The 40S Ribosomal Protein S6 Response to Blue Light by Interaction with SjAUREO in Saccharina japonica
Abstract
1. Introduction
2. Results
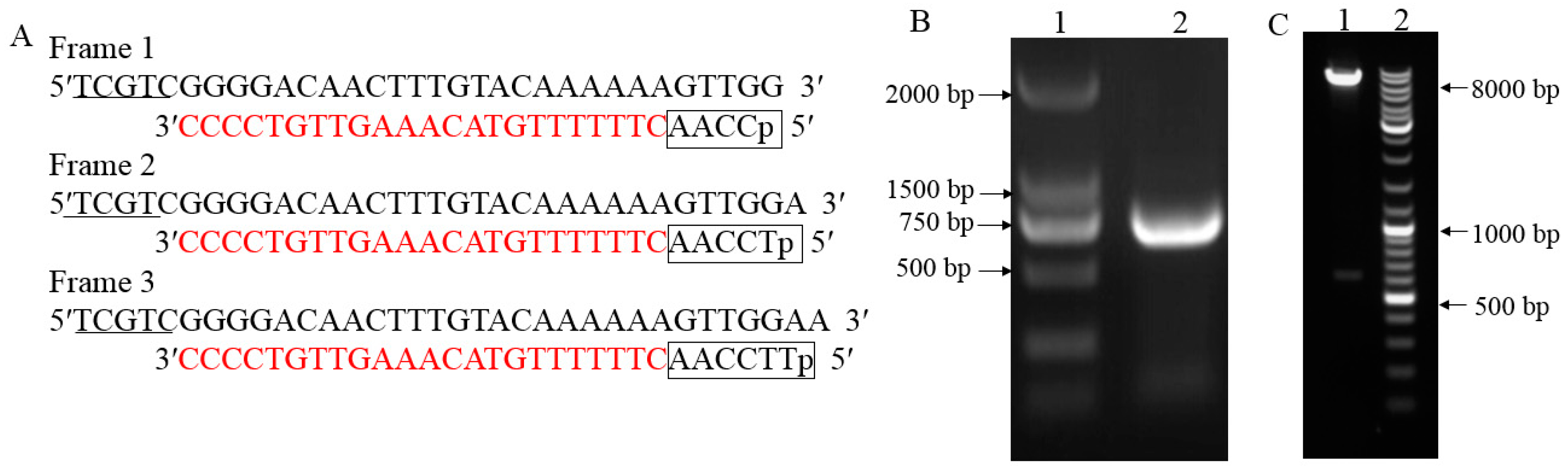
2.1. cDNA Three-Frame Library Construction and Evaluation
2.2. The Bait Construction, Toxicity and Auto-Activation Validation
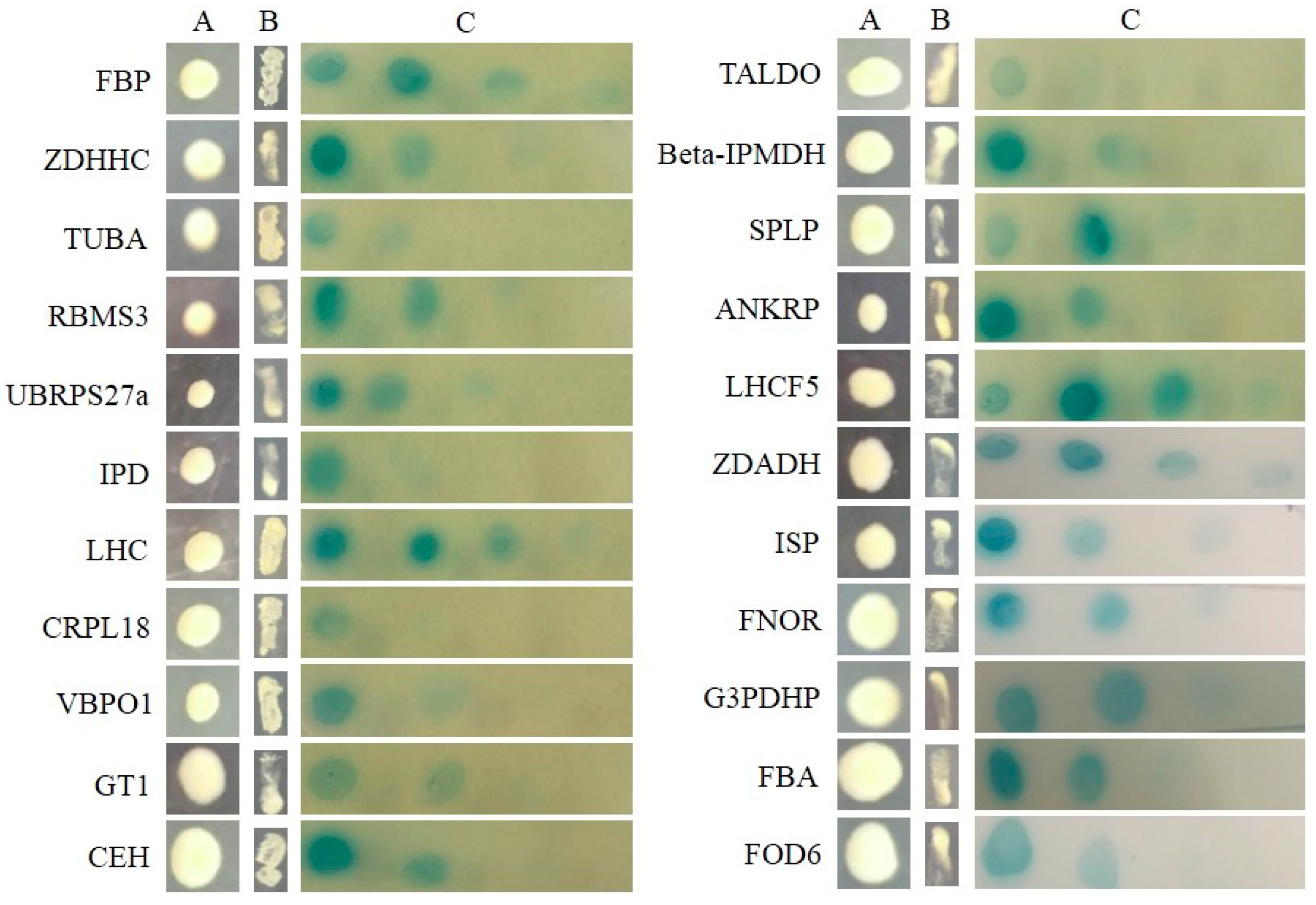
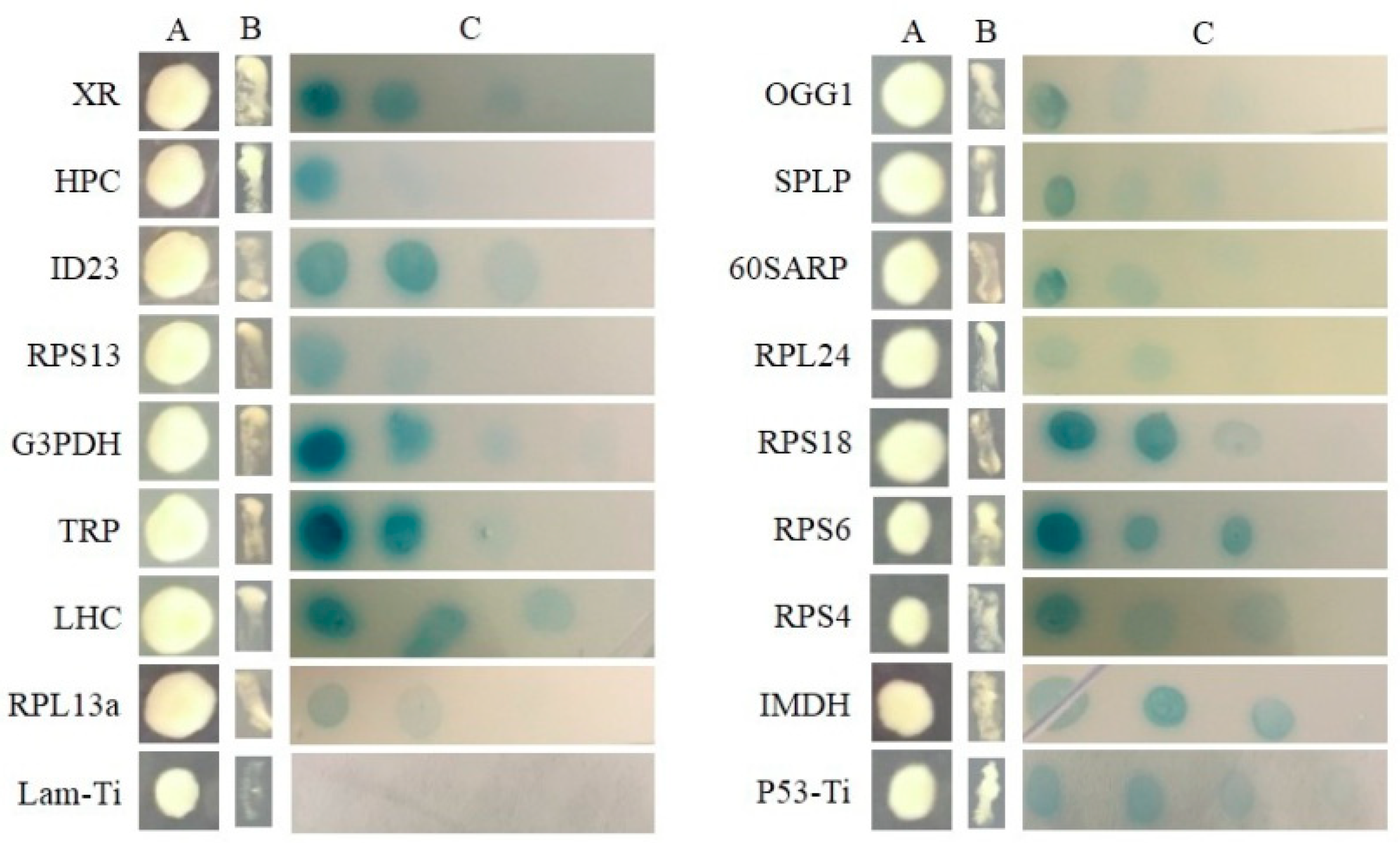
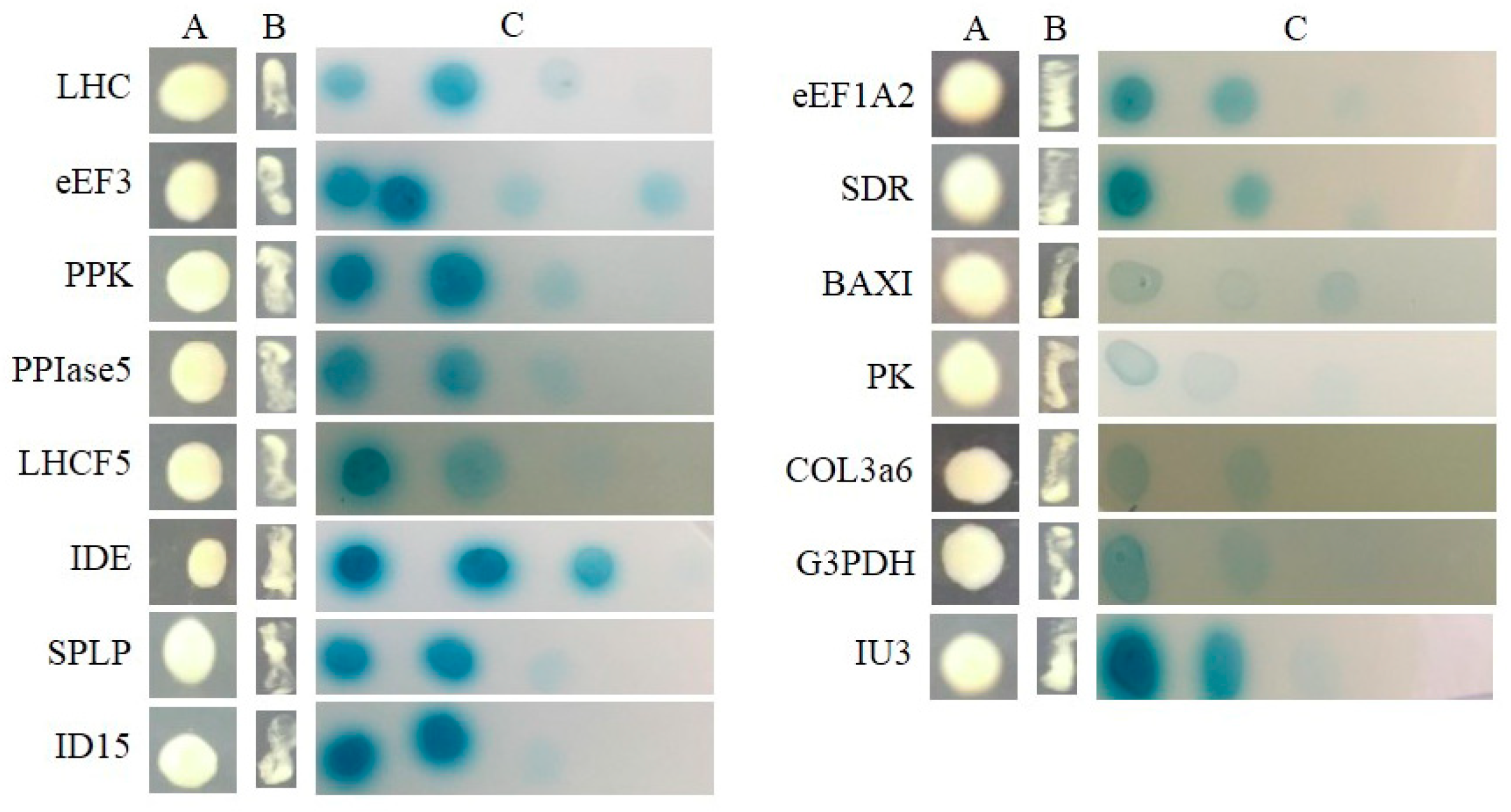
2.3. Searching for SjAUREO Interactors
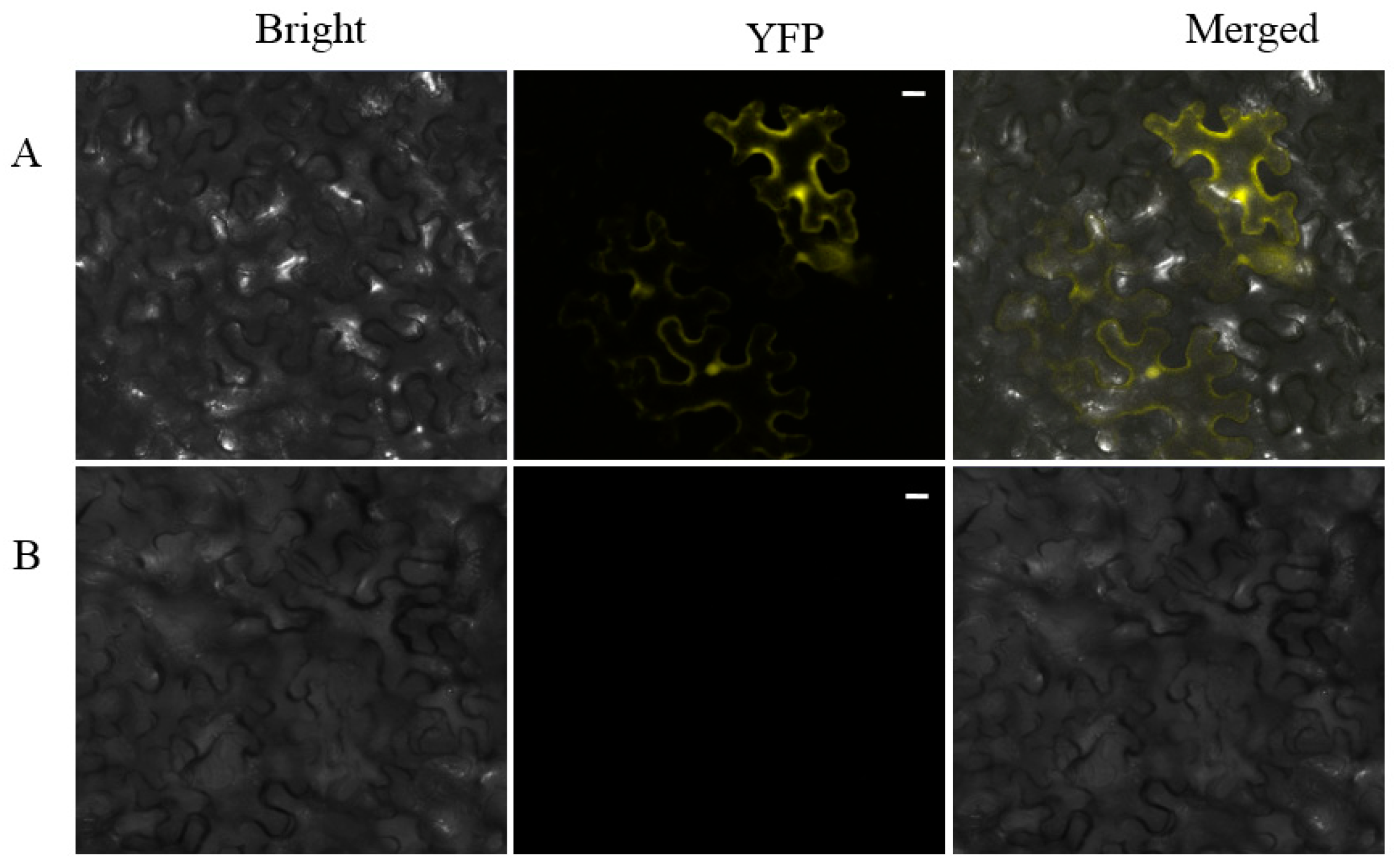
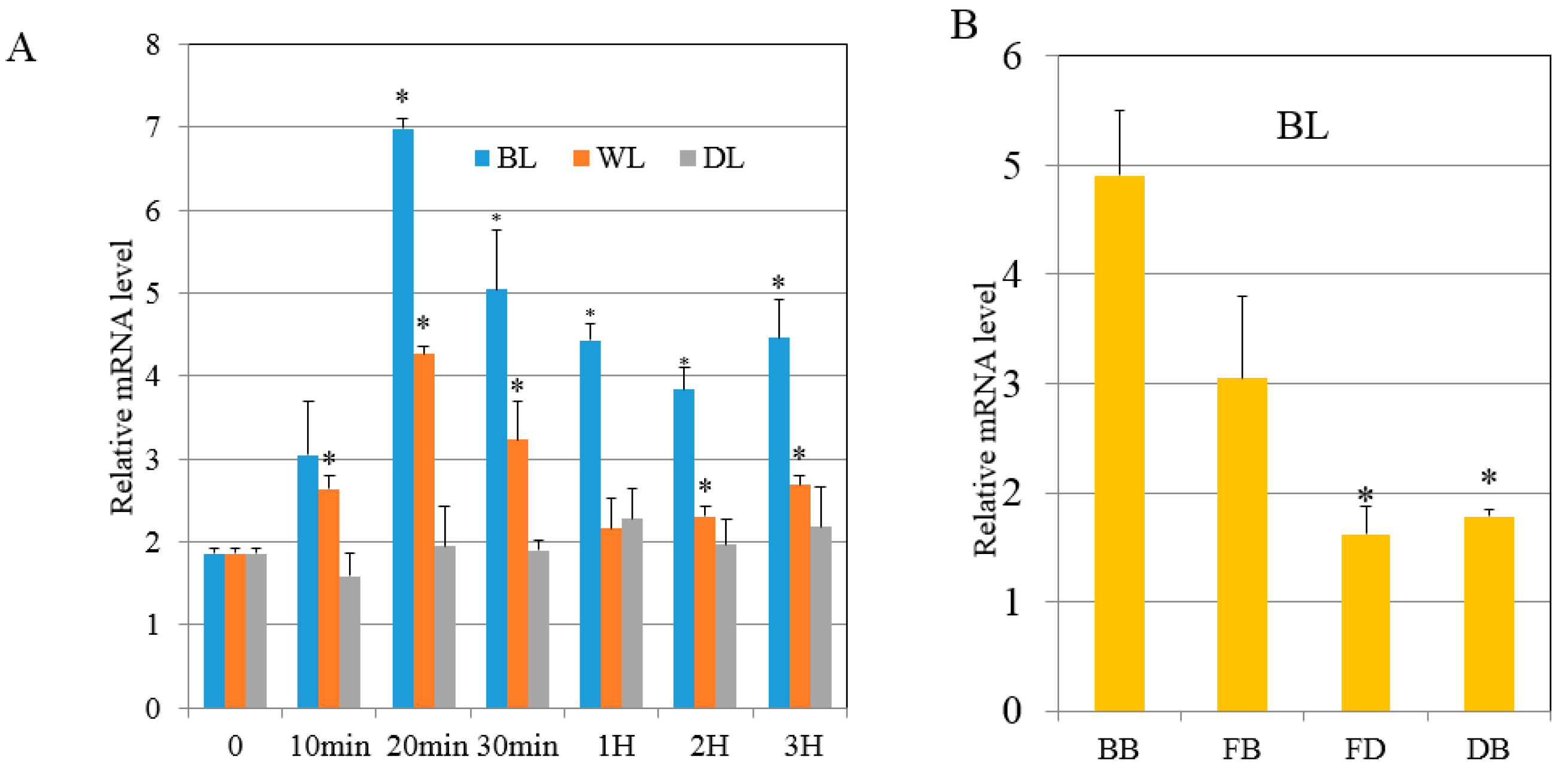
2.4. Bimolecular Fuorescence Complementation (BiFC) Assay and Expression Analyses of 40S Ribosome S6 Gene Detected with qRT-PCR
3. Discussion
4. Materials and Methods
4.1. Kelp Materials
4.2. RNA Extraction and Quantitative RT-PCR Analysis
4.3. cDNA Three-Frame Library Construction and Evaluation
4.4. The Prey Vector Construction and Application to Y2H
4.5. Yeast Two-Hybrid Library Screening and Interaction Conformation
4.6. Bimolecular Fluorescence Complementation Assays
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hay, I.D.; Rehman, Z.U.; Moradali, M.F.; Wang, Y.; Rehm, B.H.A. Microbial alginate production, modification and its applications. Microb. Biotechnol. 2013, 6, 6. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Shao, Z.; Jin, W.; Duan, D. Comparative characterization of two GDP-mannose dehydrogenase genes from Saccharina japonica (Laminariales, Phaeophyceae). BMC Plant. Bio. 2016, 16, 1. [Google Scholar] [CrossRef]
- Vishchuk, O.S.; Ermakova, S.P.; Zvyagintseva, T.N. Sulfated polysaccharides from brown seaweeds Saccharina japonica and Undariapinnatifida: Isolation, structural characteristics, and antitumor activity. Carbohydrate Res. 2011, 346, 2769–2776. [Google Scholar] [CrossRef] [PubMed]
- Hegemann, P. Algal sensory photoreceptors. Annual Rev. Plant Biol. 2008, 59, 167–189. [Google Scholar] [CrossRef] [PubMed]
- Ragni, M.; d’Alcalà, M.R. Circadian variability in the photobiology of Phaeodactylumtricornutum: Pigment content. J. Plankton Res. 2007, 29, 141–156. [Google Scholar] [CrossRef]
- Depauw, F.A.; Rogato, A.; Ribera d’Alcalá, M.; Falciatore, A. Exploring the molecular basis of responses to light in marine diatoms. J. Exp. Bot. 2012, 63, 1575–1591. [Google Scholar] [CrossRef]
- Matiiv, A.B.; Chekunova, E.M. Aureochromes–Blue Light Receptors. Biochemistry 2018, 83, 662–673. [Google Scholar] [CrossRef]
- Jiao, Y.; Lau, O.S.; Deng, X.W. Light-regulated transcriptional networks in higher plants. Nat. Rev. Genet. 2007, 8, 217. [Google Scholar] [CrossRef]
- Masuda, S.; Hasegawa, K.; Ohta, H.; Ono, T.A. Crucial role in light signal transduction for the conserved Met93 of the BLUF protein PixD/Slr1694. Plant Cell Physiol. 2008, 49, 1600–1606. [Google Scholar] [CrossRef][Green Version]
- Rizzini, L.; Favory, J.J.; Cloix, C.; Faggionato, D.; O”Hara, A.; Kaiserli, E.; Baumeister, R.; Schäfer, E.; Nagy, F.; Jenkins, G.I.; et al. Perception of uv-b by the arabidopsis uvr8 protein. Science. 2011, 332, 103–106. [Google Scholar] [CrossRef]
- Sharrock, R.A. The phytochrome red/far-red photoreceptor superfamily. Genome Bio. 2008, 9, 230. [Google Scholar] [CrossRef] [PubMed]
- Chaves, I.; Pokorny, R.; Byrdin, M.; Hoang, N.; Ritz, T.; Brettel, K.; Essen, L.O.; van der Horst, G.T.; Batschauer, A.; Ahmad, M. The cryptochromes: Blue light photoreceptors in plants and animals. Annu. Rev. Plant Biol. 2011, 62, 335–364. [Google Scholar] [CrossRef] [PubMed]
- Galvão, V.C.; Fankhauser, C. Sensing the light environment in plants: Photoreceptors and early signaling steps. Curr. Opin. Neurobiol. 2015, 1, 13. [Google Scholar] [CrossRef]
- Kottke, T.; Oldemeyer, S.; Wenzel, S.; Zou, Y.; Mittag, M. Cryptochrome photoreceptors in green algae: Unexpected versatility of mechanisms and functions. J. Plant Physiol. 2017, 217, 4–14. [Google Scholar] [CrossRef] [PubMed]
- Suetsugu, N.; Wada, M. Evolution of three lov blue light receptor families in green plants and photosynthetic stramenopiles: Phototropin, ztl/fkf1/lkp2 and aureochrome. Plant & Cell Physiol. 2013, 54, 8–23. [Google Scholar]
- Jeong, R.D.; Chandra, A.C.; Barman, S.R.; Navarre, D.; Klessig, D.F.; Kachroo, A.; Kachroo, P. Cryptochrome 2 and phototropin 2 regulate resistance protein-mediated viral defense by negatively regulating an E3 ubiquitin ligase. Proc. Natl. Acad. Sci. USA 2010, 107, 13538–13543. [Google Scholar] [CrossRef] [PubMed]
- Cavalier, S.T.; Robertson, D.L. The kingdom chromista origin and systemmatics. Progr. Phycol. Res. 1986, 4, 309–347. [Google Scholar]
- Takahashi, F.; Yamagata, D.; Ishikawa, M.; Fukamatsu, Y.; Ogura, Y.; Kasahara, M.; Kiyosue, T.; Kikuyama, M.; Wada, M.; Kataoka, H. Aureochrome, a photoreceptor required for photomorphogenesis in stramenopiles. Proc. Natl. Acad. Sci. USA 2007, 104, 19625–19630. [Google Scholar] [CrossRef] [PubMed]
- Ishikawa, M.; Takahashi, F.; Nozaki, H.; Nagasato, C.; Motomura, T.; Kataoka, H. Distribution and phylogeny of the blue light receptors aureochromes in eukaryotes. Planta 2009, 230, 543–552. [Google Scholar] [CrossRef]
- Deng, Y.; Yao, J.; Fu, G.; Guo, H.; Duan, D. Isolation, expression, and characterization of blue light receptor aureochrome gene from Saccharina japonica (laminariales, phaeophyceae). Mar. Biotechnol. 2014, 16, 135–143. [Google Scholar] [CrossRef]
- Bowler, C.; Allen, A.E.; Badger, J.H.; Grimwood, J.; Jabbari, K.; Kuo, A.; Maheswari, U.; Martens, C.; Maumus, F.; Otillar, R.P.; et al. The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature 2008, 456, 239–244. [Google Scholar] [CrossRef] [PubMed]
- Serif, M.; Lepetit, B.; Weißert, K.; Kroth, P.G.; Bartulos, R. A fast and reliable strategy to generate TALEN-mediated gene knockouts in the diatom Phaeodactylumtricornutum. Algal Res. 2017, 23, 186–195. [Google Scholar] [CrossRef]
- Kroth, P.G.; Wilhelm, C.; Kottke, T. An update on aureochromes: Phylogeny–mechanism–function. J. Plant Physiol. 2017, 217, 20–26. [Google Scholar] [CrossRef]
- Herrou, J.; Crosson, S. Function, structure and mechanism of bacterial photosensory LOV proteins. Nat. Rev. Microbiol. 2011, 9, 713–723. [Google Scholar] [CrossRef] [PubMed]
- Toyooka, T.; Hisatomi, O.; Takahashi, F.; Kataoka, H.; Terazima, M. Photoreactions of aureochrome-1. Biophys. J. 2011, 100, 2801–2809. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Herman, E.; Kottke, T. Allosterically regulated unfolding of the A′ α helix exposes the dimerization site of the blue-light-sensing aureochrome-LOV domain. Biochemistry 2015, 54, 1484–1492. [Google Scholar] [CrossRef]
- Heintz, U.; Schlichting, I. Blue light-induced LOV domain dimerization enhances the affinity of Aureochrome 1a for its target DNA sequence. Elife 2015, 5, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Hisatomi, O.; Takeuchi, K.; Zikihara, K.; Ookubo, Y.; Nakatani, Y.; Takahashi, F.; Kataoka, H. Blue light-induced conformational changes in a light-regulated transcription factor, aureochrome-1. Plant Cell Physiol. 2012, 54, 93–106. [Google Scholar] [CrossRef]
- Huysman, M.J.; Fortunato, A.E.; Matthijs, M.; Costa, B.S.; Vanderhaeghen, R.; Van, D.H.; Sachse, M.; Inzé, D.; Bowler, C.; Kroth, P.G.; et al. Aureochrome1a-mediated induction of the diatom-specific cyclin dscyc2 controls the onset of cell division in diatoms (phaeodactylum tricornutum). Plant Cell 2013, 25, 215–228. [Google Scholar] [CrossRef]
- Volarević, S.; Stewart, M.J.; Ledermann, B.; Zilberman, F.; Terracciano, L.; Montini, E.; Grompe, M.; Kozma, S.C.; Thomas, G. Proliferation, but not growth, blocked by conditional deletion of 40S ribosomal protein S6. Science 2000, 288, 2045–2047. [Google Scholar] [CrossRef]
- Stynen, B.; Tournu, H.; Tavernier, J.; Dijck, P.V. Diversity in genetic in vivo methods for protein-protein interaction studies: From the yeast two-hybrid system to the mammalian split-luciferase system. Microbiol Molecul. Biol. R. 2012, 76, 331–382. [Google Scholar] [CrossRef] [PubMed]
- Jacquier, A. Two-hybrid systems — methods and protocols. edited by paul n. macdonald, published by humana press. Biochimie 2002, 84, 349. [Google Scholar] [CrossRef]
- Ruvinsky, I.; Meyuhas, O. Ribosomal protein s6 phosphorylation: From protein synthesis to cell size. Trends Biochem. Sci. 2006, 31, 342–348. [Google Scholar] [CrossRef] [PubMed]
- Ruvinsky, I.; Sharon, N.; Lerer, T.; Cohen, H.; Stolovich, R.M.; Nir, T.; Dor, Y.; Zisman, P.; Meyuhas, O. Ribosomal protein S6 phosphorylation is a determinant of cell size and glucose homeostasis. Genes Dev. 2005, 19, 2199–2211. [Google Scholar] [CrossRef]
- Enganti, R.; Cho, S.K.; Toperzer, J.D.; Urquidi, C.R.A.; Cakir, O.S.; Ray, A.P.; Abraham, P.E.; Hettich, R.L.; Arnim, A.G. Phosphorylation of ribosomal protein RPS6 integrates light signals and circadian clock signals. Front. Plant Sci. 2018, 8, 2210–2228. [Google Scholar] [CrossRef]
- Yao, J.; Fu, W.; Wang, X.; Duan, D. Improved RNA isolation from Laminaria japonica Aresch (Laminariaceae, Phaeophyta). J. Appl. Phycol. 2009, 21, 233–238. [Google Scholar] [CrossRef]
- Shao, Z.; Liu, F.; Li, Q.; Yao, J.; Duan, D. Characterization of ribulose-1, 5-bisphosphate carboxylase/oxygenase and transcriptional analysis of its related genes in Saccharina japonica (Laminariales, Phaeophyta). Chin. J. Oceanol. Limn. 2014, 32, 377–389. [Google Scholar] [CrossRef]
- Zhao, W.; Li, X.; Liu, W.H.; Zhao, J.; Jin, Y.M.; Sui, T.T. Construction of high-quality caco-2 three-frame cDNA library and its application to yeast two-hybrid for the human astrovirus protein–protein interaction. J. Virol. Methods. 2014, 205, 104–109. [Google Scholar] [CrossRef]
| Group | CFU | RR |
|---|---|---|
| DK | 8.4 × 106 | 100% |
| BL | 1.2 × 107 | 100% |
| WL | 7.8 × 106 | 100% |
| Accession No. | Characteristics | Homology | Clone Number |
|---|---|---|---|
| CBN75354.1 | Fructose 1,6-bisphosphatase (FBP) | 54% | 2 |
| CBJ48329.1 | Fructose-1,6-bisphosphate aldolase (FBA) | 96% | 1 |
| CAQ51444.1 | Putative vanadium-dependent bromoperoxidase 6 (VBP 6) | 55% | 1 |
| CAD37191.1 | Vanadium-dependent bromoperoxidase 1 (VBP 1) | 64% | 1 |
| EJY76073.1 | DHHC zinc finger domain containing protein (ZDHHC) | 65% | 1 |
| CBJ28184.1 | Tubulin alpha (TUBA) | 76% | 1 |
| KKF25031.1 | RNA-binding motif,Single-stranded-interacting protein 3-like (RBMS3) | 97% | 2 |
| CBN74156.1 | Similar to ubiquitin and ribosomal protein S27a precursor (UBRPS27a) | 86% | 1 |
| OPX53870.1 | 3-isopropylmalate dehydrogenase (IMDH) | 99% | 3 |
| CBJ25909.1 | Cytoplasmic ribosomal protein L18 (CRPL18) | 95% | 1 |
| CBN77006.1 | Light harvesting complex protein (LHC) | 75% | 1 |
| ACE80197.1 | Chloroplast light harvesting protein lhcf5 (LHCF5) | 100% | 2 |
| CBN75623.1 | Phosphoglycerate kinase (PGK) | 93% | 1 |
| CAD37191.1 | Vanadium-dependent bromoperoxidase 1 (VBPO1) | 89% | 2 |
| WP_026719310.1 | Glycosyltransferase family 1 protein (GT1) | 46% | 1 |
| CBJ31450.1 | CAB/ELIP/HLIP superfamily protein (CEH) | 86% | 3 |
| WP_094674345.1 | Transaldolase (TALDO) | 45% | 1 |
| AAA66917.1 | Beta-isopropylmalate dehydrogenase (Beta-IPMDH) | 95% | 1 |
| ABP93411.1 | Sporulation-like protein (SPLP) | 90% | 1 |
| CBJ30026.1 | Ankyrin repeat protein (ANKRP) | 46% | 1 |
| EGA60350.1 | Zn-dependent alcohol dehydrogenase family (ZDADH) | 51% | 1 |
| CBJ29582.1 | Cytochrome b6-f complex iron-sulfur subunit (ISP) | 83% | 1 |
| ABU96658.1 | glyceraldehyde-3-phosphate dehydrogenase precursor (G3PDHP) | 99% | 1 |
| CBN78345.1 | Ferredoxin-NADP oxidoreductase (FNOR) | 86% | 1 |
| CBJ31978.1 | Flagellar outer dynein arm light chain 6 (FOD6) | 99% | 1 |
| Accession No. | Characteristics | Homology | Clone Number |
|---|---|---|---|
| EWM23897.1 | 40S ribosomal protein S18 (RPS18) | 74% | 1 |
| EWM21878.1 | 40S ribosomal protein S6 (RPS6) | 75% | 3 |
| CBJ29820.1 | 40S ribosomal protein S4 (RPS4) | 87% | 1 |
| OPX53870.1 | 3-isopropylmalate dehydrogenase (IMDH) | 95% | 4 |
| CBJ28016.1 | Similar to L-xylulose reductase (XR) | 94% | 1 |
| XP_005844401.1 | Hypothetical protein chlncdraft_139059 (HPC) | 68% | 3 |
| CBJ27783.1 | Imm downregulated 23 (ID 23) | 72% | 1 |
| ACN09872.1 | 40S ribosomal protein S13 (RPS13) | 79% | 1 |
| CBJ31240.1 | Chloroplast glyceraldehyde-3-phosphate dehydrogenase (G3PDH) | 83% | 1 |
| CBJ31434.1 | Transient receptor potential channel fragment (TRP) | 63% | 1 |
| CBN78494.1 | Light harvesting complex protein (LHC) | 89% | 2 |
| CBJ25662.1 | 60s ribosomal protein L13a (RPL13a) | 91% | 4 |
| CBJ27158.1 | Endonuclease III/similar to 8-oxoguanine DNA glycosylase isoform 1b (OGG1) | 60% | 1 |
| ABP93411.1 | Sporulation-like protein (SPLP) | 96% | 2 |
| EWM29947.1 | 60s acidic ribosomal protein (60SARP) | 50% | 1 |
| CBJ28054.1 | Ribosomal protein l24 (RPL24) | 89% | 1 |
| Accession No. | Characteristics | Homology | Clone Number |
|---|---|---|---|
| CBJ33313.1 | Light harvesting complex protein (LHC) | 83% | 3 |
| CBJ25582.1 | Elongation factor EF-3 (eEF3) | 94% | 1 |
| CBN75623.1 | Phosphoglycerate kinase (PPK) | 93% | 3 |
| CBJ26174.1 | FKBP-type peptidyl-prolyl cis-trans isomerase 5 (PPIase5) | 76% | 1 |
| ACE80197.1 | Chloroplast light harvesting protein lhcf5 (LHCF5) | 100% | 6 |
| CBN77215.1 | insulin-degrading enzyme (IDE) | 69% | 1 |
| ABP93411.1 | Sporulation-like protein (SPLP) | 99% | 1 |
| CBJ25754.1 | Imm downregulated 15 (ID15) | 87% | 1 |
| CBN74763.1 | Imm upregulated 3 (IU3) | 63% | 1 |
| CBJ31240.1 | Glyceraldehyde 3-phosphate dehydrogenase (G3PDH) | 71% | 1 |
| CBJ32893.1 | eukaryotic translation elongation factor 1 alpha (eEF1A2) | 93% | 3 |
| CBN74950.1 | Short-chain dehydrogenase/reductase (SDR) | 56% | 1 |
| CBN79712.1 | Putative BAX inhibitor (BAXI) | 81% | 1 |
| AIT70004.1 | Pyruvate kinase (PK) | 77% | 1 |
| XP_014788837.1 | collagen alpha-3(VI) chain-like (COL3a6) | 29% | 1 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luan, H.; Yao, J.; Chen, Z.; Duan, D. The 40S Ribosomal Protein S6 Response to Blue Light by Interaction with SjAUREO in Saccharina japonica. Int. J. Mol. Sci. 2019, 20, 2414. https://doi.org/10.3390/ijms20102414
Luan H, Yao J, Chen Z, Duan D. The 40S Ribosomal Protein S6 Response to Blue Light by Interaction with SjAUREO in Saccharina japonica. International Journal of Molecular Sciences. 2019; 20(10):2414. https://doi.org/10.3390/ijms20102414
Chicago/Turabian StyleLuan, Hexiang, Jianting Yao, Zhihang Chen, and Delin Duan. 2019. "The 40S Ribosomal Protein S6 Response to Blue Light by Interaction with SjAUREO in Saccharina japonica" International Journal of Molecular Sciences 20, no. 10: 2414. https://doi.org/10.3390/ijms20102414
APA StyleLuan, H., Yao, J., Chen, Z., & Duan, D. (2019). The 40S Ribosomal Protein S6 Response to Blue Light by Interaction with SjAUREO in Saccharina japonica. International Journal of Molecular Sciences, 20(10), 2414. https://doi.org/10.3390/ijms20102414




