A Snapshot of The Tumor Microenvironment in Colorectal Cancer: The Liquid Biopsy
Abstract
1. Introduction
2. Cross-Talk between Tumor and Microenvironmental Cells by Exosome Transference
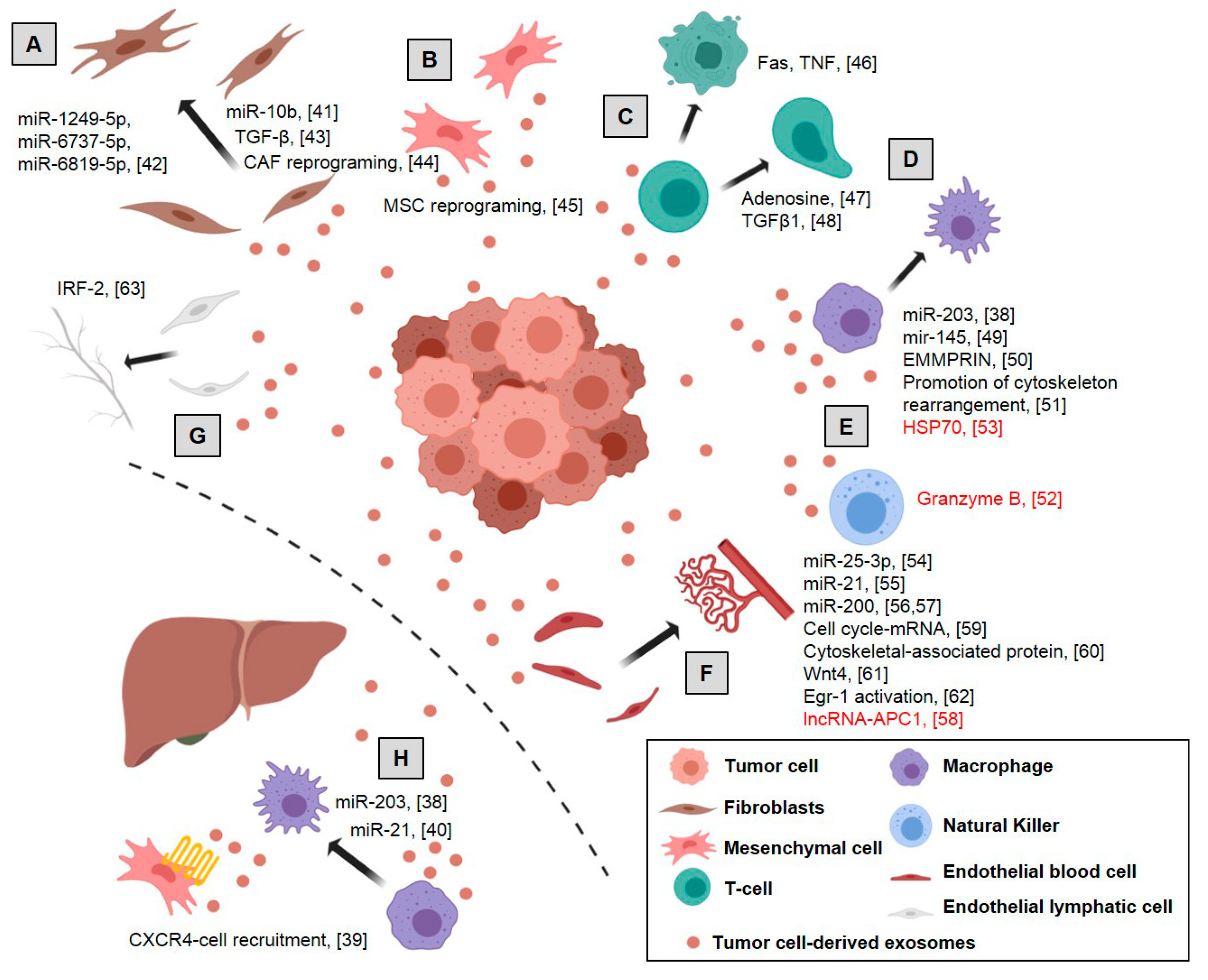
2.1. Tumor Cell-Derived Exosomes’ Effect on Microenvironmental Cells
2.1.1. Pre-Metastatic Niche
2.1.2. Fibroblasts Switch to Cancer-Associated Fibroblasts and ECM Remodeling
2.1.3. Mesenchymal Stem Cell Reprogramming
2.1.4. Immune Cells
2.1.5. Vascular Cells
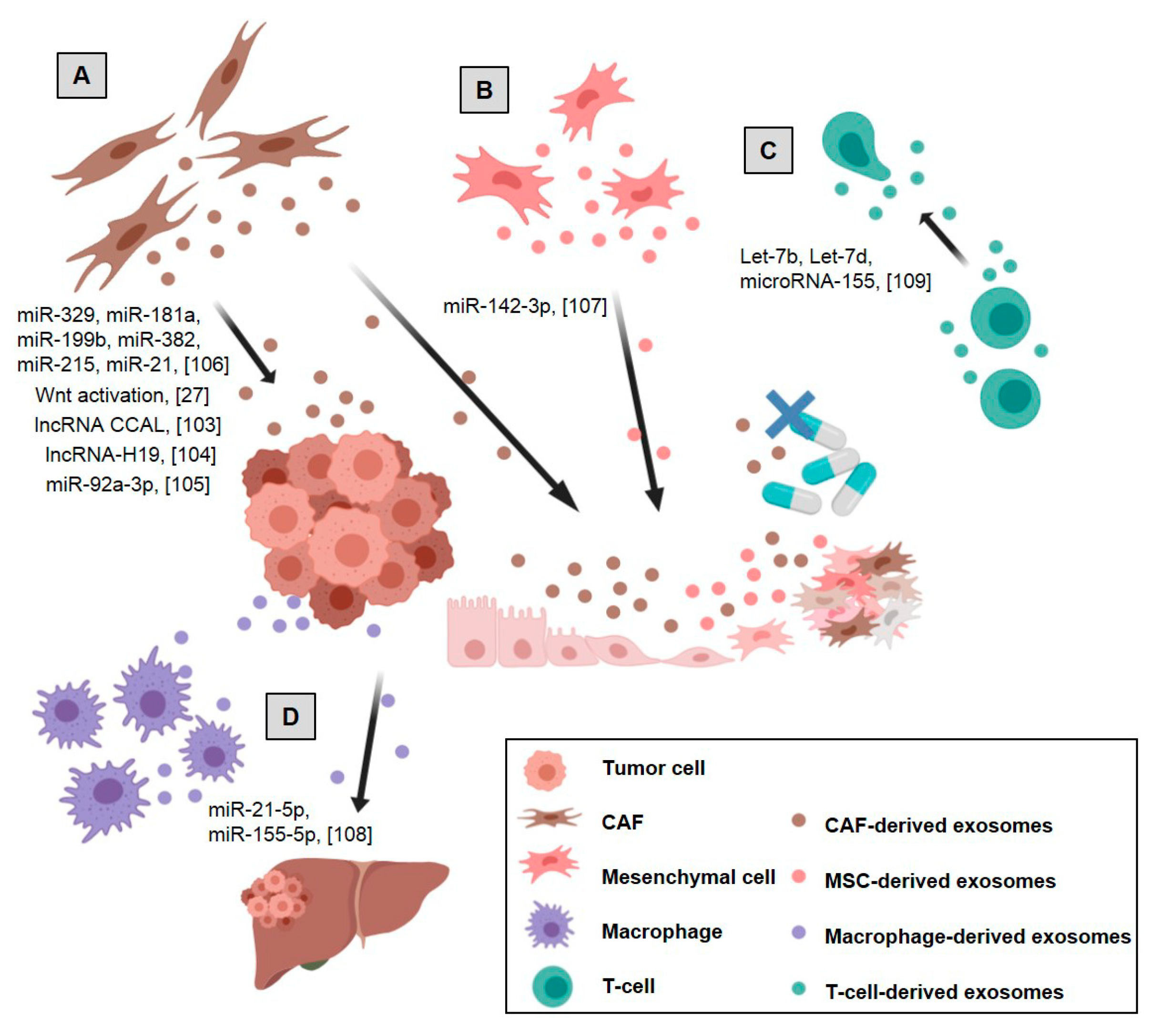
2.2. Stromal Cell-Derived Exosomes’ Effects on Tumor and Stromal Cells
2.2.1. CAF-Derived Exosomes
2.2.2. Mesenchymal Stem Cell-Derived Exosomes
2.2.3. Endothelial Cell-Derived Exosomes
2.2.4. Immune Cell-Derived Exosomes
3. Microenvironment-Derived Components as Liquid Biopsy Biomarkers
3.1. Soluble Factors
3.2. Exosomes as Biomarkers
3.3. Tumor-Related Circulating Cells
3.3.1. Circulating Endothelial Cells
3.3.2. CAFs and Mesenchymal Circulating Cells
3.3.3. Circulating Immune Cells
4. Therapeutic Applications
5. Concluding Remarks
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| 5-FU | 5-Fluorouracil |
| CAFs | Cancer-Associated Fibroblasts |
| CECs | Circulating Endothelial Cells |
| CRC | Colorectal Cancer |
| CTCs | Circulating Tumor Cells |
| ctDNA | circulating tumor DNA |
| ECM | Extracellular Matrix |
| lncRNAs | long non-coding RNAs |
| miRNAs | micro RNAs |
| MSCs | Mesenchymal Stem Cells |
| TGF-β | Transforming Growth Factor beta |
References
- Crowley, E.; Di Nicolantonio, F.; Loupakis, F.; Bardelli, A. Liquid biopsy: Monitoring cancer-genetics in the blood. Nat. Rev. Clin. Oncol. 2013, 10, 472. [Google Scholar] [CrossRef] [PubMed]
- Cheung, A.H.-K.; Chow, C.; To, K.-F. Latest development of liquid biopsy. J. Thorac. Dis. 2018, 10, S1645–S1651. [Google Scholar] [CrossRef] [PubMed]
- Yamada, T.; Matsuda, A.; Koizumi, M.; Shinji, S.; Takahashi, G.; Iwai, T.; Takeda, K.; Ueda, K.; Yokoyama, Y.; Hara, K.; et al. Liquid Biopsy for the Management of Patients with Colorectal Cancer. Digestion 2019, 99, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Giannopoulou, L.; Zavridou, M.; Kasimir-Bauer, S.; Lianidou, E.S. Liquid biopsy in ovarian cancer: The potential of circulating miRNAs and exosomes. Transl. Res. 2019, 205, 77–91. [Google Scholar] [CrossRef] [PubMed]
- Alix-Panabières, C.; Schwarzenbach, H.; Pantel, K. Circulating Tumor Cells and Circulating Tumor DNA. Annu. Rev. Med. 2012, 63, 199–215. [Google Scholar] [CrossRef]
- Friel, A.M.; Corcoran, C.; Crown, J.; O’Driscoll, L. Relevance of circulating tumor cells, extracellular nucleic acids, and exosomes in breast cancer. Breast Cancer Res. Treat. 2010, 123, 613–625. [Google Scholar] [CrossRef]
- Osumi, H.; Shinozaki, E.; Yamaguchi, K.; Zembutsu, H. Clinical utility of circulating tumor DNA for colorectal cancer. Cancer Sci. 2019, 110, 1148–1155. [Google Scholar] [CrossRef]
- Marrugo-Ramírez, J.; Mir, M.; Samitier, J. Blood-Based Cancer Biomarkers in Liquid Biopsy: A Promising Non-Invasive Alternative to Tissue Biopsy. Int. J. Mol. Sci. 2018, 19, 2877. [Google Scholar] [CrossRef]
- Bettegowda, C.; Sausen, M.; Leary, R.J.; Kinde, I.; Wang, Y.; Agrawal, N.; Bartlett, B.R.; Wang, H.; Luber, B.; Alani, R.M.; et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl. Med. 2014, 6, ra24–ra224. [Google Scholar] [CrossRef]
- Dawson, S.-J.; Tsui, D.W.Y.; Murtaza, M.; Biggs, H.; Rueda, O.M.; Chin, S.-F.; Dunning, M.J.; Gale, D.; Forshew, T.; Mahler-Araujo, B.; et al. Analysis of Circulating Tumor DNA to Monitor Metastatic Breast Cancer. N. Engl. J. Med. 2013, 368, 1199–1209. [Google Scholar] [CrossRef]
- Earl, J.; Garcia-Nieto, S.; Martinez-Avila, J.C.; Montans, J.; Sanjuanbenito, A.; Rodríguez-Garrote, M.; Lisa, E.; Mendía, E.; Lobo, E.; Malats, N.; et al. Circulating tumor cells (Ctc) and kras mutant circulating free Dna (cfdna) detection in peripheral blood as biomarkers in patients diagnosed with exocrine pancreatic cancer. BMC Cancer 2015, 15, 797. [Google Scholar] [CrossRef] [PubMed]
- Cohen, S.J.; Punt, C.J.A.; Iannotti, N.; Saidman, B.H.; Sabbath, K.D.; Gabrail, N.Y.; Picus, J.; Morse, M.A.; Mitchell, E.; Miller, M.C.; et al. Prognostic significance of circulating tumor cells in patients with metastatic colorectal cancer. Ann. Oncol. 2009, 20, 1223–1229. [Google Scholar] [CrossRef] [PubMed]
- Cristofanilli, M.; Hayes, D.F.; Budd, G.T.; Ellis, M.J.; Stopeck, A.; Reuben, J.M.; Doyle, G.V.; Matera, J.; Allard, W.J.; Miller, M.C.; et al. Circulating Tumor Cells: A Novel Prognostic Factor for Newly Diagnosed Metastatic Breast Cancer. J. Clin. Oncol. 2005, 23, 1420–1430. [Google Scholar] [CrossRef] [PubMed]
- Mong, J.; Tan, M.H. Size-Based Enrichment Technologies for Non-cancerous Tumor-Derived Cells in Blood. Trends Biotechnol. 2018, 36, 511–522. [Google Scholar] [CrossRef]
- Hinger, S.A.; Cha, D.J.; Franklin, J.L.; Higginbotham, J.N.; Dou, Y.; Ping, J.; Shu, L.; Prasad, N.; Levy, S.; Zhang, B.; et al. Diverse Long RNAs Are Differentially Sorted into Extracellular Vesicles Secreted by Colorectal Cancer Cells. Cell Rep. 2018, 25, 715–725.e4. [Google Scholar] [CrossRef]
- Sun, Z.; Yang, S.; Zhou, Q.; Wang, G.; Song, J.; Li, Z.; Zhang, Z.; Xu, J.; Xia, K.; Chang, Y.; et al. Emerging role of exosome-derived long non-coding RNAs in tumor microenvironment. Mol. Cancer 2018, 17, 82. [Google Scholar] [CrossRef]
- Li, I.; Nabet, B.Y. Exosomes in the tumor microenvironment as mediators of cancer therapy resistance. Mol. Cancer 2019, 18, 32. [Google Scholar] [CrossRef]
- Herrera, M.; Llorens, C.; Rodríguez, M.; Herrera, A.; Ramos, R.; Gil, B.; Candia, A.; Larriba, M.J.; Garre, P.; Earl, J.; et al. Differential distribution and enrichment of non-coding RNAs in exosomes from normal and Cancer-associated fibroblasts in colorectal cancer. Mol. Cancer 2018, 17, 114. [Google Scholar] [CrossRef]
- Madhavan, B.; Yue, S.; Galli, U.; Rana, S.; Gross, W.; Müller, M.; Giese, N.A.; Kalthoff, H.; Becker, T.; Büchler, M.W.; et al. Combined evaluation of a panel of protein and miRNA serum-exosome biomarkers for pancreatic cancer diagnosis increases sensitivity and specificity. Int. J. Cancer 2015, 136, 2616–2627. [Google Scholar] [CrossRef]
- Zhu, C.; Ren, C.; Han, J.; Ding, Y.; Du, J.; Dai, N.; Dai, J.; Ma, H.; Hu, Z.; Shen, H.; et al. A five-microRNA panel in plasma was identified as potential biomarker for early detection of gastric cancer. Br. J. Cancer 2014, 110, 2291–2299. [Google Scholar] [CrossRef]
- Martin, O.A.; Anderson, R.L.; Russell, P.A.; Ashley Cox, R.; Ivashkevich, A.; Swierczak, A.; Doherty, J.P.; Jacobs, D.H.M.; Smith, J.; Siva, S.; et al. Mobilization of Viable Tumor Cells into the Circulation during Radiation Therapy. Int. J. Radiat. Oncol. Biol. Phys. 2014, 88, 395–403. [Google Scholar] [CrossRef] [PubMed]
- Perumal, V.; Corica, T.; Dharmarajan, A.M.; Sun, Z.; Dhaliwal, S.S.; Dass, C.R.; Dass, J. Circulating Tumour Cells (CTC), Head and Neck Cancer and Radiotherapy; Future Perspectives. Cancers 2019, 11, 367. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, H.; Enomoto, A.; Woods, S.L.; Burt, A.D.; Takahashi, M.; Worthley, D.L. Cancer-associated fibroblasts in gastrointestinal cancer. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 282–295. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, R. The biology and function of fibroblasts in cancer. Nat. Rev. Cancer 2016, 16, 582–598. [Google Scholar] [CrossRef]
- Erez, N.; Truitt, M.; Olson, P.; Arron, S.T.; Hanahan, D. Cancer-Associated Fibroblasts Are Activated in Incipient Neoplasia to Orchestrate Tumor-Promoting Inflammation in an NF-kappaB-Dependent Manner. Cancer Cell 2010, 17, 135–147. [Google Scholar] [CrossRef]
- Erdogan, B.; Webb, D.J. Cancer-associated fibroblasts modulate growth factor signaling and extracellular matrix remodeling to regulate tumor metastasis. Biochem. Soc. Trans. 2017, 45, 229–236. [Google Scholar] [CrossRef]
- Hu, Y.; Yan, C.; Mu, L.; Huang, K.; Li, X.; Tao, D.; Wu, Y.; Qin, J. Fibroblast-Derived Exosomes Contribute to Chemoresistance through Priming Cancer Stem Cells in Colorectal Cancer. PLoS ONE 2015, 10, e0125625. [Google Scholar] [CrossRef]
- Junttila, M.R.; de Sauvage, F.J. Influence of tumour micro-environment heterogeneity on therapeutic response. Nature 2013, 501, 346–354. [Google Scholar] [CrossRef]
- Van Niel, G.; D’Angelo, G.; Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef]
- Nogués, L.; Benito-Martin, A.; Hergueta-Redondo, M.; Peinado, H. The influence of tumour-derived extracellular vesicles on local and distal metastatic dissemination. Mol. Aspects Med. 2018, 60, 15–26. [Google Scholar] [CrossRef]
- Hoshino, A.; Costa-Silva, B.; Shen, T.L.; Rodrigues, G.; Hashimoto, A.; Tesic Mark, M.; Molina, H.; Kohsaka, S.; Di Giannatale, A.; Ceder, S.; et al. Tumour exosome integrins determine organotropic metastasis. Nature 2015, 527, 329–335. [Google Scholar] [CrossRef] [PubMed]
- Simon, T.; Pinioti, S.; Schellenberger, P.; Rajeeve, V.; Wendler, F.; Cutillas, P.R.; King, A.; Stebbing, J.; Giamas, G. Shedding of bevacizumab in tumour cells-derived extracellular vesicles as a new therapeutic escape mechanism in glioblastoma. Mol. Cancer 2018, 17, 132. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Liu, R.-X.; Chan, K.-W.; Hu, J.; Zhang, J.; Wei, L.; Tan, H.; Yang, X.; Liu, H. Exosomal transfer of p-STAT3 promotes acquired 5-FU resistance in colorectal cancer cells. J. Exp. Clin. Cancer Res. 2019, 38, 320. [Google Scholar] [CrossRef] [PubMed]
- Ren, D.; Lin, B.; Zhang, X.; Peng, Y.; Ye, Z.; Ma, Y.; Liang, Y.; Cao, L.; Li, X.; Li, R.; et al. Maintenance of cancer stemness by miR-196b-5p contributes to chemoresistance of colorectal cancer cells via activating STAT3 signaling pathway. Oncotarget 2017, 8, 49807–49823. [Google Scholar] [CrossRef]
- Zhang, S.; Zhang, Y.; Qu, J.; Che, X.; Fan, Y.; Hou, K.; Guo, T.; Deng, G.; Song, N.; Li, C.; et al. Exosomes promote cetuximab resistance via the PTEN/Aktv pathway in colon cancer cells. Braz. J. Med. Biol. Res. 2018, 51. [Google Scholar] [CrossRef]
- Yang, Y.-N.; Zhang, R.; Du, J.-W.; Yuan, H.-H.; Li, Y.-J.; Wei, X.-L.; Du, X.-X.; Jiang, S.-L.; Han, Y. Predictive role of UCA1-containing exosomes in cetuximab-resistant colorectal cancer. Cancer Cell Int. 2018, 18, 164. [Google Scholar] [CrossRef]
- Wang, T.; Nasser, M.I.; Shen, J.; Qu, S.; He, Q.; Zhao, M. Functions of Exosomes in the Triangular Relationship between the Tumor, Inflammation, and Immunity in the Tumor Microenvironment. J. Immunol. Res. 2019, 2019, 1–10. [Google Scholar] [CrossRef]
- Takano, Y.; Masuda, T.; Iinuma, H.; Yamaguchi, R.; Sato, K.; Tobo, T.; Hirata, H.; Kuroda, Y.; Nambara, S.; Hayashi, N.; et al. Circulating exosomal microRNA-203 is associated with metastasis possibly via inducing tumor-associated macrophages in colorectal cancer. Oncotarget 2017, 8, 78598–78613. [Google Scholar] [CrossRef]
- Wang, X.; Ding, X.; Nan, L.; Wang, Y.; Wang, J.; Yan, Z.; Zhang, W.; Sun, J.; Zhu, W.; Ni, B.; et al. Investigation of the roles of exosomes in colorectal cancer liver metastasis. Oncol. Rep. 2015, 33, 2445–2453. [Google Scholar] [CrossRef]
- Shao, Y.; Chen, T.; Zheng, X.; Yang, S.; Xu, K.; Chen, X.; Xu, F.; Wang, L.; Shen, Y.; Wang, T.; et al. Colorectal cancer-derived small extracellular vesicles establish an inflammatory premetastatic niche in liver metastasis. Carcinogenesis 2018, 39, 1368–1379. [Google Scholar] [CrossRef]
- Dai, G.; Yao, X.; Zhang, Y.; Gu, J.; Geng, Y.; Xue, F.; Zhang, J. Colorectal cancer cell–derived exosomes containing miR-10b regulate fibroblast cells via the PI3K/Akt pathway. Bull. Cancer 2018, 105, 336–349. [Google Scholar] [CrossRef] [PubMed]
- Yoshii, S.; Hayashi, Y.; Iijima, H.; Inoue, T.; Kimura, K.; Sakatani, A.; Nagai, K.; Fujinaga, T.; Hiyama, S.; Kodama, T.; et al. Exosomal microRNAs derived from colon cancer cells promote tumor progression by suppressing fibroblast TP53 expression. Cancer Sci. 2019. [Google Scholar] [CrossRef] [PubMed]
- Webber, J.; Steadman, R.; Mason, M.D.; Tabi, Z.; Clayton, A. Cancer exosomes trigger fibroblast to myofibroblast differentiation. Cancer Res. 2010, 70, 9621–9630. [Google Scholar] [CrossRef]
- Rai, A.; Greening, D.W.; Chen, M.; Xu, R.; Ji, H.; Simpson, R.J. Exosomes Derived from Human Primary and Metastatic Colorectal Cancer Cells Contribute to Functional Heterogeneity of Activated Fibroblasts by Reprogramming Their Proteome. Proteomics 2019, 19. [Google Scholar] [CrossRef] [PubMed]
- Lugini, L.; Valtieri, M.; Federici, C.; Cecchetti, S.; Meschini, S.; Condello, M.; Signore, M.; Fais, S. Exosomes from human colorectal cancer induce a tumor-like behavior in colonic mesenchymal stromal cells. Oncotarget 2016, 7, 50086–50098. [Google Scholar] [CrossRef]
- Huber, V.; Fais, S.; Iero, M.; Lugini, L.; Canese, P.; Squarcina, P.; Zaccheddu, A.; Colone, M.; Arancia, G.; Gentile, M.; et al. Human colorectal cancer cells induce T-cell death through release of proapoptotic microvesicles: Role in immune escape. Gastroenterology 2005, 128, 1796–1804. [Google Scholar] [CrossRef]
- Clayton, A.; Al-Taei, S.; Webber, J.; Mason, M.D.; Tabi, Z. Cancer Exosomes Express CD39 and CD73, Which Suppress T Cells through Adenosine Production. J. Immunol. 2011, 187, 676–683. [Google Scholar] [CrossRef]
- Yamada, N.; Kuranaga, Y.; Kumazaki, M.; Shinohara, H.; Taniguchi, K.; Akao, Y. Colorectal cancer cell-derived extracellular vesicles induce phenotypic alteration of T cells into tumor-growth supporting cells with transforming growth factor-β1-mediated suppression. Oncotarget 2016, 7, 27033–27043. [Google Scholar] [CrossRef]
- Shinohara, H.; Kuranaga, Y.; Kumazaki, M.; Sugito, N.; Yoshikawa, Y.; Takai, T.; Taniguchi, K.; Ito, Y.; Akao, Y. Regulated Polarization of Tumor-Associated Macrophages by miR-145 via Colorectal Cancer–Derived Extracellular Vesicles. J. Immunol. 2017, 199, 1505–1515. [Google Scholar] [CrossRef]
- Menck, K.; Bleckmann, A.; Wachter, A.; Hennies, B.; Ries, L.; Schulz, M.; Balkenhol, M.; Pukrop, T.; Schatlo, B.; Rost, U.; et al. Characterisation of tumour-derived microvesicles in cancer patients’ blood and correlation with clinical outcome. J. Extracell. Vesicles 2017, 6. [Google Scholar] [CrossRef]
- Chen, Z.; Yang, L.; Cui, Y.; Zhou, Y.; Yin, X.; Guo, J.; Zhang, G.; Wang, T.; He, Q.Y. Cytoskeleton-centric protein transportation by exosomes transforms tumor-favorable macrophages. Oncotarget 2016, 7, 67387–67402. [Google Scholar] [CrossRef] [PubMed]
- Gastpar, R.; Gehrmann, M.; Bausero, M.A.; Asea, A.; Gross, C.; Schroeder, J.A.; Multhoff, G. Heat shock protein 70 surface-positive tumor exosomes stimulate migratory and cytolytic activity of natural killer cells. Cancer Res. 2005, 65, 5238–5247. [Google Scholar] [CrossRef] [PubMed]
- Guo, D.; Chen, Y.; Wang, S.; Yu, L.; Shen, Y.; Zhong, H.; Yang, Y. Exosomes from heat-stressed tumour cells inhibit tumour growth by converting regulatory T cells to Th17 cells via IL-6. Immunology 2018, 154, 132–143. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Z.; Li, Y.; Pan, Y.; Lan, X.; Song, F.; Sun, J.; Zhou, K.; Liu, X.; Ren, X.; Wang, F.; et al. Cancer-derived exosomal miR-25-3p promotes pre-metastatic niche formation by inducing vascular permeability and angiogenesis. Nat. Commun. 2018, 9. [Google Scholar] [CrossRef]
- Wang, W.; Yuan, X.; Xu, A.; Zhu, X.; Zhan, Y.; Wang, S.; Liu, M. Human cancer cells suppress behaviors of endothelial progenitor cells through miR-21 targeting IL6R. Microvasc. Res. 2018, 120, 21–28. [Google Scholar] [CrossRef] [PubMed]
- Holzner, S.; Senfter, D.; Stadler, S.; Staribacher, A.; Nguyen, C.H.; Gaggl, A.; Geleff, S.; Huttary, N.; Krieger, S.; Jäger, W.; et al. Colorectal cancer cell-derived microRNA200 modulates the resistance of adjacent blood endothelial barriers in vitro. Oncol. Rep. 2016, 36, 3065–3071. [Google Scholar] [CrossRef]
- Senfter, D.; Holzner, S.; Kalipciyan, M.; Staribacher, A.; Walzl, A.; Huttary, N.; Krieger, S.; Brenner, S.; Jäger, W.; Krupitza, G.; et al. Loss of miR-200 family in 5-fluorouracil resistant colon cancer drives lymphendothelial invasiveness in vitro. Hum. Mol. Genet. 2015, 24, 3689–3698. [Google Scholar] [CrossRef]
- Wang, F.W.; Cao, C.H.; Han, K.; Zhao, Y.X.; Cai, M.Y.; Xiang, Z.C.; Zhang, J.X.; Chen, J.W.; Zhong, L.P.; Huang, Y.; et al. APC-activated long noncoding RNA inhibits colorectal carcinoma pathogenesis through reduction of exosome production. J. Clin. Investig. 2019, 129, 727–743. [Google Scholar] [CrossRef]
- Hong, B.S.; Cho, J.H.; Kim, H.; Choi, E.J.; Rho, S.; Kim, J.; Kim, J.H.; Choi, D.S.; Kim, Y.K.; Hwang, D.; et al. Colorectal cancer cell-derived microvesicles are enriched in cell cycle-related mRNAs that promote proliferation of endothelial cells. BMC Genom. 2009, 10. [Google Scholar] [CrossRef]
- Schillaci, O.; Fontana, S.; Monteleone, F.; Taverna, S.; Di Bella, M.A.; Di Vizio, D.; Alessandro, R. Exosomes from metastatic cancer cells transfer amoeboid phenotype to non-metastatic cells and increase endothelial permeability: Their emerging role in tumor heterogeneity. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Huang, Z.; Feng, Y. Exosomes derived from hypoxic colorectal cancer cells promote angiogenesis through Wnt4-Induced β-catenin signaling in endothelial cells. Oncol. Res. 2017, 25, 651–661. [Google Scholar] [CrossRef] [PubMed]
- Yoon, Y.J.; Kim, D.K.; Yoon, C.M.; Park, J.; Kim, Y.K.; Roh, T.Y.; Gho, Y.S. Egr-1 activation by cancer-derived extracellular vesicles promotes endothelial cell migration via ERK1/2 and JNK signaling pathways. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Sun, B.; Zhou, Y.; Fang, Y.; Li, Z.; Gu, X.; Xiang, J. Colorectal cancer exosomes induce lymphatic network remodeling in lymph nodes. Int. J. Cancer 2019. [Google Scholar] [CrossRef] [PubMed]
- Peinado, H.; Alec, M.; Lavotshkin, S.; Matei, I.; Costa-silva, B.; Moreno-bueno, G.; Hergueta-redondo, M.; Williams, C.; García-santos, G.; Ghajar, C.M.; et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat. Med. 2012, 18, 883–891. [Google Scholar] [CrossRef] [PubMed]
- Sceneay, J.; Smyth, M.J.; Möller, A. The pre-metastatic niche: Finding common ground. Cancer Metastasis Rev. 2013, 32, 449–464. [Google Scholar] [CrossRef]
- Guo, Y.; Ji, X.; Liu, J.; Fan, D.; Zhou, Q.; Chen, C.; Wang, W.; Wang, G.; Wang, H.; Yuan, W.; et al. Effects of exosomes on pre-metastatic niche formation in tumors. Mol. Cancer 2019, 18. [Google Scholar] [CrossRef]
- Baroni, S.; Romero-Cordoba, S.; Plantamura, I.; Dugo, M.; D’Ippolito, E.; Cataldo, A.; Cosentino, G.; Angeloni, V.; Rossini, A.; Daidone, M.G.; et al. Exosome-mediated delivery of miR-9 induces cancer-Associated fibroblast-like properties in human breast fibroblasts. Cell Death Dis. 2016, 7. [Google Scholar] [CrossRef]
- Yan, W.; Wu, X.; Zhou, W.; Fong, M.Y.; Cao, M.; Liu, J.; Liu, X.; Chen, C.H.; Fadare, O.; Pizzo, D.P.; et al. Cancer-cell-secreted exosomal miR-105 promotes tumour growth through the MYC-dependent metabolic reprogramming of stromal cells. Nat. Cell Biol. 2018, 20, 597–609. [Google Scholar] [CrossRef]
- Zhou, X.; Yan, T.; Huang, C.; Xu, Z.; Wang, L.; Jiang, E.; Wang, H.; Chen, Y.; Liu, K.; Shao, Z.; et al. Melanoma cell-secreted exosomal miR-155-5p induce proangiogenic switch of cancer-associated fibroblasts via SOCS1/JAK2/STAT3 signaling pathway 11 Medical and Health Sciences 1112 Oncology and Carcinogenesis 06 Biological Sciences 0601 Biochemistry and Cell Biology. J. Exp. Clin. Cancer Res. 2018, 37, 242. [Google Scholar]
- García-Silva, S.; Peinado, H. Melanosomes foster a tumour niche by activating CAFs. Nat. Cell Biol. 2016, 18, 911–913. [Google Scholar] [CrossRef]
- Zhou, Y.; Ren, H.; Dai, B.; Li, J.; Shang, L.; Huang, J.; Shi, X. Hepatocellular carcinoma-derived exosomal miRNA-21 contributes to tumor progression by converting hepatocyte stellate cells to cancer-associated fibroblasts. J. Exp. Clin. Cancer Res. 2018, 37. [Google Scholar] [CrossRef]
- Wang, J.; Guan, X.; Zhang, Y.; Ge, S.; Zhang, L.; Li, H.; Wang, X.; Liu, R.; Ning, T.; Deng, T.; et al. Exosomal miR-27a derived from gastric cancer cells regulates the transformation of fibroblasts into cancer-associated fibroblasts. Cell. Physiol. Biochem. 2018, 49, 869–883. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Cai, H.; Chen, S.; Sun, D.; Zhang, D.; He, Y. Exosomal transfer of miR-124 inhibits normal fibroblasts to cancer-associated fibroblasts transition by targeting sphingosine kinase 1 in ovarian cancer. J. Cell. Biochem. 2019, 120, 13187–13201. [Google Scholar] [CrossRef] [PubMed]
- Li, C.H.; Chen, Y. Insight Into the Role of Long Noncoding RNA in Cancer Development and Progression. Int. Rev. Cell Mol. Biol. 2016, 326, 33–65. [Google Scholar] [PubMed]
- Ding, L.; Ren, J.; Zhang, D.; Li, Y.; Huang, X.; Hu, Q.; Wang, H.; Song, Y.; Ni, Y.; Hou, Y. A novel stromal lncRNA signature reprograms fibroblasts to promote the growth of oral squamous cell carcinoma via LncRNA-CAF/interleukin-33. Carcinogenesis 2018, 39, 397–406. [Google Scholar] [CrossRef]
- Ringuette Goulet, C.; Bernard, G.; Tremblay, S.; Chabaud, S.; Bolduc, S.; Pouliot, F. Exosomes Induce Fibroblast Differentiation into Cancer-Associated Fibroblasts through TGFβ Signaling. Mol. Cancer Res. 2018, 16, 1196–1204. [Google Scholar] [CrossRef]
- Ramteke, A.; Ting, H.; Agarwal, C.; Mateen, S.; Somasagara, R.; Hussain, A.; Graner, M.; Frederick, B.; Agarwal, R.; Deep, G. Exosomes secreted under hypoxia enhance invasiveness and stemness of prostate cancer cells by targeting adherens junction molecules. Mol. Carcinog. 2015, 54, 554–565. [Google Scholar] [CrossRef]
- Timaner, M.; Tsai, K.K.; Shaked, Y. The multifaceted role of mesenchymal stem cells in cancer. Semin. Cancer Biol. 2019. [Google Scholar] [CrossRef]
- Whiteside, T.L. Exosome and mesenchymal stem cell cross-talk in the tumor microenvironment. Semin. Immunol. 2018, 35, 69–79. [Google Scholar] [CrossRef]
- Lindoso, R.S.; Collino, F.; Camussi, G. Extracellular vesicles derived from renal cancer stem cells induce a pro-tumorigenic phenotype in mesenchymal stromal cells. Oncotarget 2015, 6, 7959–7969. [Google Scholar] [CrossRef]
- Li, X.; Wang, S.; Zhu, R.; Li, H.; Han, Q.; Zhao, R.C. Lung tumor exosomes induce a pro-inflammatory phenotype in mesenchymal stem cells via NFκB-TLR signaling pathway. J. Hematol. Oncol. 2016, 9. [Google Scholar] [CrossRef] [PubMed]
- Léveillé, N.; Baglio, S.R. Exosome-transferred lncRNAs at the core of cancer bone lesions. Crit. Rev. Oncol. Hematol. 2019, 139, 125–127. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Xu, H.; Han, H.; Song, S.; Zhang, X.; Ouyang, L.; Qian, C.; Hong, Y.; Qiu, Y.; Zhou, W.; et al. Exosome-mediated transfer of lncRUNX2-AS1 from multiple myeloma cells to MSCs contributes to osteogenesis. Oncogene 2018, 37, 5508–5519. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, K.; Ochi, H.; Sunamura, S.; Kosaka, N.; Mabuchi, Y.; Fukuda, T.; Yao, K.; Kanda, H.; Ae, K.; Okawa, A.; et al. Cancer-secreted hsa-miR-940 induces an osteoblastic phenotype in the bone metastatic microenvironment via targeting ARHGAP1 and FAM134A. Proc. Natl. Acad. Sci. USA 2018, 115, 2204–2209. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Li, X.; Xu, M.; Wang, J.; Zhao, R.C. Reduced adipogenesis after lung tumor exosomes priming in human mesenchymal stem cells via TGFβ signaling pathway. Mol. Cell. Biochem. 2017, 435, 59–66. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Li, X.; Zhu, R.; Han, Q.; Zhao, R.C. Lung cancer exosomes initiate global long non-coding RNA changes in mesenchymal stem cells. Int. J. Oncol. 2016, 48, 681–689. [Google Scholar] [CrossRef]
- Cho, J.A.; Park, H.; Lim, E.H.; Lee, K.W. Exosomes from breast cancer cells can convert adipose tissue-derived mesenchymal stem cells into myofibroblast-like cells. Int. J. Oncol. 2012, 40, 130–138. [Google Scholar]
- Cho, J.A.; Park, H.; Lim, E.H.; Kim, K.H.; Choi, J.S.; Lee, J.H.; Shin, J.W.; Lee, K.W. Exosomes from ovarian cancer cells induce adipose tissue-derived mesenchymal stem cells to acquire the physical and functional characteristics of tumor-supporting myofibroblasts. Gynecol. Oncol. 2011, 123, 379–386. [Google Scholar] [CrossRef]
- Gu, J.; Qian, H.; Shen, L.; Zhang, X.; Zhu, W.; Huang, L.; Yan, Y.; Mao, F.; Zhao, C.; Shi, Y.; et al. Gastric Cancer Exosomes Trigger Differentiation of Umbilical Cord Derived Mesenchymal Stem Cells to Carcinoma-Associated Fibroblasts through TGF-β/Smad Pathway. PLoS ONE 2012, 7. [Google Scholar] [CrossRef]
- Chowdhury, R.; Webber, J.P.; Gurney, M.; Mason, M.D.; Tabi, Z.; Clayton, A. Cancer exosomes trigger mesenchymal stem cell differentiation into pro-angiogenic and pro-invasive myofibroblasts. Oncotarget 2015, 6, 715–731. [Google Scholar] [CrossRef]
- Han, L.; Lam, E.W.F.; Sun, Y. Extracellular vesicles in the tumor microenvironment: Old stories, but new tales. Mol. Cancer 2019, 18. [Google Scholar] [CrossRef] [PubMed]
- Laviron, M.; Boissonnas, A. Ontogeny of Tumor-Associated Macrophages. Front. Immunol. 2019, 10, 1799. [Google Scholar] [CrossRef] [PubMed]
- Knochelmann, H.M.; Dwyer, C.J.; Bailey, S.R.; Amaya, S.M.; Elston, D.M.; Mazza-McCrann, J.M.; Paulos, C.M. When worlds collide: Th17 and Treg cells in cancer and autoimmunity. Cell. Mol. Immunol. 2018, 15, 458–469. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, N.; Whiteside, T.L. Potential roles of tumor-derived exosomes in angiogenesis. Expert Opin. Ther. Targets 2018, 22, 409–417. [Google Scholar] [CrossRef] [PubMed]
- Conigliaro, A.; Costa, V.; Lo Dico, A.; Saieva, L.; Buccheri, S.; Dieli, F.; Manno, M.; Raccosta, S.; Mancone, C.; Tripodi, M.; et al. CD90+ liver cancer cells modulate endothelial cell phenotype through the release of exosomes containing H19 lncRNA. Mol. Cancer 2015, 14. [Google Scholar] [CrossRef]
- Ma, X.; Li, Z.; Li, T.; Zhu, L.; Li, Z.; Tian, N. Long non-coding RNA HOTAIR enhances angiogenesis by induction of VEGFA expression in glioma cells and transmission to endothelial cells via glioma cell derived-extracellular vesicles. Am. J. Transl. Res. 2017, 9, 5012–5021. [Google Scholar]
- Qu, L.; Ding, J.; Chen, C.; Wu, Z.-J.; Liu, B.; Gao, Y.; Chen, W.; Liu, F.; Sun, W.; Li, X.-F.; et al. Exosome-Transmitted lncARSR Promotes Sunitinib Resistance in Renal Cancer by Acting as a Competing Endogenous RNA. Cancer Cell 2016, 29, 653–668. [Google Scholar] [CrossRef]
- Yeon, J.H.; Jeong, H.E.; Seo, H.; Cho, S.; Kim, K.; Na, D.; Chung, S.; Park, J.; Choi, N.; Kang, J.Y. Cancer-derived exosomes trigger endothelial to mesenchymal transition followed by the induction of cancer-associated fibroblasts. Acta Biomater. 2018, 76, 146–153. [Google Scholar] [CrossRef]
- Ning, X.; Zhang, H.; Wang, C.; Song, X. Exosomes released by gastric cancer cells induce transition of pericytes into cancer- associated fibroblasts. Med. Sci. Monit. 2018, 24, 2350–2359. [Google Scholar] [CrossRef]
- Zhou, C.F.; Ma, J.; Huang, L.; Yi, H.Y.; Zhang, Y.M.; Wu, X.G.; Yan, R.M.; Liang, L.; Zhong, M.; Yu, Y.H.; et al. Cervical squamous cell carcinoma-secreted exosomal miR-221-3p promotes lymphangiogenesis and lymphatic metastasis by targeting VASH1. Oncogene 2019, 38, 1256–1268. [Google Scholar] [CrossRef]
- Li, M.; Lu, Y.; Xu, Y.; Wang, J.; Zhang, C.; Du, Y.; Wang, L.; Li, L.; Wang, B.; Shen, J.; et al. Horizontal transfer of exosomal CXCR4 promotes murine hepatocarcinoma cell migration, invasion and lymphangiogenesis. Gene 2018, 676, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Carrasco-Ramírez, P.; Greening, D.W.; Andrés, G.; Gopal, S.K.; Martín-Villar, E.; Renart, J.; Simpson, R.J.; Quintanilla, M. Podoplanin is a component of extracellular vesicles that reprograms cell-derived exosomal proteins and modulates lymphatic vessel formation. Oncotarget 2016, 7, 16070–16089. [Google Scholar] [CrossRef] [PubMed]
- Deng, X.; Ruan, H.; Zhang, X.; Xu, X.; Zhu, Y.; Peng, H.; Zhang, X.; Kong, F.; Guan, M. Long noncoding RNA CCAL transferred from fibroblasts by exosomes promotes chemoresistance of colorectal cancer cells. Int. J. Cancer 2019. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.; Ding, L.; Zhang, D.; Shi, G.; Xu, Q.; Shen, S.; Wang, Y.; Wang, T.; Hou, Y. Carcinoma-associated fibroblasts promote the stemness and chemoresistance of colorectal cancer by transferring exosomal lncRNA H19. Theranostics 2018, 8, 3932–3948. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.L.; Wang, W.; Lan, X.L.; Zeng, Z.C.; Liang, Y.S.; Yan, Y.R.; Song, F.Y.; Wang, F.F.; Zhu, X.H.; Liao, W.J.; et al. CAFs secreted exosomes promote metastasis and chemotherapy resistance by enhancing cell stemness and epithelial-mesenchymal transition in colorectal cancer. Mol. Cancer 2019, 18. [Google Scholar] [CrossRef] [PubMed]
- Bhome, R.; Goh, R.W.; Bullock, M.D.; Pillar, N.; Thirdborough, S.M.; Mellone, M.; Mirnezami, R.; Galea, D.; Veselkov, K.; Gu, Q.; et al. Exosomal microRNAs derived from colorectal cancer-associated fibroblasts: Role in driving cancer progression. Aging (Albany. NY) 2017, 9, 2666–2694. [Google Scholar] [CrossRef]
- Li, H.; Li, F. Exosomes from BM-MSCs increase the population of CSCs via transfer of miR-142-3p. Br. J. Cancer 2018, 119, 744–755. [Google Scholar] [CrossRef]
- Lan, J.; Sun, L.; Xu, F.; Liu, L.; Hu, F.; Song, D.; Hou, Z.; Wu, W.; Luo, X.; Wang, J.; et al. M2 Macrophage-Derived Exosomes Promote Cell Migration and Invasion in Colon Cancer. Cancer Res. 2019, 79, 146–158. [Google Scholar] [CrossRef]
- Okoye, I.S.; Coomes, S.M.; Pelly, V.S.; Czieso, S.; Papayannopoulos, V.; Tolmachova, T.; Seabra, M.C.; Wilson, M.S. MicroRNA-containing T-regulatory-cell-derived exosomes suppress pathogenic T helper 1 cells. Immunity 2014, 41, 89–103. [Google Scholar] [CrossRef]
- Luga, V.; Wrana, J.L. Tumor-stroma interaction: Revealing fibroblast-secreted exosomes as potent regulators of Wnt-planar cell polarity signaling in cancer metastasis. Cancer Res. 2013, 73, 6843–6847. [Google Scholar] [CrossRef]
- Fu, H.; Yang, H.; Zhang, X.; Xu, W. The emerging roles of exosomes in tumor–stroma interaction. J. Cancer Res. Clin. Oncol. 2016, 142, 1897–1907. [Google Scholar] [CrossRef] [PubMed]
- Dylla, S.J.; Beviglia, L.; Park, I.K.; Chartier, C.; Raval, J.; Ngan, L.; Pickell, K.; Aguilar, J.; Lazetic, S.; Smith-Berdan, S.; et al. Colorectal cancer stem cells are enriched in xenogeneic tumors following chemotherapy. PLoS ONE 2008, 3. [Google Scholar] [CrossRef]
- Fang, Y.; Zhou, W.; Rong, Y.; Kuang, T.; Xu, X.; Wu, W.; Wang, D.; Lou, W. Exosomal miRNA-106b from cancer-associated fibroblast promotes gemcitabine resistance in pancreatic cancer. Exp. Cell Res. 2019, 383, 111543. [Google Scholar] [CrossRef] [PubMed]
- Qin, X.; Guo, H.; Wang, X.; Zhu, X.; Yan, M.; Wang, X.; Xu, Q.; Shi, J.; Lu, E.; Chen, W.; et al. Exosomal miR-196a derived from cancer-associated fibroblasts confers cisplatin resistance in head and neck cancer through targeting CDKN1B and ING5. Genome Biol. 2019, 20, 12. [Google Scholar] [CrossRef]
- Boelens, M.C.; Wu, T.J.; Nabet, B.Y.; Xu, B.; Qiu, Y.; Yoon, T.; Azzam, D.J.; Twyman-Saint Victor, C.; Wiemann, B.Z.; Ishwaran, H.; et al. Exosome transfer from stromal to breast cancer cells regulates therapy resistance pathways. Cell 2014, 159, 499–513. [Google Scholar] [CrossRef]
- Yeung, C.L.A.; Co, N.N.; Tsuruga, T.; Yeung, T.L.; Kwan, S.Y.; Leung, C.S.; Li, Y.; Lu, E.S.; Kwan, K.; Wong, K.K. Exosomal transfer of stroma-derived miR21 confers paclitaxel resistance in ovarian cancer cells through targeting APAF1. Nat. Commun. 2016, 7, 11150. [Google Scholar] [CrossRef]
- Wang, X.; Qin, X.; Yan, M.; Shi, J.; Xu, Q.; Li, Z.; Yang, W.; Zhang, J.; Chen, W. Loss of exosomal miR-3188 in cancer-associated fibroblasts contributes to HNC progression. J. Exp. Clin. Cancer Res. 2019, 38. [Google Scholar] [CrossRef]
- Richards, K.E.; Zeleniak, A.E.; Fishel, M.L.; Wu, J.; Littlepage, L.E.; Hill, R. Cancer-associated fibroblast exosomes regulate survival and proliferation of pancreatic cancer cells. Oncogene 2016. [Google Scholar] [CrossRef]
- Sun, L.; Xu, K.; Cui, J.; Yuan, D.; Zou, B.; Li, J.; Liu, J.; Li, K.; Meng, Z.; Zhang, B. Cancer-associated fibroblast-derived exosomal miR-382-5p promotes the migration and invasion of oral squamous cell carcinoma. Oncol. Rep. 2019. [Google Scholar] [CrossRef]
- WANG, J.-W.; WU, X.-F.; GU, X.-J.; JIANG, X.-H. Exosomal miR-1228 from cancer-associated fibroblasts promotes cell migration and invasion of osteosarcoma by directly targeting SCAI. Oncol. Res. Featur. Preclin. Clin. Cancer Ther. 2018. [Google Scholar] [CrossRef]
- Li, B.L.; Lu, W.; Qu, J.J.; Ye, L.; Du, G.Q.; Wan, X.P. Loss of exosomal miR-148b from cancer-associated fibroblasts promotes endometrial cancer cell invasion and cancer metastasis. J. Cell. Physiol. 2019, 234, 2943–2953. [Google Scholar] [CrossRef] [PubMed]
- Languino, L.R.; Singh, A.; Prisco, M.; Inman, G.J.; Luginbuhl, A.; Curry, J.M.; South, A.P. Exosome-mediated transfer from the tumor microenvironment increases TGFβ signaling in squamous cell carcinoma. Am. J. Transl. Res. 2016, 8, 2432–2437. [Google Scholar] [PubMed]
- Chen, Y.; Zeng, C.; Zhan, Y.; Wang, H.; Jiang, X.; Li, W. Aberrant low expression of p85α in stromal fibroblasts promotes breast cancer cell metastasis through exosome-mediated paracrine Wnt10b. Oncogene 2017. [Google Scholar] [CrossRef] [PubMed]
- Donnarumma, E.; Fiore, D.; Nappa, M.; Roscigno, G.; Adamo, A.; Iaboni, M.; Russo, V.; Affinito, A.; Puoti, I.; Quintavalle, C.; et al. Cancer-associated fibroblasts release exosomal microRNAs that dictate an aggressive phenotype in breast cancer. Oncotarget 2017, 8, 19592–19608. [Google Scholar] [CrossRef] [PubMed]
- You, J.; Li, M.; Cao, L.M.; Gu, Q.H.; Deng, P.B.; Tan, Y.; Hu, C.P. Snail1-dependent cancer-associated fibroblasts induce epithelial-mesenchymal transition in lung cancer cells via exosomes. QJM 2019, 112, 581–590. [Google Scholar] [CrossRef]
- Li, Y.-Y.; Tao, Y.-W.; Gao, S.; Li, P.; Zheng, J.-M.; Zhang, S.-E.; Liang, J.; Zhang, Y. Cancer-associated fibroblasts contribute to oral cancer cells proliferation and metastasis via exosome-mediated paracrine miR-34a-5p. EBioMedicine 2018, 36, 209–220. [Google Scholar] [CrossRef]
- Zhang, Z.; Li, X.; Sun, W.; Yue, S.; Yang, J.; Li, J.; Ma, B.; Wang, J.; Yang, X.; Pu, M.; et al. Loss of exosomal miR-320a from cancer-associated fibroblasts contributes to HCC proliferation and metastasis. Cancer Lett. 2017, 397, 33–42. [Google Scholar] [CrossRef]
- Zhao, H.; Yang, L.; Baddour, J.; Achreja, A.; Bernard, V.; Moss, T.; Marini, J.C.; Tudawe, T.; Seviour, E.G.; San Lucas, F.A.; et al. Tumor microenvironment derived exosomes pleiotropically modulate cancer cell metabolism. Elife 2016, 5, e10250. [Google Scholar] [CrossRef]
- Achreja, A.; Zhao, H.; Yang, L.; Yun, T.H.; Marini, J.; Nagrath, D. Exo-MFA—A 13C metabolic flux analysis framework to dissect tumor microenvironment-secreted exosome contributions towards cancer cell metabolism. Metab. Eng. 2017, 43, 156–172. [Google Scholar] [CrossRef]
- Shojaei, S.; Hashemi, S.M.; Ghanbarian, H.; Salehi, M.; Mohammadi-Yeganeh, S. Effect of mesenchymal stem cells-derived exosomes on tumor microenvironment: Tumor progression versus tumor suppression. J. Cell. Physiol. 2019, 234, 3394–3409. [Google Scholar] [CrossRef]
- Vakhshiteh, F.; Atyabi, F.; Ostad, S.N. Mesenchymal stem cell exosomes: A two-edged sword in cancer therapy. Int. J. Nanomed. 2019, 14, 2847–2859. [Google Scholar] [CrossRef] [PubMed]
- Qi, J.; Zhou, Y.; Jiao, Z.; Wang, X.; Zhao, Y.; Li, Y.; Chen, H.; Yang, L.; Zhu, H.; Li, Y. Exosomes Derived from Human Bone Marrow Mesenchymal Stem Cells Promote Tumor Growth Through Hedgehog Signaling Pathway. Cell. Physiol. Biochem. 2017, 42, 2242–2254. [Google Scholar] [CrossRef] [PubMed]
- Gu, H.; Ji, R.; Zhang, X.; Wang, M.; Zhu, W.; Qian, H.; Chen, Y.; Jiang, P.; Xu, W. Exosomes derived from human mesenchymal stem cells promote gastric cancer cell growth and migration via the activation of the Akt pathway. Mol. Med. Rep. 2016, 14, 3452–3458. [Google Scholar] [CrossRef] [PubMed]
- Mao, J.; Liang, Z.; Zhang, B.; Yang, H.; Li, X.; Fu, H.; Zhang, X.; Yan, Y.; Xu, W.; Qian, H. UBR2 Enriched in p53 Deficient Mouse Bone Marrow Mesenchymal Stem Cell-Exosome Promoted Gastric Cancer Progression via Wnt/β-Catenin Pathway. Stem Cells 2017, 35, 2267–2279. [Google Scholar] [CrossRef] [PubMed]
- Ji, R.; Zhang, B.; Zhang, X.; Xue, J.; Yuan, X.; Yan, Y.; Wang, M.; Zhu, W.; Qian, H.; Xu, W. Exosomes derived from human mesenchymal stem cells confer drug resistance in gastric cancer. Cell Cycle 2015, 14, 2473–2483. [Google Scholar] [CrossRef] [PubMed]
- Roccaro, A.M.; Sacco, A.; Maiso, P.; Azab, A.K.; Tai, Y.T.; Reagan, M.; Azab, F.; Flores, L.M.; Campigotto, F.; Weller, E.; et al. BM mesenchymal stromal cell-derived exosomes facilitate multiple myeloma progression. J. Clin. Investig. 2013, 123, 1542–1555. [Google Scholar] [CrossRef]
- Shi, S.; Zhang, Q.; Xia, Y.; You, B.; Shan, Y.; Bao, L.; Li, L.; You, Y.; Gu, Z. Mesenchymal stem cell-derived exosomes facilitate nasopharyngeal carcinoma progression. Am. J. Cancer Res. 2016, 6, 459–472. [Google Scholar]
- Li, W.; Han, Y.; Zhao, Z.; Ji, X.; Wang, X.; Jin, J.; Wang, Q.; Guo, X.; Cheng, Z.; Lu, M.; et al. Oral mucosal mesenchymal stem cell-derived exosomes: A potential therapeutic target in oral premalignant lesions. Int. J. Oncol. 2019, 54, 1567–1578. [Google Scholar] [CrossRef]
- Bliss, S.A.; Sinha, G.; Sandiford, O.A.; Williams, L.M.; Engelberth, D.J.; Guiro, K.; Isenalumhe, L.L.; Greco, S.J.; Ayer, S.; Bryan, M.; et al. Mesenchymal Stem Cell-Derived Exosomes Stimulate Cycling Quiescence and Early Breast Cancer Dormancy in Bone Marrow. Cancer Res. 2016, 76, 5832–5844. [Google Scholar] [CrossRef]
- Ono, M.; Kosaka, N.; Tominaga, N.; Yoshioka, Y.; Takeshita, F.; Takahashi, R.; Yoshida, M.; Tsuda, H.; Tamura, K.; Ochiya, T. Exosomes from bone marrow mesenchymal stem cells contain a microRNA that promotes dormancy in metastatic breast cancer cells. Sci. Signal. 2014, 7, ra63. [Google Scholar] [CrossRef]
- Lin, R.; Wang, S.; Zhao, R.C. Exosomes from human adipose-derived mesenchymal stem cells promote migration through Wnt signaling pathway in a breast cancer cell model. Mol. Cell. Biochem. 2013, 383, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Alzahrani, F.A.; El-Magd, M.A.; Abdelfattah-Hassan, A.; Saleh, A.A.; Saadeldin, I.M.; El-Shetry, E.S.; Badawy, A.A.; Alkarim, S. Potential Effect of Exosomes Derived from Cancer Stem Cells and MSCs on Progression of DEN-Induced HCC in Rats. Stem Cells Int. 2018, 2018, 8058979. [Google Scholar] [CrossRef] [PubMed]
- Reza, A.M.M.T.; Choi, Y.J.; Yasuda, H.; Kim, J.H. Human adipose mesenchymal stem cell-derived exosomal-miRNAs are critical factors for inducing anti-proliferation signalling to A2780 and SKOV-3 ovarian cancer cells. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Pakravan, K.; Babashah, S.; Sadeghizadeh, M.; Mowla, S.J.; Mossahebi-Mohammadi, M.; Ataei, F.; Dana, N.; Javan, M. MicroRNA-100 shuttled by mesenchymal stem cell-derived exosomes suppresses in vitro angiogenesis through modulating the mTOR/HIF-1α/VEGF signaling axis in breast cancer cells. Cell. Oncol. 2017, 40, 457–470. [Google Scholar] [CrossRef]
- Lee, J.K.; Park, S.R.; Jung, B.K.; Jeon, Y.K.; Lee, Y.S.; Kim, M.K.; Kim, Y.G.; Jang, J.Y.; Kim, C.W. Exosomes derived from mesenchymal stem cells suppress angiogenesis by down-regulating VEGF expression in breast cancer cells. PLoS ONE 2013, 8. [Google Scholar] [CrossRef]
- Lou, G.; Song, X.; Yang, F.; Wu, S.; Wang, J.; Chen, Z.; Liu, Y. Exosomes derived from miR-122-modified adipose tissue-derived MSCs increase chemosensitivity of hepatocellular carcinoma. J. Hematol. Oncol. 2015, 8, 122. [Google Scholar] [CrossRef]
- Liu, Y.; Song, B.; Wei, Y.; Chen, F.; Chi, Y.; Fan, H.; Liu, N.; Li, Z.; Han, Z.; Ma, F. Exosomes from mesenchymal stromal cells enhance imatinib-induced apoptosis in human leukemia cells via activation of caspase signaling pathway. Cytotherapy 2018, 20, 181–188. [Google Scholar] [CrossRef]
- de Araujo Farias, V.; O’Valle, F.; Serrano-Saenz, S.; Anderson, P.; Andrés, E.; López-Peñalver, J.; Tovar, I.; Nieto, A.; Santos, A.; Martín, F.; et al. Exosomes derived from mesenchymal stem cells enhance radiotherapy-induced cell death in tumor and metastatic tumor foci. Mol. Cancer 2018, 17. [Google Scholar] [CrossRef]
- van Balkom, B.W.M.; de Jong, O.G.; Smits, M.; Brummelman, J.; den Ouden, K.; de Bree, P.M.; van Eijndhoven, M.A.J.; Pegtel, D.M.; Stoorvogel, W.; Würdinger, T.; et al. Endothelial cells require miR-214 to secrete exosomes that suppress senescence and induce angiogenesis in human and mouse endothelial cells. Blood 2013, 121, 3997–4006. [Google Scholar] [CrossRef]
- Bovy, N.; Blomme, B.; Frères, P.; Dederen, S.; Nivelles, O.; Lion, M.; Carnet, O.; Martial, J.A.; Noël, A.; Thiry, M.; et al. Endothelial exosomes contribute to the antitumor response during breast cancer neoadjuvant chemotherapy via microRNA transfer. Oncotarget 2015, 6, 10253–10266. [Google Scholar] [CrossRef]
- Kim, K.-S.; Park, J.-I.; Oh, N.; Cho, H.-J.; Park, J.-H.; Park, K.-S. ELK3 expressed in lymphatic endothelial cells promotes breast cancer progression and metastasis through exosomal miRNAs. Sci. Rep. 2019, 9, 8418. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Hu, C.; Chao, H.; Zhang, Y.; Li, Y.; Hou, J.; Xu, Z.; Lu, H.; Li, H.; Chen, H. Drug-resistant endothelial cells facilitate progression, EMT and chemoresistance in nasopharyngeal carcinoma via exosomes. Cell. Signal. 2019, 63, 109385. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Y.; Yao, X.; Liu, X.; He, X.; Li, L.; Liu, X.; Yan, Z.; Wu, J.; Fu, B.M. Anti-angiogenesis triggers exosomes release from endothelial cells to promote tumor vasculogenesis. J. Extracell. Vesicles 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.-H.; Miao, Z.-W.; Jiang, Q.-Z.; Gan, D.-X.; Wei, X.-G.; Xue, X.-Z.; Li, J.-Q.; Zheng, F.; Qin, X.-X.; Fang, W.-G.; et al. Brain microvascular endothelial cell exosome-mediated S100A16 up-regulation confers small-cell lung cancer cell survival in brain. FASEB J. 2019, 33, 1742–1757. [Google Scholar] [CrossRef] [PubMed]
- Shen, M.; Ren, X. New insights into the biological impacts of immune cell-derived exosomes within the tumor environment. Cancer Lett. 2018, 431, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Zheng, P.; Chen, L.; Yuan, X.; Luo, Q.; Liu, Y.; Xie, G.; Ma, Y.; Shen, L. Exosomal transfer of tumor-associated macrophage-derived miR-21 confers cisplatin resistance in gastric cancer cells. J. Exp. Clin. Cancer Res. 2017, 36. [Google Scholar] [CrossRef]
- Wang, J.-J.; Wang, Z.-Y.; Chen, R.; Xiong, J.; Yao, Y.-L.; Wu, J.-H.; Li, G.-X. Macrophage-secreted Exosomes Delivering miRNA-21 Inhibitor can Regulate BGC-823 Cell Proliferation. Asian Pac. J. Cancer Prev. 2015, 16, 4203–4209. [Google Scholar] [CrossRef]
- Yu, X.; Zhang, Q.; Zhang, X.; Han, Q.; Li, H.; Mao, Y.; Wang, X.; Guo, H.; Irwin, D.M.; Niu, G.; et al. Exosomes from Macrophages Exposed to Apoptotic Breast Cancer Cells Promote Breast Cancer Proliferation and Metastasis. J. Cancer 2019, 10, 2892–2906. [Google Scholar] [CrossRef]
- Yin, Z.; Ma, T.; Huang, B.; Lin, L.; Zhou, Y.; Yan, J.; Zou, Y.; Chen, S. Macrophage-derived exosomal microRNA-501-3p promotes progression of pancreatic ductal adenocarcinoma through the TGFBR3-mediated TGF-β signaling pathway. J. Exp. Clin. Cancer Res. 2019, 38. [Google Scholar] [CrossRef]
- Wu, Q.; Wu, X.; Ying, X.; Zhu, Q.; Wang, X.; Jiang, L.; Chen, X.; Wu, Y.; Wang, X. Suppression of endothelial cell migration by tumor associated macrophage-derived exosomes is reversed by epithelial ovarian cancer exosomal lncRNA. Cancer Cell Int. 2017, 17. [Google Scholar] [CrossRef]
- Zhou, J.; Li, X.; Wu, X.; Zhang, T.; Zhu, Q.; Wang, X.; Wang, H.; Wang, K.; Lin, Y.; Wang, X. Exosomes released from tumor-associated macrophages transfer miRNAs that induce a Treg/Th17 cell imbalance in epithelial ovarian cancer. Cancer Immunol. Res. 2018, 6, 1578–1592. [Google Scholar] [CrossRef] [PubMed]
- Binenbaum, Y.; Fridman, E.; Yaari, Z.; Milman, N.; Schroeder, A.; Ben David, G.; Shlomi, T.; Gil, Z. Transfer of miRNA in macrophage-derived exosomes induces drug resistance in pancreatic adenocarcinoma. Cancer Res. 2018, 78, 5287–5299. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Shen, H.; Yin, X.; Yang, M.; Wei, H.; Chen, Q.; Feng, F.; Liu, Y.; Xu, W.; Li, Y. Macrophages derived exosomes deliver miR-223 to epithelial ovarian cancer cells to elicit a chemoresistant phenotype. J. Exp. Clin. Cancer Res. 2019, 38. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Kalimuthu, S.; Gangadaran, P.; Oh, J.M.; Lee, H.W.; Baek, S.H.; Jeong, S.Y.; Lee, S.-W.; Lee, J.; Ahn, B.-C. Exosomes Derived From Natural Killer Cells Exert Therapeutic Effect in Melanoma. Theranostics 2017, 7, 2732–2745. [Google Scholar] [CrossRef] [PubMed]
- Lugini, L.; Cecchetti, S.; Huber, V.; Luciani, F.; Macchia, G.; Spadaro, F.; Paris, L.; Abalsamo, L.; Colone, M.; Molinari, A.; et al. Immune Surveillance Properties of Human NK Cell-Derived Exosomes. J. Immunol. 2012, 189, 2833–2842. [Google Scholar] [CrossRef]
- Lu, J.; Wu, J.; Tian, J.; Wang, S. Role of T cell-derived exosomes in immunoregulation. Immunol. Res. 2018, 66, 313–322. [Google Scholar] [CrossRef]
- Cai, Z.; Yang, F.; Yu, L.; Yu, Z.; Jiang, L.; Wang, Q.; Yang, Y.; Wang, L.; Cao, X.; Wang, J. Activated T Cell Exosomes Promote Tumor Invasion via Fas Signaling Pathway. J. Immunol. 2012, 188, 5954–5961. [Google Scholar] [CrossRef]
- Min, H.; Sun, X.; Yang, X.; Zhu, H.; Liu, J.; Wang, Y.; Chen, G.; Sun, X. Exosomes Derived from Irradiated Esophageal Carcinoma-Infiltrating T Cells Promote Metastasis by Inducing the Epithelial–Mesenchymal Transition in Esophageal Cancer Cells. Pathol. Oncol. Res. 2018, 24, 11–18. [Google Scholar] [CrossRef]
- Wang, X.; Shen, H.; He, Q.; Tian, W.; Xia, A.; Lu, X.J. Exosomes derived from exhausted CD8 + T cells impaired the anticancer function of normal CD8 + T cells. J. Med. Genet. 2019, 56, 29–31. [Google Scholar] [CrossRef]
- Giussani, M.; Triulzi, T.; Sozzi, G.; Tagliabue, E. Tumor Extracellular Matrix Remodeling: New Perspectives as a Circulating Tool in the Diagnosis and Prognosis of Solid Tumors. Cells 2019, 8, 81. [Google Scholar] [CrossRef]
- Qiao, J.; Fang, C.Y.; Chen, S.X.; Wang, X.Q.; Cui, S.J.; Liu, X.H.; Jiang, Y.H.; Wang, J.; Zhang, Y.; Yang, P.Y.; et al. Stroma derived COL6A3 is a potential prognosis marker of colorectal carcinoma revealed by quantitative proteomics. Oncotarget 2015, 6, 29929–29946. [Google Scholar] [CrossRef]
- Kehlet, S.N.; Sanz-Pamplona, R.; Brix, S.; Leeming, D.J.; Karsdal, M.A.; Moreno, V. Excessive collagen turnover products are released during colorectal cancer progression and elevated in serum from metastatic colorectal cancer patients. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Solé, X.; Crous-Bou, M.; Cordero, D.; Olivares, D.; Guinó, E.; Sanz-Pamplona, R.; Rodriguez-Moranta, F.; Sanjuan, X.; de Oca, J.; Salazar, R.; et al. Discovery and validation of new potential biomarkers for early detection of colon cancer. PLoS ONE 2014, 9, e106748. [Google Scholar] [CrossRef] [PubMed]
- Kantola, T.; Väyrynen, J.P.; Klintrup, K.; Mäkelä, J.; Karppinen, S.M.; Pihlajaniemi, T.; Autio-Harmainen, H.; Karttunen, T.J.; Mäkinen, M.J.; Tuomisto, A. Serum endostatin levels are elevated in colorectal cancer and correlate with invasion and systemic inflammatory markers. Br. J. Cancer 2014, 111, 1605–1613. [Google Scholar] [CrossRef]
- Lee, J.-H.; Choi, J.-W.; Kim, Y.-S. Plasma or serum TIMP-1 is a predictor of survival outcomes in colorectal cancer: A meta-analysis. J. Gastrointestin. Liver Dis. 2011, 20, 287–291. [Google Scholar] [PubMed]
- Manzoni, M.; Comolli, G.; Torchio, M.; Mazzini, G.; Danova, M. Circulating endothelial cells and their subpopulations: Role as predictive biomarkers in antiangiogenic therapy for colorectal cancer. Clin. Colorectal Cancer 2015, 14, 11–17. [Google Scholar] [CrossRef]
- Rahbari, N.N.; Schölch, S.; Bork, U.; Kahlert, C.; Schneider, M.; Rahbari, M.; Büchler, M.W.; Weitz, J.; Reissfelder, C. Prognostic value of circulating endothelial cells in metastatic colorectal cancer. Oncotarget 2017, 8, 37491–37501. [Google Scholar] [CrossRef]
- Ramcharan, K.S.; Lip, G.Y.H.; Stonelake, P.S.; Blann, A.D. Increased pre-surgical numbers of endothelial progenitor cells and circulating endothelial cells in colorectal cancer fail to predict outcome. Int. J. Colorectal Dis. 2015, 30, 315–321. [Google Scholar] [CrossRef]
- Kraan, J.; Van Den Broek, P.; Verhoef, C.; Grunhagen, D.J.; Taal, W.; Gratama, J.W.; Sleijfer, S. Endothelial CD276 (B7-H3) expression is increased in human malignancies and distinguishes between normal and tumour-derived circulating endothelial cells. Br. J. Cancer 2014, 111, 149–156. [Google Scholar] [CrossRef]
- Cima, I.; Kong, S.L.; Sengupta, D.; Tan, I.B.; Phyo, W.M.; Lee, D.; Hu, M.; Iliescu, C.; Alexander, I.; Goh, W.L.; et al. Circulating_Tumor-endo-cell_SciTraMed2016. Sci. Transl. Med. 2016, 8, 345ra89. [Google Scholar] [CrossRef]
- Avallone, A.; Piccirillo, M.C.; Aloj, L.; Nasti, G.; Delrio, P.; Izzo, F.; Di Gennaro, E.; Tatangelo, F.; Granata, V.; Cavalcanti, E.; et al. A randomized phase 3 study on the optimization of the combination of bevacizumab with FOLFOX/OXXEL in the treatment of patients with metastatic colorectal cancer-OBELICS (Optimization of BEvacizumab scheduLIng within Chemotherapy Scheme). BMC Cancer 2016, 16. [Google Scholar] [CrossRef] [PubMed]
- Hagland, H.R.; Lea, D.; Watson, M.M.; Søreide, K. Correlation of blood T-cells to intratumoural density and location of CD3+ and CD8+ T-cells in colorectal cancer. Anticancer Res. 2017, 37, 675–684. [Google Scholar] [CrossRef] [PubMed]
- Yun; Lee; Kim; Chun; Engleman; Kim; Kang A Novel Type of Blood Biomarker: Distinct Changes of Cytokine-Induced STAT Phosphorylation in Blood T Cells Between Colorectal Cancer Patients and Healthy Individuals. Cancers 2019, 11, 1157. [CrossRef] [PubMed]
- Choi, J.; Maeng, H.G.; Lee, S.J.; Kim, Y.J.; Kim, D.W.; Lee, H.N.; Namgung, J.H.; Oh, H.M.; Kim, T.J.; Jeong, J.E.; et al. Diagnostic value of peripheral blood immune profiling in colorectal cancer. Ann. Surg. Treat. Res. 2018, 94, 312–321. [Google Scholar] [CrossRef]
- Mahalingam, J.; Lin, C.Y.; Chiang, J.M.; Su, P.J.; Chu, Y.Y.; Lai, H.Y.; Fang, J.H.; Huang, C.T.; Lin, Y.C. CD4+ T cells expressing latency-associated peptide and Foxp3 are an activated subgroup of regulatory T cells enriched in patients with colorectal cancer. PLoS ONE 2014, 9. [Google Scholar] [CrossRef]
- Xu, B.; Yuan, L.; Gao, Q.; Yuan, P.; Zhao, P.; Yuan, H.; Fan, H.; Li, T.; Qin, P.; Han, L.; et al. Circulating and tumor-infiltrating Tim-3 in patients with colorectal cancer. Oncotarget 2015, 6, 20592–20603. [Google Scholar] [CrossRef]
- Won, E.J.; Ju, J.K.; Cho, Y.N.; Jin, H.M.; Park, K.J.; Kim, T.J.; Kwon, Y.S.; Kee, H.J.; Kim, J.C.; Kee, S.J.; et al. Clinical relevance of circulating mucosal-associated invariant T cell levels and their anti-cancer activity in patients with mucosal-associated cancer. Oncotarget 2016, 7, 76274–76290. [Google Scholar] [CrossRef]
- Nissen, N.I.; Karsdal, M.; Willumsen, N. Collagens and Cancer associated fibroblasts in the reactive stroma and its relation to Cancer biology. J. Exp. Clin. Cancer Res. 2019, 38. [Google Scholar] [CrossRef]
- Principe, S.; Mejia-Guerrero, S.; Ignatchenko, V.; Sinha, A.; Ignatchenko, A.; Shi, W.; Pereira, K.; Su, S.; Huang, S.H.; O’Sullivan, B.; et al. Proteomic Analysis of Cancer-Associated Fibroblasts Reveals a Paracrine Role for MFAP5 in Human Oral Tongue Squamous Cell Carcinoma. J. Proteome Res. 2018, 17, 2045–2059. [Google Scholar] [CrossRef]
- Arroyo, J.D.; Chevillet, J.R.; Kroh, E.M.; Ruf, I.K.; Pritchard, C.C.; Gibson, D.F.; Mitchell, P.S.; Bennett, C.F.; Pogosova-Agadjanyan, E.L.; Stirewalt, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 2011, 108, 5003–5008. [Google Scholar] [CrossRef]
- György, B.; Módos, K.; Pállinger, E.; Pálóczi, K.; Pásztói, M.; Misják, P.; Deli, M.A.; Sipos, A.; Szalai, A.; Voszka, I.; et al. Detection and isolation of cell-derived microparticles are compromised by protein complexes resulting from shared biophysical parameters. Blood 2011, 117, e39–e48. [Google Scholar] [CrossRef] [PubMed]
- Vickers, K.C.; Palmisano, B.T.; Shoucri, B.M.; Shamburek, R.D.; Remaley, A.T. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat. Cell Biol. 2011, 13, 423–435. [Google Scholar] [CrossRef] [PubMed]
- Witwer, K.W.; Buzás, E.I.; Bemis, L.T.; Bora, A.; Lässer, C.; Lötvall, J.; Hoen, E.N.N.; Piper, M.G.; Sivaraman, S.; Skog, J.; et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research Need for standardization ISEV 2012 workshop on evRNA. J. Extracell. Vesicles 2014, 2, 1–29. [Google Scholar]
- Lötvall, J.; Hill, A.F.; Hochberg, F.; Buzás, E.I.; Di Vizio, D.; Gardiner, C.; Gho, Y.S.; Kurochkin, I.V.; Mathivanan, S.; Quesenberry, P.; et al. Minimal experimental requirements for definition of extracellular vesicles and their functions: A position statement from the International Society for Extracellular Vesicles. J. Extracell. Vesicles 2014, 3. [Google Scholar] [CrossRef] [PubMed]
- Mateescu, B.; Kowal, E.J.K.; van Balkom, B.W.M.; Bartel, S.; Bhattacharyya, S.N.; Buzás, E.I.; Buck, A.H.; de Candia, P.; Chow, F.W.N.; Das, S.; et al. Obstacles and opportunities in the functional analysis of extracellular vesicle RNA - an ISEV position paper. J. Extracell. Vesicles 2017, 6, 1286095. [Google Scholar] [CrossRef] [PubMed]
- Willms, E.; Johansson, H.J.; Mäger, I.; Lee, Y.; Blomberg, K.E.M.; Sadik, M.; Alaarg, A.; Smith, C.I.E.; Lehtiö, J.; El Andaloussi, S.; et al. Cells release subpopulations of exosomes with distinct molecular and biological properties. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- DANOVA, M.; COMOLLI, G.; MANZONI, M.; TORCHIO, M.; MAZZINI, G. Flow cytometric analysis of circulating endothelial cells and endothelial progenitors for clinical purposes in oncology: A critical evaluation. Mol. Clin. Oncol. 2016, 4, 909–917. [Google Scholar] [CrossRef]
- Bhakdi, S.C.; Suriyaphol, P.; Thaicharoen, P.; Grote, S.T.K.; Komoltri, C.; Chaiyaprasithi, B.; Charnkaew, K. Accuracy of Tumour-Associated Circulating Endothelial Cells as a Screening Biomarker for Clinically Significant Prostate Cancer. Cancers 2019, 11, 1064. [Google Scholar] [CrossRef]
- Najjar, F.; Alammar, M.; Al-Massarani, G.; Almalla, N.; Japawe, A.; Ikhtiar, A. Circulating endothelial cells and microparticles as diagnostic and prognostic biomarkers in small-cell lung cancer. Lung Cancer 2018, 124, 23–30. [Google Scholar] [CrossRef]
- Najjar, F.; Alammar, M.; Al-Massarani, G.; Almalla, N.; Aljapawe, A.; Ikhtiar, A. Circulating endothelial cells and microparticles for prediction of tumor progression and outcomes in advanced non-small cell lung cancer. Cancer Biomark. 2017, 20, 333–343. [Google Scholar] [CrossRef]
- Apolo, A.B.; Karzai, F.H.; Trepel, J.B.; Alarcon, S.; Lee, S.; Lee, M.J.; Tomita, Y.; Cao, L.; Yu, Y.; Merino, M.J.; et al. A Phase II Clinical Trial of TRC105 (Anti-Endoglin Antibody) in Adults With Advanced/Metastatic Urothelial Carcinoma. Clin. Genitourin. Cancer 2017, 15, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Ueno, T.; Masuda, N.; Kamigaki, S.; Morimoto, T.; Akiyama, F.; Kurosumi, M.; Tsuda, H.; Mikami, Y.; Tanaka, S.; Morita, S.; et al. A multicenter phase II trial of neoadjuvant letrozole plus low-dose cyclophosphamide in postmenopausal patients with estrogen receptor-positive breast cancer (JBCRG-07): Therapeutic efficacy and clinical implications of circulating endothelial cells. Cancer Med. 2018, 7, 2442–2451. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, S.; Kato, T.; Ogura, T.; Sekine, A.; Oda, T.; Masuda, N.; Igawa, S.; Katono, K.; Otani, S.; Yamada, K.; et al. Phase II study of bevacizumab, cisplatin, and docetaxel plus maintenance bevacizumab as first-line treatment for patients with advanced non-squamous non-small-cell lung cancer combined with exploratory analysis of circulating endothelial cells: Thoracic Oncology Research Group (TORG)1016. BMC Cancer 2018, 18, 241. [Google Scholar]
- García-Donas, J.; Leon, L.A.; Esteban, E.; Vidal-Mendez, M.J.; Arranz, J.A.; Garcia del Muro, X.; Basterretxea, L.; González del Alba, A.; Climent, M.A.; Virizuela, J.A.; et al. A Prospective Observational Study for Assessment and Outcome Association of Circulating Endothelial Cells in Clear Cell Renal Cell Carcinoma Patients Who Show Initial Benefit from First-line Treatment. The CIRCLES (CIRCuLating Endothelial cellS) Study (SOGUG-CEC-2011-01). Eur. Urol. Focus 2017, 3, 430–436. [Google Scholar] [PubMed]
- Ao, Z.; Shah, S.H.; Machlin, L.M.; Parajuli, R.; Miller, P.C.; Rawal, S.; Williams, A.J.; Cote, R.J.; Lippman, M.E.; Datar, R.H.; et al. Identification of cancer-associated fibroblasts in circulating blood from patients with metastatic breast cancer. Cancer Res. 2015, 75, 4681–4687. [Google Scholar] [CrossRef]
- Jones, M.L.; Siddiqui, J.; Pienta, K.J.; Getzenberg, R.H. Circulating fibroblast-like cells in men with metastatic prostate cancer. Prostate 2013, 73, 176–181. [Google Scholar] [CrossRef]
- Magbanua, M.J.M.; Pugia, M.; Lee, J.S.; Jabon, M.; Wang, V.; Gubens, M.; Marfurt, K.; Pence, J.; Sidhu, H.; Uzgiris, A.; et al. A novel strategy for detection and enumeration of circulating rare cell populations in metastatic cancer patients using automated microfluidic filtration and multiplex immunoassay. PLoS ONE 2015, 10. [Google Scholar] [CrossRef]
- Adams, D.L.; Adams, D.K.; Alpaugh, R.K.; Cristofanilli, M.; Martin, S.S.; Chumsri, S.; Tang, C.M.; Marks, J.R. Circulating cancer-associated macrophage-like cells differentiate malignant breast cancer and benign breast conditions. Cancer Epidemiol. Biomarkers Prev. 2016, 25, 1037–1042. [Google Scholar] [CrossRef]
- Hung, C.-H.; Chen, F.-M.; Lin, Y.-C.; Tsai, M.-L.; Wang, S.-L.; Chen, Y.-C.; Chen, Y.-T.; Hou, M.-F. Altered monocyte differentiation and macrophage polarization patterns in patients with breast cancer. BMC Cancer 2018, 18, 366. [Google Scholar] [CrossRef]
- Adams, D.L.; Adams, D.K.; He, J.; Kalhor, N.; Zhang, M.; Xu, T.; Gao, H.; Reuben, J.M.; Qiao, Y.; Komaki, R.; et al. Sequential tracking of PD-L1 expression and RAD50 induction in circulating tumor and stromal cells of lung cancer patients undergoing radiotherapy. Clin. Cancer Res. 2017, 23, 5948–5958. [Google Scholar] [CrossRef]
- Sutton, T.L.; Walker, B.S.; Wong, M.H. Circulating Hybrid Cells Join the Fray of Circulating Cellular Biomarkers. Cell. Mol. Gastroenterol. Hepatol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Marleau, A.M.; Chen, C.S.; Joyce, J.A.; Tullis, R.H. Exosome removal as a therapeutic adjuvant in cancer. J. Transl. Med. 2012, 10. [Google Scholar] [CrossRef] [PubMed]
- De La Fuente, A.; Alonso-Alconada, L.; Costa, C.; Cueva, J.; Garcia-Caballero, T.; Lopez-Lopez, R.; Abal, M. M-Trap: Exosome-Based Capture of Tumor Cells as a New Technology in Peritoneal Metastasis. J. Natl. Cancer Inst. 2015, 107. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Su, C. Design strategies and application progress of therapeutic exosomes. Theranostics 2019, 9, 1015–1028. [Google Scholar] [CrossRef]
- Park, K.-S.; Bandeira, E.; Shelke, G.V.; Lässer, C.; Lötvall, J. Enhancement of therapeutic potential of mesenchymal stem cell-derived extracellular vesicles. Stem Cell Res. Ther. 2019, 10, 288. [Google Scholar] [CrossRef]
- Yuan, Z.; Kolluri, K.K.; Gowers, K.H.C.; Janes, S.M. TRAIL delivery by MSC-derived extracellular vesicles is an effective anticancer therapy. J. Extracell. Vesicles 2017, 6, 1265291. [Google Scholar] [CrossRef]
- Altanerova, U.; Jakubechova, J.; Benejova, K.; Priscakova, P.; Pesta, M.; Pitule, P.; Topolcan, O.; Kausitz, J.; Zduriencikova, M.; Repiska, V.; et al. Prodrug suicide gene therapy for cancer targeted intracellular by mesenchymal stem cell exosomes. Int. J. Cancer 2019, 144, 897–908. [Google Scholar] [CrossRef]
- Cheng, L.; Wang, Y.; Huang, L. Exosomes from M1-Polarized Macrophages Potentiate the Cancer Vaccine by Creating a Pro-inflammatory Microenvironment in the Lymph Node. Mol. Ther. 2017, 25, 1665–1675. [Google Scholar] [CrossRef]
- Choo, Y.W.; Kang, M.; Kim, H.Y.; Han, J.; Kang, S.; Lee, J.R.; Jeong, G.J.; Kwon, S.P.; Song, S.Y.; Go, S.; et al. M1 Macrophage-Derived Nanovesicles Potentiate the Anticancer Efficacy of Immune Checkpoint Inhibitors. ACS Nano 2018, 12, 8977–8993. [Google Scholar] [CrossRef]
- Kamerkar, S.; Lebleu, V.S.; Sugimoto, H.; Yang, S.; Ruivo, C.F.; Melo, S.A.; Lee, J.J.; Kalluri, R. Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature 2017, 546, 498–503. [Google Scholar] [CrossRef]
- Abak, A.; Abhari, A.; Rahimzadeh, S. Exosomes in cancer: Small vesicular transporters for cancer progression and metastasis, biomarkers in cancer therapeutics. PeerJ 2018, 2018. [Google Scholar] [CrossRef] [PubMed]
- Kalimuthu, S.; Gangadaran, P.; Rajendran, R.L.; Zhu, L.; Oh, J.M.; Lee, H.W.; Gopal, A.; Baek, S.H.; Jeong, S.Y.; Lee, S.-W.; et al. A New Approach for Loading Anticancer Drugs Into Mesenchymal Stem Cell-Derived Exosome Mimetics for Cancer Therapy. Front. Pharmacol. 2018, 9, 1116. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.S.; Haney, M.J.; Zhao, Y.; Yuan, D.; Deygen, I.; Klyachko, N.L.; Kabanov, A.V.; Batrakova, E.V. Engineering macrophage-derived exosomes for targeted paclitaxel delivery to pulmonary metastases: In vitro and in vivo evaluations. Nanomed. Nanotechnol. Biol. Med. 2018, 14, 195–204. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Wang, H.; Huang, Q.; Peng, C.; Yao, L.; Chen, H.; Qiu, Z.; Wu, Y.; Wang, L.; Chen, W. Exosomes from M1-polarized macrophages enhance paclitaxel antitumor activity by activating macrophages-mediated inflammation. Theranostics 2019, 9, 1714–1727. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics of American 2019. CA Cancer J. Clin. 2019, 69, 7–34. [Google Scholar] [CrossRef]
| Soluble Factors in Blood | Involvement | References |
|---|---|---|
| Matrix metalloproteinase | With diagnostic value | [26,170] |
| Collagens | Higher levels in CRC patients than in healthy controls | [171,172,173] |
| Endostatin | Higher levels in advanced CRC patients | [174] |
| TIMP-1 + metalloproteinases | Prediction of patients survival | [175] |
| CAF Exosomes as Biomarkers | ||
| Non-coding RNAs signature | Regulatory elements specifically packaged in CAF-derived exosomes | [18] |
| miR-92a-3p | Higher levels associated with metastases and chemoresistance | [105] |
| microRNA signature | Regulation of tumor cell, proliferation, and chemoresistance | [106] |
| Circulating Endothelial Cells (CECs) | ||
| Identification and quantification of CECs | Monitoring clinical response and outcome | [176,177,178] |
| CD276 | Increase expression in tumor-derived endothelial cells | [179] |
| Transcriptomic analysis | Differentiation between healthy controls and CRC early stages | [180] |
| Quantification of CECs | Identification of early predictors of response to bevacizumab and FOLFOX/OXXEL | [181] |
| Circulating Immune Cells | ||
| Treg, myeloid-derived suppressor cells, and neutrophil-to-lymphocyte rate | Identification of CRC patients versus healthy controls | [182,183,184] |
| Immune checkpoints and clinical outcome | Association with diagnosis and metastasis | [185,186] |
| Mucosal-associated invariant T cells | Increase number in CRC patients versus healthy controls | [187] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Herrera, M.; Galindo-Pumariño, C.; García-Barberán, V.; Peña, C. A Snapshot of The Tumor Microenvironment in Colorectal Cancer: The Liquid Biopsy. Int. J. Mol. Sci. 2019, 20, 6016. https://doi.org/10.3390/ijms20236016
Herrera M, Galindo-Pumariño C, García-Barberán V, Peña C. A Snapshot of The Tumor Microenvironment in Colorectal Cancer: The Liquid Biopsy. International Journal of Molecular Sciences. 2019; 20(23):6016. https://doi.org/10.3390/ijms20236016
Chicago/Turabian StyleHerrera, Mercedes, Cristina Galindo-Pumariño, Vanesa García-Barberán, and Cristina Peña. 2019. "A Snapshot of The Tumor Microenvironment in Colorectal Cancer: The Liquid Biopsy" International Journal of Molecular Sciences 20, no. 23: 6016. https://doi.org/10.3390/ijms20236016
APA StyleHerrera, M., Galindo-Pumariño, C., García-Barberán, V., & Peña, C. (2019). A Snapshot of The Tumor Microenvironment in Colorectal Cancer: The Liquid Biopsy. International Journal of Molecular Sciences, 20(23), 6016. https://doi.org/10.3390/ijms20236016





