1. Introduction
Long-chain (≥C
20) polyunsaturated fatty acids (LC-PUFA), such as eicosapentaenoic (EPA; 20:5n−3), arachidonic (ARA, 20:4n−6), and docosahexaenoic (DHA; 22:6n−3) acids play important roles in growth, development, and reproduction in vertebrates, being specifically involved in maintenance of cellular membrane structure, energy metabolism, gene regulation and cellular signaling, and promoting cardiovascular health and immune function [
1,
2]. Fish, especially marine species, are major sources of LC-PUFA in the human diet [
3]. However, with overfishing and the degradation of the marine environment, natural wild fishery stocks have reduced sharply. The declining capture fisheries has turned attention to farmed marine fish as the major source of LC-PUFA. Thus, much attention has been focused on elucidating the regulatory mechanisms of LC-PUFA biosynthesis, in order to maximize endogenous production in marine fish.
While LC-PUFA are important for normal growth and development of all fish, the biosynthetic capacity differs between species [
4]. All the teleost fatty acyl desaturases (
fads2) genes cloned to date are homologous to mammalian
fads2, but their substrate specificities differ among species and monofunctional and bifunctional desaturases with Δ4, Δ5, and Δ6 activities have been described [
5]. Moreover, elongases of very long-chain fatty acids
(elovl) encoding genes with relevant roles in the biosynthesis of LC-PUFA in teleosts include
elovl2,
elovl4, and
elovl5, of which
elovl4 and
elovl5 are present in virtually all teleosts [
5]. In general, freshwater fish and salmonid species can convert the C
18 PUFA precursors, α-linolenic acid (18:3n−3; ALA) and linoleic acid (18:2n−6; LA), to LC-PUFA through a series of desaturation and elongation reactions catalyzed by Fads2 and Elovl, whereas most marine teleosts lack or have very limited capability [
6,
7,
8]. Consequently, essential fatty acid (EFA) requirements of freshwater fish can be satisfied by ALA and LA, while marine fish require dietary LC-PUFA. Accordingly, in aquaculture production, vegetable oils rich in ALA and LA can be used as dietary lipid sources for freshwater fish, while fish oil rich in LC-PUFA is required in feed for marine fish to meet EFA requirements for normal growth. The limited supplies of fish oil resources and their high price restricts the sustainable development of the mariculture industry. Therefore, it is necessary and important to understand the regulatory mechanisms of LC-PUFA biosynthesis in fish so as to develop methods to optimize the endogenous production (biosynthesis) of LC-PUFA with the aim to reduce the reliance of the aquaculture industry on fish oil.
It is understood that the capability of fish for LC-PUFA biosynthesis depends largely on the expression and/or activities of key enzymes involved in the biosynthetic pathway [
5,
9,
10]. At a transcriptional level, sterol regulatory element binding proteins 1 (Srebp-1) and peroxisome proliferator-activated receptors (Ppars) are major transcription factors (TF) of genes for key enzyme involved in lipid metabolism including LC-PUFA biosynthesis [
11]. While two forms of Srebp-1, i.e., Srebp-1a and -1c, have been characterized in mammals [
12], only a single form of Srebp-1 was characterized in fish, and this demonstrated to be involving two subtypes of Pparα (namely Pparα1 and Pparα2) in some fishes [
13,
14,
15], four Pparβ subtypes in Atlantic salmon (
Salmo salar) [
16], and three Ppar subtypes in rabbitfish (
Siganus canaliculatus) (Pparα, Pparβ, and Pparγ) [
17]. It was reported that Pparα up-regulated
fads2 promoter activity in rainbow trout (
Oncorhynchus mykiss) and Japanese seabass (
Lateolabrax japonicus) [
18], and Pparγ is involved in the transcriptional regulation of Δ6/Δ5
fads2 in the liver of
S. canaliculatus [
19].
In the recent years, stimulatory protein 1 (Sp1) binding sites were found in the gene promoter of human Δ6
fads2 [
6], pig
elovl6 [
20], and bovine
elovl7 [
21]. In fish, the Δ6
fads2 promoter of Atlantic salmon showed stronger promoter activity than that of Atlantic cod (
Gadus morhua) associated with the presence of the Sp1 binding site in the former [
22]. Furthermore, the lack of Sp1 binding sites in the promoters of the
fads2 gene of
L. japonicus,
Dicentrarchus labrax, and
Epinephelus coioides was associated with lower activity of the promoters [
23]. These results suggested that Sp1 could be involved in the regulation of LC-PUFA biosynthesis in teleost fish by activating promoter activities of genes encoding key enzymes. However, no direct evidence has been presented, and the underlying functions of Sp-1 and the mechanisms involved are not clear.
Rabbitfish
S. canaliculatus is a commercially important marine teleost fish widespread along the Indo-West Pacific coast and also known as one of the mainly harvested fish species. It is naturally herbivorous, consuming algae and seagrass; however, they can also feed on compound feed or trash fishes after brief domestication with them. It is noteworthy that rabbitfish was the first marine teleost demonstrated to have capability for LC-PUFA biosynthesis from C
18 precursors [
24]. Genes encoding key enzymes for LC-PUFA biosynthesis including Δ4 Fads2, bifunctional Δ6/Δ5 Fads2, Elovl4, and Elovl5 were functionally characterized in this species, which provides a good model for studying the regulatory mechanisms of LC-PUFA biosynthesis in teleosts [
24,
25]. In addition, bioinformatic analysis predicted Sp1 binding sites in the promoters of rabbitfish Δ6/Δ5
fads2 and
elovl5, but Sp1 binding sites were absent in the promoters of Δ4
fads and
elovl4. Furthermore, in a recent study, inserting the Sp1 binding site of rabbitfish Δ6/Δ5
fads2 promoter into the corresponding region of
E. coioides fads2 promoter demonstrated the importance of the Sp1 binding site in determining
fads2 promoter activity [
23]. However, until now, no study has directly demonstrated the role of Sp1 in the regulation of LC-PUFA biosynthesis in any vertebrate including fish, and thus the present study aimed to clarify this in rabbitfish. Therefore, the
sp1 gene was cloned, and its function in the regulation of LC-PUFA biosynthesis was investigated by determining the effects of Sp1 on the expression of Δ6/Δ5
fads2 and
elovl5 genes, and on the conversion of C
18 fatty acid precursors to LC-PUFA. The data obtained increased our understanding of the regulatory mechanisms of LC-PUFA biosynthesis in vertebrates and will contribute to the optimization and/or enhancement of LC-PUFA biosynthesis in teleosts.
3. Discussion
Sp1 is a transactivation molecule belonging to the family of Sp or Krüppel-like factor (KLF) proteins [
26], and the Sp family of transcription factors is characterized by a particular combination of three conserved Cys2His2 zinc fingers [
27]. In the present study, we cloned the rabbitfish
sp1 gene whose amino acid sequence shared high similarity and typical structural characteristics with those of other of other species. The C-terminus domain had the family marker region, featuring three Cys2His2 zinc fingers, required for sequence-specific DNA binding to GC-rich promoter elements [
28,
29]. Moreover, the sequences and structure were very similar between rabbitfish and zebrafish zinc finger domains, which suggested that rabbitfish Sp1 might also interact with GC sequences as found previously with zebrafish Sp1 [
30]. Since the initial discovery of Sp1, it has generally been defined as a ‘basal’ transcription factor as single or multiple Sp1 binding sites have been mapped in promoters and enhancers of genes involved in almost all cellular processes. Besides, Sp1 plays an extremely important role in growth and metastasis of many tumors by regulating oncogenes, tumor suppressor genes, cell cycle control molecules, growth-related signal transduction, angiogenesis related factors, as well as apoptosis [
28,
31,
32,
33]. It is reported that suppression of
sp1 expression reduced the growth of colon cancer stem cells (CCSC) and induced apoptosis in vitro and in nude mouse xenografts, and the proportion of CCSC markers, CD44+/CD166+, was decreased following
sp1 knock-down [
34]. Nevertheless, knowledge of the binding specificities of various Sp1 proteins for GC-boxes in promoter/enhancer DNA, or for other transcriptional and epigenetic regulators, is rather incomplete [
35,
36].
Several studies reported that Sp1 may be involved in the regulation of LC-PUFA biosynthesis in teleost fish by activating the promoter activities of genes encoding key enzymes although direct evidence was absent [
22,
23]. For instance, Sp1 elements were found in the
fads2 promoter regions of some fish species with LC-PUFA biosynthetic ability such as
S. salar [
22],
D. rerio, and
S. canaliculatus [
23]. However, Sp1 elements were lacking in
fads2 promoter regions of carnivorous marine fish species like
G. morhua,
D. labrax,
L. japonicus,
L. crocea and
E. coioides, in which LC-PUFA biosynthetic ability is lacking or very low. These data suggested that the lack of Sp1 binding sites may lead to low promoter activity of
fads2, and thus result in low hepatic
fads2 expression in carnivorous marine teleosts, as recently shown in
E. coioides [
23]. Furthermore, it is reported that Sp3 is structurally similar to Sp1, with similar affinities for the Sp1-binding site [
37]. Even so, their DNA-binding properties and regulatory functions are different [
38]. There are several studies suggesting that Sp1 is responsible for basal transcription, and Sp3 is important for the induced transcription activation [
39,
40,
41]. For example, the binding of Sp3 at the PKR promoter in vivo was interferon dependent, whereas the binding of Sp1 was constitutive [
41]. Considering the complexity of the interaction between Sp1 and Sp3, the roles of Sp3 in LC-PUFA biosynthesis deserves further study.
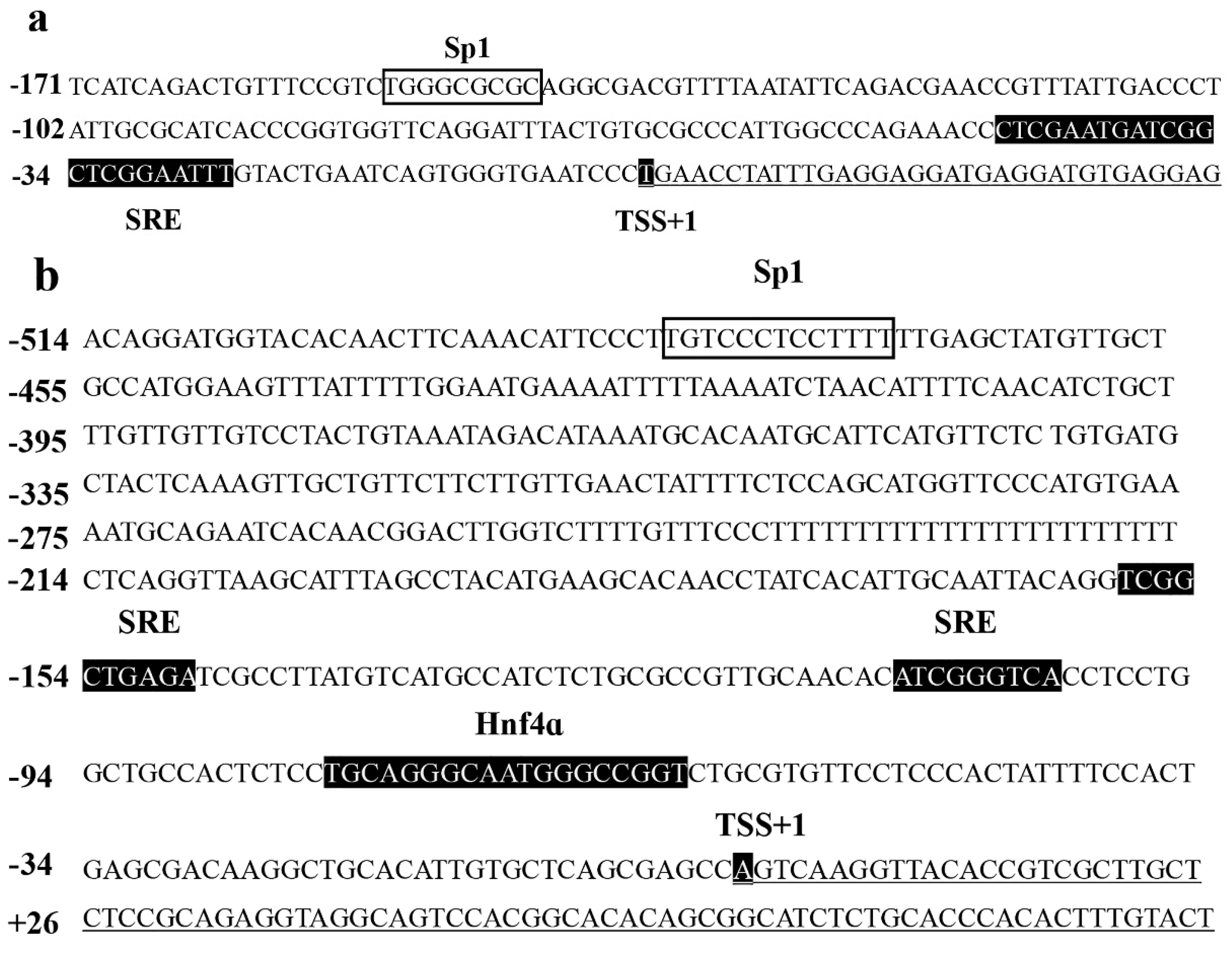
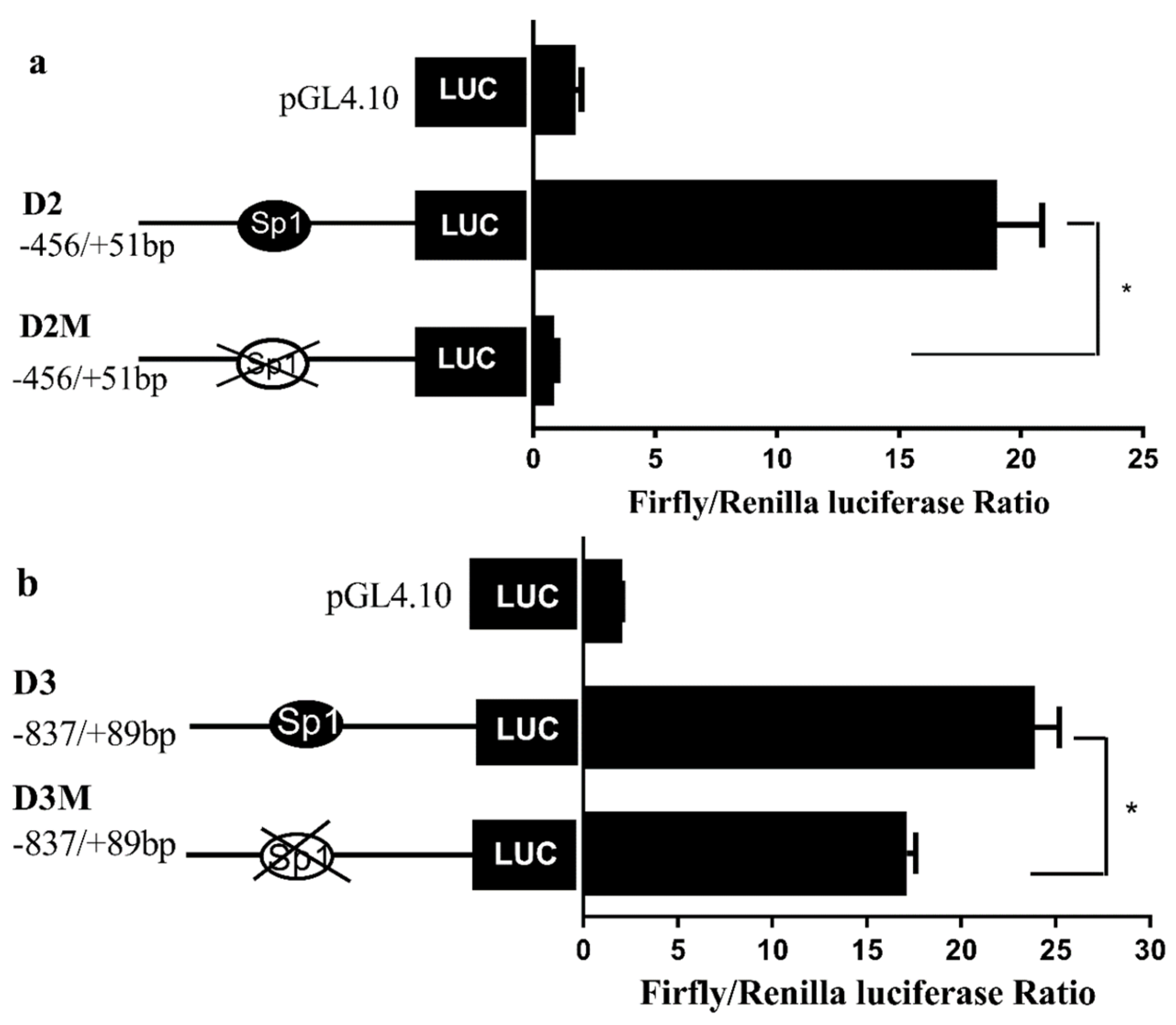
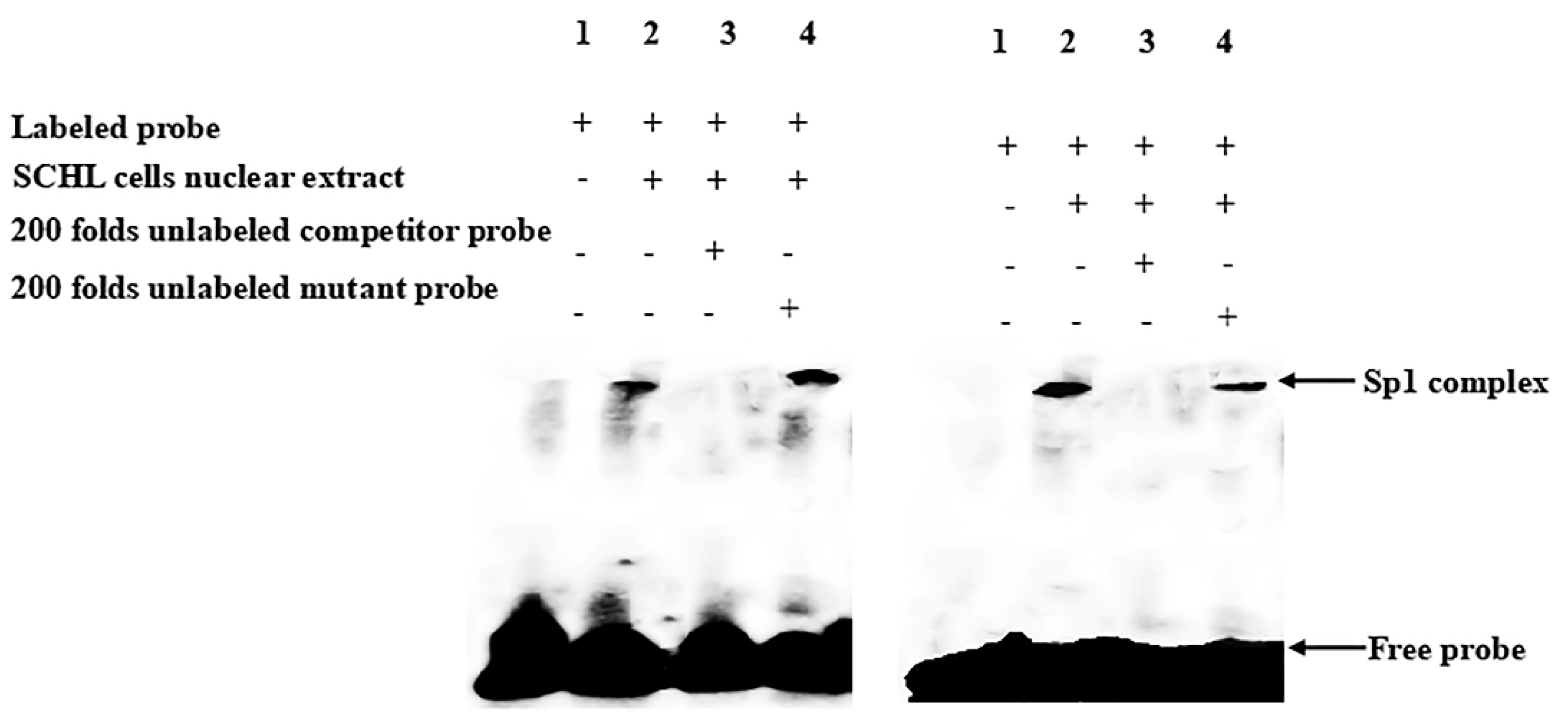
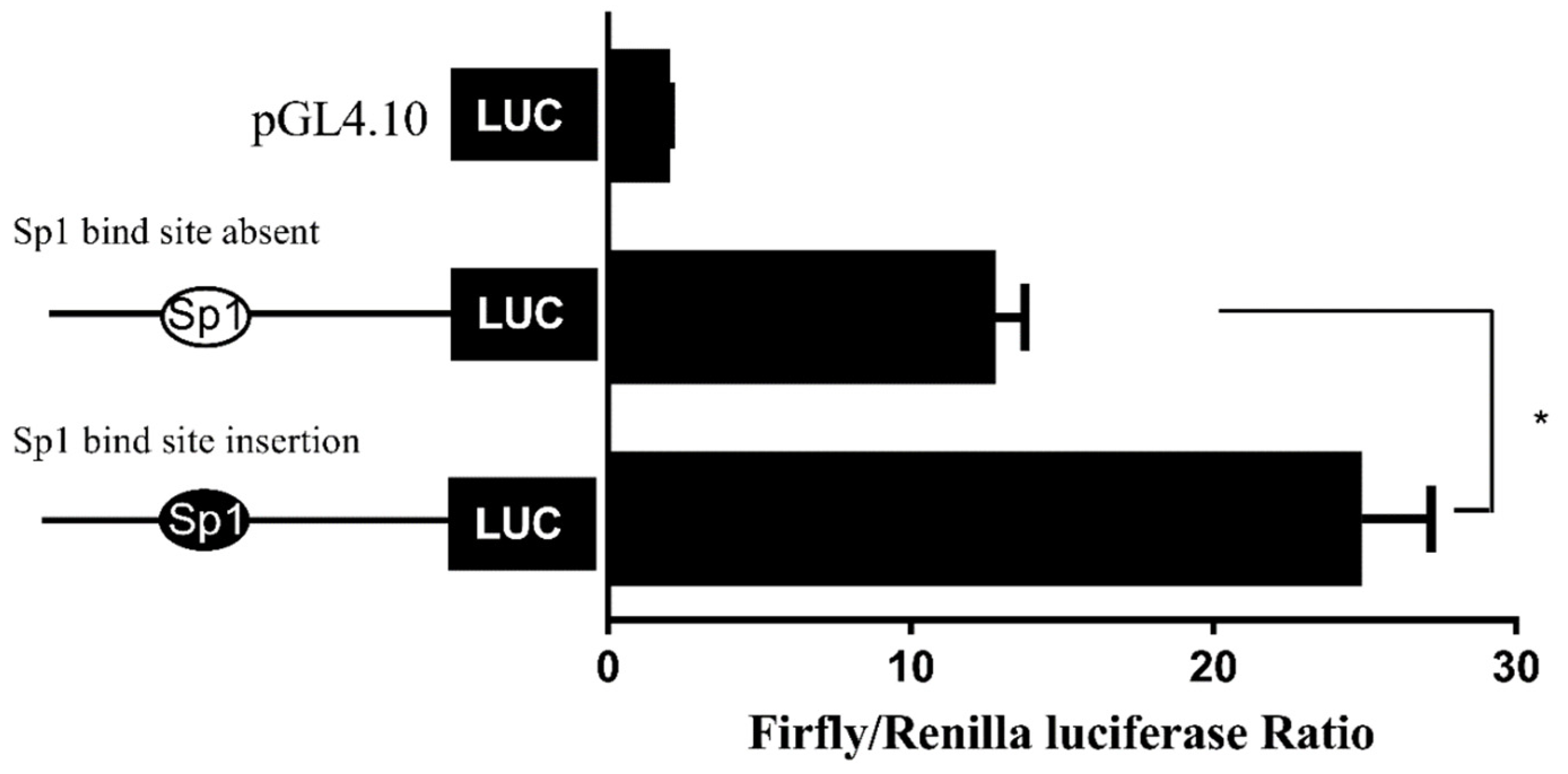
While the above data suggested the importance of the Sp1-binding site in determining fads2 promoter activity, the role of Sp1 in the transcriptional regulation of LC-PUFA biosynthesis in vertebrates was not directly shown. In the present study, potential Sp1 binding sites were found in rabbitfish S. canaliculatus Δ6/Δ5 fads2 and elovl5 promoters, but were not predicted in the Δ4 fads2 promoter region. After the Sp1 binding site was inserted into the rabbitfish Δ4 fads2 promoter, its activity was increased. Moreover, mutation of the Sp1 sites of Δ6/Δ5 fads2 and elovl5 promoters resulted in decreased promoter activities. These data provided direct evidence that Sp1 plays an important role in determining Δ6/Δ5 fads2 and elovl5 promoter activity in S. canaliculatus, and the weak activity of Δ4 fads2 promoter may be, at least partly, due to the lack of Sp1 binding sites.
Sp1 can promote the expression of its target genes [
42]. Generally, the level of gene transcription in eukaryotic cells is dependent on the binding of RNA polymerase and transcription factors to specific sequences in gene promoters [
43]. Sp1 functions by interacting with the TATA-box binding protein complex (TFIID) and facilitating binding of TFIID to the promoter, which, in turn, recruits RNA polymerase II (Pol II) [
29,
44]. In addition, however, Sp1 also plays a key role in maintaining expression of genes that lack a TATA-box in the promoter [
29,
45]. As there was no TATA box in the promoters of rabbitfish
S. canaliculatus Δ6/Δ5
fads2 and
elovl5, the significant changes in expression of Δ6/Δ5
fads2 and
elovl5 when overexpressing or inhibiting
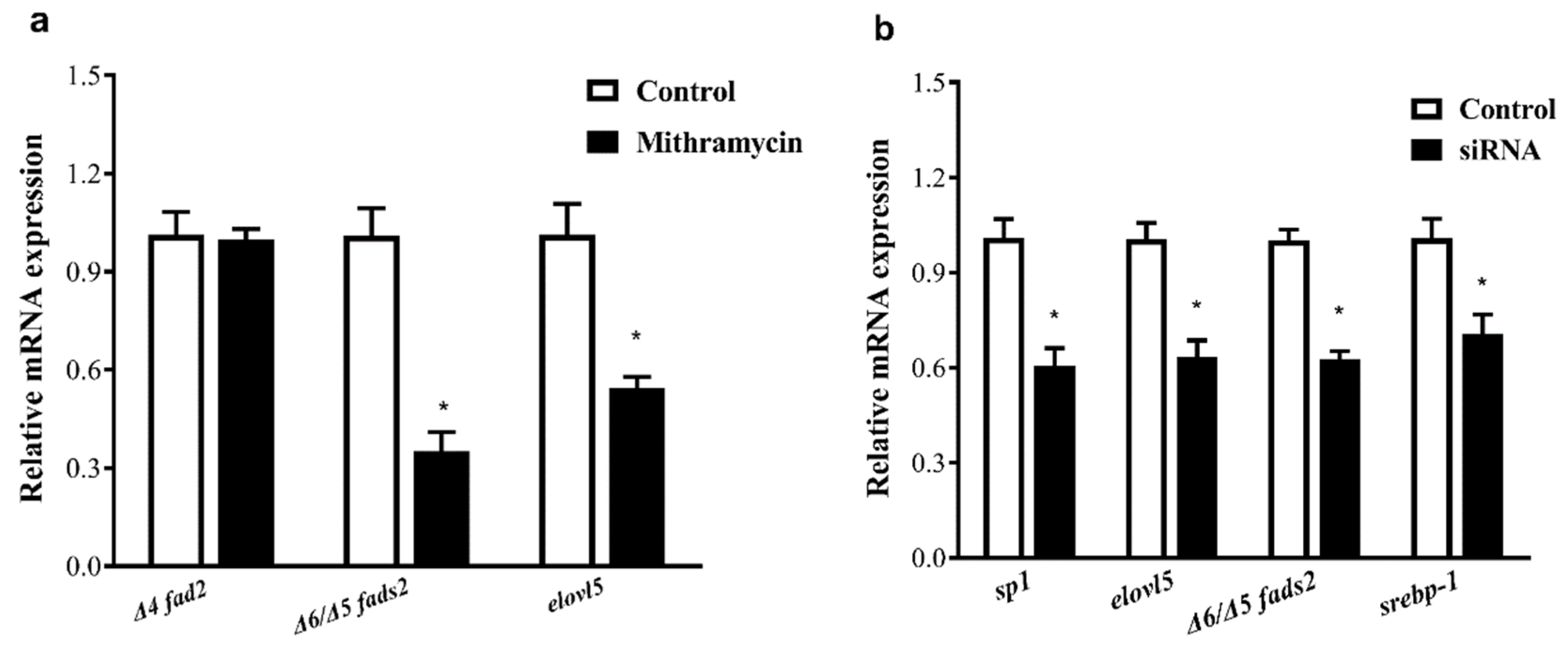
sp1 indicated that Sp1 can stimulate the expression of these two genes via the regulation of transcription activity. Further research is required to investigate the detailed regulation mechanisms of Sp1 on the expression of the rabbitfish
S. canaliculatus Δ6/Δ5
fads2 and
elovl5 gene.
It is reported that the fatty acid synthase (Fas) gene promoter is regulated by Sp1 and Srebp transcription factors [
46]. Sp1 maintains the expression of
fas directly and also has been shown to regulate Srebp-1c in colon cancer [
47]. Srebp-1 is a member of the basic helix–loop–helix–leucine zipper family of transcription factors that regulate the biosynthesis of both cholesterol and fatty acids [
48,
49,
50]. Previous studies indicated that Srebp-1 is a weak activator of transcription and only functions efficiently when activated by co-activating transcription factors such as SRE, E-box, LXR, NF-Y, and Sp1 [
50,
51,
52]. Regulation of Srebp-1 by Sp1 has also been reported previously. For example, Sp1 functioned together with Srebp-1 to synergistically activate the
fas promoter [
53,
54,
55]. Similarly, studies of Srebps in fish including
S. salar [
22,
56] and
D. labrax [
57] have reported previously that Srebp-1 mediated the expression of Δ6
fads2, and thus Srebp-1 may be involved in the transcriptional regulation of LC-PUFA biosynthesis in fish [
22,
58,
59]. Interestingly, the highly conserved NF-Y and SRE elements were demonstrated in rabbitfish
S. canaliculatus Δ6/Δ5
fads2 promoter, suggesting Srebps as a major regulator of Δ6/Δ5
fads2 expression [
23,
60]. The present study also indicated that Srebp-1 expression was changed when overexpressing or inhibiting Sp1. Therefore, Sp1 might also be indirectly involved in stimulating the expression of Srebp-1 to activate Δ6/Δ5
fads2 and
elovl5 gene expression.
Sp1 enhanced LC-PUFA biosynthesis in SCHL cells by increasing Δ6/Δ5
fads2 and
elovl5 gene expression. Another study demonstrated that Sp1 binds to bovine
elovl7 promoter and activities the expression of
elovl7 in bovine mammary epithelial cells (bMECs) [
21]. In rabbitfish, functional characterization showed that Δ6/Δ5
fads2 could efficiently convert 18:2n−6 to 18:3n−6 [
24] and the ratio of 18:3n−6/18:2n−6 is an index of Δ6
fads2 activity [
61]. In the present study, the expression of Δ6/Δ5
fads2 and
elovl5 was increased by overexpression of
sp1 and, correspondingly, cell fatty acid profiles were changed. Overexpression of
sp1 was associated with increased levels of Δ6 desaturation products such as 18:3n−6 and elongation products such as 20:2n−6 and 20:3n−3, or with further downstream products in the LC-PUFA biosynthetic pathway such as ARA, EPA, and DHA. Moreover, overexpression of
sp1 increased the 18:3n−6/18:2n−6, 20:2n−6/18:2n−6, and 20:3n−3/18:3n−3 ratios in SCHL cells, which indicated that Sp1 could stimulate LC-PUFA synthesis in liver.
In summary, the present study demonstrated that Sp1 positively regulated the biosynthesis of LC-PUFA in rabbitfish, and functioned mainly through binding to Δ6/Δ5 fads2 and elovl5 promoters and activating their expression. To our knowledge, this is the first report of the direct involvement of Sp1 in the regulation of LC-PUFA biosynthesis at transcriptional and metabolic level in vertebrates, and this knowledge may contribute to efforts to enhance LC-PUFA biosynthesis in farmed fish.