1. Introduction
Plants possess a variety of defense mechanisms for preventing pathogen infection, including the accumulation of specialized metabolites. In response to pathogen infection, many plant species accumulate antimicrobial metabolites, i.e., phytoalexins, which provide chemical defense [
1]. The importance of chemical defense in plants has been indicated by various kinds of evidence: the lack of phytoalexin resulted in the increased susceptibility of plants [
2], and accumulation of phytoalexin in other plants by introduction of biosynthetic genes enhanced resistance levels [
3]. In addition, some pathogens have been found to counteract to phytoalexin production by acquisition of the degradation and excretion mechanism of phytoalexins [
4].
Phenylamides are amides of hydroxycinnamic and benzoic acids, with various biogenic amines, and have been reported to contribute to plant defense responses. Representative hydroxycinnamic acids components of phenylamides include cinnamic (Cin),
p-coumaric (Cou), caffeic (Caf), ferulic (Fer), and benzoic (Ben) acids, whereas representative amine components include tryptamine (Try), serotonin (Ser), tyramine (Tyr), agmatine (Agm), and putrescine (Put). Hereafter, specific phenylamides will be referred to using combined abbreviations of their constituent acids and amines, e.g., CinTry (cinnamoyltryptamine). Phenylamide accumulation has been reported to occur in a wide range of plant species. In the Poaceae species, various phenylamides have been reported to accumulate in response to pathogen attack and have been indicated to function as phytoalexins in plant chemical defenses. In rice (
Oryza sativa), for example, plants have been reported to accumulate BenTry and CinTry, as phytoalexins, in response to attack by rice blast fungus (
Pyricularia oryzae) [
5] and to accumulate FerTry, CouSer, FerSer, and Ser in response to infection by brown spot fungus (
Cochliobolus miyabeanus) [
6]. In addition, rice leaves have even been reported to accumulate phenylamides (CinTry, CinTyr, CouSer, and BenTry) in response to ultraviolet (UV) light exposure [
7,
8]. Meanwhile, in oats, avenanthramides, which are hydroxycinnamic acid amides with anthranilic acids, function as phytoalexins [
9] and as substrates for the reinforcement of cell walls [
10]. Furthermore, wheat (
Triticum aestivum) leaves have recently been reported to accumulate
N-cinnamoyl-8-oxotryptamine and
N-cinnamoyl-9-hydorxy-8-oxotryptamine, as well as CouAgm, FerAgm, CouPut, FerPut, and FerSer in response to attack by
Bipolaris sorokiniana, which is the causal agent of brown spot [
11], and it is possible that
N-cinnamoyl-8-oxotryptamine and
N-cinnamoyl-9-hydorxy-8-oxotryptamine function as phytoalexins, on the basis of their anti-microbial properties.
Phenylamides are biosynthesized via the condensation of hydroxycinnamoyl-CoA esters and amines, and this reaction is catalyzed by a variety of hydroxycinnamoyl transferases [
12,
13]. In rice, for example, tryptamine benzoyl transferase (TBT), tryptamine hydroxycinnamoyl transferase (THT), and putrescine hydroxycinnamoyl transferase (PHT) contribute to the biosynthesis of the phenylamides that are constitutively present in rice leaves [
14,
15], and in oats, hydroxyanthranilate
N-hydroxycinnamoyl transferase (HHT) is involved in the biosynthesis of avenanthramide phytoalexins [
16,
17]. The enzymes involved in the synthesis of these phenylamides are classified into the BAHD acyltransferase family, which was named for the first letter of each of the first four biochemically characterized enzymes in the family, namely BEAT (benzoyl alcohol
O-acetyltransferase), AHCT (anthocyanin
O-hydroxycinnamoyl transferase), HCBT (anthranilate
N-hydroxycinnamoyl/benzoyl transferase), and DAT (deacetyl vindoline 4-
O-acetyltransferase) [
12]. In barley, the role of agmatine
N-coumaroyltransferase (ACT), which is also included in the BAHD acyltransferase family, in the biosynthetic of CouAgm has been well studied [
18,
19]. However, the enzymes and genes involved in the synthesis of phenylamides from aromatic amines have not been investigated, despite that tyramine hydroxycinnamoyl transferase activity has been detected in young seedling roots [
20].
The accumulation of specialized metabolites has been investigated in pathogen-challenged barley plants.
Bipolaris sorokiniana-infected barley leaves, for example, were reported to accumulate the dimer of serotonin and 3-(2-aminoethyl)-3-hydroxyindolin-2-one, which may function as phytoalexins [
21], and
Fusarium graminearum-infected barley roots were reported to exude cinnamic,
p-coumaric, ferulic, syringic, and vanillic acids [
22]. Furthermore, the metabolomics analysis performed by Karre et al. [
23] revealed differences in the accumulation of metabolites in the spikelets of
F. graminearum-resistant and -susceptible cultivars, and the authors reported that phenolics, lignans, hydroxycinnamic acid amides (phenylamides), flavonoids, signaling molecule-related compounds, terpenoids, indole alkaloids, and methionine biosynthesis-related compounds contributed to
Fusarium head blight resistance that was mediated by chitin elicitor receptor kinase. Even more recently, barley leaves have been reported to accumulate cinnamoyl-9-hydroxy-8-oxotryptamine and cinnamoyl-8-oxotryptamine, which function as phytoalexins in wheat, in response to CuCl
2 treatment [
11]. In barley, phenylamides that contain Agm and Put are reportedly induced by pathogen infection [
23,
24]. However, no other inducible phenylamides have been characterized, despite the characterization of multiple phenylamides from the poaceous species. Accordingly, the aim of the present study was to investigate the inducible phenylamides of barley, in the hopes of identifying an undiscovered chemical defense mechanism. As a result, three phenylamide phytoalexins, including a previously undescribed compound, were identified, and acyltransferase genes that encode the enzymes that catalyze inducible phenylamide synthesis were characterized. On the basis of these findings, the present study provides novel insight into the chemical defense mechanisms of barley, as well as a framework for the biosynthesis of phenylamide phytoalexins.
3. Discussion
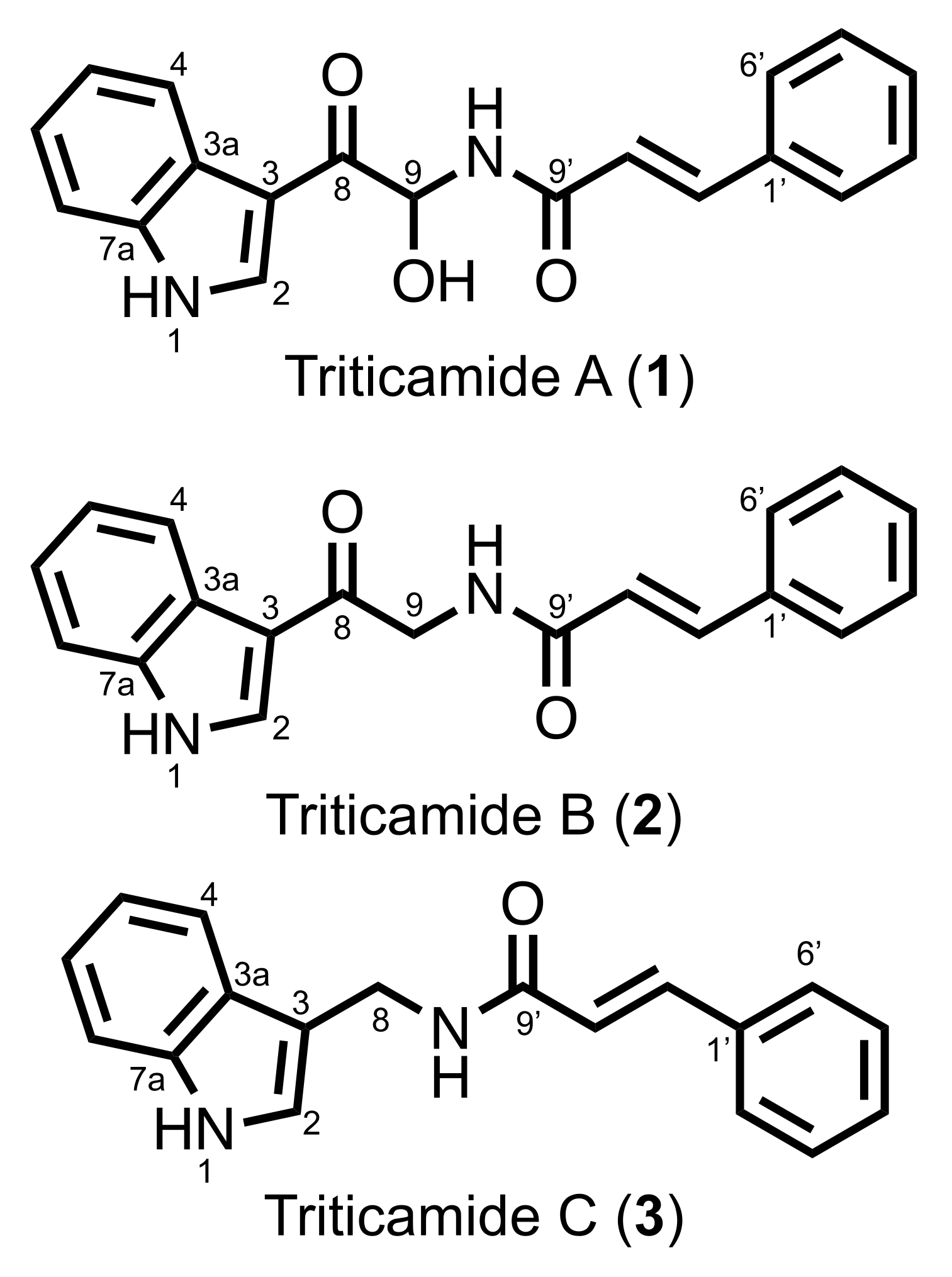
The present study demonstrated that barley roots accumulate three inducible compounds, namely
N-cinnamoyl-9-hydroxy-8-oxotryptamine (
1) and
N-cinnamoyl-8-oxotryptamine (
2), and
N-cinnamoyl-(1
H-indol-3-yl)methylamine (
3), in response to pathogen attack (
Figure 2). Compounds
1 and
2 had already been characterized as phytoalexins in wheat [
11], but compound
3 was a previously undescribed compound. Because both barley and wheat belong to the Triticeae tribe, the compounds
1–
3 were named triticamides A–C, respectively. The antifungal activities of triticamides A and B have been reported previously [
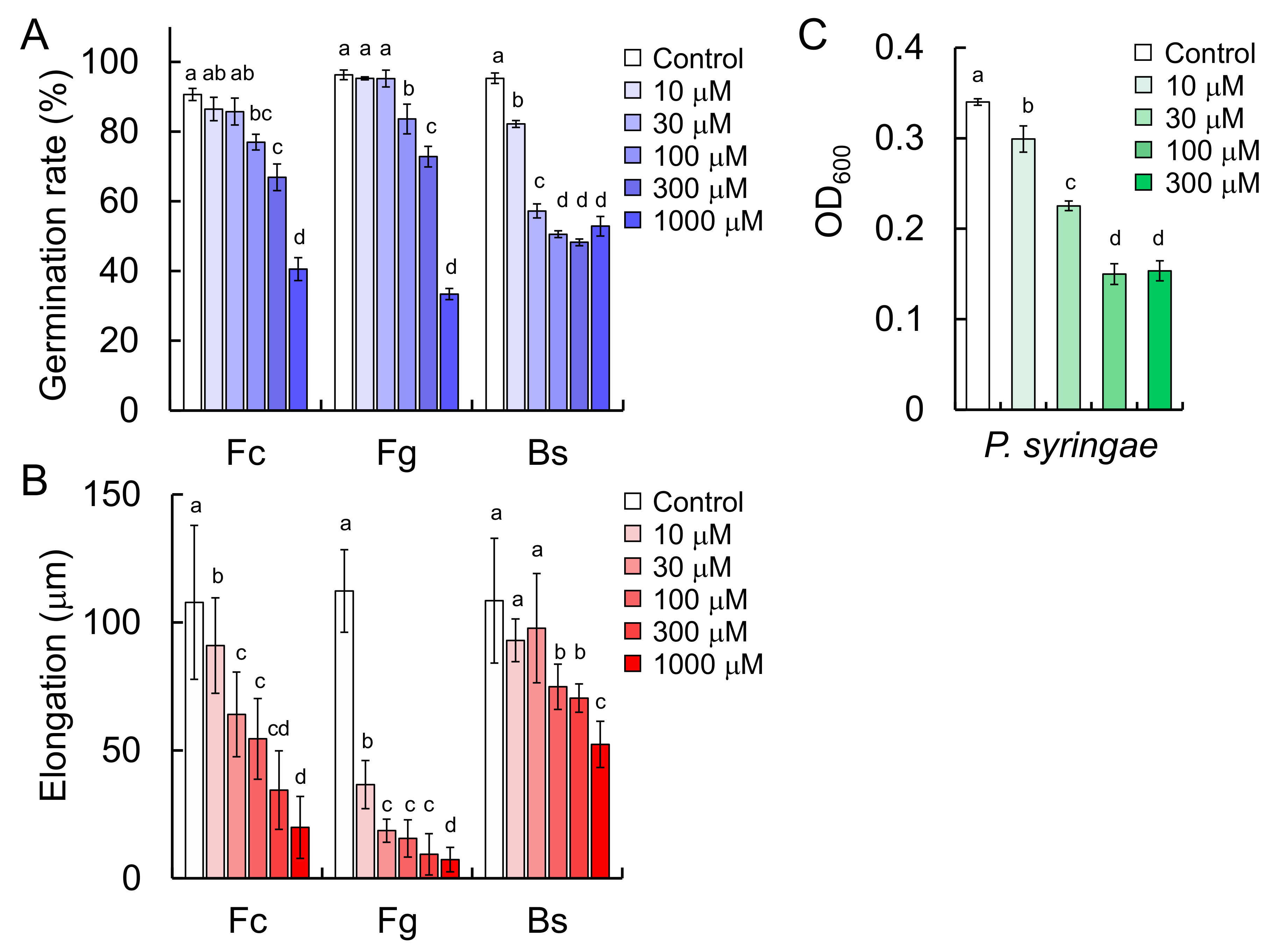
11], and the present study demonstrated that triticamide C also possesses antifungal activity against several species of phytopathogenic fungi (
F. culmorum,
F. graminearum, and
B. sorokiniana). It is of interest to note that triticamide C inhibited the growth of the bacterial pathogen
P. syringae at 100 µM because triticamides A and B did not affect the growth of
P. syringae [
11]. Triticamides function as phytoalexins in barley as previously demonstrated in wheat.
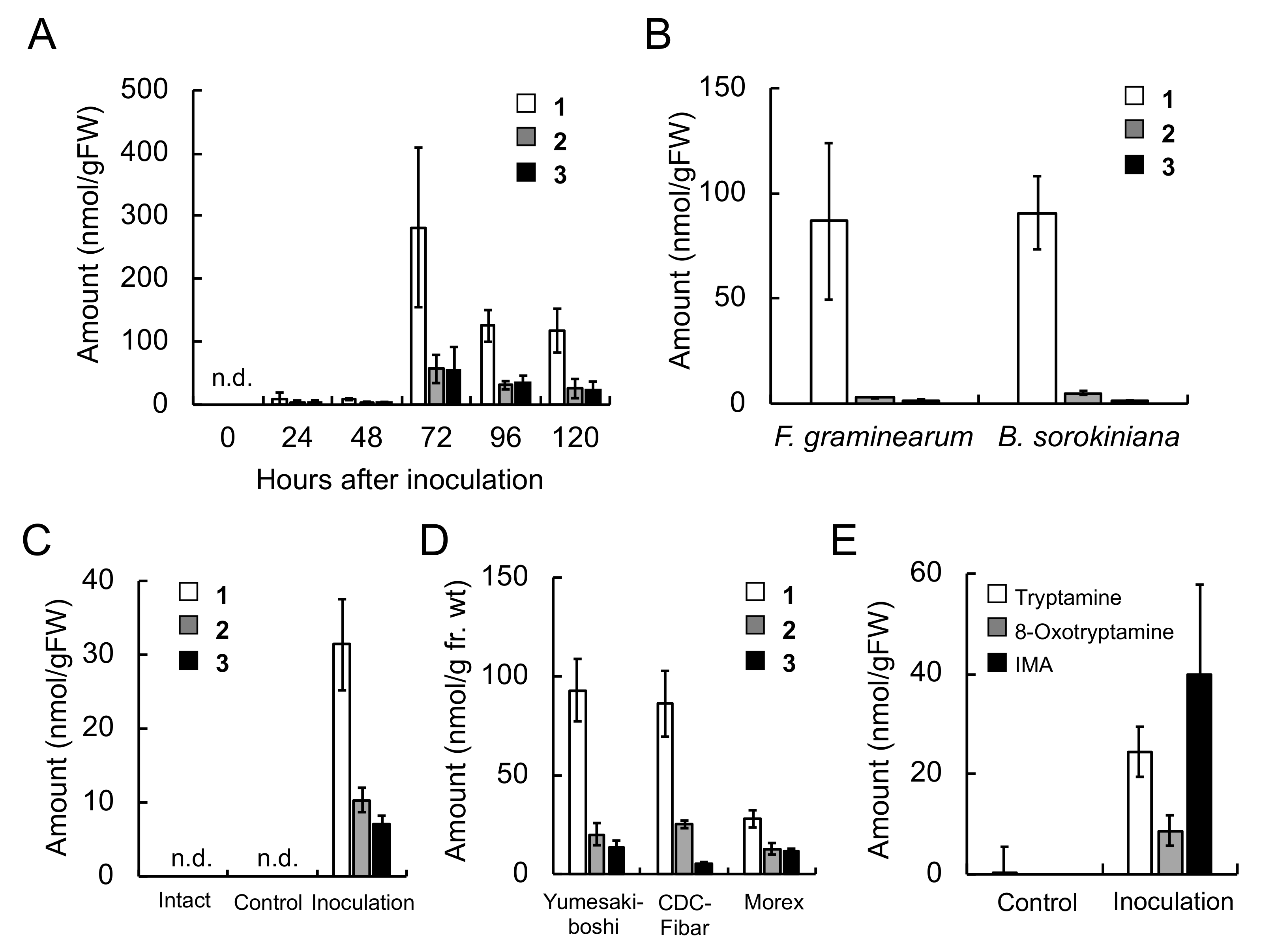
The concentrations of triticamides A–C were 282, 56.7 and 55.0 nmol/g FW, respectively, in
F. culmorum-infected roots 72 h after inoculation, and 31.4, 10.3, and 7.0 nmol/g FW, respectively, in
B. sorokiniana-infected leaves 48 h after inoculation (
Figure 3). These concentrations are similar to the concentrations at which the triticamides exert antifungal activities. For example, triticamides B and C significantly inhibited the germ tube elongation of
F. graminearum at 10 µM, and triticamides B and C inhibited the germination of
B. sorokiniana conidia at 10 and 100 µM, respectively (
Figure 8). The accumulation of triticamides likely affects the growth of these pathogens in plant tissues.
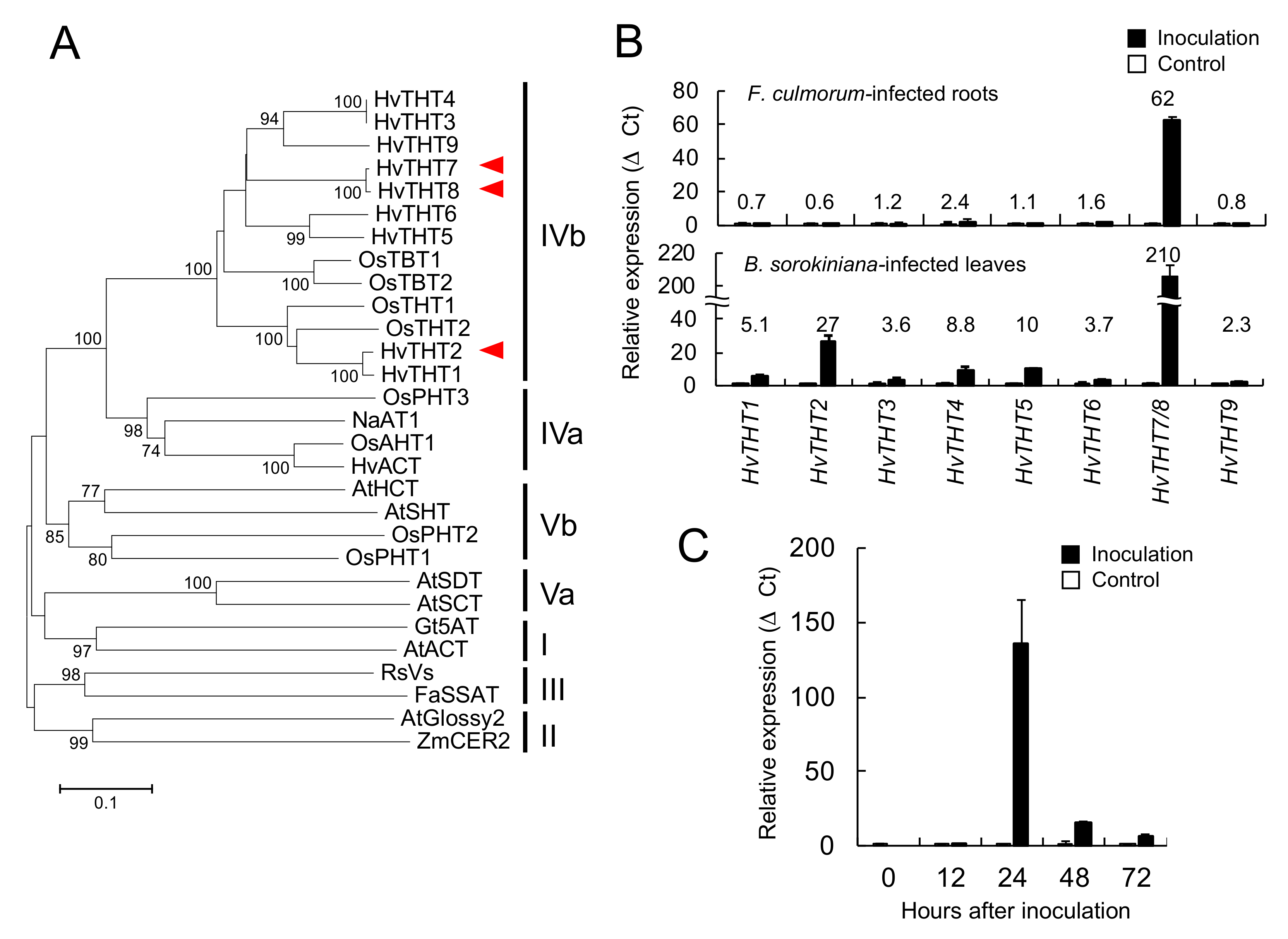
Phenylamides are biosynthesized by the condensation of hydroxycinnamic acid CoA thioesters and amines. Among the THT- and TBT-related genes that were identified in the barley genome, the combined expression levels of
HvTHT7/
8 were markedly higher in pathogen-infected roots and leaves and reached maximum levels 24 h after inoculation, which preceded the accumulation of triticamides (
Figure 3). Generally speaking, the accumulation of inducible metabolites is preceded by enhanced gene expression. For instance, in rice leaves, the amount of rice phytoalexin sakuranetin reached at a maximum 72 h after the start of treatment whereas the transcript amount of
NOMT that encodes naringenin-7-
O-methyltransfearase reached at a maximum 6 h after the treatment [
26]. Similarly, the accumulation of avenanthramides in oat leaves was about 24 h behind the expression of biosynthetic genes [
17]. To explain this gap of timings of
HvTHT7/8 expression and triticamide accumulation, however, the kinetic analysis of expression of other biosynthetic genes is needed. Some of remaining
HvTHT genes showed enhanced expression in
F. culmorum-infected root, but the fold changes were relatively small (
Figure S2). In addition, the characterization of heterologously expressed HvTHT7 and HvTHT8 revealed that both enzymes favored Cin-CoA as acyl donors and tryptamine as an acyl acceptor (
Table 2). Together, these findings suggest that HvTHT7/8 are involved in triticamide biosynthesis.
The genomic sequences of
HvTHT7 and
HvTHT8 were different only at one nucleotide at 497 bps from the start codon. In the present study, we mainly used the cultivar ‘Shunrei’. In this cultivar, we detected only the transcript of
HvTHT8 and the genomic sequence corresponding to
HvTHT8. Thus, ‘Shunrei’ probably poses only
HvTHT8. By contrast, we detected the transcripts and genomic sequences corresponding to both
HvTHT7 and
HvTHT8 in ‘Morex’. This is consistent with the result of database search–both
HvTHT7 and
HvTHT8 were deposited to ‘Morex’ genome database (URL:
https://webblast.ipk-gatersleben.de/barley_ibsc/). The ‘Morex’ probably poses both genes in its genome. Furthermore, only
HvTHT7 was deposited in ‘Barke’ genome database, whereas only
HvTHT8 was in ‘Bowman’ genome database (URL:
https://webblast.ipk-gatersleben.de/barley_ibsc/). Thus, the barley genome is polymorphic at this specific nucleotide in
HvTHT7/8 but some cultivars such as ‘Morex’ acquired both
HvTHT7 and
HvTHT8 during the breeding process. Because the enzymatic function of HvTHT7 and HvTHT8 were not largely different, this single nucleotide polymorphism does not affect the triticamide production.
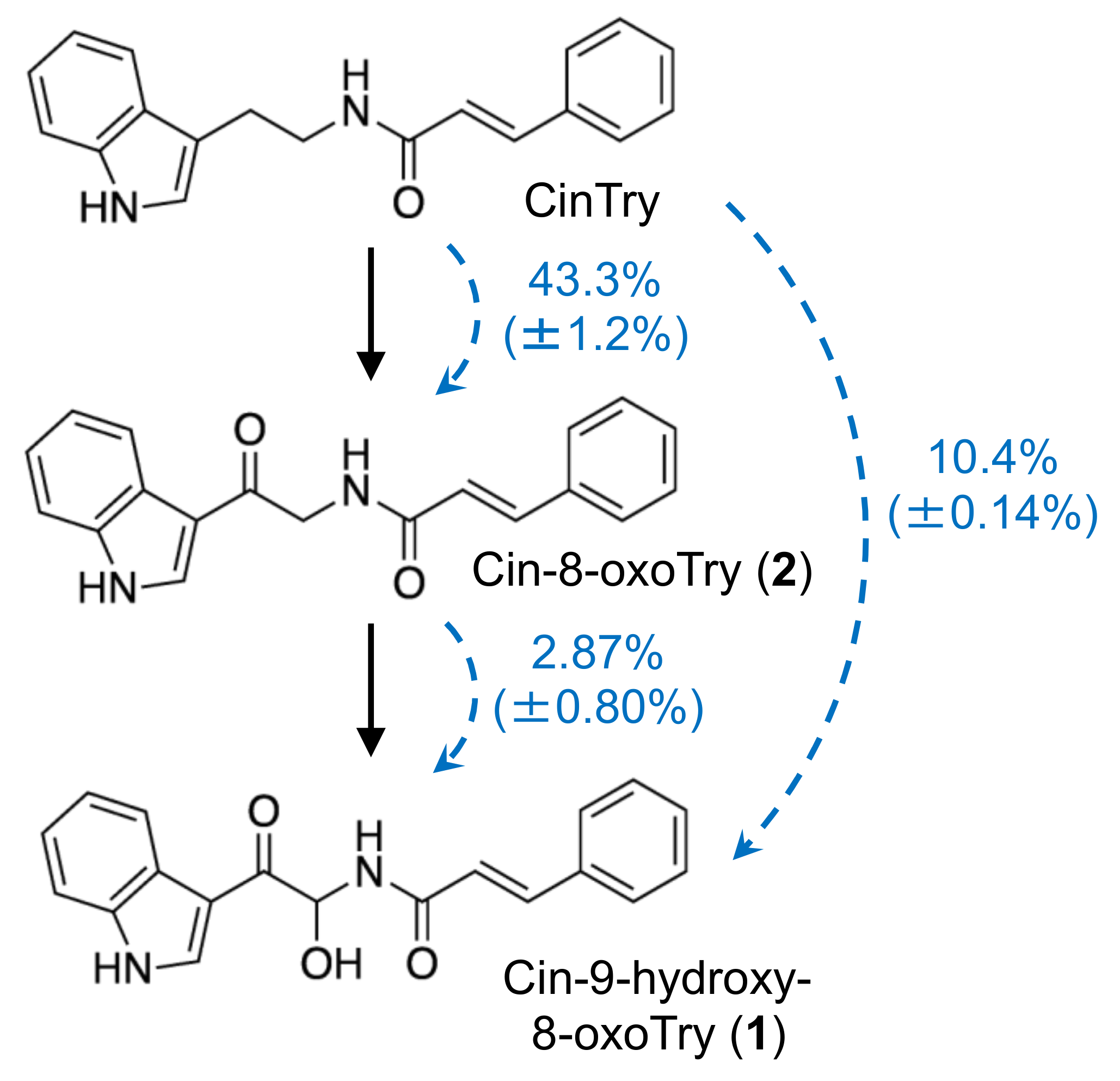
There are two hypothetical pathways for the biosynthesis of
1 and
2: the direct condensation of 9-hydroxy-8-oxotryptamine and 8-oxotryptamine with Cin-CoA thioester or the condensation of tryptamine and Cin-CoA into CinTry and the subsequent oxidation of the tryptamine part of the compound. The high substrate specificity of HvTHT7 and HvTHT8 for tryptamine suggests that the second hypothesis is more likely (
Table 2). In addition, observed accumulation of tryptamine was much greater than that of 8-oxotryptamine (
Figure 3). Indeed, subsequent feeding experiments confirmed that labeled CinTry was effectively incorporated into triticamides A and B (
Figure 7). Thus, it is most likely that
1 and
2 are biosynthesized by this route. However, 8-oxotryptamine was also detected in the pathogen-infected roots and HvTHT7 and HvTHT8 accepted 8-oxotryptamine to some extent. Thus, the possibility that triticamides are synthesized by the direct condensation should not be excluded, and the pathways may form a metabolic grid. To obtain a whole picture of the pathways for triticamide biosynthesis, the concentrations and kinetics of 9-hydroxy-8-oxotryptamine in barley roots should be investigated.
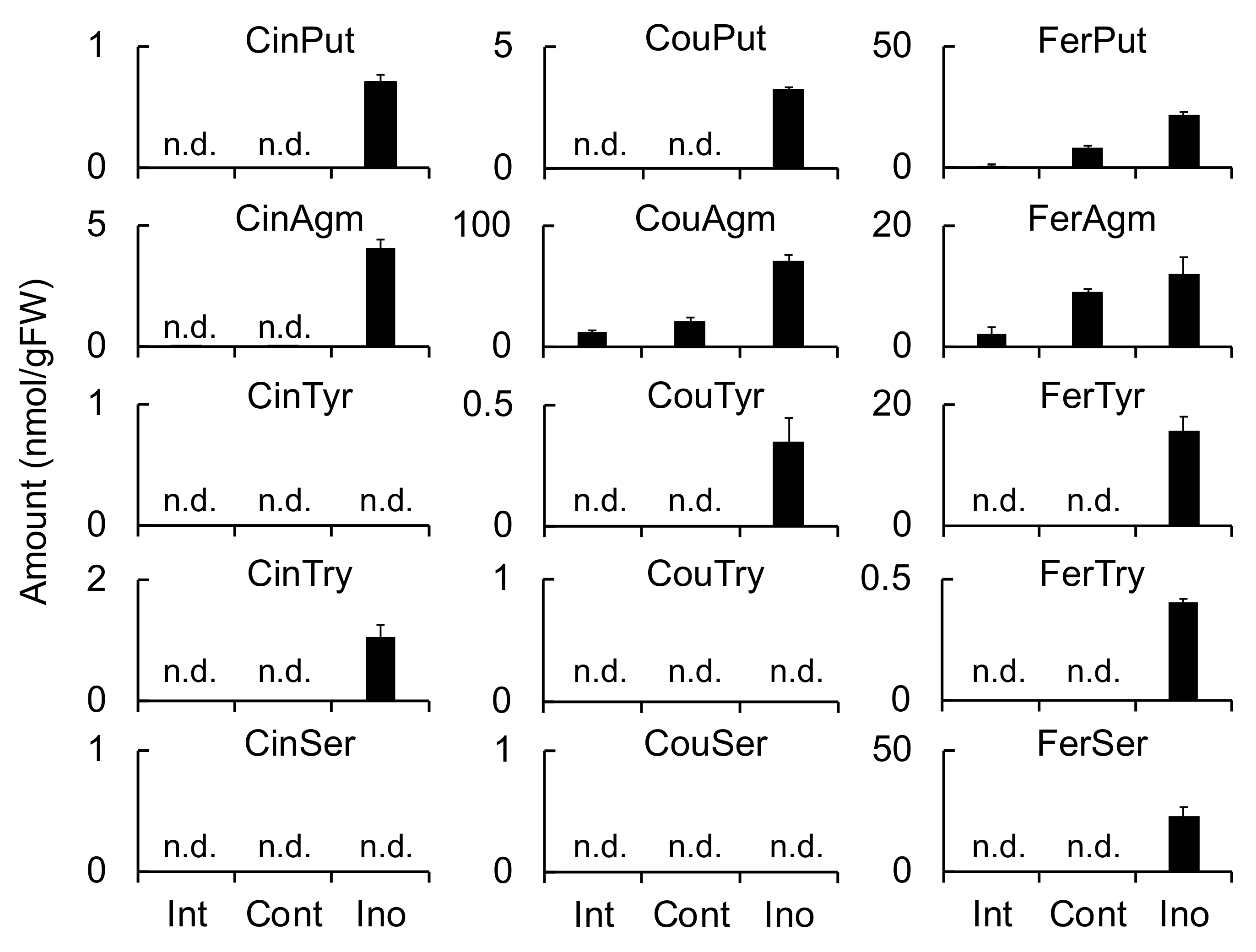
Meanwhile, HvTHT2 preferred Fer-CoA as an acyl donor and tryptamine and serotonin as acyl acceptors. The
kcat value of HvTHT2 for Cin-CoA was much smaller than that for Fer-CoA, and the relative efficiency for Cin-CoA was only 0.033% that for Fer-CoA (
Table 3). Therefore, it is unlikely that HvTHT2 contributes substantially to the biosynthesis of triticamides. The expression of
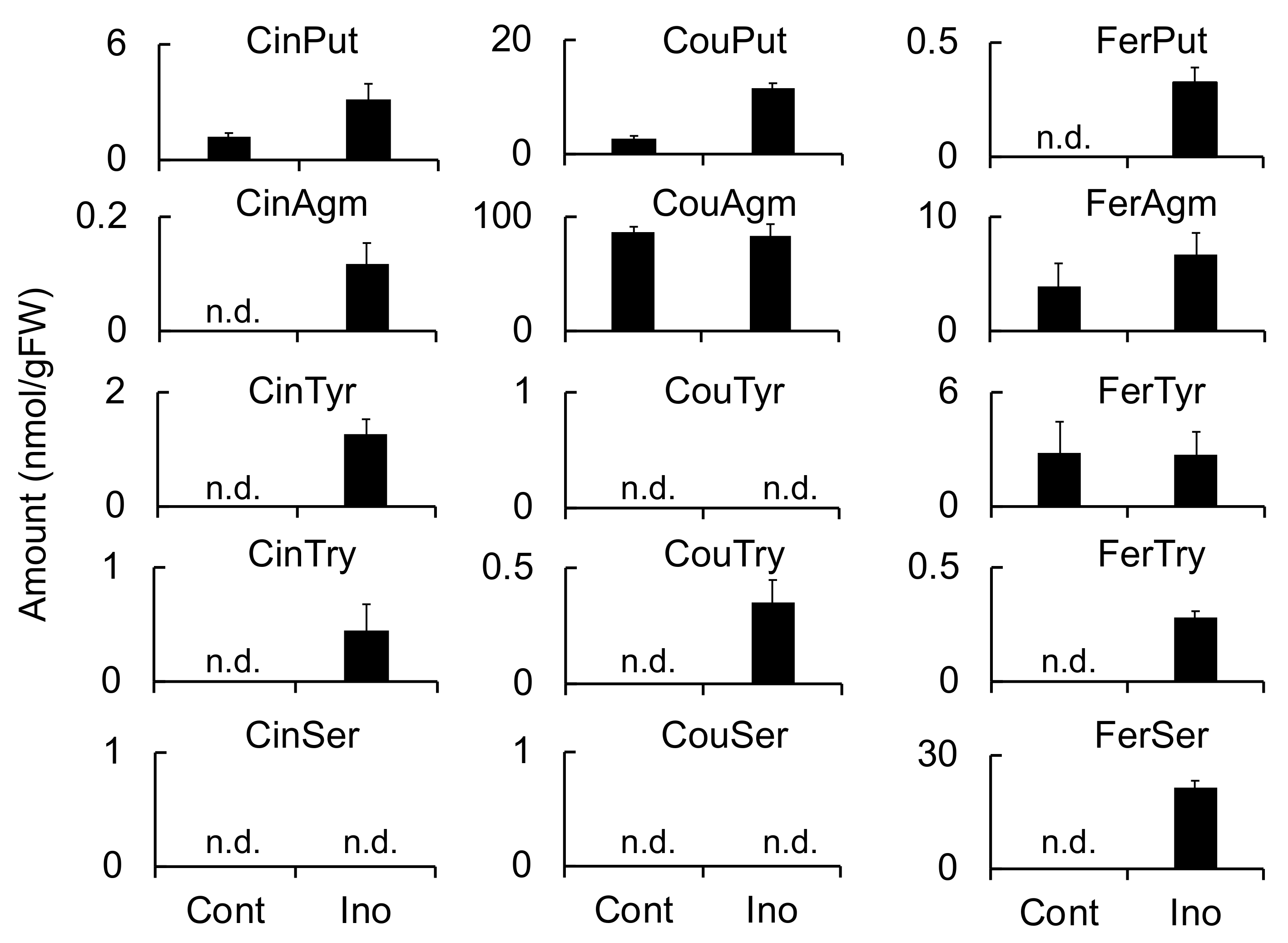
HvTHT2 was enhanced by
B. sorokiniana infection in leaves but not by
F. culmorum infection in roots, and both FerTyr and FerSer were observed to accumulate to high levels in
B. sorokiniana-infected leaves (
Figure 4,
Figure 5 and
Figure 6). Because HvTHT2 accepted tyramine and serotonin as an acyl acceptor, it is likely that
HvTHT2 is involved in the formation of such amides, rather than that of triticamides.
Triticamide C is considered to be biosynthesized by the condensation of Cin-CoA and IMA. Indeed, the recombinant enzymes HvTHT7 and HvTHT8 catalyzed this condensation reaction efficiently, and IMA accumulation was induced by
F. culmorum infection to 39.9 nmol/g FW in infected roots (
Table 2 and
Figure 3). Therefore, the present study confirms the hypothesis that triticamide C is formed by the direct condensation of Cin-CoA and IMA. This amine is also a precursor of gramine, a specialized metabolite that is also present in barley [
27]. Gramine is formed by the successive methylation of IMA by a methyltransferase [
28]. However, the biosynthetic pathway from tryptophan to IMA has yet to be fully elucidated. Interestingly, several barley cultivars (e.g., ‘Morex’) do not accumulate gramine [
28]. Therefore, it is worth noting that, in the present study, all the examined cultivars including ‘Morex’ accumulated triticamide C and, thus, produce IMA in response to pathogen infection.
The BAHD acyltransferase family is separated into five clades (Clades I-V), and Clade IV is further divided into Clades IVa and IVb (
Figure 6) [
12,
14]. In the present study, HvTHT2, HvTHT7, and HvTHT8 were mapped to Clade IVb along with OsTHTs and OsTBTs. The catalytic activities of enzymes in this subfamily have been studied in several grass species, and all the studied enzymes have been reported to catalyze the condensation of tryptamine with either Ben-CoA or Cou-CoA [
14]. In the present study, HvTHT2, HvTHT7, and HvTHT8 also accepted tryptamine as a substrate. Thus, it is reasonable to infer that enzymes in this subclade similarly prefer indole amines as substrates. Peng et al. [
14] also identified Clade IV- (EVDSWL and VLWAFP) and Clade IVb-specific (VRVAVNC and RRRR) motifs in enzymes from rice (
Figure S5). These motifs were also present in HvTHT2, HvTHT7, and HvTHT8, with the exception of the RRRR motif. Analyzing the functions of these motifs may help elucidate the structural factors that affect substrate specificity.
On the other hand, HvTHT7 and HvTHT8 showed different substrate specificity for CoA thioesters in comparison with OsTHTs and OsTBTs. HvTHT7 and HvTHT8 preferred Fer-CoA as well as Cin-CoA while OsTBTs and OsTHT1 showed low specificity to Fer-CoA although specificity of OsTHT2 for Fer-CoA has not been examined. OsTBTs and OsTHTs showed high specificities for Cou- and Ben-CoAs. We could obtain the sequences of 115 BAHD family acyltransferases by BLAST search using HvTHT7 amino acid sequences as query from the species in Poaceae family (
Figure S6). Phylogenetic analysis of the acyltransferases indicated that HvTHT7 and HvTHT8 formed a cluster together with THTs in
T. aestivum,
T. urartu,
T. durum, Aegilops tauschii, Secale cereale, and Brachypodium distachion. All of these species belong to the Pooideae subfamily. Thus, the acyltransferases in this cluster may have branched off from the cluster containing HvTHT2, OsTHT1/2, and OsTBT1/2, and play a specific role in Pooideae species. In this context, it is of interest to investigate the distribution of triticamide phytoalexins in these species as well as to analyze the substrate specificity of THTs in this cluster.
4. Materials and Methods
4.1. General Experimental Procedures
Both 1H and 13C NMR spectra and 2D COSY, HMQC, and HMBC spectra were recorded using an Avance II instrument (Bruker, Billerica, MA, USA). High-resolution mass spectra were measured using an Exactive mass spectrometer (Thermo Fisher Scientific, Waltham, MA, USA), and ESI-MS measurements were performed using a Quattro Micro API mass spectrometer (Waters, Milford, MA, USA) that was connected to an Acquity UPLC system (Waters). HPLC was performed using a 10A HPLC system (Shimadzu, Kyoto, Japan).
4.2. Plant Materials and Pathogenic Fungi
Barley (Hordeum vulgare ‘Shunrei’) and wheat (Triticum aestivum ‘Norin 61′) seeds were purchased from JA Inaba Tottori (Tottori, Japan) and Tsurushin Shubyo (Kyoto, Japan), respectively. Seeds of barley ‘Yumesakiboshi’, ‘CDC fibar’ and ‘Morex’ were stocks in the Institute of Plant Science and Resources, Okayama University, Japan. For the experiments that involved roots, seeds were placed on filter paper in a plant culture container (ф120 mm diammeter × 80 mm hight, SPL Life Science, Gyeonggi-do, Korea) that contained 5 mL sterile water, incubated at 4 °C in the dark for 24 h, and then incubated at 25 °C with a 14-h photoperiod for 3 days. For the experiments that involved leaves, seeds were immersed in distilled water for one night, sown on a 1:1 (v:v) mixture of vermiculite (Shoei Sangyo, Okayama, Japan) and culture soil (Iris Ohyama, Sendai, Japan), and then incubated at 25 °C with a 14-h photoperiod for three weeks.
Fusarium culmorum (MAFF 236454), the causal agent of
Fusarium root rot, was obtained from the National Institute of Agrobiological Sciences Genebank (
http://www.gene.affrc.go.jp/index_en.php), whereas
F. graminearum was obtained from the National Agriculture and Food Research Organization, and
B. sorokiniana (OB-25-1), the causal agent of barley spot blotch, was a stock in Natural Product Chemistry Laboratory, Faculty of Agriculture, Tottori University. The
F. culmorum,
F.
graminearum, and
B.
sorokiniana were inoculated onto V8 agar plates. To inoculate plant roots, the
F. culmorum, F. graminearum, and
B.
sorokiniana were cultured for 5 d at 25 °C under black light (FL15BL-B; Hitachi, Tokyo, Japan), and plugs from the V8 agar plates were used as inoculum. Meanwhile, conidia suspensions were used to inoculate plant leaves. To obtain conidia,
F. culmorum and
F. graminearum were cultured for one week under the conditions described above, whereas
B.
sorokiniana was cultured for two weeks.
The causal agent of bacterial black node, Pseudomonas syringae pv. japonica (MAFF 301072), was obtained from the National Institute of Agrobiological Sciences Genebank. The pathogen was cultured on potato semi-synthetic medium (300 g potato, 2 g Na2PO4·12H2O, 15 g sucrose, 5 g peptone, 0.5 g Ca(NO3)2·4H2O, and 1 L distilled water) at 25 °C with shaking (200 rpm) for 24 h.
4.3. Inoculation of Pathogenic Fungi
The inoculation of pathogens onto barley and wheat roots was performed according to the methods of Covarelli et al. [
29] with slight modification. Briefly, mycelial agar plugs (0.5- to 0.6-cm diameter) were taken from the growing edges of a 5-d-old culture plates and placed on the roots of the 96-h-old barley seedlings at 1.5 cm away from the seed. The culture dishes that contained the barley seedlings were then incubated at 25 °C with a 14-h photoperiod for 24–120 h. The inoculated roots were extracted using 10 volumes of 80% methanol, and the extracts were analyzed by HPLC, as follows: column, Cosmosil 5C
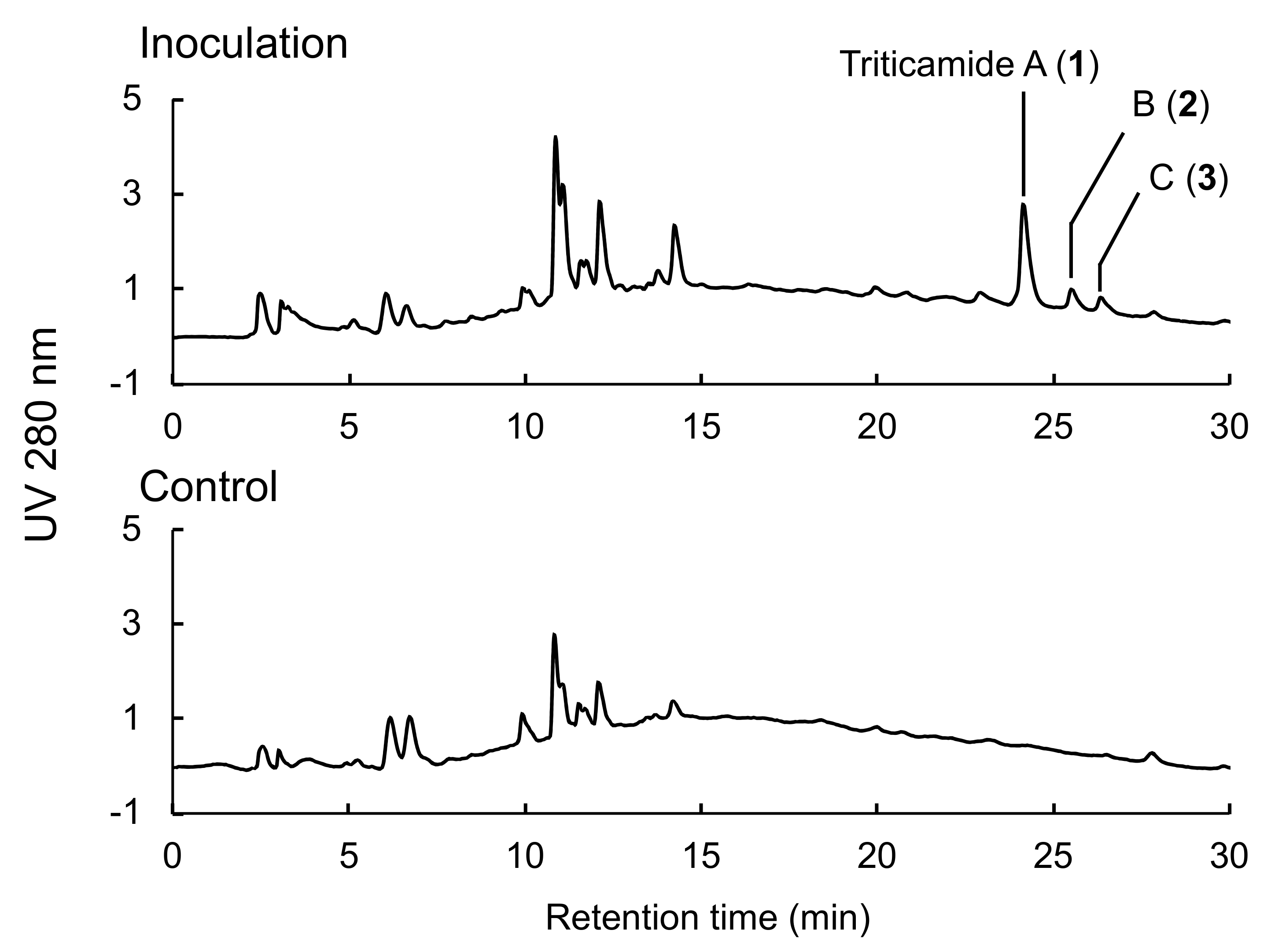
18-AR-II 4.6 I.D. × 150 mm (Nacalai Tesque, Kyoto, Japan); gradient, 5–70% B/(A+B) within 30 min, A: 0.1% trifluoroacetic acid aq., B: acetonitrile; flow rate, 0.8 mL/min; column temperature, 40 °C; detection, 280 nm. Compounds
1,
2, and
3 were eluted at 24.2, 25.5, and 26.5 min, respectively.
To inoculate barley and wheat leaves, droplets (5 μL) of the B. sorokiniana conidial suspension (5 × 105 conidia/mL) were placed on the leaves of 3-week-old seedlings at 1.0-cm intervals. The inoculated seedlings were then kept in a moist air-tight bag for 24 h, removed from the bag and further incubated at 25 °C with a 14-h photoperiod for 48 h, and then extracted using 10 volumes of 80% methanol. The extracts were analyzed by HPLC.
4.4. Purification and Identification of 3
The roots (1.5 kg) of barley ‘Shunrei’ seedlings were inoculated with F. culmorum and incubated at 23 °C with a 14-h photoperiod. After 72 h, the root metabolites were extracted in 80% methanol for 24 h, and the obtained extract was evaporated in vacuo. The resulting residue was subjected to ODS column chromatography (Cosmosil 75C18-PREP; Nacalai Tesque, Kyoto, Japan), using 20%, 40%, 50%, 60%, 70%, and 80% methanol. Because 3 was eluted in both the 50% and 60% methanol fractions, the two fractions were combined, evaporated to dryness, and subjected to silica gel column chromatography, using mixtures of acetone and hexane (20%, 30%, 40%, 50%, and 60% acetone; 1.0 L each). Because compound 3 was detected in both the 30 and 40% acetone fractions, the two fractions were concentrated, and the resulting residue (35.8 mg) was dissolved in methanol and then subjected to preparative HPLC. Conditions were as follows: column, Cosmosil 5C18-AR-II 10 mm × 250 mm; solvents, water (A) and acetonitrile (B); elution, 40% B/(A+B); flow rate, 7.0 mL/min, detection, 280 nm; column temperature, 40 °C. Compound 3 was eluted at 31.1 min.
Compound
3 (
N-cinnamoyl-(1
H-indol-3-yl)methylamine) 2.6 mg, HR MS (positive ESI):
m/z 277.1331 [M+H]
+ (calcd. for C
18H
17ON
2,
m/z 277.1341); UV-Vis (acetonitrile-water containing 0.1% formic acid): λ
max 220 and 280 nm;
1H- and
13C-NMR data are shown in
Table 1.
4.5. Synthesis of 3
N-Hydroxysuccinimide ester of cinnamic acid was synthesized from cinnamic acid according to the methods described by Stöckigt and Zenk [
30], and
N-cinnamoyl-(1
H-indol-3-yl)methylamine was synthesized from
N-hydroxysuccinimide ester of cinnamic acid and IMA (Sigma-Aldrich, St. Louis, MO, USA) according to the methods described by Negrel and Smith [
31]. IMA (73 mg, 1.0 mmoL) was dissolved in distilled water (10 mL), and the pH of the solution was adjusted to 8.0 using NaHCO
3. Meanwhile,
N-hydroxysuccinimide ester of cinnamic acid (123 mg, 0.5 mmoL) was dissolved in acetone (10 mL) and mixed with IMA solution at room temperature for 24 h. The resulting solution was then acidified using acetic acid (1.5 mL), concentrated by removing the acetone via evaporation, and then extracted three times using 20 mL ethyl acetate. The ethyl acetate layer was dried over Na
2SO
4 overnight and concentrated to dryness, and the resulting residue was subjected to silica gel column chromatography (Daisogel IR-60-63-210, 80 g; Osaka Soda, Osaka, Japan), using a 3:7 (
v:
v) mixtures of acetone and hexane and a fraction volume of 5 mL. The fractions that contained
3 were combined and concentrated.
Compound 3 (119 mg, yield 86.4%). ESI-MS: m/z: 277.1 [M+H]+. NMR data were identical with the compound isolated from barley roots.
4.6. Analyses of Phenylamides and Amines
Plant materials were immersed in 80% methanol for 24 h, and the resulting extracts were subjected to LC-MS/MS analysis with MRM, according to the methods described by Morimoto et al. [
25] and Ube et al. [
11], using the following LC conditions: column, Acquity UPLC BEH C18 column 2.1 × 50 mm (1.7 μm; Waters); flow rate, 0.2 mL/min; column temperature, 40 °C; solvents, 0.1% formic acid in water (A) and 0.1% formic acid in acetonitrile (B); gradient, 5–70% A/(A + B) within 10 min. The MRM conditions were optimized using authentic compounds.
4.7. BLAST Analysis and Phylogenetic Analysis
Blast search was performed using rice THT1/2 and TBT1/2 protein sequences as queries on the EnsemblPlants database (
http://plants.ensembl.org/Hordeum_vulgare/Info/Index?db=core). In the blast analysis, HvTHT1-9 showed more than 65% of identities % for OsTHT1/2 or TBT1/2. The HvTHT sequences were obtained from the EnsemblPlants database, whereas the sequences of BAHD acyltransferases from other plant species were extracted from GenBank (
https://www.ncbi.nlm.nih.gov/genbank/). The amino acid sequences were aligned using ClustalW, and then a dendrogram was generated using the neighbor-joining method with
p-distance in MEGA 7 (
https://www.megasoftware.net/). The robustness of the tree branches was assessed using bootstrap analysis with 1000 replicates.
Blast search was also performed using HvTHT8 protein sequences as query on GenBank and IPK Rye Blast Server (
https://webblast.ipk-gatersleben.de/ryeselect/). The alignment and dendrogram were generated though the methods described above.
4.8. RNA and DNA Extraction and qRT-PCR Analysis
Total RNA was extracted from barley ‘Shunrei’ tissues using ISOGEN II (Nippon Gene, Tokyo, Japan) and reverse-transcribed using the PrimeScript RT Reagent Kit with gDNA Eraser (Takara Bio, Kusatsu, Japan). Using total cDNA as a template, qRT-PCR was then performed using SYBR Green Realtime PCR Master Mix (Toyobo, Osaka, Japan) and gene-specific primers (
Table S2). The transcript levels of genes of interest were normalized to the expression of the ADP-ribosylation factor-like protein (ADP) gene [
32], and relative expression (log2) was calculated by subtracting the Cq values of genes of interest from those of the ADP gene. Fold changes were calculated as 2
log2, where
log2 represents relative expression.
Genomic DNA was extracted from barley ‘Shunrei’ and ‘Morex’ tissues by the modified cetyltrimethylammonium bromide method of Murray and Thompson [
33] and stored at −20 °C until use.
4.9. Molecular Cloning of Candidate Genes
The cDNA and
pCold GST vectors (
pGST, Takara Bio) were amplified using
HvTHT2,
HvTHT8, and linearized
pGST primers, respectively (
Table S2), Prime STAR MAX (Takara Bio), and the following PCR conditions: initial denaturation at 98 °C for 30 s, followed by 35 cycles of 98 °C for 10 s, 60 °C for 10 s, and 72 °C for 10 or 15 s. PCR products of the desired size were gel-purified (Promega, Wisconsin, USA) and inserted into the linearized
pCold GST using the In-Fusion cloning kit (Takara Bio). The resulting plasmids (
HvTHT2/
pGST and
HvTHT8/
pGST) were transformed into
E. coli DH5, and then clones were individually purified using a Miniprep kit from Qiagen (Venlo, Netherlands) and, finally, sequenced. The sequences of the inserted region in the
pGST and the PCR products of HvTHT7/8 were verified by DNA sequencing with Applied Bio Systems 3500xl Genetic Analyzer (Thermo Fisher Scientific).
To confirm the genomic sequence of HvTHT7/8, the genomic DNA of ‘Shunrei’ and ‘Morex’ were amplified using the HvTHT7 primers and the PCR products were purified with the same method described above. The sequence of the genomic HvTHT7/8 fragment1 were sequenced.
Meanwhile, the
HvTHT7/
pGST plasmid was generated by site-directed mutation of the
HvTHT8/
pGST plasmid. Briefly, the
HvTHT8/
pGST plasmid was amplified using mutagenic primers (
HvTHT7/
pGST;
Table S2) and the following PCR conditions: 98 °C for 10 s, followed by 30 cycles of 65 °C for 10 s, and 72 °C for 30 s. The PCR product was transformed into
E. coli DH5, and the
HvTHT7/
pGST plasmid was obtained though the methods described above.
4.10. Recombinant Protein Expression and Enzyme Assays
The HvTHT2/pGST, HvTHT7/pGST, HvTHT8/pGST, and pCold-GST plasmids were transformed into E. coli BL21 (Takara Bio) according to the manufacturer’s instructions. The resulting transformants were grown on LB medium (per liter: 10 g yeast extract, 10 g peptone, 5 g sodium chloride, and 15 g agar) that contained 100 ppm ampicillin and then selected using colony PCR. Positive colonies were grown in LB liquid medium with the antibiotics, incubated at 37 °C for 5–6 h (OD600 0.5–0.6), maintained at 15 °C for 30 min, mixed with IPTG (final conc, 500 µM), maintained at 15 °C for 24 h, and centrifuged at 5000× g for 10 min. The resulting centrifugate (pelleted cells) was resuspended in extraction buffer (20 mM Na-Pi buffer pH 8.0, 10% glycerol), sonicated, centrifuged at 15,000× g for 10 min, and then subjected to TALON metal affinity resin immobilization (Takara Bio). The His-GST-tagged proteins were then purified according to the manufacturer’s instructions, and imidazole was removed from the purified fraction using a PD-10 column (GE Healthcare, Chicago, IL, USA).
To determine the kinetic parameters of HvTHT7 and HvTHT8, enzyme reactions (50 μL) were performed at 35 °C in 100 mM Tris-HCl buffer (pH 8.0) that contained 5 μL of diluted enzyme solution and different concentrations of acyl donor and acyl acceptor. To determine the kinetic parameters for the acyl donors, the reactions were performed using Cin-, Cou-, and Fer-CoAs (1–200 μM), with tryptamine (1 mM) as the acyl acceptor, and to determine the kinetic parameters for the acyl acceptors, the reactions were performed using tryptamine, serotonin, 8-oxotryptamine, and IMA (10–2000 μM), with Cin-CoA (200 μM) as the acyl donor. To determine the kinetic parameters of HvTHT2 for the acyl acceptors, reactions were performed using tryptamine, serotonin, and tyramine (10–2000 μM), with feruloyl-CoA (200 μM) as the acyl donor. Enzyme concentration and reaction time were adjusted to ensure that the reaction proceeded linearly, and after an incubation at 35 °C for 30 min, the reactions were stopped by adding 5 µL 6N HCl and 45 µL methanol. Finally, the reaction mixtures were subjected to LC-MS/MS analysis.
4.11. Administration of Deuterium-Labeled Compounds
Both [
2H
5]-CinTry and [
2H
5]-
2 were synthesized from [
2H
5-phenyl] cinnamic acid and unlabeled corresponding amines using the same procedure that was used to synthesize unlabeled compound
3. [
2H
5-phenyl]-Cinnamic acid was a stock in the laboratory of Natural Product Chemistry, Faculty of Agriculture, Tottori University (Tottori, Japan). In the feeding experiments, barley roots were inoculated with
F. culmorum, and after a 48-h incubation, the roots were immersed in 1 mM solutions (5 mL) of labeled [
2H
5]-CinTry and [
2H
5]-
2 in 50-mL conical tubes. The treated roots were incubated for 24 h under the growth conditions and extracted using methanol. The extracts were analyzed by LC-MS/MS in MRM methods (
Table S3). MRM method of deuterium labeled-phenylamides was developed according to Li et al. [
34]
4.12. Anti-Microbial Activity Assay
The anti-microbial activities of
3 were evaluated using the methods described by Ube et al. [
11]. Conidial-germination, germ tube-elongation, and growth-inhibition assays were performed using 10, 30, 100, 300, and 1000 μM solutions of
3. Results are shown as the mean ± standard deviation. The significance of differences was estimated by analysis of variance using Tukey-Kramer tests. Findings of
p < 0.05 were considered significant. Tukey-Kramer tests were performed by using Microsoft Excel software.