Bifidobacterium and Lactobacillus Composition at Species Level and Gut Microbiota Diversity in Infants before 6 Weeks
Abstract
1. Introduction
2. Results
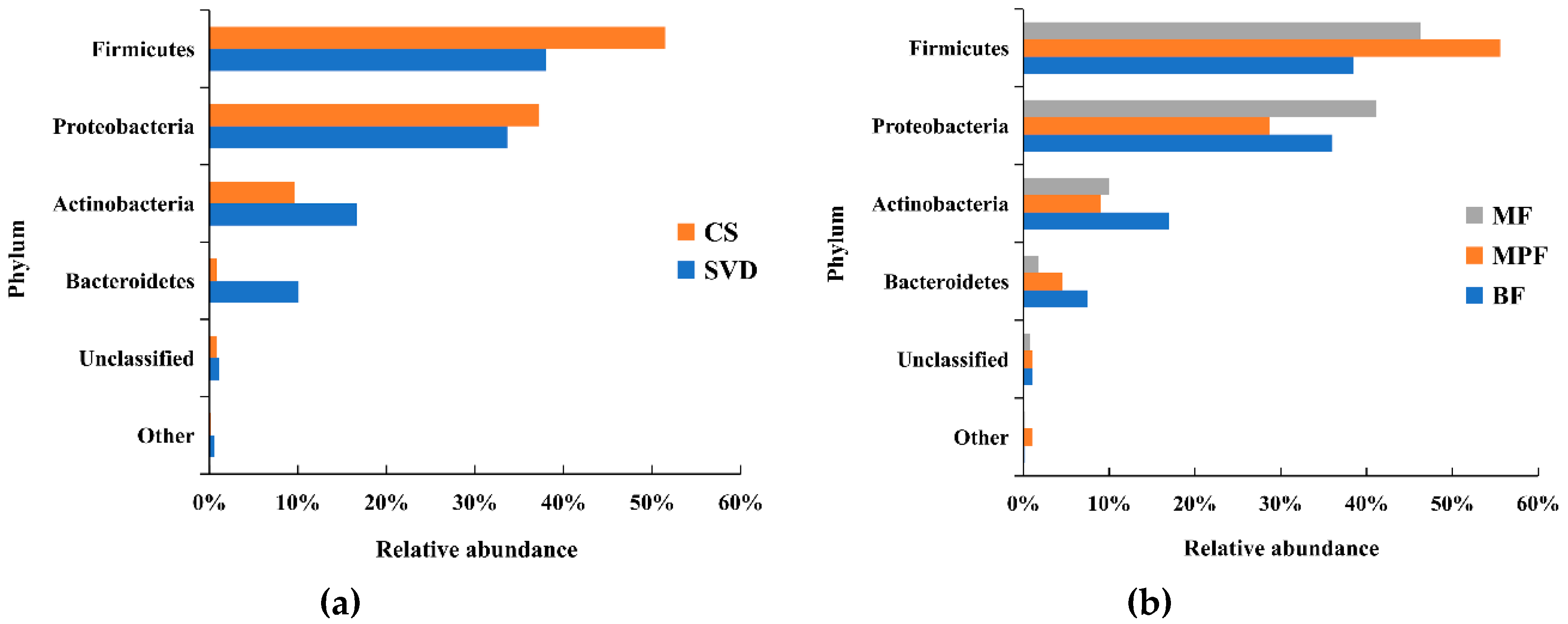
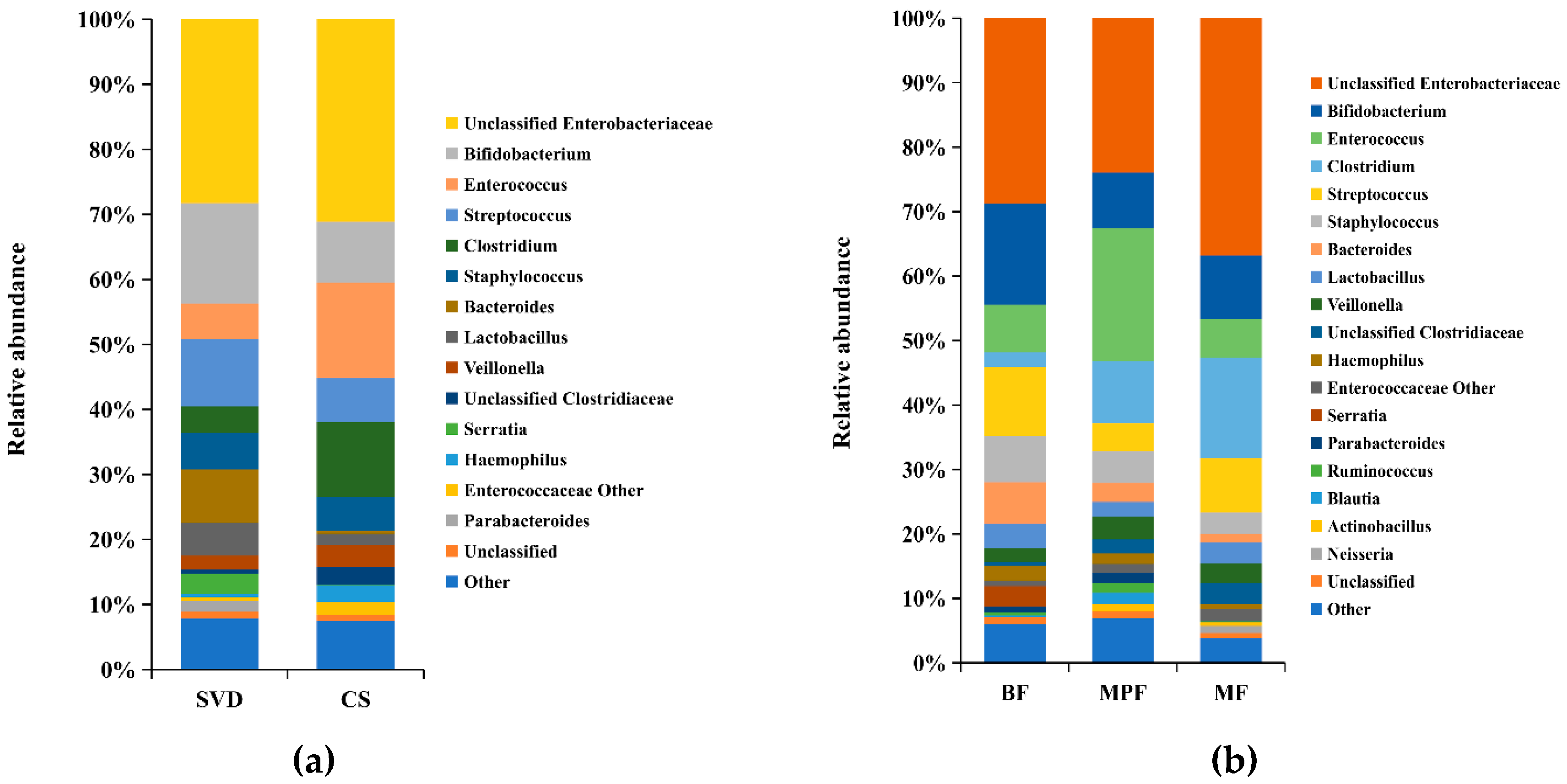
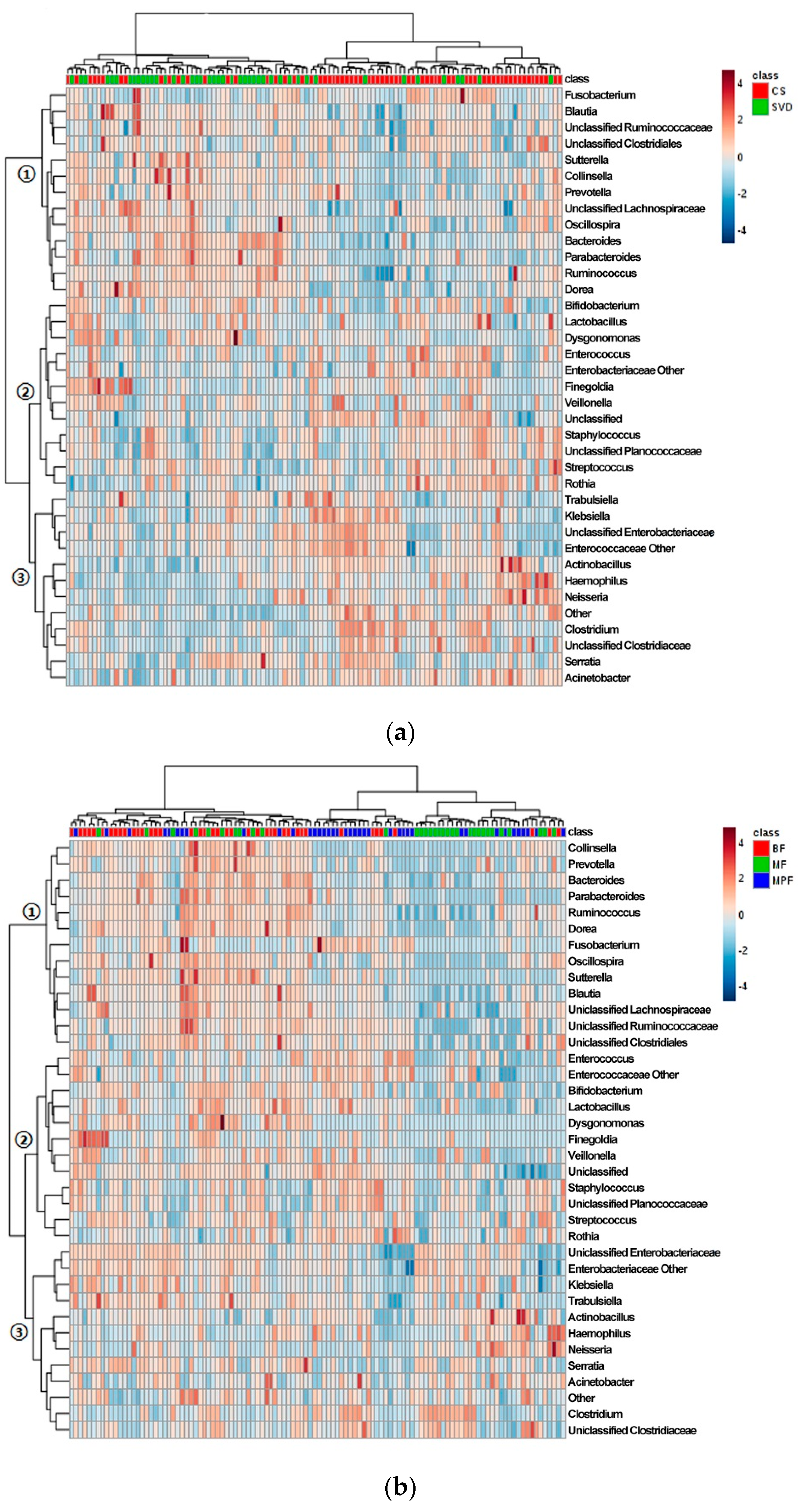
2.1. Gut Microbiota Composition in Infants
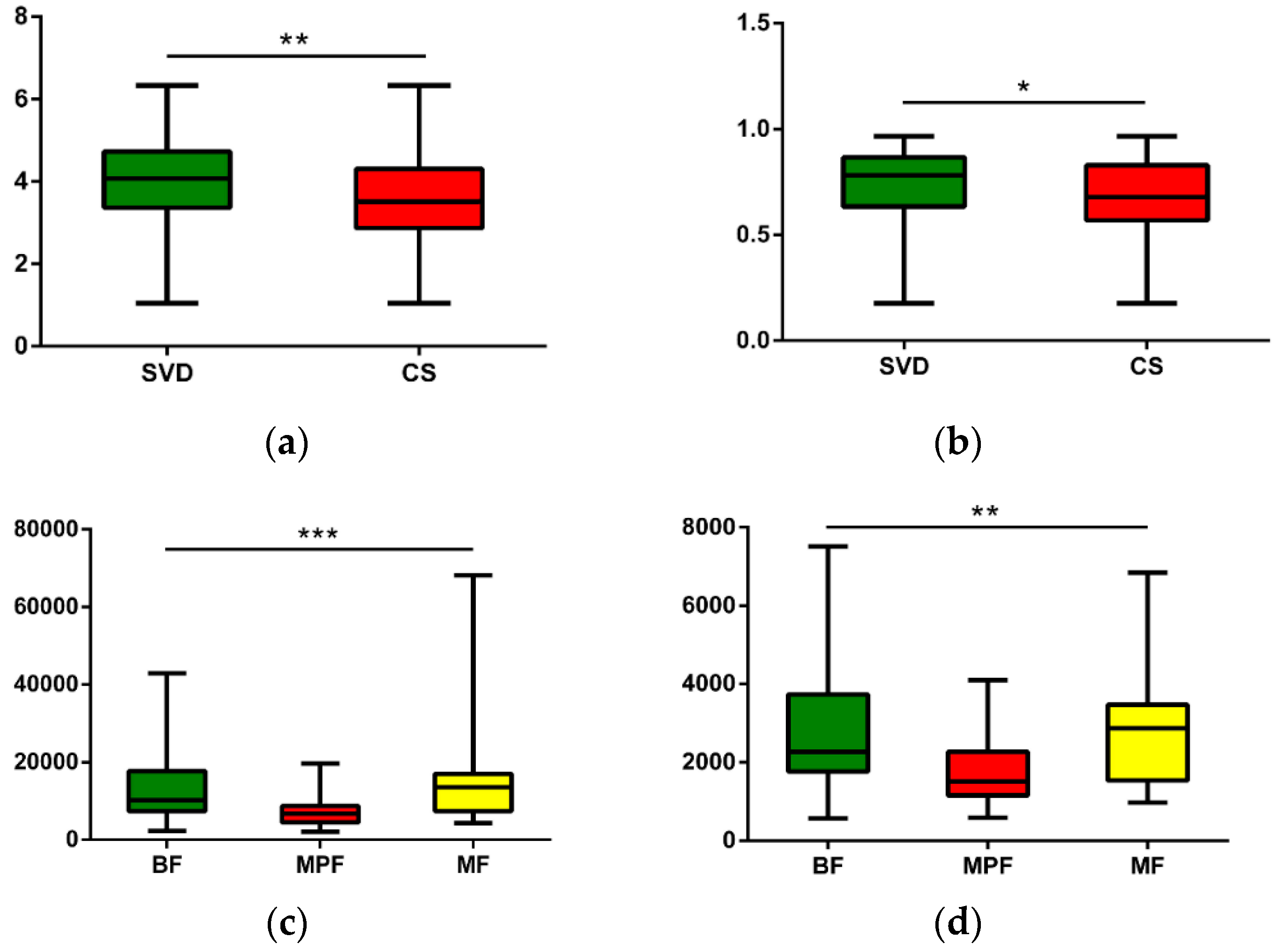
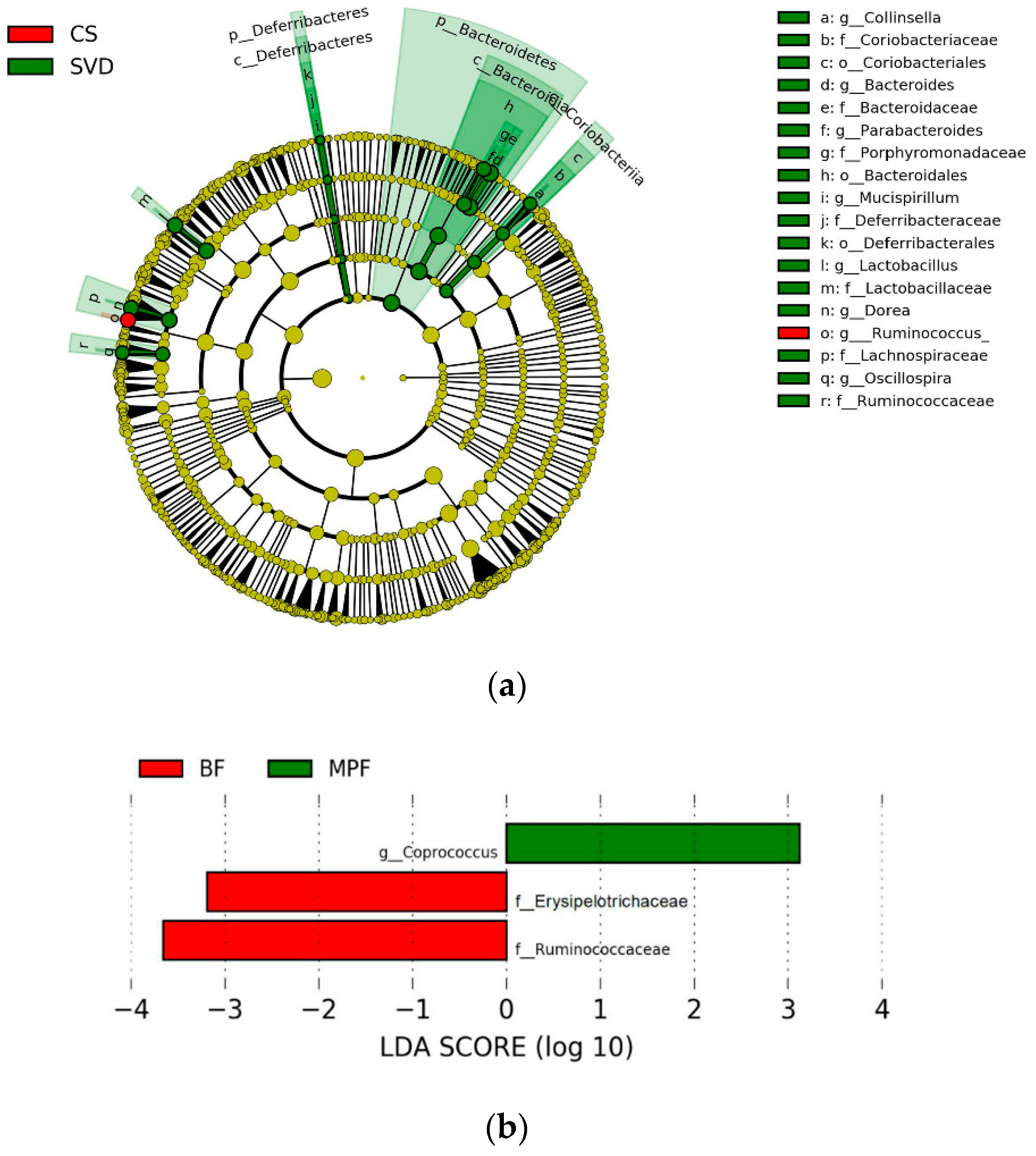
2.2. Gut Microbiota Diversity in Early Infants
2.3. Diversity of Bifidobacterium in Early Infant Gut
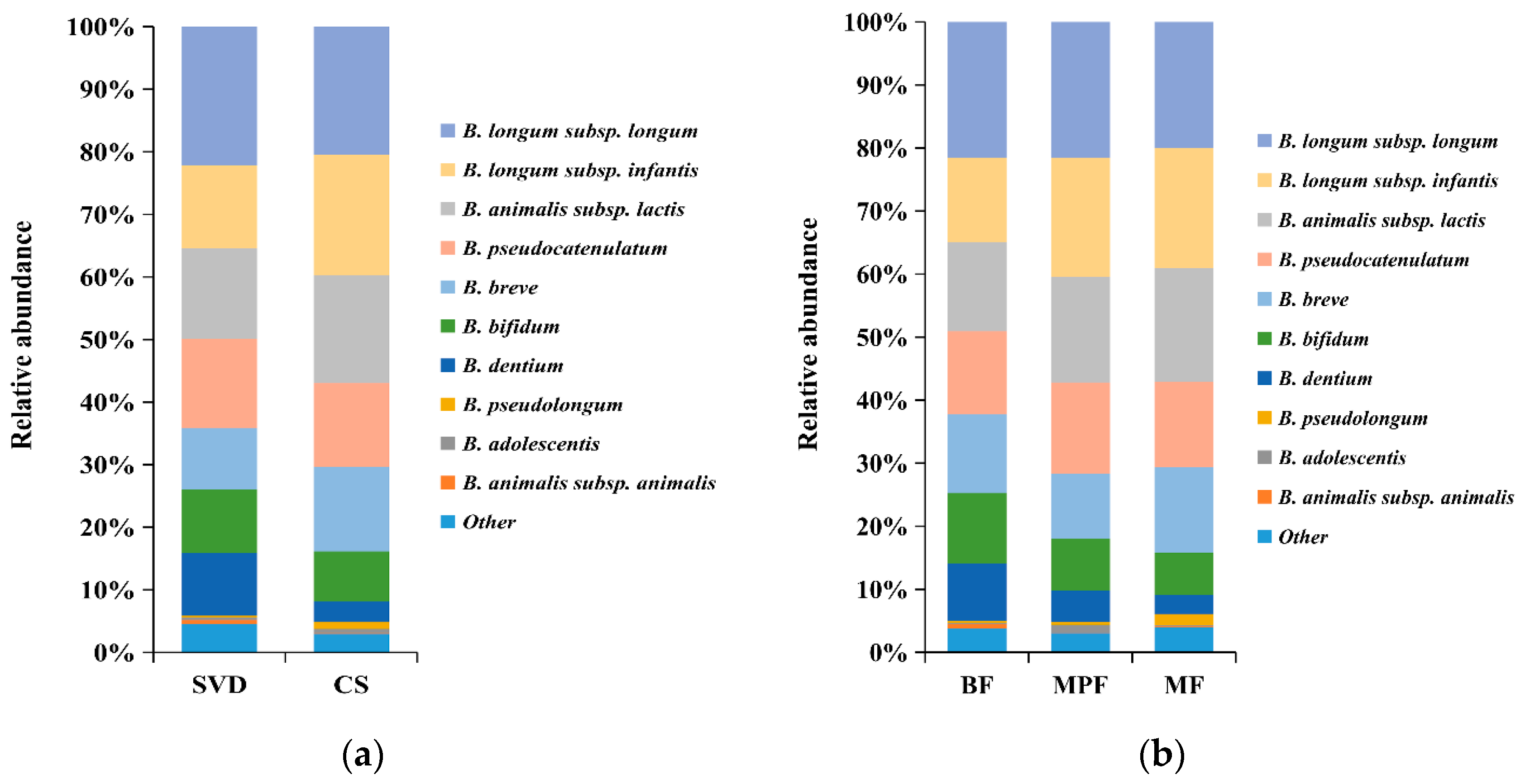
2.3.1. Bifidobacterial Composition in Infants
2.3.2. Analysis of Bifidobacterial Diversity and Related Factors in Early Infants
2.4. Diversity of Lactobacillus in Early Infant Gut
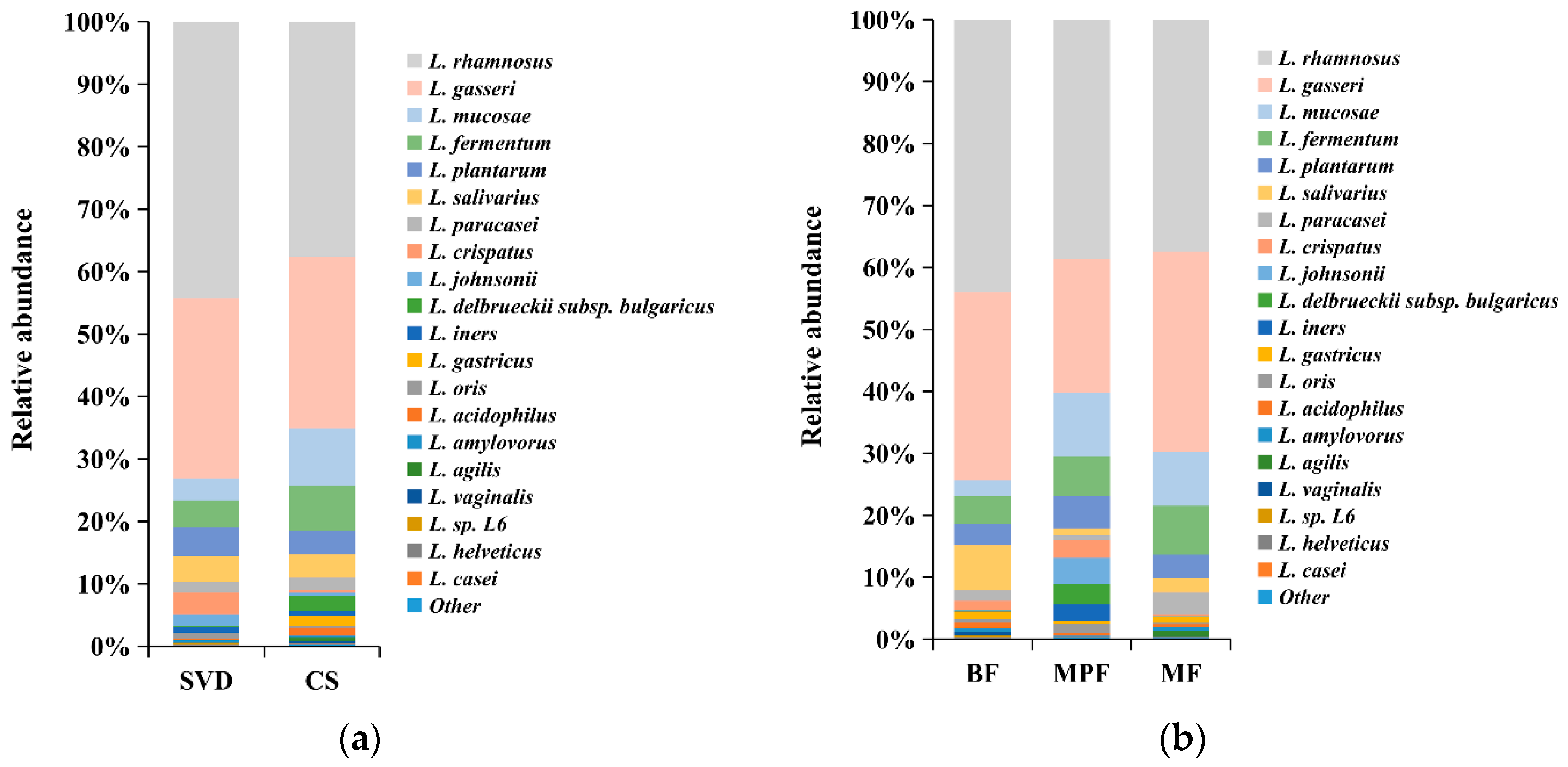
2.4.1. Lactobacillus Composition in Infants
2.4.2. Lactobacillus Diversity and Related Factors Analysis in Early Infants
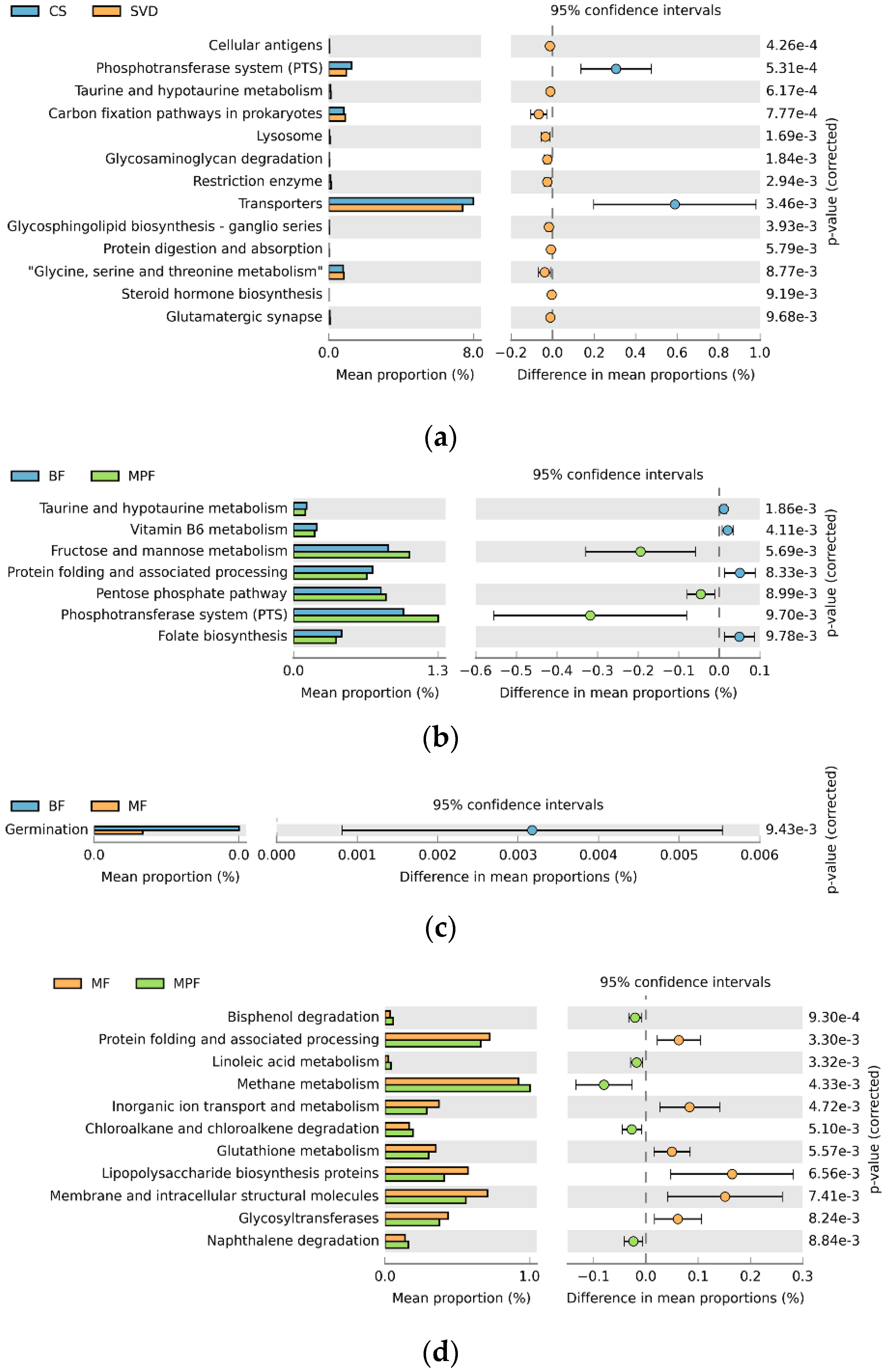
2.5. Functional Gene Composition of Gut Microbiota
3. Discussion
4. Materials and Methods
4.1. Participants and Sample Collection
4.2. Fecal Genomic DNA Extraction
4.3. DNA Amplification
4.4. Library Preparation and Sequencing
4.5. Bioinformatics Analysis
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| BF | Breast-Fed |
| Bif-GroEL | Bifidobacterial groEL Gene |
| CS | Caesarean Section |
| HMOs | Human milk oligosaccharides |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| Lac-GroEL | Lactobacillus groEL Gene |
| LDA | Linear Discriminant Analysis |
| LEfSe | Linear discriminant analysis effect size |
| MF | Mixed-Fed |
| MPF | Milk-Powder-Fed |
| OTU | Operational Taxonomic Unit |
| PICURSt | Phylogenetic Investigation of Communities by Reconstruction of Unobserved States |
| PCA | Principal component analysis |
| PCoA | Principal coordinate analysis |
| PTS | Phosphotransferase System |
| PyNAST | Python Nearest Alignment Space Termination |
| QIIME | Quantitative Insights into Microbial Ecology |
| RDP | Ribosomal Database Project |
| SVD | Standard Vaginally Delivered |
References
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Matamoros, S.; Gras-Leguen, C.; Le Vacon, F.; Potel, G.; de La Cochetiere, M.F. Development of intestinal microbiota in infants and its impact on health. Trends Microbiol. 2013, 21, 167–173. [Google Scholar] [CrossRef] [PubMed]
- Stearns, J.C.; Zulyniak, M.A.; de Souza, R.J.; Campbell, N.C.; Fontes, M.; Shaikh, M.; Sears, M.R.; Becker, A.B.; Mandhane, P.J.; Subbarao, P. Ethnic and diet-related differences in the healthy infant microbiome. Genome Med. 2017, 9, 32. [Google Scholar] [CrossRef] [PubMed]
- Lagier, J.C.; Hugon, P.; Khelaifia, S.; Fournier, P.E.; La Scola, B.; Raoult, D. The rebirth of culture in microbiology through the example of culturomics to study human gut microbiota. Clin. Microbiol. Rev. 2015, 28, 237–264. [Google Scholar] [CrossRef] [PubMed]
- Tamburini, S.; Shen, N.; Wu, H.C.; Clemente, J.C. The microbiome in early life: Implications for health outcomes. Nat. Med. 2016, 22, 713–722. [Google Scholar] [CrossRef] [PubMed]
- Arboleya, S.; Sánchez, B.; Milani, C.; Duranti, S.; Solís, G.; Fernández, N.; de los Reyes-Gavilán, C.G.; Ventura, M.; Margolles, A.; Gueimonde, M. Intestinal microbiota development in preterm neonates and effect of perinatal antibiotics. J. Pediatr. 2015, 166, 538–544. [Google Scholar] [CrossRef] [PubMed]
- Azad, M.B.; Konya, T.; Persaud, R.R.; Guttman, D.S.; Chari, R.S.; Field, C.J.; Sears, M.R.; Mandhane, P.J.; Turvey, S.E.; Subbarao, P.; et al. Impact of maternal intrapartum antibiotics, method of birth and breastfeeding on gut microbiota during the first year of life: A prospective cohort study. BJOG 2016, 123, 983–993. [Google Scholar] [CrossRef]
- Madan, J.C.; Hoen, A.G.; Lundgren, S.N.; Farzan, S.F.; Cottingham, K.L.; Morrison, H.G.; Sogin, M.L.; Li, H.; Moore, J.H.; Karagas, M.R. Association of cesarean delivery and formula supplementation with the intestinal microbiome of 6-week-old infants. JAMA Pediatr. 2016, 170, 212–219. [Google Scholar] [CrossRef] [PubMed]
- Rutayisire, E.; Huang, K.; Liu, Y.; Tao, F. The mode of delivery affects the diversity and colonization pattern of the gut microbiota during the first year of infants' life: A systematic review. BMC Gastroenterol. 2016, 16, 86. [Google Scholar] [CrossRef] [PubMed]
- Indrio, F.; Mauro, A.D.; Riezzo, G.; Mauro, F.D.; Francavilla, R. Microbiota in healthy term infant. Early Human Develop. 2013, 89, S15–S17. [Google Scholar] [CrossRef]
- Azad, M.B.; Konya, T.; Maughan, H.; Guttman, D.S.; Field, C.J.; Chari, R.S.; Sears, M.R.; Becker, A.B.; Scott, J.A.; Kozyrskyj, A.L. Gut microbiota of healthy Canadian infants: Profiles by mode of delivery and infant diet at 4 months. Canad. Med. Assoc. J. 2013, 185, 385–394. [Google Scholar] [CrossRef] [PubMed]
- Oozeer, R.; Van, L.K.; Ludwig, T.; Ben, A.K.; Martin, R.; Wind, R.D.; Boehm, G.; Knol, J. Intestinal microbiology in early life: Specific prebiotics can have similar functionalities as human-milk oligosaccharides. Am. J. Clin. Nutr. 2013, 98, 561S–571S. [Google Scholar] [CrossRef] [PubMed]
- Bergström, A.; Skov, T.H.; Bahl, M.I.; Roager, H.M.; Christensen, L.B.; Ejlerskov, K.T.; Mølgaard, C.; Michaelsen, K.F.; Licht, T.R. Establishment of intestinal microbiota during early life: A longitudinal, explorative study of a large cohort of Danish infants. Appl. Environ. Microbiol. 2014, 80, 2889–2900. [Google Scholar] [CrossRef] [PubMed]
- Fernández, L.; Langa, S.; Martín, V.; Jiménez, E.; Martín, R.; Rodríguez, J.M. The microbiota of human milk in healthy women. Cell. Mol. Biol. 2013, 59, 31–42. [Google Scholar]
- Martin, R.; Makino, H.; Cetinyurek Yavuz, A.; Ben-Amor, K.; Roelofs, M.; Ishikawa, E.; Kubota, H.; Swinkels, S.; Sakai, T.; Oishi, K.; et al. Early-life events, including mode of delivery and type of feeding, siblings and gender, shape the developing gut microbiota. PLoS ONE 2016, 11, e0158498. [Google Scholar] [CrossRef]
- Hill, C.J.; Lynch, D.B.; Murphy, K.; Ulaszewska, M.; Jeffery, I.B.; O’Shea, C.A.; Watkins, C.; Dempsey, E.; Mattivi, F.; Tuohy, K. Evolution of gut microbiota composition from birth to 24 weeks in the Infantmet cohort. Microbiome 2017, 5, 21. [Google Scholar] [CrossRef] [PubMed]
- Dogra, S.; Sakwinska, O.; Soh, S.E.; Ngombru, C.; Brück, W.M.; Berger, B.; Brüssow, H.; Lee, Y.S.; Yap, F.; Chong, Y.S. Dynamics of infant gut microbiota are influenced by delivery mode and gestational duration and are associated with subsequent adiposity. mBio 2015, 6, 312–314. [Google Scholar] [CrossRef]
- Valles, Y.; Gosalbes, M.J.; Vries, L.E.D.; Abelián, J.J.; Francino, M.P. Metagenomics and development of the gut microbiota in infants. Clin. Microbiol. Infect. 2012, 18, 21–26. [Google Scholar] [CrossRef]
- Miniello, V.L.; Colasanto, A.; Diaferio, L.; Ficele, L.; Leggi, M.S.; Santoiemma, V. Mode of delivery and gut microbiota. Ital. J. Pediatr. 2014, 40, A17. [Google Scholar] [CrossRef]
- Nagpal, R.; Kurakawa, T.; Tsuji, H.; Takahashi, T.; Kawashima, K.; Nagata, S.; Nomoto, K.; Yamashiro, Y. Evolution of gut Bifidobacterium population in healthy Japanese infants over the first three years of life: A quantitative assessment. Sci. Rep. 2017, 7, 10097. [Google Scholar] [CrossRef]
- Sirilun, S.; Takahashi, H.; Boonyaritichaikij, S.; Chaiyasut, C.; Lertruangpanya, P.; Koga, Y.; Mikami, K. Impact of maternal bifidobacteria and the mode of delivery on Bifidobacterium microbiota in infants. Benef. Microbes 2015, 6, 767–774. [Google Scholar] [CrossRef]
- Mikami, K.; Kimura, M.; Takahashi, H. Influence of maternal bifidobacteria on the development of gut bifidobacteria in infants. Pharmaceuticals 2012, 5, 629–642. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Li, M.; Wu, S.; Lebrilla, C.B.; Chapkin, R.S.; Ivanov, I.; Donovan, S.M. Fecal microbiota composition of breast-fed infants is correlated with human milk oligosaccharides consumed. J. Pediatr. Gastroenterol. Nutr. 2015, 60, 825–833. [Google Scholar] [CrossRef]
- Zivkovic, A.M.; German, J.B.; Lebrilla, C.B.; Mills, D.A. Human milk glycobiome and its impact on the infant gastrointestinal microbiota. Proc. Natl. Acad. Sci. USA 2011, 108, 4653–4658. [Google Scholar] [CrossRef] [PubMed]
- Ashida, H.; Miyake, A.; Kiyohara, M.; Wada, J.; Yoshida, E.; Kumagai, H.; Katayama, T.; Yamamoto, K. Two distinct alpha-L-fucosidases from Bifidobacterium bifidum are essential for the utilization of fucosylated milk oligosaccharides and glycoconjugates. Glycobiology 2009, 19, 1010–1017. [Google Scholar] [CrossRef] [PubMed]
- Martín, R.; Heilig, G.H.J.; Zoetendal, E.G.; Smidt, H.; Rodríguez, J.M. Diversity of the Lactobacillus group in breast milk and vagina of healthy women and potential role in the colonization of the infant gut. J. Appl. Microbiol. 2007, 103, 2638–2644. [Google Scholar] [CrossRef] [PubMed]
- Soto, A.; Martín, V.; Jiménez, E.; Mader, I.; Rodríguez, J.M.; Fernández, L. Lactobacilli and bifidobacteria in human breast milk: Influence of antibiotherapy and other host and clinical factors. J. Pediatr. Gastroenterol. Nutr. 2014, 59, 78–88. [Google Scholar] [CrossRef]
- Vestman, N.R.; Timby, N.; Holgerson, P.L.; Kressirer, C.A.; Claesson, R.; Domellöf, M.; Öhman, C.; Tanner, A.C.; Hernell, O.; Johansson, I. Characterization and in vitro properties of oral lactobacilli in breastfed infants. BMC Microbiol. 2013, 13, 193. [Google Scholar] [CrossRef]
- Salminen, S.; Endo, A.; Isolauri, E.; Scalabrin, D. Early gut colonization with lactobacilli and Staphylococcus in infants: The hygiene hypothesis extended. J. Pediatr. Gastroenterol. Nutr. 2016, 62, 80–86. [Google Scholar] [CrossRef]
- Andriantsoanirina, V.; Allano, S.; Butel, M.J.; Aires, J. Tolerance of Bifidobacterium human isolates to bile, acid and oxygen. Anaerobe 2013, 21, 39–42. [Google Scholar] [CrossRef]
- Zhao, J.; Cheung, P.C. Comparative proteome analysis of Bifidobacterium longum subsp. infantis grown on β-glucans from different sources and a model for their utilization. J. Agric. Food Chem. 2013, 61, 4360–4370. [Google Scholar]
- Valeriano, V.; Oh, J.K.; Bagon, B.B.; Kim, H.; Kang, D.K. Comparative genomic analysis of Lactobacillus mucosae LM1 identifies potential niche-specific genes and pathways for gastrointestinal adaptation. Genomics 2019, 111, 24–33. [Google Scholar] [CrossRef] [PubMed]
- Nagasaki, H.; Ito, K.; Matsuzaki, S.; Tanaka, S. Existence of phosphoenolpyruvate: Carbohydrate phosphotransferase systems in Lactobacillus fermentum, an obligate heterofermenter. Microbiol. Immunol. 2013, 36, 533–538. [Google Scholar] [CrossRef]
- Lönnerdal, B. Infant formula and infant nutrition: Bioactive proteins of human milk and implications for composition of infant formulas. Am. J. Clin. Nutr. 2014, 99, 712S–717S. [Google Scholar] [CrossRef] [PubMed]
- He, Y.B.; Ren, H.W.; Cao, Y.T.; Li, H.J.; Zhang, Z.; Liu, N. Comparing the composition and trend of fatty acid in human milk with bovine milk and infant formula in northeast region of China. CyTA-J. Food 2016, 14, 632–638. [Google Scholar] [CrossRef]
- Leonard, M.T.; Valladares, R.B.; Ardissone, A.; Gonzalez, C.F.; Lorca, G.L.; Triplett, E.W. Complete genome sequences of Lactobacillus johnsonii strain N6.2 and Lactobacillus reuteri strain TD1. Genome Announc. 2014, 2, e00397. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.R.; Geng, L.L.; Li, Y.H.; Qiang, W.; Diao, Q.Y.; Zhou, T.; Dai, P.L. The effects of Bt Cry1le toxin on bacterial diversity in the midgut of Apis mellifera ligustica (Hymenoptera: Apidae). Sci. Rep. 2016, 6, 24664. [Google Scholar] [CrossRef]
- Hu, L.; Lu, W.; Wang, L.; Pan, M.; Zhang, H.; Zhao, J.; Chen, W. Assessment of Bifidobacterium species using groEL gene on the basis of Illumina Miseq high-throughput sequencing. Genes 2017, 8, 336. [Google Scholar] [CrossRef]
- Mao, B.; Li, D.; Zhao, J.; Liu, X.; Gu, Z.; Chen, Y.Q.; Zhang, H.; Chen, W. Metagenomic insights into the effects of fructo-oligosaccharides (fos) on the composition of fecal microbiota in mice. J. Agric. Food Chem. 2015, 63, 856–863. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Bittinger, K.; Bushman, F.D.; Desantis, T.Z.; Andersen, G.L.; Knight, R. PyNAST: A flexible tool for aligning sequences to a template alignment. Bioinformatics 2010, 26, 266–267. [Google Scholar] [CrossRef]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree: Computing large minimum evolution trees with profiles instead of a distance matrix. Mol. Biol. Evol. 2009, 26, 1641–1650. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Giardine, B.; Riemer, C.; Hardison, R.C.; Burhans, R.; Elnitski, L.; Shah, P.; Zhang, Y.; Blankenberg, D.; Albert, I.; Taylor, J.; et al. Galaxy: A platform for interactive large-scale genome analysis. Genome Res. 2005, 15, 1451–1455. [Google Scholar] [CrossRef] [PubMed]
- Langille, M.G.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Vega Thurber, R.L.; Knight, R.; et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, B.; Chen, Y.; Stanton, C.; Ross, R.P.; Lee, Y.-K.; Zhao, J.; Zhang, H.; Chen, W. Bifidobacterium and Lactobacillus Composition at Species Level and Gut Microbiota Diversity in Infants before 6 Weeks. Int. J. Mol. Sci. 2019, 20, 3306. https://doi.org/10.3390/ijms20133306
Yang B, Chen Y, Stanton C, Ross RP, Lee Y-K, Zhao J, Zhang H, Chen W. Bifidobacterium and Lactobacillus Composition at Species Level and Gut Microbiota Diversity in Infants before 6 Weeks. International Journal of Molecular Sciences. 2019; 20(13):3306. https://doi.org/10.3390/ijms20133306
Chicago/Turabian StyleYang, Bo, Yingqi Chen, Catherine Stanton, R. Paul Ross, Yuan-Kun Lee, Jianxin Zhao, Hao Zhang, and Wei Chen. 2019. "Bifidobacterium and Lactobacillus Composition at Species Level and Gut Microbiota Diversity in Infants before 6 Weeks" International Journal of Molecular Sciences 20, no. 13: 3306. https://doi.org/10.3390/ijms20133306
APA StyleYang, B., Chen, Y., Stanton, C., Ross, R. P., Lee, Y.-K., Zhao, J., Zhang, H., & Chen, W. (2019). Bifidobacterium and Lactobacillus Composition at Species Level and Gut Microbiota Diversity in Infants before 6 Weeks. International Journal of Molecular Sciences, 20(13), 3306. https://doi.org/10.3390/ijms20133306




