Rothmund-Thomson Syndrome: Insights from New Patients on the Genetic Variability Underpinning Clinical Presentation and Cancer Outcome
Abstract
1. Introduction
2. Results
2.1. Clinical Report of RTS Patients
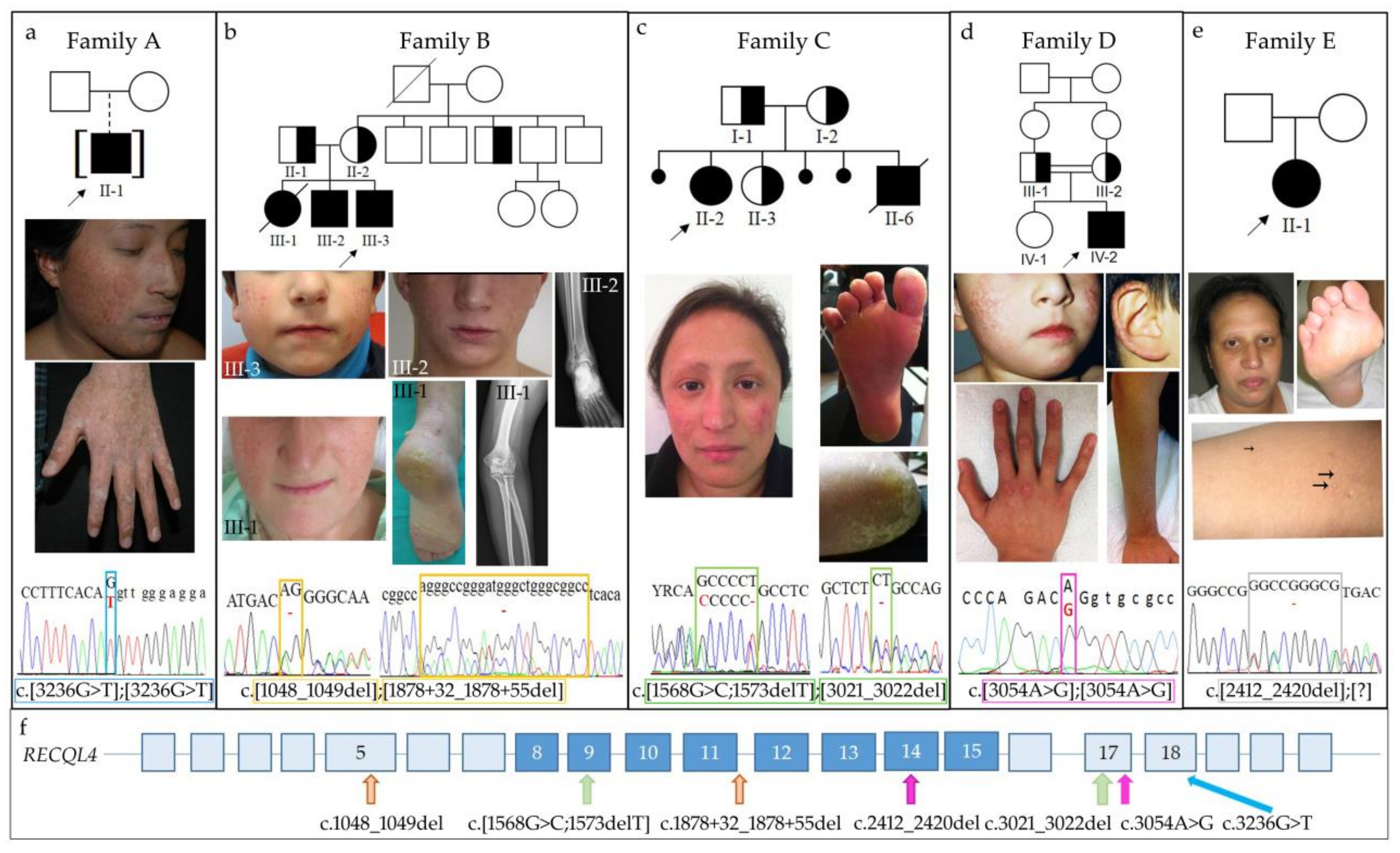
2.1.1. Family A
2.1.2. Family B
2.1.3. Family C
2.1.4. Family D
2.1.5. Family E
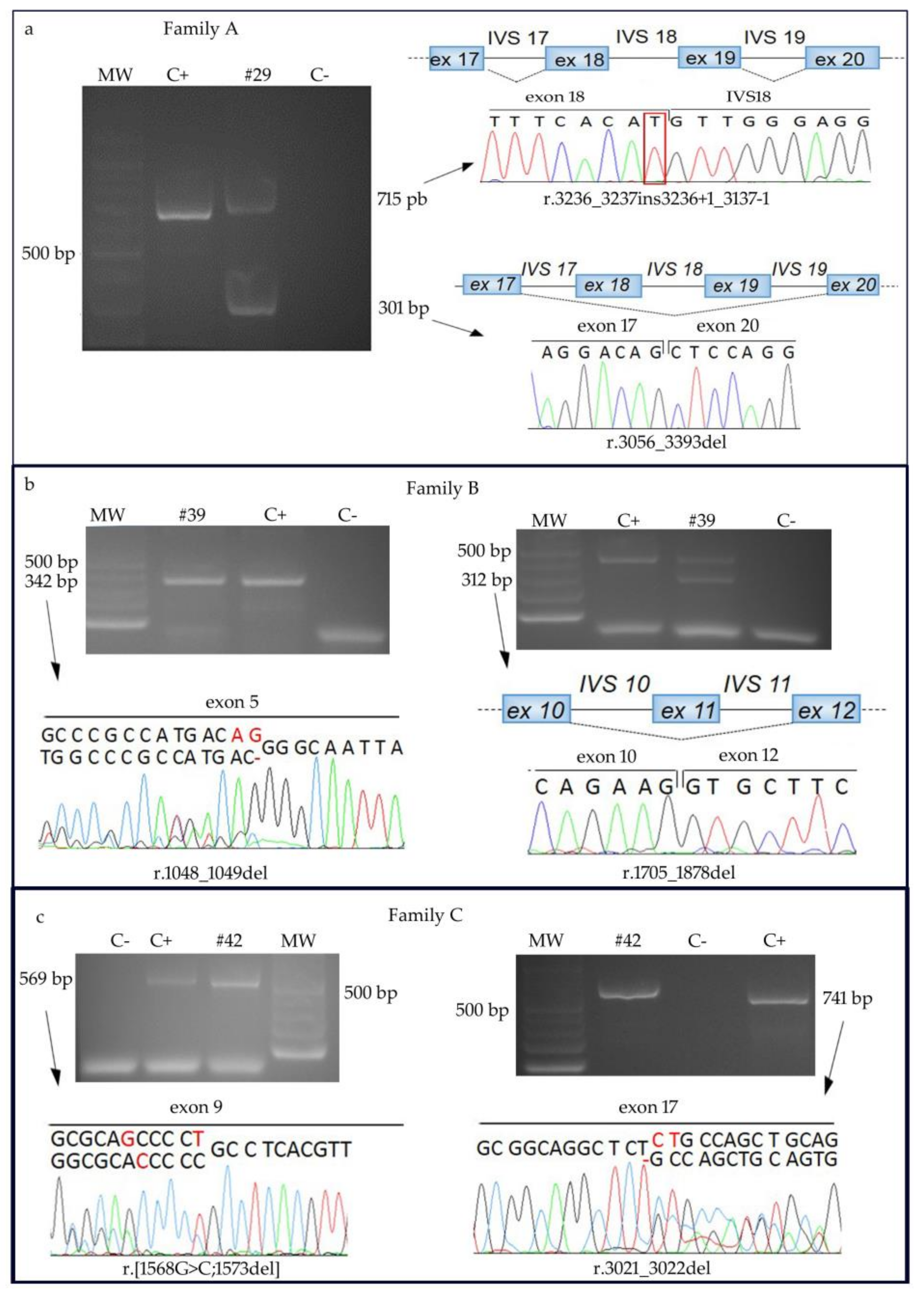
2.2. Molecular Characterization of RTS Patients
3. Discussion
4. Materials and Methods
4.1. Biological Material from Affected and Healthy Individuals from RTS Families
4.2. DNA Isolation and RECQL4 Mutational Analysis
4.3. Cell Cultures
4.4. RNA Isolation, RT-PCR and cDNA Analysis
4.5. Bioinformatic Analyses
4.6. High-Resolution CGH-Array
4.7. SNP-Array
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Larizza, L.; Roversi, G.; Volpi, L. Rothmund-Thomson syndrome. Orphanet J. Rare Dis. 2010, 5, 2. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.L.; Plon, S.E. Rothmund-Thomson Syndrome. In GeneReviews®; Adam, M.P., Ardinger, H.H., Pagon, R.A., Wallace, S.E., Bean, L.J., Stephens, K., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Croteau, D.L.; Singh, D.K.; Hoh Ferrarelli, L.; Lu, H.; Bohr, V.A. RECQL4 in genomic instability and aging. Trends Genet. 2012, 28, 624–631. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Jin, W.; Liu, H.; Wang, L.L. RECQ DNA helicases and osteosarcoma. Adv. Exp. Med. Biol. 2014, 804, 129–145. [Google Scholar] [CrossRef] [PubMed]
- Stinco, G.; Governatori, G.; Mattighello, P.; Patrone, P. Multiple cutaneous neoplasms in a patient with Rothmund-Thomson syndrome: Case report and published work review. J. Dermatol. 2008, 35, 154–161. [Google Scholar] [CrossRef] [PubMed]
- Chu, W.K.; Hickson, I.D. RecQ helicases: Multifunctional genome caretakers. Nat. Rev. Cancer 2009, 9, 644–654. [Google Scholar] [CrossRef] [PubMed]
- Van Maldergem, L.; Piard, J.; Larizza, L.; Wang, L.L. RECQL4-Related Recessive Conditions. In Epstein’s Inborn Errors of Development: The Molecular Basis of Clinical Disorders of Morphogenesis; Erickson, R.P., Wynshaw-Boris, A.J., Eds.; Oxford University Press: New York, NY, USA, 2016; pp. 1141–1147. ISBN 978-0-19-027542-6. [Google Scholar]
- Siitonen, H.A.; Sotkasiira, J.; Biervliet, M.; Benmansour, A.; Capri, Y.; Cormier-Daire, V.; Crandall, B.; Hannula-Jouppi, K.; Hennekam, R.; Herzog, D.; et al. The mutation spectrum in RECQL4 diseases. Eur. J. Hum. Genet. 2009, 17, 151–158. [Google Scholar] [CrossRef] [PubMed]
- Van Maldergem, L. Baller-Gerold Syndrome. In GeneReviews®; Adam, M.P., Ardinger, H.H., Pagon, R.A., Wallace, S.E., Bean, L.J., Stephens, K., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Sangrithi, M.N.; Bernal, J.A.; Madine, M.; Philpott, A.; Lee, J.; Dunphy, W.G.; Venkitaraman, A.R. Initiation of DNA replication requires the RECQL4 protein mutated in Rothmund-Thomson syndrome. Cell 2005, 121, 887–898. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Shamanna, R.A.; Keijzers, G.; Anand, R.; Rasmussen, L.J.; Cejka, P.; Croteau, D.L.; Bohr, V.A. RECQL4 Promotes DNA End Resection in Repair of DNA Double-Strand Breaks. Cell Rep. 2016, 16, 161–173. [Google Scholar] [CrossRef] [PubMed]
- Fan, W.; Luo, J. RecQ4 facilitates UV light-induced DNA damage repair through interaction with nucleotide excision repair factor xeroderma pigmentosum group A (XPA). J. Biol. Chem. 2008, 283, 29037–29044. [Google Scholar] [CrossRef] [PubMed]
- Schurman, S.H.; Hedayati, M.; Wang, Z.; Singh, D.K.; Speina, E.; Zhang, Y.; Becker, K.; Macris, M.; Sung, P.; Wilson, D.M.; et al. Direct and indirect roles of RECQL4 in modulating base excision repair capacity. Hum. Mol. Genet. 2009, 18, 3470–3483. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, A.K.; Rossi, M.L.; Singh, D.K.; Dunn, C.; Ramamoorthy, M.; Croteau, D.L.; Liu, Y.; Bohr, V.A. RECQL4, the protein mutated in Rothmund-Thomson syndrome, functions in telomere maintenance. J. Biol. Chem. 2012, 287, 196–209. [Google Scholar] [CrossRef] [PubMed]
- De, S.; Kumari, J.; Mudgal, R.; Modi, P.; Gupta, S.; Futami, K.; Goto, H.; Lindor, N.M.; Furuichi, Y.; Mohanty, D.; et al. RECQL4 is essential for the transport of p53 to mitochondria in normal human cells in the absence of exogenous stress. J. Cell Sci. 2012, 125, 2509–2522. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; De, S.; Srivastava, V.; Hussain, M.; Kumari, J.; Muniyappa, K.; Sengupta, S. RECQL4 and p53 potentiate the activity of polymerase γ and maintain the integrity of the human mitochondrial genome. Carcinogenesis 2014, 35, 34–45. [Google Scholar] [CrossRef] [PubMed]
- Burks, L.M.; Yin, J.; Plon, S.E. Nuclear import and retention domains in the amino terminus of RECQL4. Gene 2007, 391, 26–38. [Google Scholar] [CrossRef] [PubMed]
- Dietschy, T.; Shevelev, I.; Pena-Diaz, J.; Hühn, D.; Kuenzle, S.; Mak, R.; Miah, M.F.; Hess, D.; Fey, M.; Hottiger, M.O.; et al. p300-mediated acetylation of the Rothmund-Thomson-syndrome gene product RECQL4 regulates its subcellular localization. J. Cell Sci. 2009, 122, 1258–1267. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kaiser, S.; Sauer, F.; Kisker, C. The structural and functional characterization of human RecQ4 reveals insights into its helicase mechanism. Nat. Commun. 2017, 8, 15907. [Google Scholar] [CrossRef] [PubMed]
- Mojumdar, A.; De March, M.; Marino, F.; Onesti, S. The Human RecQ4 Helicase Contains a Functional RecQ C-terminal Region (RQC) That Is Essential for Activity. J. Biol. Chem. 2017, 292, 4176–4184. [Google Scholar] [CrossRef] [PubMed]
- Chi, Z.; Nie, L.; Peng, Z.; Yang, Q.; Yang, K.; Tao, J.; Mi, Y.; Fang, X.; Balajee, A.S.; Zhao, Y. RecQL4 cytoplasmic localization: Implications in mitochondrial DNA oxidative damage repair. Int. J. Biochem. Cell Biol. 2012, 44, 1942–1951. [Google Scholar] [CrossRef] [PubMed]
- Abe, T.; Yoshimura, A.; Hosono, Y.; Tada, S.; Seki, M.; Enomoto, T. The N-terminal region of RECQL4 lacking the helicase domain is both essential and sufficient for the viability of vertebrate cells. Role of the N-terminal region of RECQL4 in cells. Biochim. Biophys. Acta 2011, 1813, 473–479. [Google Scholar] [CrossRef] [PubMed]
- Wessels, J.A.M.; de Vries-Bouwstra, J.K.; Heijmans, B.T.; Slagboom, P.E.; Goekoop-Ruiterman, Y.P.M.; Allaart, C.F.; Kerstens, P.J.S.M.; van Zeben, D.; Breedveld, F.C.; Dijkmans, B.A.C.; et al. Efficacy and toxicity of methotrexate in early rheumatoid arthritis are associated with single-nucleotide polymorphisms in genes coding for folate pathway enzymes. Arthritis Rheum. 2006, 54, 1087–1095. [Google Scholar] [CrossRef] [PubMed]
- De Somer, L.; Wouters, C.; Morren, M.-A.; De Vos, R.; Van Den Oord, J.; Devriendt, K.; Meyts, I. Granulomatous skin lesions complicating Varicella infection in a patient with Rothmund-Thomson syndrome and immune deficiency: Case report. Orphanet J. Rare Dis. 2010, 5, 37. [Google Scholar] [CrossRef] [PubMed]
- Guerrero-González, G.A.; Martínez-Cabriales, S.A.; Hernández-Juárez, A.A.; de Jesús Lugo-Trampe, J.; Espinoza-González, N.A.; Gómez-Flores, M.; Ocampo-Candiani, J. Rothmund-thomson syndrome: A 13-year follow-up. Case Rep. Dermatol. 2014, 6, 176–179. [Google Scholar] [CrossRef] [PubMed]
- Suter, A.-A.; Itin, P.; Heinimann, K.; Ahmed, M.; Ashraf, T.; Fryssira, H.; Kini, U.; Lapunzina, P.; Miny, P.; Sommerlund, M.; et al. Rothmund-Thomson Syndrome: Novel pathogenic mutations and frequencies of variants in the RECQL4 and USB1 (C16orf57) gene. Mol. Genet. Genom. Med. 2016, 4, 359–366. [Google Scholar] [CrossRef] [PubMed]
- Van Rij, M.C.; Grijsen, M.L.; Appelman-Dijkstra, N.M.; Hansson, K.B.M.; Ruivenkamp, C.L.; Mulder, K.; van Doorn, R.; Oranje, A.P.; Kant, S.G. Rothmund-Thomson syndrome and osteoma cutis in a patient previously diagnosed as COPS syndrome. Eur. J. Pediatr. 2017, 176, 279–283. [Google Scholar] [CrossRef] [PubMed]
- Cabral, R.E.C.; Queille, S.; Bodemer, C.; de Prost, Y.; Neto, J.B.C.; Sarasin, A.; Daya-Grosjean, L. Identification of new RECQL4 mutations in Caucasian Rothmund-Thomson patients and analysis of sensitivity to a wide range of genotoxic agents. Mutat. Res. 2008, 643, 41–47. [Google Scholar] [CrossRef] [PubMed]
- Piard, J.; Aral, B.; Vabres, P.; Holder-Espinasse, M.; Mégarbané, A.; Gauthier, S.; Capra, V.; Pierquin, G.; Callier, P.; Baumann, C.; et al. Search for ReCQL4 mutations in 39 patients genotyped for suspected Rothmund-Thomson/Baller-Gerold syndromes. Clin. Genet. 2015, 87, 244–251. [Google Scholar] [CrossRef] [PubMed]
- Salih, A.; Inoue, S.; Onwuzurike, N. Rothmund-Thomson syndrome (RTS) with osteosarcoma due toRECQL4mutation. BMJ Case Rep. 2018, 2018. [Google Scholar] [CrossRef] [PubMed]
- Van Maldergem, L.; Siitonen, H.A.; Jalkh, N.; Chouery, E.; De Roy, M.; Delague, V.; Muenke, M.; Jabs, E.W.; Cai, J.; Wang, L.L.; et al. Revisiting the craniosynostosis-radial ray hypoplasia association: Baller-Gerold syndrome caused by mutations in the RECQL4 gene. J. Med. Genet. 2006, 43, 148–152. [Google Scholar] [CrossRef] [PubMed]
- Simon, T.; Kohlhase, J.; Wilhelm, C.; Kochanek, M.; De Carolis, B.; Berthold, F. Multiple malignant diseases in a patient with Rothmund-Thomson syndrome with RECQL4 mutations: Case report and literature review. Am. J. Med. Genet. A 2010, 152, 1575–1579. [Google Scholar] [CrossRef] [PubMed]
- Padhy, D.; Madhuri, V.; Pulimood, S.A.; Danda, S.; Walter, N.M.; Wang, L.L. Metatarsal osteosarcoma in Rothmund-Thomson syndrome: A case report. J. Bone Joint Surg. Am. 2010, 92, 726–730. [Google Scholar] [CrossRef] [PubMed]
- Debeljak, M.; Zver, A.; Jazbec, J. A patient with Baller-Gerold syndrome and midline NK/T lymphoma. Am. J. Med. Genet. A 2009, 149, 755–759. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.L.; Gannavarapu, A.; Kozinetz, C.A.; Levy, M.L.; Lewis, R.A.; Chintagumpala, M.M.; Ruiz-Maldanado, R.; Contreras-Ruiz, J.; Cunniff, C.; Erickson, R.P.; et al. Association between osteosarcoma and deleterious mutations in the RECQL4 gene in Rothmund-Thomson syndrome. J. Natl. Cancer Inst. 2003, 95, 669–674. [Google Scholar] [CrossRef] [PubMed]
- Siitonen, H.A.; Kopra, O.; Kääriäinen, H.; Haravuori, H.; Winter, R.M.; Säämänen, A.-M.; Peltonen, L.; Kestilä, M. Molecular defect of RAPADILINO syndrome expands the phenotype spectrum of RECQL diseases. Hum. Mol. Genet. 2003, 12, 2837–2844. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Yan, M.; Cheng, R.; Ni, C.; Liang, J.; Li, M.; Yao, Z. A rare RECQL4 indel mutation in a Chinese patient with Rothmund-Thomson syndrome. J. Eur. Acad. Dermatol. Venereol. 2016, 30, e159–e161. [Google Scholar] [CrossRef] [PubMed]
- Colombo, E.A.; Fontana, L.; Roversi, G.; Negri, G.; Castiglia, D.; Paradisi, M.; Zambruno, G.; Larizza, L. Novel physiological RECQL4 alternative transcript disclosed by molecular characterisation of Rothmund-Thomson Syndrome sibs with mild phenotype. Eur. J. Hum. Genet. 2014, 22, 1298–1304. [Google Scholar] [CrossRef] [PubMed]
- Jensen, M.B.; Dunn, C.A.; Keijzers, G.; Kulikowicz, T.; Rasmussen, L.J.; Croteau, D.L.; Bohr, V.A. The helicase and ATPase activities of RECQL4 are compromised by mutations reported in three human patients. Aging 2012, 4, 790–802. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Den Dunnen, J.T.; Dalgleish, R.; Maglott, D.R.; Hart, R.K.; Greenblatt, M.S.; McGowan-Jordan, J.; Roux, A.-F.; Smith, T.; Antonarakis, S.E.; Taschner, P.E.M. HGVS Recommendations for the Description of Sequence Variants: 2016 Update. Hum. Mutat. 2016, 37, 564–569. [Google Scholar] [CrossRef] [PubMed]
- LOVD v.2.0: The Next Generation in Gene Variant Databases. Wiley Online Library. 2011. Available online: http://onlinelibrary.wiley.com/doi/10.1002/humu.21438/abstract (accessed on 22 February 2018).
- The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar] [CrossRef]
- Kumar, P.; Henikoff, S.; Ng, P.C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 2009, 4, 1073–1081. [Google Scholar] [CrossRef] [PubMed]
- Adzhubei, I.A.; Schmidt, S.; Peshkin, L.; Ramensky, V.E.; Gerasimova, A.; Bork, P.; Kondrashov, A.S.; Sunyaev, S.R. A method and server for predicting damaging missense mutations. Nat. Methods 2010, 7, 248–249. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Sims, G.E.; Murphy, S.; Miller, J.R.; Chan, A.P. Predicting the Functional Effect of Amino Acid Substitutions and Indels. PLoS ONE 2012, 7, e46688. [Google Scholar] [CrossRef] [PubMed]
- Reva, B.; Antipin, Y.; Sander, C. Predicting the functional impact of protein mutations: Application to cancer genomics. Nucleic Acids Res. 2011, 39, e118. [Google Scholar] [CrossRef] [PubMed]
- Wildeman, M.; van Ophuizen, E.; den Dunnen, J.T.; Taschner, P.E.M. Improving sequence variant descriptions in mutation databases and literature using the Mutalyzer sequence variation nomenclature checker. Hum. Mutat. 2008, 29, 6–13. [Google Scholar] [CrossRef] [PubMed]
- Smith, P.J.; Zhang, C.; Wang, J.; Chew, S.L.; Zhang, M.Q.; Krainer, A.R. An increased specificity score matrix for the prediction of SF2/ASF-specific exonic splicing enhancers. Hum. Mol. Genet. 2006, 15, 2490–2508. [Google Scholar] [CrossRef] [PubMed]
- Desmet, F.-O.; Hamroun, D.; Lalande, M.; Collod-Béroud, G.; Claustres, M.; Béroud, C. Human Splicing Finder: An online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 2009, 37, e67. [Google Scholar] [CrossRef] [PubMed]
- Hebsgaard, S.M.; Korning, P.G.; Tolstrup, N.; Engelbrecht, J.; Rouzé, P.; Brunak, S. Splice site prediction in Arabidopsis thaliana pre-mRNA by combining local and global sequence information. Nucleic Acids Res. 1996, 24, 3439–3452. [Google Scholar] [CrossRef] [PubMed]
- Miller, D.T.; Adam, M.P.; Aradhya, S.; Biesecker, L.G.; Brothman, A.R.; Carter, N.P.; Church, D.M.; Crolla, J.A.; Eichler, E.E.; Epstein, C.J.; et al. Consensus statement: Chromosomal microarray is a first-tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies. Am. J. Hum. Genet. 2010, 86, 749–764. [Google Scholar] [CrossRef] [PubMed]
- Kearney, H.M.; Thorland, E.C.; Brown, K.K.; Quintero-Rivera, F.; South, S.T.; Working Group of the American College of Medical Genetics Laboratory Quality Assurance Committee. American College of Medical Genetics standards and guidelines for interpretation and reporting of postnatal constitutional copy number variants. Genet. Med. 2011, 13, 680–685. [Google Scholar] [CrossRef] [PubMed]

| Family | A | B | C | D | E | |||
|---|---|---|---|---|---|---|---|---|
| Pedigree position | II-1 | III-1 | III-2 | III-3 | II-2 | II-6 | IV-2 | II-1 |
| Index case code | #29 | - | - | #39 | #42 | - | #19 | #38 |
| Birth/death | 1995 | 1989-2013 | 1993 | 2008 | 1986 | 1994-2012 | 1997 | 1980 |
| Sex | M | F | M | M | F | M | M | F |
| Origin | Ecuador | Spain | Italy | Turkey | Belgium | |||
| Growth delay | + | + | + | - | + | + | - | + |
| Poikiloderma | + | + | + | + | + | + | + | - * |
| Onset (age) | 6–7 m | 1 y | <1 y | 18 m | At birth | At birth | 2 y | - |
| First localization | Sun-exposed areas | Cheeks | Cheeks | Cheeks | Face | Cheeks | Sun-exposed areas | - |
| Hyperkeratosis | Palmo-plantar | Plantar | - | - | Plantar | Palmo-plantar | Palms and joints | Plantar |
| Photosensitivity | Only in infancy | + | + | - | + | + | - | - |
| Hair | Normal | Thin | Thin | Normal | Sparse | Sparse | Normal | Sparse |
| Eyelashes | Normal | Sparse | Sparse | Normal | Absent | Absent | Normal | Sparse |
| Eyebrows | Sparse | Normal | Absent | Normal | Sparse | Sparse | Normal | Sparse |
| Onychodystrophy | + | - | - | - | + | + | Only in infancy | - |
| Dental defects | - | - | - | - | + | + | Irregular end | Enamel defect |
| Skeletal anomalies | Low bone density | n.a. | Low bone density | - | Osteosclerosis; Cystic-like lesion | - | n.a. | Osteopenia |
| Gastrointestinal | - | Constipation GER | - | - | Diarrhea; Food intolerance in infancy | Diarrhea; Food intolerance in infancy | - | Diarrhea in infancy |
| Cancer (onset age) | - | Olecranon OS (23 y) | Ankle OS (19 y) | - | - | Ulnar OS (14 y); Femur OS (17 y) | - | Alveolar rhabdomyosarcoma (12 y) |
| Others | CD4/CD8 = 2.5 keratoconus | - | Tibiotalar joint degenerative changes | - | - | - | Recurrent middle ear infections; IgA deficiency; Knee arthritis | Hypogonadism; Chronic anemia; Hyper-ferritinemia; Hyper-cholesterolemia; Insulin resistance |
| RECQL4 genotype | c.[3236G>T]; [3236G>T] | c.[1048_1049del]; [1878+32_1878+55del] | c.[1568G>C;1573delT]; [3021_3022del] | c.[3054A>G]; [3054A>G] | c.[2412_2420del]; [?] | |||
| Patient Code | Age at Analysis | Cancer | Pathogenic Variant I | Pathogenic Variant II | Reference | ||
|---|---|---|---|---|---|---|---|
| FCP-195 | 1 y | - | c.1048_1049del | ex 5 | p.(Q757Ter) | ex 14 | [8] |
| Pt 10 | 13 y | - | c.1048_1049del | ex 5 | p.(Q757Ter) | ex 14 | [8] |
| Pt 13 | 4 y | - | c.1048_1049del | ex 5 | p.(Gln800Ter) | ex 14 | [8] |
| RTS | 13 y | - | c.1048_1049del | ex 5 | p.(Gln757Ter) | ex 14 | [25] |
| RTS | 6 y | - | c.1048_1049del | ex 5 | c.1391-1G>A | IVS7 | [24] |
| Pt 13 | 5 y | - | c.1048_1049del | ex 5 | p.(Gln800Ter) | ex 14 | [26] |
| RTS 1 | 34 y | HL 35 y | c.1048_1049del | ex 5 | c.1391-1G>A | IVS7 | [27] |
| RTS 2 | 5 y | - | |||||
| #39 III-1 | 24 y † | OS 23 y | c.1048_1049del | ex 5 | c.1878+32_1878+55del | IVS11 | This work |
| #39 III-2 | 22 y | OS 19 y | |||||
| #39 III-3 | 7 y | - | |||||
| FCP-210 | - | OS 8 y | c.1718delA | ex 11 | c.1878+32_1878+55del | IVS11 | [8] |
| #42 II-2 | 31 y | - | c.[1568G>C;1573delT] | ex 9 | c.3021_3022del | ex 17 | This work |
| #42 II-6 | 18 y † | OS 14 y, 17 y | |||||
| AS517 | - | OS 13 y | c.[1568G>C;1573delT] | ex 9 | p.(Leu926Arg) | ex 16 | [28] |
| AS518 | 10 y | - | |||||
| AS287 | 32 y | - | c.[1568G>C;1573delT] | ex 9 | p.(Gln757Ter) | ex 14 | [28] |
| RTS | 14 y | OS 10 y | c.[1568G>C;1573delT] | ex 9 | p.(Gln757Ter) | ex 14 | [30] |
| II-1 | 21 y | OS 21 y | c.[1568G>C;1573delT] | ex 9 | c.1391-1G>A | IVS7 | [8] |
| II-2 | 9 y † | OS 7 y | |||||
| FCP-129 | - | OS 4 y | c.[1568G>C;1573delT] | ex 9 | p.(Gln757Ter) | ex 14 | [8] |
| FCP-153 | - | OS 20 y | c.[1568G>C;1573delT] | ex 9 | c.1391-1G>A | IVS7 | [8] |
| FCP-153 sibling | - | OS 9 y | |||||
| Pt 1 | 19 y | - | c.[1568G>C;1573delT] | ex 9 | c.2059-1G>C | IVS12 | [8] |
| FCP-157 | 10 y | - | c.[1568G>C;1573delT] | ex 9 | p.(Gln757Ter) | ex 14 | [8] |
| FCP-167 | 14 y | - | c.[1568G>C;1573delT] | ex 9 | c.3270delG | ex 19 | [8] |
| FCP-175 | 2 y | - | c.[1568G>C;1573delT] | ex 9 | p.(Gln1175Ter) | ex 21 | [8] |
| Pt 1 | 9 y | - | c.[1568G>C;1573delT] | ex 9 | p.(Arg1021Trp) | ex 18 | [8] |
| Pt 9 | 12 y | - | c.[1568G>C;1573delT] | ex 9 | c.84+6del16 | IVS1 | [8] |
| Pt 11 | 11 y | - | c.[1568G>C;1573delT] | ex 9 | p.(Gln821Ter) | ex 14 | [8] |
| Pt 2 | 6 y | - | c.[1568G>C;1573delT] | ex 9 | p.(Cys511Arg) | ex 9 | [29] |
| Pt 3 | 6 y | - | c.[1568G>C;1573delT] | ex 9 | p.(Trp412Ter) | ex 6 | [29] |
| Pt 6 | 21 y | - | c.[1568G>C;1573delT] | ex 9 | c.2059-1G>A | IVS12 | [29] |
| Pt 9 | 3 y | - | c.[1568G>C;1573delT] | ex 9 | p.(Arg1021Trp) | ex 18 | [29] |
| Pt 14 | 8 y | - | c.[1568G>C;1573delT] | ex 9 | c.1930_1935dup | ex 12 | [26] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Colombo, E.A.; Locatelli, A.; Cubells Sánchez, L.; Romeo, S.; Elcioglu, N.H.; Maystadt, I.; Esteve Martínez, A.; Sironi, A.; Fontana, L.; Finelli, P.; et al. Rothmund-Thomson Syndrome: Insights from New Patients on the Genetic Variability Underpinning Clinical Presentation and Cancer Outcome. Int. J. Mol. Sci. 2018, 19, 1103. https://doi.org/10.3390/ijms19041103
Colombo EA, Locatelli A, Cubells Sánchez L, Romeo S, Elcioglu NH, Maystadt I, Esteve Martínez A, Sironi A, Fontana L, Finelli P, et al. Rothmund-Thomson Syndrome: Insights from New Patients on the Genetic Variability Underpinning Clinical Presentation and Cancer Outcome. International Journal of Molecular Sciences. 2018; 19(4):1103. https://doi.org/10.3390/ijms19041103
Chicago/Turabian StyleColombo, Elisa A., Andrea Locatelli, Laura Cubells Sánchez, Sara Romeo, Nursel H. Elcioglu, Isabelle Maystadt, Altea Esteve Martínez, Alessandra Sironi, Laura Fontana, Palma Finelli, and et al. 2018. "Rothmund-Thomson Syndrome: Insights from New Patients on the Genetic Variability Underpinning Clinical Presentation and Cancer Outcome" International Journal of Molecular Sciences 19, no. 4: 1103. https://doi.org/10.3390/ijms19041103
APA StyleColombo, E. A., Locatelli, A., Cubells Sánchez, L., Romeo, S., Elcioglu, N. H., Maystadt, I., Esteve Martínez, A., Sironi, A., Fontana, L., Finelli, P., Gervasini, C., Pecile, V., & Larizza, L. (2018). Rothmund-Thomson Syndrome: Insights from New Patients on the Genetic Variability Underpinning Clinical Presentation and Cancer Outcome. International Journal of Molecular Sciences, 19(4), 1103. https://doi.org/10.3390/ijms19041103







