Insights into Gene Regulatory Networks in Chondrocytes
Abstract
1. The Concept of Genomic Control of Development
2. The Discovery of Cis-Trans Regulatory Mechanisms in Chondrocytes
3. Genome-Scale Analysis of Sox9 Action in Chondrogenesis
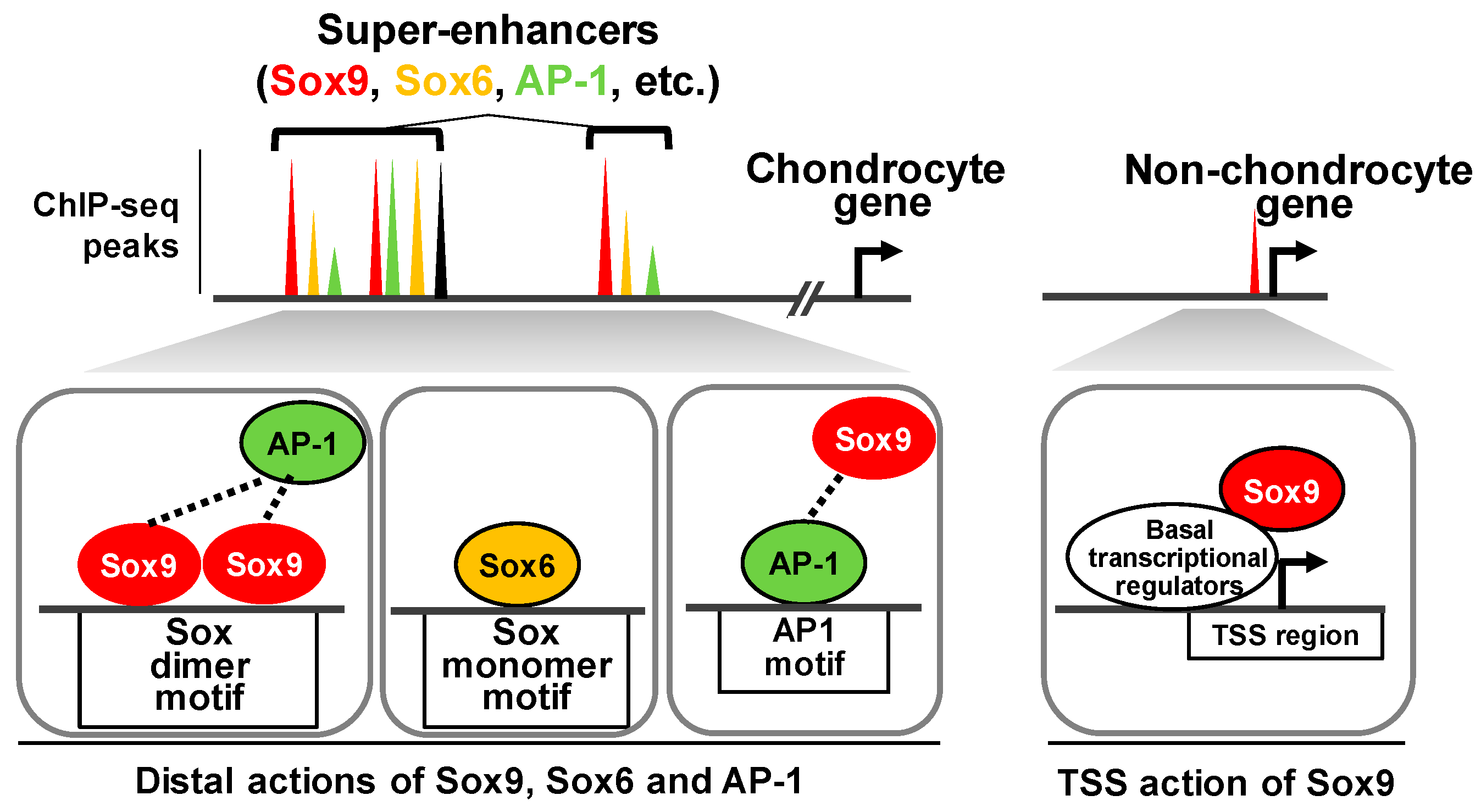
4. Cooperative Actions of Multiple Transcription Factors to Establish a Chondrocyte Program
5. Pioneer Factors for Establishing Cell Type-Distinct Gene Regulatory Networks
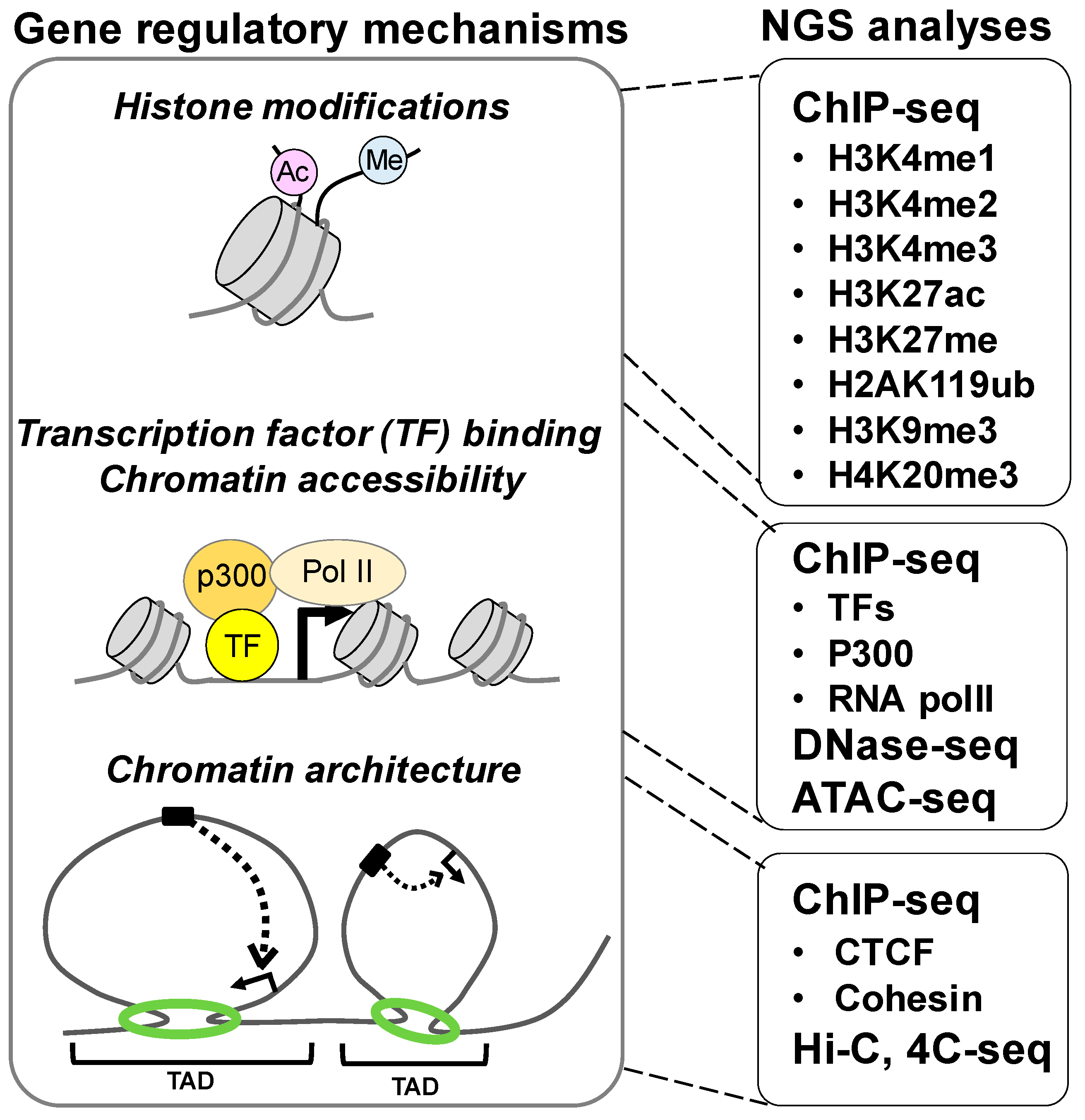
6. Higher-Order Gene Regulatory Machineries
7. Gene Regulatory Networks Underlying the Association of Human Genetic Variants with Chondrocyte-Related Diseases
8. Concluding Remarks and Future Perspective
Funding
Conflicts of Interest
References
- Peter, I.; Davidson, E.H. Genom. Control. Process: Development and Evolution; Academic Press: Cambridge, MA, USA, 2015. [Google Scholar]
- Waddington, C.H. The Strategy of the Genes: A Discussion of Some Aspects of Theoretical Biology; Allen & Unwin: London, UK, 1957. [Google Scholar]
- Britten, R.J.; Davidson, E.H. Gene regulation for higher cells: A theory. Science 1969, 165, 349–357. [Google Scholar] [CrossRef] [PubMed]
- Banerji, J.; Rusconi, S.; Schaffner, W. Expression of a beta-globin gene is enhanced by remote SV40 DNA sequences. Cell 1981, 27, 299–308. [Google Scholar] [CrossRef]
- Schaffner, W. Enhancers, enhancers—From their discovery to today’s universe of transcription enhancers. Biol. Chem. 2015, 396, 311–327. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, A.; Rossi, P.; de Crombrugghe, B. Transcriptional control of the mouse alpha 2(I) collagen gene: Functional deletion analysis of the promoter and evidence for cell-specific expression. Mol. Cell. Biol. 1986, 6, 347–354. [Google Scholar] [CrossRef]
- Horton, W.; Miyashita, T.; Kohno, K.; Hassell, J.R.; Yamada, Y. Identification of a phenotype-specific enhancer in the first intron of the rat collagen II gene. Proc. Natl. Acad. Sci. USA 1987, 84, 8864–8868. [Google Scholar] [CrossRef]
- Lefebvre, V.; Zhou, G.; Mukhopadhyay, K.; Smith, C.N.; Zhang, Z.; Eberspaecher, H.; Zhou, X.; Sinha, S.; Maity, S.N.; de Crombrugghe, B. An 18-base-pair sequence in the mouse proalpha1(II) collagen gene is sufficient for expression in cartilage and binds nuclear proteins that are selectively expressed in chondrocytes. Mol. Cell. Biol. 1996, 16, 4512–4523. [Google Scholar] [CrossRef]
- Lefebvre, V.; Huang, W.; Harley, V.R.; Goodfellow, P.N.; de Crombrugghe, B. SOX9 is a potent activator of the chondrocyte-specific enhancer of the pro alpha1(II) collagen gene. Mol. Cell. Biol. 1997, 17, 2336–2346. [Google Scholar] [CrossRef]
- Bell, D.M.; Leung, K.K.; Wheatley, S.C.; Ng, L.J.; Zhou, S.; Ling, K.W.; Sham, M.H.; Koopman, P.; Tam, P.P.; Cheah, K.S.; et al. SOX9 directly regulates the type-II collagen gene. Nat. Genet. 1997, 16, 174–178. [Google Scholar] [CrossRef]
- Ng, L.J.; Wheatley, S.; Muscat, G.E.; Conway-Campbell, J.; Bowles, J.; Wright, E.; Bell, D.M.; Tam, P.P.; Cheah, K.S.; Koopman, P.; et al. SOX9 binds DNA, activates transcription, and coexpresses with type II collagen during chondrogenesis in the mouse. Dev. Biol. 1997, 183, 108–121. [Google Scholar] [CrossRef]
- Akiyama, H.; Lefebvre, V. Unraveling the transcriptional regulatory machinery in chondrogenesis. J. Bone Miner. Metab. 2011, 29, 390–395. [Google Scholar] [CrossRef]
- Lefebvre, V.; Dvir-Ginzberg, M. SOX9 and the many facets of its regulation in the chondrocyte lineage. Connect. Tissue Res. 2016, 58, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Foster, J.W.; Dominguez-Steglich, M.A.; Guioli, S.; Kwok, C.; Weller, P.A.; Stevanovic, M.; Weissenbach, J.; Mansour, S.; Young, I.D.; Goodfellow, P.N.; et al. Campomelic dysplasia and autosomal sex reversal caused by mutations in an SRY-related gene. Nature 1994, 372, 525–530. [Google Scholar] [CrossRef] [PubMed]
- Wagner, T.; Wirth, J.; Meyer, J.; Zabel, B.; Held, M.; Zimmer, J.; Pasantes, J.; Bricarelli, F.D.; Keutel, J.; Hustert, E.; et al. Autosomal sex reversal and campomelic dysplasia are caused by mutations in and around the SRY-related gene SOX9. Cell 1994, 79, 1111–1120. [Google Scholar] [CrossRef]
- Bi, W.; Deng, J.M.; Zhang, Z.; Behringer, R.R.; de Crombrugghe, B. Sox9 is required for cartilage formation. Nat. Genet. 1999, 22, 85–89. [Google Scholar] [CrossRef]
- Akiyama, H.; Chaboissier, M.C.; Martin, J.F.; Schedl, A.; de Crombrugghe, B. The transcription factor Sox9 has essential roles in successive steps of the chondrocyte differentiation pathway and is required for expression of Sox5 and Sox6. Genes Dev. 2002, 16, 2813–2828. [Google Scholar] [CrossRef]
- Ikegami, D.; Akiyama, H.; Suzuki, A.; Nakamura, T.; Nakano, T.; Yoshikawa, H.; Tsumaki, N. Sox9 sustains chondrocyte survival and hypertrophy in part through Pik3ca-Akt pathways. Development 2011, 138, 1507–1519. [Google Scholar] [CrossRef]
- Dy, P.; Wang, W.; Bhattaram, P.; Wang, Q.; Wang, L.; Ballock, R.T.; Lefebvre, V. Sox9 directs hypertrophic maturation and blocks osteoblast differentiation of growth plate chondrocytes. Dev. Cell 2012, 22, 597–609. [Google Scholar] [CrossRef]
- Akiyama, H.; Lyons, J.P.; Mori-Akiyama, Y.; Yang, X.; Zhang, R.; Zhang, Z.; Deng, J.M.; Taketo, M.M.; Nakamura, T.; Behringer, R.R.; et al. Interactions between Sox9 and beta-catenin control chondrocyte differentiation. Genes Dev. 2004, 18, 1072–1087. [Google Scholar] [CrossRef]
- Kozhemyakina, E.; Lassar, A.B.; Zelzer, E. A pathway to bone: Signaling molecules and transcription factors involved in chondrocyte development and maturation. Development 2015, 142, 817–831. [Google Scholar] [CrossRef]
- Liu, C.F.; Samsa, W.E.; Zhou, G.; Lefebvre, V. Transcriptional control of chondrocyte specification and differentiation. Semin. Cell Dev. Biol. 2017, 62, 34–49. [Google Scholar] [CrossRef]
- Perino, M.; Veenstra, G.J. Chromatin Control of Developmental Dynamics and Plasticity. Dev. Cell 2016, 38, 610–620. [Google Scholar] [CrossRef] [PubMed]
- Ohba, S.; He, X.; Hojo, H.; McMahon, A.P. Distinct Transcriptional Programs Underlie Sox9 Regulation of the Mammalian Chondrocyte. Cell Rep. 2015, 12, 229–243. [Google Scholar] [CrossRef]
- Oh, C.D.; Lu, Y.; Liang, S.; Mori-Akiyama, Y.; Chen, D.; de Crombrugghe, B.; Yasuda, H. SOX9 regulates multiple genes in chondrocytes, including genes encoding ECM proteins, ECM modification enzymes, receptors, and transporters. PLoS ONE 2014, 9, 107577. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.F.; Lefebvre, V. The transcription factors SOX9 and SOX5/SOX6 cooperate genome-wide through super-enhancers to drive chondrogenesis. Nucleic Acids Res. 2015, 43, 8183–8203. [Google Scholar] [CrossRef]
- Furumatsu, T.; Tsuda, M.; Yoshida, K.; Taniguchi, N.; Ito, T.; Hashimoto, M.; Asahara, H. Sox9 and p300 cooperatively regulate chromatin-mediated transcription. J. Biol. Chem. 2005, 280, 35203–35208. [Google Scholar] [CrossRef] [PubMed]
- Whyte, W.A.; Orlando, D.A.; Hnisz, D.; Abraham, B.J.; Lin, C.Y.; Kagey, M.H.; Rahl, P.B.; Lee, T.I.; Young, R.A. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 2013, 153, 307–319. [Google Scholar] [CrossRef] [PubMed]
- Sabari, B.R.; Dall’Agnese, A.; Boija, A.; Klein, I.A.; Coffey, E.L.; Shrinivas, K.; Abraham, B.J.; Hannett, N.M.; Zamudio, A.V.; Manteiga, J.C.; et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 2018, 361. [Google Scholar] [CrossRef] [PubMed]
- Hnisz, D.; Shrinivas, K.; Young, R.A.; Chakraborty, A.K.; Sharp, P.A. A Phase Separation Model for Transcriptional Control. Cell 2017, 169, 13–23. [Google Scholar] [CrossRef]
- Bernard, P.; Tang, P.; Liu, S.; Dewing, P.; Harley, V.R.; Vilain, E. Dimerization of SOX9 is required for chondrogenesis, but not for sex determination. Hum. Mol. Genet. 2003, 12, 1755–1765. [Google Scholar] [CrossRef]
- Farley, E.K.; Olson, K.M.; Zhang, W.; Brandt, A.J.; Rokhsar, D.S.; Levine, M.S. Suboptimization of developmental enhancers. Science 2015, 350, 325–328. [Google Scholar] [CrossRef]
- Long, H.K.; Prescott, S.L.; Wysocka, J. Ever-Changing Landscapes: Transcriptional Enhancers in Development and Evolution. Cell 2016, 167, 1170–1187. [Google Scholar] [CrossRef] [PubMed]
- Rickels, R.; Shilatifard, A. Enhancer Logic and Mechanics in Development and Disease. Trends Cell Biol. 2018, 28, 608–630. [Google Scholar] [CrossRef] [PubMed]
- Smits, P.; Li, P.; Mandel, J.; Zhang, Z.; Deng, J.M.; Behringer, R.R.; de Crombrugghe, B.; Lefebvre, V. The transcription factors L-Sox5 and Sox6 are essential for cartilage formation. Dev. Cell 2001, 1, 277–290. [Google Scholar] [CrossRef]
- Ikeda, T.; Kamekura, S.; Mabuchi, A.; Kou, I.; Seki, S.; Takato, T.; Nakamura, K.; Kawaguchi, H.; Ikegawa, S.; Chung, U.I. The combination of SOX5, SOX6, and SOX9 (the SOX trio) provides signals sufficient for induction of permanent cartilage. Arthritis Rheum. 2004, 50, 3561–3573. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Ohba, S.; Hojo, H.; McMahon, A.P. AP-1 family members act with Sox9 to promote chondrocyte hypertrophy. Development 2016, 143, 3012–3023. [Google Scholar] [CrossRef] [PubMed]
- Karreth, F.; Hoebertz, A.; Scheuch, H.; Eferl, R.; Wagner, E.F. The AP1 transcription factor Fra2 is required for efficient cartilage development. Development 2004, 131, 5717–5725. [Google Scholar] [CrossRef] [PubMed]
- Behrens, A.; Haigh, J.; Mechta-Grigoriou, F.; Nagy, A.; Yaniv, M.; Wagner, E.F. Impaired intervertebral disc formation in the absence of Jun. Development 2003, 130, 103–109. [Google Scholar] [CrossRef]
- Tan, Z.; Niu, B.; Tsang, K.Y.; Melhado, I.G.; Ohba, S.; He, X.; Huang, Y.; Wang, C.; McMahon, A.P.; Jauch, R.; et al. Synergistic co-regulation and competition by a SOX9-GLI-FOXA phasic transcriptional network coordinate chondrocyte differentiation transitions. PLoS Genet. 2018, 14, 1007346. [Google Scholar] [CrossRef]
- Iwafuchi-Doi, M.; Zaret, K.S. Cell fate control by pioneer transcription factors. Development 2016, 143, 1833–1837. [Google Scholar] [CrossRef]
- Liu, C.F.; Angelozzi, M.; Haseeb, A.; Lefebvre, V. SOX9 is dispensable for the initiation of epigenetic remodeling and the activation of marker genes at the onset of chondrogenesis. Development 2018, 145. [Google Scholar] [CrossRef]
- Adam, R.C.; Yang, H.; Rockowitz, S.; Larsen, S.B.; Nikolova, M.; Oristian, D.S.; Polak, L.; Kadaja, M.; Asare, A.; Zheng, D.; et al. Pioneer factors govern super-enhancer dynamics in stem cell plasticity and lineage choice. Nature 2015, 521, 366–370. [Google Scholar] [CrossRef] [PubMed]
- Andrey, G.; Mundlos, S. The three-dimensional genome: Regulating gene expression during pluripotency and development. Development 2017, 144, 3646–3658. [Google Scholar] [CrossRef] [PubMed]
- Will, A.J.; Cova, G.; Osterwalder, M.; Chan, W.L.; Wittler, L.; Brieske, N.; Heinrich, V.; de Villartay, J.P.; Vingron, M.; Klopocki, E.; et al. Composition and dosage of a multipartite enhancer cluster control developmental expression of Ihh (Indian hedgehog). Nat. Genet. 2017, 49, 1539–1545. [Google Scholar] [CrossRef] [PubMed]
- Osterwalder, M.; Barozzi, I.; Tissieres, V.; Fukuda-Yuzawa, Y.; Mannion, B.J.; Afzal, S.Y.; Lee, E.A.; Zhu, Y.; Plajzer-Frick, I.; Pickle, C.S.; et al. Enhancer redundancy provides phenotypic robustness in mammalian development. Nature 2018, 554, 239–243. [Google Scholar] [CrossRef] [PubMed]
- Dixon, J.R.; Selvaraj, S.; Yue, F.; Kim, A.; Li, Y.; Shen, Y.; Hu, M.; Liu, J.S.; Ren, B. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 2012, 485, 376–380. [Google Scholar] [CrossRef] [PubMed]
- Lupianez, D.G.; Kraft, K.; Heinrich, V.; Krawitz, P.; Brancati, F.; Klopocki, E.; Horn, D.; Kayserili, H.; Opitz, J.M.; Laxova, R.; et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 2015, 161, 1012–1025. [Google Scholar] [CrossRef] [PubMed]
- Kragesteen, B.K.; Spielmann, M.; Paliou, C.; Heinrich, V.; Schopflin, R.; Esposito, A.; Annunziatella, C.; Bianco, S.; Chiariello, A.M.; Jerkovic, I.; et al. Dynamic 3D chromatin architecture contributes to enhancer specificity and limb morphogenesis. Nat. Genet. 2018, 50, 1463–1473. [Google Scholar] [CrossRef]
- Lettice, L.A.; Horikoshi, T.; Heaney, S.J.; van Baren, M.J.; van der Linde, H.C.; Breedveld, G.J.; Joosse, M.; Akarsu, N.; Oostra, B.A.; Endo, N.; et al. Disruption of a long-range cis-acting regulator for Shh causes preaxial polydactyly. Proc. Natl. Acad. Sci. USA 2002, 99, 7548–7553. [Google Scholar] [CrossRef]
- Capellini, T.D.; Chen, H.; Cao, J.; Doxey, A.C.; Kiapour, A.M.; Schoor, M.; Kingsley, D.M. Ancient selection for derived alleles at a GDF5 enhancer influencing human growth and osteoarthritis risk. Nat. Genet. 2017, 49, 1202–1210. [Google Scholar] [CrossRef]
- Guo, M.; Liu, Z.; Willen, J.; Shaw, C.P.; Richard, D.; Jagoda, E.; Doxey, A.C.; Hirschhorn, J.; Capellini, T.D. Epigenetic profiling of growth plate chondrocytes sheds insight into regulatory genetic variation influencing height. eLife 2017, 6. [Google Scholar] [CrossRef]
- Maurano, M.T.; Humbert, R.; Rynes, E.; Thurman, R.E.; Haugen, E.; Wang, H.; Reynolds, A.P.; Sandstrom, R.; Qu, H.; Brody, J.; et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 2012, 337, 1190–1195. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Chang, J.C.; Hon, C.C.; Fukui, N.; Tanaka, N.; Zhang, Z.; Lee, M.T.M.; Minoda, A. Chromatin accessibility landscape of articular knee cartilage reveals aberrant enhancer regulation in osteoarthritis. Sci. Rep. 2018, 8, 15499. [Google Scholar] [CrossRef] [PubMed]
- Kawata, M.; Mori, D.; Kanke, K.; Hojo, H.; Ohba, S.; Chung, U.I.; Yano, F.; Masaki, H.; Otsu, M.; Nakauchi, H.; et al. Simple and Robust Differentiation of Human Pluripotent Stem Cells toward Chondrocytes by Two Small-Molecule Compounds. Stem Cell Rep. 2019, 13, 530–544. [Google Scholar] [CrossRef] [PubMed]
- Buenrostro, J.D.; Wu, B.; Litzenburger, U.M.; Ruff, D.; Gonzales, M.L.; Snyder, M.P.; Chang, H.Y.; Greenleaf, W.J. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 2015, 523, 486–490. [Google Scholar] [CrossRef]
- Li, J.; Luo, H.; Wang, R.; Lang, J.; Zhu, S.; Zhang, Z.; Fang, J.; Qu, K.; Lin, Y.; Long, H.; et al. Systematic Reconstruction of Molecular Cascades Regulating GP Development Using Single-Cell RNA-Seq. Cell Rep. 2016, 15, 1467–1480. [Google Scholar] [CrossRef]
- Mizuhashi, K.; Nagata, M.; Matsushita, Y.; Ono, W.; Ono, N. Growth Plate Borderline Chondrocytes Behave as Transient Mesenchymal Precursor Cells. J. Bone Miner. Res. 2019, 34, 1387–1392. [Google Scholar] [CrossRef]
- Feregrino, C.; Sacher, F.; Parnas, O.; Tschopp, P. A single-cell transcriptomic atlas of the developing chicken limb. BMC Genom. 2019, 20, 401. [Google Scholar] [CrossRef]
- Ji, Q.; Zheng, Y.; Zhang, G.; Hu, Y.; Fan, X.; Hou, Y.; Wen, L.; Li, L.; Xu, Y.; Wang, Y.; et al. Single-cell RNA-seq analysis reveals the progression of human osteoarthritis. Ann. Rheum. Dis. 2019, 78, 100–110. [Google Scholar] [CrossRef]
- Hilton, I.B.; D’Ippolito, A.M.; Vockley, C.M.; Thakore, P.I.; Crawford, G.E.; Reddy, T.E.; Gersbach, C.A. Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat. Biotechnol. 2015, 33, 510–517. [Google Scholar] [CrossRef]
- Kearns, N.A.; Pham, H.; Tabak, B.; Genga, R.M.; Silverstein, N.J.; Garber, M.; Maehr, R. Functional annotation of native enhancers with a Cas9-histone demethylase fusion. Nat. Methods 2015, 12, 401–403. [Google Scholar] [CrossRef]
- Gilbert, L.A.; Larson, M.H.; Morsut, L.; Liu, Z.; Brar, G.A.; Torres, S.E.; Stern-Ginossar, N.; Brandman, O.; Whitehead, E.H.; Doudna, J.A.; et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 2013, 154, 442–451. [Google Scholar] [CrossRef] [PubMed]
- Gasperini, M.; Hill, A.J.; McFaline-Figueroa, J.L.; Martin, B.; Kim, S.; Zhang, M.D.; Jackson, D.; Leith, A.; Schreiber, J.; Noble, W.S.; et al. A Genome-wide Framework for Mapping Gene Regulation via Cellular Genetic Screens. Cell 2019, 176, 377–390. [Google Scholar] [CrossRef] [PubMed]
- Mochizuki, Y.; Chiba, T.; Kataoka, K.; Yamashita, S.; Sato, T.; Kato, T.; Takahashi, K.; Miyamoto, T.; Kitazawa, M.; Hatta, T.; et al. Combinatorial CRISPR/Cas9 Approach to Elucidate a Far-Upstream Enhancer Complex for Tissue-Specific Sox9 Expression. Dev. Cell 2018, 46, 794–806. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hojo, H.; Ohba, S. Insights into Gene Regulatory Networks in Chondrocytes. Int. J. Mol. Sci. 2019, 20, 6324. https://doi.org/10.3390/ijms20246324
Hojo H, Ohba S. Insights into Gene Regulatory Networks in Chondrocytes. International Journal of Molecular Sciences. 2019; 20(24):6324. https://doi.org/10.3390/ijms20246324
Chicago/Turabian StyleHojo, Hironori, and Shinsuke Ohba. 2019. "Insights into Gene Regulatory Networks in Chondrocytes" International Journal of Molecular Sciences 20, no. 24: 6324. https://doi.org/10.3390/ijms20246324
APA StyleHojo, H., & Ohba, S. (2019). Insights into Gene Regulatory Networks in Chondrocytes. International Journal of Molecular Sciences, 20(24), 6324. https://doi.org/10.3390/ijms20246324



