Abstract
Mitochondria are vital cellular organelles involved in a plethora of cellular processes such as energy conversion, calcium homeostasis, heme biogenesis, regulation of apoptosis and ROS reactive oxygen species (ROS) production. Although they are frequently depicted as static bean-shaped structures, our view has markedly changed over the past few decades as many studies have revealed a remarkable dynamicity of mitochondrial shapes and sizes both at the cellular and intra-mitochondrial levels. Aberrant changes in mitochondrial dynamics and cristae structure are associated with ageing and numerous human diseases (e.g., cancer, diabetes, various neurodegenerative diseases, types of neuro- and myopathies). Another unique feature of mitochondria is that they harbor their own genome, the mitochondrial DNA (mtDNA). MtDNA exists in several hundreds to thousands of copies per cell and is arranged and packaged in the mitochondrial matrix in structures termed mt-nucleoids. Many human diseases are mechanistically linked to mitochondrial dysfunction and alteration of the number and/or the integrity of mtDNA. In particular, several recent studies identified remarkable and partly unexpected links between mitochondrial structure, fusion and fission dynamics, and mtDNA. In this review, we will provide an overview about these recent insights and aim to clarify how mitochondrial dynamics, cristae ultrastructure and mtDNA structure influence each other and determine mitochondrial functions.
1. Mitochondrial Membrane Structure and Cristae Biogenesis
Mitochondrial shape is constantly adapting at the cellular and intra-mitochondrial levels in response to energetic and/or developmental cues. Mitochondria are enclosed by two membranes where the inner membrane (IM) characteristically folds inwards to form cristae. Cristae house the respiratory chain complexes and the F1FO ATP synthase representing the major functional units for energy conversion. The rest of the IM, located parallel to the outer membrane (OM), is termed the inner boundary membrane (IBM). The current knowledge views the cristae membrane (CM) not just as mere invaginations of the IM into the matrix; instead, the CM is a structurally and functionally distinct subcompartment of the IM. This distinction is in part due to the recent rediscovery of crista junctions (CJs) with a diameter of 12–40 nm at the neck of cristae [1,2,3]. Based on this unique structure, CJs were proposed to restrict the normal passage of proteins, metabolites and protons both towards and away from cristae, thus separating the mitochondria into several subcompartments [4,5]. This implies that a mitochondrion has three aqueous subcompartments, namely, the intermembrane space (IMS) between the IM and OM, the intracristal space (ICS) and the matrix.
The shape, size and number of cristae are constantly changing based on varying energy demands or other physiological changes. Almost half a century ago, Charles Hackenbrock reported ultrastructural alterations in mitochondrial cristae to occur upon metabolic changes [6,7]. This is classically described as the transition between orthodox (matrix expanded, state IV respiration, low ADP levels) and condensed (matrix compacted, state III respiration, high ADP levels) states of mitochondria, which were suggested to be critical for ATP production. Remodelling of cristae also occurs during apoptosis ensuring release of cytochrome c from the ICS to the cytosol resulting in a downstream cascade of caspase activation and cell death [8]. It is evident that aberrant and altered cristae are associated with several human diseases [3] but what are the molecular players required for cristae formation and maintenance? Albeit several factors are reported to modulate cristae morphology, we will first focus on three major protein complexes that are shown to be involved in the biogenesis of cristae and/or CJs: OPA1 (Optic Atrophy 1), F1FO ATP synthase and the MICOS (Mitochondrial Contact Site and Cristae Organizing System) complex.
1.1. Optic Atrophy Type 1 (OPA1)
OPA1 is a large dynamin-related GTPase present at the IM which plays a dual role in IM fusion as well as cristae biogenesis. Heterozygous mutations in OPA1 cause optic atrophy associated with deafness and dementia [9,10,11], while a rare homozygous mutation is reported to be associated with early encephalomyopathy, cardiomyopathy and early death [12]. The regulation of OPA1 is complicated as it comprises 8 alternate splice variants and two proteolytic cleavage sites, which generate several forms of OPA1 broadly classified as long, IM-anchored forms (full length) or short, proteolytically cleaved forms [13,14]. Although the long forms of OPA1 are sufficient to manage its mitochondrial fusion activity [15], joint action of long and short OPA1 forms is proposed to be necessary to keep CJs closed, thereby limiting the diffusion/release of metabolites including cytochrome c [16,17]. The proposed role of s-OPA1 is to manage mitochondrial fission, cristae biogenesis or energetics [15,18,19]. Deletion of Opa1 was characterized by the presence of balloon-like swollen cristae and widening of CJs modulating the release of cytochrome c to initiate apoptosis independent of its fusion activity [16]. How does OPA1 dually regulate IM fusion and cristae biogenesis? Based on elegant experiments, Walter Neupert and colleagues proposed recently that the formation of lamellar sheets of cristae is dependent on a preceding IM fusion event mediated by Mgm1, the baker’s yeast ortholog of OPA1 [20]. They suggest that post-OM fusion, the juxtaposed IM are tethered by Mgm1 to initiate the IM fusion along the IM-OM contact sites, which would generate a cristae-shaped sac protruding into the matrix. In contrast to this, tubular cristae are presumably formed by invaginations of IM. Similar and other models of cristae formation were discussed earlier [3], yet further experimental evidence supporting or excluding different models is urgently needed.
1.2. F1FO ATP Synthase
The F1FO ATP synthase is best known for its essential role in generating ATP using energy stored in an electrochemical proton gradient across the IM. A rather novel role of this complex is to determine cristae ultrastructure. Monomeric F1FO ATP synthase consists of F1 (catalytic) domain facing the matrix and FO embedded in the CM. Although the monomeric enzyme is sufficient for ATP production, the F1FO ATP synthase is able to form dimers and oligomers, consisting of rows of dimers, which not only control the stability of this multiprotein complex but also shape the IM and thus contribute to cristae formation. Formation of F1FO ATP synthase dimers occurs via distinct interactions between adjacent FO domains, e.g., via the dimer-specific subunits e and g or the subunit ATP4 of this complex [21,22,23]. The stable association of two monomers at a defined angle leads to bending of the IM creating protrusions and a strong positive curvature at cristae tips [24,25,26,27,28]. Indeed, absence of dimerization of this complex by deleting dimer-specific subunits e or g results in perturbed cristae structure [22].
1.3. Mitochondrial Contact Site and Cristae Organizing System (MICOS) Complex
In baker’s yeast, Mic60/Fcj1 located at CJs was shown to be required for their formation [29]. Overexpression of Mic60/Fcj1 resulted in branched cristae resembling internal CJs not in close proximity to the OM. Mic60 is an important subunit of the MICOS complex; a large oligomeric complex present at CJs [30,31,32]. It is required for the formation of CJs and contact sites between the IM and OM and is highly conserved across phyla [33,34]. The various subunits of the MICOS complex are identified in yeast and mammalian model systems and are termed as MicX/MICX (with X indicating the approximate molecular weight) according to a uniform nomenclature proposed in 2014 [35]. Hitherto, seven main subunits of the MICOS complex have been identified in mammals, namely, MIC60/Mitofilin/Fcj1, MIC10/Minos1, MIC19/CHCHD3, MIC25/CHCHD6, MIC13/Qil1, MIC26/APOO and MIC27/APOOL [36,37]. Lack of individual subunits of MICOS complex results in loss or reduction of CJs and cristae detaching from the IBM forming concentric rings resembling onion-like slices in sections of electron micrographs [30,31,32]. The extent of the phenotype varies with the subunit deleted, so that they are classified as essential/core or peripheral components of the MICOS complex. MIC60 is considered as the pioneer of the MICOS complex as it is required for the formation of CJs and contact sites between IM and OM owing to its interaction with several OM proteins (TOM complex, SAM/TOB, SCL25A46, metaxin) [38,39,40,41,42,43,44]. Mic10 is considered the other core component as its deletion also leads to a virtual loss of CJs culminating in the appearance of cristae stacks appearing as onion-like slices. Mic10 is a small protein with a conserved glycine-motif in the transmembrane domain required for self-oligomerization and CJs formation [45,46,47]. Mic60 and Mic10 have the capability to bend membranes that control the formation of CJs [45,48,49]. MIC13/Qil1 is essential for formation of CJs in mammalian cells [50,51]. Loss of MIC13 causes a destabilization of MIC10, MIC26 and MIC27 while other subunits MIC60, MIC19 and MIC25 remain intact and can form a stable subcomplex. This indicates a hierarchy in the assembly pathway of the MICOS complex, suggesting two distinct subcomplexes, the MIC60-19-25 and the MIC10-13-26-27 subcomplex. MIC26 and MIC27 belong to the protein family of apolipoproteins and are deemed non-core components of the MICOS complex [52,53,54]. MIC27 can bind to cardiolipin, the signature lipid of mitochondria [53]. MIC26 exists in two forms: the non-glycosylated form is mitochondrially targeted and a subunit of the MICOS complex, while the glycosylated form is translocated to the ER and subsequently secreted [52]. Both MIC26 and MIC27 are required for maintenance of cristae morphology and respiration [52,53]. MIC19 and MIC25 on the other hand belong to CHCHD family of proteins. While the physiological role of MICOS is still not completely understood, depletion of its subunits affect several mitochondrial functions such as respiration, protein import, lipid transport, mtDNA organization, apoptosis, and autophagy. Altered levels of MICOS components are associated with a number of pathologies such as diabetes, cardiomyopathy, epilepsy, Down syndrome and Parkinson’s disease (PD) [3,55]. Mutations in MIC13 are reported to cause severe forms of mitochondrial encephalopathy [56,57,58,59], and coding variants of MIC60 present in the mitochondrial targeting sequence (MTS) are found in PD patients [60].
In addition to increasing number of studies revealing the individual roles of OPA1, the MICOS complex, and the F1FO ATP synthase in determining cristae morphology, there is evidence these complexes functionally interact at different levels. In baker’s yeast, it was shown that Mic10 binds to oligomeric F1FO ATP synthase and both Mic10 and Mic26 promote its oligomerization [61,62]. Mic60/Fcj1 play an antagonist role to subunits e and g of the F1FO ATP synthase [29]. It was suggested that this interplay modulates the curvature of the IM locally near CJs and thereby regulates CJ formation. In addition, it was reported that OPA1 favors oligomerization of F1FO ATP synthase to protect mitochondria from respiratory stress [63]. Moreover, OPA1 was suggested to function epistatic to the MICOS complex in order to regulate the width of CJs [42] and Opa1 was shown to physically interact with Mic60 [64]. Albeit Opa1 clearly affects CJs architecture, CJs can still be formed in the absence of Opa1 demonstrating that this complex is not absolutely required for formation of CJs per se [64]. Overall, it appears that OPA1 has an important regulatory role in determining cristae and CJ structure which acts in concert with the MICOS complex as well as the F1FO ATP synthase. Interestingly, all three complexes are reported to affect the integrity and/or inheritance of mtDNA as discussed in the next sections.
2. Organization and Structure of mtDNA
Mitochondrial DNA (mtDNA) was initially visualized as fibrous structures within the mitochondria of developing chick embryo [65,66,67]. MtDNA was shown to be of universal occurrence across phyla ranging from protozoa to plants to higher vertebrates [68] possessing a circular topology of around 5 µm circumference in osmotically disrupted mitochondria [69]. Each mitochondrion released an average of two to six circular DNA molecules present as monomer or dimers either in extended or partially coiled configuration upon osmotic stress. In addition, 80% of mtDNA monomers remained bound to mitochondrial membranes [70]. It was later shown that mtDNA was associated to membrane-like structures near the D-loop or origin of mtDNA replication [71] which was later confirmed by the observation that mtDNA is located next to the IM when visualized by immunogold labelling using an anti-DNA antibody [72]. This was further corroborated by using correlative 3D super-resolution fluorescence iPALM imaging followed by FIB (focused ion beam)-SEM (Scanning electron microscopy) which showed that mtDNA is in close proximity with the CMs [73]. Discoveries leading to the identification of mtDNA as a separate entity paved the way for the sequencing of the entire human mitochondrial genome harboring 16,569 bases encoding 2 rRNAs, 22 tRNAs and 13 mRNAs [74]. MtDNA contains very few noncoding bases between these genes and even has overlapping genes accounting for high economy of mtDNA. MtDNA can lack fully encoded termination codons where the termination is completed post-transcriptionally with addition of the 3’-polyA tail, and contains exceptions from the universal genetic code accounting for its uniqueness [74]. The tRNA punctuation model was proposed in the early 1980’s according to which tRNA processing of primary transcripts, containing mRNAs and rRNAs, occurs at sites of interspersed tRNAs. Thus, tRNAs act as breaks or punctuations for genes encoding proteins or rRNAs [75,76]. This organization and its physiological consequences on translation are elegantly described elsewhere [77,78]. In addition, a comprehensive description of mtDNA replication and transcription in mitochondria has been well covered earlier [79,80]. Here, we summarize some necessary aspects in brief. The minimal mtDNA replisome consists of the helicase TWINKLE, which unwinds the mtDNA duplex template, with the mitochondrial single-stranded DNA-binding protein (mtSSB) stabilizing this state and allowing replication by DNA polymerase γ (POLγ). The transcription apparatus in mitochondria consists of a mitochondrial transcription factor A (TFAM), mitochondrial transcription factors B1 & B2 (TFB1M and TFB2M), mitochondrial transcription elongation factor (TEFM) and mitochondrial RNA polymerase (POLRMT) [79,80,81]. POLRMT cannot bind directly to promoter DNA and requires the assistance of TFAM and TFB2M. TFAM plays an important role in mitochondrial transcription by distorting mtDNA to induce negative supercoils [82] and forces the promoter DNA to undergo a U-turn on the mtDNA [83]. TEFM is present at the promoter before transcription begins and stimulates the activity of POLRMT besides assisting POLRMT to transcribe longer transcripts [79,80,81]. A major difference to how DNA is packed and organized in mitochondria as opposed to the nucleus is the absence of histones in mitochondria. Still, mtDNA is known to exist as foci within the mitochondrial matrix termed mt-nucleoids. Using confocal microscopy, the initial studies concluded that the number of mitochondrial nucleoids per cell ranged from 450 to 800 in cultured human cells with preliminary data showing an average ranging from 2 to 10 mtDNA copies per nucleoid [72,84]. The numbers of mtDNA copies per nucleoid and cell were 2–3 times higher in tumor cell lines compared to non-tumor cell lines such as fibroblasts. Nucleoids are dynamic structures and undergo fusion and fission dynamics, yet this is not thought to involve a major exchange of mtDNA content [72]. Indeed, when two heterologous mtDNA populations from different cells containing non-overlapping deletions were fused, fully functional protein complementation was achieved without exchanging mtDNA [85]. With the advent of diffraction-unlimited imaging techniques such as STED super-resolution microscopy, individual nucleoids marked as a single entity, limited by resolution of confocal microscopy, could be further resolved [86]. Hence, the average number of mt-nucleoids in a mitochondrion revealed by confocal microscopy was not represented accurately, as mitochondria imaged using STED super-resolution microscopy revealed roughly 1.6 times more nucleoids per cell. Consequently, the number of mtDNA molecules per nucleoid had to be revised to 1.4. The diameter of the mt-nucleoids has to be adapted from 250 nm to 100 nm in various cell types although a single mtDNA molecule has a contour length of around 5 µm. Several proteins distinct from nuclear histones help to pack mtDNA. TFAM (mitochondrial transcription factor A) is an essential protein interacting with mtDNA, playing an important role in transcription of mtDNA. Intriguingly, it possesses an additional role of packaging mtDNA. Increasing the ratio of TFAM molecules to number of mtDNA bases, from 1 TFAM molecule per 150 bp to 6 bp of mtDNA, progressively and consistently increased the mtDNA compaction of nucleoids [87]. This is mediated by the ability of TFAM molecules to bind across a DNA strand in a single mtDNA molecule as deciphered from in vitro reconstitution experiments using rotary shadowing electron microscopy (EM). In yeast, Abf2 is the mitochondrial DNA-packaging protein. Nucleoids undergo dynamic remodelling by forming either open or closed structures by modulating the ratio of Abf2 to mtDNA [88].
What is known about the in vivo role of TFAM? Murine whole body knockouts (KOs) of Tfam were lethal around embryonic day e8.5 to e10.5 showing that TFAM plays a critical role in embryogenesis [89]. MtDNA was absent at e8.5 in Tfam knockouts. In addition, KO embryos at e8.5 showed accumulation of enlarged mitochondria together with disorganized cristae and respiratory chain dysfunction as revealed by COX-SDH staining pattern [89]. A conditional knockout of Tfam in the heart and skeletal muscle using Ckmm-cre resulted in dilated cardiomyopathy where the animals succumbed between 2–4 weeks of age. At an ultrastructural level, EM of myocardium revealed enlarged mitochondria and vesicle-like cristae together with respiratory deficiency as observed in the whole body knockouts of Tfam [90]. In another conditional KO model, where Tfam was deleted exclusively in the skeletal muscle, mice suffered from myopathy and electron micrographs revealed enlarged mitochondria with abnormal cristae [91]. Conditional KO of Tfam in dopamine neurons causes reduced mtDNA expression associated with progressive neurodegeneration of dopamine neurons, impairment of motor function in adulthood and formation of intraneuronal inclusions containing mitochondria. These mitochondria exhibited abnormal mitochondrial membranes with vacuolization in some cases [92]. In fact, it was shown that ageing and PD were positively correlated with high levels of mtDNA deletions in neurons of the substantia nigra [93]. Substantia nigra sections of PD patients showed a decreased number of neurons which occupied less area characterized by the presence of classical Lewy bodies. Surprisingly, unhealthy mitochondria with deranged cristae and electron-dense deposits were also found in substantia nigra [94]. Mutations in PINK1 also cause PD [95]. At a molecular level, it was recently identified that PINK1 phosphorylates MIC60 to stabilize its oligomerization placing MIC60 function downstream of PINK1 [60]. Intriguingly, some PD patients also displayed mutations in the MTS of MIC60 laying emphasis on the PINK1-MIC60 pathway. Overall, depletion of Tfam results in abnormalities in mitochondrial morphology, function, cristae organization and reduction in mtDNA level, pointing towards a complex interplay between them. In addition, whole body loss of TWINKLE also results in embryonic lethality at e8.5 [96]. Conditional KOs of Twinkle in the heart and skeletal muscle led to a decrease in the mtDNA content, mitochondrial transcripts and respiratory chain assembly. A homozygous knockin of the mutant Polγ mutator mouse model, which lacks its proof-reading activity, showed a 3 to 5 fold increase in the frequency of point mutations culminating in premature ageing, weight loss, hair loss, kyphosis, osteoporosis, and heart enlargement [97]. These studies indicate a functional link between PD, reduced mtDNA and occurrence of deformed cristae.
A variety of approaches have been used to decipher the mt-nucleoid proteome. (1) A considerable amount of proteins associated with mtDNA were discovered by classical coimmunoprecipitation (coIP), using antibodies against proteins already known to be present in mt-nucleoids (e.g., TFAM). Optimization of the methods of purifying mitochondrial nucleoids in Hela cells coupled with mild lysis using nonionic detergents and sedimentation followed by coIP using anti-TFAM and anti-mtSSB antibodies led to the identification of proteins interacting with mtDNA [98]. Coimmunoprecipitation with these two antibodies mostly yielded common proteins which were broadly classified into categories belonging to mtDNA replication and transcription, mtDNA binding and metabolism, mitochondrial chaperones and other miscellaneous proteins having a variety of other roles. (2) In a more stringent approach, formaldehyde-crosslinking was performed on mt-nucleoids in the presence of ionic detergents and high salt to release proteins non-covalently bound to mtDNA. The subset of proteins found in both studies included TFAM, mtSSB and other proteins involved in mtDNA replication and transcription. Interestingly, a nucleoid model consisting of core and peripheral layered structure was proposed. According to this, the inner core contained proteins obtained after formaldehyde-crosslinking and included proteins of mtDNA replication and transcription. The outer layer proteins obtained in native nucleoids included Hsp60, ATAD3 and prohibitins [99]. In fact, prohibitin 1 was proposed to maintain the stability of mtDNA copy number by regulating the TFAM levels [100]. ATAD3 has the capacity to bind to displacement loops. This interaction could be modulated leading to dissociation or association of mt-nucleoids [101]. (3) Recently, a proximity biotinylation assay, using Twinkle-APEX2 followed by mass spectrometry, was successfully used to decipher the nucleoid proteome [102]. Some novel proteins such as FASTKD1, C7ORF55 and NDUFS6 were found in addition to corroborating the known proteins present in nucleoids.
3. Interplay between Mitochondrial Dynamics and mtDNA
At the cellular level, mitochondria are arranged as an interconnected network with long and short tubules. Described as early as 1914 [103] and rediscovered later [104,105], mitochondria are highly dynamic organelles that constantly undergo fusion and fission events and move within cells. By using the power of yeast genetics, initial pioneering studies led to identification of several proteins localized both on the OM and IM responsible for fusion and fission [106,107,108]. In mammals, mitochondria undergo fusion with the help of large GTPases, Mitofusins 1 and 2 (Mfn1 and Mfn2) for the OM and OPA1 for the IM. Disruption of mitochondrial fusion results in fragmented mitochondria due to ongoing fission. Fusion is required for proper inheritance of mtDNA and content mixing, which helps to complement mitochondria, ensuring proper mitochondrial health [109,110]. For mitochondrial fusion, the outer membranes of apposing mitochondria are first tethered and juxtaposed via homo- or hetero-oligomerization of MFN1 and MFN2, and after GTP hydrolysis, a conformational change needed for mitochondrial fusion is induced [111,112,113,114]. When isolated mitochondria were subjected to classical protease treatment followed by western blotting using antibodies specific for certain domains of mitofusins, it was deciphered that mitofusins harbor two transmembrane segments [115]. However, recently it was also suggested that the helical repeats HR2 are present in IMS and that mitofusins only harbor one transmembrane segment [116,117]. Mutations in MFN2 are linked to Charcot-Marie-Tooth neuropathy type 2A (CMT2A) [118]. Fission in mammals is coordinated by another large dynamin-like GTPase, namely, DRP1. DNM2, MFF (mitochondrial fission factor), MID49, MID51 (mitochondrial dynamics proteins of 49 and 51 kDa) and FIS1 (mitochondrial fission 1 protein) are other factors which play a role in mitochondrial fission [119,120,121,122,123,124,125,126]. Loss of mitochondrial fission results in hyperfused mitochondria due to ongoing fusion. Mitochondrial fission has been proposed to ensure proper transport of mitochondria (e.g., in neurons) and to promote apoptosis [110,127]. Mitochondrial dysfunction results in alteration of mitochondrial dynamics and, combined with the selective removal of such damaged mitochondria by mitophagy, is critical for mitochondrial quality control [128,129,130,131,132]. DRP1 is recruited to mitochondria by several adaptors present at OM of mitochondria, namely, MFF, MID49, MID51 and FIS1 [133,134,135]. DRP1 causes mitochondrial constriction but DYN2 was proposed to work in collaboration with DRP1 to perform the final scission of the membranes [136]. However, more recent studies showed that all three dynamin proteins including DYN2 (DNM2) are dispensable for mitochondrial and peroxisomal fission [137,138]. DRP1 was shown to possess constricting as well as severing ability [137]. Apart from these molecules, ER contact with mitochondria and actin cytoskeleton also play crucial roles in mitochondria division [139,140]. Mutation in DRP1 leads to manifestation of severe defects and early death including childhood epileptic encephalopathy, microcephaly and optic atrophy [141,142,143].
Next we focus on the question ‘Why does alteration of mitochondrial dynamics result in dual problems, namely, aberrant cristae and loss of mtDNA and/or aberrancies in mt-nucleoid morphology?’ Initial observations in yeast showed that deletion of the fusion factors Fzo1 and Mgm1, located on the OM and IM, respectively, caused loss of mtDNA [144,145,146,147]. Similarly, deletion of Ugo1 which physically links Fzo1 and Mgm1 also resulted in mtDNA loss [148,149] providing evidence that defective fusion leads to loss of mtDNA. Mfn1 and Mfn2 null mice were never obtained with embryonic lethality occurring at midgestation around E12 stage [120]. It was shown that mtDNA levels were not altered in Mfn (Double Knockout) DKO mice although the mitochondrial fusion was strongly abolished resulting in fragmented mitochondria. It is possible that Mfn DKO mice exhibit lethality around the time mtDNA levels begin to drop, providing an alternate explanation for detecting no change in mtDNA when mitochondrial OM fusion is inhibited. Embryonic lethality of Mfn2 null mice encouraged researchers to study conditional KOs of Mfns in various tissues. Intriguingly, loss of Mfn2 in the cerebellum resulted in Purkinje cells (PCs) with swollen mitochondria having abnormal vesicular cristae [150]. These mice show growth defects, problems in limb coordination, and difficulties in gaining posture after being placed on their back and mostly move by writhing using abdomen as support. Also, depletion of Mfn1, Mfn2, Opa1 or Mfn1 & Mfn2 together in MEFs clearly showed loss of mitochondrial nucleoids when compared to control MEFs [150] which could have accounted for loss of membrane potential, decreased growth rates, and reduced mitochondrial oxygen consumption [151]. In accordance with the deletion of Mfns in cerebellum, where swollen mitochondria with sparse cristae were observed, loss of Mfn1 & 2 in skeletal muscle led to both interfibrillar and subsarcolemmal fragmented mitochondria containing swollen and sparse cristae [152]. Skeletal muscle specific KOs of Mfns had growth defects when compared to control littermates and displayed reduced levels of non-fasting and fasting blood glucose levels when compared to controls pointing to metabolic aberrations when the ultrastructural integrity of mitochondrial cristae is compromised. As a result, respiratory function was defective indicated by skeletal muscle COX/SDH staining revealing a low COX activity and an increased SDH activity due to mtDNA defects. Intriguingly, these mice also show a drastic decrease of mtDNA copies per nuclear genome, strongly indicating that the integrity of cristae and mtDNA are interlinked. Mfn2 DKO in the adult hearts results in progressive dilated cardiomyopathy [153]. Here, the cardiomyocytes displayed mitochondrial fragmentation and abnormal cristae morphology coupled with respiratory deficiency highlighting the intricate relationship between mitochondrial dynamics, cristae and respiratory efficiency. More recently, mitochondrial fusion was shown to be necessary for maintaining mtDNA replication and distribution [154]. Mfn2 DKO in the heart resulted in higher heart to body weight ratio, indicating hypertrophy and abnormal mitochondria with disrupted ultrastructure in the form of irregularly arranged swollen cristae. MtDNA content was reduced in Mfn DKO in the heart but there were no adverse effects on the number of breakpoints in mtDNA or other signs of impaired mtDNA integrity. In addition, the frequency of mtDNA mutations was not altered. STED super-resolution microscopy of mt-nucleoids revealed that deletion of Mfns and Opa1 led to mt-nucleoid clustering without affecting the size of mt-nucleoids [154]. Loss of fusion further resulted in a reduction of the steady-state levels of mtSSB, whereas Polγ levels and TWINKLE levels were increased indicating alteration of major proteins possibly compromising formation of the mtDNA replisome. This could explain decreased mtDNA copy numbers upon loss of mitochondrial fusion. Despite this, no impairment in mtDNA transcription was observed when mitochondrial fusion was impaired compared to controls based on in organello transcription experiments. Overall, loss of mitochondrial fusion due to loss of OM fusion leads to aberrations in mtDNA replication accompanied by defects in mitochondrial cristae organization [154].
How does a loss of mitochondrial fission lead to abnormalities in mt-nucleoid organization and to problems in cristae biogenesis? DRP1 plays an important role in the fission of the OM as discussed before. The physiological role of DRP1 is evident as mice deleted for Drp1 exhibit embryonic lethality by midgestation where KO embryos were smaller than controls [155,156]. Analysis of mitochondrial ultrastructure by EM mostly showed no change in the internal organization of the interconnected large mitochondrial network [156]. Concurrently, no changes were observed in mtDNA levels of Drp1 KO MEFS [155]. In contrast, another study reported loss of mtDNA, reduced membrane potential and cellular ATP levels in cells deleted for DRP1 [157]. Drp1 was further shown to be required for the maintenance of PCs, and mice deficient in Drp1 exhibited reduced latency when a rotarod test was used to test motor coordination ability [158]. Here, loss of Drp1 in postmitotic neurons led to formation of swollen mitochondria, respiratory impairment and enhanced ubiquitination of mitochondria colocalizing with LC3 punctae, suggesting induction of mitophagy in PCs. Isolated cerebellar neurons deficient for Drp1 in culture faithfully mimicked mitochondrial swelling and cell death. In addition, N-acetylcysteine (NAC) treatment obliterated the mitochondrial swelling phenotype of Drp1 KO PCs in culture indicating that oxidative damage promotes formation of swollen mitochondria. Thus, it could be that oxidative damage disrupts mtDNA and associated proteins forming nucleoids which may result in swelling of mitochondria and alteration of cristae morphology. In fact, depletion of Drp1 in MEFs led to formation of huge nucleoids in mitochondria [159]. In this study, it was proposed that clustering of nucleoids led to reorganization of cristae into densely packed units called mito-bulbs. Interestingly, in 96% of the cases mt-nucleoids were located next to Drp1 and Mff spots, players in IM fission. Mt-nucleoid clustering was apparently upstream of mito-bulb formation because inhibition of mtDNA replication by 2’-3’-dideoxycytidine (ddC) treatment, leading to loss of nucleoids, followed by depletion of Drp1 did not lead to formation of mito-bulbs. In addition, nucleoid clustering paired with mito-bulb formation led to a delay of cytochrome c release and apoptosis. Overall, it appears that in this context mito-bulb formation is a consequence of formation of enlarged mt-nucleoids. Death of heart and skeletal muscle specific KOs of Drp1 mice occurred within 11 days of birth, which was ascribed to dilated heart exhibiting hypertrophy and aberrant parameters deciphered by echocardiography. Drp1 KO heart exhibited enlarged mitochondria with respiratory deficiency, nucleoid clustering and densely packed cristae compared to controls [160]. Hence, loss of mitochondrial fission apparatus leads to aberrant mt-nucleoids and associated disruption of cristae in heart. Mice deficient in Mff, a protein required for mitochondrial fission, in the whole body, die around 13 weeks of age with symptoms of dilated cardiomyopathy associated with heart failure. There is a decrease of mtDNA molecules with age coinciding with abnormal internal structure of mitochondria containing vacuoles. Additionally, there is reduced mitochondrial oxygen consumption, increased LC3 and p62 punctae co-localizing with ubiquitinated mitochondria suggesting increased mitophagy. Knockout of Mfn1 in combination with Mff led to complete rescue of lifespan by rescuing the cardiac defects together with rescuing the mitochondrial oxygen consumption and autophagy. Intriguingly, the mtDNA levels were increased in DKO of Mff and Mfn1 reiterating that balanced mitochondrial dynamics maintain mtDNA at steady state [161]. This is reminiscent of data from baker’s yeast showing that strains lacking Dnm1 and Fzo1 (or Dnm1 and Mgm1) are rescued from mtDNA loss [162,163,164] strongly indicating that lack of mitochondrial fusion only led to loss of mtDNA when mitochondrial fission is functional. Loss of both processes, however, will prevent content mixing which was shown to ensure complementation of mitochondria containing different mutant mtDNA molecules. This was exemplified earlier as polykaryons formed upon PEG fusion of wild type (WT) cells and cells devoid of mtDNA (rho0 cells) resulted in the distribution of mitochondrial nucleoids from WT mitochondria to rho0 mitochondria [84]. Moreover, mtDNA synthesis was shown to couple with mitochondrial fission at the ER–mitochondria contact sites [165]. Overall, several studies support the view that impairment of mitochondrial dynamics can result in mitochondrial dysfunction often linked to loss of mtDNA, accumulation of enlarged/altered mt-nucleoids and altered cristae morphology. To decipher these interdependencies in more detail, we will now focus on the impact of cristae biogenesis on mtDNA integrity and vice versa.
4. Interplay between Cristae Biogenesis and mtDNA
We have introduced the roles of three factors on cristae biogenesis, namely, OPA1, the MICOS complex and the F1FO ATP synthase. How do these factors influence mtDNA organization? Mutations in OPA1 cause Autosomal Dominant Optic Atrophy (ADOA) characterized by optic nerve atrophy leading to progressive loss of vision [9,11]. Patients suffering from autosomal dominant optic atrophy were found to contain reduced mtDNA content [166]. Hela cells downregulated for OPA1 displayed swollen cristae and lower membrane potential than control cells [17]. Opa1+/− mice showed a delayed phenotype of abnormal cardiac function resulting in decreased cardiac output coinciding with the onset of blindness. Strikingly, the Opa1 mutant hearts also displayed loss of cristae coupled with impaired respiratory function of ETC complexes I, II and IV and reduction in mtDNA copy number [167]. Similarly, another study used a mouse model containing a splice site mutation leading to 27 amino acid residues’ deletion in the GTPase domain [168]. This led to 50% production of full length OPA1 protein where heterozygous mutant mice developed an age dependent loss of retinal ganglion cells (RGCs) and axons of the optic nerve displayed disorganized cristae in Opa1 mutant mice. Consistent with this, another study showed that heterozygous OPA1+/− mice showed an increase in the number of mitochondria harboring swollen cristae in the optic nerve and the hippocampus but did not show any change in the number of CJs [64]. A muscle specific deletion of Opa1 led to death of all Opa1 knockouts by postnatal day 9 (P9) corroborating its importance in early development of the mice [169]. Therefore, a tamoxifen-induced deletion of Opa1 was performed at 5 months of age, which resulted in muscle atrophy, weakness, kyphosis and hair greying. Mitochondria of Opa1 KO mice were smaller compared to controls and had dilated cristae. Since mtDNA depletion was not observed, the authors proposed loss of mitochondrial fusion upon Opa1 deletion leads to irregularities in cristae shape and mitochondrial supercomplex assembly [169]. This was in line with a study reporting that mtDNA depletion results from chronic fusion inhibition [170]. In the latter study, acute Opa1 ablation was performed using Opa1 floxed MAFs transduced with Cre. During acute ablation of Opa1 cristae shape and respiratory chain complexes were already impaired, while the mtDNA levels remained normal suggesting that cristae shape defects are not necessarily linked to mtDNA loss. Depletion of OPA1 and DRP1 simultaneously in the skeletal muscle of mice did not rescue accumulation of dysfunctional mitochondria where one-third of the mitochondria possessed either abnormal size or disrupted cristae structures supporting the view that a balance of mitochondrial fission and fusion is very important for regulation of mitochondrial size and cristae structure. Muscle weakness and atrophy were not rescued but a reduction of oxidative stress, denervation and FGF21 induction, which contributes to muscle atrophy, was observed [171].
Loss of MIC60 leads to loss of CJs, physically separating the IBM from the CM, and the appearance of concentric stacks of CM [29,172]. Depletion of MIC60 in both yeast and mammalian cells led to formation of enlarged nucleoids [173,174] demonstrating a rather direct link of altered cristae morphogenesis on mtDNA organization. Depletion of MIC60 in mammalian cells led to formation of enlarged mitochondria where mitochondrial fusion and fission events were reduced [174]. This was coupled to the formation of enlarged nucleoids and a drastic decrease of transcription of mt-encoded genes. This led to the hypothesis that CJs could be responsible for the distribution of nucleoids within the mitochondrial matrix in manner that is partially dependent on DRP1 [174]. In line with the antagonistic roles of Mic60/Fcj1 and dimer-specific subunits e and g of the F1FO ATP synthase [29] it was shown that in yeast enlarged mt-nucleoid size was prevented when both, Mic60/Fcj1 and subunits e (or g) were deleted simultaneously [173]. Depletion of MIC10 did not induce formation of enlarged nucleoids while deletion of MIC19 resulted in enlarged nucleoids. Also, deletion of Mic60/Fcj1 or subunits e (or g) alone was reported to cause a partial loss of mtDNA [29,175]. In addition, absence of mitochondrial fission by Dnm1 deletion led to the formation of larger nucleoids in DKO strain of Mic60 and Dnm1 compared to Mic60 KO showing that Dnm1 participated to distribute nucleoids [173].
Diabetic cardiomyopathy is a condition where patients suffering from type I diabetes are prone to increased risk of heart failure. Mic60 is significantly reduced in the interfibrillar mitochondria (IFM) of mice hearts which were insulted with streptozotocin to induce diabetes [176]. Accordingly, a Mic60 knockin mice model was used to check whether cardiomyopathy induced by diabetes could be rescued in this genetic background [177]. Cardiac contractile function was reduced in diabetic mice which was rescued in diabetic mice overexpressing Mic60 pointing out a beneficial role of high amounts of Mic60 under diabetic conditions. Interestingly, the cristae morphology altered in the IFM under diabetic conditions was rescued in Mic60 knockin mice. This further points to an important role of cristae remodelling in the development and progression of diabetic cardiomyopathy [177]. The role of mtDNA mutations in cardiac pathologies is well documented [178]. Patients harboring pathological mtDNA mutations displayed swollen mitochondria, and mitochondria with concentric cristae once again corroborating the interplay between mtDNA and mitochondrial cristae in the context of cardiac pathology [179]. Taken together, remodelling of cristae appears to have an immediate impact on mt-nucleoid structure and inheritance. Further analysis of the effect of cristae shaping proteins on the organization of mt-nucleoids is needed. A summary of deletion of factors playing a role in mitochondrial dynamics (and cristae organization) leading to aberrations in nucleoids/mtDNA content and cristae organization is provided in Table 1.

Table 1.
Summary of phenotypes linked to deletion of factors involved in mitochondrial dynamics or cristae biogenesis.
5. Conclusions
We have tried to summarize a rather complex and interdependent interplay between mitochondrial dynamics, cristae biogenesis and mtDNA maintenance and integrity (Figure 1). It is a challenging and open question whether abnormalities in mitochondrial dynamics and cristae structure give rise to aberrancies in mtDNA and nucleoids or vice-versa. Based on the data available, the answer to this puzzle is still not clear—also, each case might have a different mode of action and it is possible that these processes are interdependent and cannot be separated that easily (as shown in Figure 1). It is not always that a reduction in mtDNA copy number, variations in nucleoid size and associated clustering are associated with disorganized cristae or the other way round. Also, alteration of mitochondrial dynamics has distinct effects on cristae and mtDNA integrity. At least we can say that loss of mitochondrial dynamics frequently leads to aberrant cristae formation and that aberrations in cristae morphology frequently result in loss of mtDNA. It is obvious that these processes are interlinked at multiple levels. Future studies will have to dissect the molecular hierarchy and interdependency of these processes. This will have important implications for understanding human diseases linked to various forms of mitochondrial dysfunction.
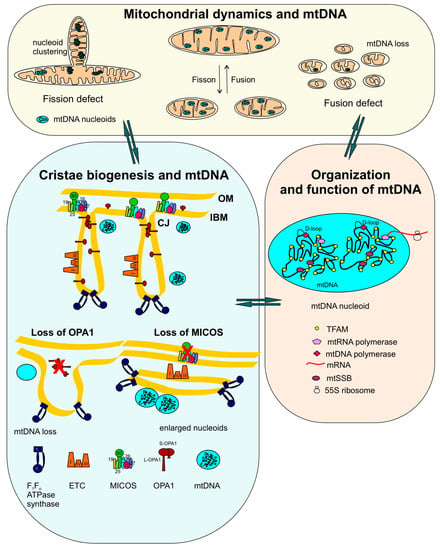
Figure 1.
Functional interplay of mitochondrial dynamics, cristae biogenesis and mtDNA integrity: The scheme describes that the balance between mitochondrial dynamics, cristae biogenesis and mtDNA structure helps to manage integrity and function of mitochondrial DNA. Mitochondrial dynamics comprising fission and fusion cycles helps in redistribution and complementation of mtDNA within mitochondria. Lack of fusion causes subsequent loss of mtDNA from the fragmented mitochondria, whereas accumulation of aggregated nucleoids is observed in elongated mitochondria generated during a fission defect. Relationship between mitochondrial dynamics and cristae biogenesis and mtDNA organization is evident by several examples where the loss of fusion or fission concurrently shows defective cristae and mtDNA loss. Altered morphology of mitochondria is observed with mtDNA aberrations during cristae biogenesis defects. The internal mitochondrial structure is modulated by the following key players regulating cristae shape: OPA1 oligomers, MICOS, and F1Fo ATPase synthase. Chronic loss of OPA1 is accompanied by cristae defects as well as mtDNA loss. Depletion of MIC60 results in loss of CJs together with accumulation of enlarged nucleoids. The function and organization of mtDNA nucleoids is schematically depicted. On average, 1.4 mtDNA molecules are present per nucleoid which also contains associated proteins required for compaction, replication and transcription. ETC, electron transport chain (complex II is not shown).
Funding
This research was funded by Deutsche Forschungsgemeinschaft (DFG), grant RE 1575/2-1 (to ASR) and by the Research Committee of the Medical faculty of Heinrich Heine University, Düsseldorf, FoKo-37/2015 (to AKK).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Perkins, G.; Renken, C.; Martone, M.E.; Young, S.J.; Ellisman, M.; Frey, T. Electron tomography of neuronal mitochondria: Three-dimensional structure and organization of cristae and membrane contacts. J. Struct. Biol. 1997, 119, 260–272. [Google Scholar] [CrossRef] [PubMed]
- Perkins, G.A.; Song, J.Y.; Tarsa, L.; Deerinck, T.J.; Ellisman, M.H.; Frey, T.G. Electron tomography of mitochondria from brown adipocytes reveals crista junctions. J. Bioenerg. Biomembr. 1998, 30, 431–442. [Google Scholar] [CrossRef] [PubMed]
- Zick, M.; Rabl, R.; Reichert, A.S. Cristae formation-linking ultrastructure and function of mitochondria. Biochim. Biophys. Acta 2009, 1793, 5–19. [Google Scholar] [CrossRef] [PubMed]
- Vogel, F.; Börnhovd, C.; Neupert, W.; Reichert, A.S. Dynamic subcompartmentalization of the mitochondrial inner membrane. J. Cell Biol. 2006, 175, 237–247. [Google Scholar] [CrossRef] [PubMed]
- Mannella, C.A.; Marko, M.; Buttle, K. Reconsidering mitochondrial structure: New views of an old organelle. Trends Biochem. Sci. 1997, 22, 37–38. [Google Scholar] [CrossRef]
- Hackenbrock, C.R. Ultrastructural bases for metabolically linked mechanical activity in mitochondria. I. Reversible ultrastructural changes with change in metabolic steady state in isolated liver mitochondria. J. Cell Biol. 1966, 30, 269–297. [Google Scholar] [CrossRef] [PubMed]
- Hackenbrock, C.R. Chemical and physical fixation of isolated mitochondria in low-energy and high-energy states. Proc. Natl. Acad. Sci. USA 1968, 61, 598–605. [Google Scholar] [CrossRef] [PubMed]
- Scorrano, L.; Ashiya, M.; Buttle, K.; Weiler, S.; Oakes, S.A.; Mannella, C.A.; Korsmeyer, S.J. A distinct pathway remodels mitochondrial cristae and mobilizes cytochrome c during apoptosis. Dev. Cell 2002, 2, 55–67. [Google Scholar] [CrossRef]
- Alexander, C.; Votruba, M.; Pesch, U.E.; Thiselton, D.L.; Mayer, S.; Moore, A.; Rodriguez, M.; Kellner, U.; Leo-Kottler, B.; Auburger, G.; et al. OPA1, encoding a dynamin-related GTPase, is mutated in autosomal dominant optic atrophy linked to chromosome 3q28. Nat. Genet. 2000, 26, 211–215. [Google Scholar] [CrossRef] [PubMed]
- Carelli, V.; Schimpf, S.; Fuhrmann, N.; Valentino, M.L.; Zanna, C.; Iommarini, L.; Papke, M.; Schaich, S.; Tippmann, S.; Baumann, B.; et al. A clinically complex form of dominant optic atrophy (OPA8) maps on chromosome 16. Hum. Mol. Genet. 2011, 20, 1893–1905. [Google Scholar] [CrossRef]
- Delettre, C.; Lenaers, G.; Griffoin, J.M.; Gigarel, N.; Lorenzo, C.; Belenguer, P.; Pelloquin, L.; Grosgeorge, J.; Turc-Carel, C.; Perret, E.; et al. Nuclear gene OPA1, encoding a mitochondrial dynamin-related protein, is mutated in dominant optic atrophy. Nat. Genet. 2000, 26, 207–210. [Google Scholar] [CrossRef] [PubMed]
- Spiegel, R.; Saada, A.; Flannery, P.J.; Burte, F.; Soiferman, D.; Khayat, M.; Eisner, V.; Vladovski, E.; Taylor, R.W.; Bindoff, L.A.; et al. Fatal infantile mitochondrial encephalomyopathy, hypertrophic cardiomyopathy and optic atrophy associated with a homozygous OPA1 mutation. J. Med. Genet. 2016, 53, 127–131. [Google Scholar] [CrossRef] [PubMed]
- Anand, R.; Langer, T.; Baker, M.J. Proteolytic control of mitochondrial function and morphogenesis. Biochim. Biophys. Acta 2013, 1833, 195–204. [Google Scholar] [CrossRef] [PubMed]
- MacVicar, T.; Langer, T. OPA1 processing in cell death and disease-the long and short of it. J. Cell Sci. 2016, 129, 2297–2306. [Google Scholar] [CrossRef] [PubMed]
- Anand, R.; Wai, T.; Baker, M.J.; Kladt, N.; Schauss, A.C.; Rugarli, E.; Langer, T. The i-AAA protease YME1L and OMA1 cleave OPA1 to balance mitochondrial fusion and fission. J. Cell Biol. 2014, 204, 919–929. [Google Scholar] [CrossRef] [PubMed]
- Frezza, C.; Cipolat, S.; Martins de Brito, O.; Micaroni, M.; Beznoussenko, G.V.; Rudka, T.; Bartoli, D.; Polishuck, R.S.; Danial, N.N.; De Strooper, B.; et al. OPA1 controls apoptotic cristae remodeling independently from mitochondrial fusion. Cell 2006, 126, 177–189. [Google Scholar] [CrossRef] [PubMed]
- Olichon, A.; Baricault, L.; Gas, N.; Guillou, E.; Valette, A.; Belenguer, P.; Lenaers, G. Loss of OPA1 perturbates the mitochondrial inner membrane structure and integrity, leading to cytochrome c release and apoptosis. J. Biol. Chem. 2003, 278, 7743–7746. [Google Scholar] [CrossRef] [PubMed]
- Del Dotto, V.; Mishra, P.; Vidoni, S.; Fogazza, M.; Maresca, A.; Caporali, L.; McCaffery, J.M.; Cappelletti, M.; Baruffini, E.; Lenaers, G.; et al. OPA1 Isoforms in the Hierarchical Organization of Mitochondrial Functions. Cell Rep. 2017, 19, 2557–2571. [Google Scholar] [CrossRef]
- Lee, H.; Smith, S.B.; Yoon, Y. The short variant of the mitochondrial dynamin OPA1 maintains mitochondrial energetics and cristae structure. J. Biol. Chem. 2017, 292, 7115–7130. [Google Scholar] [CrossRef]
- Harner, M.E.; Unger, A.K.; Geerts, W.J.; Mari, M.; Izawa, T.; Stenger, M.; Geimer, S.; Reggiori, F.; Westermann, B.; Neupert, W. An evidence based hypothesis on the existence of two pathways of mitochondrial crista formation. eLife 2016, 5, e18853. [Google Scholar] [CrossRef]
- Brunner, S.; Everard-Gigot, V.; Stuart, R.A. Su e of the yeast F1Fo-ATP synthase forms homodimers. J. Biol. Chem. 2002, 277, 48484–48489. [Google Scholar] [CrossRef] [PubMed]
- Paumard, P.; Vaillier, J.; Coulary, B.; Schaeffer, J.; Soubannier, V.; Mueller, D.M.; Brethes, D.; di Rago, J.P.; Velours, J. The ATP synthase is involved in generating mitochondrial cristae morphology. Embo J. 2002, 21, 221–230. [Google Scholar] [CrossRef]
- Soubannier, V.; Vaillier, J.; Paumard, P.; Coulary, B.; Schaeffer, J.; Velours, J. In the absence of the first membrane-spanning segment of subunit 4(b), the yeast ATP synthase is functional but does not dimerize or oligomerize. J. Biol. Chem. 2002, 277, 10739–10745. [Google Scholar] [CrossRef] [PubMed]
- Blum, T.B.; Hahn, A.; Meier, T.; Davies, K.M.; Kuhlbrandt, W. Dimers of mitochondrial ATP synthase induce membrane curvature and self-assemble into rows. Proc. Natl. Acad. Sci. USA 2019, 116, 4250–4255. [Google Scholar] [CrossRef] [PubMed]
- Wittig, I.; Karas, M.; Schagger, H. High resolution clear native electrophoresis for in-gel functional assays and fluorescence studies of membrane protein complexes. Mol. Cell. Proteom. 2007, 6, 1215–1225. [Google Scholar] [CrossRef] [PubMed]
- Strauss, M.; Hofhaus, G.; Schroder, R.R.; Kuhlbrandt, W. Dimer ribbons of ATP synthase shape the inner mitochondrial membrane. Embo J. 2008, 27, 1154–1160. [Google Scholar] [CrossRef] [PubMed]
- Davies, K.M.; Strauss, M.; Daum, B.; Kief, J.H.; Osiewacz, H.D.; Rycovska, A.; Zickermann, V.; Kuhlbrandt, W. Macromolecular organization of ATP synthase and complex I in whole mitochondria. Proc. Natl. Acad. Sci. USA 2011, 108, 14121–14126. [Google Scholar] [CrossRef]
- Hahn, A.; Parey, K.; Bublitz, M.; Mills, D.J.; Zickermann, V.; Vonck, J.; Kuhlbrandt, W.; Meier, T. Structure of a Complete ATP Synthase Dimer Reveals the Molecular Basis of Inner Mitochondrial Membrane Morphology. Mol. Cell 2016, 63, 445–456. [Google Scholar] [CrossRef]
- Rabl, R.; Soubannier, V.; Scholz, R.; Vogel, F.; Mendl, N.; Vasiljev-Neumeyer, A.; Körner, C.; Jagasia, R.; Keil, T.; Baumeister, W.; et al. Formation of cristae and crista junctions in mitochondria depends on antagonism between Fcj1 and Su e/g. J. Cell Biol. 2009, 185, 1047–1063. [Google Scholar] [CrossRef]
- Von der Malsburg, K.; Muller, J.M.; Bohnert, M.; Oeljeklaus, S.; Kwiatkowska, P.; Becker, T.; Loniewska-Lwowska, A.; Wiese, S.; Rao, S.; Milenkovic, D.; et al. Dual role of mitofilin in mitochondrial membrane organization and protein biogenesis. Dev. Cell 2011, 21, 694–707. [Google Scholar] [CrossRef]
- Harner, M.; Korner, C.; Walther, D.; Mokranjac, D.; Kaesmacher, J.; Welsch, U.; Griffith, J.; Mann, M.; Reggiori, F.; Neupert, W. The mitochondrial contact site complex, a determinant of mitochondrial architecture. Embo J. 2011, 30, 4356–4370. [Google Scholar] [CrossRef] [PubMed]
- Hoppins, S.; Collins, S.R.; Cassidy-Stone, A.; Hummel, E.; Devay, R.M.; Lackner, L.L.; Westermann, B.; Schuldiner, M.; Weissman, J.S.; Nunnari, J. A mitochondrial-focused genetic interaction map reveals a scaffold-like complex required for inner membrane organization in mitochondria. J. Cell Biol. 2011, 195, 323–340. [Google Scholar] [CrossRef] [PubMed]
- Huynen, M.A.; Muhlmeister, M.; Gotthardt, K.; Guerrero-Castillo, S.; Brandt, U. Evolution and structural organization of the mitochondrial contact site (MICOS) complex and the mitochondrial intermembrane space bridging (MIB) complex. Biochim. Biophys. Acta 2016, 1863, 91–101. [Google Scholar] [CrossRef] [PubMed]
- Munoz-Gomez, S.A.; Slamovits, C.H.; Dacks, J.B.; Baier, K.A.; Spencer, K.D.; Wideman, J.G. Ancient homology of the mitochondrial contact site and cristae organizing system points to an endosymbiotic origin of mitochondrial cristae. Curr. Biol. 2015, 25, 1489–1495. [Google Scholar] [CrossRef] [PubMed]
- Pfanner, N.; van der Laan, M.; Amati, P.; Capaldi, R.A.; Caudy, A.A.; Chacinska, A.; Darshi, M.; Deckers, M.; Hoppins, S.; Icho, T.; et al. Uniform nomenclature for the mitochondrial contact site and cristae organizing system. J. Cell Biol. 2014, 204, 1083–1086. [Google Scholar] [CrossRef] [PubMed]
- Wollweber, F.; von der Malsburg, K.; van der Laan, M. Mitochondrial Contact Site and Cristae Organizing System: A Central Player in Membrane Shaping and Crosstalk. Biochim. Biophys. Acta 2017, 1864, 1481–1489. [Google Scholar] [CrossRef] [PubMed]
- Van der Laan, M.; Horvath, S.E.; Pfanner, N. Mitochondrial contact site and cristae organizing system. Curr. Opin. Cell Biol. 2016, 41, 33–42. [Google Scholar] [CrossRef]
- Xie, J.; Marusich, M.F.; Souda, P.; Whitelegge, J.; Capaldi, R.A. The mitochondrial inner membrane protein mitofilin exists as a complex with SAM50, metaxins 1 and 2, coiled-coil-helix coiled-coil-helix domain-containing protein 3 and 6 and DnaJC11. FEBS Lett. 2007, 581, 3545–3549. [Google Scholar] [CrossRef]
- Körner, C.; Barrera, M.; Dukanovic, J.; Eydt, K.; Harner, M.; Rabl, R.; Vogel, F.; Rapaport, D.; Neupert, W.; Reichert, A.S. The C-terminal domain of Fcj1 is required for formation of crista junctions and interacts with the TOB/SAM complex in mitochondria. Mol. Biol. Cell 2012, 23, 2143–2155. [Google Scholar] [CrossRef]
- Ott, C.; Ross, K.; Straub, S.; Thiede, B.; Gotz, M.; Goosmann, C.; Krischke, M.; Mueller, M.J.; Krohne, G.; Rudel, T.; et al. Sam50 functions in mitochondrial intermembrane space bridging and biogenesis of respiratory complexes. Mol. Cell. Biol. 2012, 32, 1173–1188. [Google Scholar] [CrossRef]
- Abrams, A.J.; Hufnagel, R.B.; Rebelo, A.; Zanna, C.; Patel, N.; Gonzalez, M.A.; Campeanu, I.J.; Griffin, L.B.; Groenewald, S.; Strickland, A.V.; et al. Mutations in SLC25A46, encoding a UGO1-like protein, cause an optic atrophy spectrum disorder. Nat. Genet. 2015, 47, 926–932. [Google Scholar] [CrossRef] [PubMed]
- Glytsou, C.; Calvo, E.; Cogliati, S.; Mehrotra, A.; Anastasia, I.; Rigoni, G.; Raimondi, A.; Shintani, N.; Loureiro, M.; Vazquez, J.; et al. Optic Atrophy 1 Is Epistatic to the Core MICOS Component MIC60 in Mitochondrial Cristae Shape Control. Cell Rep. 2016, 17, 3024–3034. [Google Scholar] [CrossRef] [PubMed]
- Pinero-Martos, E.; Ortega-Vila, B.; Pol-Fuster, J.; Cisneros-Barroso, E.; Ruiz-Guerra, L.; Medina-Dols, A.; Heine-Suner, D.; Llado, J.; Olmos, G.; Vives-Bauza, C. Disrupted in schizophrenia 1 (DISC1) is a constituent of the mammalian mitochondrial contact site and cristae organizing system (MICOS) complex, and is essential for oxidative phosphorylation. Hum. Mol. Genet. 2016, 25, 4157–4169. [Google Scholar] [CrossRef] [PubMed]
- Genin, E.C.; Plutino, M.; Bannwarth, S.; Villa, E.; Cisneros-Barroso, E.; Roy, M.; Ortega-Vila, B.; Fragaki, K.; Lespinasse, F.; Pinero-Martos, E.; et al. CHCHD10 mutations promote loss of mitochondrial cristae junctions with impaired mitochondrial genome maintenance and inhibition of apoptosis. Embo Mol. Med. 2015, 8, 58–72. [Google Scholar] [CrossRef] [PubMed]
- Barbot, M.; Jans, D.C.; Schulz, C.; Denkert, N.; Kroppen, B.; Hoppert, M.; Jakobs, S.; Meinecke, M. Mic10 oligomerizes to bend mitochondrial inner membranes at cristae junctions. Cell Metab. 2015, 21, 756–763. [Google Scholar] [CrossRef] [PubMed]
- Bohnert, M.; Zerbes, R.M.; Davies, K.M.; Muhleip, A.W.; Rampelt, H.; Horvath, S.E.; Boenke, T.; Kram, A.; Perschil, I.; Veenhuis, M.; et al. Central role of Mic10 in the mitochondrial contact site and cristae organizing system. Cell Metab. 2015, 21, 747–755. [Google Scholar] [CrossRef]
- Milenkovic, D.; Larsson, N.G. Mic10 Oligomerization Pinches off Mitochondrial Cristae. Cell Metab. 2015, 21, 660–661. [Google Scholar] [CrossRef][Green Version]
- Hessenberger, M.; Zerbes, R.M.; Rampelt, H.; Kunz, S.; Xavier, A.H.; Purfurst, B.; Lilie, H.; Pfanner, N.; van der Laan, M.; Daumke, O. Regulated membrane remodeling by Mic60 controls formation of mitochondrial crista junctions. Nat. Commun. 2017, 8, 15258. [Google Scholar] [CrossRef]
- Tarasenko, D.; Barbot, M.; Jans, D.C.; Kroppen, B.; Sadowski, B.; Heim, G.; Mobius, W.; Jakobs, S.; Meinecke, M. The MICOS component Mic60 displays a conserved membrane-bending activity that is necessary for normal cristae morphology. J. Cell Biol. 2017, 216, 889–899. [Google Scholar] [CrossRef]
- Anand, R.; Strecker, V.; Urbach, J.; Wittig, I.; Reichert, A.S. Mic13 Is Essential for Formation of Crista Junctions in Mammalian Cells. PLoS ONE 2016, 11, e0160258. [Google Scholar] [CrossRef]
- Guarani, V.; McNeill, E.M.; Paulo, J.A.; Huttlin, E.L.; Frohlich, F.; Gygi, S.P.; Van Vactor, D.; Harper, J.W. QIL1 is a novel mitochondrial protein required for MICOS complex stability and cristae morphology. eLife 2015, 4, e06265. [Google Scholar] [CrossRef] [PubMed]
- Koob, S.; Barrera, M.; Anand, R.; Reichert, A.S. The non-glycosylated isoform of MIC26 is a constituent of the mammalian MICOS complex and promotes formation of crista junctions. Biochim. Biophys. Acta 2015, 1853, 1551–1563. [Google Scholar] [CrossRef] [PubMed]
- Weber, T.A.; Koob, S.; Heide, H.; Wittig, I.; Head, B.; van der Bliek, A.; Brandt, U.; Mittelbronn, M.; Reichert, A.S. APOOL is a cardiolipin-binding constituent of the Mitofilin/MINOS protein complex determining cristae morphology in mammalian mitochondria. PLoS ONE 2013, 8, e63683. [Google Scholar] [CrossRef] [PubMed]
- Koob, S.; Reichert, A.S. Novel intracellular functions of apolipoproteins: The ApoO protein family as constituents of the Mitofilin/MINOS complex determines cristae morphology in mitochondria. Biol. Chem. 2014, 395, 285–296. [Google Scholar] [CrossRef]
- Zerbes, R.M.; van der Klei, I.J.; Veenhuis, M.; Pfanner, N.; van der Laan, M.; Bohnert, M. Mitofilin complexes: Conserved organizers of mitochondrial membrane architecture. Biol. Chem. 2012, 393, 1247–1261. [Google Scholar] [CrossRef] [PubMed]
- Russell, B.E.; Whaley, K.G.; Bove, K.E.; Labilloy, A.; Lombardo, R.C.; Hopkin, R.J.; Leslie, N.D.; Prada, C.; Assouline, Z.; Barcia, G.; et al. Expanding and Underscoring the Hepato-Encephalopathic Phenotype of QIL1/MIC13. Hepatol. (Baltim. Md.) 2019, 70, 1066–1070. [Google Scholar] [CrossRef] [PubMed]
- Godiker, J.; Gruneberg, M.; DuChesne, I.; Reunert, J.; Rust, S.; Westermann, C.; Wada, Y.; Classen, G.; Langhans, C.D.; Schlingmann, K.P.; et al. QIL1-dependent assembly of MICOS complex-lethal mutation in C19ORF70 resulting in liver disease and severe neurological retardation. J. Hum. Genet. 2018, 63, 707–716. [Google Scholar] [CrossRef] [PubMed]
- Zeharia, A.; Friedman, J.R.; Tobar, A.; Saada, A.; Konen, O.; Fellig, Y.; Shaag, A.; Nunnari, J.; Elpeleg, O. Mitochondrial hepato-encephalopathy due to deficiency of QIL1/MIC13 (C19orf70), a MICOS complex subunit. Eur. J. Hum. Genet. 2016, 24, 1778–1782. [Google Scholar] [CrossRef]
- Guarani, V.; Jardel, C.; Chretien, D.; Lombes, A.; Benit, P.; Labasse, C.; Lacene, E.; Bourillon, A.; Imbard, A.; Benoist, J.F.; et al. QIL1 mutation causes MICOS disassembly and early onset fatal mitochondrial encephalopathy with liver disease. eLife 2016, 5, e17163. [Google Scholar] [CrossRef]
- Tsai, P.I.; Lin, C.H.; Hsieh, C.H.; Papakyrikos, A.M.; Kim, M.J.; Napolioni, V.; Schoor, C.; Couthouis, J.; Wu, R.M.; Wszolek, Z.K.; et al. PINK1 Phosphorylates MIC60/Mitofilin to Control Structural Plasticity of Mitochondrial Crista Junctions. Mol. Cell 2018, 69, 744–756. [Google Scholar] [CrossRef]
- Eydt, K.; Davies, K.M.; Behrendt, C.; Wittig, I.; Reichert, A.S. Cristae architecture is determined by an interplay of the MICOS complex and the F1FO ATP synthase via Mic27 and Mic10. Microb. Cell (GrazAustria) 2017, 4, 259–272. [Google Scholar] [CrossRef] [PubMed]
- Rampelt, H.; Bohnert, M.; Zerbes, R.M.; Horvath, S.E.; Warscheid, B.; Pfanner, N.; van der Laan, M. Mic10, a Core Subunit of the Mitochondrial Contact Site and Cristae Organizing System, Interacts with the Dimeric F1Fo-ATP Synthase. J. Mol. Biol. 2017, 429, 1162–1170. [Google Scholar] [CrossRef] [PubMed]
- Quintana-Cabrera, R.; Quirin, C.; Glytsou, C.; Corrado, M.; Urbani, A.; Pellattiero, A.; Calvo, E.; Vazquez, J.; Enriquez, J.A.; Gerle, C.; et al. The cristae modulator Optic atrophy 1 requires mitochondrial ATP synthase oligomers to safeguard mitochondrial function. Nat. Commun. 2018, 9, 3399. [Google Scholar] [CrossRef] [PubMed]
- Barrera, M.; Koob, S.; Dikov, D.; Vogel, F.; Reichert, A.S. OPA1 functionally interacts with MIC60 but is dispensable for crista junction formation. FEBS Lett. 2016, 590, 3309–3322. [Google Scholar] [CrossRef] [PubMed]
- Nass, M.M.; Nass, S. Fibrous Structures within the Matrix of Developing Chick Embryo Mitochondria. Exp. Cell Res. 1962, 26, 424–427. [Google Scholar] [CrossRef]
- Nass, M.M.; Nass, S. Intramitochondrial Fibers with DNA Characteristics: I. Fixation and Electron Staining Reactions. J. Cell Biol. 1963, 19, 593–611. [Google Scholar] [CrossRef] [PubMed]
- Nass, S.; Nass, M.M. Intramitochondrial Fibers with DNA Characteristics: II. Enzymatic and Other Hydrolytic Treatments. J. Cell Biol. 1963, 19, 613–629. [Google Scholar] [CrossRef] [PubMed]
- Nass, M.M.; Nass, S.; Afzelius, B.A. The general occurrence of mitochondrial DNA. Exp. Cell Res. 1965, 37, 516–539. [Google Scholar] [CrossRef]
- Nass, M.M. The circularity of mitochondrial DNA. Proc. Natl. Acad. Sci. USA 1966, 56, 1215–1222. [Google Scholar] [CrossRef]
- Nass, M.M. Mitochondrial DNA. I. Intramitochondrial distribution and structural relations of single- and double-length circular DNA. J. Mol. Biol. 1969, 42, 521–528. [Google Scholar] [CrossRef]
- Albring, M.; Griffith, J.; Attardi, G. Association of a protein structure of probable membrane derivation with HeLa cell mitochondrial DNA near its origin of replication. Proc. Natl. Acad. Sci. USA 1977, 74, 1348–1352. [Google Scholar] [CrossRef]
- Iborra, F.J.; Kimura, H.; Cook, P.R. The functional organization of mitochondrial genomes in human cells. BMC Biol. 2004, 2, 9. [Google Scholar] [CrossRef] [PubMed]
- Kopek, B.G.; Shtengel, G.; Xu, C.S.; Clayton, D.A.; Hess, H.F. Correlative 3D superresolution fluorescence and electron microscopy reveal the relationship of mitochondrial nucleoids to membranes. Proc. Natl. Acad. Sci. USA 2012, 109, 6136–6141. [Google Scholar] [CrossRef]
- Anderson, S.; Bankier, A.T.; Barrell, B.G.; de Bruijn, M.H.; Coulson, A.R.; Drouin, J.; Eperon, I.C.; Nierlich, D.P.; Roe, B.A.; Sanger, F.; et al. Sequence and organization of the human mitochondrial genome. Nature 1981, 290, 457–465. [Google Scholar] [CrossRef] [PubMed]
- Ojala, D.; Merkel, C.; Gelfand, R.; Attardi, G. The tRNA genes punctuate the reading of genetic information in human mitochondrial DNA. Cell 1980, 22, 393–403. [Google Scholar] [CrossRef]
- Ojala, D.; Montoya, J.; Attardi, G. tRNA punctuation model of RNA processing in human mitochondria. Nature 1981, 290, 470–474. [Google Scholar] [CrossRef]
- Bruni, F.; Lightowlers, R.N.; Chrzanowska-Lightowlers, Z.M. Human mitochondrial nucleases. FEBS J. 2017, 284, 1767–1777. [Google Scholar] [CrossRef]
- Sanchez, M.I.; Mercer, T.R.; Davies, S.M.; Shearwood, A.M.; Nygard, K.K.; Richman, T.R.; Mattick, J.S.; Rackham, O.; Filipovska, A. RNA processing in human mitochondria. Cell Cycle 2011, 10, 2904–2916. [Google Scholar] [CrossRef]
- Falkenberg, M.; Larsson, N.G.; Gustafsson, C.M. DNA replication and transcription in mammalian mitochondria. Annu. Rev. Biochem. 2007, 76, 679–699. [Google Scholar] [CrossRef]
- Gustafsson, C.M.; Falkenberg, M.; Larsson, N.G. Maintenance and Expression of Mammalian Mitochondrial DNA. Annu. Rev. Biochem. 2016, 85, 133–160. [Google Scholar] [CrossRef] [PubMed]
- Bonekamp, N.A.; Larsson, N.G. SnapShot: Mitochondrial Nucleoid. Cell 2018, 172, 388-388.e1. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Dierckx, A.; Wanrooij, P.H.; Wanrooij, S.; Larsson, N.G.; Wilhelmsson, L.M.; Falkenberg, M.; Gustafsson, C.M. Mammalian transcription factor A is a core component of the mitochondrial transcription machinery. Proc. Natl. Acad. Sci. USA 2012, 109, 16510–16515. [Google Scholar] [CrossRef] [PubMed]
- Ngo, H.B.; Kaiser, J.T.; Chan, D.C. The mitochondrial transcription and packaging factor Tfam imposes a U-turn on mitochondrial DNA. Nat. Struct. Mol. Biol. 2011, 18, 1290–1296. [Google Scholar] [CrossRef] [PubMed]
- Legros, F.; Malka, F.; Frachon, P.; Lombes, A.; Rojo, M. Organization and dynamics of human mitochondrial DNA. J. Cell Sci. 2004, 117, 2653–2662. [Google Scholar] [CrossRef] [PubMed]
- Gilkerson, R.W.; Schon, E.A.; Hernandez, E.; Davidson, M.M. Mitochondrial nucleoids maintain genetic autonomy but allow for functional complementation. J. Cell Biol. 2008, 181, 1117–1128. [Google Scholar] [CrossRef] [PubMed]
- Kukat, C.; Wurm, C.A.; Spahr, H.; Falkenberg, M.; Larsson, N.G.; Jakobs, S. Super-resolution microscopy reveals that mammalian mitochondrial nucleoids have a uniform size and frequently contain a single copy of mtDNA. Proc. Natl. Acad. Sci. USA 2011, 108, 13534–13539. [Google Scholar] [CrossRef] [PubMed]
- Kukat, C.; Davies, K.M.; Wurm, C.A.; Spahr, H.; Bonekamp, N.A.; Kuhl, I.; Joos, F.; Polosa, P.L.; Park, C.B.; Posse, V.; et al. Cross-strand binding of TFAM to a single mtDNA molecule forms the mitochondrial nucleoid. Proc. Natl. Acad. Sci. USA 2015, 112, 11288–11293. [Google Scholar] [CrossRef]
- Kucej, M.; Kucejova, B.; Subramanian, R.; Chen, X.J.; Butow, R.A. Mitochondrial nucleoids undergo remodeling in response to metabolic cues. J. Cell Sci. 2008, 121, 1861–1868. [Google Scholar] [CrossRef]
- Larsson, N.G.; Wang, J.; Wilhelmsson, H.; Oldfors, A.; Rustin, P.; Lewandoski, M.; Barsh, G.S.; Clayton, D.A. Mitochondrial transcription factor A is necessary for mtDNA maintenance and embryogenesis in mice. Nat. Genet. 1998, 18, 231–236. [Google Scholar] [CrossRef]
- Wang, J.; Wilhelmsson, H.; Graff, C.; Li, H.; Oldfors, A.; Rustin, P.; Bruning, J.C.; Kahn, C.R.; Clayton, D.A.; Barsh, G.S.; et al. Dilated cardiomyopathy and atrioventricular conduction blocks induced by heart-specific inactivation of mitochondrial DNA gene expression. Nat. Genet. 1999, 21, 133–137. [Google Scholar] [CrossRef]
- Wredenberg, A.; Wibom, R.; Wilhelmsson, H.; Graff, C.; Wiener, H.H.; Burden, S.J.; Oldfors, A.; Westerblad, H.; Larsson, N.G. Increased mitochondrial mass in mitochondrial myopathy mice. Proc. Natl. Acad. Sci. USA 2002, 99, 15066–15071. [Google Scholar] [CrossRef] [PubMed]
- Ekstrand, M.I.; Terzioglu, M.; Galter, D.; Zhu, S.; Hofstetter, C.; Lindqvist, E.; Thams, S.; Bergstrand, A.; Hansson, F.S.; Trifunovic, A.; et al. Progressive parkinsonism in mice with respiratory-chain-deficient dopamine neurons. Proc. Natl. Acad. Sci. USA 2007, 104, 1325–1330. [Google Scholar] [CrossRef]
- Bender, A.; Krishnan, K.J.; Morris, C.M.; Taylor, G.A.; Reeve, A.K.; Perry, R.H.; Jaros, E.; Hersheson, J.S.; Betts, J.; Klopstock, T.; et al. High levels of mitochondrial DNA deletions in substantia nigra neurons in aging and Parkinson disease. Nat. Genet. 2006, 38, 515–517. [Google Scholar] [CrossRef] [PubMed]
- Zilocchi, M.; Finzi, G.; Lualdi, M.; Sessa, F.; Fasano, M.; Alberio, T. Mitochondrial alterations in Parkinson’s disease human samples and cellular models. Neurochem. Int. 2018, 118, 61–72. [Google Scholar] [CrossRef] [PubMed]
- Pickrell, A.M.; Youle, R.J. The roles of PINK1, parkin, and mitochondrial fidelity in Parkinson’s disease. Neuron 2015, 85, 257–273. [Google Scholar] [CrossRef]
- Milenkovic, D.; Matic, S.; Kuhl, I.; Ruzzenente, B.; Freyer, C.; Jemt, E.; Park, C.B.; Falkenberg, M.; Larsson, N.G. TWINKLE is an essential mitochondrial helicase required for synthesis of nascent D-loop strands and complete mtDNA replication. Hum. Mol. Genet. 2013, 22, 1983–1993. [Google Scholar] [CrossRef] [PubMed]
- Trifunovic, A.; Wredenberg, A.; Falkenberg, M.; Spelbrink, J.N.; Rovio, A.T.; Bruder, C.E.; Bohlooly, Y.M.; Gidlof, S.; Oldfors, A.; Wibom, R.; et al. Premature ageing in mice expressing defective mitochondrial DNA polymerase. Nature 2004, 429, 417–423. [Google Scholar] [CrossRef]
- Wang, Y.; Bogenhagen, D.F. Human mitochondrial DNA nucleoids are linked to protein folding machinery and metabolic enzymes at the mitochondrial inner membrane. J. Biol. Chem. 2006, 281, 25791–25802. [Google Scholar] [CrossRef]
- Bogenhagen, D.F.; Rousseau, D.; Burke, S. The layered structure of human mitochondrial DNA nucleoids. J. Biol. Chem. 2008, 283, 3665–3675. [Google Scholar] [CrossRef]
- Kasashima, K.; Sumitani, M.; Satoh, M.; Endo, H. Human prohibitin 1 maintains the organization and stability of the mitochondrial nucleoids. Exp. Cell Res. 2008, 314, 988–996. [Google Scholar] [CrossRef]
- He, J.; Mao, C.C.; Reyes, A.; Sembongi, H.; Di Re, M.; Granycome, C.; Clippingdale, A.B.; Fearnley, I.M.; Harbour, M.; Robinson, A.J.; et al. The AAA+ protein ATAD3 has displacement loop binding properties and is involved in mitochondrial nucleoid organization. J. Cell Biol. 2007, 176, 141–146. [Google Scholar] [CrossRef]
- Han, S.; Udeshi, N.D.; Deerinck, T.J.; Svinkina, T.; Ellisman, M.H.; Carr, S.A.; Ting, A.Y. Proximity Biotinylation as a Method for Mapping Proteins Associated with mtDNA in Living Cells. Cell Chem. Biol. 2017, 24, 404–414. [Google Scholar] [CrossRef] [PubMed]
- Lewis, M.R.; Lewis, W.H. Mitochondria in Tissue Culture. Science 1914, 39, 330–333. [Google Scholar] [CrossRef] [PubMed]
- Bereiter-Hahn, J.; Voth, M. Dynamics of mitochondria in living cells: Shape changes, dislocations, fusion, and fission of mitochondria. Microsc. Res. Tech. 1994, 27, 198–219. [Google Scholar] [CrossRef] [PubMed]
- Nunnari, J.; Marshall, W.F.; Straight, A.; Murray, A.; Sedat, J.W.; Walter, P. Mitochondrial transmission during mating in Saccharomyces cerevisiae is determined by mitochondrial fusion and fission and the intramitochondrial segregation of mitochondrial DNA. Mol. Biol. Cell 1997, 8, 1233–1242. [Google Scholar] [CrossRef]
- Hermann, G.J.; Shaw, J.M. Mitochondrial dynamics in yeast. Annu. Rev. Cell Dev. Biol. 1998, 14, 265–303. [Google Scholar] [CrossRef] [PubMed]
- Shaw, J.M.; Nunnari, J. Mitochondrial dynamics and division in budding yeast. Trends Cell Biol. 2002, 12, 178–184. [Google Scholar] [CrossRef]
- Osteryoung, K.W.; Nunnari, J. The division of endosymbiotic organelles. Science 2003, 302, 1698–1704. [Google Scholar] [CrossRef] [PubMed]
- Ono, T.; Isobe, K.; Nakada, K.; Hayashi, J.I. Human cells are protected from mitochondrial dysfunction by complementation of DNA products in fused mitochondria. Nat. Genet. 2001, 28, 272–275. [Google Scholar] [CrossRef] [PubMed]
- Westermann, B. Mitochondrial fusion and fission in cell life and death. Nat. Rev. Mol. Cell Biol. 2010, 11, 872–884. [Google Scholar] [CrossRef] [PubMed]
- Qi, Y.; Yan, L.; Yu, C.; Guo, X.; Zhou, X.; Hu, X.; Huang, X.; Rao, Z.; Lou, Z.; Hu, J. Structures of human mitofusin 1 provide insight into mitochondrial tethering. J. Cell Biol. 2016, 215, 621–629. [Google Scholar] [CrossRef] [PubMed]
- Koshiba, T.; Detmer, S.A.; Kaiser, J.T.; Chen, H.; McCaffery, J.M.; Chan, D.C. Structural basis of mitochondrial tethering by mitofusin complexes. Science 2004, 305, 858–862. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Qi, Y.; Huang, X.; Yu, C.; Lan, L.; Guo, X.; Rao, Z.; Hu, J.; Lou, Z. Structural basis for GTP hydrolysis and conformational change of MFN1 in mediating membrane fusion. Nat. Struct. Mol. Biol. 2018, 25, 233–243. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.L.; Meng, S.; Chen, Y.; Feng, J.X.; Gu, D.D.; Yu, B.; Li, Y.J.; Yang, J.Y.; Liao, S.; Chan, D.C.; et al. MFN1 structures reveal nucleotide-triggered dimerization critical for mitochondrial fusion. Nature 2017, 542, 372–376. [Google Scholar] [CrossRef] [PubMed]
- Rojo, M.; Legros, F.; Chateau, D.; Lombes, A. Membrane topology and mitochondrial targeting of mitofusins, ubiquitous mammalian homologs of the transmembrane GTPase Fzo. J. Cell Sci. 2002, 115, 1663–1674. [Google Scholar] [PubMed]
- Mattie, S.; Krols, M.; McBride, H.M. The enigma of an interconnected mitochondrial reticulum: New insights into mitochondrial fusion. Curr. Opin. Cell Biol. 2019, 59, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Mattie, S.; Riemer, J.; Wideman, J.G.; McBride, H.M. A new mitofusin topology places the redox-regulated C terminus in the mitochondrial intermembrane space. J. Cell Biol. 2018, 217, 507–515. [Google Scholar] [CrossRef]
- Zuchner, S.; Mersiyanova, I.V.; Muglia, M.; Bissar-Tadmouri, N.; Rochelle, J.; Dadali, E.L.; Zappia, M.; Nelis, E.; Patitucci, A.; Senderek, J.; et al. Mutations in the mitochondrial GTPase mitofusin 2 cause Charcot-Marie-Tooth neuropathy type 2A. Nat. Genet. 2004, 36, 449–451. [Google Scholar] [CrossRef]
- Smirnova, E.; Griparic, L.; Shurland, D.L.; van der Bliek, A.M. Dynamin-related protein Drp1 is required for mitochondrial division in mammalian cells. Mol. Biol. Cell 2001, 12, 2245–2256. [Google Scholar] [CrossRef]
- Chen, H.; Detmer, S.A.; Ewald, A.J.; Griffin, E.E.; Fraser, S.E.; Chan, D.C. Mitofusins Mfn1 and Mfn2 coordinately regulate mitochondrial fusion and are essential for embryonic development. J. Cell Biol. 2003, 160, 189–200. [Google Scholar] [CrossRef]
- Chan, D.C. Mitochondrial fusion and fission in mammals. Annu. Rev. Cell Dev. Biol. 2006, 22, 79–99. [Google Scholar] [CrossRef]
- Chan, D.C. Fusion and fission: Interlinked processes critical for mitochondrial health. Annu. Rev. Genet. 2012, 46, 265–287. [Google Scholar] [CrossRef]
- Tilokani, L.; Nagashima, S.; Paupe, V.; Prudent, J. Mitochondrial dynamics: Overview of molecular mechanisms. Essays Biochem. 2018, 62, 341–360. [Google Scholar] [CrossRef]
- Van der Bliek, A.M.; Shen, Q.; Kawajiri, S. Mechanisms of mitochondrial fission and fusion. Cold Spring Harb. Perspect. Biol. 2013, 5, 6. [Google Scholar] [CrossRef] [PubMed]
- Meeusen, S.; McCaffery, J.M.; Nunnari, J. Mitochondrial fusion intermediates revealed in vitro. Science 2004, 305, 1747–1752. [Google Scholar] [CrossRef]
- Cipolat, S.; Martins de Brito, O.; Dal Zilio, B.; Scorrano, L. OPA1 requires mitofusin 1 to promote mitochondrial fusion. Proc. Natl. Acad. Sci. USA 2004, 101, 15927–15932. [Google Scholar] [CrossRef] [PubMed]
- Frank, S.; Gaume, B.; Bergmann-Leitner, E.S.; Leitner, W.W.; Robert, E.G.; Catez, F.; Smith, C.L.; Youle, R.J. The role of dynamin-related protein 1, a mediator of mitochondrial fission, in apoptosis. Dev. Cell 2001, 1, 515–525. [Google Scholar] [CrossRef]
- Duvezin-Caubet, S.; Jagasia, R.; Wagener, J.; Hofmann, S.; Trifunovic, A.; Hansson, A.; Chomyn, A.; Bauer, M.F.; Attardi, G.; Larsson, N.G.; et al. Proteolytic Processing of OPA1 Links Mitochondrial Dysfunction to Alterations in Mitochondrial Morphology. J. Biol. Chem. 2006, 281, 37972–37979. [Google Scholar] [CrossRef] [PubMed]
- Herlan, M.; Bornhovd, C.; Hell, K.; Neupert, W.; Reichert, A.S. Alternative topogenesis of Mgm1 and mitochondrial morphology depend on ATP and a functional import motor. J. Cell Biol. 2004, 165, 167–173. [Google Scholar] [CrossRef]
- Twig, G.; Elorza, A.; Molina, A.J.; Mohamed, H.; Wikstrom, J.D.; Walzer, G.; Stiles, L.; Haigh, S.E.; Katz, S.; Las, G.; et al. Fission and selective fusion govern mitochondrial segregation and elimination by autophagy. Embo J. 2008, 27, 433–446. [Google Scholar] [CrossRef]
- Zimmermann, M.; Reichert, A.S. How to get rid of mitochondria: Crosstalk and regulation of multiple mitophagy pathways. Biol. Chem. 2017, 399, 29–45. [Google Scholar] [CrossRef] [PubMed]
- Lyamzaev, K.G.; Pletjushkina, O.Y.; Saprunova, V.B.; Bakeeva, L.E.; Chernyak, B.V.; Skulachev, V.P. Selective elimination of mitochondria from living cells induced by inhibitors of bioenergetic functions. Biochem. Soc. Trans. 2004, 32, 1070–1071. [Google Scholar] [CrossRef] [PubMed]
- Otera, H.; Wang, C.; Cleland, M.M.; Setoguchi, K.; Yokota, S.; Youle, R.J.; Mihara, K. Mff is an essential factor for mitochondrial recruitment of Drp1 during mitochondrial fission in mammalian cells. J. Cell Biol. 2010, 191, 1141–1158. [Google Scholar] [CrossRef] [PubMed]
- Otera, H.; Miyata, N.; Kuge, O.; Mihara, K. Drp1-dependent mitochondrial fission via MiD49/51 is essential for apoptotic cristae remodeling. J. Cell Biol. 2016, 212, 531–544. [Google Scholar] [CrossRef] [PubMed]
- Loson, O.C.; Song, Z.; Chen, H.; Chan, D.C. Fis1, Mff, MiD49, and MiD51 mediate Drp1 recruitment in mitochondrial fission. Mol. Biol. Cell 2013, 24, 659–667. [Google Scholar] [CrossRef]
- Lee, J.E.; Westrate, L.M.; Wu, H.; Page, C.; Voeltz, G.K. Multiple dynamin family members collaborate to drive mitochondrial division. Nature 2016, 540, 139–143. [Google Scholar] [CrossRef] [PubMed]
- Kamerkar, S.C.; Kraus, F.; Sharpe, A.J.; Pucadyil, T.J.; Ryan, M.T. Dynamin-related protein 1 has membrane constricting and severing abilities sufficient for mitochondrial and peroxisomal fission. Nat. Commun. 2018, 9, 5239. [Google Scholar] [CrossRef]
- Fonseca, T.B.; Sanchez-Guerrero, A.; Milosevic, I.; Raimundo, N. Mitochondrial fission requires DRP1 but not dynamins. Nature 2019, 570, E34–E42. [Google Scholar] [CrossRef]
- Friedman, J.R.; Lackner, L.L.; West, M.; DiBenedetto, J.R.; Nunnari, J.; Voeltz, G.K. ER tubules mark sites of mitochondrial division. Science 2011, 334, 358–362. [Google Scholar] [CrossRef]
- Korobova, F.; Ramabhadran, V.; Higgs, H.N. An actin-dependent step in mitochondrial fission mediated by the ER-associated formin INF2. Science 2013, 339, 464–467. [Google Scholar] [CrossRef]
- Fahrner, J.A.; Liu, R.; Perry, M.S.; Klein, J.; Chan, D.C. A novel de novo dominant negative mutation in DNM1L impairs mitochondrial fission and presents as childhood epileptic encephalopathy. Am. J. Med Genet. 2016, 170, 2002–2011. [Google Scholar] [CrossRef] [PubMed]
- Sheffer, R.; Douiev, L.; Edvardson, S.; Shaag, A.; Tamimi, K.; Soiferman, D.; Meiner, V.; Saada, A. Postnatal microcephaly and pain insensitivity due to a de novo heterozygous DNM1L mutation causing impaired mitochondrial fission and function. Am. J. Med. Genet. 2016, 170, 1603–1607. [Google Scholar] [CrossRef] [PubMed]
- Waterham, H.R.; Koster, J.; van Roermund, C.W.; Mooyer, P.A.; Wanders, R.J.; Leonard, J.V. A lethal defect of mitochondrial and peroxisomal fission. N. Engl. J. Med. 2007, 356, 1736–1741. [Google Scholar] [CrossRef] [PubMed]
- Jones, B.A.; Fangman, W.L. Mitochondrial DNA maintenance in yeast requires a protein containing a region related to the GTP-binding domain of dynamin. Genes Dev. 1992, 6, 380–389. [Google Scholar] [CrossRef] [PubMed]
- Guan, K.; Farh, L.; Marshall, T.K.; Deschenes, R.J. Normal mitochondrial structure and genome maintenance in yeast requires the dynamin-like product of the MGM1 gene. Curr. Genet. 1993, 24, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Hermann, G.J.; Thatcher, J.W.; Mills, J.P.; Hales, K.G.; Fuller, M.T.; Nunnari, J.; Shaw, J.M. Mitochondrial fusion in yeast requires the transmembrane GTPase Fzo1p. J. Cell Biol. 1998, 143, 359–373. [Google Scholar] [CrossRef]
- Rapaport, D.; Brunner, M.; Neupert, W.; Westermann, B. Fzo1p is a mitochondrial outer membrane protein essential for the biogenesis of functional mitochondria in Saccharomyces cerevisiae. J. Biol. Chem. 1998, 273, 20150–20155. [Google Scholar] [CrossRef] [PubMed]
- Sesaki, H.; Jensen, R.E. UGO1 encodes an outer membrane protein required for mitochondrial fusion. J. Cell Biol. 2001, 152, 1123–1134. [Google Scholar] [CrossRef]
- Sesaki, H.; Jensen, R.E. Ugo1p links the Fzo1p and Mgm1p GTPases for mitochondrial fusion. J. Biol. Chem. 2004, 279, 28298–28303. [Google Scholar] [CrossRef]
- Chen, H.; McCaffery, J.M.; Chan, D.C. Mitochondrial fusion protects against neurodegeneration in the cerebellum. Cell 2007, 130, 548–562. [Google Scholar] [CrossRef]
- Chen, H.; Chomyn, A.; Chan, D.C. Disruption of fusion results in mitochondrial heterogeneity and dysfunction. J. Biol. Chem. 2005, 280, 26185–26192. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Vermulst, M.; Wang, Y.E.; Chomyn, A.; Prolla, T.A.; McCaffery, J.M.; Chan, D.C. Mitochondrial fusion is required for mtDNA stability in skeletal muscle and tolerance of mtDNA mutations. Cell 2010, 141, 280–289. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Liu, Y.; Dorn, G.W., 2nd. Mitochondrial fusion is essential for organelle function and cardiac homeostasis. Circ. Res. 2011, 109, 1327–1331. [Google Scholar] [CrossRef] [PubMed]
- Silva Ramos, E.; Motori, E.; Bruser, C.; Kuhl, I.; Yeroslaviz, A.; Ruzzenente, B.; Kauppila, J.H.K.; Busch, J.D.; Hultenby, K.; Habermann, B.H.; et al. Mitochondrial fusion is required for regulation of mitochondrial DNA replication. PLoS Genet. 2019, 15, e1008085. [Google Scholar] [CrossRef] [PubMed]
- Ishihara, N.; Nomura, M.; Jofuku, A.; Kato, H.; Suzuki, S.O.; Masuda, K.; Otera, H.; Nakanishi, Y.; Nonaka, I.; Goto, Y.; et al. Mitochondrial fission factor Drp1 is essential for embryonic development and synapse formation in mice. Nat. Cell Biol. 2009, 11, 958–966. [Google Scholar] [CrossRef] [PubMed]
- Wakabayashi, J.; Zhang, Z.; Wakabayashi, N.; Tamura, Y.; Fukaya, M.; Kensler, T.W.; Iijima, M.; Sesaki, H. The dynamin-related GTPase Drp1 is required for embryonic and brain development in mice. J. Cell Biol. 2009, 186, 805–816. [Google Scholar] [CrossRef] [PubMed]
- Parone, P.A.; Da Cruz, S.; Tondera, D.; Mattenberger, Y.; James, D.I.; Maechler, P.; Barja, F.; Martinou, J.C. Preventing mitochondrial fission impairs mitochondrial function and leads to loss of mitochondrial DNA. PLoS ONE 2008, 3, e3257. [Google Scholar] [CrossRef] [PubMed]
- Kageyama, Y.; Zhang, Z.; Roda, R.; Fukaya, M.; Wakabayashi, J.; Wakabayashi, N.; Kensler, T.W.; Reddy, P.H.; Iijima, M.; Sesaki, H. Mitochondrial division ensures the survival of postmitotic neurons by suppressing oxidative damage. J. Cell Biol. 2012, 197, 535–551. [Google Scholar] [CrossRef] [PubMed]
- Ban-Ishihara, R.; Ishihara, T.; Sasaki, N.; Mihara, K.; Ishihara, N. Dynamics of nucleoid structure regulated by mitochondrial fission contributes to cristae reformation and release of cytochrome c. Proc. Natl. Acad. Sci. USA 2013, 110, 11863–11868. [Google Scholar] [CrossRef]
- Ishihara, T.; Ban-Ishihara, R.; Maeda, M.; Matsunaga, Y.; Ichimura, A.; Kyogoku, S.; Aoki, H.; Katada, S.; Nakada, K.; Nomura, M.; et al. Dynamics of mitochondrial DNA nucleoids regulated by mitochondrial fission is essential for maintenance of homogeneously active mitochondria during neonatal heart development. Mol. Cell. Biol. 2015, 35, 211–223. [Google Scholar] [CrossRef]
- Chen, H.; Ren, S.; Clish, C.; Jain, M.; Mootha, V.; McCaffery, J.M.; Chan, D.C. Titration of mitochondrial fusion rescues Mff-deficient cardiomyopathy. J. Cell Biol. 2015, 211, 795–805. [Google Scholar] [CrossRef] [PubMed]
- Jensen, R.E.; Hobbs, A.E.; Cerveny, K.L.; Sesaki, H. Yeast mitochondrial dynamics: Fusion, division, segregation, and shape. Microsc. Res. Tech. 2000, 51, 573–583. [Google Scholar] [CrossRef]
- Bleazard, W.; McCaffery, J.M.; King, E.J.; Bale, S.; Mozdy, A.; Tieu, Q.; Nunnari, J.; Shaw, J.M. The dynamin-related GTPase Dnm1 regulates mitochondrial fission in yeast. Nat. Cell Biol. 1999, 1, 298–304. [Google Scholar] [CrossRef] [PubMed]
- Wong, E.D.; Wagner, J.A.; Gorsich, S.W.; McCaffery, J.M.; Shaw, J.M.; Nunnari, J. The dynamin-related GTPase, Mgm1p, is an intermembrane space protein required for maintenance of fusion competent mitochondria. J. Cell Biol. 2000, 151, 341–352. [Google Scholar] [CrossRef] [PubMed]
- Lewis, S.C.; Uchiyama, L.F.; Nunnari, J. ER-mitochondria contacts couple mtDNA synthesis with mitochondrial division in human cells. Science 2016, 353, aaf5549. [Google Scholar] [CrossRef]
- Kim, J.Y.; Hwang, J.M.; Ko, H.S.; Seong, M.W.; Park, B.J.; Park, S.S. Mitochondrial DNA content is decreased in autosomal dominant optic atrophy. Neurology 2005, 64, 966–972. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Liu, T.; Tran, A.; Lu, X.; Tomilov, A.A.; Davies, V.; Cortopassi, G.; Chiamvimonvat, N.; Bers, D.M.; Votruba, M.; et al. OPA1 mutation and late-onset cardiomyopathy: Mitochondrial dysfunction and mtDNA instability. J. Am. Heart Assoc. 2012, 1, e003012. [Google Scholar] [CrossRef]
- Alavi, M.V.; Bette, S.; Schimpf, S.; Schuettauf, F.; Schraermeyer, U.; Wehrl, H.F.; Ruttiger, L.; Beck, S.C.; Tonagel, F.; Pichler, B.J.; et al. A splice site mutation in the murine Opa1 gene features pathology of autosomal dominant optic atrophy. Brain 2007, 130, 1029–1042. [Google Scholar] [CrossRef]
- Tezze, C.; Romanello, V.; Desbats, M.A.; Fadini, G.P.; Albiero, M.; Favaro, G.; Ciciliot, S.; Soriano, M.E.; Morbidoni, V.; Cerqua, C.; et al. Age-Associated Loss of OPA1 in Muscle Impacts Muscle Mass, Metabolic Homeostasis, Systemic Inflammation, and Epithelial Senescence. Cell Metab. 2017, 25, 1374–1389. [Google Scholar] [CrossRef]
- Cogliati, S.; Frezza, C.; Soriano, M.E.; Varanita, T.; Quintana-Cabrera, R.; Corrado, M.; Cipolat, S.; Costa, V.; Casarin, A.; Gomes, L.C.; et al. Mitochondrial cristae shape determines respiratory chain supercomplexes assembly and respiratory efficiency. Cell 2013, 155, 160–171. [Google Scholar] [CrossRef]
- Romanello, V.; Scalabrin, M.; Albiero, M.; Blaauw, B.; Scorrano, L.; Sandri, M. Inhibition of the Fission Machinery Mitigates OPA1 Impairment in Adult Skeletal Muscles. Cells 2019, 8, 597. [Google Scholar] [CrossRef] [PubMed]
- John, G.B.; Shang, Y.; Li, L.; Renken, C.; Mannella, C.A.; Selker, J.M.; Rangell, L.; Bennett, M.J.; Zha, J. The mitochondrial inner membrane protein mitofilin controls cristae morphology. Mol. Biol. Cell 2005, 16, 1543–1554. [Google Scholar] [CrossRef] [PubMed]
- Itoh, K.; Tamura, Y.; Iijima, M.; Sesaki, H. Effects of Fcj1-Mos1 and mitochondrial division on aggregation of mitochondrial DNA nucleoids and organelle morphology. Mol. Biol. Cell 2013, 24, 1842–1851. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Ruan, Y.; Zhang, K.; Jian, F.; Hu, C.; Miao, L.; Gong, L.; Sun, L.; Zhang, X.; Chen, S.; et al. Mic60/Mitofilin determines MICOS assembly essential for mitochondrial dynamics and mtDNA nucleoid organization. Cell Death Differ. 2015, 22, 380–392. [Google Scholar] [CrossRef] [PubMed]
- Bornhovd, C.; Vogel, F.; Neupert, W.; Reichert, A.S. Mitochondrial membrane potential is dependent on the oligomeric state of F1F0-ATP synthase supracomplexes. J. Biol. Chem. 2006, 281, 13990–13998. [Google Scholar] [CrossRef] [PubMed]
- Baseler, W.A.; Dabkowski, E.R.; Williamson, C.L.; Croston, T.L.; Thapa, D.; Powell, M.J.; Razunguzwa, T.T.; Hollander, J.M. Proteomic alterations of distinct mitochondrial subpopulations in the type 1 diabetic heart: Contribution of protein import dysfunction. Am. J. Physiol. 2011, 300, R186–R200. [Google Scholar] [CrossRef] [PubMed]
- Thapa, D.; Nichols, C.E.; Lewis, S.E.; Shepherd, D.L.; Jagannathan, R.; Croston, T.L.; Tveter, K.J.; Holden, A.A.; Baseler, W.A.; Hollander, J.M. Transgenic overexpression of mitofilin attenuates diabetes mellitus-associated cardiac and mitochondria dysfunction. J. Mol. Cell. Cardiol. 2015, 79, 212–223. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.R.; Kim, N.; Noh, Y.H.; Xu, Z.; Ko, K.S.; Rhee, B.D.; Han, J. Mitochondrial DNA, mitochondrial dysfunction, and cardiac manifestations. Front. Biosci. 2017, 22, 1177–1194. [Google Scholar]
- Arbustini, E.; Diegoli, M.; Fasani, R.; Grasso, M.; Morbini, P.; Banchieri, N.; Bellini, O.; Dal Bello, B.; Pilotto, A.; Magrini, G.; et al. Mitochondrial DNA mutations and mitochondrial abnormalities in dilated cardiomyopathy. Am. J. Pathol. 1998, 153, 1501–1510. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).