Common microRNA–mRNA Interactions in Different Newcastle Disease Virus-Infected Chicken Embryonic Visceral Tissues
Abstract
1. Introduction
2. Results
2.1. NDV Infection, Clinical Symptoms, and Sample Collection
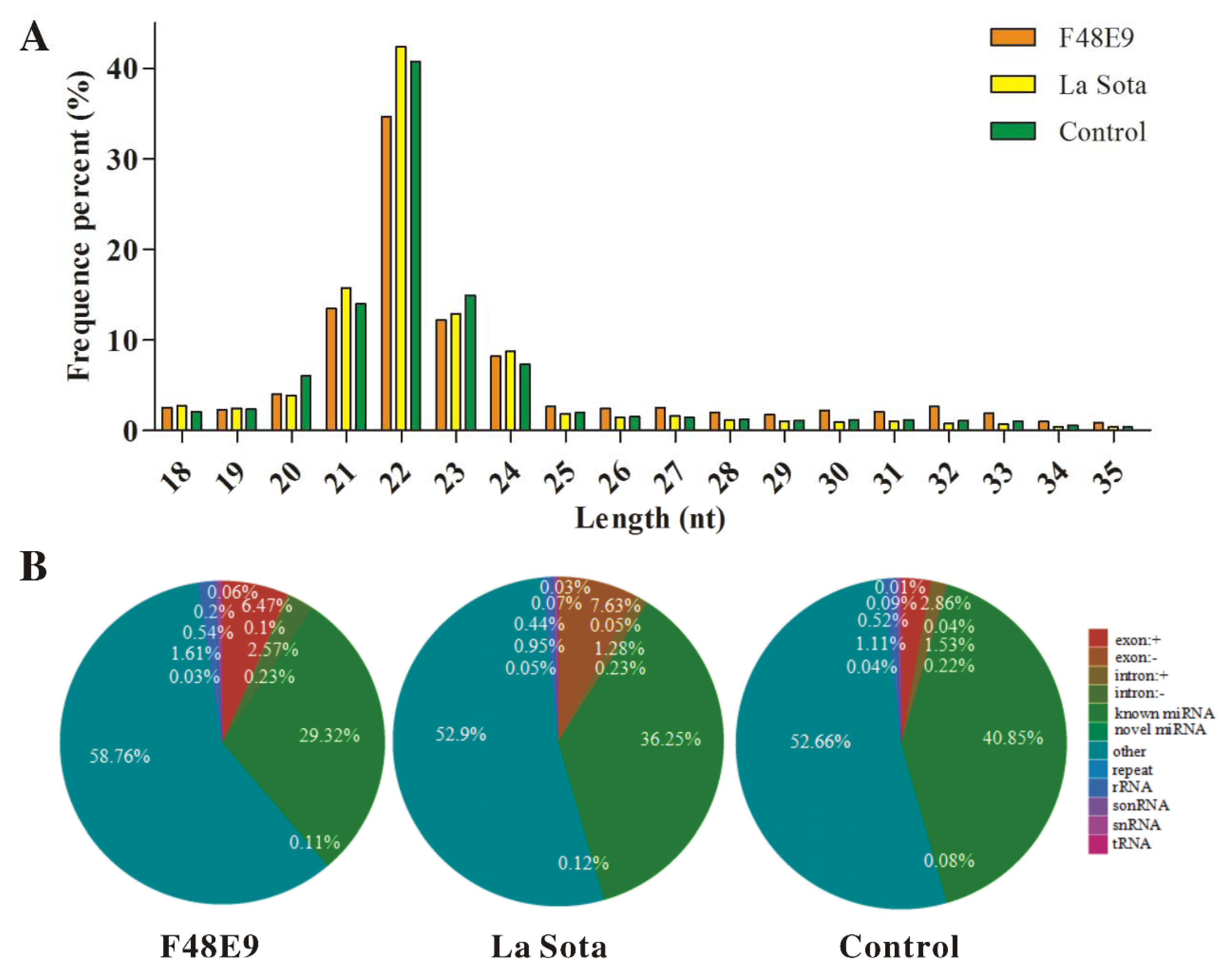
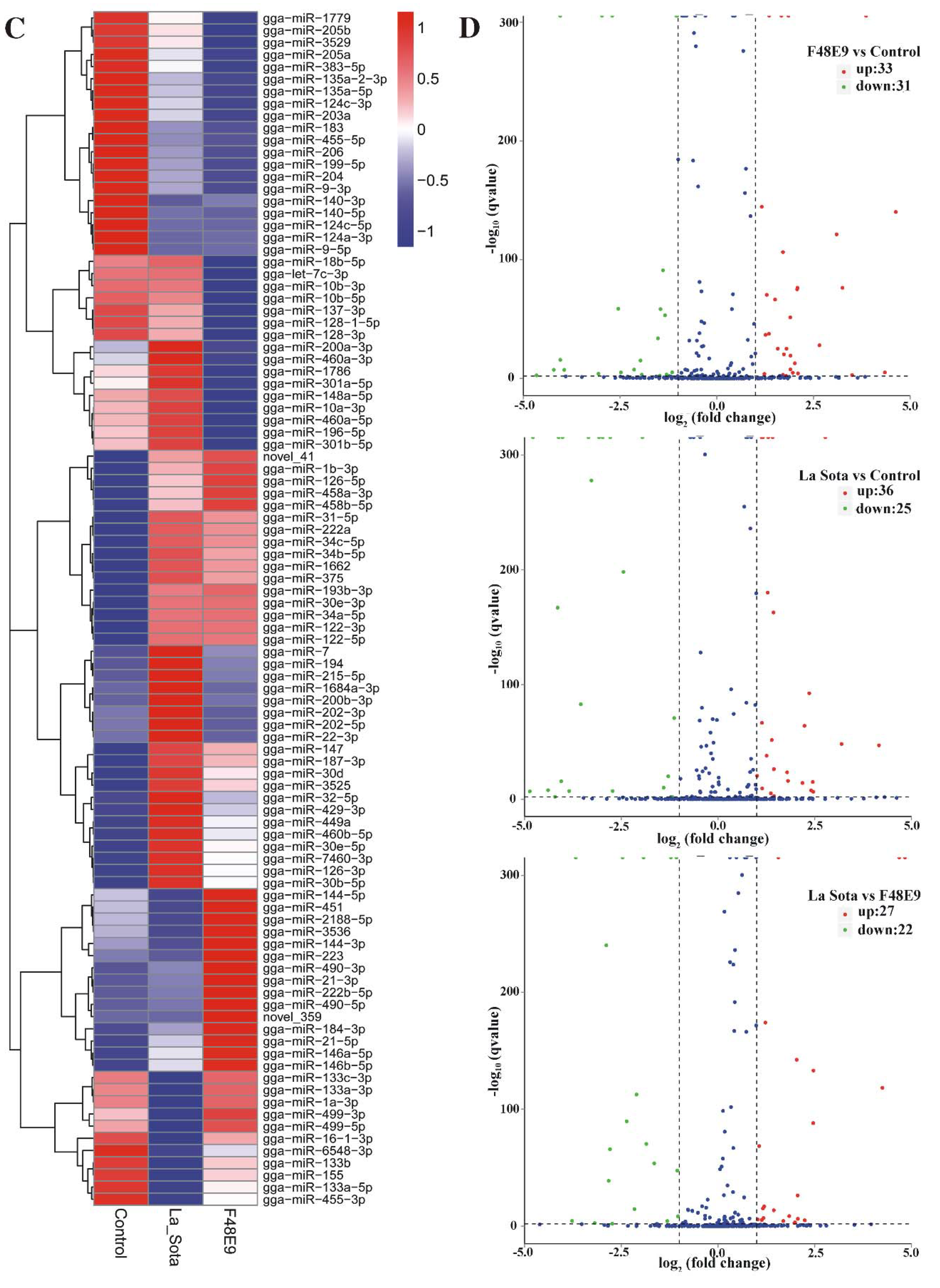
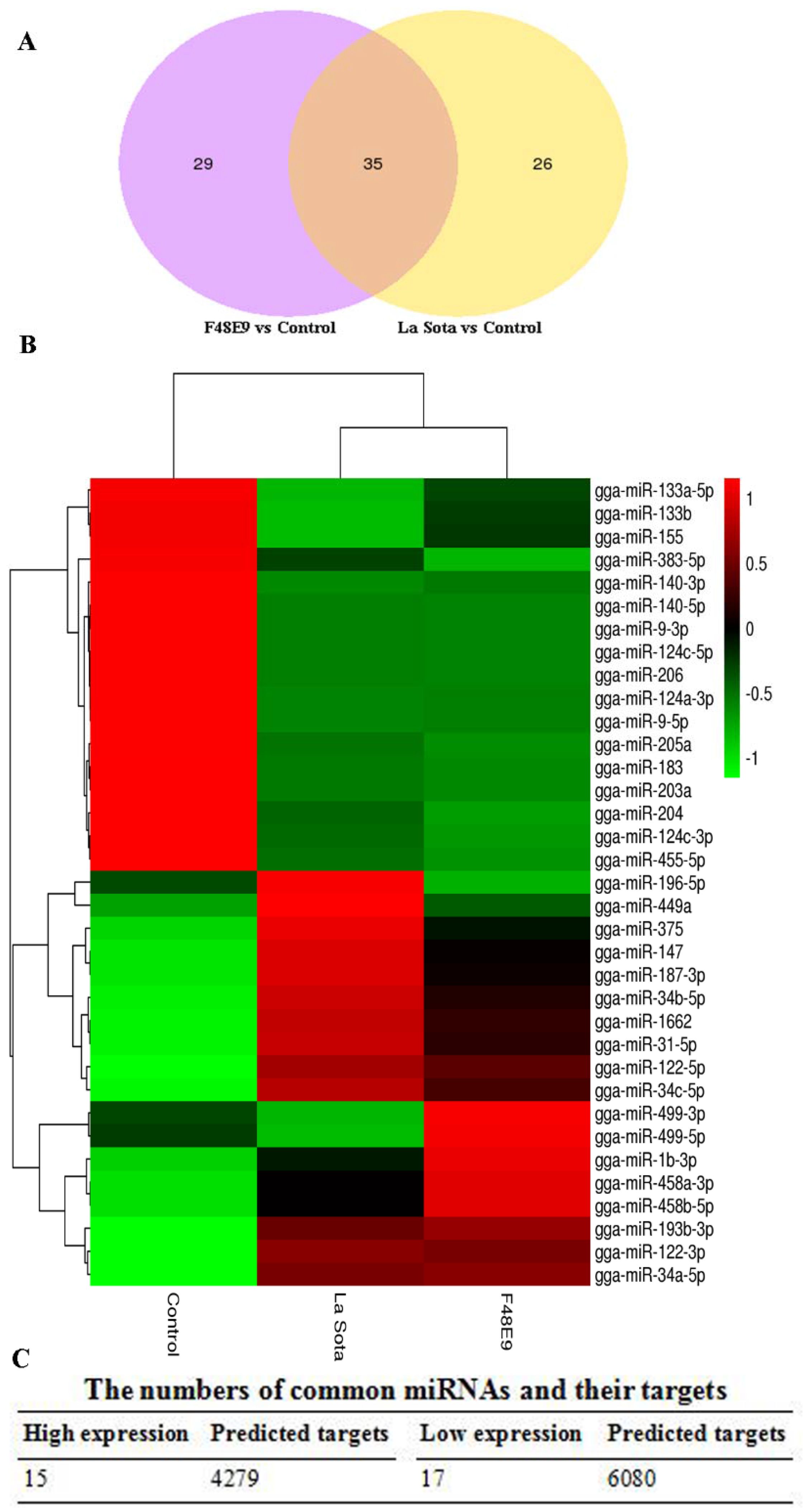
2.2. Global miRNA Expression Pattern in Visceral Tissues from Chicken Embryos
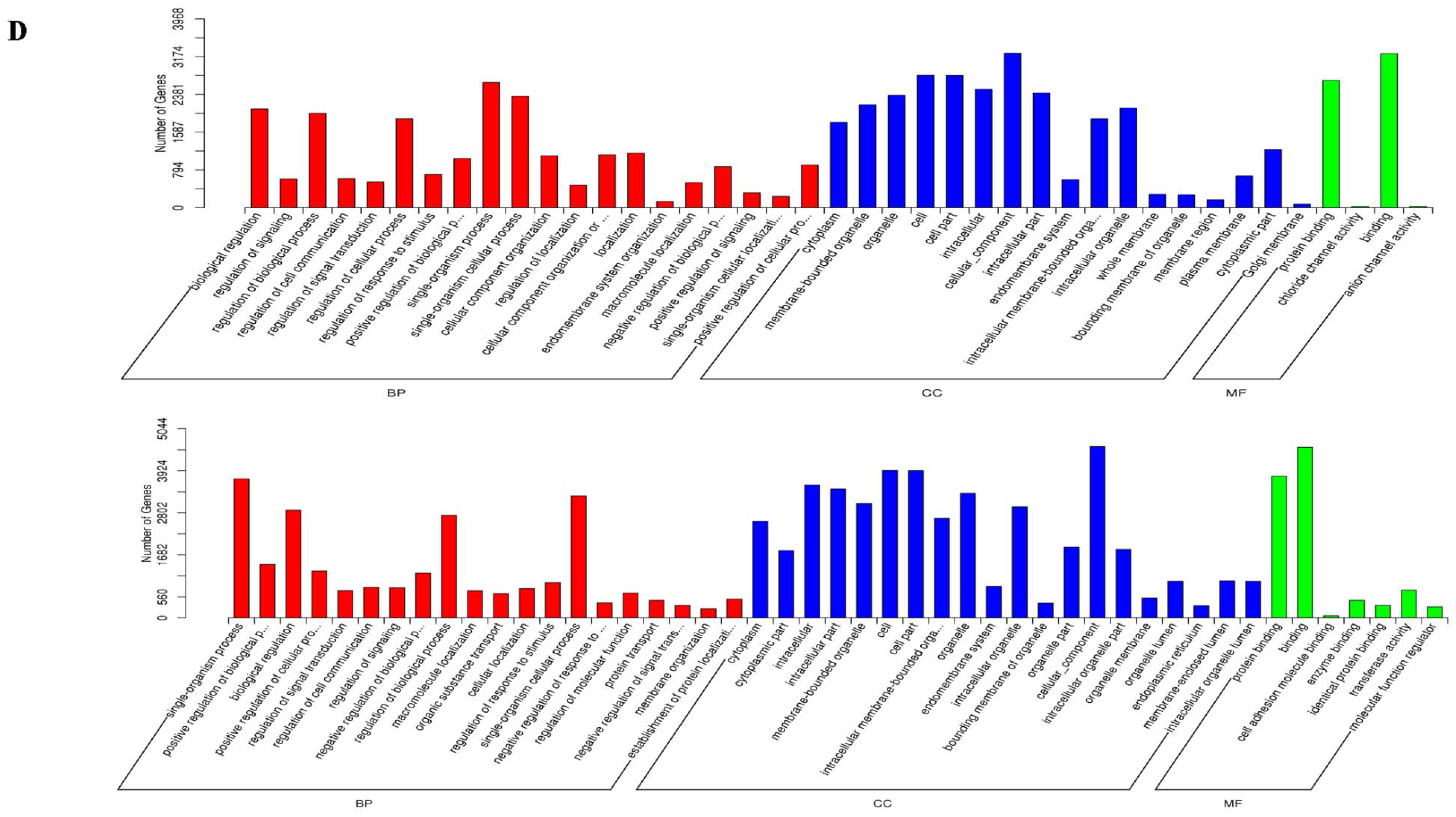
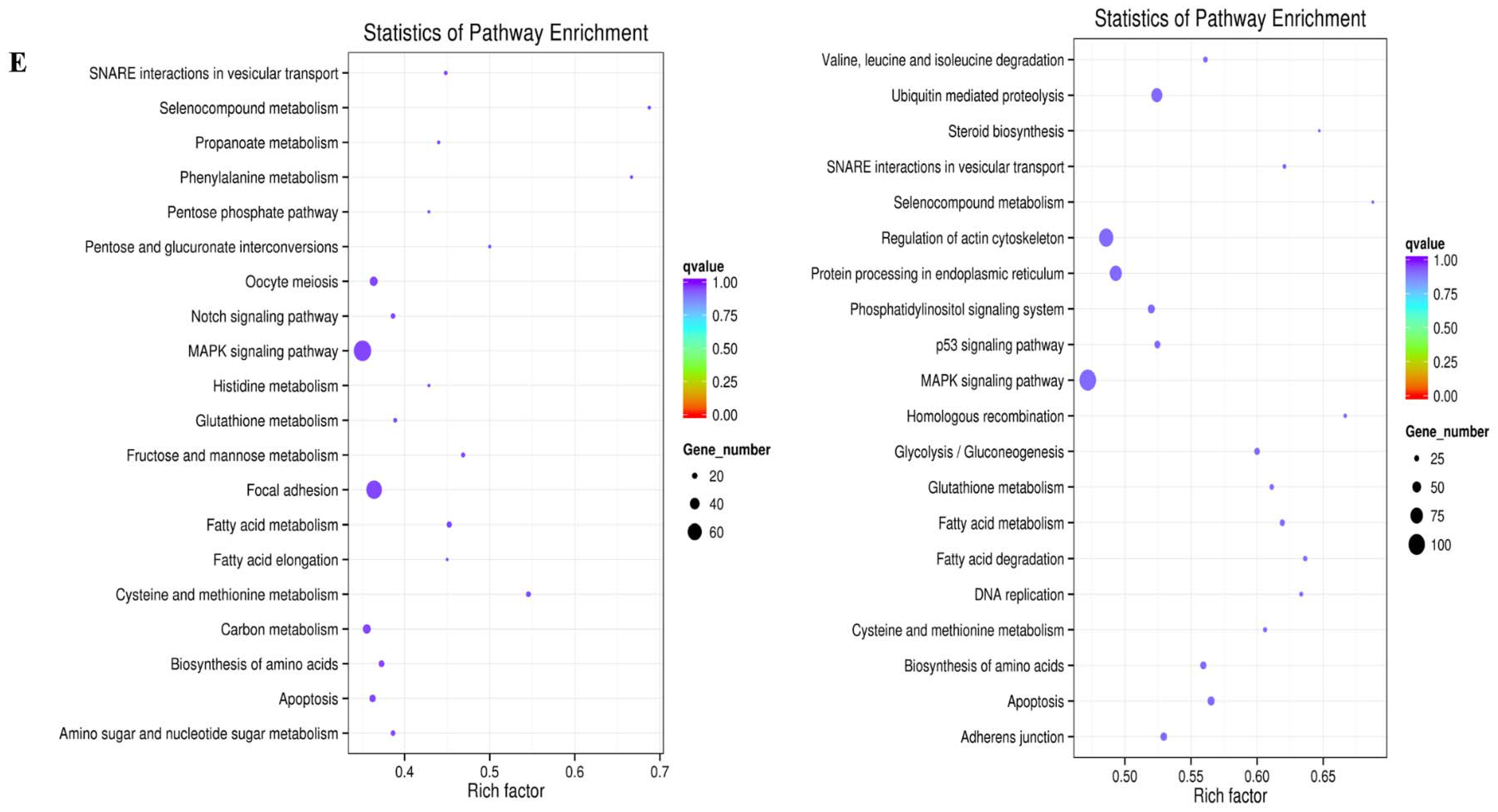
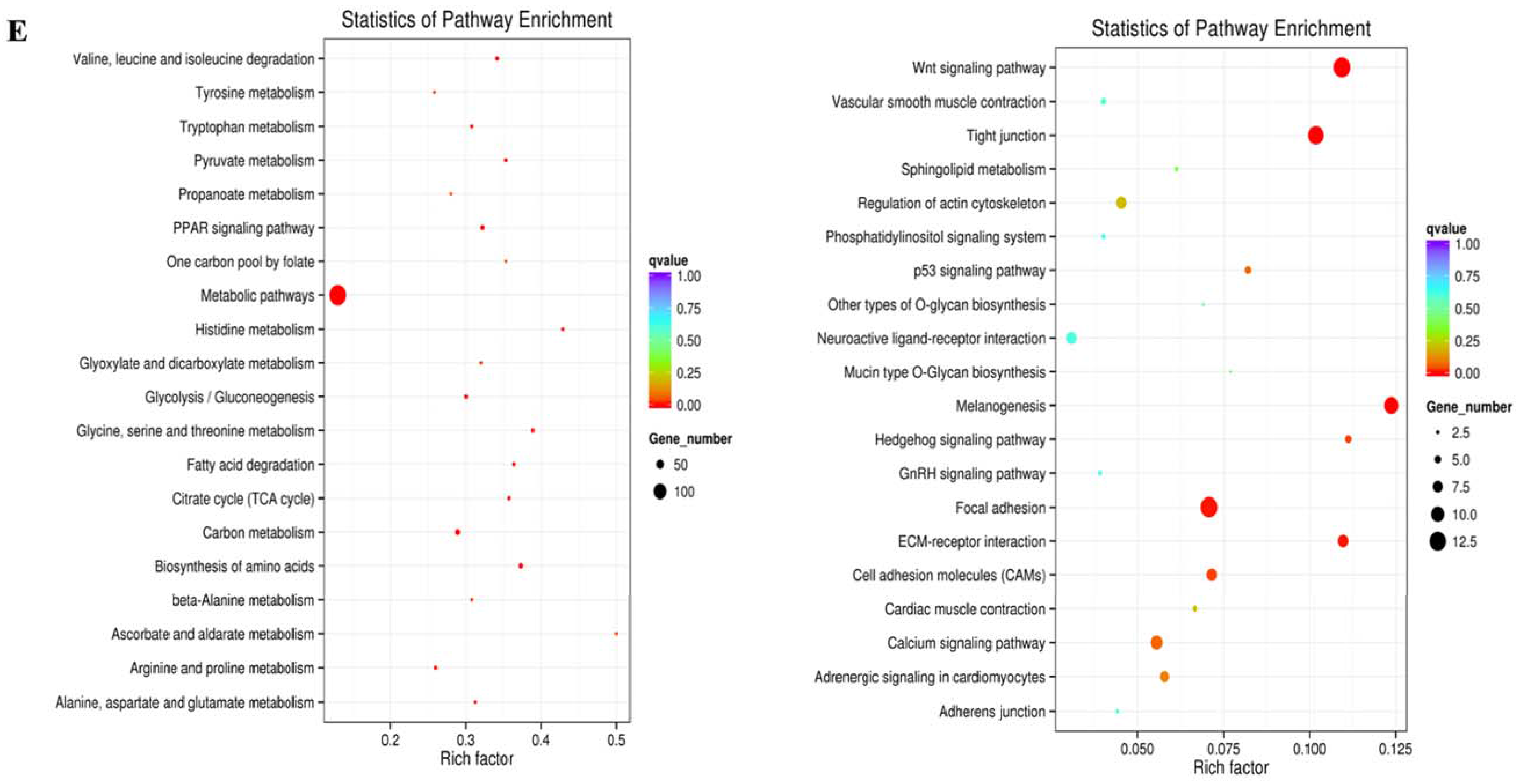
2.3. Prediction and Functional Characterization of Potential Target mRNAs of Common miRNAs during NDV Infection
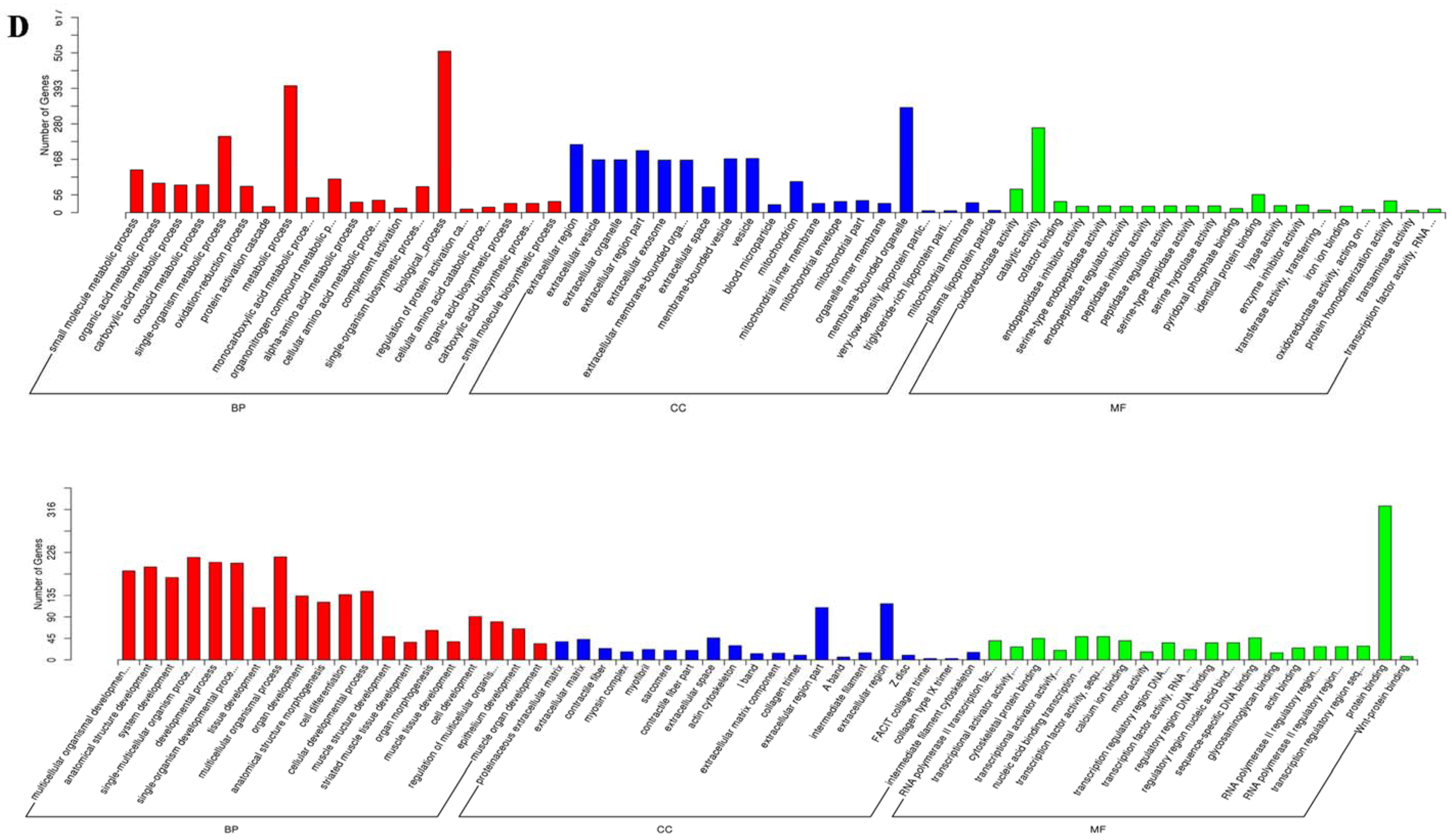
2.4. Global mRNA Expression Pattern in Visceral Tissues from Chicken Embryos during NDV Infection
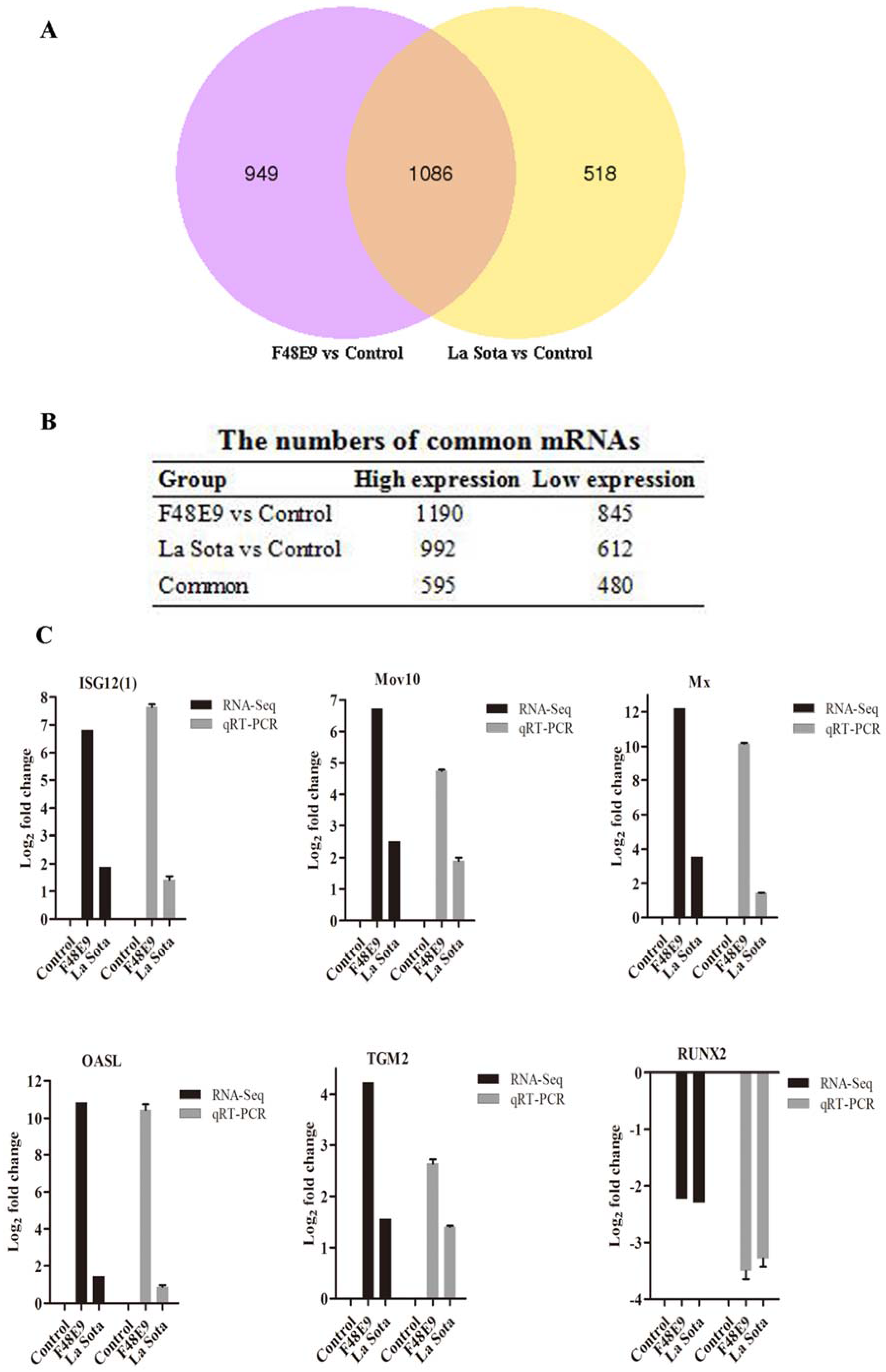
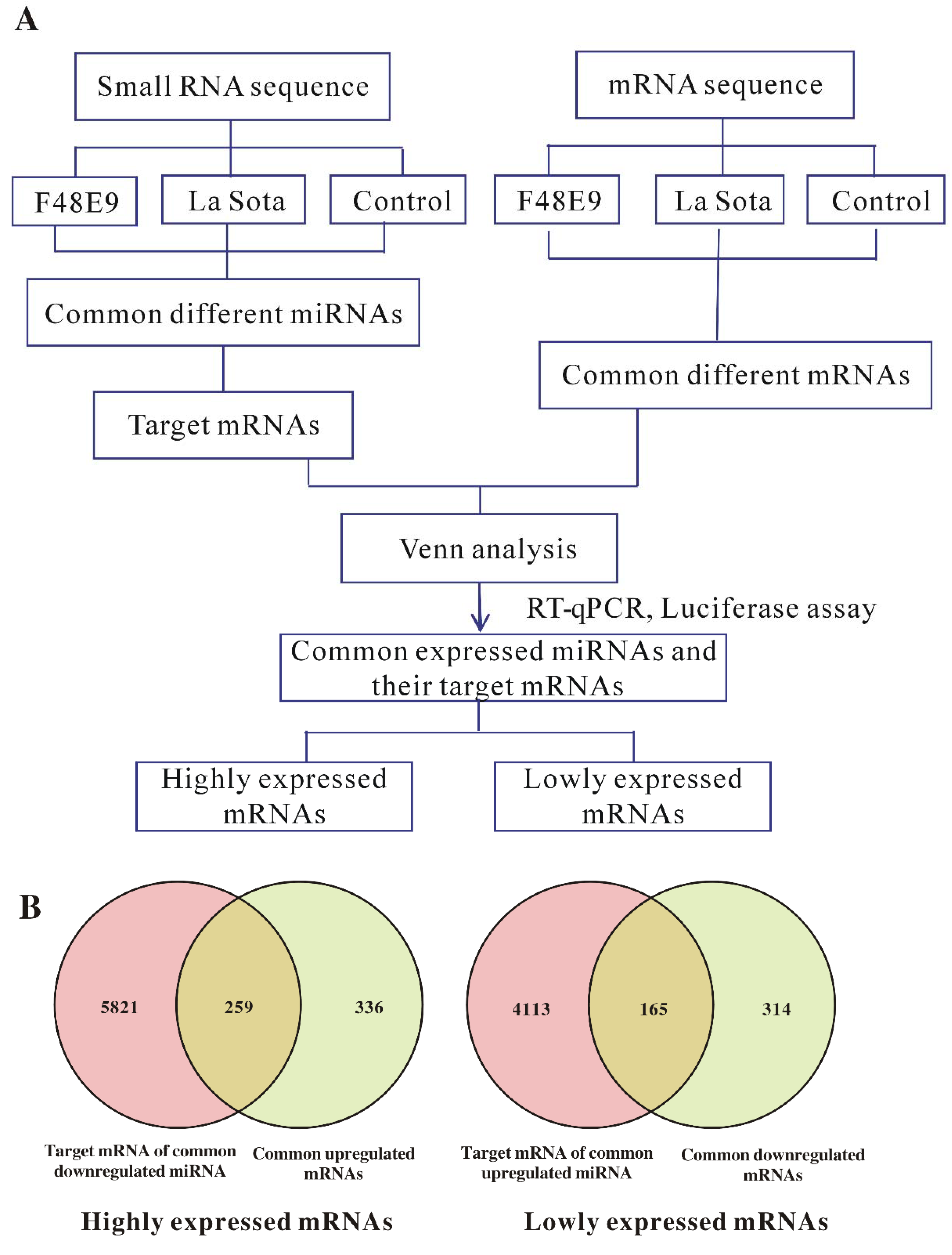
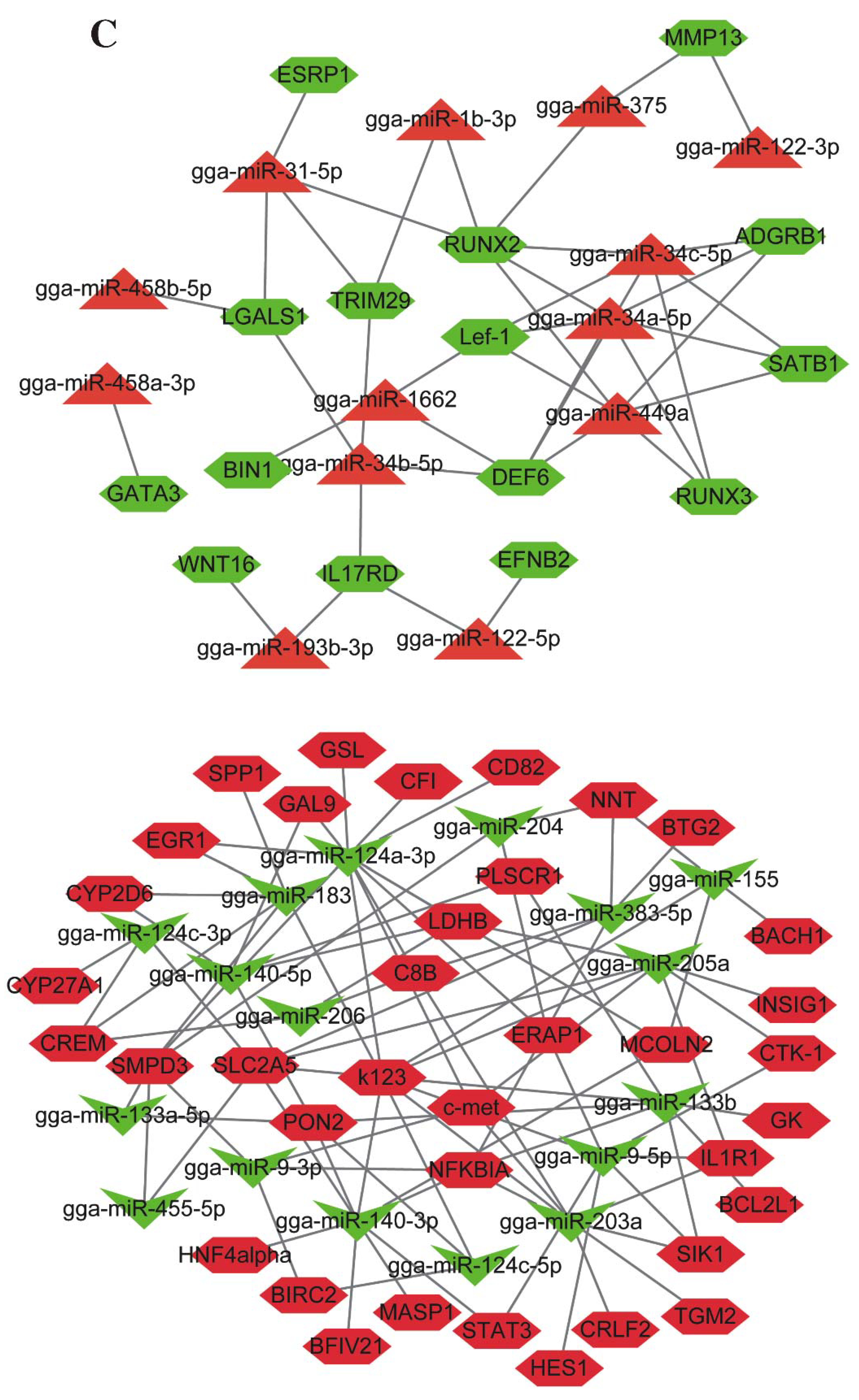
2.5. Conjoint Analysis of Small RNA-Seq and mRNA-Seq
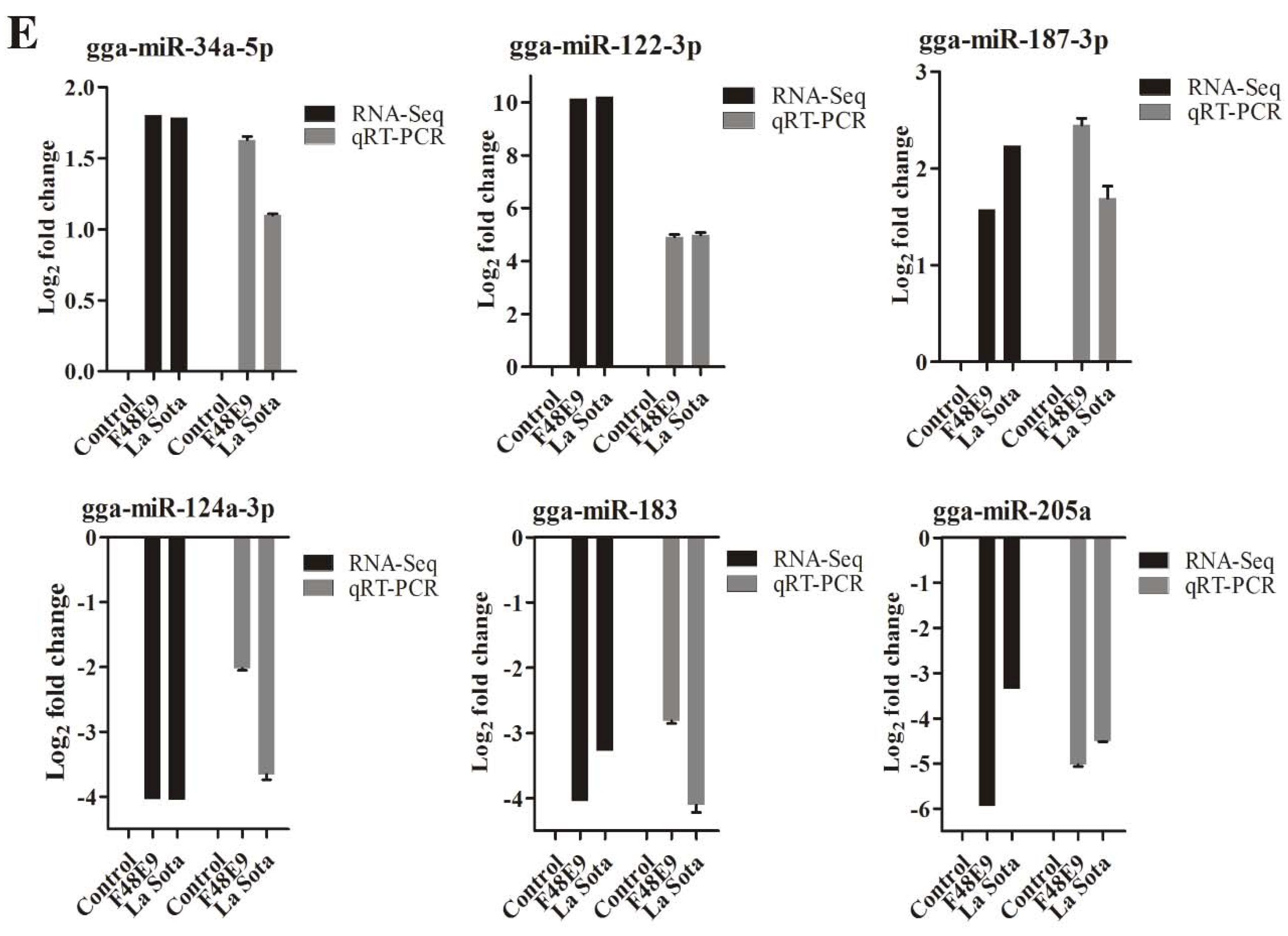
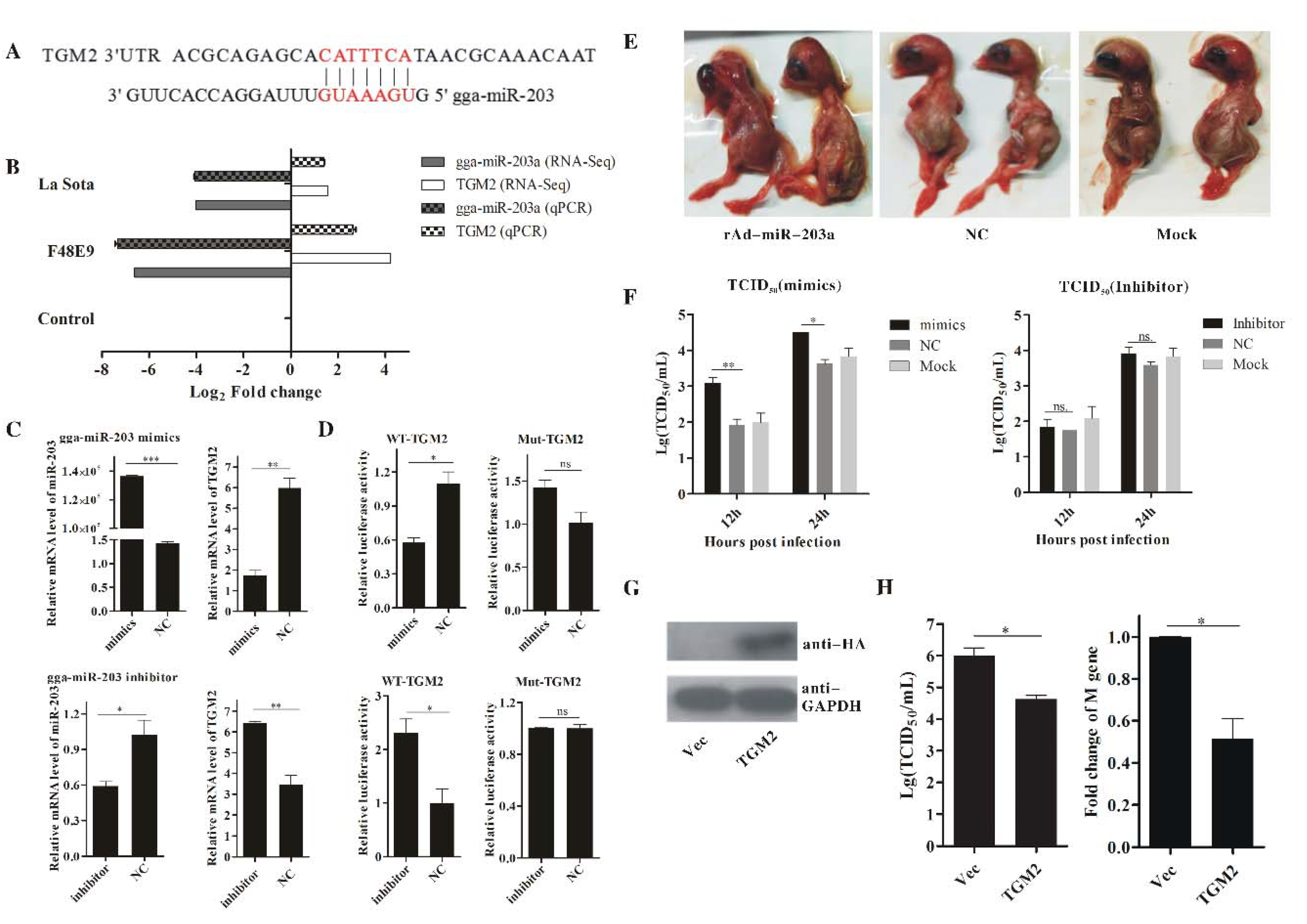
2.6. Validation of the miRNA–mRNA Interactions and Example of a Related Biological Function in NDV Replication
3. Discussion
4. Materials and Methods
4.1. Ethics Statement
4.2. Virus, Cells, and Chicken Embryos
4.3. NDV Infection, Sample Collection, and Preparation
4.4. Library Construction and Deep Sequencing
4.5. Screening for Differentially Expressed miRNAs and mRNAs
4.6. Functional Analysis
4.7. Correlation Analysis of miRNAs and mRNAs
4.8. Real Time qPCR (RT-qPCR)
4.9. Gene Clone and Vector Construction
4.10. Cell Transfection and Luciferase Assay
4.11. Virus Titration
4.12. rAd-miR-203a Construction, Packaging, and Infection
4.13. Western Blotting
4.14. Statistical Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| NDV | Newcastle disease virus |
| miRNA | microRNA |
| 3′UTR | 3-untranslated region |
| AIV | Avian influenza virus |
| IBDV | Infectious Bursal disease virus |
| MDV | Marek’s disease virus |
| RIG-I | Retinoic acid inducible-gene I |
| HCV | Hepatitis C virus |
| CEFs | Chicken embryonic fibroblasts |
| SPF | Specific pathogen-free |
| HA | Hemagglutinin |
| GO | Gene Ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| MAPK | Mitogen-activated protein kinase |
| CSRP3 | Cysteine- and glycine-rich protein 3 |
| MYL2 | Myosin light chain 2 |
| MYBPC3 | Myosin binding protein C, cardiac |
| MME | Membrane metalloendopeptidase |
| POPDC2 | Popeye domain containing 2 |
| BVES | Blood vessel epicardial substance |
| Gadd45 | Growth arrest and DNA damage-inducible 45 |
| Mov10 | Mov10 RISC complex RNA helicase |
| Mx | Interferon-induced GTP-binding protein Mx |
| OASL | 2′-5′-oligoadenylate synthetase like |
| RUNX2 | Runt-related transcription factor 2 |
| ISG12(1) | Interferon alpha inducible protein 27 like 1 (IFI27L1) |
| RT-qPCR | Real-time quantitative polymerase chain reaction |
| IRF2 | Interferon regulatory factor 2 |
| TWIST1 | Twist family bHLH transcription factor 1 |
| BCL-2 | B cell leukemia/lymphoma 2 |
| DEGs | Differentially expressed genes |
| OPN | Osteopontin |
| TLR9 | Toll-like receptor 9 |
| STAT5 | Signal transducer and activator of transcription 5 |
| IAP | Inhibitor of apoptosis protein |
| NR5A2 | Nuclear receptor subfamily 5 group A member 2 |
| HNF1B | HNF1 homeobox B |
| FOXF1 | Forkhead box F1 |
| GATA4 | GATA binding protein 4 |
| VEGFA | Vascular endothelial growth factor A |
| PROX1 | Prospero homeobox 1 |
| TNFR2 | TNF receptor 2 |
| EBV | Epstein–Barr virus |
| IL-12 | Interleukin-12 |
| TGM2 | Transglutaminase 2 |
| TGF-β | Transforming growth factor-β |
| DMEM | Dulbecco’s modified Eagle’s medium |
| PFU | Plaque-forming unit |
| NC | Negative control |
| TCID50 | 50% tissue culture infection doses |
| NOD | Nucleotide.bindingoligomerization domain |
| Jak-STAT | Janus kinase-Signal transducers and activators of transcription |
| PPAR | Peroxisome proliferators-activated receptor |
| NF-κB | Nuclear factor κ-light-chain-enhancer of activated B cells |
| IκB | Inhibitor of NF-κB |
| PMSF | Phenylmethanesulfonyl fluoride |
References
- Chen, S.; Hao, H.; Liu, Q.; Wang, R.; Zhang, P.; Wang, X.; Du, E.; Yang, Z. Phylogenetic and pathogenic analyses of two virulent Newcastle disease viruses isolated from Crested Ibis (Nipponia nippon) in China. Virus Genes 2013, 46, 447–453. [Google Scholar] [CrossRef] [PubMed]
- Ganar, K.; Das, M.; Sinha, S.; Kumar, S. Newcastle disease virus: Current status and our understanding. Virus Res. 2014, 184, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Brown, V.R.; Bevins, S.N. A review of virulent Newcastle disease viruses in the United States and the role of wild birds in viral persistence and spread. Vet. Res. 2017, 48, 68. [Google Scholar] [CrossRef] [PubMed]
- Miller, P.J.; Kim, L.M.; Ip, H.S.; Afonso, C.L. Evolutionary dynamics of Newcastle disease virus. Virology 2009, 391, 64–72. [Google Scholar] [CrossRef] [PubMed]
- Dortmans, J.C.; Koch, G.; Rottier, P.J.; Peeters, B.P. Virulence of Newcastle disease virus: What is known so far? Vet. Res. 2011, 42, 122. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.S.; Kye, S.J.; Kim, J.Y.; To, T.L.; Nguyen, D.T.; Lee, Y.J.; Choi, J.G.; Kang, H.M.; Kim, K.I.; Song, B.M.; et al. Molecular epidemiology of Newcastle disease viruses in Vietnam. Trop. Anim. Health Prod. 2014, 46, 271–277. [Google Scholar] [CrossRef] [PubMed]
- Capraro, G.A.; Johnson, J.B.; Kock, N.D.; Parks, G.D. Virus growth and antibody responses following respiratory tract infection of ferrets and mice with WT and P/V mutants of the paramyxovirus Simian Virus 5. Virology 2008, 376, 416–428. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Fu, Q.; Meng, C.; Yu, S.; Zhan, Y.; Dong, L.; Song, C.; Sun, Y.; Tan, L.; Hu, S.; et al. Newcastle Disease Virus V Protein Targets Phosphorylated STAT1 to Block IFN-I Signaling. PLoS ONE 2016, 11, e0148560. [Google Scholar] [CrossRef] [PubMed]
- Rue, C.A.; Susta, L.; Cornax, I.; Brown, C.C.; Kapczynski, D.R.; Suarez, D.L.; King, D.J.; Miller, P.J.; Afonso, C.L. Virulent Newcastle disease virus elicits a strong innate immune response in chickens. J. Gen.Virol. 2011, 92 Pt 4, 931–939. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Li, H.; Xie, Q.; Shang, H.; Ji, J.; Bai, S.; Cao, Y.; Ma, Y.; Bi, Y. Transcriptional profiling of host gene expression in chicken liver tissues infected with oncogenic Marek’s disease virus. J. Gen. Virol. 2011, 92 Pt 12, 2724–2733. [Google Scholar] [CrossRef] [PubMed]
- Glennon, N.B.; Jabado, O.; Lo, M.K.; Shaw, M.L. Transcriptome Profiling of the Virus-Induced Innate Immune Response in Pteropus vampyrus and Its Attenuation by Nipah Virus Interferon Antagonist Functions. J. Virol. 2015, 89, 7550–7566. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Mardis, E.R. The impact of next-generation sequencing technology on genetics. Trends Genet. TIG 2008, 24, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef] [PubMed]
- Bruscella, P.; Bottini, S.; Baudesson, C.; Pawlotsky, J.M.; Feray, C.; Trabucchi, M. Viruses and miRNAs: More Friends than Foes. Front. Microbiol. 2017, 8, 824. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Du, G.; Wang, Y.; Shi, L.; Mi, J.; Tang, G. Integrative analysis of mRNA and miRNA expression profiles in oral lichen planus: Preliminary results. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. 2017, 124, 390–402. [Google Scholar] [CrossRef] [PubMed]
- Krutzfeldt, J.; Stoffel, M. MicroRNAs: A new class of regulatory genes affecting metabolism. Cell Metab. 2006, 4, 9–12. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Lin, J.; Xia, J.; Chen, Y.T.; Zhang, K.Y.; Zeng, Y.; Yang, Q. H9N2 avian influenza virus enhances the immune responses of BMDCs by down-regulating miR29c. Vaccine 2017, 35, 729–737. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, W.; Wang, Y.S.; Du, X.N.; Liu, H.J.; Zhang, H.B. gga-miR-9* inhibits IFN production in antiviral innate immunity by targeting interferon regulatory factor 2 to promote IBDV replication. Vet. Microbiol. 2015, 178, 41–49. [Google Scholar] [CrossRef] [PubMed]
- Chi, J.Q.; Teng, M.; Yu, Z.H.; Xu, H.; Su, J.W.; Zhao, P.; Xing, G.X.; Liang, H.D.; Deng, R.G.; Qu, L.H.; et al. Marek’s disease virus-encoded analog of microRNA-155 activates the oncogene c-Myc by targeting LTBP1 and suppressing the TGF-β signaling pathway. Virology 2015, 476, 72–84. [Google Scholar] [CrossRef] [PubMed]
- Conrad, K.D.; Giering, F.; Erfurth, C.; Neumann, A.; Fehr, C.; Meister, G.; Niepmann, M. microRNA-122 Dependent Binding of Ago2 Protein to Hepatitis C Virus RNA Is Associated with Enhanced RNA Stability and Translation Stimulation. PLoS ONE 2013, 8, e56272. [Google Scholar] [CrossRef] [PubMed]
- Trobaugh, D.W.; Klimstra, W.B. MicroRNA Regulation of RNA Virus Replication and Pathogenesis. Trends Mol. Med. 2017, 23, 80–93. [Google Scholar] [CrossRef] [PubMed]
- Thibault, P.A.; Huys, A.; Amador-Canizares, Y.; Gailius, J.E.; Pinel, D.E.; Wilson, J.A. Regulation of Hepatitis C Virus Genome Replication by Xrn1 and MicroRNA-122 Binding to Individual Sites in the 5’ Untranslated Region. J. Virol. 2015, 89, 6294–6311. [Google Scholar] [CrossRef] [PubMed]
- Ingle, H.; Kumar, S.; Raut, A.A.; Mishra, A.; Kulkarni, D.D.; Kameyama, T.; Takaoka, A.; Akira, S.; Kumar, H. The microRNA miR-485 targets host and influenza virus transcripts to regulate antiviral immunity and restrict viral replication. Sci. Signal 2015, 8, 406. [Google Scholar] [CrossRef] [PubMed]
- Bandiera, S.; Pernot, S.; El Saghire, H.; Durand, S.C.; Thumann, C.; Crouchet, E.; Ye, T.; Fofana, I.; Oudot, M.A.; Barths, J.; et al. Hepatitis C Virus-Induced Upregulation of MicroRNA miR-146a-5p in Hepatocytes Promotes Viral Infection and Deregulates Metabolic Pathways Associated with Liver Disease Pathogenesis. J. Virol. 2016, 90, 6387–6400. [Google Scholar] [CrossRef] [PubMed]
- Munir, S.; Sharma, J.M.; Kapur, V. Transcriptional response of avian cells to infection with Newcastle disease virus. Virus Res. 2005, 107, 103–108. [Google Scholar] [CrossRef] [PubMed]
- Balasubramaniam, V.R.; Wai, T.H.; Omar, A.R.; Othman, I.; Hassan, S.S. Cellular transcripts of chicken brain tissues in response to H5N1 and Newcastle disease virus infection. Virol. J. 2012, 9, 53. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.; Hu, J.; Hu, S.; Song, Q.; Ding, P.; Zhu, J.; Liu, X.; Wang, X.; Liu, X. High levels of virus replication and an intense inflammatory response contribute to the severe pathology in lymphoid tissues caused by Newcastle disease virus genotype VIId. Arch. Virol. 2015, 160, 639–648. [Google Scholar] [CrossRef] [PubMed]
- Krutzfeldt, J.; Poy, M.N.; Stoffel, M. Strategies to determine the biological function of microRNAs. Nat. Genet. 2006, 38, S14–S19. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Xie, Y.; Cao, H.; Wang, H. Common microRNA-mRNA interactions exist among distinct porcine iPSC lines independent of their metastable pluripotent states. Cell Death Dis. 2017, 8, e3027. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Jing, J.; Li, X.; Wang, J.; Feng, X.; Cao, R.; Chen, P. Integration analysis of miRNA and mRNA expression profiles in swine testis cells infected with Japanese encephalitis virus. Infect. Genet. Evol. J. Mol. Epidemiol. Evolut. Genet. Infecti. Dis. 2015, 32, 342–347. [Google Scholar] [CrossRef] [PubMed]
- Fiorucci, G.; Chiantore, M.V.; Mangino, G.; Romeo, G. MicroRNAs in virus-induced tumorigenesis and IFN system. Cytokine Growth Factor Rev. 2015, 26, 183–194. [Google Scholar] [CrossRef] [PubMed]
- Thakar, J.; Hartmann, B.M.; Marjanovic, N.; Sealfon, S.C.; Kleinstein, S.H. Comparative analysis of anti-viral transcriptomics reveals novel effects of influenza immune antagonism. BMC Immunol. 2015, 16, 46. [Google Scholar] [CrossRef] [PubMed]
- Trejo-Avila, L.M.; Elizondo-Gonzalez, R.; Rodriguez-Santillan, P.; Aguilar-Briseno, J.A.; Ricque-Marie, D.; Rodriguez-Padilla, C.; Cruz-Suarez, L.E. Innocuity and anti-Newcastle-virus-activity of Cladosiphon okamuranus fucoidan in chicken embryos. Poult. Sci. 2016, 95, 2795–2802. [Google Scholar] [CrossRef] [PubMed]
- Bolha, L.; Bencina, D.; Cizelj, I.; Oven, I.; Slavec, B.; Rojs, O.Z.; Narat, M. Effect of Mycoplasma synoviae and lentogenic Newcastle disease virus coinfection on cytokine and chemokine gene expression in chicken embryos. Poul. Sci. 2013, 92, 3134–3143. [Google Scholar] [CrossRef] [PubMed]
- Primo, M.N.; Bak, R.O.; Schibler, B.; Mikkelsen, J.G. Regulation of pro-inflammatory cytokines TNFα and IL24 by microRNA-203 in primary keratinocytes. Cytokine 2012, 60, 741–748. [Google Scholar] [CrossRef] [PubMed]
- Miao, L.; Liu, K.; Xie, M.; Xing, Y.; Xi, T. miR-375 inhibits Helicobacter pylori-induced gastric carcinogenesis by blocking JAK2-STAT3 signaling. Cancer Immunol. Immunother. CII 2014, 63, 699–711. [Google Scholar] [CrossRef] [PubMed]
- Shinohara, M.L.; Lu, L.; Bu, J.; Werneck, M.B.; Kobayashi, K.S.; Glimcher, L.H.; Cantor, H. Osteopontin expression is essential for interferon-α production by plasmacytoid dendritic cells. Nat. Immunol. 2006, 7, 498–506. [Google Scholar] [CrossRef] [PubMed]
- Rani, A.; Murphy, J.J. STAT5 in Cancer and Immunity. J. Interferon Cytokine Res. 2016, 36, 226–237. [Google Scholar] [CrossRef] [PubMed]
- Villarino, A.; Laurence, A.; Robinson, G.W.; Bonelli, M.; Dema, B.; Afzali, B.; Shih, H.Y.; Sun, H.W.; Brooks, S.R.; Hennighausen, L.; et al. Signal transducer and activator of transcription 5 (STAT5) paralog dose governs T cell effector and regulatory functions. eLife 2016, 5, e08384. [Google Scholar] [CrossRef] [PubMed]
- Beug, S.T.; Cheung, H.H.; LaCasse, E.C.; Korneluk, R.G. Modulation of immune signalling by inhibitors of apoptosis. Trends Immunol. 2012, 33, 535–545. [Google Scholar] [CrossRef] [PubMed]
- Anderton, H.; Rickard, J.A.; Varigos, G.A.; Lalaoui, N.; Silke, J. Inhibitor of Apoptosis Proteins (IAPs) Limit RIPK1-Mediated Skin Inflammation. J. Investig. Dermatol. 2017, 137, 2371–2379. [Google Scholar] [CrossRef] [PubMed]
- Mellett, M.; Atzei, P.; Bergin, R.; Horgan, A.; Floss, T.; Wurst, W.; Callanan, J.J.; Moynagh, P.N. Orphan receptor IL-17RD regulates Toll-like receptor signalling via SEFIR/TIR interactions. Nat. Commun. 2015, 6, 6669. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Wang, Y.; Mei, K.; Zhang, S.; Sun, X.; Ren, F.; Liu, S.; Yang, Z.; Wang, X.; Qin, Z.; et al. Tumor necrosis factor receptor 2 (TNFR2).interleukin-17 receptor D (IL-17RD) heteromerization reveals a novel mechanism for NF-κB activation. J. Biol. Chem. 2015, 290, 861–871. [Google Scholar] [CrossRef] [PubMed]
- Moriya, N.; Shibasaki, S.; Karasaki, M.; Iwasaki, T. The Impact of MicroRNA-223-3p on IL-17 Receptor D Expression in Synovial Cells. PLoS ONE 2017, 12, e0169702. [Google Scholar] [CrossRef] [PubMed]
- Chang, Z.; Wang, Y.; Bian, L.; Liu, Q.; Long, J.E. Enterovirus 71 antagonizes the antiviral activity of host STAT3 and IL-6R with partial dependence on virus-induced miR-124. J. Gen. Virol. 2017, 98, 3008–3025. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.; Luo, D.; Wang, D.; Ma, L.; Zhao, Y.; Li, L. IL-17 Activates the IL-6/STAT3 Signal Pathway in the Proliferation of Hepatitis B Virus-Related Hepatocellular Carcinoma. Cell. Physiol. Biochem. 2017, 43, 2379–2390. [Google Scholar] [CrossRef] [PubMed]
- Skinner, C.M.; Ivanov, N.S.; Barr, S.A.; Chen, Y.; Skalsky, R.L. An Epstein-Barr Virus MicroRNA Blocks Interleukin-1 (IL-1) Signaling by Targeting IL-1 Receptor 1. J. Virol. 2017, 91, e00530-17. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; Chen, J.; Zhou, L.; Chen, W.; Ding, G.; Cao, L. Pancreatic cancer derived exosomes regulate the expression of TLR4 in dendritic cells via miR-203. Cell. Immunol. 2014, 292, 65–69. [Google Scholar] [CrossRef] [PubMed]
- Tovar-Vidales, T.; Clark, A.F.; Wordinger, R.J. Focus on molecules: Transglutaminase 2. Exp. Eye Res. 2011, 93, 2–3. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Hong, J.M.; Jeong, E.M.; Lee, W.J.; Kim, H.R.; Kang, J.S.; Kim, I.G.; Hwang, Y.I. Lack of transglutaminase 2 diminished T-cell responses in mice. Immunology 2014, 142, 506–516. [Google Scholar] [CrossRef] [PubMed]
- Klock, C.; Diraimondo, T.R.; Khosla, C. Role of transglutaminase 2 in celiac disease pathogenesis. Semin. Immunopathol. 2012, 34, 513–522. [Google Scholar] [CrossRef] [PubMed]
- Nunes, I.; Gleizes, P.E.; Metz, C.N.; Rifkin, D.B. Latent transforming growth factor-β binding protein domains involved in activation and transglutaminase-dependent cross-linking of latent transforming growth factor-β. J. Cell Biol. 1997, 136, 1151–1163. [Google Scholar] [CrossRef] [PubMed]
- Tkach, M.; Thery, C. Communication by Extracellular Vesicles: Where We Are and Where We Need to Go. Cell 2016, 164, 1226–1232. [Google Scholar] [CrossRef] [PubMed]
- Holm, C.K.; Rahbek, S.H.; Gad, H.H.; Bak, R.O.; Jakobsen, M.R.; Jiang, Z.; Hansen, A.L.; Jensen, S.K.; Sun, C.; Thomsen, M.K.; et al. Influenza A virus targets a cGAS-independent STING pathway that controls enveloped RNA viruses. Nat. Commun. 2016, 7, 10680. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.K.; Feng, Z.X.; Wang, X.; Wang, X.W.; Zhang, X.G. DEGseq: An R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 2010, 26, 136–138. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Mao, X.; Huang, J.; Ding, Y.; Wu, J.; Dong, S.; Kong, L.; Gao, G.; Li, C.Y.; Wei, L. KOBAS 2.0: A web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 2011, 39, W316–W322. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.L.; Li, Y.F.; Qi, W.; Ma, X.L.; Yu, K.X.; Huang, B.; Liao, M.; Li, F.; Pan, J.; Song, M.X. Comparative analysis of selected innate immune-related genes following infection of immortal DF-1 cells with highly pathogenic (H5N1) and low pathogenic (H9N2) avian influenza viruses. Virus Genes 2015, 50, 189–199. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.F.; Soung do, Y.; Schwarz, E.M.; O’Keefe, R.J.; Drissi, H. Wnt induction of chondrocyte hypertrophy through the Runx2 transcription factor. J. Cell. Physiol. 2006, 208, 77–86. [Google Scholar] [CrossRef] [PubMed]
- Xiao, S.Q.; Wang, X.; Ni, H.B.; Li, N.; Zhang, A.K.; Liu, H.L.; Pu, F.X.; Xu, L.L.; Gao, J.M.; Zhao, Q.; et al. MicroRNA miR-24-3p Promotes Porcine Reproductive and Respiratory Syndrome Virus Replication through Suppression of Heme Oxygenase-1 Expression. J. Virol. 2015, 89, 4494–4503. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pizzi, M. Sampling variation of the fifty percent end-point, determined by the Reed-Muench (Behrens) method. Hum. Biol. 1950, 22, 151–190. [Google Scholar] [PubMed]
| Primers | Sequences (5′-3′) | Methods | References |
|---|---|---|---|
| Mx | F: AAGCCTGAGCATGAGCAGAA | Real-time qPCR | [62] |
| R: TCTCAGGCTGTCAACAAGATCAA | |||
| OASL | F: AGATGTTGAAGCCGAAGTACCC | Real-time qPCR | [62] |
| R: CTGAAGTCCTCCCTGCCTGT | |||
| MOV10 | F: CCAGGACTGACATAAGAACACC | Real-time qPCR | |
| R: TTCCCACGGATGAACTCG | |||
| TGM2 | F: CACGACACCAACGGCAACC | Real-time qPCR | |
| R: GACCTCCGCAAAGACGAA | |||
| RUNX2 | F: ACTTTGACAATAACTGTCCT | Real-time qPCR | [63] |
| R: GACACCTACTCTCATACTGG | |||
| ISG12(1) | F: GGGTTCCATAGCAGCCAAG | Real-time qPCR | |
| R: CAACGAAAGAGAGCCCCGC | |||
| gga-miR-122-3p | F: AACGCCATTATCACACTAAATA | Real-time qPCR | |
| gga-miR-187-3p | F: TCGTGTCTTGTGTTGCAGCC | Real-time qPCR | |
| gga-miR-34a-5p | F: TGGCAGTGTCTTAGCTGGTTGTT | Real-time qPCR | |
| gga-miR-124a-3p | F: TTAAGGCACGCGGTGAATGCCA | Real-time qPCR | |
| gga-miR-183 | F: TATGGCACTGGTAGAATTCACTG | Real-time qPCR | |
| gga-miR-205a | F: TCCTTCATTCCACCGGAGTCTG | Real-time qPCR | |
| 28S | F: GGTATGGGCCCGACGCT | Real-time qPCR | |
| R: CCGATGCCGACGCTCAT | |||
| gga-miR-203a-RT | CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGCAAGTGGT | Real-time qPCR | |
| gga-miR-203a-qF | ACACTCCAGCTGGGGTGAAATGTTTAGGAC | ||
| URP (unified reverse primer) | TGGTGTCGTGGAGTCG | ||
| Luc-TGM2 | F: CCGCTCGAGGGGCCCCCCGAGCCCCCACCCT | Luciferase | |
| R: TTGCGGCCGCTGGCTTTAAAAGTGAAAAATAGAGGT | |||
| Mut-TGM2 | F: TCGAGCCCCTGGGACGCAGAGCAGTAAAGTTAACGCAAACAATAGAAAGACAATCGC | Luciferase | |
| R: GGCCGCGATTGTCTTTCTATTGTTTGCGTTAACTTTACTGCTCTGCGTCCCAGGGGC | |||
| pCAGGS-HA-TGM2 | F: GGAATTCATGGGTGGACCGGGACCGGACGG | Gene clone | |
| R: CGGCTAGCTCAAGCGTAGTCTGGGACGTCGTATGGGTACTTGGGCAGGGGTGCGATG |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jia, Y.-Q.; Wang, X.-L.; Wang, X.-W.; Yan, C.-Q.; Lv, C.-J.; Li, X.-Q.; Chu, Z.-L.; Adam, F.E.A.; Xiao, S.; Zhang, S.-X.; et al. Common microRNA–mRNA Interactions in Different Newcastle Disease Virus-Infected Chicken Embryonic Visceral Tissues. Int. J. Mol. Sci. 2018, 19, 1291. https://doi.org/10.3390/ijms19051291
Jia Y-Q, Wang X-L, Wang X-W, Yan C-Q, Lv C-J, Li X-Q, Chu Z-L, Adam FEA, Xiao S, Zhang S-X, et al. Common microRNA–mRNA Interactions in Different Newcastle Disease Virus-Infected Chicken Embryonic Visceral Tissues. International Journal of Molecular Sciences. 2018; 19(5):1291. https://doi.org/10.3390/ijms19051291
Chicago/Turabian StyleJia, Yan-Qing, Xing-Long Wang, Xiang-Wei Wang, Chuan-Qi Yan, Chang-Jie Lv, Xiao-Qin Li, Zhi-Li Chu, Fathalrhman Eisa Addoma Adam, Sa Xiao, Shu-Xia Zhang, and et al. 2018. "Common microRNA–mRNA Interactions in Different Newcastle Disease Virus-Infected Chicken Embryonic Visceral Tissues" International Journal of Molecular Sciences 19, no. 5: 1291. https://doi.org/10.3390/ijms19051291
APA StyleJia, Y.-Q., Wang, X.-L., Wang, X.-W., Yan, C.-Q., Lv, C.-J., Li, X.-Q., Chu, Z.-L., Adam, F. E. A., Xiao, S., Zhang, S.-X., & Yang, Z.-Q. (2018). Common microRNA–mRNA Interactions in Different Newcastle Disease Virus-Infected Chicken Embryonic Visceral Tissues. International Journal of Molecular Sciences, 19(5), 1291. https://doi.org/10.3390/ijms19051291







