Proteomic Analysis in Morquio A Cells Treated with Immobilized Enzymatic Replacement Therapy on Nanostructured Lipid Systems
Abstract
1. Introduction
2. Results
2.1. Qualitative Analysis Liquid Chromatography—Mass Spectrometry (DDA-LC-MS/MS)
2.2. Protein Quantification by SWATH-MS
3. Discussion
4. Materials and Methods
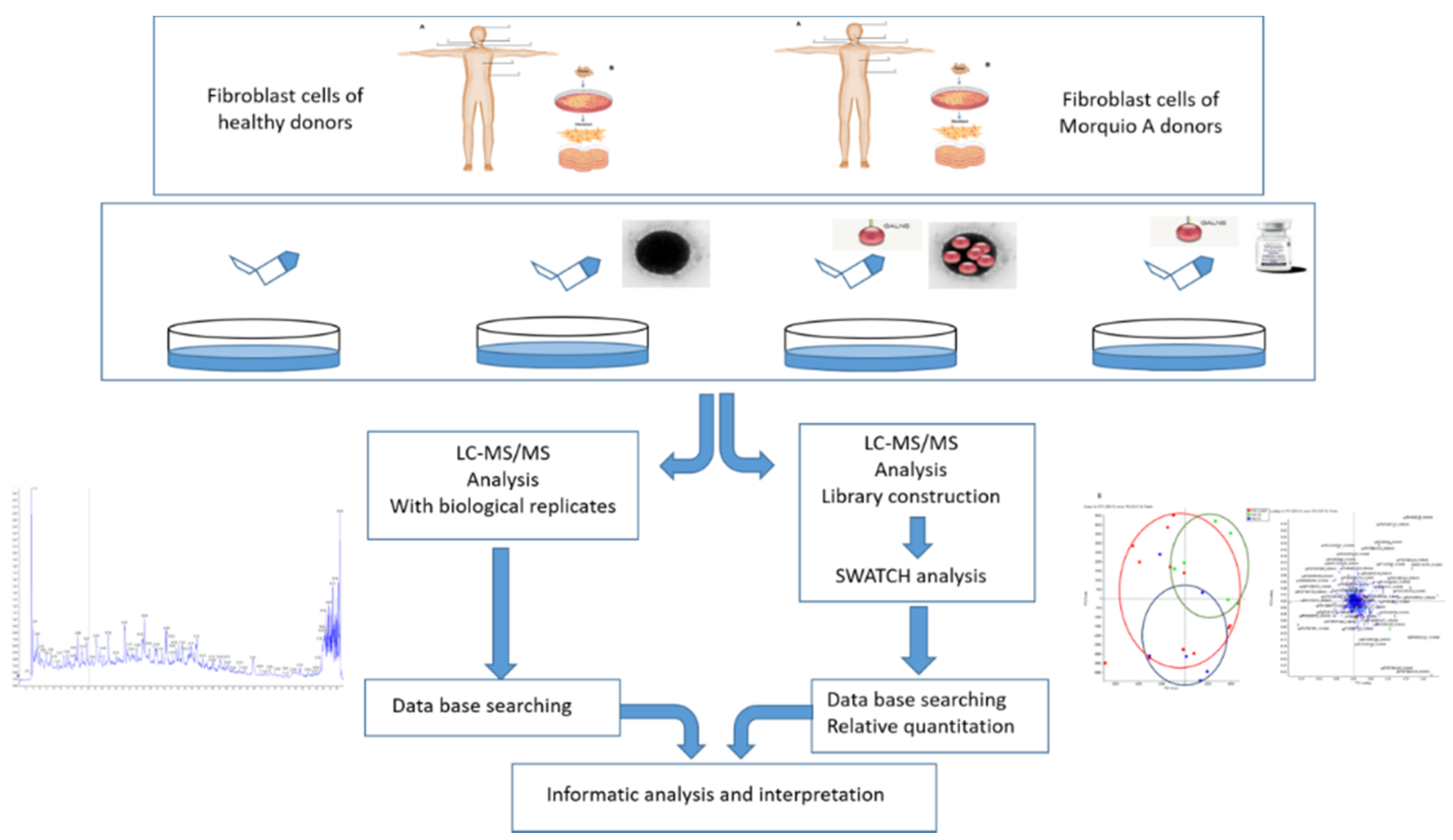
4.1. The Work Scheme of This Study
4.2. Sample Resources
4.3. Cell Cultures in Fibroblasts
4.4. Materials and Preparation of NLC
4.5. Protein Extraction
4.6. Enzyme Activity Test
4.7. Proteomic Analysis
4.7.1. In Gel Protein Digestion
4.7.2. Mass Spectrometric Analysis (DDA acquisition)
4.7.3. Data Analysis
4.8. Protein Quantification by SWATH-MS
4.8.1. Creation of the Spectral Library
4.8.2. Relative Quantification by SWATH-MS Acquisition
4.8.3. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Conflicts of Interest
Abbreviations
| C6S | Chondroitin 6 sulphate |
| ERT | Enzyme replacement therapy |
| FC | Fold change |
| GAG | Glycosaminoglycan |
| GALNS | N-acetylgalactosamine-6-sulfatase |
| KS | Keratan sulphateJ |
| LC | Liquid chromatography |
| MPS IVA | Mucopolysaccharidosis type IVA |
| MRM | Multiple reaction monitoring |
| MS | Mass spectrometry |
| NLC | Nanostructure lipid carrier |
| S | Sample |
| SDC | Sodium dodecyl sulfate |
| SRM | Selected reaction monitoring |
| SNARE | soluble N-ethylmaleimide-sensitive factor attachment protein receptor |
| SWATH-MS | Sequential window acquisition of all theoretical mass spectra |
| UMoC | Untreated Morquio A cells |
References
- Hendriksz, C.J.; Berger, K.I.; Giugliani, R.; Harmatz, P.; Kampmann, C.; Mackenzie, W.G.; Raiman, J.; Villarreal, M.S.; Savarirayan, R. International Guidelines for the Management and Treatment of Morquio A Syndrome. Am. J. Med. Genet. A 2015, 167, 11–25. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.; Alméciga-Díaz, C.J.; Sawamoto, K.; Mackenzie, W.G.; Theroux, M.C.; Pizarro, C.; Mason, R.W.; Orii, T.; Tomatsu, S. Mucopolysaccharidosis IVA and glycosaminoglycans. Mol. Genet. Metab. 2017, 120, 78–95. [Google Scholar] [CrossRef]
- Montaño, A.M.; Tomatsu, S.; Gottesman, G.S.; Smith, M.; Orii, T. International MorquioA Registry: Clinical manifestation and natural course of Morquio A disease. J. Inherit. Metab. Dis. 2007, 30, 165–174. [Google Scholar] [CrossRef] [PubMed]
- Tomatsu, S.; Montaño, A.M.; Oikawa, H.; Smith, M.; Barrera, L.; Chinen, Y.; Thacker, M.M.; Mackenzie, W.G.; Suzuki, Y.; Orii, T. Mucopolysaccharidosis type IVA (Morquio A disease): Clinical review and current treatment. Curr. Pharm. Biotechnol. 2011, 12, 931–945. [Google Scholar] [CrossRef]
- Lavery, C.; Hendriksz, C. Mortality in patients with Morquio Syndrome A. JIMD Rep. 2015, 15, 59–66. [Google Scholar]
- Hendriksz, C.J. Elosulfasealfa (BMN 110) for the treatment of mucopolysaccharidosis IVA (Morquio A Syndrome). Expert Rev. Clin. Pharm. 2016, 9, 1521–1532. [Google Scholar] [CrossRef] [PubMed]
- Hendriksz, C.; Santra, S.; Jones, S.A.; Geberhiwot, T.; Jesaitis, L.; Long, B.; Qi, Y.; Hawley, S.M.; Decker, C. Safety, immunogenicity, and clinical outcomes in patients with Morquio A syndrome participating in 2 sequential open-label studies of elosulfase alfa enzyme replacement therapy (MOR-002/MOR-100), representing 5 years of treatment. Mol. Genet. Metab. 2018, 123, 479–487. [Google Scholar] [CrossRef] [PubMed]
- Doherty, C.; Stapleton, M.; Piechnik, M.; Mason, R.W.; Mackenzie, W.G.; Yamaguchi, S.; Kobayashi, H.; Suzuki, Y.; Tomatsu, S. Effect of enzyme replacement therapy on the growth of patients with Morquio A. J. Hum. Genet. 2019, 64, 625–635. [Google Scholar] [CrossRef]
- Tomatsu, S.; Averill, L.W.; Sawamoto, K.; Mackenzie, W.G.; Bober, M.B.; Pizarro, C.; Goff, C.J.; Xie, L.; Orii, T.; Theroux, M. Obstructive airway in Morquio A syndrome, the past, the present and the future. Mol. Genet. Metab. 2016, 117, 150–156. [Google Scholar] [CrossRef]
- Tomatsu, S.; Sawamoto, K.; Shimada, T.; Bober, M.B.; Kubaski, F.; Yasuda, E.; Mason, R.W.; Khan, S.; Alméciga-Díaz, C.J.; Barrera, L.A.; et al. Enzyme replacement therapy for treating mucopolysaccharidosis type IVA (Morquio A syndrome): Effect and limitations. Expert Opin. Orphan Drugs 2015, 3, 1279–1290. [Google Scholar] [CrossRef]
- Tomatsu, S.; Sawamoto, K.; Alméciga-Díaz, C.J.; Shimada, T.; Bober, M.B.; Chinen, Y.; Yabe, H.; Montaño, A.M.; Giugliani, R.; Kubaski, F.; et al. Impact of enzyme replacement therapy and hematopoietic stem cell transplantation in patients with Morquio A syndrome. Drug Des. Devel. 2015, 9, 1937–1953. [Google Scholar] [CrossRef] [PubMed]
- Qi, Y.; Musson, D.G.; Schweighardt, B.; Tompkins, T.; Jesaitis, L.; Shaywitz, A.J.; Yang, K.; O’Neill, C.A. Pharmacokinetic and pharmacodynamic evaluation of Elosulfase Alfa, an enzyme replacement therapy in patients with MorquioA Syndrome. Clin. Pharm. 2014, 53, 1137–1147. [Google Scholar] [CrossRef] [PubMed]
- Kenth, J.J.; Thompson, G.; Fullwood, C.; Wilkinson, S.; Jones, S.; Bruce, I.A. The characterisation of pulmonary function in patients with mucopolysaccharidoses IVA: A longitudinal analysis. Mol. Genet. Metab. Rep. 2019, 20, 100487. [Google Scholar] [CrossRef] [PubMed]
- Tomatsu, S.; Montaño, A.M.; Dung, V.C.; Ohashi, A.; Oikawa, H.; Oguma, T.; Orii, T.; Barrera, L.; Sly, W.S. Enhancement of Drug Delivery: Enzyme-replacement therapy for murine Morquio A Syndrome. Mol. Ther. 2010, 18, 1094–1102. [Google Scholar] [CrossRef] [PubMed]
- Long, B.; Tompkins, T.; Decker, C.; Jesaitis, L.; Khan, S.; Slasor, P.; Harmatz, P.; O’Neill, C.A.; Schweighardt, B. Long-term Immunogenicity of Elosulfase Alfa in the Treatment of MorquioA Syndrome: Results From MOR-005, a Phase III Extension Study. Clin. Ther. 2017, 39, 118–129. [Google Scholar] [CrossRef]
- Sawamoto, K.; Stapleton, M.; Alméciga-Díaz, C.J.; Espejo-Mojica, A.J.; Losada, J.C.; Suarez, D.A.; Tomatsu, S. Therapeutic options for Mucopolysaccharidoses: Current and emerging treatments. Drugs 2019, 79, 1103–1134. [Google Scholar] [CrossRef] [PubMed]
- Lavillette, D.; Russell, S.J.; Cosset, F.L. Retargeting gene delivery using surface-engineered retroviral vector particles. Curr. Opin. Biotechnol. 2001, 12, 461–466. [Google Scholar] [CrossRef]
- St George, J.A. Gene therapy progress and prospects: Adenoviral vectors. Gene 2003, 10, 1135–1141. [Google Scholar] [CrossRef]
- Buning, H.; Ried, M.U.; Perabo, L.; Gerner, F.M.; Huttner, N.A.; Enssle, J.; Hallek, M. Receptor targeting of adeno-associated virus vectors. Gene 2003, 10, 1142–1151. [Google Scholar]
- Loiler, S.A.; Conlon, T.J.; Song, S.; Tang, Q.; Warrington, K.H.; Agarwal, A.; Kapturczak, M.; Li, C.; Ricordi, C.; Atkinson, M.A.; et al. Targeting recombinant adeno-associated virus vectors to enhance gene transfer to pancreatic islets and liver. Gene 2003, 10, 1551–1558. [Google Scholar] [CrossRef]
- Choi, V.W.; McCarty, D.M.; Samulski, R.J. AAV hybrid serotypes: Improved vectors for gene delivery. Curr. Gene 2005, 5, 299–310. [Google Scholar] [CrossRef]
- Rabilloud, T.; Chevallet, M.; Luche, S.; Lelong, C. Two-dimensional gel electrophoresis in proteomics: Past, present and future. J. Proteom. 2010, 73, 2064–2077. [Google Scholar] [CrossRef] [PubMed]
- Kapphahn, R.J.; Richards, M.J.; Ferrington, D.A.; Fliesler, S.J. Lipid-derived and other oxidative modifications of retinal proteins in a rat model of Smith-Lemli-Opitz syndrome. Exp. Eye Res. 2019, 178, 247–254. [Google Scholar] [CrossRef] [PubMed]
- Santana, A.M.; Thomas, F.C.; Silva, D.G.; McCulloch, E.; Vidal, A.M.C.; Burchmore, R.J.S.; Fagliari, J.J.; Eckersall, P.D. Reference 1D and 2D electrophoresis maps for potential disease related proteins in milk whey from lactating buffaloes and blood serum from buffalo calves (Water buffalo, Bubalusbubalis). Res. Vet. Sci. 2018, 118, 449–465. [Google Scholar] [CrossRef] [PubMed]
- Vidova, V.; Spacil, Z. A review on mass spectrometry-based quantitative proteomics: Targeted and data independent acquisition. Anal. Chim. Acta 2017, 964, 7–23. [Google Scholar] [CrossRef] [PubMed]
- Couselo-Seijas, M.; López-Canoa, J.N.; Agra-Bermejo, R.M.; Díaz-Rodriguez, E.; Fernandez, A.L.; Martinez-Cereijo, J.M.; Durán-Muñoz, D.; Bravo, S.B.; Velo, A.; González-Melchor, L.; et al. Cholinergic activity regulates the secretome of epicardial adipose tissue: Association with atrial fibrillation. J. Cell. Physiol. 2019, 234, 10512–11522. [Google Scholar] [CrossRef] [PubMed]
- Del Pilar Chantada-Vázquez, M.; López, A.C.; Bravo, S.B.; Vázquez-Estévez, S.; Acea-Nebril, B.; Núñez, C. Proteomic analysis of the bio-corona formed on the surface of (Au, Ag, Pt)-nanoparticles in human serum. Colloids Surf. B Biointerfaces 2019, 177, 141–148. [Google Scholar] [CrossRef]
- Shilov, I.V.; Seymour, S.L.; Patel, A.A.; Loboda, A.; Tang, W.H.; Keating, S.P.; Hunter, C.L.; Nuwaysir, L.M.; Schaeffer, D.A. The Paragon Algorithm, a Next Generation Search Engine That Uses Sequence Temperature Values and Feature Probabilities to Identify Peptides from Tandem Mass Spectra. Mol. Cell. Proteom. 2007, 6, 1638–1655. [Google Scholar] [CrossRef]
- Pathan, M.; Keerthikumar, S.; Ang, C.S.; Gangoda, L.; Quek, C.Y.J.; Williamson, N.A.; Mouradov, D.; Sieber, O.M.; Simpson, R.J.; Salim, A.; et al. FunRich: An open access standalone functional enrichment and interaction network analysis tool. Proteomics 2015, 15, 2597–2601. [Google Scholar] [CrossRef]
- Pathan, M.; Keerthikumar, S.; Chisanga, D.; Alessandro, R.; Ang, C.-S.; Askenase, P.; Batagov, A.O.; Benito-Martin, A.; Camussi, G.; Clayton, A.; et al. A novel community driven software for functional enrichment analysis of extracellular vesicles data. J. Extracell. Vesicles 2017, 6, 1321455. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009, 37, 1–13. [Google Scholar] [CrossRef]
- Barallobre-Barreiro, J.; Chung, Y.L.; Mayr, M. Proteomics and Metabolomics for Mechanistic Insights and Biomarker Discovery in Cardiovascular Disease. Rev. Esp. Cardiol. (Engl. Ed.) 2013, 66, 657–661. [Google Scholar] [CrossRef]
- Gregorich, Z.R.; Ge, Y. Top-down proteomics in health and disease: Challenges and opportunities. Proteomics 2014, 14, 1195–1210. [Google Scholar] [CrossRef]
- Tan, H.T.; Chung, M.C.M. Label-Free quantitative phosphoproteomics reveals regulation of vasodilator-stimulated fhosphoprotein upon Stathmin-1 Silencing in a pair of isogenic colorectal cancer cell lines. Proteomics 2018, 18, 1–30. [Google Scholar] [CrossRef]
- Jayabalan, N.; Lai, A.; Nair, S.; Guanzon, D.; Scholz-Romero, K.; Palma, P.; McIntyre, H.D.; Lappas, M.; Salomon, C. Quantitative proteomics by SWATH-MS suggest an association between circulating exosomes and maternal metabolic changes in gestational diabetes mellitus. Proteomics 2019, 19, e1800164. [Google Scholar] [CrossRef]
- Braccia, C.; Espinal, M.P.; Pini, M.; De Pietri Tonelli, D.; Armirotti, A. A new SWATH ion library for mouse adult hippocampal neural stem cells. Data Br. 2018, 18, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Jamwal, R.; Barlock, B.J.; Adusumalli, S.; Ogasawara, K.; Simons, B.L.; Akhlaghi, F. Multiplex and label-Free relative quantification approach for studying protein abundance of drug metabolizing enzymes in human liver microsomes using SWATH-MS. J. Proteome Res. 2017, 16, 4134–4143. [Google Scholar] [CrossRef]
- Zhu, Y.; Zhu, J.; Lu, C.; Zhang, Q.; Xie, W.; Sun, P.; Dong, X.; Yue, L.; Sun, Y.; Yi, X.; et al. Identification of protein abundance changes in Hepatocellular Carcinoma Tissues using PCT–SWATH. Proteom. Clin. Appl. 2019, 13, e1700179. [Google Scholar] [CrossRef]
- Li, H.; Mao, Y.; Xiong, Y.; Zhao, H.H.; Shen, F.; Gao, X.; Yang, P.; Liu, X.; Fu, D. A Comprehensive proteome analysis of peripheral blood mononuclear cells (PBMCs) to identify candidate biomarkers of Pancreatic Cancer. Cancer Genom. Proteom. 2019, 16, 81–89. [Google Scholar] [CrossRef]
- Chang, R.Y.K.; Etheridge, N.; Nouwens, A.S.; Dodd, P.R. SWATH analysis of the synaptic proteome in Alzheimer’s disease. Neurochem. Int. 2015, 87, 1–12. [Google Scholar] [CrossRef]
- Ebhardt, H.A.; Degen, S.; Tadini, V.; Schilb, A.; Johns, N.; Greig, C.A.; Fearon, K.C.H.; Aebersold, R.; Jacobi, C. Comprehensive proteome analysis of human skeletal muscle in cachexia and sarcopenia: A pilot study. J. Cachexia Sarcopenia Muscle 2017, 8, 567–582. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Walsh, T.; Atherton, J.J.; Kostner, K.; Schulz, B.; Punyadeera, C. Identification and validation of a salivary protein panel to detect heart failure early. Theranostics 2017, 7, 4350–4358. [Google Scholar] [CrossRef] [PubMed]
- Coutinho, M.F.; Prata, M.J.; Alves, S. Mannose-6-phosphate pathway: A review on its role in lysosomal function and dysfunction. Mol. Genet. Metab. 2012, 105, 542–550. [Google Scholar] [CrossRef] [PubMed]
- Gary-Bobo, M.; Nirdé, P.; Jeanjean, A.; Morère, A.; Garcia, M. Mannose 6-phosphate receptor targeting and its applications in human diseases. Curr. Med. Chem. 2007, 14, 2945–2953. [Google Scholar] [CrossRef] [PubMed]
- Sleat, D.E.; Ding, L.; Wang, S.; Zhao, C.; Wang, Y.; Xin, W.; Zheng, H.; Moore, D.F.; Sims, K.B.; Lobel, P. Mass spectrometry-based protein profiling to determine the cause of lysosomal storage diseases of unknown etiology. Mol. Cell. Proteom. 2009, 8, 1708–1718. [Google Scholar] [CrossRef] [PubMed]
- Ortea, I.; Rodríguez-Ariza, A.; Chicano-Gálvez, E.; Arenas Vacas, M.S.; Jurado Gámez, B. Discovery of potential protein biomarkers of lung adenocarcinoma in bronchoalveolar lavage fluid by SWATH MS data-independent acquisition and targeted data extraction. J. Proteom. 2016, 138, 106–114. [Google Scholar] [CrossRef]
- Garrido-Rodríguez, M.; Ortea, I.; Calzado, M.A.; Muñoz, E.; García, V. SWATH proteomic profiling of prostate cancer cells identifies NUSAP1 as a potential molecular target for Galiellalactone. J. Proteom. 2019, 193, 217–229. [Google Scholar] [CrossRef]
- Appelqvist, H.; Waster, P.; Kagedal, K.; Ollinger, K. The lysosome: From waste bag to potential therapeutic target. J. Mol. Cell Biol. 2013, 5, 214–226. [Google Scholar] [CrossRef]
- Ballabio, A. The awesome lysosome. EMBO Mol. Med. 2016, 8, 73–76. [Google Scholar] [CrossRef]
- Todkar, K.; Ilamathi, H.S.; Germain, M. Mitochondria and Lysosomes: Discovering Bonds. Front. Cell Dev. Biol. 2017, 5, 106. [Google Scholar] [CrossRef]
- Das, A.; Nag, S.; Mason, A.B.; Barroso, M.M. Endosome mitocondria interactions are modulated by iron release from transferrin. J. Cell Biol. 2016, 214, 831–845. [Google Scholar] [CrossRef] [PubMed]
- Lieberman, A.P.; Puertollano, R.; Raben, N.; Slaugenhaupt, S.; Walkley, S.U.; Ballabio, A. Autophagy in lysosomal storage disorders. Autophagy 2012, 8, 719–730. [Google Scholar] [CrossRef] [PubMed]
- Luzio, J.P.; Gray, S.R.; Bright, N.A. Endosome-lysosome fusión. Biochem. Soc. Trans. 2010, 38, 1413–1416. [Google Scholar] [CrossRef] [PubMed]
- Fraldi, A.; Annunziata, F.; Lombardi, A.; Kaiser, H.-J.; Medina, D.L.; Spampanato, C.; Fedele, A.O.; Polishchuk, R.; Sorrentino, N.C.; Simons, K.; et al. Lysosomal fusion and SNARE function are impaired by cholesterol accumulation in lysosomal storage disorders. EMBO J. 2010, 29, 3607–3620. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Pluhackova, K.; Böckmann, R.A. The multifaceted role of SNARE proteins in membrane fusion. Front. Physiol. 2017, 8, 5. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Liu, Q.; Wang, K.; Dang, X.; Zhang, F. Comparative analysis of gene expression profiles in normal hip human cartilage and cartilage from patients with necrosis of the femoral head. Arthritis Res. 2016, 18, 98. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lamandé, S.R.; Bateman, J.F.; Hutchison, W.; McKinlay Gardner, R.J.; Bower, S.P.; Byrne, E.; Dahl, H.H. Reduced collagen VI causes Bethlem myopathy: A heterozygous COL6A1 nonsense mutation results in mRNA decay and functional haploinsufficiency. Hum. Mol. Genet. 1998, 7, 981–989. [Google Scholar] [CrossRef][Green Version]
- Camelier, M.V.; Burin, M.G.; De Mari, J.; Vieira, T.A.; Marasca, G.; Giugliani, R. Practical and reliable enzyme test for the detection of mucopolysaccharidosis IVA (Morquio Syndrome type A) in dried blood samples. Clin. Chim. Acta 2011, 412, 1805–1808. [Google Scholar] [CrossRef]
- Bonzon-Kulichenko, E.; Pérez-Hernández, D.; Núñez, E.; Martínez-Acedo, P.; Navarro, P.; Trevisan-Herraz, M.; del Carmen Ramos, M.; Sierra, S.; Martínez-Martínez, S.; Ruiz-Meana, M.; et al. A robust method for quantitative high-throughput analysis of proteomes by 18O labeling. Mol. Cell. Proteom. 2011, 10, M110.003335. [Google Scholar] [CrossRef]
- Perez-Hernandez, D.; Gutiérrez-Vázquez, C.; Jorge, I.; López-Martín, S.; Ursa, A.; Sánchez-Madrid, F.; Vázquez, J.; Yáñez-Mó, M. The intracellular interactome of tetraspanin-enriched microdomains reveals their function as sorting machineries toward exosomes. J. Biol. Chem. 2013, 288, 11649–11661. [Google Scholar] [CrossRef]
- Shevchenko, A.; Wilm, M.; Vorm, O.; Mann, M. Mass Spectrometric Sequencing of Proteins from Silver-Stained Polyacrylamide Gels. Anal. Chem. 1996, 68, 850–858. [Google Scholar] [CrossRef] [PubMed]
- Ortea, I.; Ruiz-Sánchez, I.; Cañete, R.; Caballero-Villarraso, J.; Cañete, M.D. Identification of candidate serum biomarkers of childhood-onset growth hormone deficiency using SWATH-MS and feature selection. J. Proteom. 2018, 175, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Meyer, J.G.; Schilling, B. Clinical applications of quantitative proteomics using targeted and untargeted data-independent acquisition techniques. Expert Rev. Proteom. 2017, 14, 419–429. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Mok, T.S.; Lin, X.; Zhang, W.; Cui, Y.; Guo, J.; Chen, X.; Zhang, T.; Wang, T. SWATH-based proteomics identified carbonic anhydrase 2 as a potential diagnosis biomarker for nasopharyngeal carcinoma. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Karpievitch, Y.V.; Dabney, A.R.; Smith, R.D. Normalization and missing value imputation for label-free LC-MS analysis. BMC Bioinform. 2012, 13, S5. [Google Scholar] [CrossRef] [PubMed]





| Number of Proteins. 1st Analysis | Number of Proteins. 2nd Analysis | Number of Proteins. 3rd Analysis | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Samples | No Treat | NLC + ERT | ERT | No Treat | NLC + ERT | ERT | No Treat | NLC + ERT | ERT |
| HC | 683 | 1395 | 1612 | 1561 | 1668 | 919 | 737 | 1183 | |
| S.1 MoC | 730 | 960 | 923 | 1304 | 1623 | 1817 | |||
| S.2 MoC | 1220 | 1255 | 832 | 1022 | 793 | 1078 | |||
| S.3 MoC | 793 | 1658 | 1251 | 1594 | 1419 | 1104 | |||
| S.4 MoC | 611 | 1445 | 287 | 870 | 1403 | 586 | |||
| A. Upregulated Proteins in UMoC, Compared in MoC with ERT | |||
| Protein | Group | p-Value | FC |
| P29692 | Elongation factor 1-delta | 0.0341 | 2.1053 |
| P54727 | UV excision repair protein RAD23 homolog B | 0.0340 | 1.9325 |
| P16401 | Histone H1.5 | 0.0357 | 1.8159 |
| P26373 | 60S ribosomal protein L13 | 0.0449 | 1.8140 |
| O15143 | Actin-related protein 2/3 complex subunit 1B | 0.0329 | 1.7815 |
| P31942 | Heterogeneous nuclear ribonucleoprotein H3 | 0.0234 | 1.6284 |
| P07437 | Tubulin beta chain | 0.0361 | 1.6138 |
| Q07020 Q9Y4L1 | 60S ribosomal protein L18 Hypoxia up-regulated protein 1 | 0.0477 0.0104 | 1.4902 1.4313 |
| P13674 | Prolyl 4-hydroxylase subunit alpha-1 | 0.0299 | 1.3145 |
| P68104 | Elongation factor 1-alpha 1 | 0.0342 | 1.2451 |
| B. Upregulated Proteins in MoC with ERT, Compared in UMoC | |||
| Protein | Group | p-Value | FC |
| P07237 | Protein disulfide-isomerase | 0.0408 | 1.2058 |
| P78417 | Glutathione S-transferase omega-1 | 0.0425 | 1.2224 |
| O00299 | Chloride intracellular channel protein 1 | 0.0195 | 1.2303 |
| P23526 | Adenosylhomocysteinase | 0.0443 | 1.2358 |
| Q96S97 | Myeloid-associated differentiation marker | 0.0496 | 1.2446 |
| P62258 | 14-3-3 protein epsilon | 0.0079 | 1.2498 |
| P16152 | Carbonyl reductase [NADPH] 1 | 0.0199 | 1.2521 |
| P32119 | Peroxiredoxin-2 | 0.0385 | 1.2540 |
| P00387 | NADH-cytochrome b5 reductase 3 | 0.0087 | 1.2629 |
| P17655 | Calpain-2 catalytic subunit | 0.0287 | 1.2708 |
| P61981 | 14-3-3 Protein gamma | 0.0482 | 1.2743 |
| P60900 | Proteasome subunit alpha type-6 | 0.0270 | 1.2756 |
| O43707 | Alpha-actinin-4 | 0.0309 | 1.2781 |
| P27824 | Calnexin | 0.0254 | 1.2798 |
| P11413 | Glucose-6-phosphate 1-dehydrogenase | 0.0215 | 1.2906 |
| P08133 | Annexin A6 | 0.0055 | 1.2937 |
| P12814 | Alpha-actinin-1 | 0.0146 | 1.3017 |
| P22314 | Ubiquitin-like modifier-activating enzyme 1 | 0.0015 | 1.3163 |
| P54578 | Ubiquitin carboxyl-terminal hydrolase 14 | 0.0386 | 1.3214 |
| P49721 | Proteasome subunit beta type-2 | 0.0414 | 1.3568 |
| Q96AG4 | Leucine-rich repeat-containing protein 59 | 0.0085 | 1.3642 |
| P67812 | Signal peptidase complex catalytic subunit SEC11A | 0.0153 | 1.3671 |
| Q15404 | Ras suppressor protein 1 | 0.0172 | 1.3808 |
| P07996 | Thrombospondin-1 | 0.0379 | 1.3992 |
| O00629 | Importin subunit alpha-3 | 0.0386 | 1.4218 |
| Q9BS26 | Endoplasmic reticulum resident protein 44 | 0.0170 | 1.4754 |
| Q99798 | Aconitate hydratase, mitocondrial | 0.0357 | 1.4799 |
| P60953 | Cell division control protein 42 homolog | 0.0215 | 1.4806 |
| Q9Y3I0 | tRNA-splicing ligase RtcB homolog | 0.0506 | 1.4807 |
| P2484 | Myosin regulatory light polypeptide 9 | 0.0344 | 1.5135 |
| Q15758 | Neutral amino acid transporter B(0) | 0.0076 | 1.5970 |
| Q13724 | Mannosyl-oligosaccharide glucosidase | 0.0351 | 1.6011 |
| P08727 | Keratin, type I cytoskeletal 19 | 0.0315 | 1.6256 |
| P24941 | Cyclin-dependent kinase 2 | 0.0114 | 1.6367 |
| P61619 | Protein transport protein Sec61 subunit alpha isoform 1 | 0.0398 | 1.7619 |
| P08195 | 4F2 cell-surface antigen heavy chain | 0.0151 | 1.7870 |
| Q16881 | Thioredoxin reductase 1, cytoplasmic | 0.0472 | 1.8402 |
| P55060 | Exportin-2 | 0.0229 | 1.9550 |
| P04179 | Superoxide dismutase [Mn], mitocondrial | 0.0066 | 2.3668 |
| Q9NVD7 | Alpha-parvin | 0.0339 | 2.3802 |
| Q9HDC9 | Adipocyte plasma membrane-associated protein | 0.0129 | 2.4386 |
| P62314 | Small nuclear ribonucleoprotein Sm D1 | 0.0349 | 2.6814 |
| P78344 | Eukaryotic translation initiation factor 4 gamma 2 | 0.0332 | 10.9652 |
| P07237 | Protein disulfide-isomerase | 0.0408 | 1.2058 |
| P78417 | Glutathione S-transferase omega-1 | 0.0425 | 1.2224 |
| O00299 | Chloride intracellular channel protein 1 | 0.0195 | 1.2303 |
| P23526 | Adenosylhomocysteinase | 0.0443 | 1.2358 |
| Q96S97 | Myeloid-associated differentiation marker | 0.0496 | 1.2446 |
| P62258 | 14-3-3 protein epsilon | 0.0079 | 1.2498 |
| P16152 | Carbonyl reductase [NADPH] 1 | 0.0199 | 1.2521 |
| P32119 | Peroxiredoxin-2 | 0.0385 | 1.2540 |
| P00387 | NADH-cytochrome b5 reductase 3 | 0.0087 | 1.2629 |
| P17655 | Calpain-2 catalytic subunit | 0.0287 | 1.2708 |
| P61981 | 14-3-3 Protein gamma | 0.0482 | 1.2743 |
| P60900 | Proteasome subunit alpha type-6 | 0.0270 | 1.2756 |
| O43707 | Alpha-actinin-4 | 0.0309 | 1.2781 |
| P27824 | Calnexin | 0.0254 | 1.2798 |
| P11413 | Glucose-6-phosphate 1-dehydrogenase | 0.0215 | 1.2906 |
| P08133 | Annexin A6 | 0.0055 | 1.2937 |
| P12814 | Alpha-actinin-1 | 0.0146 | 1.3017 |
| P22314 | Ubiquitin-like modifier-activating enzyme 1 | 0.0015 | 1.3163 |
| P54578 | Ubiquitin carboxyl-terminal hydrolase 14 | 0.0386 | 1.3214 |
| P49721 | Proteasome subunit beta type-2 | 0.0414 | 1.3568 |
| Q96AG4 | Leucine-rich repeat-containing protein 59 | 0.0085 | 1.3642 |
| P67812 | Signal peptidase complex catalytic subunit SEC11A | 0.0153 | 1.3671 |
| Q15404 | Ras suppressor protein 1 | 0.0172 | 1.3808 |
| P07996 | Thrombospondin-1 | 0.0379 | 1.3992 |
| O00629 | Importin subunit alpha-3 | 0.0386 | 1.4218 |
| Q9BS26 | Endoplasmic reticulum resident protein 44 | 0.0170 | 1.4754 |
| Q99798 | Aconitate hydratase, mitocondrial | 0.0357 | 1.4799 |
| P60953 | Cell division control protein 42 homolog | 0.0215 | 1.4806 |
| Q9Y3I0 | tRNA-splicing ligase RtcB homolog | 0.0506 | 1.4807 |
| P2484 | Myosin regulatory light polypeptide 9 | 0.0344 | 1.5135 |
| Q15758 | Neutral amino acid transporter B(0) | 0.0076 | 1.5970 |
| Q13724 | Mannosyl-oligosaccharide glucosidase | 0.0351 | 1.6011 |
| P08727 | Keratin, type I cytoskeletal 19 | 0.0315 | 1.6256 |
| P24941 | Cyclin-dependent kinase 2 | 0.0114 | 1.6367 |
| P61619 | Protein transport protein Sec61 subunit alpha isoform 1 | 0.0398 | 1.7619 |
| P08195 | 4F2 cell-surface antigen heavy chain | 0.0151 | 1.7870 |
| Q16881 | Thioredoxin reductase 1, cytoplasmic | 0.0472 | 1.8402 |
| P55060 | Exportin-2 | 0.0229 | 1.9550 |
| P04179 | Superoxide dismutase [Mn], mitocondrial | 0.0066 | 2.3668 |
| Q9NVD7 | Alpha-parvin | 0.0339 | 2.3802 |
| Q9HDC9 | Adipocyte plasma membrane-associated protein | 0.0129 | 2.4386 |
| P62314 | Small nuclear ribonucleoprotein Sm D1 | 0.0349 | 2.6814 |
| P78344 | Eukaryotic translation initiation factor 4 gamma 2 | 0.0332 | 10.9652 |
| A. Upregulated Proteins in UMoC, Compared in MoC with NLC + ERT | |||
| Protein | Group | p-Value | FC |
| P04843 | Dolichyl-diphosphooligosaccharide-protein glycosyltransferase subunit 1 | 0.0416 | 1.2453 |
| Q7KZF4 | Staphylococcal nuclease domain-containing protein 1 | 0.0308 | 1.3226 |
| Q99829 | Copine-1 | 0.0098 | 1.5958 |
| P43686 | 26S proteasome regulatory subunit 6B | 0.0054 | 1.8527 |
| O75436 | Vacuolar protein sorting-associated protein 26A | 0.0499 | 1.2358 |
| P01024 | Complement C3 | 0.0246 | 2.0811 |
| P02788 | Lactotransferrin | 0.0130 | 2.2108 |
| Q9P219 | Protein Daple | 0.0205 | 2.3711 |
| P30084 | Enoyl-CoA hydratase, mitochondrial | 0.0377 | 2.3790 |
| P19823 | Inter-alpha-trypsin inhibitor heavy chain H2 | 0.0491 | 2.3791 |
| P00367 | Glutamate dehydrogenase 1, mitocondrial | 0.0225 | 2.5313 |
| Q7L1Q6 | Basic leucine zipper and W2 domain-containing protein 1 | 0.0495 | 2.5351 |
| Q7LBC6 | Lysine-specific demethylase 3B | 0.0124 | 2.5605 |
| P02765 | Alpha-2-HS-glycoprotein | 0.0175 | 2.6800 |
| B. Upregulated Proteins in MoC with NLC + ERT, Compared in UMoC | |||
| Protein | Group | p-Value | FC |
| P22090 | 40S ribosomal protein S4, Y isoform 1 | 0.0134 | 2.6287 |
| Q9BQB6 | Vitamin K epoxide reductase complex subunit 1 | 0.0413 | 2.5833 |
| P05783 | Keratin, type I cytoskeletal 18 | 0.0048 | 2.4465 |
| P51665 | 26S proteasome non-ATPase regulatory subunit 7 | 0.0118 | 2,1633 |
| O75368 | SH3 domain-binding glutamic acid-rich-like protein | 0.0004 | 2.0888 |
| Q96AT9 | Ribulose-phosphate 3-epimerase | 0.0115 | 2.0690 |
| P48147 | Prolyl endopeptidase | 0.0276 | 2.0209 |
| P30498 | HLA class I histocompatibility antigen, B-78 alpha chain | 0.0315 | 1.8355 |
| P04179 | Superoxide dismutase [Mn], mitochondrial | 0.0153 | 1.8015 |
| Q7Z4H8 | KDEL motif-containing protein 2 | 0.0292 | 1.7891 |
| Q12797 | Aspartyl/asparaginyl beta-hydroxylase | 0.0031 | 1.7517 |
| Q06323 | Proteasome activator complex subunit 1 | 0.0064 | 1.7109 |
| O15260 | Surfeit locus protein 4 | 0.0346 | 1.6967 |
| Q86UP2 | Kinectin | 0.0338 | 1.6722 |
| P84077 | ADP-ribosylation factor 1 | 0.0069 | 1.5850 |
| P62857 | 40S ribosomal protein S28 | 0.0021 | 1.5181 |
| P17987 | T-complex protein 1 subunit alpha | 0.0435 | 1.5070 |
| P36578 | 60S ribosomal protein L4 | 0.0492 | 1.4908 |
| P09651 | Heterogeneous nuclear ribonucleoprotein A1 | 0.0033 | 1.4863 |
| O00629 | Importin subunit alpha-3 | 0.0228 | 1.4728 |
| Q15293 | Reticulocalbin-1 | 0.0212 | 1.4704 |
| P62805 | Histone H4 | 0.0123 | 1.4701 |
| P60953 | Cell division control protein 42 homolog | 0.0168 | 1.4677 |
| P18669 | Phosphoglycerate mutase 1 | 0.0088 | 1.4651 |
| P48643 | T-complex protein 1 subunit epsilon | 0.0003 | 1.4511 |
| Q99798 | Aconitate hydratase, mitochondrial | 0.0225 | 1.4494 |
| P00558 | Phosphoglycerate kinase 1 | 0.0053 | 1.4426 |
| P60174 | Triosephosphate isomerase | 0.0062 | 1.4168 |
| Q16222 | UDP-N-acetylhexosamine pyrophosphorylase | 0.0459 | 1.4093 |
| P11021 | Endoplasmic reticulum chaperone BiP | 0.0005 | 1.3987 |
| P26599 | Polypyrimidine tract-binding protein 1 | 0.0205 | 1.3867 |
| Q14152 | Eukaryotic translation initiation factor 3 subunit A | 0.0272 | 1.3757 |
| P09936 | Ubiquitin carboxyl-terminal hydrolase isozyme L1 | 0.0402 | 1.3509 |
| P34897 | Serine hydroxymethyltransferase, mitochondrial | 0.0384 | 1.3406 |
| P38646 | Stress-70 protein, mitochondrial | 0.0023 | 1.3405 |
| P30101 | Protein disulfide-isomerase A3 | 0.0012 | 1.3319 |
| P17655 | Calpain-2 catalytic subunit | 0.0433 | 1.3318 |
| P07237 | Protein disulfide-isomerase | 0.0020 | 1.3316 |
| P18124 | 60S ribosomal protein L7 | 0.0196 | 1.3239 |
| P06733 | Alpha-enolase | 0.0031 | 1.3183 |
| P27797 | Calreticulin | 0.0271 | 1.3103 |
| Q96AY3 | Peptidyl-prolyl cis-trans isomerase FKBP10 | 0.0103 | 1.2954 |
| P62269 | 40S ribosomal protein S18 | 0.0455 | 1.2947 |
| Q9NR31 | GTP-binding protein SAR1a | 0.0097 | 1.2880 |
| Q13148 | TAR DNA-binding protein 43 | 0.0483 | 1.2741 |
| P61978 | Heterogeneous nuclear ribonucleoprotein K | 0.0396 | 1.2736 |
| P46777 | 60S ribosomal protein L5 | 0.0015 | 1.2716 |
| P62701 | 40S ribosomal protein S4, X isoform | 0.0060 | 1.2709 |
| P62826 | GTP-binding nuclear protein Ran | 0.0164 | 1.2660 |
| P05388 | 60S acidic ribosomal protein P0 | 0.0071 | 1.2656 |
| O95394 | Phosphoacetylglucosamine mutase | 0.0247 | 1.2547 |
| O60701 | UDP-glucose 6-dehydrogenase | 0.0232 | 1.2538 |
| P62820 | Ras-related protein Rab-1A OS=Homo sapiens SV=3 | 0.0094 | 1.2514 |
| Q8NBJ5 | Procollagen galactosyltransferase 1 | 0.0389 | 1.2479 |
| Q9ULV4 | Coronin-1C OS=Homo sapiens | 0.0338 | 1.2455 |
| Q14315 | Filamin-C OS=Homo sapiens | 0.0186 | 1.2394 |
| Q14697 | Neutral alpha-glucosidase AB | 0.0435 | 1.2386 |
| P00338 | L-lactate dehydrogenase A chain | 0.0260 | 1.2371 |
| Q6NZI2 | Caveolae-associated protein 1 | 0.0295 | 1.2301 |
| P07195 | L-lactate dehydrogenase B chain | 0.0202 | 1.2247 |
| O00299 | Chloride intracellular channel protein 1 | 0.0501 | 1.2232 |
| O43707 | Alpha-actinin-4 | 0.0311 | 1.2167 |
| P12111 | Collagen alpha-3(VI) chain | 0.0378 | 1.2093 |
| Samples | Sex | Diagnosis | Phenotype |
|---|---|---|---|
| S.1 MoC | Male | MPS IVA | Classic |
| S.2 MoC | Female | MPS IVA | Classic |
| S.3 MoC | Male | MPS IVA | Classic |
| S.4 MoC | Female | MPS-IVA | Classic |
| Samples | Untreated | NLC + ERT | ERT |
|---|---|---|---|
| H.C | X | X | X |
| S.1 MoC | X | X | |
| S.2 MoC | X | X | |
| S.3 MoC | X | X | |
| S.4 MoC | X | X |
| Samples | Before Treatment (nM/h/mg) | After Treatment (nM/h/mg) |
|---|---|---|
| S.1 MoC | 0.0 | 1.6 |
| S.2 MoC | 0.0 | 4.8 |
| S.3 MoC | 0.0 | 6.5 |
| S.4 MoC | 0.0 | 8.9 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Álvarez, J.V.; Bravo, S.B.; García-Vence, M.; De Castro, M.J.; Luzardo, A.; Colón, C.; Tomatsu, S.; Otero-Espinar, F.J.; Couce, M.L. Proteomic Analysis in Morquio A Cells Treated with Immobilized Enzymatic Replacement Therapy on Nanostructured Lipid Systems. Int. J. Mol. Sci. 2019, 20, 4610. https://doi.org/10.3390/ijms20184610
Álvarez JV, Bravo SB, García-Vence M, De Castro MJ, Luzardo A, Colón C, Tomatsu S, Otero-Espinar FJ, Couce ML. Proteomic Analysis in Morquio A Cells Treated with Immobilized Enzymatic Replacement Therapy on Nanostructured Lipid Systems. International Journal of Molecular Sciences. 2019; 20(18):4610. https://doi.org/10.3390/ijms20184610
Chicago/Turabian StyleÁlvarez, J. Víctor, Susana B. Bravo, María García-Vence, María J. De Castro, Asteria Luzardo, Cristóbal Colón, Shunji Tomatsu, Francisco J. Otero-Espinar, and María L. Couce. 2019. "Proteomic Analysis in Morquio A Cells Treated with Immobilized Enzymatic Replacement Therapy on Nanostructured Lipid Systems" International Journal of Molecular Sciences 20, no. 18: 4610. https://doi.org/10.3390/ijms20184610
APA StyleÁlvarez, J. V., Bravo, S. B., García-Vence, M., De Castro, M. J., Luzardo, A., Colón, C., Tomatsu, S., Otero-Espinar, F. J., & Couce, M. L. (2019). Proteomic Analysis in Morquio A Cells Treated with Immobilized Enzymatic Replacement Therapy on Nanostructured Lipid Systems. International Journal of Molecular Sciences, 20(18), 4610. https://doi.org/10.3390/ijms20184610








