Beneficial Phytochemicals with Anti-Tumor Potential Revealed through Metabolic Profiling of New Red Pigmented Lettuces (Lactuca sativa L.)
Abstract
:1. Introduction
2. Results and Discussion
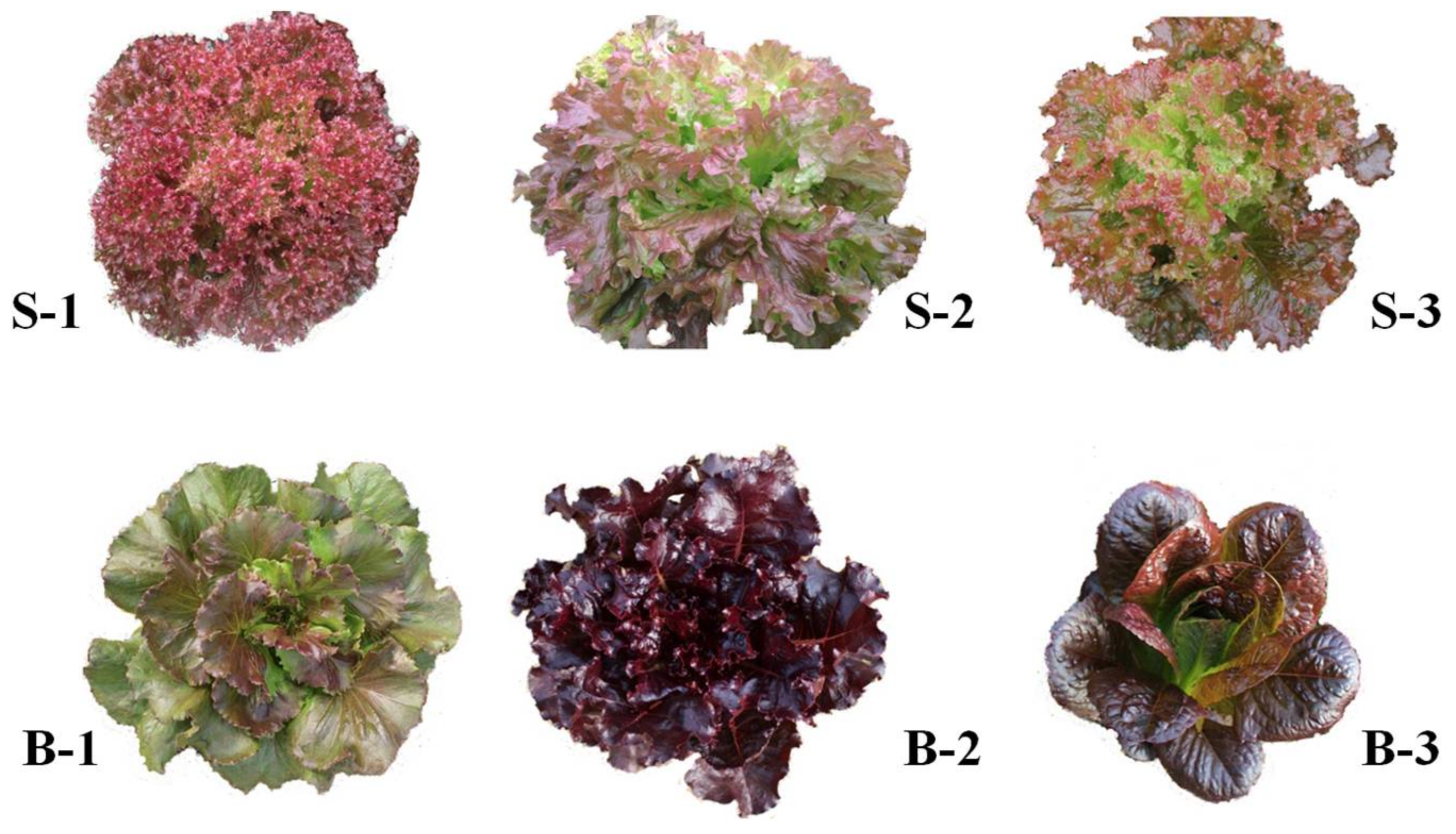
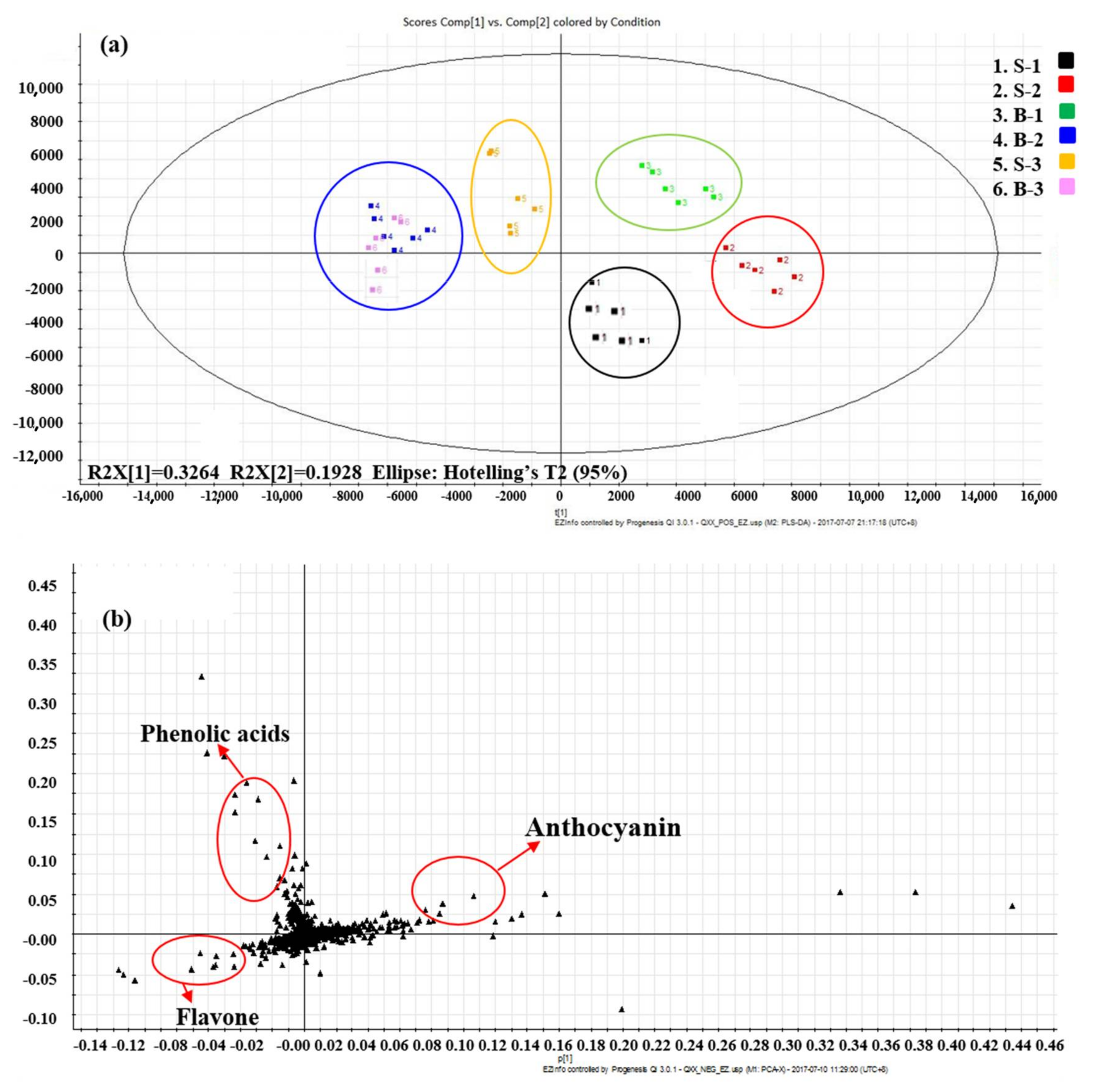
2.1. Phytochemical Composition in Different Red Lettuce Cultivars
2.2. Mineral Content in Different Red Lettuce Cultivars
2.3. Comparison of Antioxidant Activity in Different Red-Lettuce Cultivars
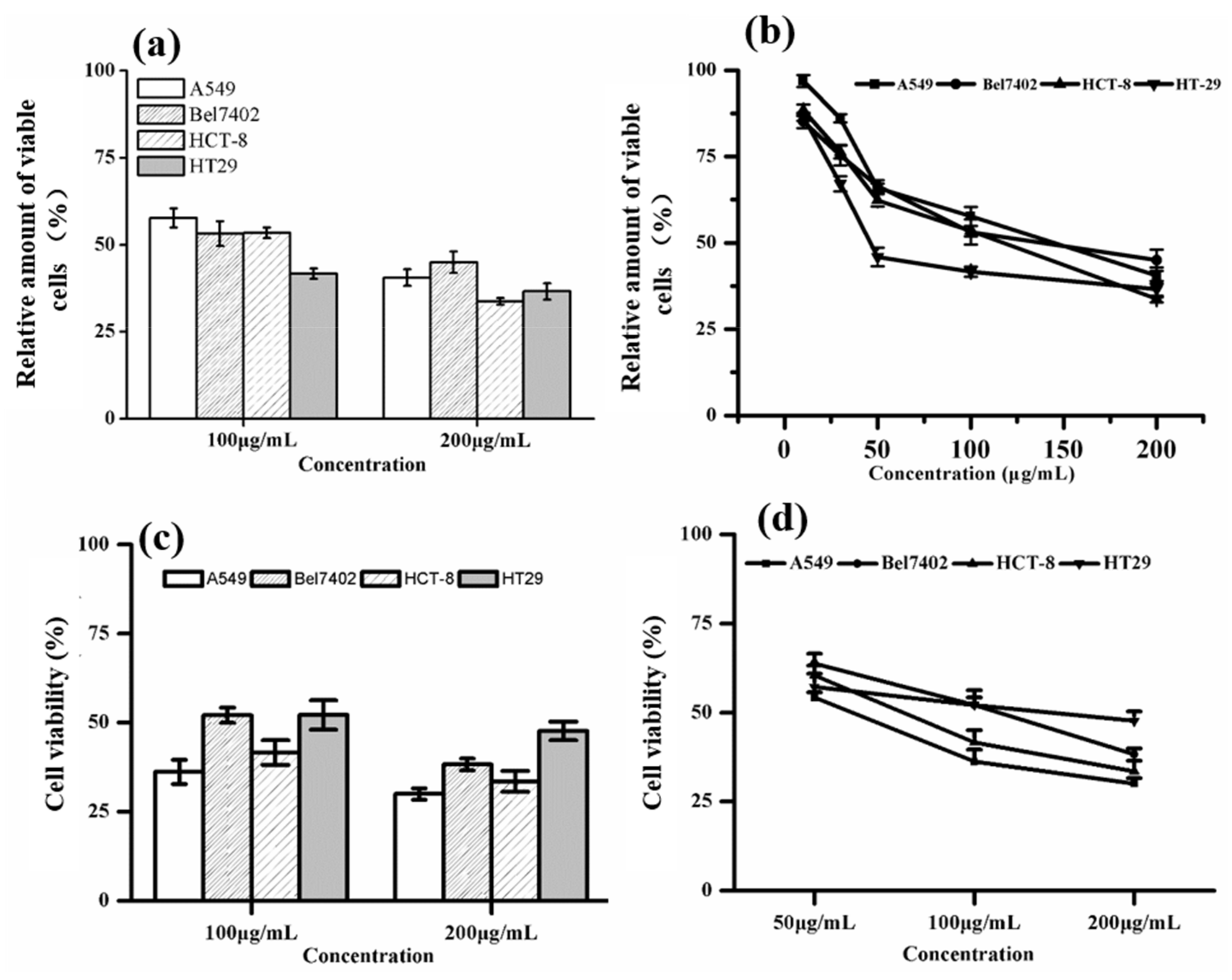
2.4. Analysis of In Vitro Anticancer Effects Activity in the New Dark-Red Lettuce Cultivar (B-2)
3. Materials and Methods
3.1. Plant Materials
3.2. UPLC–QTOF-MS Analysis Phytochemical Composition
3.3. Analysis of Mineral Content
3.4. Analysis of Antioxidant Activity
3.5. Extraction of Phytochemical Composition in Red-Lettuce B-2 Cultivar
3.6. Analysis of Tumor Cell Growth Inhibition In Vitro
3.7. Statistical Analysis
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Llorach, R.; Martínez-Sánchez, A.; Tomás-Barberán, F.A.; Gil, M.I.; Ferreres, F. Characterisation of polyphenols and antioxidant properties of five lettuce varieties and escarole. Food Chem. 2008, 108, 1028–1038. [Google Scholar] [CrossRef] [PubMed]
- Nicolle, C.; Cardinault, N.; Gueux, E.; Jaffrelo, L.; Rock, E.; Mazur, A.; Amouroux, P.; Rémésy, C. Health effect of vegetable-based diet: Lettuce consumption improves cholesterol metabolism and antioxidant status in the rat. Clin. Nutr. 2004, 23, 605–614. [Google Scholar] [CrossRef] [PubMed]
- Urszula, Z.; Micha, Ś.; Anna, J. Effect of abiotic elicitation on main health-promoting compounds, antioxidant activity and commercial quality of butter lettuce (Lactuca sativa L.). Food Chem. 2014, 148, 253–260. [Google Scholar]
- Kim, M.J.; Moon, Y.; Tou, J.C.; Mou, B.; Waterland, N.L. Nutritional value, bioactive compounds and health benefits of lettuce (Lactuca sativa L.). J. Food Compos. Anal. 2016, 49, 19–34. [Google Scholar] [CrossRef]
- Mulabagal, V.; Ngouajio, M.; Nair, A.; Zhang, Y.J.; Gottumukkala, A.L.; Nair, M.G. In vitro evaluation of red and green lettuce (Lactuca sativa) for functional food properties. Food Chem. 2010, 118, 300–306. [Google Scholar] [CrossRef]
- Siti Azima, A.M.; Noriham, A.; Manshoor, N. Anthocyanin content in relation to the antioxidant activity and colour properties of Garcinia mangostana peel, Syzigium cumini and Clitoria ternatea extracts. Int. Food Res. J. 2014, 26, 347–356. [Google Scholar]
- Han, K.H.; Sekikawa, M.; Shimada, K.; Hashimoto, M.; Hashimoto, N.; Noda, T.; Tanaka, H.; Fukushima, M. Anthocyanin-rich purple potato flake extract has antioxidant capacity and improves antioxidant potential in rats. Br. J. Nutr. 2006, 96, 1125–1133. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Deng, Z.; Zhu, H.; Hu, C.; Liu, R.; Young, J.C.; Tsao, R. Highly pigmented vegetables: Anthocyanin compositions and their role in antioxidant activities. Food Res. Int. 2012, 46, 250–259. [Google Scholar] [CrossRef]
- Ferreres, F.; Gil, M.I.; Castaner, M.; Tomasbarberan, F.A. Phenolic metabolites in red pigmented lettuce (Lactuca sativa). Changes with minimal processing and cold storage. J. Agric. Food Chem. 1997, 45, 4249–4254. [Google Scholar] [CrossRef]
- Garcia, C.J.; García-Villalba, R.; Garrido, Y.; Gil, M.I.; Tomás-Barberán, F.A. Untargeted metabolomics approach using UPLC-ESI-QTOF-MS to explore the metabolome of fresh-cut iceberg lettuce. Metabolomics 2016, 12, 138. [Google Scholar] [CrossRef]
- Becker, C. Flavonoids and phenolic acids in lettuce: How can we maximize their concentration? And why should we? Acta Hortic. 2016, 1–10. [Google Scholar] [CrossRef]
- Khoo, H.E.; Azlan, A.; Tang, S.T.; Lim, S.M. Anthocyanidins and anthocyanins: Colored pigments as food, pharmaceutical ingredients, and the potential health benefits. Food Nutr. Res. 2017, 61, 1361779. [Google Scholar] [CrossRef] [PubMed]
- Fang, J. Classification of fruits based on anthocyanin types and relevance to their health effects. Nutrition 2015, 31, 1301–1306. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Li, H.; Yang, S.; Ma, W.; Liu, M.; Guo, S.; Zhan, J.; Zhang, H.; Tsang, S.Y.; Zhang, Z. Cyanidin-3-o-glucoside directly binds to ERα36 and inhibits EGFR-positive triple-negative breast cancer. Oncotarget 2016, 7, 68864–68882. [Google Scholar] [PubMed]
- Viacava, G.E.; Roura, S.I.; Berrueta, L.A.; Iriondo, C.; Gallo, B.; Alonso-Salces, R.M. Characterization of phenolic compounds in green and red oak-leaf lettuce cultivars by UHPLC-DAD-ESI-QToF/MS using MSE scan mode. J. Mass Spectrom. 2017, 52, 873–902. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Arasu, M.V.; Lee, M.K.; Chun, J.H.; Seo, J.M.; Lee, S.W.; Al-Dhabi, N.A.; Kim, S.J. Quantification of glucosinolates, anthocyanins, free amino acids, and vitamin C in inbred lines of cabbage (Brassica oleracea L.). Food Chem. 2014, 145, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Marín, A.; Rubio, J.S.; Martínez, V.; Gil, M.I. Antioxidant compounds in green and red peppers as affected by irrigation frequency, salinity and nutrient solution composition. J. Sci. Food Agric. 2009, 89, 1352–1359. [Google Scholar]
- Kim, D.E.; Shang, X.; Assefa, A.D.; Keum, Y.S.; Saini, R.K. Metabolite profiling of green, green/red, and red lettuce cultivars: Variation in health beneficial compounds and antioxidant potential. Food Res. Int. 2018, 105, 361–370. [Google Scholar] [CrossRef] [PubMed]
- Baslam, M.; Morales, F.; Garmendia, I.; Goicoechea, N. Nutritional quality of outer and inner leaves of green and red pigmented lettuces (Lactuca sativa L.) consumed as salads. Sci. Hortic. 2013, 151, 103–111. [Google Scholar] [CrossRef]
- Tsao, R.; Khanizadeh, S.; Dale, A. Designer fruits and vegetables with enriched phytochemicals for human health. Can. J. Plant Sci. 2006, 86, 773–786. [Google Scholar] [CrossRef]
- Morales-Soto, A.; García-Salas, P.; Rodríguez-Pérez, C.; Jiménez-Sánchez, C.; de la Luz Cádiz-Gurrea, M.; Segura-Carretero, A.; Fernández-Gutiérrez, A. Antioxidant capacity of 44 cultivars of fruits and vegetables grown in Andalusia (Spain). Food Res. Int. 2014, 58, 35–46. [Google Scholar] [CrossRef]
- Heo, H.J.; Kim, Y.J.; Chung, D.; Kim, D.O. Antioxidant capacities of individual and combined phenolics in a model system. Food Chem. 2007, 104, 87–92. [Google Scholar] [CrossRef]
- Sofo, A.; Lundegårdh, B.; Mårtensson, A.; Manfra, M.; Pepe, G.; Sommella, E.; Nisco, M.D.; Tenore, G.C.; Campiglia, P.; Scopa, A. Different agronomic and fertilization systems affect polyphenolic profile, antioxidant capacity and mineral composition of lettuce. Sci. Hortic. 2016, 204, 106–115. [Google Scholar] [CrossRef]
- Alarcón-Flores, M.I.; Romero-González, R.; Martínez Vidal, J.L.; Garrido Frenich, A. Multiclass Determination of Phenolic Compounds in Different Varieties of Tomato and Lettuce by Ultra High Performance Liquid Chromatography Coupled to Tandem Mass Spectrometry. Int. J. Food Prop. 2016, 19, 494–507. [Google Scholar] [CrossRef]
- Romani, A.; Pinelli, P.; Galardi, C.; Sani, G.; Cimato, A.; Heimler, D. Polyphenols in greenhouse and open-air-grown lettuce. Food Chem. 2002, 79, 337–342. [Google Scholar] [CrossRef]
- Llorach, R.; And, F.A.T.; Ferreres, F. Lettuce and Chicory Byproducts as a Source of Antioxidant Phenolic Extracts. J. Agric. Food Chem. 2004, 52, 5109–5116. [Google Scholar] [CrossRef] [PubMed]
- Plumb, R.; Mazzeo, J.R.; Grumbach, E.S.; Rainville, P.; Jones, M.; Wheat, T.; Neue, U.D.; Smith, B.; Johnson, K.A. The application of small porous particles, high temperatures, and high pressures to generate very high resolution LC and LC/MS separations. J. Sep. Sci. 2007, 30, 1158–1166. [Google Scholar] [CrossRef] [PubMed]
- Jaiswal, R.; Kiprotich, J.; Kuhnert, N. Determination of the hydroxycinnamate profile of 12 members of the Asteraceae family. Phytochemistry 2011, 72, 781–790. [Google Scholar] [CrossRef] [PubMed]
- Arzu, A.; Vural, G. Effect of various anti-browning agents on phenolic compounds profile of fresh lettuce (L. sativa). Food Chem. 2009, 117, 122–126. [Google Scholar]
- Becker, C.; Klaering, H.P.; Kroh, L.W.; Krumbein, A. Cool-cultivated red leaf lettuce accumulates cyanidin-3-O-(6”-O-malonyl)-glucoside and caffeoylmalic acid. Food Chem. 2014, 146, 404–411. [Google Scholar] [CrossRef] [PubMed]
- Fallovo, C.; Rouphael, Y.; Cardarelli, M.; Rea, E.; Battistelli, A.; Colla, G. Yield and quality of leafy lettuce in response to nutrient solution composition and growing season. Int. J. Food Agric. Environ. 2009, 7, 456–462. [Google Scholar]
- Mengjiao, X.; Yingyan, H.; Xiaoxiao, Q.; Zhifan, S.; Shuangxi, F. Study of nutritional quality and antioxidant activity in varieties of leaf lettuce. J. Beijing Univ. Agric. 2017, 32, 46–51. [Google Scholar]
- Song, J.; Zhao, M.; Liu, X.; Zhu, Y.; Hu, X.; Chen, F. Protection of cyanidin-3-glucoside against oxidative stress induced by acrylamide in human MDA-MB-231 cells. Food Chem. Toxicol. 2013, 58, 306–310. [Google Scholar] [CrossRef] [PubMed]
- Kong, J.M.; Chia, L.S.; Goh, N.K.; Chia, T.F.; Brouillard, R. Analysis and biological activities of anthocyanins. Phytochemistry 2003, 64, 923–933. [Google Scholar] [CrossRef]
- Bub, A.; Watzl, B.; Heeb, D.; Rechkemmer, G.; Briviba, K. Malvidin-3-glucoside bioavailability in humans after ingestion of red wine, dealcoholized red wine and red grape juice. Eur. J. Nutr. 2001, 40, 113–120. [Google Scholar] [CrossRef] [PubMed]
- Manach, C.; Morand, C.; Gilizquierdo, A.; Bouteloupdemange, C.; Rémésy, C. Bioavailability in humans of the flavanones hesperidin and narirutin after the ingestion of two doses of orange juice. Eur. J. Clin. Nutr. 2003, 57, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Erlund, I.; Meririnne, E.G.; Aro, A. Plasma kinetics and urinary excretion of the flavanones naringenin and hesperetin in humans after ingestion of orange juice and grapefruit juice. J. Nutr. 2001, 131, 235–241. [Google Scholar] [CrossRef] [PubMed]
- Serafini, M.; Bugianesi, R.; Salucci, M.; Azzini, E.; Raguzzini, A.; Maiani, G. Effect of acute ingestion of fresh and stored lettuce (Lactuca sativa) on plasma total antioxidant capacity and antioxidant levels in human subjects. Br. J. Nutr. 2002, 88, 615–623. [Google Scholar] [CrossRef] [PubMed]
- Suffness, M.; Pezzuto, J.M. Assays related to cancer drug discovery. In Methods in Plant Biochemistry: Assays for Bioactivity; Hostettmann, K., Ed.; Academic Press: London, UK, 1990; Volume 6, pp. 71–133. [Google Scholar]
- John, K.M.; Jung, E.S.; Lee, S.; Kim, J.-S.; Lee, C.H. Primary and secondary metabolites variation of soybean contaminated with Aspergillus sojae. Food Res. Int. 2013, 54, 487–494. [Google Scholar] [CrossRef]
- Lee, S.; Do, S.-G.; Kim, S.Y.; Kim, J.; Jin, Y.; Lee, C.H. Mass spectrometry-based metabolite profiling and antioxidant activity of Aloe vera (Aloe barbadensis Miller) in different growth stages. J. Agric. Food Chem. 2012, 60, 11222–11228. [Google Scholar] [CrossRef] [PubMed]
- Qin, X.; Xing, Y.F.; Zhou, Z.; Yao, Y. Dihydrochalcone Compounds Isolated from Crabapple Leaves Showed Anticancer Effects on Human Cancer Cell Lines. Molecules 2015, 20, 21193–21203. [Google Scholar] [CrossRef] [PubMed]
- Dai, Q.; Yin, Q.; Zhao, Y.; Guo, R.; Li, Z.; Ma, S.; Lu, N. III-10, a newly synthesized flavonoid, induces cell apoptosis with the involvement of reactive oxygen species-mitochondria pathway in human hepatocellular carcinoma cells. Eur. J. Pharmacol. 2015, 764, 353–362. [Google Scholar] [CrossRef] [PubMed]
- Strober, W. Trypan blue exclusion test of cell viability. Curr. Protoc. Immunol. 2001, 111. [Google Scholar] [CrossRef]
- Abel, S.D.A.; Baird, S.K. Honey is cytotoxic towards prostate cancer cells but interacts with the MTT reagent: considerations for the choice of cell viability assay. Food Chem. 2017, 241, 70–78. [Google Scholar] [CrossRef] [PubMed]

| Compounds | Rt (min) | Molecular Weight (Da) | Ion Mode | Q1 (Da) | Q3 (Da) | Class | VIP | Semi Quantitative |
|---|---|---|---|---|---|---|---|---|
| Cyanidin 3-O-glucoside | 2.58 | 448.3 | Negative | 447.3 | 284.1 | Anthocyanin | 1.07 | + |
| Malvidin 3-O-glucoside (Oenin) | 2.86 | 493.2 | Positive | 493.2 | 331.6 | Anthocyanin | 2.01 | ++ |
| Cyanidin 3-O-glucosyl-malonylglucoside | 3.06 | 697.1 | Positive | 697.1 | 686.9 | Anthocyanin | 2.59 | ++ |
| Cyanidin-3,5-O-diglucoside (Cyanin) | 3.17 | 611 | Positive | 611 | 287.7 | Anthocyanin | 3.47 | +++ |
| Delphinidin 3-O-glucoside (Mirtillin) | 4.73 | 465.1 | Positive | 465.1 | 303.1 | Anthocyanin | 1.02 | + |
| Cyanidin O-malonyl-malonylhexoside | 3.11 | 621.1 | Positive | 621.1 | 287.4 | Anthocyanin | 1.03 | + |
| Myricetin O-hexoside | 3.5 | 480.1 | Negative | 479.1 | 435.3 | Flavone | <1 | ---- |
| Apigenin -glucuronide | 2.88 | 446.076 | Negative | 445.07 | 269.2 | Flavone | 3.96 | +++ |
| Apigenin 7-O-glucoside (Cosmosiin) | 4.11 | 432.1 | Negative | 431.097 | 269.044 | Flavone | 1.33 | + |
| Apigenin 7-O-neohesperidoside (Rhoifolin) | 4.4 | 580.1792 | Negative | 579.178 | 135 | Flavone | 1.91 | + |
| Apigenin O-malonylhexoside | 4.36 | 518.0 | Negative | 519 | 282.1 | Flavone | 1.23 | + |
| Apigenin 7-rutinoside (Isorhoifolin) | 4.42 | 578.1636 | Positive | 579.164 | 271.1 | Flavone | <1 | ---- |
| Chrysoeriol 5-O-hexoside | 3.96 | 462.1 | Negative | 463.123 | 285.2 | Flavone | 1.13 | + |
| Luteolin O-rutinoside | 3.59 | 594.2 | Positive | 595.2 | 271.7 | Flavone | <1 | ---- |
| Luteolin 7-O-glucoside (Cynaroside) | 3.61 | 448.101 | Positive | 449.1 | 287.1 | Flavone | 3.64 | +++ |
| Luteolin O-hexosyl-O-gluconic acid | 3.85 | 625.1 | Negative | 624.1 | 285.3 | Flavone | 2.25 | ++ |
| Luteolin O-malonylhexoside | 4.15 | 534.0 | Positive | 535 | 301.7 | Flavone | <1 | ---- |
| Luteolin | 4.91 | 286.2363 | Negative | 285 | 133.1 | Flavone | <1 | ---- |
| Quercetin O-hexoside | 3.45 | 464.3 | Positive | 465.3 | 447.7 | Flavonol | <1 | ---- |
| Quercetin-3-O-rutinoside (Rutin) | 3.51 | 610.5175 | Positive | 609.153 | 300.1 | Flavonol | <1 | ---- |
| Quercetin O-hexosyl-O-malonylhexoside | 2.83 | 712.14 | Positive | 713.1 | 303.05 | Flavonol | <1 | ---- |
| Quercetin-3-O-malonylglucoside | 3.93 | 550.1031 | Negative | 549.089 | 505.099 | Flavonol | 1.01 | + |
| Quercetin-3-β-O-galactoside | 4.26 | 464.1 | Positive | 465.1 | 303.6 | Flavonol | <1 | ---- |
| Quercetin 3-O-glucoside (Isotrifoliin) | 3.72 | 464.096 | Negative | 463 | 301.1 | Flavonol | 1.61 | + |
| Chrysoeriol O-malonylhexoside | 4.43 | 548.1 | Positive | 549.1 | 286.7 | Flavone | <1 | ---- |
| Chrysin O-hexoside | 4.67 | 416.2 | Positive | 417.2 | 167.5 | Flavone | <1 | ---- |
| Chrysoeriol O-ferulic acid | 4.12 | 476.1 | Positive | 477.1 | 286.8 | Flavone | <1 | ---- |
| Chrysoeriol 7-O-rutinoside | 3.92 | 608.2 | Positive | 609.2 | 325.8 | Flavone | <1 | ---- |
| Chrysoeriol O-hexoside | 4.14 | 462.2 | Positive | 463.2 | 287.7 | Flavone | <1 | ---- |
| Chrysoeriol O-Glucuronic acid | 4.12 | 476.1 | Negative | 475.1 | 299 | Flavone | <1 | ---- |
| Chrysin O-malonylhexoside | 5.24 | 502.0 | Positive | 503 | 485.1 | Flavone | <1 | ---- |
| Chrysoeriol | 5.6 | 300.0634 | Negative | 299.063 | 284 | Flavone | <1 | ---- |
| Isorhamnetin O-acetyl-hexoside | 4.22 | 520.1 | Negative | 519.1 | 314.2 | Flavone | <1 | ---- |
| Tricin di-O-hexoside | 3.49 | 654.1 | Positive | 655.1 | 316.2 | Flavone | <1 | ---- |
| Genistein (4′,5,7-Trihydroxyisoflavone) | 5.45 | 270.2369 | Negative | 269.053 | 151 | Flavone | <1 | ---- |
| Tangeretin | 6.32 | 372.3686 | Positive | 373.121 | 343 | Flavone | <1 | ---- |
| Eriodictyol | 4.85 | 288.2522 | Negative | 287.063 | 151 | Flavanone | <1 | ---- |
| Geranyl acetate | 5.91 | 196.28 | Positive | 197 | 161.5 | Vitamins | 1.92 | + |
| Cinnamic acid | 5.22 | 148.1586 | Negative | 147.052 | 61.8 | Hydroxycinnamoyl derivatives | <1 | ---- |
| Quinic acid | 1.97 | 192.16658 | Negative | 191.1 | 85.1 | Quinate and its derivatives | 2.12 | ++ |
| Gallic acid O-hexoside | 1.18 | 332 | Negative | 331 | 313.7 | Benzoic acid derivatives | 1.04 | |
| Gallic acid | 1.81 | 170.1195 | Negative | 169.022 | 125.1 | Benzoic acid | 2.41 | ++ |
| Benzamidine | 1.31 | 120.154 | Positive | 121 | 77 | Phenylpropanoids | <1 | ---- |
| protocatechuic acid | 2.42 | 154.027 | Negative | 153.1 | 109.1 | Catechin derivatives | <1 | ---- |
| Neochlorogenic acid (5-O-Caffeoylquinic acid) | 2.53 | 354.1087 | Positive | 355.1 | 145.4 | Quinate and its derivatives | 3.16 | +++ |
| Chlorogenic acid (3-O-Caffeoylquinic acid) | 2.69 | 354.0951 | Negative | 353.095 | 191.1 | Quinate and its derivatives | 3.05 | +++ |
| 3,5-Dimethoxy-4-hydroxycinnamic acid | 2.51 | 223.085 | Negative | 222.1 | 163 | Amino acid derivatives | 1.15 | + |
| Coumaric acid O-glucoside | 2.8 | 326 | Negative | 325 | 119.2 | Hydroxycinnamoyl derivatives | 1.23 | + |
| Caffeic acid O-glucoside | 3.03 | 342.0867 | Negative | 341.0876 | 179.033 | Hydroxycinnamoyl derivatives | 3.09 | +++ |
| Sinapoyl O-hexoside | 3.27 | 403.155 | Positive | 404.155 | 175.4 | Phenolamides | <1 | ---- |
| 6-Hydroxymethylherniarin | 3.1 | 206.1 | Positive | 207.1 | 147.4 | Hydroxycinnamoyl derivatives | <1 | ---- |
| Caffeic acid | 3.14 | 180.0423 | Negative | 179.042 | 134.1 | Hydroxycinnamoyl derivatives | 3.11 | +++ |
| 5-O-p-Coumaroylquinic acid | 3.18 | 338.0 | Negative | 337 | 275.8 | Quinate and its derivatives | <1 | ---- |
| 1-O-β-d-Glucopyranosyl sinapate | 3.1 | 386.1 | Negative | 385.1 | 223.2 | Hydroxycinnamoyl derivatives | <1 | ---- |
| Syringic acid O-glucoside | 2.34 | 360.1 | Negative | 359.1 | 197.1 | Benzoic acid derivatives | <1 | ---- |
| 1-O-Feruloyl quinic acid | 2.94 | 368.1 | Positive | 369.1 | 207.5 | Quinate and its derivatives | 1.25 | + |
| Coumaroylquinic acid | 3.43 | 338.0 | Negative | 337 | 275.8 | Quinate and its derivatives | 2.19 | ++ |
| Esculin (6,7-Dihydroxycoumarin-6-glucoside) | 2.73 | 340.1 | Negative | 339.1 | 177.1 | Coumarins | <1 | ---- |
| p-Coumaric acid | 3.71 | 164.158 | Negative | 163.047 | 119 | Hydroxycinnamoyl derivatives | 3.11 | +++ |
| Protocatechuic acid O-glucoside | 4.99 | 316.0 | Negative | 315.071 | 153.018 | Catechin derivatives | <1 | ---- |
| Ferulic acid | 3.92 | 193.1766 | Negative | 193.058 | 134.1 | Hydroxycinnamoyl derivatives | 2.31 | ++ |
| Naringenin 7-O-neohesperidoside (Naringin) | 3.38 | 580.1792 | Positive | 581.1 | 449.2 | Flavanone | <1 | ---- |
| Selgin O-malonylhexoside | 3.45 | 564.1 | Positive | 565.1 | 317.8 | Flavone | <1 | ---- |
| Luteolin 6-C-glucoside | 3.47 | 448.1 | Positive | 449.1 | 300 | Flavone C-glycosides | <1 | ---- |
| Luteolin C-hexoside | 3.32 | 448.1 | Positive | 449.1 | 329.6 | Flavone C-glycosides | 1.04 | + |
| Luteolin 8-C-hexosyl-O-hexoside | 3.67 | 610.2 | Positive | 611.2 | 299.8 | Flavone C-glycosides | <1 | ---- |
| Chrysoeriol C-hexosyl | 3.31 | 462.1 | Positive | 463.1 | 301.8 | Flavone C-glycosides | <1 | ---- |
| Nobiletin | 3.89 | 402.1 | Positive | 403.1 | 373.1 | Flavone | <1 | ---- |
| lactucopicrin | 5.26 | 410.1366 | Negative | 409.137 | 213.1 | Others | <1 | ---- |
| lactucin | 3.38 | 276.2845 | Negative | 275.1 | 213 | Others | <1 | ---- |
| Eriodictyol 7-O-glucoside | 3.75 | 450.3928 | Negative | 449 | 151 | Flavone C-glycosides | <1 | ---- |
| Cultivar | S-1 | S-2 | B-1 | B-2 | S-3 | B-3 | |
|---|---|---|---|---|---|---|---|
| Ions | |||||||
| Ca | 8.275 ± 0.0760 e | 8.3375 ± 0.4904 e | 9.0875 ± 0.0661 d | 11.2167 ± 0.0711 c | 14.7915 ± 0.1909 a | 14.25 ± 0.0250 b | |
| Cu | 0.0148 ± 0.0002 a | 0.0148 ± 0.0001 a | 0.0157 ± 0.0001 a | 0.0149 ± 0.0001 a | 0.0152 ± 0.0002 a | 0.0151 ± 0.0002 a | |
| Fe | 0.1842 ± 0.0040 a | 0.1631 ± 0.0022 b | 0.1625 ± 0.0022 b | 0.1850 ± 0.0057 a | 0.1638 ± 0.0078 b | 0.1846 ± 0.0058 a | |
| K | 53.1667 ± 0.2887 b | 44.6250 ± 4.0292 e | 55.8750 ± 0.1250 a | 50.3750 ± 0.3307 c | 47.4167 ± 0.4732 d | 42.7083 ± 0.5052 e | |
| Mg | 4.6917 ± 0.0505 e | 4.6667 ± 0.0289 e | 5.8875 ± 0.0217 d | 6.7500 ± 0.0331 c | 8.6583 ± 0.1778 a | 8.5292 ± 0.0315 b | |
| Mn | 0.0301 ± 0.0002 d | 0.0379 ± 0.0008 a | 0.0384 ± 0.0002 a | 0.0291 ± 0.0001 e | 0.0323 ± 0.0008 c | 0.0348 ± 0.0001 b | |
| Na | 1.7083 ± 0.1512 dc | 1.4983 ± 0.1118 d | 2.5500 ± 0.0976 a | 2.0458 ± 0.1553 b | 1.8875 ± 0.0125 cb | 1.8417 ± 0.1697 cb | |
| P | 4.0500 ± 0.0500 c | 4.6000 ± 0.1083 a | 4.6750 ± 0.0125 a | 4.1750 ± 0.0331 b | 3.3333 ± 0.0473 d | 3.3208 ± 0.0072 d | |
| Zn | 0.0545 ± 0.0003 c | 0.0584 ± 0.0004 b | 0.0661 ± 0.0006 a | 0.0300 ± 0.0001 e | 0.0350 ± 0.0009 d | 0.0360 ± 0.0010 d | |
| Cultivar | DPPH Index a | ABTS Index a | T-AOC-FRAP Index a | Antioxidant Potency Composite Index b |
|---|---|---|---|---|
| S-1 | 78.49 | 62.09 | 46.31 | 62.30 |
| S-2 | 39.33 | 21.12 | 31.62 | 30.69 |
| B-1 | 48.58 | 29.01 | 23.40 | 33.66 |
| B-2 | 100.00 | 100.00 | 97.25 | 99.08 |
| S-3 | 71.87 | 61.58 | 49.97 | 61.14 |
| B-3 | 98.97 | 96.44 | 100.00 | 98.47 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qin, X.-X.; Zhang, M.-Y.; Han, Y.-Y.; Hao, J.-H.; Liu, C.-J.; Fan, S.-X. Beneficial Phytochemicals with Anti-Tumor Potential Revealed through Metabolic Profiling of New Red Pigmented Lettuces (Lactuca sativa L.). Int. J. Mol. Sci. 2018, 19, 1165. https://doi.org/10.3390/ijms19041165
Qin X-X, Zhang M-Y, Han Y-Y, Hao J-H, Liu C-J, Fan S-X. Beneficial Phytochemicals with Anti-Tumor Potential Revealed through Metabolic Profiling of New Red Pigmented Lettuces (Lactuca sativa L.). International Journal of Molecular Sciences. 2018; 19(4):1165. https://doi.org/10.3390/ijms19041165
Chicago/Turabian StyleQin, Xiao-Xiao, Ming-Yue Zhang, Ying-Yan Han, Jing-Hong Hao, Chao-Jie Liu, and Shuang-Xi Fan. 2018. "Beneficial Phytochemicals with Anti-Tumor Potential Revealed through Metabolic Profiling of New Red Pigmented Lettuces (Lactuca sativa L.)" International Journal of Molecular Sciences 19, no. 4: 1165. https://doi.org/10.3390/ijms19041165
APA StyleQin, X.-X., Zhang, M.-Y., Han, Y.-Y., Hao, J.-H., Liu, C.-J., & Fan, S.-X. (2018). Beneficial Phytochemicals with Anti-Tumor Potential Revealed through Metabolic Profiling of New Red Pigmented Lettuces (Lactuca sativa L.). International Journal of Molecular Sciences, 19(4), 1165. https://doi.org/10.3390/ijms19041165