HtrA1 Is Specifically Up-Regulated in Active Keloid Lesions and Stimulates Keloid Development
Abstract
:1. Introduction
2. Results
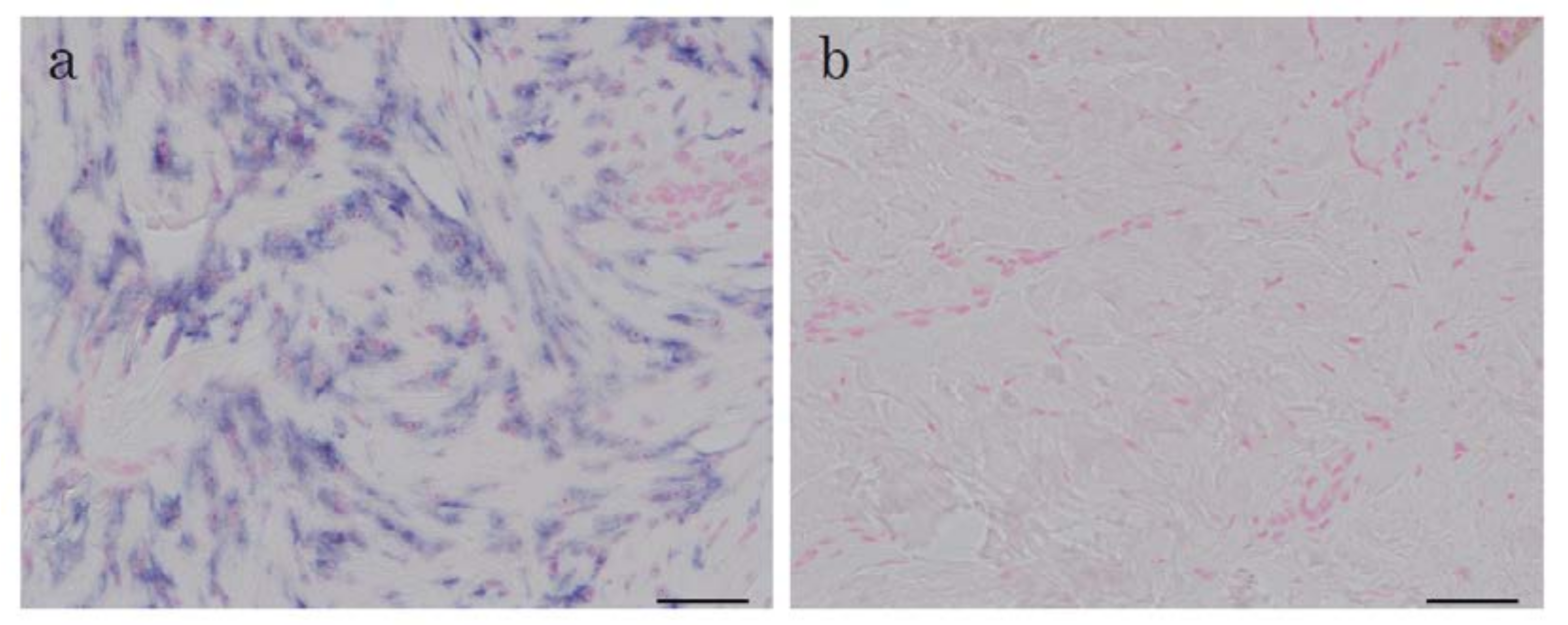
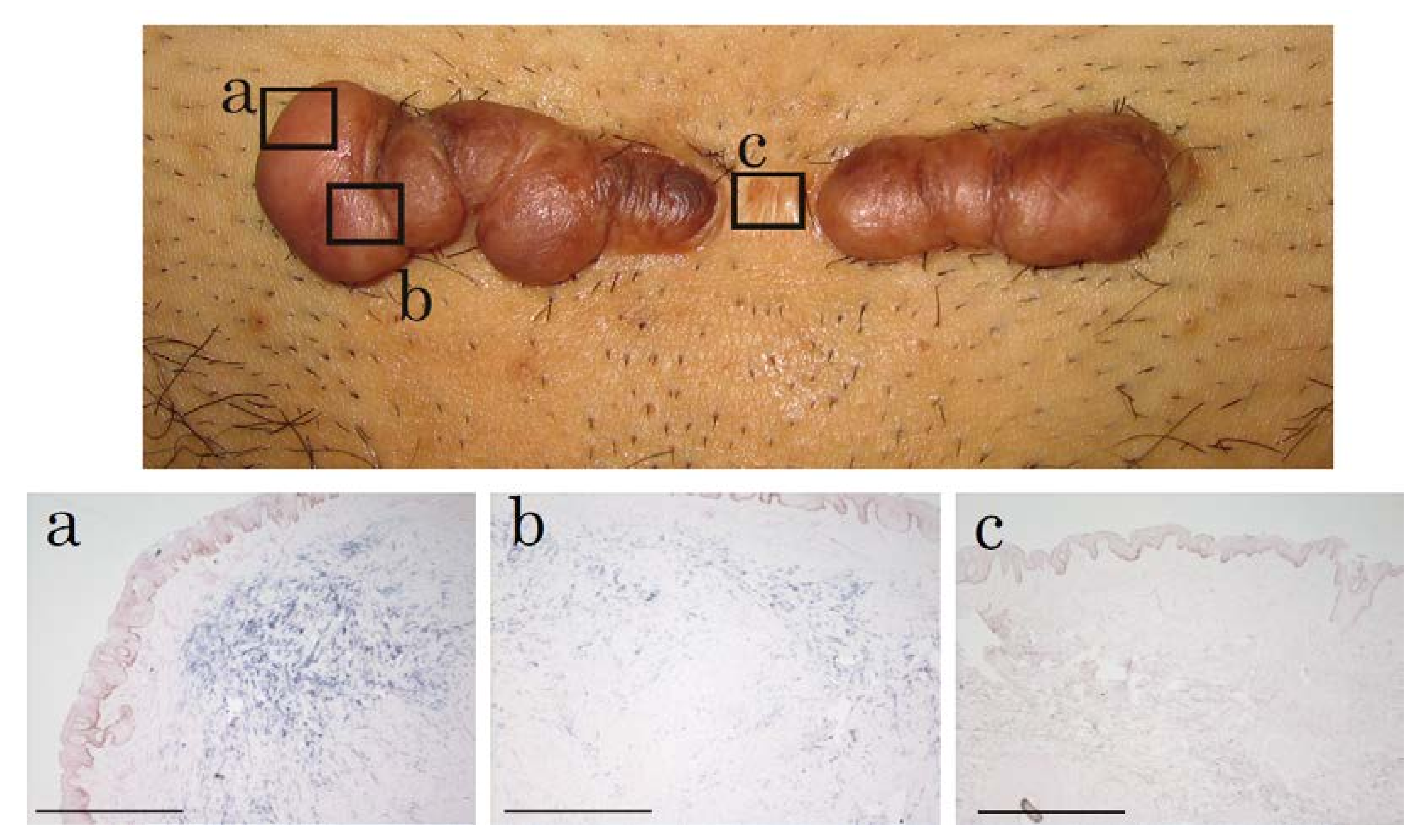
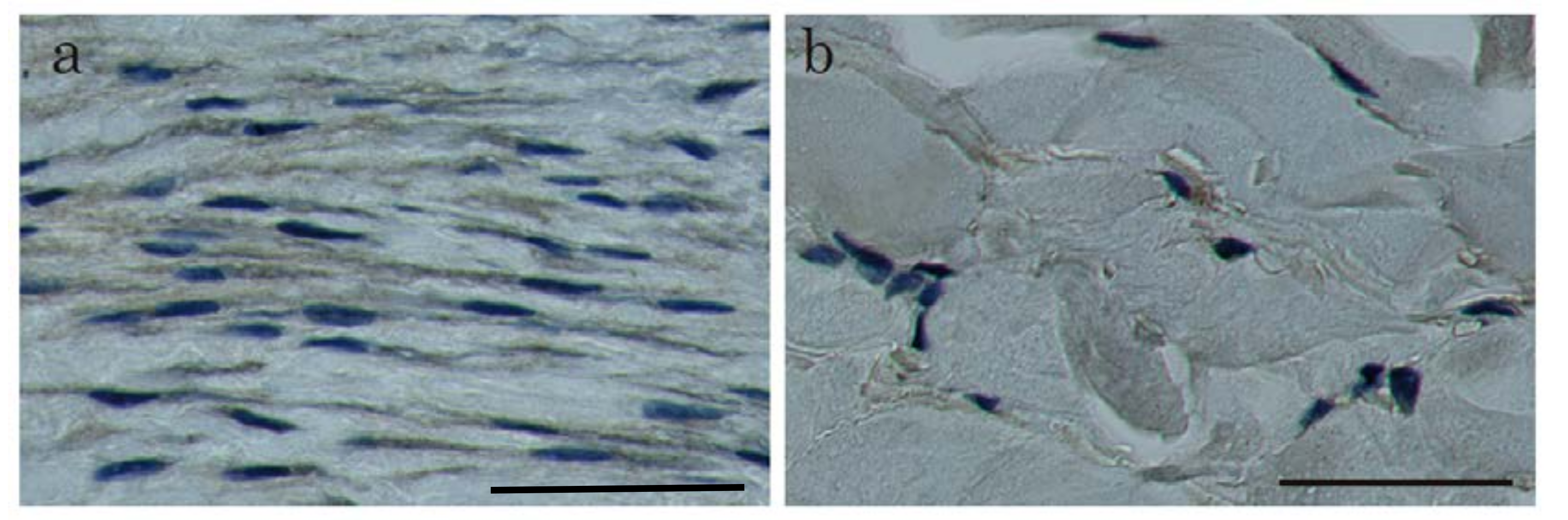
2.1. In Situ Hybridization of HtrA1 mRNA in Keloid Lesions and Normal Skin
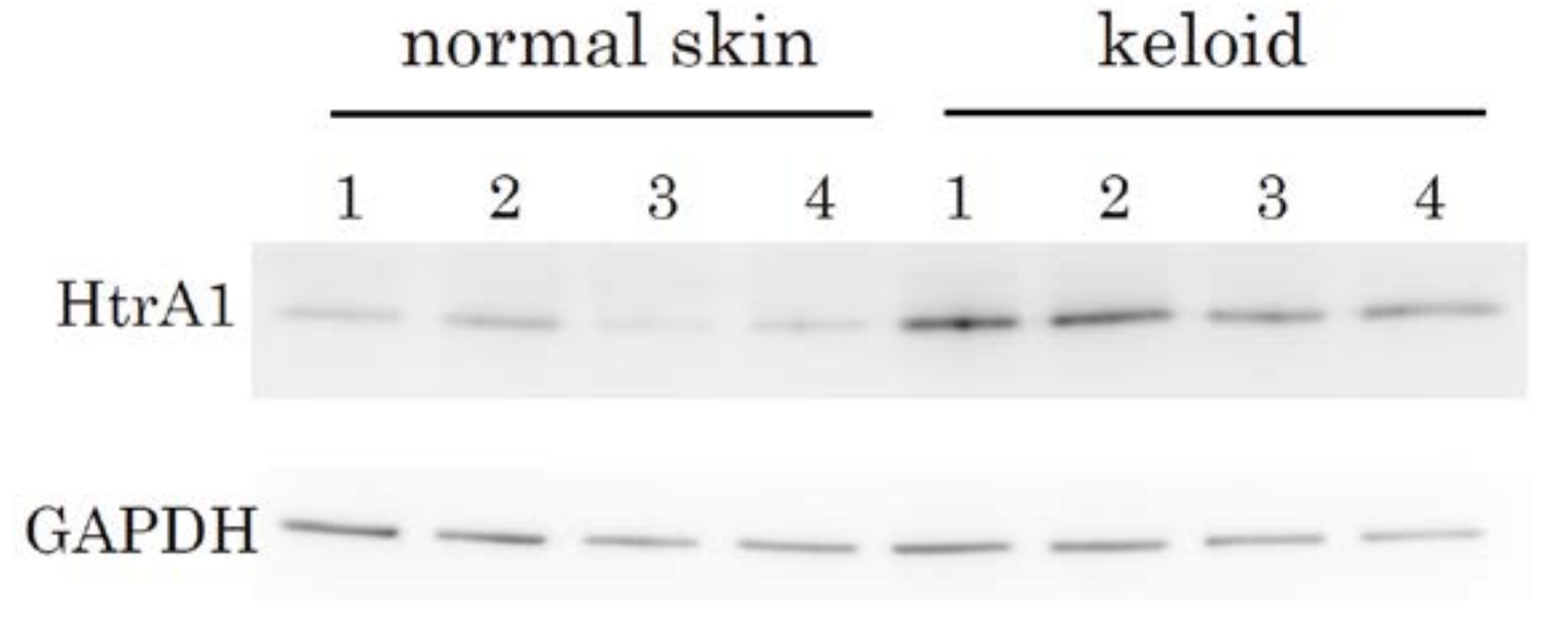
2.2. Immunohistochemical Staining and Western Blot Analysis of HtrA1
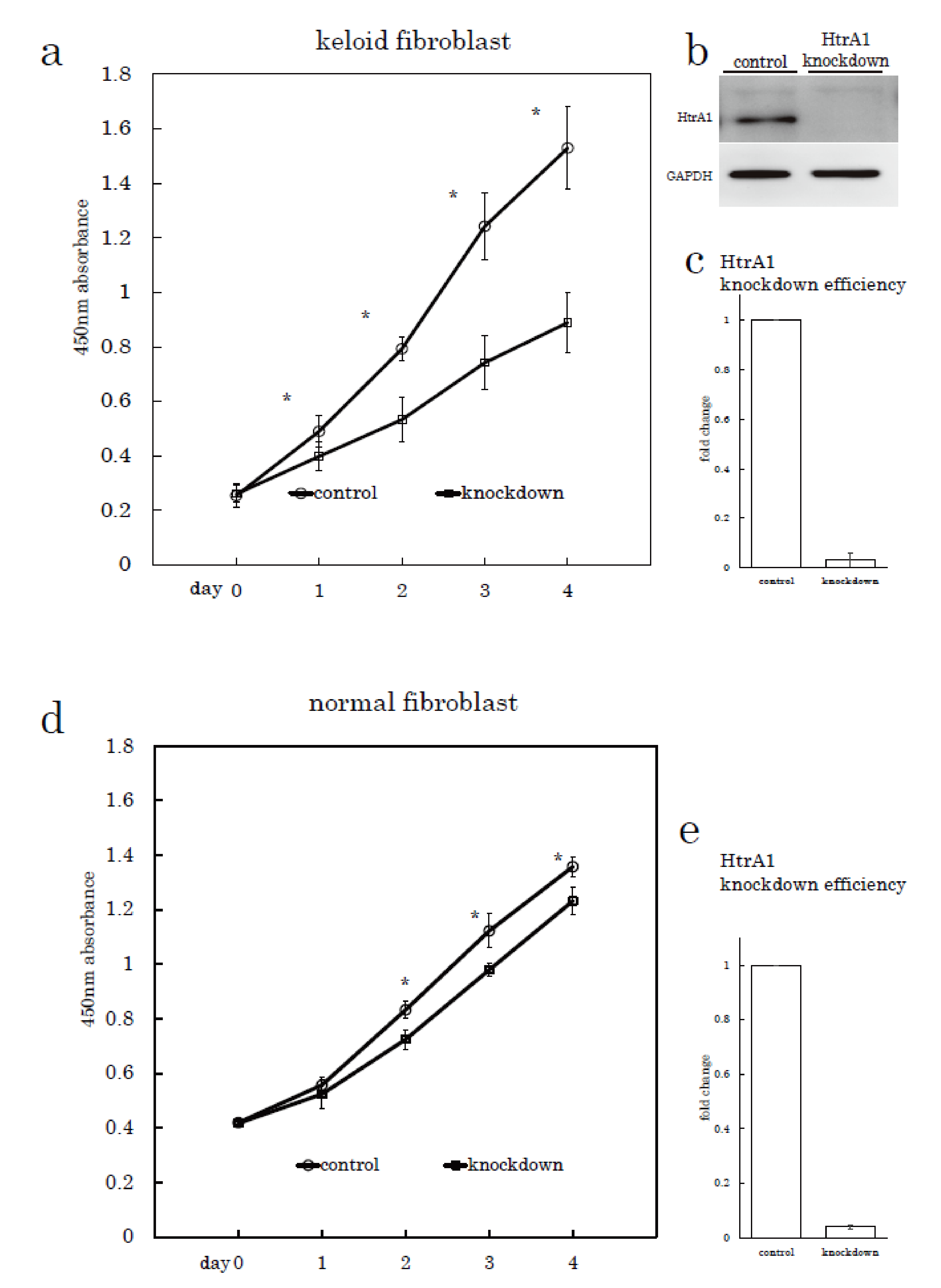
2.3. HtrA1 Knockdown Inhibits Keloid Cell Proliferation
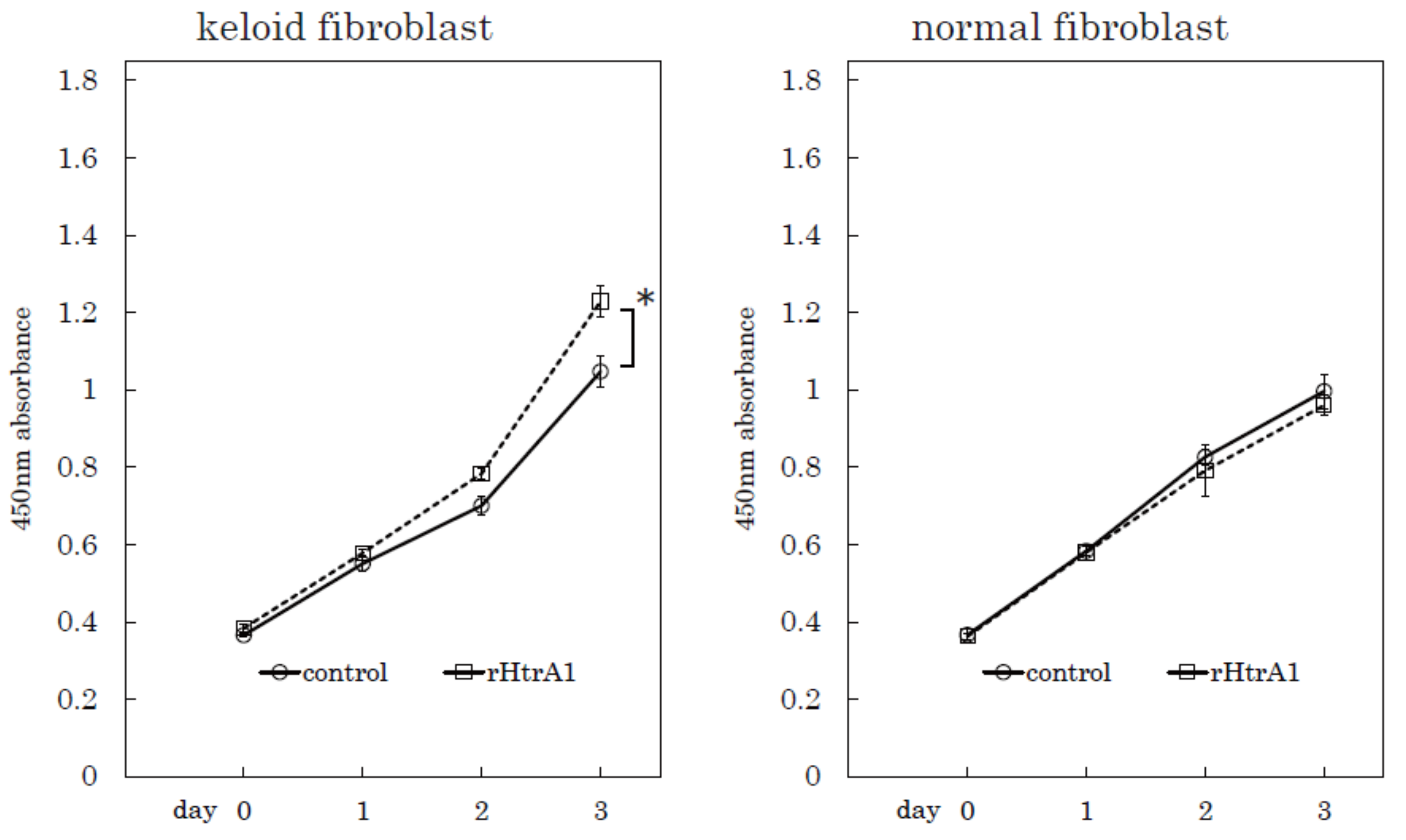
2.4. Additional HtrA1 in Culture Medium Stimulates Keloid Cell Proliferation
3. Discussion
4. Materials and Methods
4.1. Tissue Specimens
4.2. Antibodies
4.3. In Situ Hybridization
4.4. Immunohistochemical Analysis
4.5. Statistical Analysis
4.6. Western Blot Analysis
4.7. Knockdown of HtrA1 Gene Expression and Cell Proliferation Assay
4.8. Real-Time PCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
Abbreviations
| HtrA1 | High Temperature Requirement Factor A1 |
References
- Abergel, R.P.; Pizzurro, D.; Meeker, C.A.; Lask, G.; Matsuoka, L.Y.; Minor, R.R.; Chu, M.L.; Uitto, J. Biochemical composition of the connective tissue in keloids and analysis of collagen metabolism in keloid fibroblast cultures. J.Investig. Dermatol. 1985, 84, 384–390. [Google Scholar] [CrossRef] [PubMed]
- Sidgwick, G.P.; Bayat, A. Extracellular matrix molecules implicated in hypertrophic and keloid scarring. J. Eur. Acad. Dermatol. Venereol. 2012, 26, 141–152. [Google Scholar] [CrossRef] [PubMed]
- Mustoe, T.A.; Cooter, R.D.; Gold, M.H.; Hobbs, F.D.; Ramelet, A.A.; Shakespeare, P.G.; Stella, M.; Teot, L.; Wood, F.M.; Ziegler, U.E.; et al. International clinical recommendations on scar management. Plast. Reconstruct. Surg. 2002, 110, 560–571. [Google Scholar] [CrossRef]
- Alster, T.S.; Tanzi, E.L. Hypertrophic scars and keloids: Etiology and management. Am. J. Clin. Dermatol. 2003, 4, 235–243. [Google Scholar] [CrossRef] [PubMed]
- Van de Kar, A.L.; Kreulen, M.; van Zuijlen, P.P.; Oldenburger, F. The results of surgical excision and adjuvant irradiation for therapy-resistant keloids: A prospective clinical outcome study. Plast. Reconstruct. Surg. 2007, 119, 2248–2254. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, R.; Miyashita, T.; Hyakusoku, H.; Akaishi, S.; Kuribayashi, S.; Tateno, A. Postoperative radiation protocol for keloids and hypertrophic scars: Statistical analysis of 370 sites followed for over 18 months. Ann. Plast. Surg. 2007, 59, 688–691. [Google Scholar] [CrossRef] [PubMed]
- Yamawaki, S.; Naitoh, M.; Ishiko, T.; Muneuchi, G.; Suzuki, S. Keloids can be forced into remission with surgical excision and radiation, followed by adjuvant therapy. Ann. Plast. Surg. 2011, 67, 402–406. [Google Scholar] [CrossRef] [PubMed]
- Van Leeuwen, M.C.; Stokmans, S.C.; Bulstra, A.E.; Meijer, O.W.; Heymans, M.W.; Ket, J.C.; Ritt, M.J.; van Leeuwen, P.A.; Niessen, F.B. Surgical excision with adjuvant irradiation for treatment of keloid scars: A systematic review. Plast. Reconstruct. Surg. 2015, 3, e440. [Google Scholar] [CrossRef] [PubMed]
- Naitoh, M.; Kubota, H.; Ikeda, M.; Tanaka, T.; Shirane, H.; Suzuki, S.; Nagata, K. Gene expression in human keloids is altered from dermal to chondrocytic and osteogenic lineage. Genes Cells 2005, 10, 1081–1091. [Google Scholar] [CrossRef] [PubMed]
- Chien, J.; Campioni, M.; Shridhar, V.; Baldi, A. HtrA serine proteases as potential therapeutic targets in cancer. Curr. Cancer Drug Targets 2009, 9, 451–468. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Huang, L.; Yu, W.; Wu, X.; Zhou, P.; Li, X. Overexpression of HTRA1 leads to down-regulation of fibronectin and functional changes in RF/6A cells and HUVECs. PLoS ONE 2012, 7, e46115. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Lim, S.L.; Du, H.; Zhang, M.; Kozak, I.; Hannum, G.; Wang, X.; Ouyang, H.; Hughes, G.; Zhao, L.; et al. High temperature requirement factor A1 (HTRA1) gene regulates angiogenesis through transforming growth factor-β family member growth differentiation factor 6. J. Biol. Chem. 2012, 287, 1520–1526. [Google Scholar] [CrossRef] [PubMed]
- Beaufort, N.; Scharrer, E.; Kremmer, E.; Lux, V.; Ehrmann, M.; Huber, R.; Houlden, H.; Werring, D.; Haffner, C.; Dichgans, M. Cerebral small vessel disease-related protease HtrA1 processes latent TGF-β binding protein 1 and facilitates TGF-β signaling. Proc. Natl. Acad. Sci. USA 2014, 111, 16496–16501. [Google Scholar] [CrossRef] [PubMed]
- Grau, S.; Richards, P.J.; Kerr, B.; Hughes, C.; Caterson, B.; Williams, A.S.; Junker, U.; Jones, S.A.; Clausen, T.; Ehrmann, M. The role of human HtrA1 in arthritic disease. J. Biol. Chem. 2006, 281, 6124–6129. [Google Scholar] [CrossRef] [PubMed]
- Jones, A.; Kumar, S.; Zhang, N.; Tong, Z.; Yang, J.H.; Watt, C.; Anderson, J.; Amrita, J.; Fillerup, H.; McCloskey, M.; et al. Increased expression of multifunctional serine protease, HTRA1, in retinal pigment epithelium induces polypoidal choroidal vasculopathy in mice. Proc. Natl. Acad. Sci. USA 2011, 108, 14578–14583. [Google Scholar] [CrossRef] [PubMed]
- Vierkotten, S.; Muether, P.S.; Fauser, S. Overexpression of HTRA1 leads to ultrastructural changes in the elastic layer of Bruch’s membrane via cleavage of extracellular matrix components. PLoS ONE 2011, 6, e22959. [Google Scholar] [CrossRef] [PubMed]
- Filliat, G.; Mirsaidi, A.; Tiaden, A.N.; Kuhn, G.A.; Weber, F.E.; Oka, C.; Richards, P.J. Role of HTRA1 in bone formation and regeneration: In vitro and in vivo evaluation. PLoS ONE 2017, 12, e0181600. [Google Scholar] [CrossRef] [PubMed]
- Tsuchiya, A.; Yano, M.; Tocharus, J.; Kojima, H.; Fukumoto, M.; Kawaichi, M.; Oka, C. Expression of mouse HtrA1 serine protease in normal bone and cartilage and its upregulation in joint cartilage damaged by experimental arthritis. Bone 2005, 37, 323–336. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Ortega, M.; Rodriguez-Vita, J.; Sanchez-Lopez, E.; Carvajal, G.; Egido, J. TGF-β signaling in vascular fibrosis. Cardiovasc. Res. 2007, 74, 196–206. [Google Scholar] [CrossRef] [PubMed]
- Taipale, J.; Miyazono, K.; Heldin, C.H.; Keski-Oja, J. Latent transforming growth factor-β 1 associates to fibroblast extracellular matrix via latent TGF-β binding protein. J. Cell Biol. 1994, 124, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Tesseur, I.; Zou, K.; Berber, E.; Zhang, H.; Wyss-Coray, T. Highly sensitive and specific bioassay for measuring bioactive TGF-β. BMC Cell Biol. 2006, 7, 15. [Google Scholar] [CrossRef] [PubMed]
- Todorovic, V.; Rifkin, D.B. LTBPs, more than just an escort service. J. Cell. Biochem. 2012, 113, 410–418. [Google Scholar] [CrossRef] [PubMed]
- Dallas, S.L.; Rosser, J.L.; Mundy, G.R.; Bonewald, L.F. Proteolysis of latent transforming growth factor-β (TGF-β)-binding protein-1 by osteoclasts. A cellular mechanism for release of TGF-β from bone matrix. J. Cell. Biochem. 2002, 277, 21352–21360. [Google Scholar] [CrossRef]
- Ge, G.; Greenspan, D.S. BMP1 controls TGF-β1 activation via cleavage of latent TGF-β-binding protein. J. Cell Biol. 2006, 175, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Tatti, O.; Vehvilainen, P.; Lehti, K.; Keski-Oja, J. MT1-MMP releases latent TGF-β1 from endothelial cell extracellular matrix via proteolytic processing of LTBP-1. Exp. Cell Res. 2008, 314, 2501–2514. [Google Scholar] [CrossRef] [PubMed]
- Border, W.A.; Noble, N.A. Transforming growth factor β in tissue fibrosis. N. Engl. J. Med. 1994, 331, 1286–1292. [Google Scholar] [CrossRef] [PubMed]
- Peltonen, J.; Hsiao, L.L.; Jaakkola, S.; Sollberg, S.; Aumailley, M.; Timpl, R.; Chu, M.L.; Uitto, J. Activation of collagen gene expression in keloids: Co-localization of type I and VI collagen and transforming growth factor-β 1 mRNA. J. Investig. Dermatol. 1991, 97, 240–248. [Google Scholar] [CrossRef] [PubMed]
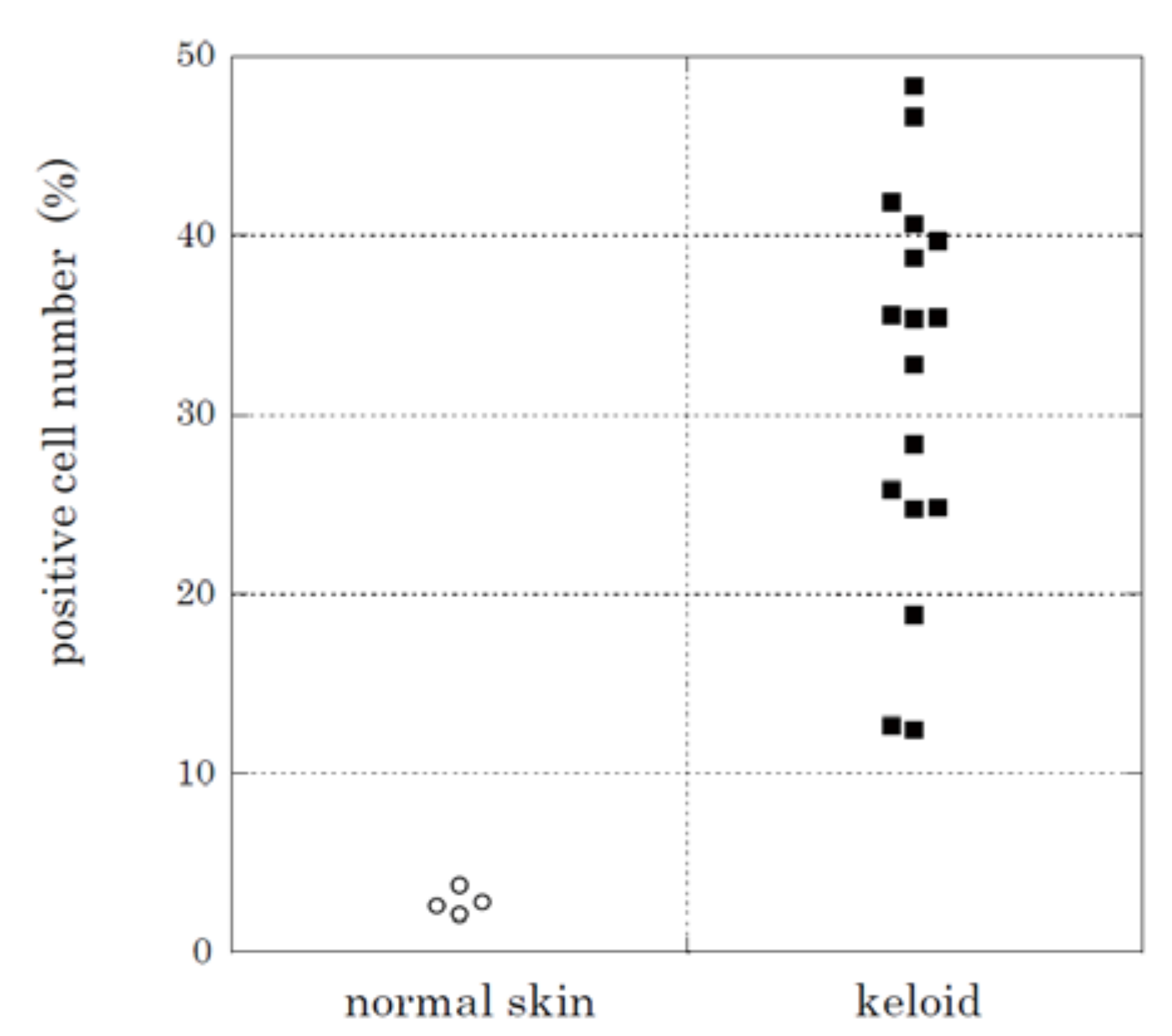
| Tissue Source | No. | Age | Sex | Region | HtrA1-Positve Cells (%) 1 | Assays 2 |
|---|---|---|---|---|---|---|
| Keloid | 1 | 75 | M | shoulder | 18.83 | A |
| 2 | 51 | F | chest | 12.40 | A | |
| 3 | 49 | M | neck | 38.74 | A, B | |
| 4 | 67 | F | abdomen | 43.52 | A | |
| 5 | 32 | M | chest | 24.77 | A | |
| 6 | 34 | M | abdomen | 40.64 | A | |
| 7 | 67 | M | abdomen | 32.80 | A | |
| 8 | 16 | F | chest | 41.90 | A, C | |
| 9 | 64 | M | shoulder | 25.82 | A | |
| 10 | 28 | M | chest | 24.85 | A | |
| 11 | 24 | F | chest | 28.36 | A, C | |
| 12 | 20 | F | shoulder | 35.43 | A | |
| 13 | 62 | F | chest | 12.68 | A | |
| 14 | 30 | M | shoulder | 35.34 | A | |
| 15 | 20 | F | chest | 48.37 | A | |
| 16 | 65 | M | back | 35.55 | A | |
| 17 | 38 | F | chest | 39.68 | A | |
| 18 | 39 | F | abdomen | B | ||
| 19 | 75 | M | chest | B | ||
| 20 | 21 | M | back | B | ||
| 21 | 20 | M | back | B | ||
| 22 | 31 | M | shoulder | D | ||
| 23 | 20 | F | shoulder | D | ||
| 24 | 20 | F | chest | C, D | ||
| 25 | 58 | F | shoulder | D | ||
| 26 | 24 | M | chest | C | ||
| 27 | 41 | F | abdomen | B | ||
| 28 | 27 | M | chest | C | ||
| 29 | 17 | F | chest | C | ||
| 30 | 24 | F | chest | C | ||
| Normal skin | 1 | 51 | F | back | 3.76 | A, D |
| 2 | 45 | F | abdomen | 2.83 | A | |
| 3 | 47 | F | shoulder | 2.60 | A | |
| 4 | 51 | F | thigh | 2.13 | A | |
| 5 | 88 | M | back | D | ||
| 6 | 49 | F | abdomen | D | ||
| 7 | 51 | F | abdomen | D | ||
| 8 | 52 | F | chest | C | ||
| 9 | 53 | F | chest | C | ||
| 10 | 40 | F | abdomen | C | ||
| 11 | 31 | F | abdomen | C | ||
| 12 | 52 | F | chest | C |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yamawaki, S.; Naitoh, M.; Kubota, H.; Aya, R.; Katayama, Y.; Ishiko, T.; Tamura, T.; Yoshikawa, K.; Enoshiri, T.; Ikeda, M.; et al. HtrA1 Is Specifically Up-Regulated in Active Keloid Lesions and Stimulates Keloid Development. Int. J. Mol. Sci. 2018, 19, 1275. https://doi.org/10.3390/ijms19051275
Yamawaki S, Naitoh M, Kubota H, Aya R, Katayama Y, Ishiko T, Tamura T, Yoshikawa K, Enoshiri T, Ikeda M, et al. HtrA1 Is Specifically Up-Regulated in Active Keloid Lesions and Stimulates Keloid Development. International Journal of Molecular Sciences. 2018; 19(5):1275. https://doi.org/10.3390/ijms19051275
Chicago/Turabian StyleYamawaki, Satoko, Motoko Naitoh, Hiroshi Kubota, Rino Aya, Yasuhiro Katayama, Toshihiro Ishiko, Taku Tamura, Katsuhiro Yoshikawa, Tatsuki Enoshiri, Mika Ikeda, and et al. 2018. "HtrA1 Is Specifically Up-Regulated in Active Keloid Lesions and Stimulates Keloid Development" International Journal of Molecular Sciences 19, no. 5: 1275. https://doi.org/10.3390/ijms19051275
APA StyleYamawaki, S., Naitoh, M., Kubota, H., Aya, R., Katayama, Y., Ishiko, T., Tamura, T., Yoshikawa, K., Enoshiri, T., Ikeda, M., & Suzuki, S. (2018). HtrA1 Is Specifically Up-Regulated in Active Keloid Lesions and Stimulates Keloid Development. International Journal of Molecular Sciences, 19(5), 1275. https://doi.org/10.3390/ijms19051275





