Molecular Characterization of ZosmaNRT2, the Putative Sodium Dependent High-Affinity Nitrate Transporter of Zostera marina L.
Abstract
1. Introduction
2. Results
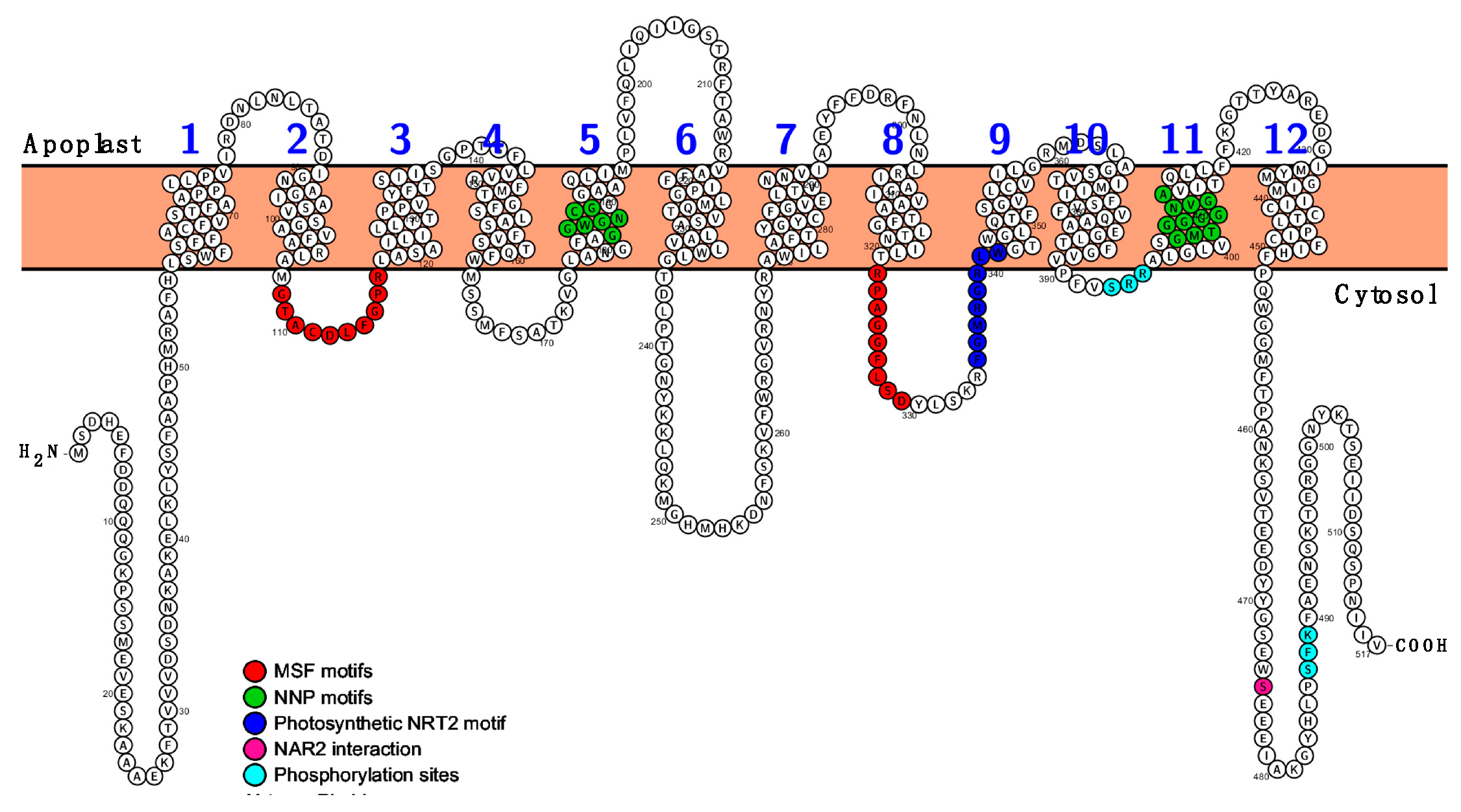
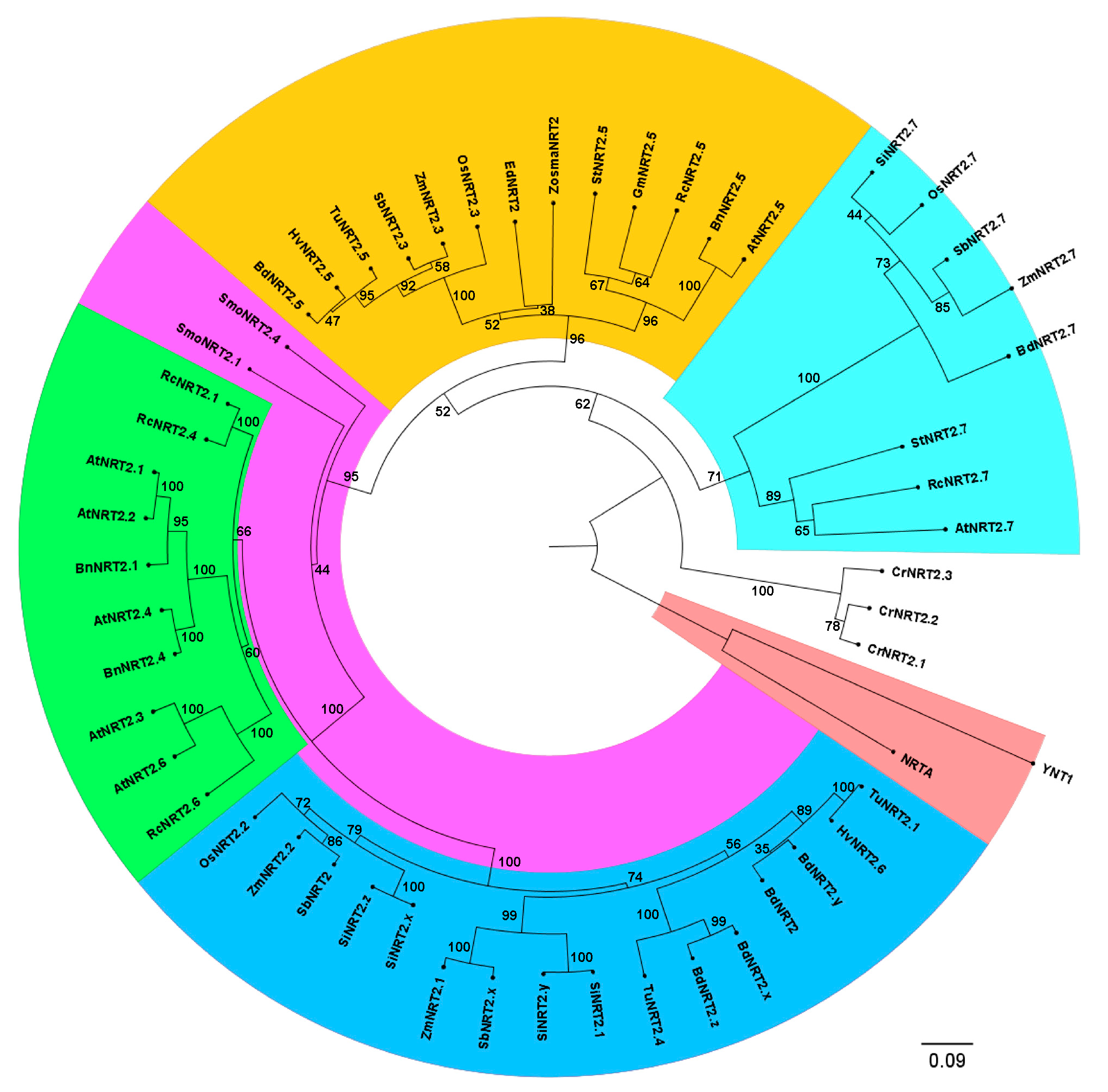
2.1. Zostera marina High-Affinity NO3− Transporter Secuence Identification
2.2. The Coding Sequences of ZosmaNRT2 and ZosmaNAR2 were Isolated from Z. marina Leaves mRNA
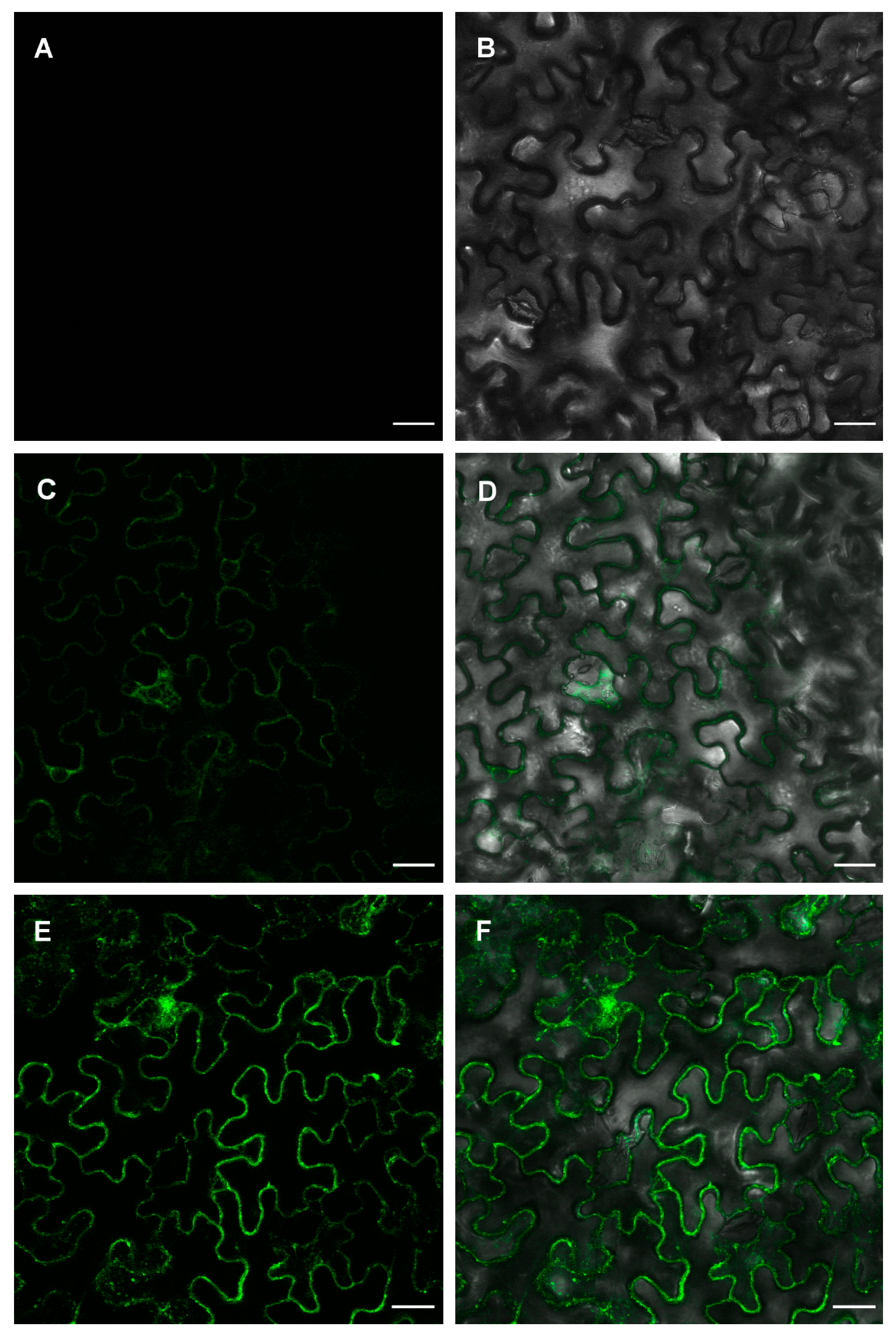
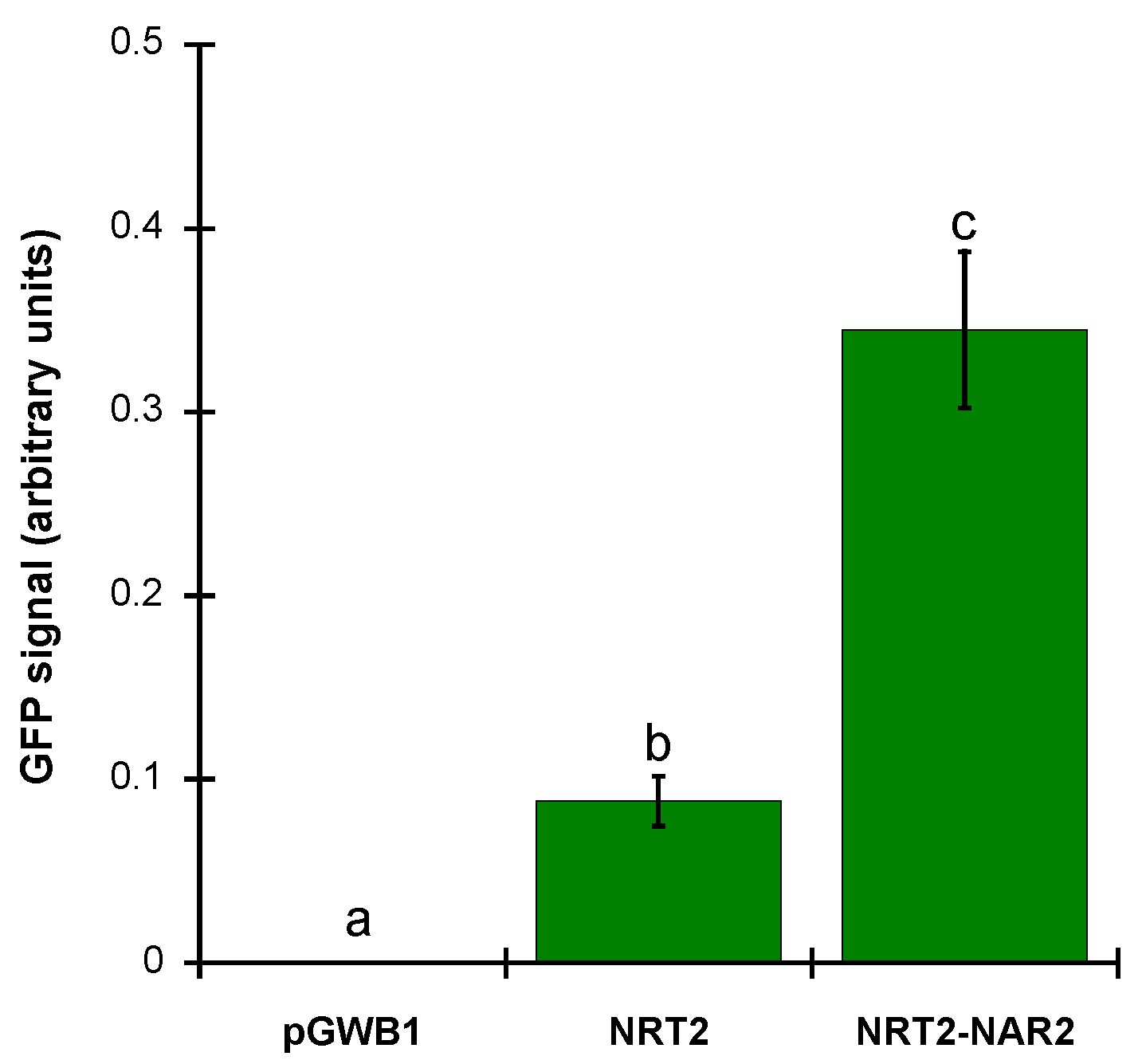
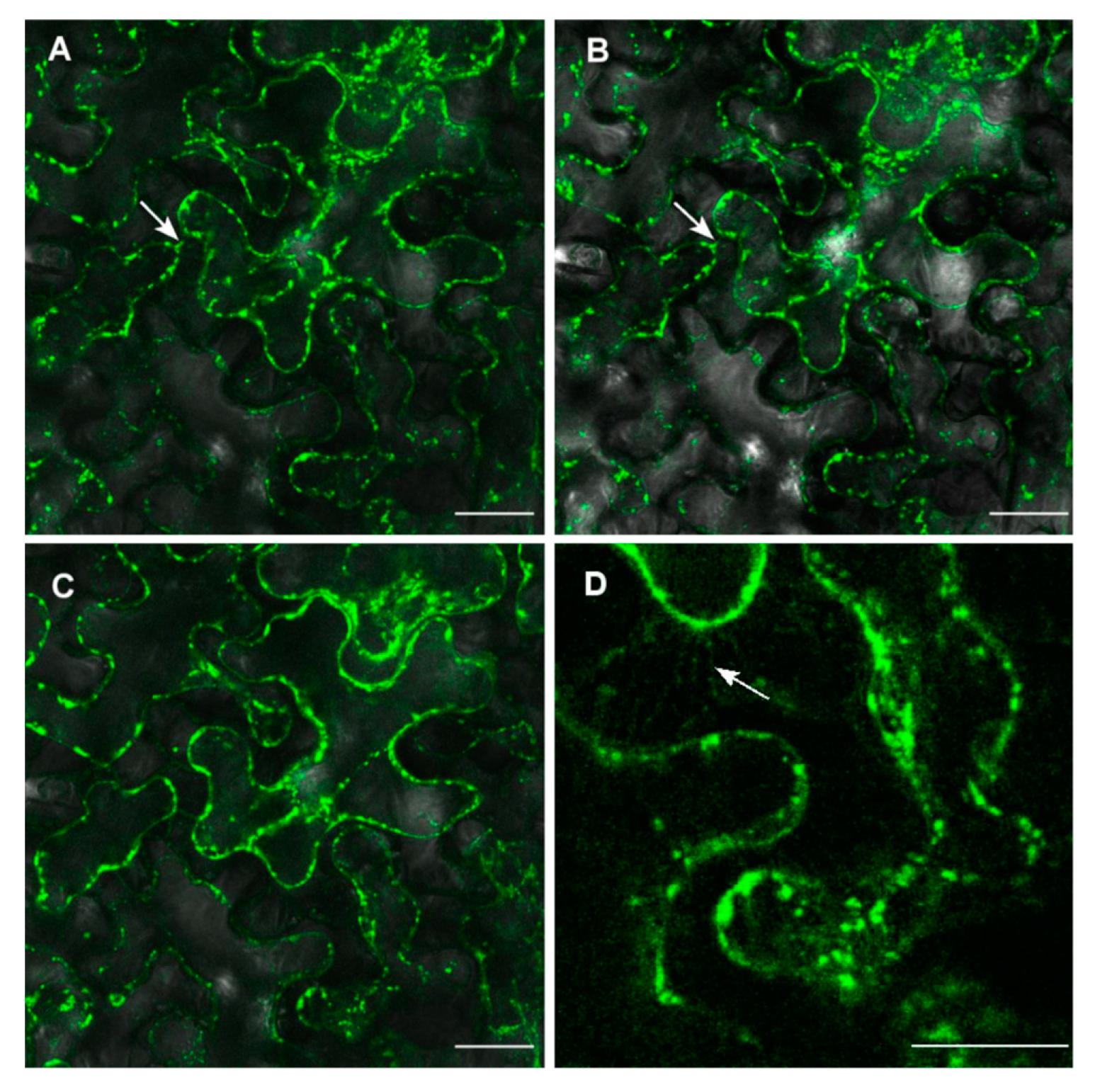
2.3. ZosmaNAR2 Stabilizes the Location of ZosmaNRT2 at the Plasma Membrane
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Sequence and Phylogenetic Analysis
4.3. Obtaining NRT2 and NAR2 cDNAs from Z. marina Leaves.
4.4. Molecular Cloning
4.5. Transient Expression in N. benthamiana Leaves and Subcelular Localization
4.6. Image Processing and Fluorescence Intensity Quantification
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| HAT | High-affinity transporter |
| MFS | Mayor facilitator superfamily |
| NNP | Nitrate nitrite porter |
| NCBI | National center of biotechnology information |
| NSW | Natural sea water |
| NRT2 | Nitrate Transporter 2 |
References
- Touchette, B.W.; Burkholder, J.M. Review of nitrogen and phosphorus metabolism in seagrasses. J. Exp. Mar. Biol. Ecol. 2000, 250, 133–167. [Google Scholar] [CrossRef]
- Fukuhara, T.; Pak, J.Y.; Ohwaki, Y.; Tsujimura, H.; Nitta, T. Tissue-specific expression of the gene for a putative plasma membrane H+-ATPase in a seagrass. Plant Physiol. 1996, 110, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Fernández, J.A.; García-Sánchez, M.J.; Felle, H.H. Physiological evidence for a proton pump and sodium exclusion mechanisms at the plasma membrane of the marine angiosperm Zostera marina L. J. Exp. Bot. 1999, 50, 1763–1768. [Google Scholar] [CrossRef]
- García-Sánchez, M.J.; Jaime, M.P.; Ramos, A.; Sanders, D.; Fernández, J.A. Sodium-dependent nitrate transport at the plasma membrane of leaf cells of the marine higher plant Zostera marina L. Plant Physiol. 2000, 122, 879–886. [Google Scholar] [CrossRef] [PubMed]
- Rubio, L.; Linares-Rueda, A.; García-Sánchez, M.J.; Fernández, J.A. Physiological evidence for a sodium-dependent high-affinity phosphate and nitrate transport at the plasma membrane of leaf and root cells of Zostera marina L. J. Exp. Bot. 2005, 56, 613–622. [Google Scholar] [CrossRef] [PubMed]
- Rubio, L.; García-Pérez, D.; García-Sánchez, M.J.; Fernández, J.A. Na+-dependent high-affinity nitrate; phosphate and amino acids transport in leaf cells of the seagrass Posidonia oceanica (L.) Delile. Int. J. Mol. Sci. 2018, 19, 1570. [Google Scholar] [CrossRef] [PubMed]
- Olsen, J.L.; Rouzé, P.; Verhelst, B.; Lin, Y.C.; Bayer, T.; Collen, J.; Dattolo, E.; De Paoli, E.; Dittami, S.; Maumus, F.; et al. The genome of the seagrass Zostera marina reveals angiosperm adaptation to the sea. Nature 2016, 530, 331–335. [Google Scholar] [CrossRef]
- Lee, H.; Golicz, A.A.; Bayer, P.E.; Jiao, Y.; Tang, H.; Paterson, A.H.; Sablok, G.; Krishnaraj, R.R.; Chan, C.K.; Batley, J.; et al. The Genome of a Southern Hemisphere Seagrass Species (Zostera muelleri). Plant Physiol. 2016, 172, 272–283. [Google Scholar] [CrossRef]
- Williams, S.L. Genomics: From sea to sea. Nature 2016, 530, 290–291. [Google Scholar] [CrossRef]
- Fan, X.; Naz, M.; Fan, X.; Xuan, W.; Miller, A.J.; Xu, G. Plant nitrate transporters: From gene function to application. J. Exp. Bot. 2017, 68, 2463–2475. [Google Scholar] [CrossRef]
- Siddiqi, M.Y.; Glass, A.D.M.; Shaff, J.E.; Kochian, L.V. Studies of the Uptake of Nitrate in Barley: IV. Electrophysiology. Plant Physiol. 2008, 99, 456–463. [Google Scholar] [CrossRef]
- Miller, A.J.; Fan, X.; Orsel, M.; Smith, S.J.; Wells, D.M. Nitrate transport and signalling. J. Exp. Bot. 2007, 58, 2297–2306. [Google Scholar] [CrossRef] [PubMed]
- Léran, S.; Varala, K.; Boyer, J.C.; Chiurazzi, M.; Crawford, N.; Daniel-Vedele, F.; David, L.; Dickstein, R.; Fernandez, E.; Forde, B.; et al. A unified nomenclature of NITRATE TRANSPORTER 1/PEPTIDE TRANSPORTER family members in plants. Trends Plant Sci. 2014, 19, 5–9. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Tsay, Y. Switching between the two action modes of the dual-affinity nitrate transporter CHL1 by phosphorylation. EMBO J. 2003, 22, 1005–1013. [Google Scholar] [CrossRef] [PubMed]
- Quesada, A.; Galvan, A.; Fernandez, E. Identification of nitrate transporter genes in Chlamydomonas reinhardtii. Plant J. 1994, 5, 407–419. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Hsu, P.; Tsay, Y. Uptake; allocation and signaling of nitrate. Trends Plant Sci. 2012, 17, 458–467. [Google Scholar] [CrossRef] [PubMed]
- Bergsdorf, E.Y.; Zdebik, A.A.; Jentsch, T.J. Residues Important for Nitrate/Proton Coupling in Plant and Mammalian CLC Transporters. J. Biol. Chem. 2009, 284, 11184–11193. [Google Scholar] [CrossRef] [PubMed]
- Geiger, D.; Maierhofer, T.; Al-Rasheid, K.A.; Scherzer, S.; Mumm, P.; Liese, A.; Ache, P.; Wellmann, C.; Marten, I.; Grill, E.; et al. Stomatal closure by fast abscisic acid signaling is mediated by the guard cell anion channel SLAH3 and the receptor RCAR1. Sci. Signal. 2011, 4, ra32. [Google Scholar] [CrossRef] [PubMed]
- Maierhofer, T.; Diekmann, M.; Offenborn, J.N.; Lind, C.; Bauer, H.; Hashimoto, K.S.; Al-Rasheid, K.A.; Luan, S.; Kudla, J.; Geiger, D.; et al. Site- and kinase-specific phosphorylation-mediated activation of SLAC1, a guard cell anion channel stimulated by abscisic acid. Sci. Signal. 2014, 7, ra86. [Google Scholar] [CrossRef]
- Chopin, F.; Orsel, M.; Dorbe, M.F.; Chardon, F.; Truong, H.N.; Miller, A.J.; Krapp, A.; Daniel-Vedele, F. The Arabidopsis ATNRT2.7 nitrate transporter controls nitrate content in seeds. Plant Cell 2007, 19, 1590–1602. [Google Scholar] [CrossRef]
- Li, W.; Wang, Y.; Okamoto, M.; Crawford, N.M.; Siddiqi, M.Y.; Glass, A.D.M. Dissection of the AtNRT2.1:AtNRT2.2 inducible high-affinity nitrate transporter gene cluster. Plant Phisiol. 2007, 143, 425–433. [Google Scholar] [CrossRef] [PubMed]
- Kotur, Z.; Mackenzie, N.; Ramesh, S.; Tyerman, S.D.; Kaiser, B.N.; Glass, A.D.M. Nitrate transport capacity of the Arabidopsis thaliana NRT2 family members and their interactions with AtNAR2.1. New Phytol. 2012, 194, 724–731. [Google Scholar] [CrossRef] [PubMed]
- Orsel, M.; Chopin, F.; Leleu, O.; Smith, S.J.; Krapp, A.; Daniel-Vedele, F.; Miller, A.J. Nitrate signaling and the two component high affinity uptake system in Arabidopsis. Plant Signal. Behav. 2007, 2, 260–262. [Google Scholar] [CrossRef] [PubMed]
- Lezhneva, L.; Kiba, T.; Feria-Bourrellier, A.B.; Lafouge, F.; Boutet-Mercey, S.; Zoufan, P.; Sakakibara, H.; Daniel-Vedele, F.; Krapp, A. The Arabidopsis nitrate transporter NRT2.5 plays a role in nitrate acquisition and remobilization in nitrogen-starved plants. Plant J. 2014, 80, 230–241. [Google Scholar] [CrossRef] [PubMed]
- Kiba, T.; Feria-Bourrellier, A.B.; Lafouge, F.; Lezhneva, L.; Boutet-Mercey, S.; Orsel, M.; Bréhaut, V.; Miller, A.; Daniel-Vedele, F.; Sakakibara, H.; et al. The Arabidopsis nitrate transporter NRT2.4 plays a double role in roots and shoots of nitrogen-starved plants. Plant Cell 2012, 24, 245–258. [Google Scholar] [CrossRef]
- Orsel, M.; Krapp, A.; Daniel-Vedele, F. Analysis of the NRT2 nitrate transporter family in Arabidopsis. Structure and gene expression. Plant Phisiol. 2002, 129, 886–896. [Google Scholar] [CrossRef]
- Plett, D.; Toubia, J.; Garnett, T.; Tester, M.; Kaiser, B.N.; Baumann, U. Dichotomy in the NRT gene families of dicots and grass species. PLoS ONE 2010, 5, e15289. [Google Scholar] [CrossRef]
- Forde, B.G. Nitrate transporters in plants: Structure; function and regulation. Biochim. Biophys. Acta 2000, 1465, 219–235. [Google Scholar] [CrossRef]
- Jacquot, A.; Li, Z.; Gojon, A.; Schulze, W.; Lejay, L. Post-translational regulation of nitrogen transporters in plants and microorganisms. J. Exp. Bot. 2017, 68, 2567–2580. [Google Scholar] [CrossRef]
- Pao, S.S.; Paulsen, I.T.; Saier, M.H., Jr. Major facilitator superfamily. Microbiol. Mol. Biol. Rev. 1998, 62, 1–34. [Google Scholar] [CrossRef]
- Okamoto, M.; Vidmar, J.J.; Glass, A.D.M. Regulation of NRT1 and NRT2 gene families of Arabidopsis thaliana: Responses to nitrate provision. Plant Cell Phisiol. 2003, 44, 304–317. [Google Scholar] [CrossRef] [PubMed]
- Cabrera, E.; González-Montelongo, R.; Giraldez, T.; de la Rosa, D.A.; Siverio, J.M. Molecular components of nitrate and nitrite efflux in yeast. Eukaryot. Cell 2014, 13, 267–278. [Google Scholar] [CrossRef] [PubMed]
- Hresko, R.C.; Kruse, M.; Strube, M.; Muecklers, M. Topology of the Glut 1 glucose transporter deduced from glycosylation scanning mutagenesis. J. Biol. Chem. 1994, 269, 20482–20488. [Google Scholar] [PubMed]
- Marsh, B.J.; Alm, R.A.; McIntosh, S.R.; James, D.E. Molecular regulation of GLUT-4 targeting in 3T3-L1 adipocytes. J. Cell Biol. 1995, 130, 1081–1091. [Google Scholar] [CrossRef] [PubMed]
- Higuera, J.; Calatrava, V.; González, Z.; Mariscal, V.; Siverio, J.M.; Fernández, E.; Galván, A. NRT2.4 and NRT2.5 are two half-size transporters from the Chlamydomonas NRT2 Family. Agronomy 2016, 6, 20. [Google Scholar] [CrossRef]
- Ishikawa, S.; Ito, Y.; Sato, Y.; Fukaya, Y.; Takahashi, M.; Morikawa, H.; Ohtake, N.; Ohyama, T.; Sueyoshi, K. Two-component high-affinity nitrate transport system in barley: Membrane localization; protein expression in roots and a direct protein-protein interaction. Plant Biotechnol. 2009, 26, 197–205. [Google Scholar] [CrossRef]
- Kotur, Z.; Glass, A.D.M. A 150kDa plasma membrane complex of AtNRT2.5 and AtNAR2.1 is the major contributor to constitutive high-affinity nitrate influx in Arabidopsis thaliana. Plant Cell Environ. 2015, 38, 1490–1502. [Google Scholar] [CrossRef]
- Von Wittgenstein, N.J.; Le, C.H.; Hawkins, B.J.; Ehlting, J. Evolutionary classification of ammonium; nitrate; and peptide transporters in land plants. BMC Evol. Biol. 2014, 14, 11. [Google Scholar] [CrossRef]
- Touchette, B.W. Seagrass-salinity interactions: Physiological mechanisms used by submersed marine angiosperms for a life at sea. J. Exp. Mar. Biol. Ecol. 2007, 350, 194–215. [Google Scholar] [CrossRef]
- Crawford, N.M.; Glass, A.D.M. Molecular and physiological aspects of nitrate uptake in plants. Trends Plant Sci. 1998, 3, 389–395. [Google Scholar] [CrossRef]
- Wang, Y.Y.; Cheng, Y.H.; Chen, K.E.; Tsay, Y.F. Nitrate transport, signaling, and use efficiency. Annu. Rev. Plant Biol. 2018, 69, 85–122. [Google Scholar] [CrossRef]
- Garnett, T.; Conn, V.; Plett, D.; Conn, S.; Zanghellini, J.; Mackenzie, N.; Enju, A.; Francis, K.; Holtham, L.; Roessner, U.; et al. The response of the maize nitrate transport system to nitrogen demand and supply across the lifecycle. New Phytol. 2013, 198, 82–94. [Google Scholar] [CrossRef]
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; et al. Phytozome: A comparative platform for green plant genomics. Nucleic Acids Res. 2012, 40, D1178–D1186. [Google Scholar] [CrossRef] [PubMed]
- Corpet, F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 1988, 16, 10881–10890. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef] [PubMed]
- Gouy, M.; Guindon, S.; Gascuel, O. SeaView Version 4: A multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol. 2010, 27, 221–224. [Google Scholar] [CrossRef]
- Krogh, A.; Larsson, B.; von Heijne, G.; Sonnhammer, E.L. Predicting transmembrane protein topology with a hidden markov model: Application to complete genomes. J. Mol. Biol. 2001, 305, 567–580. [Google Scholar] [CrossRef] [PubMed]
- De Castro, E.; Sigrist, C.J.; Gattiker, A.; Bulliard, V.; Langendijk-Genevaux, P.S.; Gasteiger, E.; Bairoch, A.; Hulo, N. ScanProsite: Detection of PROSITE signature matches and ProRule-associated functional and structural residues in proteins. Nucleic Acids Res. 2006, 34, W362–W365. [Google Scholar] [CrossRef] [PubMed]
- Boden, M.; Hawkins, J. Prediction of subcellular localization using sequence-biased recurrent networks. Bioinformatics 2005, 21, 2279–2286. [Google Scholar] [CrossRef]
- Horton, P.; Park, K.J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef] [PubMed]
- Gambino, G.; Perrone, I.; Gribaudo, I. A Rapid and effective method for RNA extraction from different tissues of grapevine and other woody plants. Phytochem. Anal. 2008, 19, 520–525. [Google Scholar] [CrossRef] [PubMed]
- Voinnet, O.; Rivas, S.; Mestre, P.; Baulcombe, D. Retracted: An enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J. 2003, 33, 949–956. [Google Scholar] [CrossRef] [PubMed]
- Lang-Pauluzzi, I.; Gunning, B.E.S. A plasmolytic cycle: The fate of cytoskeletal elements. Protoplasma 2000, 212, 174–185. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rubio, L.; Díaz-García, J.; Amorim-Silva, V.; Macho, A.P.; Botella, M.A.; Fernández, J.A. Molecular Characterization of ZosmaNRT2, the Putative Sodium Dependent High-Affinity Nitrate Transporter of Zostera marina L. Int. J. Mol. Sci. 2019, 20, 3650. https://doi.org/10.3390/ijms20153650
Rubio L, Díaz-García J, Amorim-Silva V, Macho AP, Botella MA, Fernández JA. Molecular Characterization of ZosmaNRT2, the Putative Sodium Dependent High-Affinity Nitrate Transporter of Zostera marina L. International Journal of Molecular Sciences. 2019; 20(15):3650. https://doi.org/10.3390/ijms20153650
Chicago/Turabian StyleRubio, Lourdes, Jordi Díaz-García, Vítor Amorim-Silva, Alberto P. Macho, Miguel A. Botella, and José A. Fernández. 2019. "Molecular Characterization of ZosmaNRT2, the Putative Sodium Dependent High-Affinity Nitrate Transporter of Zostera marina L." International Journal of Molecular Sciences 20, no. 15: 3650. https://doi.org/10.3390/ijms20153650
APA StyleRubio, L., Díaz-García, J., Amorim-Silva, V., Macho, A. P., Botella, M. A., & Fernández, J. A. (2019). Molecular Characterization of ZosmaNRT2, the Putative Sodium Dependent High-Affinity Nitrate Transporter of Zostera marina L. International Journal of Molecular Sciences, 20(15), 3650. https://doi.org/10.3390/ijms20153650





