Genome-Wide Analysis and Characterization of the Aux/IAA Family Genes Related to Floral Scent Formation in Hedychium coronarium
Abstract
1. Introduction
2. Results
2.1. Identification and Sequence Analysis of the Aux/IAA Gene Family in H. coronarium
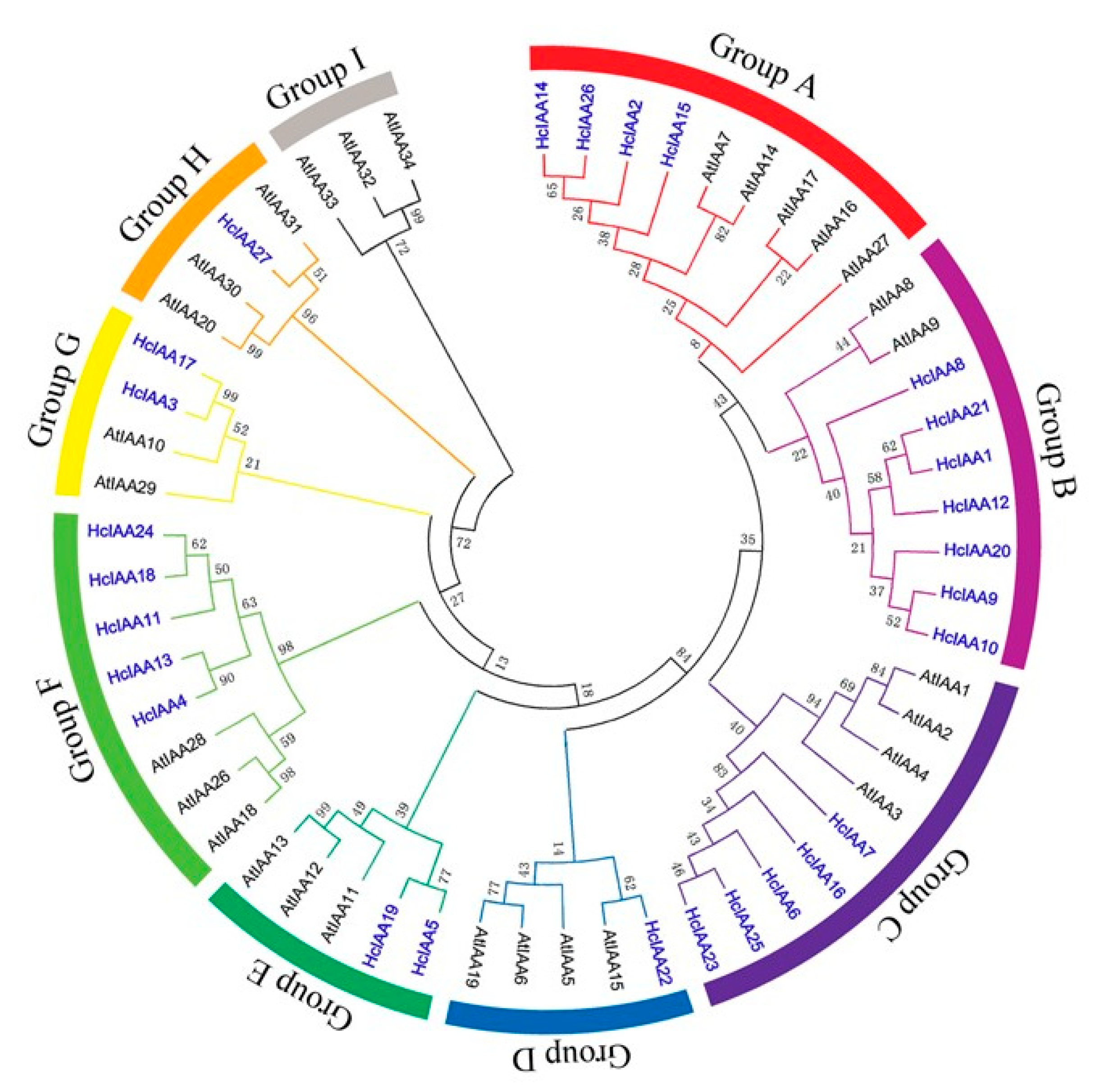
2.2. Multiple Sequence Alignment and Phylogenetic Analysis of HcIAA Genes
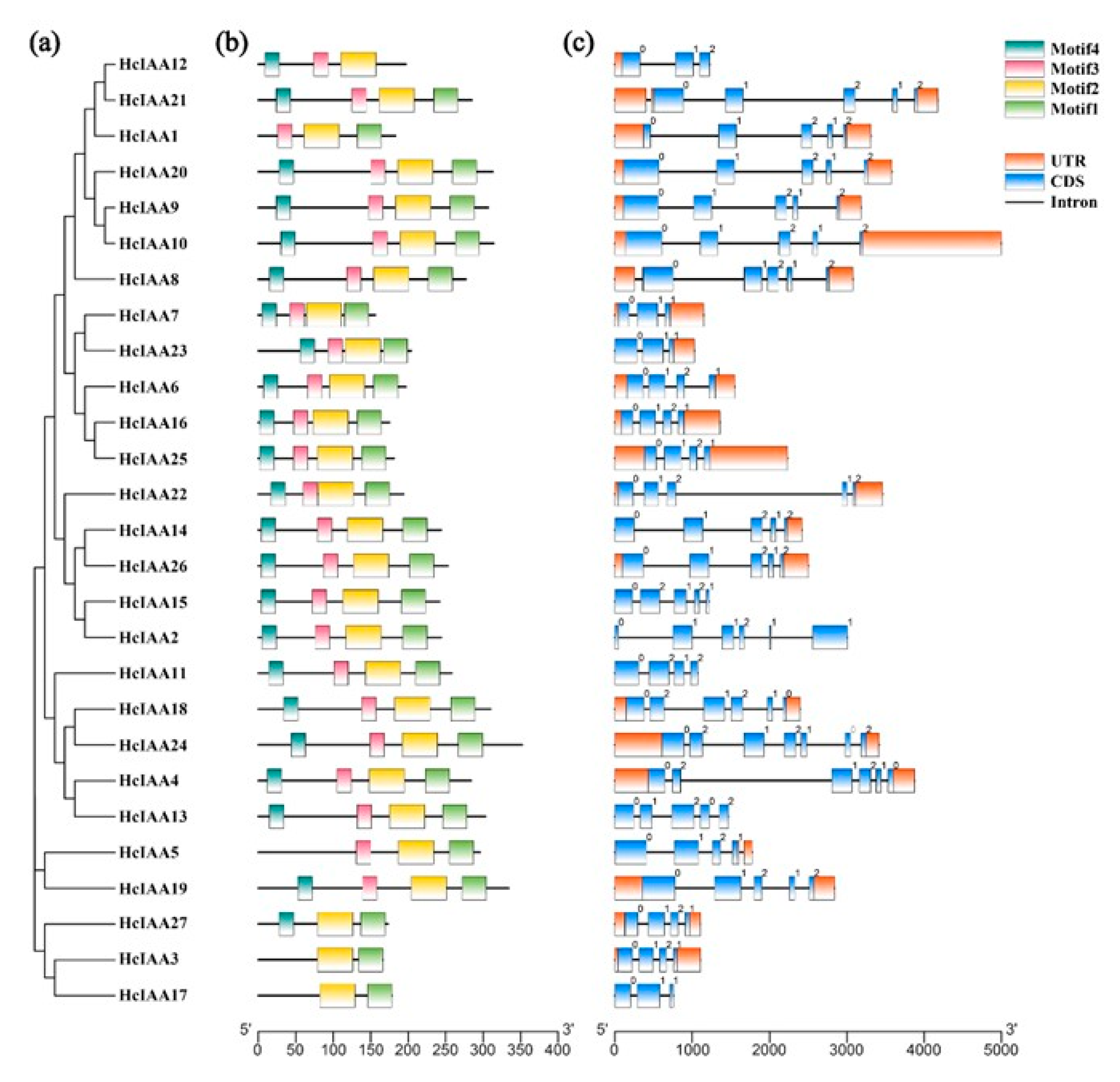
2.3. Gene Structure and Motif Composition of HcIAA Genes
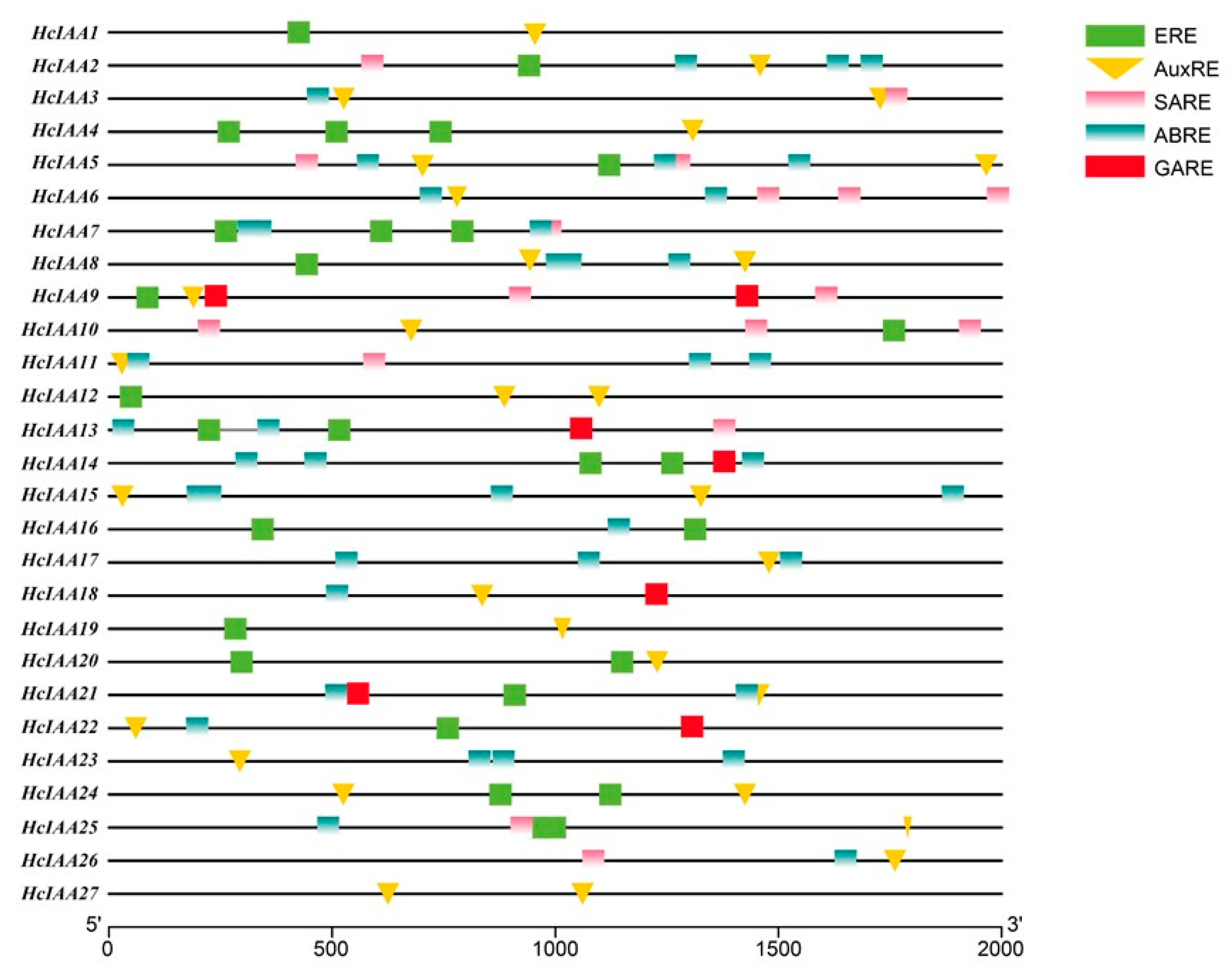
2.4. Analysis of Hormone-Related cis-Elements in the Promoter Regions of HcIAA Genes
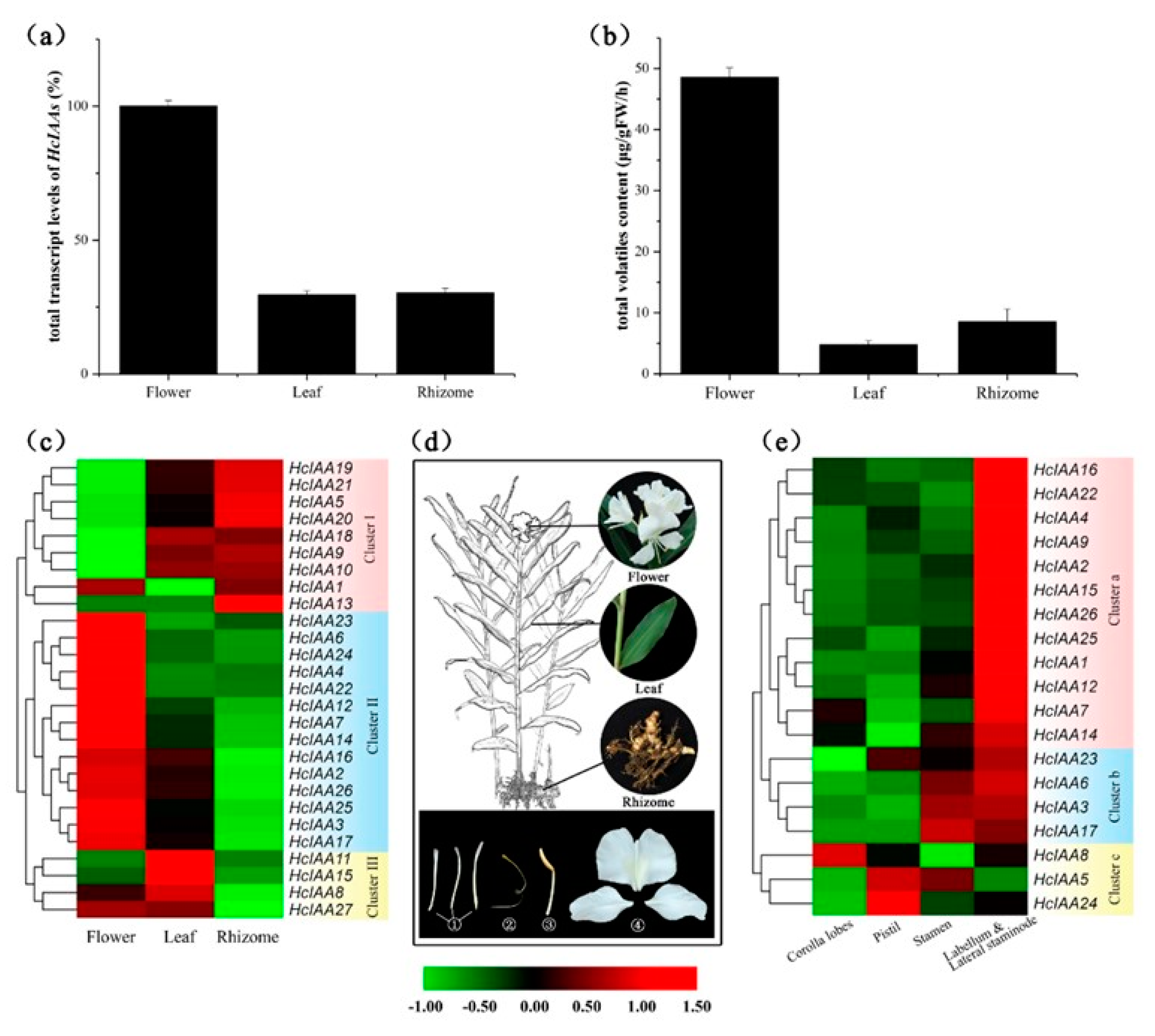
2.5. Expression Profiling of HcIAA Genes in Different Organs/Tissues
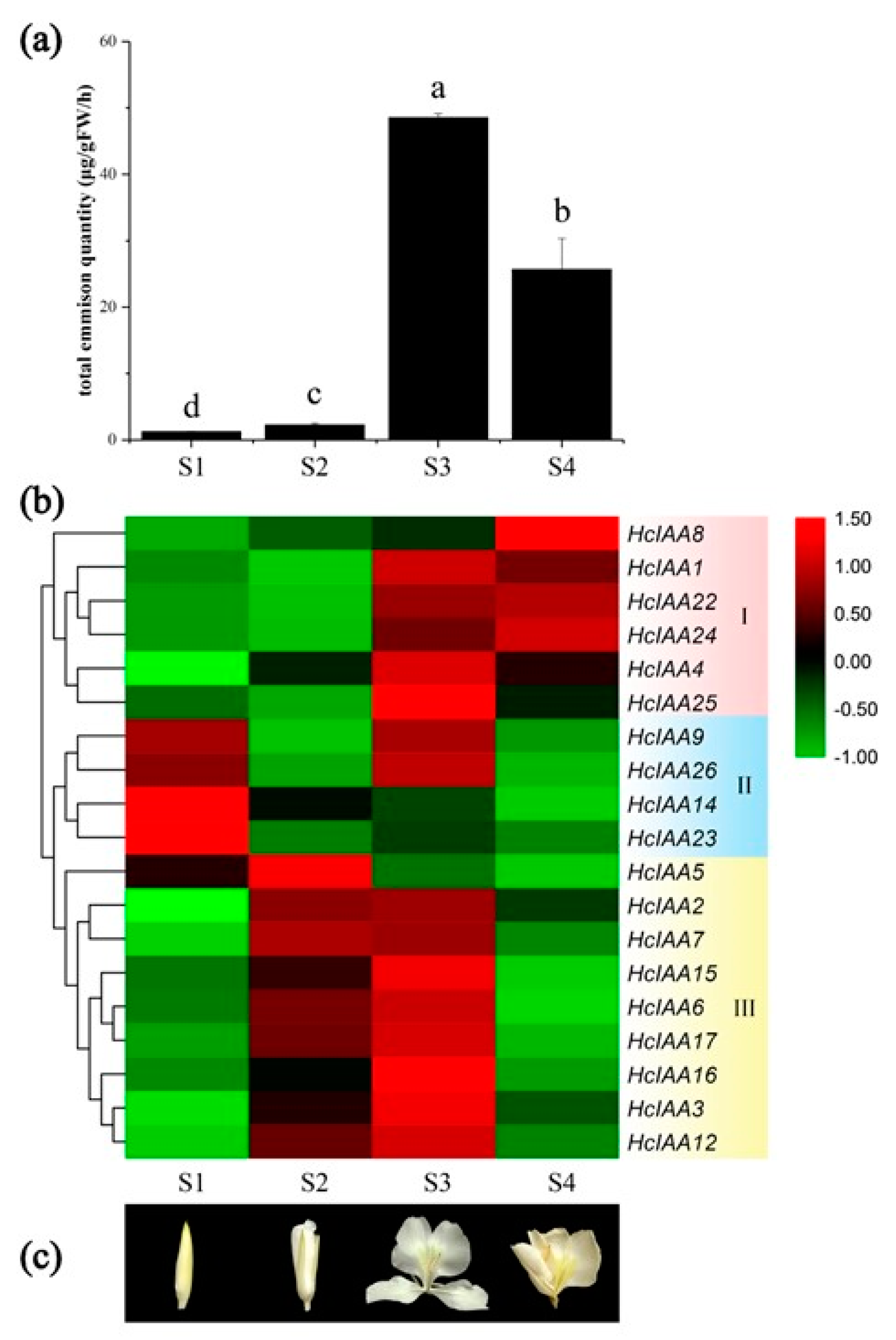
2.6. Expression of HcIAA Genes at Different Flower Developmental Stages
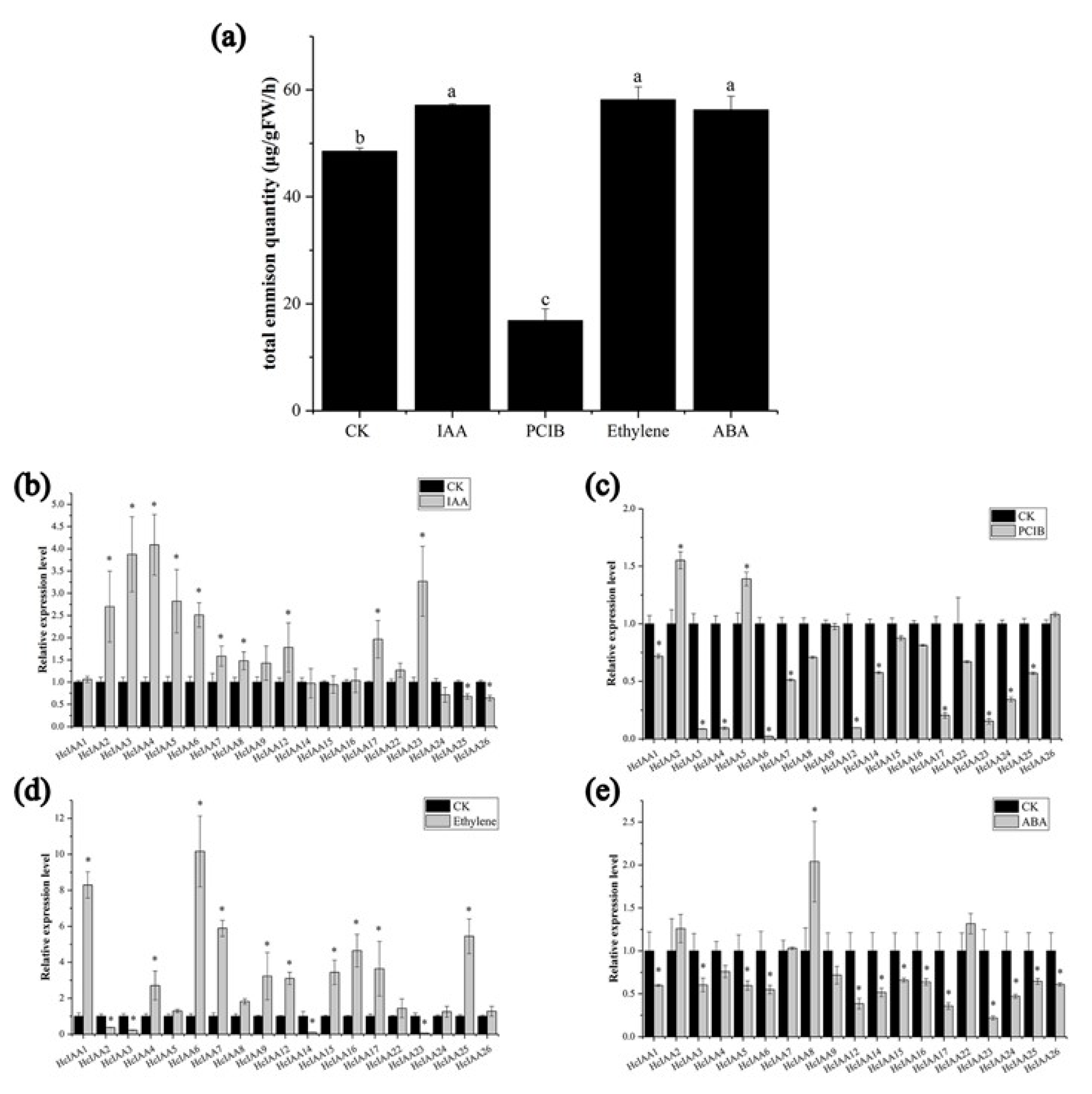
2.7. Expression of HcIAA Genes in Response to Hormone Treatments
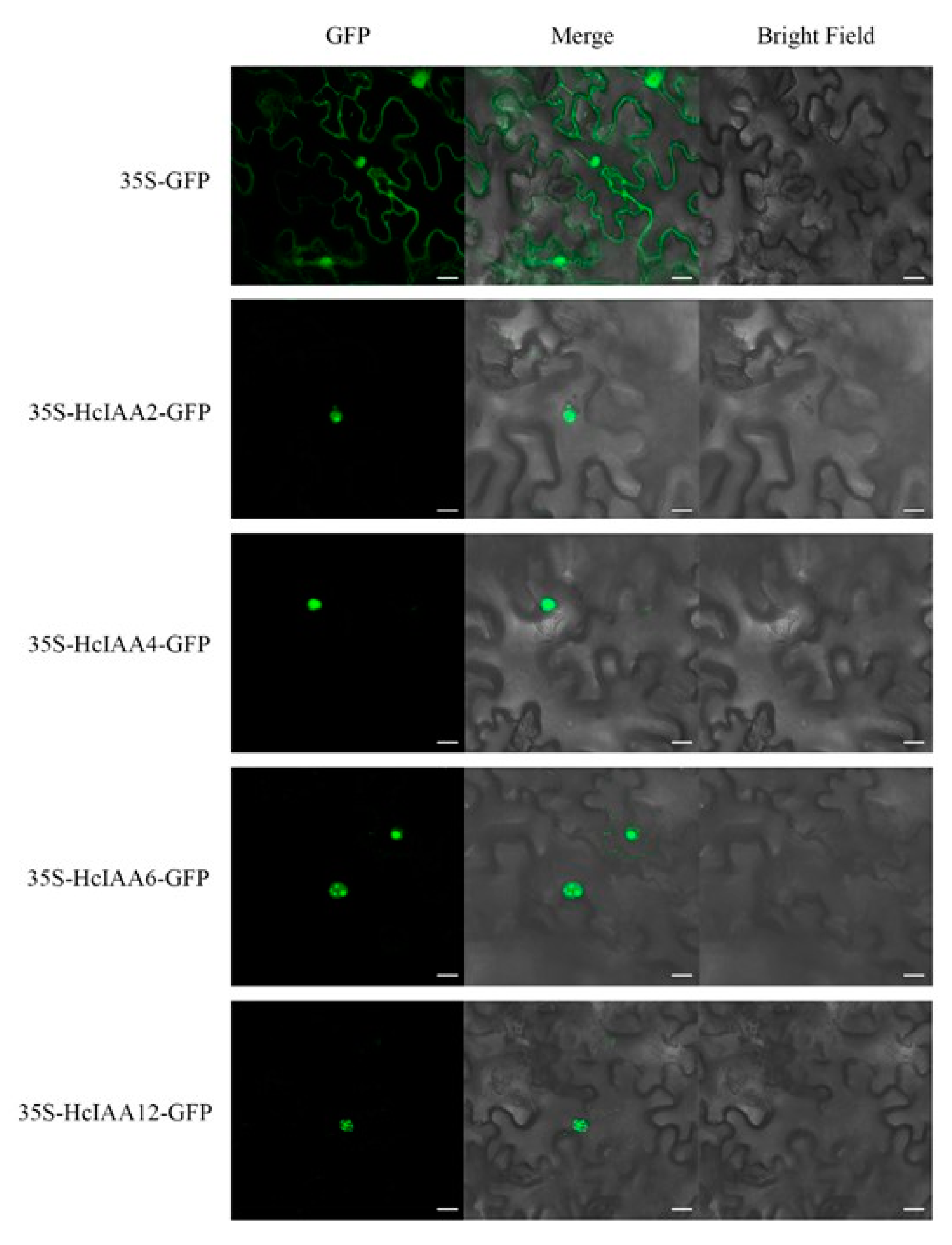
2.8. Subcellular Localization of HcIAA Candidate Genes
2.9. Silencing of HcIAA2 and HcIAA4 Altered the Flower Volatile Compound Amount
3. Discussion
4. Materials and Methods
4.1. Plant Material, Growth Conditions and Hormone Treatment
4.2. RNA Isolation, cDNA Synthesis and Quantitative Real-Time PCR (qRT-PCR)
4.3. Genome-Wide Identification of HcIAA Genes
4.4. Phylogenetic Tree Construction, Gene Structure and Motif Prediction
4.5. Analysis of Hormone-Related cis-Elements
4.6. Subcellular Localization of HcIAA Genes
4.7. Headspace Analysis of Floral Volatiles
4.8. Virus-Induced Gene Silencing (VIGS)
4.9. Statistical Analysis
4.10. Data Availability
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| Aux/IAA | Auxin/indole-3-acetic acid |
| SAUR | Small Auxin UP RNA |
| GH3 | Gretchen Hagen 3 |
| ARF | Auxin response factor |
| AuxREs | Auxin-responsive cis-elements |
| MS | Murashige and Skoog |
| ABA | Abscisic acid |
| PCIB | 2-(4-chlorophenoxy)-isobutyric acid |
| IAA | Indole-3-Acetic Acid |
| GSDS | Gene Structure Display Server |
| MEME | Multiple Expectation Maximization for Motif Elicitation |
| AuxRE | Auxin-responsive Element |
| ERE | Ethylene-responsive element |
| GARE | Gibberellin-responsive element |
| SARE | SA-responsive element |
| ABRE | ABA-responsive element |
| ORFs | Open reading frame |
| LB | Luria-Bertani |
| PDMS | Polydimethylsiloxane |
| GC-MS | Gas chromatography-mass spectrometry |
| VIGS | Virus-induced gene silencing |
| BSMV | Barley stripe mosaic virus |
| MW | Molecular weight |
| pI | Isoelectric point |
| qRT-PCR | Quantitative real-time polymerase chain reaction |
| GFP | Green fluorescence protein |
References
- Hagen, G.; Guilfoyle, T. Auxin-responsive gene expression: Genes, promoters and regulatory factors. Plant Mol. Biol. 2002, 49, 373–385. [Google Scholar] [CrossRef] [PubMed]
- Friml, J.; Vieten, A.; Sauer, M.; Weijers, D.; Scwartz, H.; Hamann, T.; Offringa, R.; Jürgens, G. Efflux-dependent auxin gradients establish the apical-basal axis of Arabidopsis. Nature 2003, 426, 147–153. [Google Scholar] [CrossRef] [PubMed]
- Jain, M.; Kaur, N.; Garg, R.; Thakur, J.K.; Tyagi, A.K.; Khurana, J.P. Structure and expression analysis of early auxin-responsive Aux/IAA gene family in rice (Oryza sativa). Funct. Integr. Genom. 2006, 6, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Theologis, A.; Huynh, T.V.; Davis, R.W. Rapid induction of specific mRNAs by auxin in pea epicotyl tissue. J. Mol. Biol. 1985, 183, 53–68. [Google Scholar] [CrossRef]
- Oeller, P.W.; Keller, J.A.; Parks, J.E.; Silbert, J.E.; Theologis, A. Structural characterization of the early indoleacetic acid-inducible genes, PS-IAA4/5 and PS-IAA6, of pea (Pisum sativum L.). J. Mol. Biol. 1993, 233, 789–798. [Google Scholar] [CrossRef] [PubMed]
- Dharmasiri, S.; Estelle, M. The role of regulated protein degradation in auxin response. Plant Mol. Biol. 2002, 49, 401–408. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, S.B.; Wang, X.J.; Hagen, G.; Guilfoyle, T.J. Aux/IAA proteins are active repressors, and their stability and activity are modulated by auxin. Plant Cell 2001, 13, 2809–2822. [Google Scholar] [CrossRef] [PubMed]
- Abel, S.; Oeller, P.W.; Theologis, A. Early auxin-induced genes encode short-lived nuclear proteins. Proc. Natl. Acad. Sci. USA 1994, 91, 326–330. [Google Scholar] [CrossRef]
- Arase, F.; Nishitani, H.; Egusa, M.; Nishimoto, N.; Sakurai, S.; Sakamoto, N.; Kaminaka, H. IAA8 involved in lateral root formation interacts with the TIR1 auxin receptor and ARF transcription factors in Arabidopsis. PLoS ONE 2012, 7, e43414. [Google Scholar] [CrossRef]
- Tiwari, S.B. Aux/IAA proteins contain a potent transcriptional repression domain. Plant Cell 2004, 16, 533–543. [Google Scholar] [CrossRef]
- Szemenyei, H.; Hannon, M.; Long, J.A. TOPLESS Mediates Auxin-Dependent Transcriptional Repression During Arabidopsis Embryogenesis. Science 2008, 319, 1384–1386. [Google Scholar] [CrossRef] [PubMed]
- Dharmasiri, N.; Dharmasiri, S.; Estelle, M. The F-box protein TIR1 is an auxin receptor. Nature 2005, 435, 441–445. [Google Scholar] [CrossRef] [PubMed]
- Kepinski, S.; Leyser, O. The Arabidopsis F-box protein TIR1 is an auxin receptor. Nature 2005, 435, 446–451. [Google Scholar] [CrossRef] [PubMed]
- Tan, X.; Calderon-Villalobos, L.I.A.; Sharon, M.; Zheng, C.; Robinson, C.V.; Estelle, M.; Zheng, N. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 2007, 446, 640–645. [Google Scholar] [CrossRef] [PubMed]
- Leyser, O. Auxin Signaling. Plant Physiol. 2018, 176, 465–479. [Google Scholar] [CrossRef] [PubMed]
- Remington, D.L. Contrasting modes of diversification in the Aux/IAA and ARF gene families. Plant Physiol. 2004, 135, 1738–1752. [Google Scholar] [CrossRef] [PubMed]
- Overvoorde, P.J. Functional genomic analysis of the Auxin/indole-3-acetic acid gene family members in Arabidopsis thaliana. Plant Cell 2005, 17, 3282–3300. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, S.B. The Roles of Auxin Response Factor Domains in Auxin-Responsive Transcription. Plant Cell. 2003, 15, 533–543. [Google Scholar] [CrossRef] [PubMed]
- Deng, W.; Yan, F.; Liu, M.; Wang, X.; Li, Z. Down-regulation of SlIAA15 in tomato altered stem xylem development and production of volatile compounds in leaf exudates. Plant Signal. Behav. 2014, 7, 911–913. [Google Scholar] [CrossRef]
- Bassa, C.; Mila, I.; Bouzayen, M.; Audran-Delalande, C. Phenotypes associated with down-regulation of Sl-IAA27 support functional diversity among AUX/IAA family members in tomato. Plant Cell Physiol. 2012, 53, 1583–1595. [Google Scholar] [CrossRef]
- Guillotin, B.; Etemadi, M.; Audran, C.; Bouzayen, M.; Bécard, G.; Combier, J. Sl-IAA27 regulates strigolactone biosynthesis and mycorrhization in tomato (var. Micro Tom). New Phytol. 2016, 213, 1124–1132. [Google Scholar] [CrossRef] [PubMed]
- Woodward, A.W.; Bartel, B. Auxin: Regulation, action, and interaction. Ann. Bot. 2005, 95, 707–735. [Google Scholar] [CrossRef] [PubMed]
- Zhu, B.Q.; Xu, X.Q.; Wu, Y.W.; Duan, C.Q.; Pan, Q.H. Isolation and characterization of two hydroperoxide lyase genes from grape berries: HPL isogenes in Vitis vinifera grapes. Mol. Biol. Rep. 2012, 39, 7443–7455. [Google Scholar] [CrossRef] [PubMed]
- Kitomi, Y.; Inahashi, H.; Takehisa, H.; Sato, Y.; Inukai, Y. OsIAA13-mediated auxin signaling is involved in lateral root initiation in rice. Plant Sci. 2012, 190, 116–122. [Google Scholar] [CrossRef] [PubMed]
- Liscum, E.; Reed, J.W. Genetics of Aux/IAA and ARF action in plant growth and development. Plant Mol. Biol. 2002, 49, 387–400. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Deng, D.; Bian, Y.; Lv, Y.; Xie, Q. Genome-wide analysis of primary auxin-responsive Aux/IAA gene family in maize (Zea mays. L.). Mol. Biol. Rep. 2010, 37, 3991–4001. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Peng, Z.; Liu, S.Y.; He, Y.J.; Cheng, L.; Kong, F.; Wang, J.; Lu, G. Genome-wide analysis of Aux/IAA gene family in Solanaceae species using tomato as a model. Mol. Genet. Genom. 2012, 287, 295–311. [Google Scholar] [CrossRef] [PubMed]
- Çakir, B.; Kiliçkaya, O.; Olcay, A. Genome-wide analysis of Aux/IAA genes in Vitis vinifera: Cloning and expression profiling of a grape Aux/IAA gene in response to phytohormone and abiotic stresses. Acta Physiol. Plant. 2013, 35, 365–377. [Google Scholar] [CrossRef]
- Yue, Y.; Yu, R.; Fan, Y. Transcriptome profiling provides new insights into the formation of floral scent in Hedychium coronarium. BMC Genom. 2015, 16, 470. [Google Scholar] [CrossRef]
- Liu, K.; Yuan, C.; Feng, S.; Zhong, S.; Li, H.; Zhong, J.; Shen, C.; Liu, J. Genome-wide analysis and characterization of Aux/IAA family genes related to fruit ripening in papaya (Carica papaya L.). BMC Genom. 2017, 18, 351. [Google Scholar] [CrossRef]
- Wu, Z.Y.; Raven, H.P. Flora of China; Missouri Botanical Garden Press: Saint Louis, MO, USA, 2000; Volume 24, pp. 370–377. [Google Scholar]
- Baez, D.; Pino, J.A.; Morales, D. Floral scent composition in Hedychium coronarium J. Koenig analyzed by SPME. J. Essent. Oil Res. 2011, 23, 64–67. [Google Scholar] [CrossRef]
- Fan, Y.P.; Yu, R.C.; Huang, Y.; Chen, Y.F. Studies on the essential constituent of Hedychium flavum and H. coronarium. Acta Horti. Sinica 2003, 30, 475. [Google Scholar]
- Fan, Y.P.; Wang, X.R.; Yu, R.C.; Yang, P. Analysis on the aroma components in several species of Hedychium. Acta Horti. Sinica 2007, 34, 231–234. [Google Scholar]
- Yue, Y.; Yu, R.; Fan, Y. Characterization of two monoterpene synthases involved in floral scent formation in Hedychium coronarium. Planta 2014, 240, 745–762. [Google Scholar] [CrossRef] [PubMed]
- Pichersky, E.; Dudareva, N. Scent engineering: Toward the goal of controlling how flowers smell. Trends Biotechnol. 2007, 25, 105–110. [Google Scholar] [CrossRef]
- Audran-Delalande, C.; Bassa, C.; Mila, I.; Regad, F.; Zouine, M.; Bouzayen, M. Genome-Wide identification, functional analysis and expression profiling of the Aux/IAA gene family in tomato. Plant Cell Physiol. 2012, 53, 659–672. [Google Scholar] [CrossRef] [PubMed]
- Wils, C.R.; Kaufmann, K. Gene-regulatory networks controlling inflorescence and flower development in Arabidopsis thaliana. Biochim. Biophys. Acta Gene Regul. Mech. 2017, 1860, 95–105. [Google Scholar] [CrossRef]
- Zubo, Y.O.; Yamburenko, M.V.; Kusnetsov, V.V.; Börner, T. Methyl jasmonate, gibberellic acid, and auxin affect transcription and transcript accumulation of chloroplast genes in barley. J. Plant Physiol. 2011, 168, 1335–1344. [Google Scholar] [CrossRef]
- Agarwal, P.K.; Agarwal, P.; Custers, J.B.M.; Liu, C.; Bhojwani, S.S. PCIB an anti-auxin enhances microspore embryogenesis in microspore culture of Brassica juncea. Plant Cell Tissue Organ Cult. 2006, 86, 201–210. [Google Scholar] [CrossRef]
- Fransson, P. Studies on the interaction of antiauxin and native auxin in wheat roots. Physiol. Plantarum 1958, 11, 644–654. [Google Scholar] [CrossRef]
- Heupel, T.; Stange, L. The auxin antagonist p-chlorophenoxyisobutyric acid abolishes polar distribution of DNA synthesizing cells within the meristem of Riella helicophylla. J. Plant Physiol. 1995, 146, 757–759. [Google Scholar] [CrossRef]
- Van-Doorn, W.G.; Dole, I.; Çelikel, F.G.; Harkema, H. Opening of Iris flowers is regulated by endogenous auxins. J. Plant Physiol. 2013, 170, 161–164. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Soler, M.; San Clemente, H.; Mila, I.; Paiva, J.A.P.; Myburg, A.A.; Bouzayen, M.; Grima-Pettenati, J.; Cassan-Wang, H. Comprehensive genome-wide analysis of the Aux/IAA gene family in Eucalyptus: Evidence for the role of EgrIAA4 in wood formation. Plant Cell Physiol. 2015, 56, 700–714. [Google Scholar] [CrossRef] [PubMed]
- Tyurin, A.A.; Kabardaeva, K.V.; Berestovoy, M.A.; Sidorchuk, Y.V.; Fomenkov, A.A.; Nosov, A.V.; Goldenkova-Pavlova, I.V. Simple and reliable system for transient gene expression for the characteristic signal sequences and the estimation of the localization of target protein in plant cell. Russ. J. Plant Physiol. 2017, 64, 672–679. [Google Scholar] [CrossRef]
- Bohn-Courseau, I. Auxin: A major regulator of organogenesis. Comptes Rendus Biologies 2010, 333, 290–296. [Google Scholar] [CrossRef] [PubMed]
- Gaudinová, A.; Malbeck, J.; Dobrev, P.; Kubelková, D.; Špak, J.; Vanková, R. Cytokinin, auxin, and abscisic acid dynamics during flower development in white and red currants infected with Black currant reversion virus. Physiol. Mol. Plant Pathol. 2008, 73, 119–125. [Google Scholar] [CrossRef]
- Luo, J.; Zhou, J.; Zhang, J. Aux/IAA Gene Family in Plants: Molecular Structure, Regulation, and Function. Int. J. Mol. Sci. 2018, 19, 259. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, U.C.; Difazio, S.P.; Brunner, A.M.; Tuskan, G.A. Genome-wide analysis of Aux/IAA and ARF gene families in (Populus trichocarpa). BMC Plant Biol. 2007, 7, 59. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Yan, D.; Yuan, T.; Gao, X.; Lu, Y. A gain-of-function mutation in IAA8 alters Arabidopsis floral organ development by change of jasmonic acid level. Plant Mol. Biol. 2013, 82, 71–83. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Xu, X.; Fang, D.D.; Zhang, T.; Guo, W. Cloning and expression analysis of novel Aux/IAA family genes in Gossypium hirsutum. Gene 2012, 503, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Singh, V.K.; Jain, M. Genome-wide survey and comprehensive expression profiling of Aux/IAA gene family in chickpea and soybean. Front. Plant Sci. 2015, 6, 918. [Google Scholar] [CrossRef]
- Reed, J.W. Roles and activities of Aux/IAA proteins in Arabidopsis. Trends Plant Sci. 2001, 6, 420–425. [Google Scholar] [CrossRef]
- Jain, M.; Kaur, N.; Tyagi, A.K.; Khurana, J.P. The auxin-responsive GH3 gene family in rice (Oryza sativa). Funct. Integr. Genom. 2006, 6, 36–46. [Google Scholar] [CrossRef] [PubMed]
- Meir, S.; Philosoph-Hadas, S.; Sundaresan, S.; Selvaraj, K.S.V.; Burd, S.; Ophir, R.; Kochanek, B.; Reid, M.S.; Jiang, C.Z.; Lers, A. Microarray analysis of the abscission-related transcriptome in the tomato flower abscission zone in response to auxin depletion. Plant Physiol. 2010, 154, 1929–1956. [Google Scholar] [CrossRef] [PubMed]
- Chaabouni, S.; Jones, B.; Delalande, C.; Wang, H.; Li, Z.; Mila, I.; Frasse, P.; Latché, A.; Pech, J.; Bouzayen, M. Sl-IAA3, a tomato Aux/IAA at the crossroads of auxin and ethylene signaling involved in differential growth. J. Exp. Bot. 2009, 60, 1349–1362. [Google Scholar] [CrossRef] [PubMed]
- Mai, Y.X.; Wang, L.; Yang, H.Q. A gain-of-function mutation in IAA7/AXR2 confers late flowering under short-day light in Arabidopsis. J. Integr. Plant Biol. 2011, 53, 480–492. [Google Scholar] [CrossRef] [PubMed]
- Abbas, F.; Ke, Y.; Yu, R.; Yue, Y.; Amanullah, S.; Jahangir, M.M.; Fan, Y. Volatile terpenoids: Multiple functions, biosynthesis, modulation and manipulation by genetic engineering. Planta 2017, 246, 803–816. [Google Scholar] [CrossRef]
- Abbas, F.; Ke, Y.; Yu, R.; Fan, Y. Functional characterization and expression analysis of two terpene synthases involved in floral scent formation in Lilium ‘Siberia’. Planta 2019, 249, 71–93. [Google Scholar] [CrossRef]
- Dudareva, N.; Negre, F.; Nagegowda, D.A.; Orlova, I. Plant volatiles: Recent advances and future perspectives. Crit. Rev. Plant Sci. 2006, 25, 417–440. [Google Scholar] [CrossRef]
- Dudareva, N.; Klempien, A.; Muhlemann, J.K.; Kaplan, I. Biosynthesis, function and metabolic engineering of plant volatile organic compounds. New Phytol. 2013, 198, 16–32. [Google Scholar] [CrossRef]
- Kwangjin, K.; Mijung, K.; Heungdeug, K.; Song, J.S.; Eunha, Y.; Junggun, C. The effect of flower scent and essential oils on reduction of concentration of cortisol, a stress hormone. Korean J. Hortic. Sci. 2006, 30, 1198–1209. [Google Scholar]
- Ramya, M.; Kwon, O.K.; An, H.R.; Park, P.M.; Baek, Y.S.; Park, P.H. Floral scent: Regulation and role of MYB transcription factors. Phytochem. Lett. 2017, 19, 114–120. [Google Scholar] [CrossRef]
- Schmelz, E.A.; Engelberth, J.; Alborn, H.T.; Donnell, P.; Sammons, M.; Toshima, H.; Tumlinson, J.H. Simultaneous analysis of phytohormones, phytotoxins, and volatile organic compounds in plants. Proc. Natl. Acad. Sci. USA 2003, 100, 10552–10557. [Google Scholar] [CrossRef] [PubMed]
- Livakm, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCt method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2013, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Higo, K.; Ugawa, I.M.; Korenaga, T. Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res. 1999, 27, 297–300. [Google Scholar] [CrossRef] [PubMed]
- Sainsbury, F.; Thuenemann, E.C.; Lomonossoff, G.P. pEAQ: Versatile expression vectors for easy and quick transient expression of heterologous proteins in plants. Plant Biotechnol. J. 2009, 7, 682–693. [Google Scholar] [CrossRef] [PubMed]
- Mahadevan, C.; Jaleel, A.; Deb, L.; Thomas, G.; Sakuntala, M. Development of an efficient virus induced gene silencing strategy in the non-model wild ginger Zingiber zerumbet and investigation of associated proteome changes. PLoS ONE 2014, 10, e124518. [Google Scholar] [CrossRef]
- Renner, T.; Bragg, J.; Driscoll, H.E.; Cho, J.; Jackson, A.O.; Specht, C.D. Virus-Induced Gene Silencing in the Culinary Ginger (Zingiber officinale): An effective mechanism for down-regulating gene expression in tropical monocots. Mol. Plant 2009, 2, 1084–1094. [Google Scholar] [CrossRef]
- Yuan, C.; Li, C.; Yan, L.; Jackson, A.O.; Liu, Z.; Han, C.; Yu, J.; Li, D. A high throughput barley stripe mosaic virus vector for virus induced gene silencing in monocots and dicots. PLoS ONE 2011, 6, e26468. [Google Scholar] [CrossRef]


| Gene Name | Gene ID | ORF (bp) | Deduced Polypetide | Exon No. | Genome Location | ||
|---|---|---|---|---|---|---|---|
| Length (aa) | Mol. Wt (kda) | pI | |||||
| HcIAA1 | Hc118.198 | 552 | 183 | 20.698 | 9.13 | 5 | 1298266–1301736 |
| HcIAA2 | Hc909.36 | 735 | 244 | 26.869 | 8.65 | 6 | 311277–314281 |
| HcIAA3 | Hc563.22 | 501 | 166 | 19.177 | 4.84 | 4 | 170726–171832 |
| HcIAA4 | Hc215.30 | 855 | 284 | 31.179 | 9.04 | 6 | 354002–357876 |
| HcIAA5 | Hc1096.12 | 891 | 296 | 31.617 | 5.09 | 4 | 96290–98064 |
| HcIAA6 | Hc506.60 | 594 | 197 | 21.856 | 8.57 | 4 | 520699–522248 |
| HcIAA7 | Hc717.7 | 468 | 155 | 16.881 | 9.36 | 3 | 53508–54653 |
| HcIAA8 | Hc38.101 | 852 | 283 | 30.625 | 7.42 | 5 | 1966779–1969857 |
| HcIAA9 | Hc899.17 | 924 | 307 | 32.633 | 7.01 | 5 | 251006–254190 |
| HcIAA10 | Hc357.107 | 945 | 314 | 33.141 | 6.16 | 5 | 779004–783999 |
| HcIAA11 | Hc189.134 | 777 | 258 | 28.457 | 5.97 | 4 | 1076907–1077976 |
| HcIAA12 | Hc189.121 | 594 | 197 | 21.273 | 5.30 | 3 | 987719–988944 |
| HcIAA13 | Hc107.22 | 912 | 303 | 33.216 | 9.31 | 5 | 734366–735832 |
| HcIAA14 | Hc285.12 | 735 | 244 | 26.556 | 7.93 | 5 | 146079–148497 |
| HcIAA15 | Hc440.26 | 729 | 242 | 26.643 | 9.23 | 5 | 424147–425365 |
| HcIAA16 | Hc47.162 | 528 | 175 | 19.225 | 5.11 | 4 | 1790499–1791855 |
| HcIAA17 | Hc1004.1 | 537 | 178 | 20.500 | 4.66 | 3 | 64402–73225 |
| HcIAA18 | Hc412.59.2 | 933 | 310 | 34.252 | 5.91 | 6 | 489469–493695 |
| HcIAA19 | Hc269.7 | 1005 | 334 | 35.863 | 8.23 | 5 | 386761–389599 |
| HcIAA20 | Hc41.21 | 942 | 313 | 33.433 | 6.55 | 5 | 707930–711511 |
| HcIAA21 | Hc614.34 | 858 | 285 | 30.632 | 6.55 | 5 | 348817–352993 |
| HcIAA22 | Hc42.52 | 585 | 194 | 21.522 | 5.40 | 5 | 737421–740882 |
| HcIAA23 | Hc270.58 | 615 | 204 | 23.116 | 10.15 | 3 | 459106–460132 |
| HcIAA24 | Hc484.51.1 | 1059 | 352 | 39.190 | 6.29 | 7 | 480251–483666 |
| HcIAA25 | Hc566.42 | 546 | 181 | 19.907 | 5.45 | 4 | 628269–630501 |
| HcIAA26 | Hc641.57 | 762 | 253 | 27.553 | 7.14 | 5 | 340256–342758 |
| HcIAA27 | Hc326.19 | 522 | 173 | 19.566 | 6.78 | 4 | 329556–335070 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ke, Y.; Abbas, F.; Zhou, Y.; Yu, R.; Yue, Y.; Li, X.; Yu, Y.; Fan, Y. Genome-Wide Analysis and Characterization of the Aux/IAA Family Genes Related to Floral Scent Formation in Hedychium coronarium. Int. J. Mol. Sci. 2019, 20, 3235. https://doi.org/10.3390/ijms20133235
Ke Y, Abbas F, Zhou Y, Yu R, Yue Y, Li X, Yu Y, Fan Y. Genome-Wide Analysis and Characterization of the Aux/IAA Family Genes Related to Floral Scent Formation in Hedychium coronarium. International Journal of Molecular Sciences. 2019; 20(13):3235. https://doi.org/10.3390/ijms20133235
Chicago/Turabian StyleKe, Yanguo, Farhat Abbas, Yiwei Zhou, Rangcai Yu, Yuechong Yue, Xinyue Li, Yunyi Yu, and Yanping Fan. 2019. "Genome-Wide Analysis and Characterization of the Aux/IAA Family Genes Related to Floral Scent Formation in Hedychium coronarium" International Journal of Molecular Sciences 20, no. 13: 3235. https://doi.org/10.3390/ijms20133235
APA StyleKe, Y., Abbas, F., Zhou, Y., Yu, R., Yue, Y., Li, X., Yu, Y., & Fan, Y. (2019). Genome-Wide Analysis and Characterization of the Aux/IAA Family Genes Related to Floral Scent Formation in Hedychium coronarium. International Journal of Molecular Sciences, 20(13), 3235. https://doi.org/10.3390/ijms20133235







