; Reyon, D.; Voytas, D.F.; Joung, J.K.; Dobbs, D. ZiFiT (Zinc Finger Targeter): An updated zinc finger engineering tool. Nucleic Acids Res. 2010, 38, W462–W468. [Google Scholar] [CrossRef]
- Ma, X.; Liu, Y.G. CRISPR/Cas9-based multiplex genome editing in monocot and dicot plants. Curr. Protoc. Mol. Biol. 2016, 115, 31–36. [Google Scholar] [CrossRef] [PubMed]
- Bae, S.; Park, J.; Kim, J. Cas-OFFinder: A fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 2014, 30, 1473–1475. [Google Scholar] [CrossRef] [PubMed]
- Katsiarimpa, A.; Kalinowska, K.; Anzenberger, F.; Weis, C.; Ostertag, M.; Tsutsumi, C.; Schwechheimer, C.; Brunner, F.; Hückelhoven, R.; Isono, E. The deubiquitinating enzyme AMSH1 and the ESCRT-III subunit VPS2. 1 are required for autophagic degradation in Arabidopsis. Plant Cell 2013, 25, 2236–2252. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Sawhney, V.; Steeves, T. Staining of paraffin-embedded plant material in safranin and fast green without prior removal of the paraffin. Can. J. Bot. 1993, 71, 996–999. [Google Scholar] [CrossRef]
- Yang, J.; Zhang, J.; Wang, Z.; Zhu, Q.; Wang, W. Hormonal changes in the grains of rice subjected to water stress during grain filling. Plant Physiol. 2001, 127, 315–323. [Google Scholar] [CrossRef]
- Weiler, E.; Jourdan, P.; Conrad, W. Levels of indole-3-acetic acid in intact and decapitated coleoptiles as determined by a specific and highly sensitive solid-phase enzyme immunoassay. Planta 1981, 153, 561–571. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Zhang, Y.; Chen, L. Expression and Regulation of PpEIN3b during Fruit Ripening and Senescence via Integrating SA, Glucose, and ACC Signaling in Pear (Pyrus pyrifolia Nakai. Whangkeumbae). Genes (Basel) 2019, 10, 476. [Google Scholar] [CrossRef] [PubMed]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pang, H.; Yan, Q.; Zhao, S.; He, F.; Xu, J.; Qi, B.; Zhang, Y. Knockout of the S-acyltransferase Gene, PbPAT14, Confers the Dwarf Yellowing Phenotype in First Generation Pear by ABA Accumulation. Int. J. Mol. Sci. 2019, 20, 6347. https://doi.org/10.3390/ijms20246347
Pang H, Yan Q, Zhao S, He F, Xu J, Qi B, Zhang Y. Knockout of the S-acyltransferase Gene, PbPAT14, Confers the Dwarf Yellowing Phenotype in First Generation Pear by ABA Accumulation. International Journal of Molecular Sciences. 2019; 20(24):6347. https://doi.org/10.3390/ijms20246347
Chicago/Turabian StylePang, Hongguang, Qi Yan, Shuliang Zhao, Fang He, Jianfeng Xu, Baoxiu Qi, and Yuxing Zhang. 2019. "Knockout of the S-acyltransferase Gene, PbPAT14, Confers the Dwarf Yellowing Phenotype in First Generation Pear by ABA Accumulation" International Journal of Molecular Sciences 20, no. 24: 6347. https://doi.org/10.3390/ijms20246347
APA StylePang, H., Yan, Q., Zhao, S., He, F., Xu, J., Qi, B., & Zhang, Y. (2019). Knockout of the S-acyltransferase Gene, PbPAT14, Confers the Dwarf Yellowing Phenotype in First Generation Pear by ABA Accumulation. International Journal of Molecular Sciences, 20(24), 6347. https://doi.org/10.3390/ijms20246347



