Comprehensive Analysis of the Cadmium Tolerance of Abscisic Acid-, Stress- and Ripening-Induced Proteins (ASRs) in Maize
Abstract
1. Introduction
2. Results
2.1. Candidate Cd Tolerance Gene ZmASR1 Isolated from the Maize cDNA Library
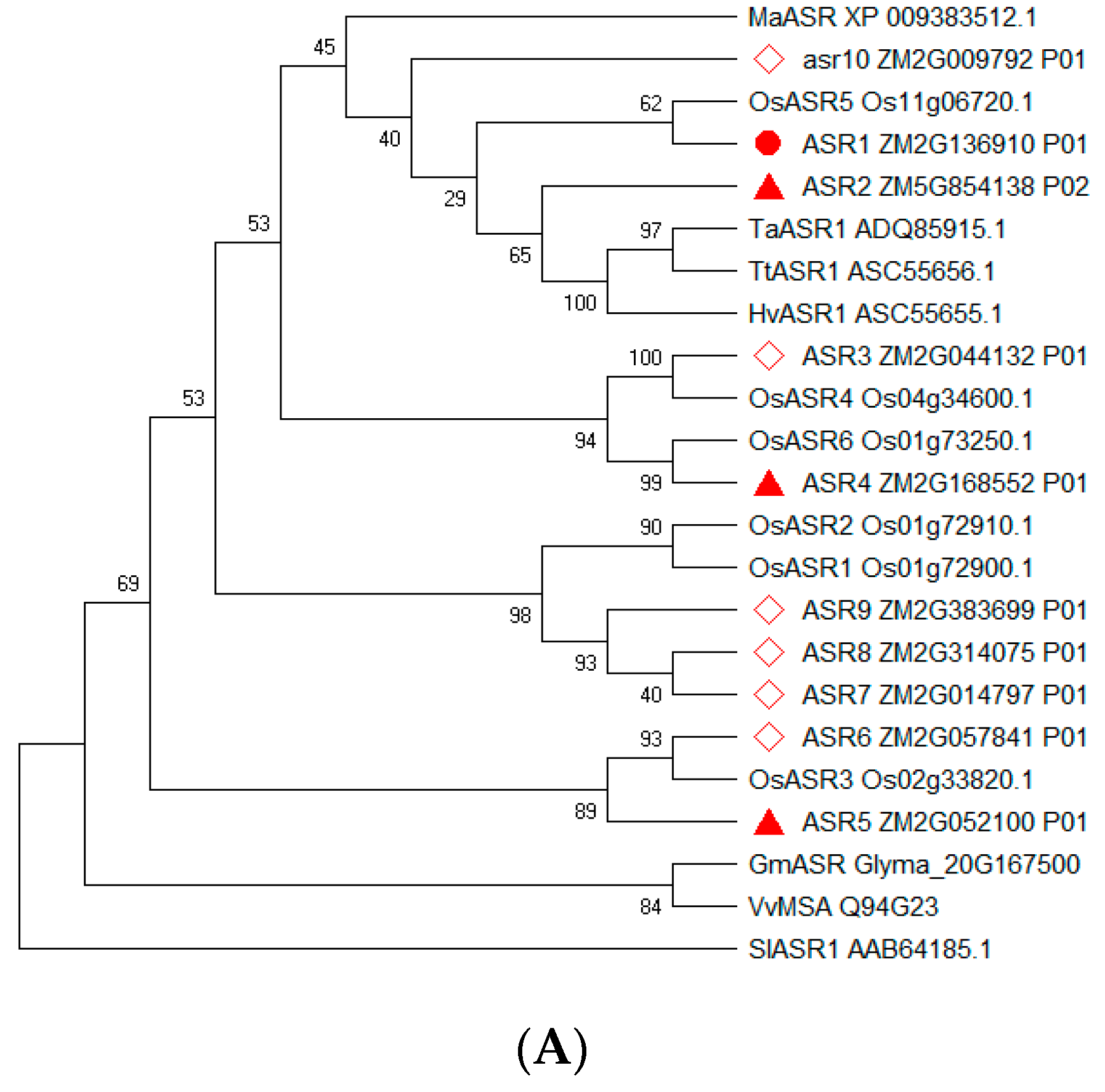
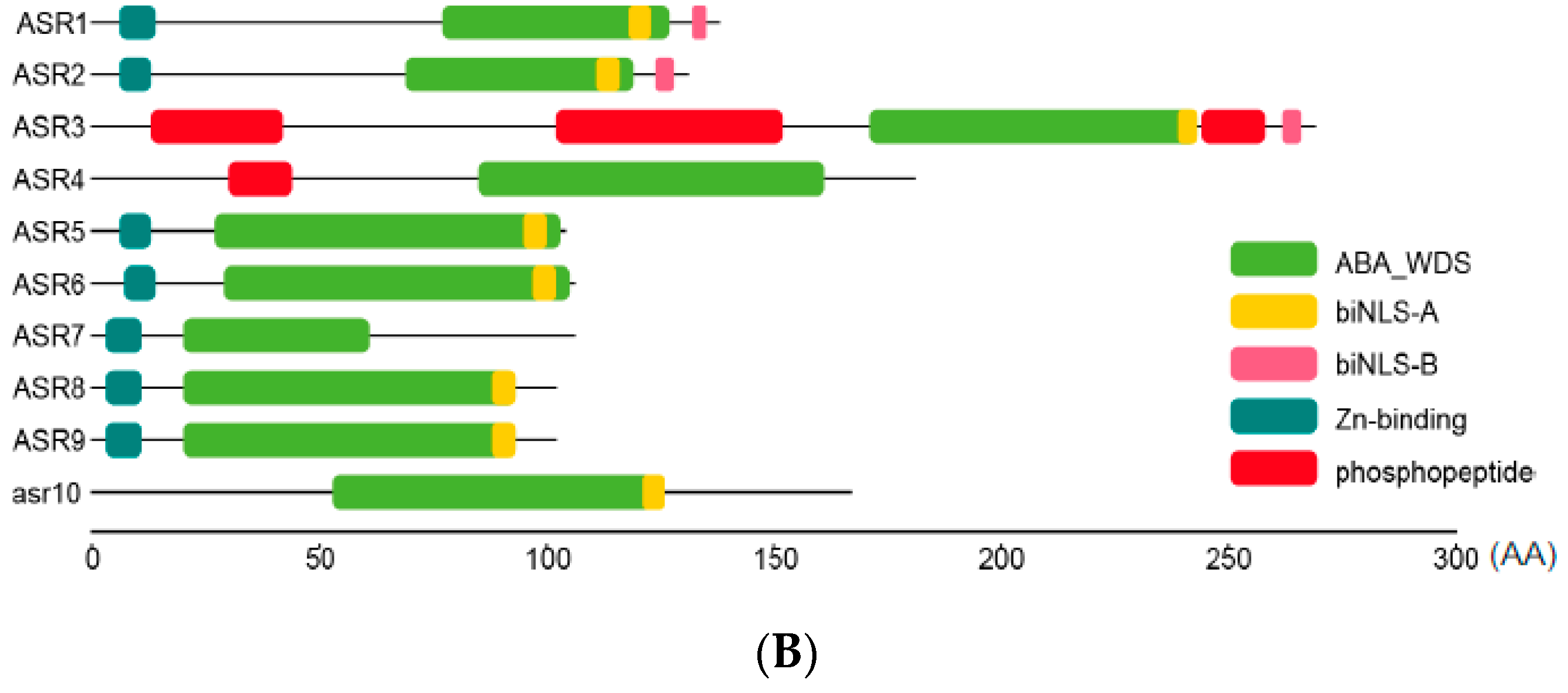
2.2. Sequence Analysis of Maize ZmASR Genes
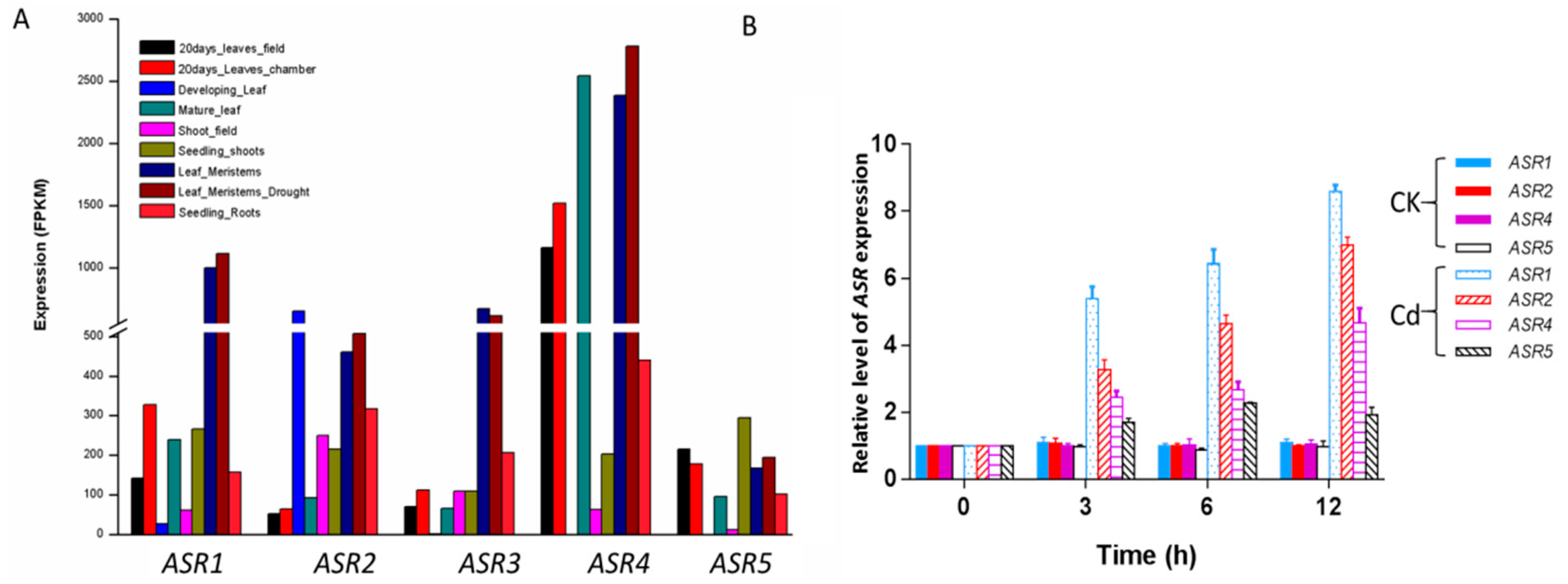
2.3. Expression Patterns of ZmASR Genes
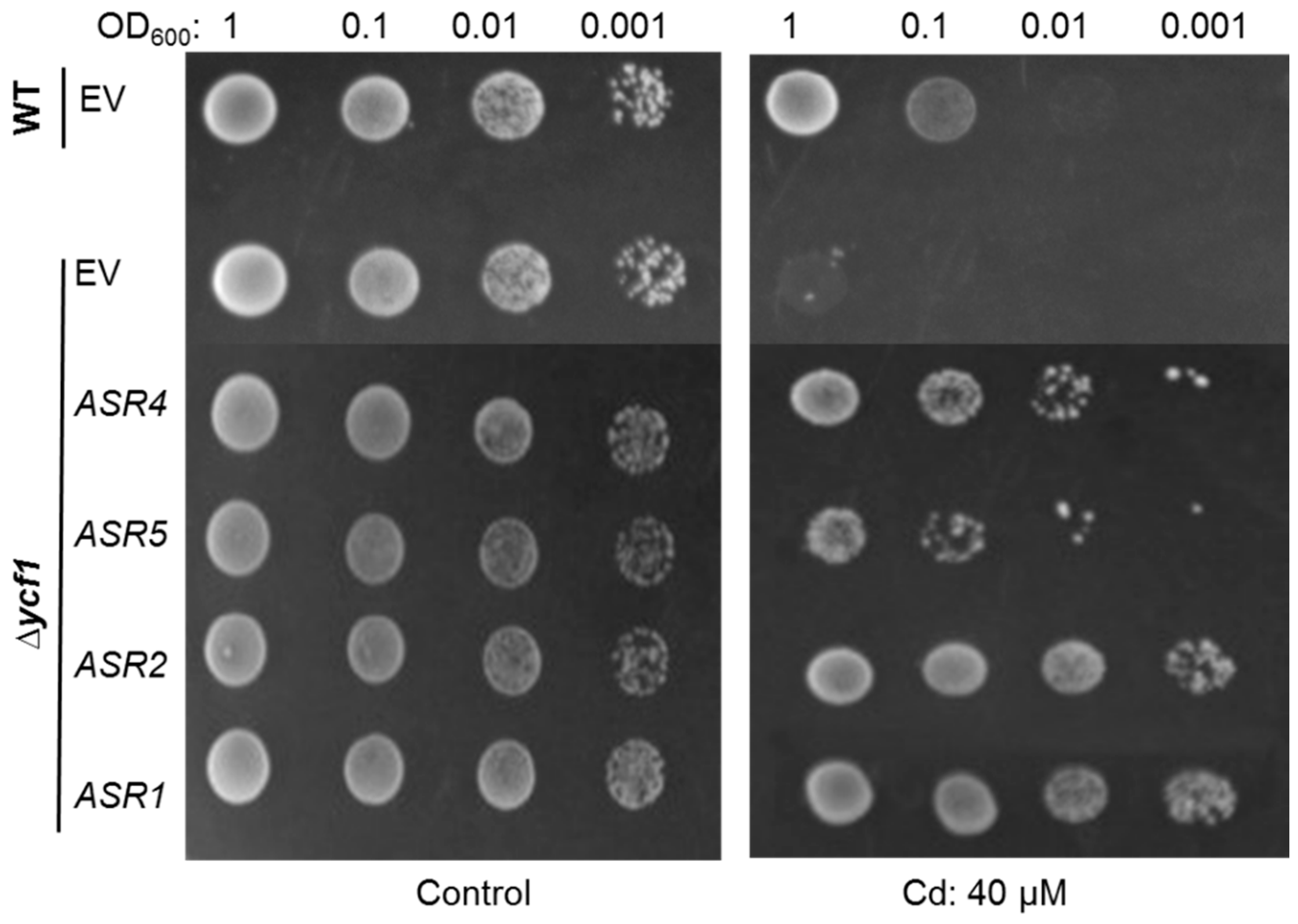
2.4. Confirmation That Maize ASRs Confer Cd Tolerance in Cd-Sensitive Yeast Mutant
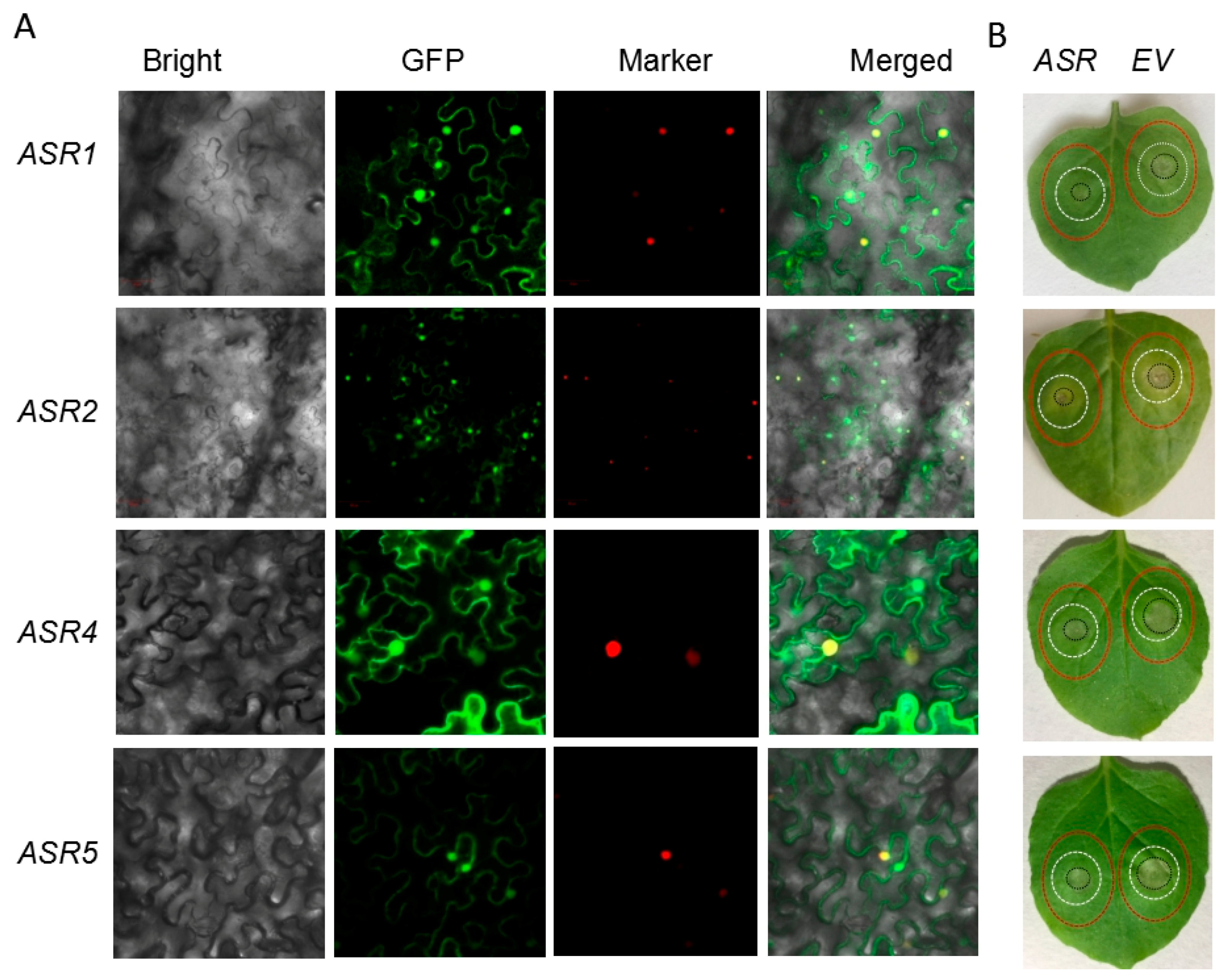
2.5. Transient Expression of Maize ASRs Conferred Cd Tolerance in Tobacco
3. Discussion
3.1. Maize ASRs Function in Cd-Tolerance
3.2. The Feature of Dual Targeting Determines the Multiple Function of ASRs
4. Materials and Methods
4.1. Isolation of Cd-Tolerance Genes from the Maize Leaf cDNA Library
4.2. Maize Seedlings Cultivation and cDNA Synthesis for Cloning ASR
4.3. Sequence Alignment and Phylogenetic Analysis
4.4. In Silico Expression Analysis
4.5. Yeast and Transient Plant Expression Vector Reconstruction
4.6. Cd Tolerance Complementation Assay in Cd-Sensitive Yeast
4.7. Agro-Infiltration and Cd Response in Tobacco Leaves
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Lou, L.; Kang, J.; Pang, H.; Li, Q.; Du, X.; Wu, W.; Chen, J.; Lv, J. Sulfur protects pakchoi (Brassica chinensis L.) seedlings against cadmium stress by regulating ascorbate-glutathione metabolism. Int. J. Mol. Sci. 2017, 18, 1628. [Google Scholar] [CrossRef]
- Clemens, S.; Ma, J.F. Toxic heavy metal and metalloid accumulation in crop plants and foods. Annu. Rev. Plant Biol. 2016, 67, 489–512. [Google Scholar] [CrossRef]
- Jian, H.; Yang, B.; Zhang, A.; Ma, J.; Ding, Y.; Chen, Z.; Li, J.; Xu, X.; Liu, L. Genome-wide identification of micrornas in response to cadmium stress in oilseed rape (Brassica napus L.) using high-throughput sequencing. Int. J. Mol. Sci. 2018, 19, 1431. [Google Scholar] [CrossRef] [PubMed]
- Guo, Q.; Meng, L.; Humphreys, M.W.; Scullion, J.; Mur, L.A.J. Expression of FlHMA3, a P1B2-ATPase from Festulolium loliaceum, correlates with response to cadmium stress. Plant Physiol. Biochem. 2017, 112, 270–277. [Google Scholar] [CrossRef]
- Ismael, M.A.; Elyamine, A.M.; Zhao, Y.Y.; Moussa, M.G.; Rana, M.S.; Afzal, J.; Imran, M.; Zhao, X.H.; Hu, C.X. Can selenium and molybdenum restrain cadmium toxicity to pollen grains in Brassica napus? Int. J. Mol. Sci. 2018, 19, 2163. [Google Scholar] [CrossRef] [PubMed]
- Adrees, M.; Ali, S.; Rizwan, M.; Zia-Ur-Rehman, M.; Ibrahim, M.; Abbas, F.; Farid, M.; Qayyum, M.F.; Irshad, M.K. Mechanisms of silicon-mediated alleviation of heavy metal toxicity in plants: A review. Ecotoxicol. Environ. Saf. 2015, 119, 186–197. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Li, Y.; Yin, Z.; Jiang, J.; Zhang, M.; Guo, X.; Ye, Z.; Zhao, Y.; Xiong, H.; Zhang, Z.; et al. OsASR5 enhances drought tolerance through a stomatal closure pathway associated with ABA and H2O2 signalling in rice. Plant Biotechnol. J. 2017, 15, 183–196. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, R.M.; Iusem, N.D. Twenty years of research on ASR (ABA-stress-ripening) genes and proteins. Planta 2014, 239, 941–949. [Google Scholar] [CrossRef]
- Sade, H.; Meriga, B.; Surapu, V.; Gadi, J.; Sunita, M.S.; Suravajhala, P.; Kavi Kishor, P.B. Toxicity and tolerance of aluminum in plants: Tailoring plants to suit to acid soils. Biometals 2016, 29, 187–210. [Google Scholar] [CrossRef]
- Ricardi, M.M.; Gonzalez, R.M.; Zhong, S.; Dominguez, P.G.; Duffy, T.; Turjanski, P.G.; Salgado Salter, J.D.; Alleva, K.; Carrari, F.; Giovannoni, J.J.; et al. Genome-wide data (ChIP-seq) enabled identification of cell wall-related and aquaporin genes as targets of tomato ASR1, a drought stress-responsive transcription factor. BMC Plant Biol. 2014, 14, 29. [Google Scholar] [CrossRef]
- Arenhart, R.A.; Schunemann, M.; Bucker Neto, L.; Margis, R.; Wang, Z.Y.; Margis-Pinheiro, M. Rice ASR1 and ASR5 are complementary transcription factors regulating aluminium responsive genes. Plant Cell Environ. 2016, 39, 645–651. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Huang, C.; Deng, X.; Zhou, S.; Chen, L.; Li, Y.; Wang, C.; Ma, Z.; Yuan, Q.; Wang, Y.; et al. TaASR1, a transcription factor gene in wheat, confers drought stress tolerance in transgenic tobacco. Plant Cell Environ. 2013, 36, 1449–1464. [Google Scholar] [CrossRef]
- Virlouvet, L.; Jacquemot, M.P.; Gerentes, D.; Corti, H.; Bouton, S.; Gilard, F.; Valot, B.; Trouverie, J.; Tcherkez, G.; Falque, M.; et al. The ZmASR1 protein influences branched-chain amino acid biosynthesis and maintains kernel yield in maize under water-limited conditions. Plant Physiol. 2011, 157, 917–936. [Google Scholar] [CrossRef]
- Joo, J.; Lee, Y.H.; Kim, Y.K.; Nahm, B.H.; Song, S.I. Abiotic stress responsive rice ASR1 and ASR3 exhibit different tissue-dependent sugar and hormone-sensitivities. Mol. Cells 2013, 35, 421–435. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Wei, S.; Chen, J.; Yang, F.; Kong, L.; Chen, C.; Ding, X.; Chu, Z. OsASR2 regulates the expression of a defence-related gene, Os2H16, by targeting the GT-1 cis-element. Plant Biotechnol. J. 2018, 16, 771–783. [Google Scholar] [CrossRef] [PubMed]
- Li, R.H.; Liu, G.B.; Wang, H.; Zheng, Y.Z. Effects of Fe3+ and Zn2+ on the structural and thermodynamic properties of a soybean ASR protein. Biosci. Biotechnol. Biochem. 2013, 77, 475–481. [Google Scholar] [CrossRef]
- Li, J.; Dong, Y.; Li, C.; Pan, Y.; Yu, J. SiASR4, the target gene of SiARDP from Setaria italica, improves abiotic stress adaption in plants. Front. Plant Sci. 2016, 7, 2053. [Google Scholar] [CrossRef]
- Feng, Z.J.; Xu, Z.S.; Sun, J.; Li, L.C.; Chen, M.; Yang, G.X.; He, G.Y.; Ma, Y.Z. Investigation of the ASR family in foxtail millet and the role of ASR1 in drought/oxidative stress tolerance. Plant Cell Rep. 2016, 35, 115–128. [Google Scholar] [CrossRef]
- Wang, L.; Hu, W.; Feng, J.; Yang, X.; Huang, Q.; Xiao, J.; Liu, Y.; Yang, G.; He, G. Identification of the ASR gene family from Brachypodium distachyon and functional characterization of BdASR1 in response to drought stress. Plant Cell Rep. 2016, 35, 1221–1234. [Google Scholar] [CrossRef]
- Zhang, L.; Hu, W.; Wang, Y.; Feng, R.; Zhang, Y.; Liu, J.; Jia, C.; Miao, H.; Zhang, J.; Xu, B.; et al. The MaASR gene as a crucial component in multiple drought stress response pathways in Arabidopsis. Funct. Integr. Genom. 2015, 15, 247–260. [Google Scholar] [CrossRef]
- Tiwari, V.; Chaturvedi, A.K.; Mishra, A.; Jha, B. Introgression of the SbASR-1 gene cloned from a halophyte Salicornia brachiate enhances salinity and drought endurance in transgenic groundnut (Arachis hypogaea)and acts as a transcription factor. PLoS ONE 2015, 10, e0131567. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Tao, J.J.; Chen, H.W.; Li, Q.T.; Zhang, W.K.; Ma, B.; Lin, Q.; Zhang, J.S.; Chen, S.Y. A histone code reader and a transcriptional activator interact to regulate genes for salt tolerance. Plant Physiol. 2017, 175, 1304–1320. [Google Scholar] [CrossRef] [PubMed]
- Golan, I.; Dominguez, P.G.; Konrad, Z.; Shkolnik-Inbar, D.; Carrari, F.; Bar-Zvi, D. Tomato abscisic acid stress ripening (ASR) gene family revisited. PLoS ONE 2014, 9, e107117. [Google Scholar] [CrossRef] [PubMed]
- Dai, J.R.; Liu, B.; Feng, D.R.; Liu, H.Y.; He, Y.M.; Qi, K.B.; Wang, H.B.; Wang, J.F. MpASR encodes an intrinsically unstructured protein and enhances osmotic tolerance in transgenic Arabidopsis. Plant Cell Rep. 2011, 30, 1219–1230. [Google Scholar] [CrossRef] [PubMed]
- Hsu, Y.F.; Yu, S.C.; Yang, C.Y.; Wang, C.S. Lily ASR protein-conferred cold and freezing resistance in Arabidopsis. Plant Physiol. Biochem. 2011, 49, 937–945. [Google Scholar] [CrossRef] [PubMed]
- Cakir, B.; Agasse, A.; Gaillard, C.; Saumonneau, A.; Delrot, S.; Atanassova, R. A grape ASR protein involved in sugar and abscisic acid signaling. Plant Cell 2003, 15, 2165–2180. [Google Scholar] [CrossRef] [PubMed]
- Goldgur, Y.; Rom, S.; Ghirlando, R.; Shkolnik, D.; Shadrin, N.; Konrad, Z.; Bar-Zvi, D. Desiccation and zinc binding induce transition of tomato abscisic acid stress ripening 1, a water stress- and salt stress-regulated plant-specific protein, from unfolded to folded state. Plant Physiol. 2007, 143, 617–628. [Google Scholar] [CrossRef] [PubMed]
- Konrad, Z.; Bar-Zvi, D. Synergism between the chaperone-like activity of the stress regulated ASR1 protein and the osmolyte glycine-betaine. Planta 2008, 227, 1213–1219. [Google Scholar] [CrossRef]
- Dominguez, P.G.; Carrari, F. ASR1 transcription factor and its role in metabolism. Plant Signal. Behav. 2015, 10, e992751. [Google Scholar] [CrossRef]
- Hamdi, K.; Salladini, E.; O’Brien, D.P.; Brier, S.; Chenal, A.; Yacoubi, I.; Longhi, S. Structural disorder and induced folding within two cereal, ABA stress and ripening (ASR) proteins. Sci. Rep. 2017, 7, 15544. [Google Scholar] [CrossRef]
- Arenhart, R.A.; Bai, Y.; de Oliveira, L.F.; Neto, L.B.; Schunemann, M.; Maraschin Fdos, S.; Mariath, J.; Silverio, A.; Sachetto-Martins, G.; Margis, R.; et al. New insights into aluminum tolerance in rice: The ASR5 protein binds the star1 promoter and other aluminum-responsive genes. Mol. Plant 2014, 7, 709–721. [Google Scholar] [CrossRef] [PubMed]
- Wetzler, D.E.; Fuchs Wightman, F.; Bucci, H.A.; Rinaldi, J.; Caramelo, J.J.; Iusem, N.D.; Ricardi, M.M. Conformational plasticity of the intrinsically disordered protein ASR1 modulates its function as a drought stress-responsive gene. PLoS ONE 2018, 13, e0202808. [Google Scholar] [CrossRef]
- Gallego, S.M.; Pena, L.B.; Barcia, R.A.; Azpilicueta, C.E.; Iannone, M.F.; Rosales, E.P.; Zawoznik, M.S.; Groppa, M.D.; Benavides, M.P. Unravelling cadmium toxicity and tolerance in plants: Insight into regulatory mechanisms. Environ. Exp. Bot. 2012, 83, 33–46. [Google Scholar] [CrossRef]
- Kumar, R.; Mustafiz, A.; Sahoo, K.K.; Sharma, V.; Samanta, S.; Sopory, S.K.; Pareek, A.; Singla-Pareek, S.L. Functional screening of cDNA library from a salt tolerant rice genotype Pokkali identifies mannose-1-phosphate guanyl transferase gene (OsMPG1) as a key member of salinity stress response. Plant Mol. Biol. 2012, 79, 555–568. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.S.; Szczypka, M.; Lu, Y.P.; Thiele, D.J.; Rea, P.A. The yeast cadmium factor protein (YCF1) is a vacuolar glutathione S-conjugate pump. J. Biol. Chem. 1996, 271, 6509–6517. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Mo, H.; Sun, W.; Guo, Y.; Li, J. Systematic isolation and characterization of cadmium tolerant genes in tobacco: A cDNA library construction and screening approach. PLoS ONE 2016, 11, e0161147. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Chen, C.; Tan, Z.; Liu, J.; Zhuang, L.; Yang, Z.; Huang, B. Functional identification and characterization of genes cloned from halophyte Seashore paspalum conferring salinity and cadmium tolerance. Front. Plant Sci. 2016, 7, 102. [Google Scholar] [CrossRef]
- Andorf, C.M.; Cannon, E.K.; Portwood, J.L., II; Gardiner, J.M.; Harper, L.C.; Schaeffer, M.L.; Braun, B.L.; Campbell, D.A.; Vinnakota, A.G.; Sribalusu, V.V.; et al. MaizeGDB update: New tools, data and interface for the maize model organism database. Nucleic Acids Res. 2016, 44, D1195–D1201. [Google Scholar] [CrossRef]
- Walley, J.W.; Sartor, R.C.; Shen, Z.; Schmitz, R.J.; Wu, K.J.; Urich, M.A.; Nery, J.R.; Smith, L.G.; Schnable, J.C.; Ecker, J.R.; et al. Integration of omic networks in a developmental atlas of maize. Science 2016, 353, 814–818. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Ochiai, H.; Sugawara, T.; Yamamoto, T. Simultaneous live imaging of the transcription and nuclear position of specific genes. Nucleic Acids Res. 2015, 43, e127. [Google Scholar] [CrossRef] [PubMed]
- Majeran, W.; Friso, G.; Ponnala, L.; Connolly, B.; Huang, M.; Reidel, E.; Zhang, C.; Asakura, Y.; Bhuiyan, N.H.; Sun, Q.; et al. Structural and metabolic transitions of C4 leaf development and differentiation defined by microscopy and quantitative proteomics in maize. Plant Cell 2010, 22, 3509–3542. [Google Scholar] [CrossRef] [PubMed]
- Friso, G.; Majeran, W.; Huang, M.; Sun, Q.; van Wijk, K.J. Reconstruction of metabolic pathways, protein expression, and homeostasis machineries across maize bundle sheath and mesophyll chloroplasts: Large-scale quantitative proteomics using the first maize genome assembly. Plant Physiol. 2010, 152, 1219–1250. [Google Scholar] [CrossRef] [PubMed]
- Kim, I.S.; Kim, Y.S.; Yoon, H.S. Rice ASR1 protein with reactive oxygen species scavenging and chaperone-like activities enhances acquired tolerance to abiotic stresses in Saccharomyces cerevisiae. Mol. Cells 2012, 33, 285–293. [Google Scholar] [CrossRef] [PubMed]
- Rom, S.; Gilad, A.; Kalifa, Y.; Konrad, Z.; Karpasas, M.M.; Goldgur, Y.; Bar-Zvi, D. Mapping the DNA- and zinc-binding domains of ASR1 (abscisic acid stress ripening), an abiotic-stress regulated plant specific protein. Biochimie 2006, 88, 621–628. [Google Scholar] [CrossRef] [PubMed]
- Kharenko, O.A.; Ogawa, M.Y. Metal-induced folding of a designed metalloprotein. J. Inorg. Biochem. 2004, 98, 1971–1974. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Yan, J.; Liu, W.; Liu, L.; Sheng, Y.; Sun, Y.; Li, Y.; Scheller, H.V.; Jiang, M.; Hou, X.; et al. Phosphorylation of a NAC transcription factor by a calcium/calmodulin-dependent protein kinase regulates abscisic acid-induced antioxidant defense in maize. Plant Physiol. 2016, 171, 1651–1664. [Google Scholar] [CrossRef]
- Shim, D.; Hwang, J.U.; Lee, J.; Lee, S.; Choi, Y.; An, G.; Martinoia, E.; Lee, Y. Orthologs of the class A4 heat shock transcription factor HsfA4a confer cadmium tolerance in wheat and rice. Plant Cell 2009, 21, 4031–4043. [Google Scholar] [CrossRef]
- Winzeler, E.A.; Shoemaker, D.D.; Astromoff, A.; Liang, H.; Anderson, K.; Andre, B.; Bangham, R.; Benito, R.; Boeke, J.D.; Bussey, H.; et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 1999, 285, 901–906. [Google Scholar] [CrossRef]
- Chen, X.; Li, J.; Wang, L.; Ma, G.; Zhang, W. A mutagenic study identifying critical residues for the structure and function of rice manganese transporter OsMTP8.1. Sci. Rep. 2016, 6, 32073. [Google Scholar] [CrossRef]
- Yan, J.; Wang, P.; Yang, M.; Lian, X.; Tang, Z.; Huang, C.F.; Salt, D.E.; Zhao, F.J. A loss-of-function allele of OsHMA3 associated with high cadmium accumulation in shoots and grain of Japonica rice cultivars. Plant Cell Environ. 2016, 39, 1941–1954. [Google Scholar] [CrossRef] [PubMed]
- Kunihiro, S.; Saito, T.; Matsuda, T.; Inoue, M.; Kuramata, M.; Taguchi-Shiobara, F.; Youssefian, S.; Berberich, T.; Kusano, T. Rice DEP1, encoding a highly cysteine-rich G protein gamma subunit, confers cadmium tolerance on yeast cells and plants. J. Exp. Bot. 2013, 64, 4517–4527. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.S.; Jiang, J.; Han, X.J.; Zhang, Y.X.; Zhuo, R.Y. Identification, expression analysis of the Hsf family, and characterization of class A4 in Sedum alfredii hance under cadmium stress. Int. J. Mol. Sci. 2018, 19, 1216. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.H.; Yu, G.; Li, J.T.; Jia, P.; Zhang, J.C.; Jia, C.G.; Zhang, Y.H.; Pan, H.Y. A heavy metal-associated protein (AcHMA1) from the halophyte, Atriplex canescens (Pursh) nutt., confers tolerance to iron and other abiotic stresses when expressed in Saccharomyces cerevisiae. Int. J. Mol. Sci. 2014, 15, 14891–14906. [Google Scholar] [CrossRef]
- Cheng, D.; Tan, M.; Yu, H.; Li, L.; Zhu, D.; Chen, Y.; Jiang, M. Comparative analysis of Cd-responsive maize and rice transcriptomes highlights Cd co-modulated orthologs. BMC Genom. 2018, 19, 709. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Qiu, X.; Kang, J.; Wang, Y.; Chen, H.; Huang, J.; Qiu, M.; Zhao, Y.; Kong, G.; Ma, Z.; et al. A phytophthora effector manipulates host histone acetylation and reprograms defense gene expression to promote infection. Curr. Biol. 2017, 27, 981–991. [Google Scholar] [CrossRef]
- Lin, T.; Yang, W.; Lu, W.; Wang, Y.; Qi, X. Transcription factors PvERF15 and PvMTF-1 form a cadmium stress transcriptional pathway. Plant Physiol. 2017, 173, 1565–1573. [Google Scholar] [CrossRef]
- Song, T.; Ma, Z.; Shen, D.; Li, Q.; Li, W.; Su, L.; Ye, T.; Zhang, M.; Wang, Y.; Dou, D. An oomycete CRN effector reprograms expression of plant HSP genes by targeting their promoters. PLoS Pathog. 2015, 11, e1005348. [Google Scholar] [CrossRef]
| Gene | Gene Model * | pYES2-F (5′-3′) | pYES2-R (5′-3′) |
| ZmASR1 | ZM2G136910 | AAGCTT-AATTGTCACTTGCTCTCC | GGATCC-GCTCGATATCACTCTCAC |
| ZmASR2 | ZM5G854138 | AAGCTT-CCAGCCATCCTACTGTCACA | GGTACC-TCAGCCGAAGAGGTGGTGGT |
| ZmASR4 | ZM2G168552 | AAGCTT-TGAGAGCCTGAGACGATGG | GGTACC-TTCAGTCGCAGTAGTAGGAGTA |
| ZmASR5 | ZM2G052100 | AAGCTT-GGAGCCATGTCTGAGGAGAAG | GGTACC-CGACGATGTGCTGCTGCTT |
| pSuper-1300-F (5′-3′) | pSuper-1300-R (5′-3′) | ||
| ZmASR1 | ZM2G136910 | TCTAGA-ATGGCGGAGGAGAAG | GGTACC-GCCGAAGAAGTGGTG |
| ZmASR2 | ZM5G854138 | AAGCTT-CCAGCCATCCTACTGTCACA | GGTACC-GCCGAAGAGGTGGTGGTG |
| ZmASR4 | ZM2G168552 | AAGCTT-TGAGAGCCTGAGACGATGG | GGTACC-GTCGCAGTAGTAGGAGT |
| ZmASR5 | ZM2G052100 | AAGCTT-GGAGCCATGTCTGAGGAGAAG | GGTACC-GTTGTGGGCGTGCTTCTT |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, J.; Zhu, Q.; Yu, H.; Li, L.; Zhang, G.; Chen, X.; Jiang, M.; Tan, M. Comprehensive Analysis of the Cadmium Tolerance of Abscisic Acid-, Stress- and Ripening-Induced Proteins (ASRs) in Maize. Int. J. Mol. Sci. 2019, 20, 133. https://doi.org/10.3390/ijms20010133
Zhang J, Zhu Q, Yu H, Li L, Zhang G, Chen X, Jiang M, Tan M. Comprehensive Analysis of the Cadmium Tolerance of Abscisic Acid-, Stress- and Ripening-Induced Proteins (ASRs) in Maize. International Journal of Molecular Sciences. 2019; 20(1):133. https://doi.org/10.3390/ijms20010133
Chicago/Turabian StyleZhang, Jie, Qiusha Zhu, Haijuan Yu, Liang Li, Guoqiang Zhang, Xi Chen, Mingyi Jiang, and Mingpu Tan. 2019. "Comprehensive Analysis of the Cadmium Tolerance of Abscisic Acid-, Stress- and Ripening-Induced Proteins (ASRs) in Maize" International Journal of Molecular Sciences 20, no. 1: 133. https://doi.org/10.3390/ijms20010133
APA StyleZhang, J., Zhu, Q., Yu, H., Li, L., Zhang, G., Chen, X., Jiang, M., & Tan, M. (2019). Comprehensive Analysis of the Cadmium Tolerance of Abscisic Acid-, Stress- and Ripening-Induced Proteins (ASRs) in Maize. International Journal of Molecular Sciences, 20(1), 133. https://doi.org/10.3390/ijms20010133






