Dynamics of Plant Metabolism during Cold Acclimation
Abstract
1. Introduction
2. Perception of Low Temperature
3. Tight Regulation of Photosynthesis and Carbohydrate Metabolism Prevents ROS Generation under Low Temperature
4. Carbohydrate Dynamics of Plant Cold Acclimation
5. Subcellular Metabolic Regulation during Cold Acclimation
6. Sucrose Cycling—Stabilization of Metabolism in a Changing Environment
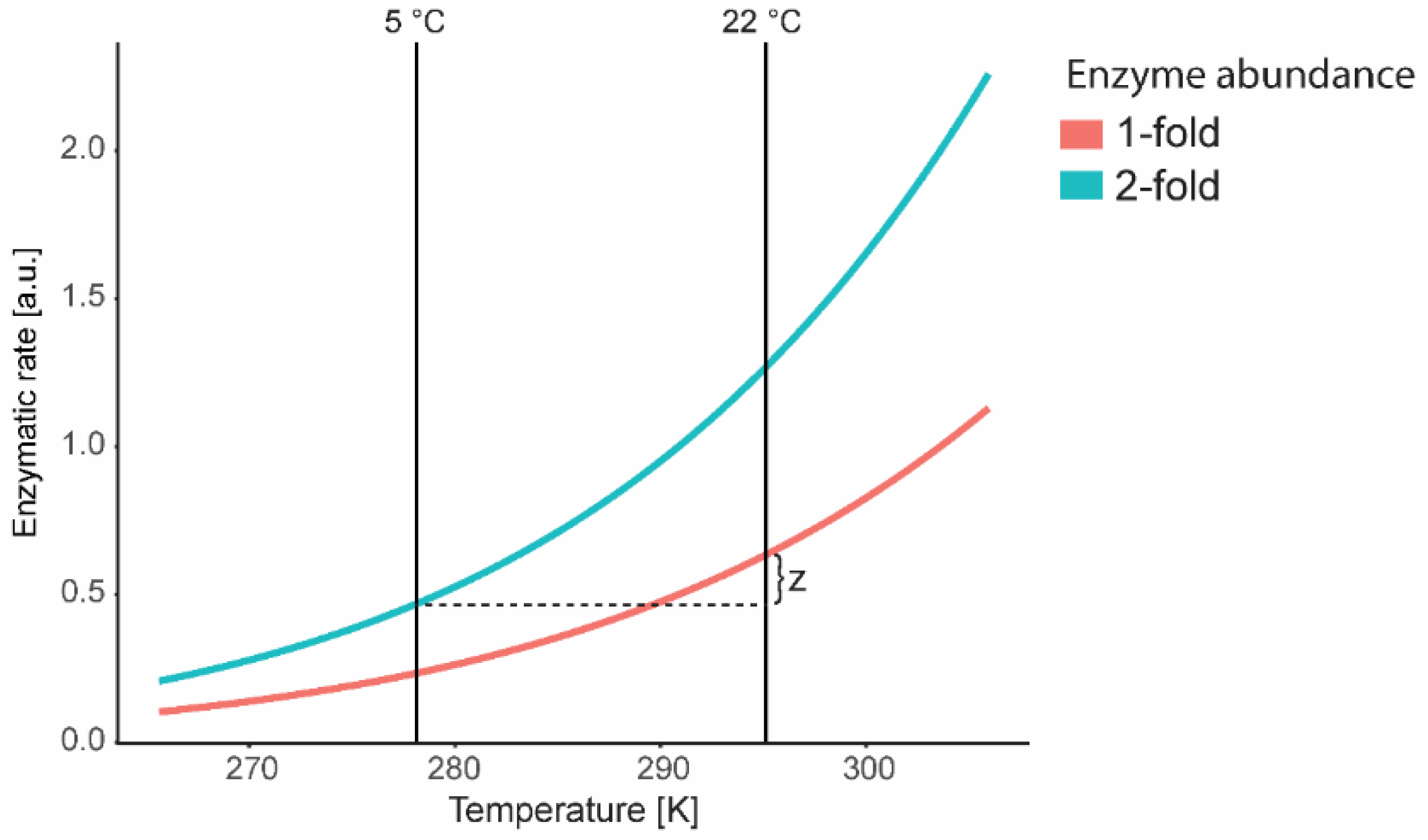
7. Mathematical Modelling at Low Temperature: Kinetics and Thermodynamic Constraints
8. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Herrmann, H.A.; Schwartz, J.-M.; Johnson, G.N. Metabolic acclimation—A key to enhancing photosynthesis in changing environments? J. Exp. Bot. 2019, 70, 3043–3056. [Google Scholar] [CrossRef]
- Hoffmann, M.H. Biogeography of Arabidopsis thaliana (L.) Heynh. (Brassicaceae). J. Biogeogr. 2002, 29, 125–134. [Google Scholar] [CrossRef]
- Hurry, V. Metabolic reprogramming in response to cold stress is like real estate, it’s all about location. Plant Cell Environ. 2017, 40, 599–601. [Google Scholar] [CrossRef] [PubMed]
- Huner, N.P.A.; Öquist, G.; Sarhan, F. Energy balance and acclimation to light and cold. Trends Plant Sci. 1998, 3, 224–230. [Google Scholar] [CrossRef]
- Scarth, W.G.; Levitt, J. The frost-hardening mechanism of plant cells. Plant Physiol. 1937, 12, 51–78. [Google Scholar] [CrossRef] [PubMed]
- Unterberger, C.; Brunner, L.; Nabernegg, S.; Steininger, K.W.; Steiner, A.K.; Stabentheiner, E.; Monschein, S.; Truhetz, H. Spring frost risk for regional apple production under a warmer climate. PLoS ONE 2018, 13, e0200201. [Google Scholar] [CrossRef]
- Horton, M.W.; Willems, G.; Sasaki, E.; Koornneef, M.; Nordborg, M. The genetic architecture of freezing tolerance varies across the range of Arabidopsis thaliana. Plant Cell Environ. 2016, 39, 2570–2579. [Google Scholar] [CrossRef]
- Ristic, Z.; Ashworth, E.N. Changes in leaf ultrastructure and carbohydrates in Arabidopsis thaliana L. (Heyn) cv. Columbia during rapid cold-acclimation. Protoplasma 1993, 172, 111–123. [Google Scholar] [CrossRef]
- Zhen, Y.; Ungerer, M.C. Clinal variation in freezing tolerance among natural accessions of Arabidopsis thaliana. New Phytol. 2008, 177, 419–427. [Google Scholar] [CrossRef]
- Korn, M.; Gärtner, T.; Erban, A.; Kopka, J.; Selbig, J.; Hincha, D.K. Predicting Arabidopsis freezing tolerance and heterosis in freezing tolerance from metabolite composition. Mol. Plant 2010, 3, 224–235. [Google Scholar] [CrossRef]
- Liu, Y.; Dang, P.; Liu, L.; He, C. Cold acclimation by the CBF–COR pathway in a changing climate: Lessons from Arabidopsis thaliana. Plant Cell Rep. 2019, 38, 511–519. [Google Scholar] [CrossRef] [PubMed]
- Hannah, M.A.; Wiese, D.; Freund, S.; Fiehn, O.; Heyer, A.G.; Hincha, D.K. Natural genetic variation of freezing tolerance in Arabidopsis. Plant Physiol. 2006, 142, 98–112. [Google Scholar] [CrossRef] [PubMed]
- Zuther, E.; Lee, Y.P.; Erban, A.; Kopka, J.; Hincha, D.K. Natural variation in freezing tolerance and cold acclimation response in Arabidopsis thaliana and related species. In Survival Strategies in Extreme Cold and Desiccation: Adaptation Mechanisms and Their Applications; Iwaya-Inoue, M., Sakurai, M., Uemura, M., Eds.; Springer: Singapore, 2018; pp. 81–98. [Google Scholar] [CrossRef]
- Litvan, G.G. Mechanism of cryoinjury in biological-systems. Cryobiology 1972, 9, 182–191. [Google Scholar] [CrossRef][Green Version]
- Hincha, D.K.; Zuther, E.; Heyer, A.G. The preservation of liposomes by raffinose family oligosaccharides during drying is mediated by effects on fusion and lipid phase transitions. Biochim. Biophys. Acta Biomembr. 2003, 1612, 172–177. [Google Scholar] [CrossRef]
- Thomashow, M.F. Molecular genetics of cold acclimation in higher plants. Adv. Genet. 1990, 28, 99–131. [Google Scholar]
- Thomashow, M.F. So what’s new in the field of plant cold acclimation? Lots! Plant Physiol. 2001, 125, 89–93. [Google Scholar] [CrossRef] [PubMed]
- Vaultier, M.-N.; Cantrel, C.; Vergnolle, C.; Justin, A.-M.; Demandre, C.; Benhassaine-Kesri, G.; Çiçek, D.; Zachowski, A.; Ruelland, E. Desaturase mutants reveal that membrane rigidification acts as a cold perception mechanism upstream of the diacylglycerol kinase pathway in Arabidopsis cells. FEBS Lett. 2006, 580, 4218–4223. [Google Scholar] [CrossRef]
- Guo, X.; Liu, D.; Chong, K. Cold signaling in plants: Insights into mechanisms and regulation. J. Integr. Plant Biol. 2018, 60, 745–756. [Google Scholar] [CrossRef]
- Sangwan, V.; Foulds, I.; Singh, J.; Dhindsa, R.S. Cold-activation of Brassica napus BN115 promoter is mediated by structural changes in membranes and cytoskeleton, and requires Ca2+ influx. Plant J. 2001, 27, 1–12. [Google Scholar] [CrossRef]
- Ding, Y.; Shi, Y.; Yang, S. Advances and challenges in uncovering cold tolerance regulatory mechanisms in plants. New Phytol. 2019, 222, 1690–1704. [Google Scholar] [CrossRef]
- Liu, Z.; Jia, Y.; Ding, Y.; Shi, Y.; Li, Z.; Guo, Y.; Gong, Z.; Yang, S. Plasma membrane CRPK1-mediated phosphorylation of 14-3-3 proteins induces their nuclear import to fine-tune CBF signaling during cold response. Mol. Cell 2017, 66, 117–128. [Google Scholar] [CrossRef] [PubMed]
- Tan, W.-J.; Yang, Y.-C.; Zhou, Y.; Huang, L.-P.; Xu, L.; Chen, Q.-F.; Yu, L.-J.; Xiao, S. DIACYLGLYCEROL ACYLTRANSFERASE and DIACYLGLYCEROL KINASE modulate triacylglycerol and phosphatidic acid production in the plant response to freezing stress. Plant Physiol. 2018, 177, 1303–1318. [Google Scholar] [CrossRef] [PubMed]
- Thomashow, M.F. Molecular basis of plant cold acclimation: Insights gained from studying the CBF cold response pathway. Plant Physiol. 2010, 154, 571–577. [Google Scholar] [CrossRef] [PubMed]
- Gilmour, S.J.; Zarka, D.G.; Stockinger, E.J.; Salazar, M.P.; Houghton, J.M.; Thomashow, M.F. Low temperature regulation of the Arabidopsis CBF family of AP2 transcriptional activators as an early step in cold-induced COR gene expression. Plant J. 1998, 16, 433–442. [Google Scholar] [CrossRef]
- Zhao, C.; Zhang, Z.; Xie, S.; Si, T.; Li, Y.; Zhu, J.-K. Mutational evidence for the critical role of CBF genes in cold acclimation in Arabidopsis. Plant Physiol. 2016, 171, 2744–2759. [Google Scholar] [CrossRef]
- Fowler, S.; Thomashow, M.F. Arabidopsis transcriptome profiling indicates that multiple regulatory pathways are activated during cold acclimation in addition to the CBF cold response pathway. Plant Cell 2002, 14, 1675–1690. [Google Scholar] [CrossRef]
- Jaglo-Ottosen, K.R.; Gilmour, S.J.; Zarka, D.G.; Schabenberger, O.; Thomashow, M.F. Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science 1998, 280, 104–106. [Google Scholar] [CrossRef]
- Gehan, M.A.; Park, S.; Gilmour, S.J.; An, C.; Lee, C.M.; Thomashow, M.F. Natural variation in the C-repeat binding factor cold response pathway correlates with local adaptation of Arabidopsis ecotypes. Plant J. Cell Mol. Biol. 2015, 84, 682–693. [Google Scholar] [CrossRef]
- Kurepin, L.V.; Dahal, K.P.; Savitch, L.V.; Singh, J.; Bode, R.; Ivanov, A.G.; Hurry, V.; Huner, N.P. Role of CBFs as integrators of chloroplast redox, phytochrome and plant hormone signaling during cold acclimation. Int. J. Mol. Sci. 2013, 14, 12729–12763. [Google Scholar] [CrossRef]
- Shi, Y.; Ding, Y.; Yang, S. Cold signal transduction and its interplay with phytohormones during cold acclimation. Plant Cell Physiol. 2015, 56, 7–15. [Google Scholar] [CrossRef]
- Shi, Y.; Ding, Y.; Yang, S. Molecular regulation of CBF signaling in cold acclimation. Trends Plant Sci. 2018, 23, 623–637. [Google Scholar] [CrossRef]
- Rekarte-Cowie, I.; Ebshish, O.S.; Mohamed, K.S.; Pearce, R.S. Sucrose helps regulate cold acclimation of Arabidopsis thaliana. J. Exp. Bot. 2008, 59, 4205–4217. [Google Scholar] [CrossRef] [PubMed]
- Taji, T.; Ohsumi, C.; Iuchi, S.; Seki, M.; Kasuga, M.; Kobayashi, M.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Important roles of drought-and cold-inducible genes for galactinol synthase in stress tolerance in Arabidopsis thaliana. Plant J. 2002, 29, 417–426. [Google Scholar] [CrossRef] [PubMed]
- Claeys, H.; de Bodt, S.; Inze, D. Gibberellins and DELLAs: Central nodes in growth regulatory networks. Trends Plant Sci. 2014, 19, 231–239. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Gong, F.; Cheminant, S.; Alioua, M.; Hedden, P.; Genschik, P. The cold-inducible CBF1 factor-dependent signaling pathway modulates the accumulation of the growth-repressing DELLA proteins via its effect on gibberellin metabolism. Plant Cell 2008, 20, 2117–2129. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; van den Ende, W.; Rolland, F. Sucrose induction of anthocyanin biosynthesis is mediated by DELLA. Mol. Plant 2014, 7, 570–572. [Google Scholar] [CrossRef] [PubMed]
- Schulz, E.; Tohge, T.; Zuther, E.; Fernie, A.R.; Hincha, D.K. Flavonoids are determinants of freezing tolerance and cold acclimation in Arabidopsis thaliana. Sci. Rep. 2016, 6, 34027. [Google Scholar] [CrossRef]
- Kosová, K.; Vítámvás, P.; Urban, M.O.; Prášil, I.T.; Renaut, J. Plant abiotic stress proteomics: The major factors determining alterations in cellular proteome. Front. Plant Sci. 2018, 9, 122. [Google Scholar] [CrossRef]
- Maier, T.; Güell, M.; Serrano, L. Correlation of mRNA and protein in complex biological samples. FEBS Lett. 2009, 583, 3966–3973. [Google Scholar] [CrossRef]
- Janmohammadi, M.; Zolla, L.; Rinalducci, S. Low temperature tolerance in plants: Changes at the protein level. Phytochemistry 2015, 117, 76–89. [Google Scholar] [CrossRef]
- Hannah, M.A.; Heyer, A.G.; Hincha, D.K. A global survey of gene regulation during cold acclimation in Arabidopsis thaliana. PLoS Genet. 2005, 1, e26. [Google Scholar] [CrossRef] [PubMed]
- Nagler, M.; Nukarinen, E.; Weckwerth, W.; Nägele, T. Integrative molecular profiling indicates a central role of transitory starch breakdown in establishing a stable C/N homeostasis during cold acclimation in two natural accessions of Arabidopsis thaliana. BMC Plant Biol. 2015, 15, 284. [Google Scholar] [CrossRef] [PubMed]
- Fürtauer, L.; Pschenitschnigg, A.; Scharkosi, H.; Weckwerth, W.; Nägele, T. Combined multivariate analysis and machine learning reveals a predictive module of metabolic stress response in Arabidopsis thaliana. Mol. Omics 2018, 14, 437–449. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Song, X.; Shi, K.; Xia, X.; Zhou, Y.; Yu, J. Changes in electron transport, superoxide dismutase and ascorbate peroxidase isoenzymes in chloroplasts and mitochondria of cucumber leaves as influenced by chilling. Photosynthetica 2008, 46, 581. [Google Scholar] [CrossRef]
- Nägele, T.; Kandel, B.A.; Frana, S.; Meißner, M.; Heyer, A.G. A systems biology approach for the analysis of carbohydrate dynamics during acclimation to low temperature in Arabidopsis thaliana. FEBS J. 2011, 278, 506–518. [Google Scholar] [CrossRef] [PubMed]
- Crafts-Brandner, S.J.; van de Loo, F.J.; Salvucci, M.E. The two forms of ribulose-1, 5-bisphosphate carboxylase/oxygenase activase differ in sensitivity to elevated temperature. Plant Physiol. 1997, 114, 439–444. [Google Scholar] [CrossRef] [PubMed]
- Yamori, W.; Suzuki, K.; Noguchi, K.; Nakai, M.; Terashima, I. Effects of Rubisco kinetics and Rubisco activation state on the temperature dependence of the photosynthetic rate in spinach leaves from contrasting growth temperatures. Plant Cell Environ. 2006, 29, 1659–1670. [Google Scholar] [CrossRef]
- Hashida, S.-N.; Kawai-Yamada, M. Inter-organelle NAD metabolism underpinning light responsive NADP dynamics in plants. Front. Plant Sci. 2019, 10, 960. [Google Scholar] [CrossRef]
- Poolman, M.G.; Fell, D.A.; Thomas, S. Modelling photosynthesis and its control. J. Exp. Bot. 2000, 51, 319–328. [Google Scholar] [CrossRef]
- Flügge, U.I.; Heldt, H.W. Metabolite translocators of the chloroplast envelope. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1991, 42, 129–144. [Google Scholar] [CrossRef]
- Schneider, A.; Häusler, R.E.; Kolukisaoglu, Ü.; Kunze, R.; van der Graaff, E.; Schwacke, R.; Catoni, E.; Desimone, M.; Flügge, U.I. An Arabidopsis thaliana knock-out mutant of the chloroplast triose phosphate/phosphate translocator is severely compromised only when starch synthesis, but not starch mobilisation is abolished. Plant J. 2002, 32, 685–699. [Google Scholar] [CrossRef] [PubMed]
- Strand, A.; Foyer, C.; Gustafsson, P.; Gardeström, P.; Hurry, V. Altering flux through the sucrose biosynthesis pathway in transgenic Arabidopsis thaliana modifies photosynthetic acclimation at low temperatures and the development of freezing tolerance. Plant Cell Environ. 2003, 26, 523–535. [Google Scholar] [CrossRef]
- Nägele, T.; Stutz, S.; Hörmiller, I.I.; Heyer, A.G. Identification of a metabolic bottleneck for cold acclimation in Arabidopsis thaliana. Plant J. Cell Mol. Biol. 2012, 72, 102–114. [Google Scholar] [CrossRef] [PubMed]
- Stitt, M. Limitation of photosynthesis by carbon metabolism. Plant Physiol. 1986, 81, 1115–1122. [Google Scholar] [CrossRef]
- Leegood, R.C.; Furbank, R.T. Stimulation of photosynthesis by 2% oxygen at low temperatures is restored by phosphate. Planta 1986, 168, 84–93. [Google Scholar] [CrossRef]
- Bagnall, D.J.; King, R.W.; Farquhar, G.D. Temperature-dependent feedback inhibition of photosynthesis in peanut. Planta 1988, 175, 348–354. [Google Scholar] [CrossRef]
- Lundmark, M.; Cavaco, A.M.; Trevanion, S.; Hurry, V. Carbon partitioning and export in transgenic Arabidopsis thaliana with altered capacity for sucrose synthesis grown at low temperature: A role for metabolite transporters. Plant Cell Environ. 2006, 29, 1703–1714. [Google Scholar] [CrossRef]
- Dreyer, A.; Dietz, K.J. Reactive oxygen species and the redox-regulatory network in cold stress acclimation. Antioxidants 2018, 7, 169. [Google Scholar] [CrossRef]
- Heiber, I.; Cai, W.; Baier, M. Linking chloroplast antioxidant defense to carbohydrate availability: The transcript abundance of stromal ascorbate peroxidase is sugar-controlled via ascorbate biosynthesis. Mol. Plant 2014, 7, 58–70. [Google Scholar] [CrossRef]
- Yamori, W.; Hikosaka, K.; Way, D.A. Temperature response of photosynthesis in C3, C4, and CAM plants: Temperature acclimation and temperature adaptation. Photosynth. Res. 2014, 119, 101–117. [Google Scholar] [CrossRef]
- Kaplan, F.; Kopka, J.; Haskell, D.W.; Zhao, W.; Schiller, K.C.; Gatzke, N.; Sung, D.Y.; Guy, C.L. Exploring the temperature-stress metabolome of Arabidopsis. Plant Physiol. 2004, 136, 4159–4168. [Google Scholar] [CrossRef] [PubMed]
- Winkel-Shirley, B. Biosynthesis of flavonoids and effects of stress. Curr. Opin. Plant Biol. 2002, 5, 218–223. [Google Scholar] [CrossRef]
- Graf, A.; Schlereth, A.; Stitt, M.; Smith, A.M. Circadian control of carbohydrate availability for growth in Arabidopsis plants at night. Proc. Natl. Acad. Sci. USA 2010, 107, 9458–9463. [Google Scholar] [CrossRef] [PubMed]
- Pommerrenig, B.; Ludewig, F.; Cvetkovic, J.; Trentmann, O.; Klemens, P.A.W.; Neuhaus, H.E. In concert: Orchestrated changes in carbohydrate homeostasis are critical for plant abiotic stress tolerance. Plant Cell Physiol. 2018, 59, 1290–1299. [Google Scholar] [CrossRef] [PubMed]
- Miura, K.; Furumoto, T. Cold signaling and cold response in plants. Int. J. Mol. Sci. 2013, 14, 5312–5337. [Google Scholar] [CrossRef] [PubMed]
- Haydon, M.J.; Mielczarek, O.; Robertson, F.C.; Hubbard, K.E.; Webb, A.A.R. Photosynthetic entrainment of the Arabidopsis thaliana circadian clock. Nature 2013, 502, 689–692. [Google Scholar] [CrossRef]
- Wanner, L.A.; Junttila, O. Cold-induced freezing tolerance in Arabidopsis. Plant Physiol. 1999, 120, 391–400. [Google Scholar] [CrossRef]
- Soitamo, A.J.; Piippo, M.; Allahverdiyeva, Y.; Battchikova, N.; Aro, E.M. Light has a specific role in modulating Arabidopsis gene expression at low temperature. BMC Plant Biol. 2008, 8, 13. [Google Scholar] [CrossRef]
- Strauss, G.; Hauser, H. Stabilization of lipid bilayer vesicles by sucrose during freezing. Proc. Natl. Acad. Sci. USA 1986, 83, 2422–2426. [Google Scholar] [CrossRef]
- Sami, F.; Yusuf, M.; Faizan, M.; Faraz, A.; Hayat, S. Role of sugars under abiotic stress. Plant Physiol. Biochem. 2016, 109, 54–61. [Google Scholar] [CrossRef]
- Tarkowski, L.P.; van den Ende, W. Cold tolerance triggered by soluble sugars: A multifaceted countermeasure. Front. Plant Sci. 2015, 6, 203. [Google Scholar] [CrossRef] [PubMed]
- Valluru, R.; Lammens, W.; Claupein, W.; van den Ende, W. Freezing tolerance by vesicle-mediated fructan transport. Trends Plant Sci. 2008, 13, 409–414. [Google Scholar] [CrossRef] [PubMed]
- Zuther, E.; Büchel, K.; Hundertmark, M.; Stitt, M.; Hincha, D.K.; Heyer, A.G. The role of raffinose in the cold acclimation response of Arabidopsis thaliana. FEBS Lett. 2004, 576, 169–173. [Google Scholar] [CrossRef] [PubMed]
- Nishizawa, A.; Yabuta, Y.; Shigeoka, S. Galactinol and raffinose constitute a novel function to protect plants from oxidative damage. Plant Physiol. 2008, 147, 1251–1263. [Google Scholar] [CrossRef]
- Schneider, T.; Keller, F. Raffinose in chloroplasts is synthesized in the cytosol and transported across the chloroplast envelope. Plant Cell Physiol. 2009, 50, 2174–2182. [Google Scholar] [CrossRef]
- Knaupp, M.; Mishra, K.B.; Nedbal, L.; Heyer, A.G. Evidence for a role of raffinose in stabilizing photosystem II during freeze-thaw cycles. Planta 2011, 234, 477–486. [Google Scholar] [CrossRef]
- Allison, S.D.; Chang, B.; Randolph, T.W.; Carpenter, J.F. Hydrogen bonding between sugar and protein is responsible for inhibition of dehydration-induced protein unfolding. Arch. Biochem. Biophys. 1999, 365, 289–298. [Google Scholar] [CrossRef]
- Skryhan, K.; Gurrieri, L.; Sparla, F.; Trost, P.; Blennow, A. Redox regulation of starch metabolism. Front. Plant Sci. 2018, 9, 1344. [Google Scholar] [CrossRef]
- Tiessen, A.; Hendriks, J.H.M.; Stitt, M.; Branscheid, A.; Gibon, Y.; Farre, E.M.; Geigenberger, P. Starch synthesis in potato tubers is regulated by post-translational redox modification of ADP-glucose pyrophosphorylase: A novel regulatory mechanism linking starch synthesis to the sucrose supply. Plant Cell 2002, 14, 2191–2213. [Google Scholar] [CrossRef]
- Thalmann, M.; Santelia, D. Starch as a determinant of plant fitness under abiotic stress. New Phytol. 2017, 214, 943–951. [Google Scholar] [CrossRef]
- Sicher, R. Carbon partitioning and the impact of starch deficiency on the initial response of Arabidopsis to chilling temperatures. Plant Sci. 2011, 181, 167–176. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, F.; Guy, C.L. β-Amylase induction and the protective role of maltose during temperature shock. Plant Physiol. 2004, 135, 1674–1684. [Google Scholar] [CrossRef] [PubMed]
- Zanella, M.; Borghi, G.L.; Pirone, C.; Thalmann, M.; Pazmino, D.; Costa, A.; Santelia, D.; Trost, P.; Sparla, F. β-amylase 1 (BAM1) degrades transitory starch to sustain proline biosynthesis during drought stress. J. Exp. Bot. 2016, 67, 1819–1826. [Google Scholar] [CrossRef] [PubMed]
- Zeeman, S.C.; Thorneycroft, D.; Schupp, N.; Chapple, A.; Weck, M.; Dunstan, H.; Haldimann, P.; Bechtold, N.; Smith, A.M.; Smith, S.M. Plastidial α-glucan phosphorylase is not required for starch degradation in Arabidopsis leaves but has a role in the tolerance of abiotic stress. Plant Physiol. 2004, 135, 849–858. [Google Scholar] [CrossRef]
- Dong, S.; Beckles, D.M. Dynamic changes in the starch-sugar interconversion within plant source and sink tissues promote a better abiotic stress response. J. Plant Physiol. 2019, 234–235, 80–93. [Google Scholar] [CrossRef]
- Klotke, J.; Kopka, J.; Gatzke, N.; Heyer, A.G. Impact of soluble sugar concentrations on the acquisition of freezing tolerance in accessions of Arabidopsis thaliana with contrasting cold adaptation—Evidence for a role of raffinose in cold acclimation. Plant Cell Environ. 2004, 27, 1395–1404. [Google Scholar] [CrossRef]
- Szabados, L.; Savoure, A. Proline: A multifunctional amino acid. Trends Plant Sci. 2010, 15, 89–97. [Google Scholar] [CrossRef]
- Verslues, P.E.; Sharma, S. Proline metabolism and its implications for plant-environment interaction. Arab. Book/Am. Soc. Plant Biol. 2010, 8, e0140. [Google Scholar] [CrossRef]
- Usadel, B.; Blaesing, O.E.; Gibon, Y.; Poree, F.; Hoehne, M.; Guenter, M.; Trethewey, R.; Kamlage, B.; Poorter, H.; Stitt, M. Multilevel genomic analysis of the response of transcripts, enzyme activities and metabolites in Arabidopsis rosettes to a progressive decrease of temperature in the non-freezing range. Plant Cell Environ. 2008, 31, 518–547. [Google Scholar] [CrossRef]
- Kaplan, F.; Kopka, J.; Sung, D.Y.; Zhao, W.; Popp, M.; Porat, R.; Guy, C.L. Transcript and metabolite profiling during cold acclimation of Arabidopsis reveals an intricate relationship of cold-regulated gene expression with modifications in metabolite content. Plant J. Cell Mol. Biol. 2007, 50, 967–981. [Google Scholar] [CrossRef]
- Doerfler, H.; Lyon, D.; Nagele, T.; Sun, X.L.; Fragner, L.; Hadacek, F.; Egelhofer, V.; Weckwerth, W. Granger causality in integrated GC-MS and LC-MS metabolomics data reveals the interface of primary and secondary metabolism. Metabolomics 2013, 9, 564–574. [Google Scholar] [CrossRef] [PubMed]
- Tohge, T.; de Souza, L.P.; Fernie, A.R. Current understanding of the pathways of flavonoid biosynthesis in model and crop plants. J. Exp. Bot. 2017, 68, 4013–4028. [Google Scholar] [CrossRef] [PubMed]
- Haslam, E. Shikimic Acid: Metabolism and Metabolites; John Wiley & Sons Inc.: Hoboken, NJ, USA, 1993. [Google Scholar]
- Harborne, J. Plant polyphenols—XIV.: Characterization of flavonoid glycosides by acidic and enzymic hydrolyses. Phytochemistry 1965, 4, 107–120. [Google Scholar] [CrossRef]
- Petrussa, E.; Braidot, E.; Zancani, M.; Peresson, C.; Bertolini, A.; Patui, S.; Vianello, A. Plant flavonoids-biosynthesis, transport and involvement in stress responses. Int. J. Mol. Sci. 2013, 14, 14950–14973. [Google Scholar] [CrossRef]
- Schulz, E.; Tohge, T.; Zuther, E.; Fernie, A.R.; Hincha, D.K. Natural variation in flavonol and anthocyanin metabolism during cold acclimation in Arabidopsis thaliana accessions. Plant Cell Environ. 2015, 38, 1658–1672. [Google Scholar] [CrossRef]
- Linka, N.; Weber, A. Intracellular metabolite transporters in plants. Mol. Plant 2010, 3, 21–53. [Google Scholar] [CrossRef]
- Martinoia, E.; Maeshima, M.; Neuhaus, H.E. Vacuolar transporters and their essential role in plant metabolism. J. Exp. Bot. 2007, 58, 83–102. [Google Scholar] [CrossRef]
- Wormit, A.; Trentmann, O.; Feifer, I.; Lohr, C.; Tjaden, J.; Meyer, S.; Schmidt, U.; Martinoia, E.; Neuhaus, H.E. Molecular identification and physiological characterization of a novel monosaccharide transporter from Arabidopsis involved in vacuolar sugar transport. Plant Cell 2006, 18, 3476–3490. [Google Scholar] [CrossRef]
- Jung, B.; Ludewig, F.; Schulz, A.; Meissner, G.; Wostefeld, N.; Flugge, U.I.; Pommerrenig, B.; Wirsching, P.; Sauer, N.; Koch, W.; et al. Identification of the transporter responsible for sucrose accumulation in sugar beet taproots. Nat. Plants 2015, 1, 14001. [Google Scholar] [CrossRef]
- Klemens, P.A.W.; Patzke, K.; Deitmer, J.; Spinner, L.; le Hir, R.; Bellini, C.; Bedu, M.; Chardon, F.; Krapp, A.; Neuhaus, H.E. Overexpression of the vacuolar sugar carrier AtSWEET16 modifies germination, growth, and stress tolerance in Arabidopsis. Plant Physiol. 2013, 163, 1338–1352. [Google Scholar] [CrossRef]
- Patzke, K.; Prananingrum, P.; Klemens, P.A.W.; Trentmann, O.; Rodrigues, C.M.; Keller, I.; Fernie, A.R.; Geigenberger, P.; Bölter, B.; Lehmann, M.; et al. The plastidic sugar transporter pSuT influences flowering and affects cold responses. Plant Physiol. 2019, 179, 569–587. [Google Scholar] [CrossRef] [PubMed]
- Gerhardt, R.; Heldt, H.W. Measurement of subcellular metabolite levels in leaves by fractionation of freeze-stopped material in nonaqueous media. Plant Physiol. 1984, 75, 542–547. [Google Scholar] [CrossRef] [PubMed]
- Fürtauer, L.; Küstner, L.; Weckwerth, W.; Heyer, A.G.; Nägele, T. Resolving subcellular plant metabolism. Plant J. 2019. [Google Scholar] [CrossRef] [PubMed]
- Fürtauer, L.; Weckwerth, W.; Nägele, T. A benchtop fractionation procedure for subcellular analysis of the plant metabolome. Front. Plant Sci. 2016, 7, 1912. [Google Scholar] [CrossRef]
- Talts, P.; Pärnik, T.; Gardeström, P.; Keerberg, O. Respiratory acclimation in Arabidopsis thaliana leaves at low temperature. J. Plant Physiol. 2004, 161, 573–579. [Google Scholar] [CrossRef]
- Nägele, T.; Heyer, A.G. Approximating subcellular organisation of carbohydrate metabolism during cold acclimation in different natural accessions of Arabidopsis thaliana. New Phytol. 2013, 198, 777–787. [Google Scholar] [CrossRef]
- Hoermiller, I.I.; Naegele, T.; Augustin, H.; Stutz, S.; Weckwerth, W.; Heyer, A.G. Subcellular reprogramming of metabolism during cold acclimation in Arabidopsis thaliana. Plant Cell Environ. 2017, 40, 602–610. [Google Scholar] [CrossRef]
- Küstner, L.; Fürtauer, L.; Weckwerth, W.; Nägele, T.; Heyer, A.G. Subcellular dynamics of proteins and metabolites under abiotic stress reveal deferred response of the Arabidopsis thaliana hexokinase-1 mutant gin2-1 to high light. Plant J. 2019. [Google Scholar] [CrossRef]
- Fürtauer, L.; Nägele, T. Approximating the stabilization of cellular metabolism by compartmentalization. Theory Biosci. 2016, 135, 73–87. [Google Scholar] [CrossRef]
- Weiszmann, J.; Fürtauer, L.; Weckwerth, W.; Nägele, T. Vacuolar sucrose cleavage prevents limitation of cytosolic carbohydrate metabolism and stabilizes photosynthesis under abiotic stress. FEBS J. 2018, 285, 4082–4098. [Google Scholar] [CrossRef]
- Ruan, Y.-L. Sucrose metabolism: Gateway to diverse carbon use and sugar signaling. Annu. Rev. Plant Biol. 2014, 65, 33–67. [Google Scholar] [CrossRef] [PubMed]
- Stitt, M.; Wilke, I.; Feil, R.; Heldt, H.W. Coarse control of sucrose-phosphate synthase in leaves—Alterations of the kinetic-properties in response to the rate of photosynthesis and the accumulation of sucrose. Planta 1988, 174, 217–230. [Google Scholar] [CrossRef] [PubMed]
- Huber, J.L.A.; Huber, S.C.; Nielsen, T.H. Protein-phosphorylation as a mechanism for regulation of spinach leaf sucrose-phosphate synthase activity. Arch. Biochem. Biophys. 1989, 270, 681–690. [Google Scholar] [CrossRef]
- Huber, S.C.; Huber, J.L. Role of sucrose-phosphate synthase in sucrose metabolism in leaves. Plant Physiol. 1992, 99, 1275–1278. [Google Scholar] [CrossRef]
- Doehlert, D.C.; Huber, S.C. Spinach leaf sucrose phosphate synthase. FEBS Lett. 1983, 153, 293–297. [Google Scholar] [CrossRef]
- Tiessen, A.; Padilla-Chacon, D. Subcellular compartmentation of sugar signaling: Links among carbon cellular status, route of sucrolysis, sink-source allocation, and metabolic partitioning. Front. Plant Sci. 2013, 3, 306. [Google Scholar] [CrossRef]
- Wan, H.; Wu, L.; Yang, Y.; Zhou, G.; Ruan, Y.-L. Evolution of sucrose metabolism: The dichotomy of invertases and beyond. Trends Plant Sci. 2018, 23, 163–177. [Google Scholar] [CrossRef]
- Stein, O.; Granot, D. An overview of sucrose synthases in plants. Front. Plant Sci. 2019, 10, 95. [Google Scholar] [CrossRef]
- Nägele, T.; Henkel, S.; Hormiller, I.; Sauter, T.; Sawodny, O.; Ederer, M.; Heyer, A.G. Mathematical modeling of the central carbohydrate metabolism in Arabidopsis reveals a substantial regulatory influence of vacuolar invertase on whole plant carbon metabolism. Plant Physiol. 2010, 153, 260–272. [Google Scholar] [CrossRef]
- Geigenberger, P.; Stitt, M. A futile cycle of sucrose synthesis and degradation is involved in regulating partitioning between sucrose, starch and respiration in cotyledons of germinating Ricinus communis L. seedlings when phloem transport is inhibited. Planta 1991, 185, 81–90. [Google Scholar] [CrossRef]
- Huber, S.C. Biochemical-mechanism for regulation of sucrose accumulation in leaves during photosynthesis. Plant Physiol. 1989, 91, 656–662. [Google Scholar] [CrossRef] [PubMed]
- Henkel, S.; Nägele, T.; Hörmiller, I.; Sauter, T.; Sawodny, O.; Ederer, M.; Heyer, A.G. A systems biology approach to analyse leaf carbohydrate metabolism in Arabidopsis thaliana. EURASIP J. Bioinform. Syst. Biol. 2011, 2011, 2. [Google Scholar] [CrossRef] [PubMed]
- Brauner, K.; Stutz, S.; Paul, M.; Heyer, A.G. Measuring whole plant CO2 exchange with the environment reveals opposing effects of the gin2-1 mutation in shoots and roots of Arabidopsis thaliana. Plant Signal. Behav. 2015, 10, e973822. [Google Scholar] [CrossRef] [PubMed]
- Küstner, L.; Nägele, T.; Heyer, A.G. Mathematical modeling of diurnal patterns of carbon allocation to shoot and root in Arabidopsis thaliana. NPJ Syst. Biol. Appl. 2019, 5, 4. [Google Scholar] [CrossRef]
- Kim, J.Y.; Mahe, A.; Brangeon, J.; Prioul, J.L. A maize vacuolar invertase, IVR2, is induced by water stress. Organ/tissue specificity and diurnal modulation of expression. Plant Physiol. 2000, 124, 71–84. [Google Scholar] [CrossRef]
- Andersen, M.N.; Asch, F.; Wu, Y.; Jensen, C.R.; Naested, H.; Mogensen, V.O.; Koch, K.E. Soluble invertase expression is an early target of drought stress during the critical, abortion-sensitive phase of young ovary development in maize. Plant Physiol. 2002, 130, 591–604. [Google Scholar] [CrossRef]
- Trouverie, J.; Thevenot, C.; Rocher, J.P.; Sotta, B.; Prioul, J.L. The role of abscisic acid in the response of a specific vacuolar invertase to water stress in the adult maize leaf. J. Exp. Bot. 2003, 54, 2177–2186. [Google Scholar] [CrossRef]
- Sweetlove, L.J.; Nielsen, J.; Fernie, A.R. Engineering central metabolism—A grand challenge for plant biologists. Plant J. 2017, 90, 749–763. [Google Scholar] [CrossRef]
- Nägele, T. Linking metabolomics data to underlying metabolic regulation. Front. Mol. Biosci. 2014, 1, 22. [Google Scholar] [CrossRef]
- Ni, X.Y.; Drengstig, T.; Ruoff, P. The control of the controller: Molecular mechanisms for robust perfect adaptation and temperature compensation. Biophys. J. 2009, 97, 1244–1253. [Google Scholar] [CrossRef]
- Fürtauer, L.; Weiszmann, J.; Weckwerth, W.; Nägele, T. Mathematical modeling approaches in plant metabolomics. In Plant Metabolomics: Methods and Protocols; António, C., Ed.; Springer: New York, NY, USA, 2018; pp. 329–347. [Google Scholar] [CrossRef]
- De Oliveira Dal’Molin, C.G.; Nielsen, L.K. Plant genome-scale metabolic reconstruction and modelling. Curr. Opin. Biotechnol. 2013, 24, 271–277. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira Dal’Molin, C.G.; Nielsen, L.K. Plant genome-scale reconstruction: From single cell to multi-tissue modelling and omics analyses. Curr. Opin. Biotechnol. 2018, 49, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.G.; Lynch, J.P.; Lebauer, D.S.; Millar, A.J.; Stitt, M.; Long, S.P. Plants in silico: Why, why now and what?—An integrative platform for plant systems biology research. Plant Cell Environ. 2016, 39, 1049–1057. [Google Scholar] [CrossRef] [PubMed]
- Töpfer, N.; Kleessen, S.; Nikoloski, Z. Integration of metabolomics data into metabolic networks. Front. Plant Sci. 2015, 6, 49. [Google Scholar] [CrossRef] [PubMed]
- Steuer, R.; Gross, T.; Selbig, J.; Blasius, B. Structural kinetic modeling of metabolic networks. Proc. Natl. Acad. Sci. USA 2006, 103, 11868–11873. [Google Scholar] [CrossRef]
- Pettersson, G.; Ryde-Pettersson, U. A mathematical model of the Calvin photosynthesis cycle. Eur. J. Biochem. 1988, 175, 661–672. [Google Scholar] [CrossRef]
- Pokhilko, A.; Bou-Torrent, J.; Pulido, P.; Rodriguez-Concepcion, M.; Ebenhoh, O. Mathematical modelling of the diurnal regulation of the MEP pathway in Arabidopsis. New Phytol. 2015, 206, 1075–1085. [Google Scholar] [CrossRef]
- Pokhilko, A.; Flis, A.; Sulpice, R.; Stitt, M.; Ebenhöh, O. Adjustment of carbon fluxes to light conditions regulates the daily turnover of starch in plants: A computational model. Mol. Biosyst. 2014, 10, 613–627. [Google Scholar] [CrossRef]
- Minchin, P.; Lacointe, A. New understanding on phloem physiology and possible consequences for modelling long-distance carbon transport. New Phytol. 2005, 166, 771–779. [Google Scholar] [CrossRef]
- Van’t Hoff, J.H. Etudes de dynamique chimique. Recl. Trav. Chim. Pays-Bas. 1884, 3, 333–336. [Google Scholar] [CrossRef]
- Laidler, K.J. The development of the Arrhenius equation. J. Chem. Educ. 1984, 61, 494. [Google Scholar] [CrossRef]
- Arrhenius, S. Über die reaktionsgeschwindigkeit bei der inversion von rohrzucker durch säuren. Z. Phys. Chem. 1889, 4, 226–248. [Google Scholar] [CrossRef]
- Johnson, I.R.; Thornley, J.H.M. Temperature dependence of plant and crop process. Ann. Bot. 1985, 55, 1–24. [Google Scholar] [CrossRef]
- Bisswanger, H. Enzyme Kinetics; Wiley-VCH: Weinheim, Germany, 2001. [Google Scholar]
- Bisswanger, H. pH and temperature dependence of enzymes. In Enzyme Kinetics. 2017, 145–152. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fürtauer, L.; Weiszmann, J.; Weckwerth, W.; Nägele, T. Dynamics of Plant Metabolism during Cold Acclimation. Int. J. Mol. Sci. 2019, 20, 5411. https://doi.org/10.3390/ijms20215411
Fürtauer L, Weiszmann J, Weckwerth W, Nägele T. Dynamics of Plant Metabolism during Cold Acclimation. International Journal of Molecular Sciences. 2019; 20(21):5411. https://doi.org/10.3390/ijms20215411
Chicago/Turabian StyleFürtauer, Lisa, Jakob Weiszmann, Wolfram Weckwerth, and Thomas Nägele. 2019. "Dynamics of Plant Metabolism during Cold Acclimation" International Journal of Molecular Sciences 20, no. 21: 5411. https://doi.org/10.3390/ijms20215411
APA StyleFürtauer, L., Weiszmann, J., Weckwerth, W., & Nägele, T. (2019). Dynamics of Plant Metabolism during Cold Acclimation. International Journal of Molecular Sciences, 20(21), 5411. https://doi.org/10.3390/ijms20215411





