Signaling by Steroid Hormones in the 3D Nuclear Space
Abstract
1. Introduction
2. Genome 3D Organization
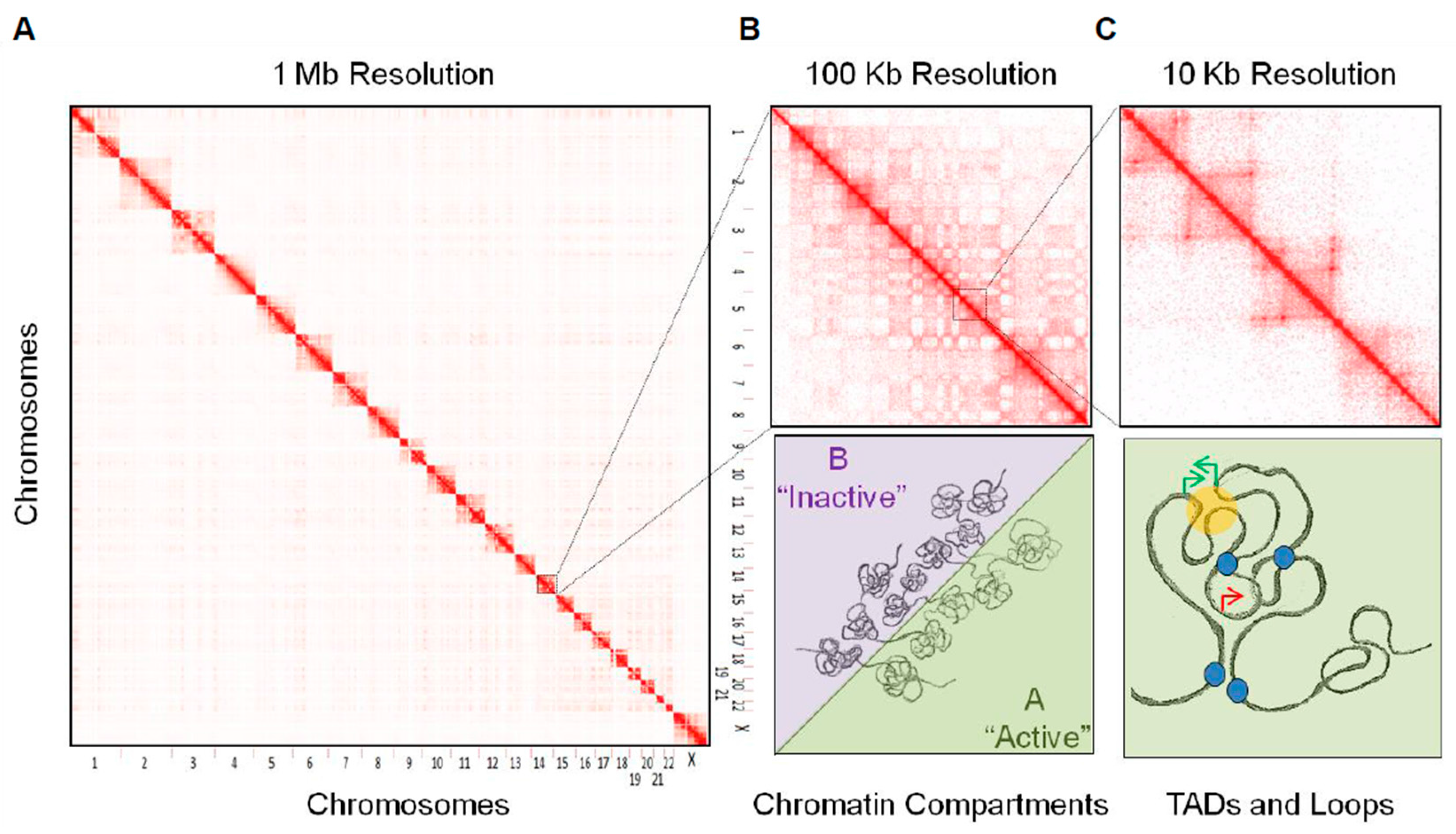
2.1. Chromosome Territories
2.2. Chromatin Compartments
2.3. Topologically Associating Domains
2.4. Sub-Domains and Chromatin Loops
3. 3D Genome Folding Modulates the Response to Steroids
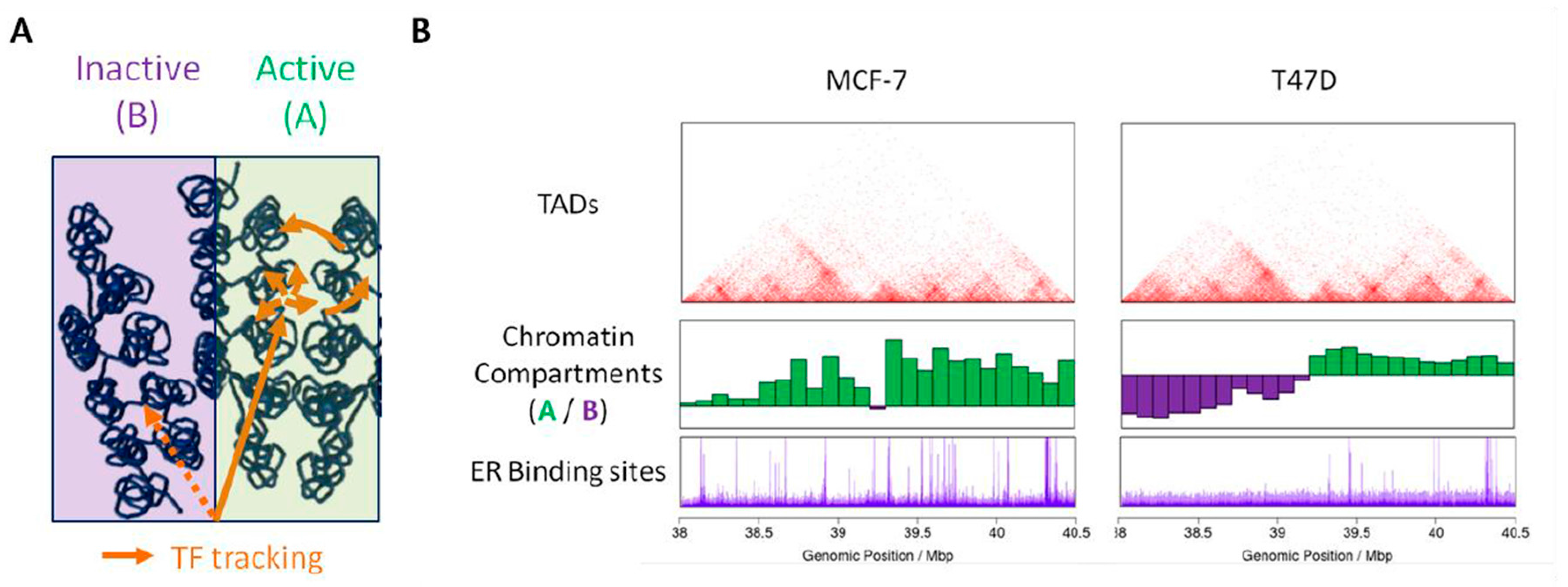
3.1. Steroid Receptors Cistrome
3.2. Differential Accessibility of Hormone Response Elements
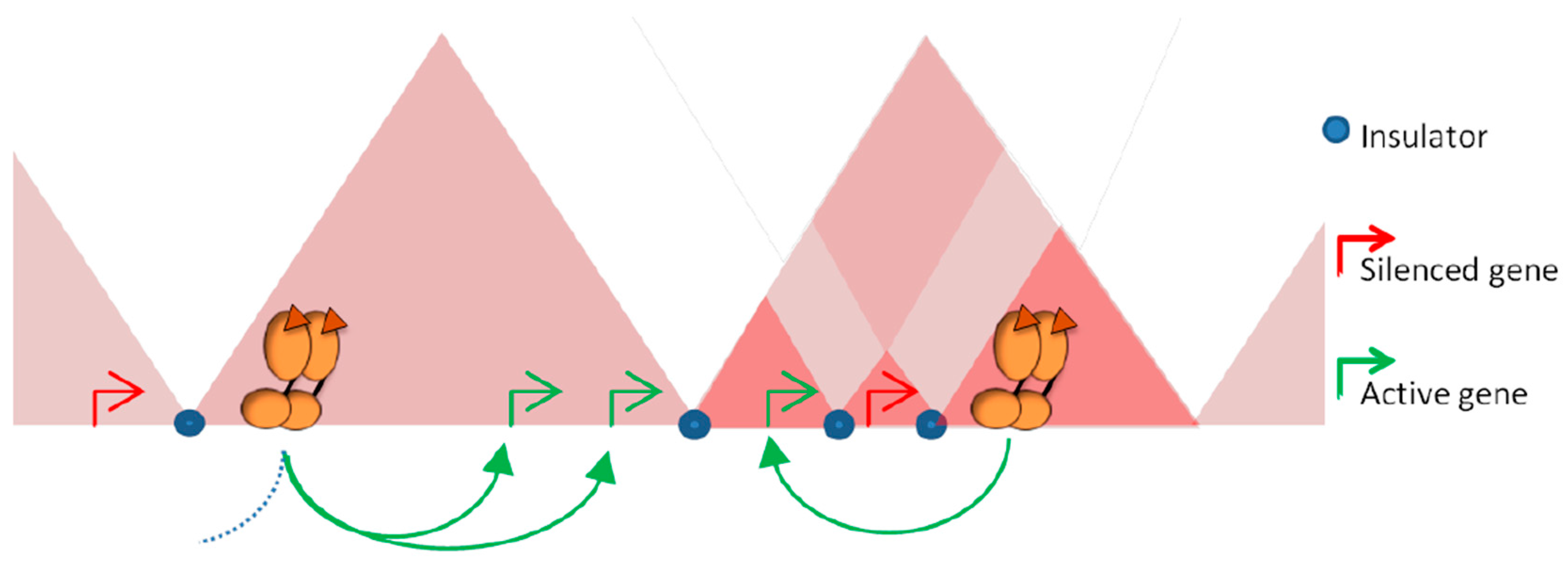
3.3. Topological Restraint of Promoter-Enhancer Looping
3.4. Influence of Architectural Proteins in Steroid Response
4. Steroid Receptors Mediated Genome (re)-Organization
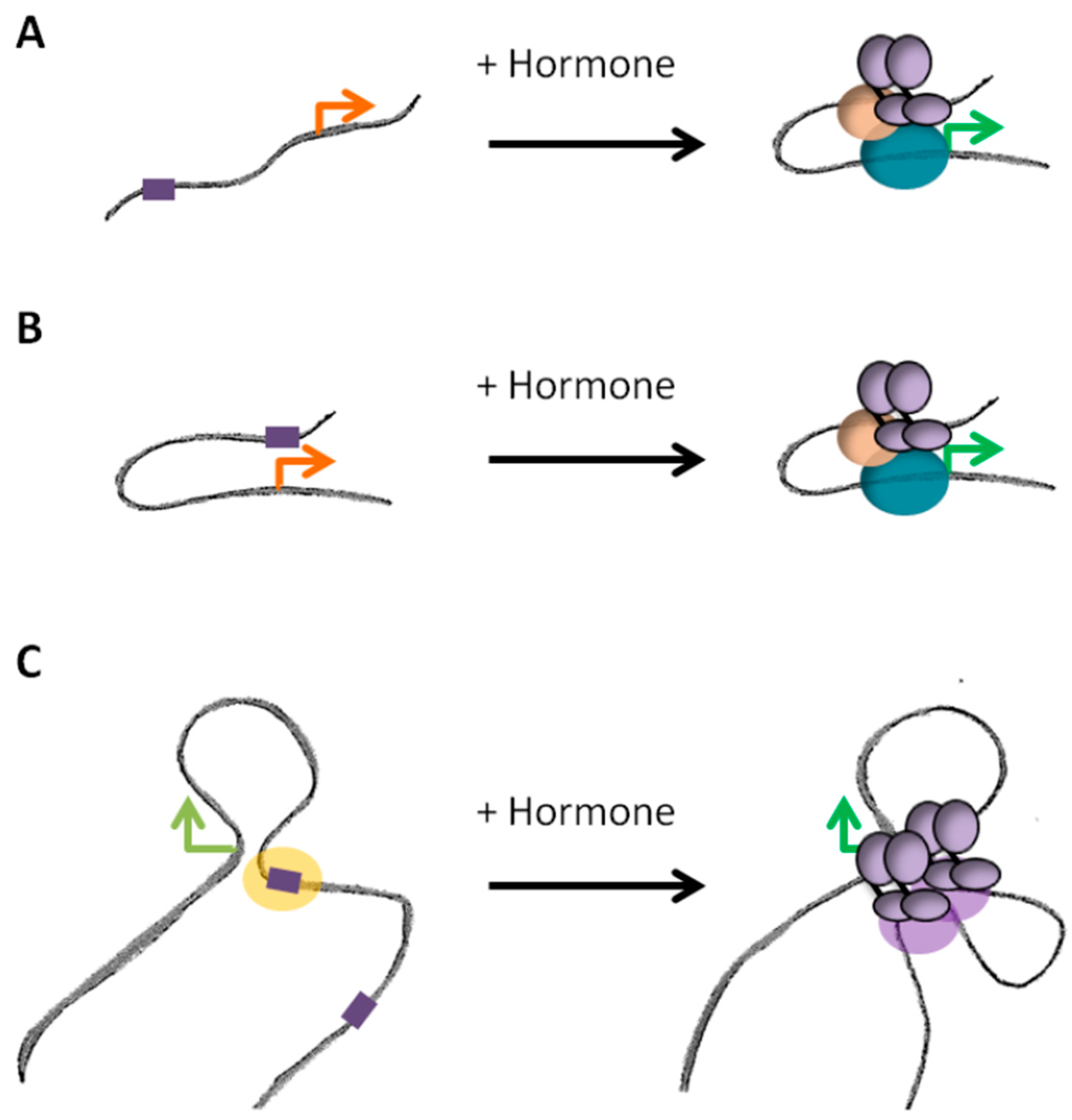
4.1. Steroid Receptors Dependent Promoter-Enhancer Loops
4.2. Dynamic and/or Pre-Settled Organization of Steroid Responsive Hubs
4.3. Steroid Induced Changes at Higher Levels of Organization
5. Future Directions
Acknowledgments
Conflicts of Interest
References
- Nilsson, S.; Makela, S.; Treuter, E.; Tujague, M.; Thomsen, J.; Andersson, G.; Enmark, E.; Pettersson, K.; Warner, M.; Gustafsson, J.A. Mechanisms of estrogen action. Physiol. Rev. 2001, 81, 1535–1565. [Google Scholar] [CrossRef] [PubMed]
- Mangelsdorf, D.J.; Thummel, C.; Beato, M.; Herrlich, P.; Schutz, G.; Umesono, K.; Blumberg, B.; Kastner, P.; Mark, M.; Chambon, P.; et al. The nuclear receptor superfamily: The second decade. Cell 1995, 83, 835–839. [Google Scholar] [CrossRef]
- Levin, E.R. Plasma membrane estrogen receptors. Trends Endocrinol. Metab. TEM 2009, 20, 477–482. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Chambon, P. The estrogen receptor binds tightly to its responsive element as a ligand-induced homodimer. Cell 1988, 55, 145–156. [Google Scholar] [CrossRef]
- Safe, S. Transcriptional activation of genes by 17β-estradiol through estrogen receptor-sp1 interactions. Vitam. Horm. 2001, 62, 231–252. [Google Scholar] [PubMed]
- Webb, P.; Nguyen, P.; Valentine, C.; Lopez, G.N.; Kwok, G.R.; McInerney, E.; Katzenellenbogen, B.S.; Enmark, E.; Gustafsson, J.A.; Nilsson, S.; et al. The estrogen receptor enhances ap-1 activity by two distinct mechanisms with different requirements for receptor transactivation functions. Mol. Endocrinol. 1999, 13, 1672–1685. [Google Scholar] [CrossRef] [PubMed]
- Perissi, V.; Rosenfeld, M.G. Controlling nuclear receptors: The circular logic of cofactor cycles. Nat. Rev. Mol. Cell Biol. 2005, 6, 542–554. [Google Scholar] [CrossRef] [PubMed]
- Metivier, R.; Penot, G.; Hubner, M.R.; Reid, G.; Brand, H.; Kos, M.; Gannon, F. Estrogen receptor-α directs ordered, cyclical, and combinatorial recruitment of cofactors on a natural target promoter. Cell 2003, 115, 751–763. [Google Scholar] [CrossRef]
- Shang, Y.; Hu, X.; DiRenzo, J.; Lazar, M.A.; Brown, M. Cofactor dynamics and sufficiency in estrogen receptor-regulated transcription. Cell 2000, 103, 843–852. [Google Scholar] [CrossRef]
- Cicatiello, L.; Scafoglio, C.; Altucci, L.; Cancemi, M.; Natoli, G.; Facchiano, A.; Iazzetti, G.; Calogero, R.; Biglia, N.; De Bortoli, M.; et al. A genomic view of estrogen actions in human breast cancer cells by expression profiling of the hormone-responsive transcriptome. J. Mol. Endocrinol. 2004, 32, 719–775. [Google Scholar] [CrossRef] [PubMed]
- Kininis, M.; Isaacs, G.D.; Core, L.J.; Hah, N.; Kraus, W.L. Postrecruitment regulation of RNA polymerase ii directs rapid signaling responses at the promoters of estrogen target genes. Mol. Cell. Biol. 2009, 29, 1123–1133. [Google Scholar] [CrossRef] [PubMed]
- Hah, N.; Danko, C.G.; Core, L.; Waterfall, J.J.; Siepel, A.; Lis, J.T.; Kraus, W.L. A rapid, extensive, and transient transcriptional response to estrogen signaling in breast cancer cells. Cell 2011, 145, 622–634. [Google Scholar] [CrossRef] [PubMed]
- Carroll, J.S.; Liu, X.S.; Brodsky, A.S.; Li, W.; Meyer, C.A.; Szary, A.J.; Eeckhoute, J.; Shao, W.; Hestermann, E.V.; Geistlinger, T.R.; et al. Chromosome-wide mapping of estrogen receptor binding reveals long-range regulation requiring the forkhead protein foxa1. Cell 2005, 122, 33–43. [Google Scholar] [CrossRef] [PubMed]
- Welboren, W.J.; van Driel, M.A.; Janssen-Megens, E.M.; van Heeringen, S.J.; Sweep, F.C.; Span, P.N.; Stunnenberg, H.G. CHIP-Seq of ERα and RNA polymerase ii defines genes differentially responding to ligands. EMBO J. 2009, 28, 1418–1428. [Google Scholar] [CrossRef] [PubMed]
- Ross-Innes, C.S.; Stark, R.; Holmes, K.A.; Schmidt, D.; Spyrou, C.; Russell, R.; Massie, C.E.; Vowler, S.L.; Eldridge, M.; Carroll, J.S. Cooperative interaction between retinoic acid receptor-α and estrogen receptor in breast cancer. Genes Dev. 2010, 24, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Ballare, C.; Castellano, G.; Gaveglia, L.; Althammer, S.; Gonzalez-Vallinas, J.; Eyras, E.; Le Dily, F.; Zaurin, R.; Soronellas, D.; Vicent, G.P.; et al. Nucleosome-driven transcription factor binding and gene regulation. Mol. Cell 2013, 49, 67–79. [Google Scholar] [CrossRef] [PubMed]
- John, S.; Sabo, P.J.; Thurman, R.E.; Sung, M.H.; Biddie, S.C.; Johnson, T.A.; Hager, G.L.; Stamatoyannopoulos, J.A. Chromatin accessibility pre-determines glucocorticoid receptor binding patterns. Nat. Genet. 2011, 43, 264–268. [Google Scholar] [CrossRef] [PubMed]
- Harmston, N.; Lenhard, B. Chromatin and epigenetic features of long-range gene regulation. Nucleic Acids Res. 2013, 41, 7185–7199. [Google Scholar] [CrossRef] [PubMed]
- Cavalli, G.; Misteli, T. Functional implications of genome topology. Nat. Struct. Mol. Biol. 2013, 20, 290–299. [Google Scholar] [CrossRef] [PubMed]
- Dekker, J.; Mirny, L. The 3d genome as moderator of chromosomal communication. Cell 2016, 164, 1110–1121. [Google Scholar] [CrossRef] [PubMed]
- Gibcus, J.H.; Dekker, J. The hierarchy of the 3d genome. Mol. Cell 2013, 49, 773–782. [Google Scholar] [CrossRef] [PubMed]
- Pope, B.D.; Ryba, T.; Dileep, V.; Yue, F.; Wu, W.; Denas, O.; Vera, D.L.; Wang, Y.; Hansen, R.S.; Canfield, T.K.; et al. Topologically associating domains are stable units of replication-timing regulation. Nature 2014, 515, 402–405. [Google Scholar] [CrossRef] [PubMed]
- Dixon, J.R.; Jung, I.; Selvaraj, S.; Shen, Y.; Antosiewicz-Bourget, J.E.; Lee, A.Y.; Ye, Z.; Kim, A.; Rajagopal, N.; Xie, W.; et al. Chromatin architecture reorganization during stem cell differentiation. Nature 2015, 518, 331–336. [Google Scholar] [CrossRef] [PubMed]
- Dixon, J.R.; Selvaraj, S.; Yue, F.; Kim, A.; Li, Y.; Shen, Y.; Hu, M.; Liu, J.S.; Ren, B. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 2012, 485, 376–380. [Google Scholar] [CrossRef] [PubMed]
- Sexton, T.; Yaffe, E.; Kenigsberg, E.; Bantignies, F.; Leblanc, B.; Hoichman, M.; Parrinello, H.; Tanay, A.; Cavalli, G. Three-dimensional folding and functional organization principles of the drosophila genome. Cell 2012, 148, 458–472. [Google Scholar] [CrossRef] [PubMed]
- Kuznetsova, T.; Wang, S.Y.; Rao, N.A.; Mandoli, A.; Martens, J.H.; Rother, N.; Aartse, A.; Groh, L.; Janssen-Megens, E.M.; Li, G.; et al. Glucocorticoid receptor and nuclear factor κ-b affect three-dimensional chromatin organization. Genome Biol. 2015, 16, 264. [Google Scholar] [CrossRef] [PubMed]
- Fullwood, M.J.; Liu, M.H.; Pan, Y.F.; Liu, J.; Xu, H.; Mohamed, Y.B.; Orlov, Y.L.; Velkov, S.; Ho, A.; Mei, P.H.; et al. An oestrogen-receptor-α-bound human chromatin interactome. Nature 2009, 462, 58–64. [Google Scholar] [CrossRef] [PubMed]
- Le Dily, F.; Bau, D.; Pohl, A.; Vicent, G.P.; Serra, F.; Soronellas, D.; Castellano, G.; Wright, R.H.; Ballare, C.; Filion, G.; et al. Distinct structural transitions of chromatin topological domains correlate with coordinated hormone-induced gene regulation. Genes Dev. 2014, 28, 2151–2162. [Google Scholar] [CrossRef] [PubMed]
- Le Dily, F.; Beato, M. Tads as modular and dynamic units for gene regulation by hormones. FEBS Lett. 2015, 589, 2885–2892. [Google Scholar] [CrossRef] [PubMed]
- Dekker, J.; Rippe, K.; Dekker, M.; Kleckner, N. Capturing chromosome conformation. Science 2002, 295, 1306–1311. [Google Scholar] [CrossRef] [PubMed]
- Lieberman-Aiden, E.; van Berkum, N.L.; Williams, L.; Imakaev, M.; Ragoczy, T.; Telling, A.; Amit, I.; Lajoie, B.R.; Sabo, P.J.; Dorschner, M.O.; et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 2009, 326, 289–293. [Google Scholar] [CrossRef] [PubMed]
- Nora, E.P.; Lajoie, B.R.; Schulz, E.G.; Giorgetti, L.; Okamoto, I.; Servant, N.; Piolot, T.; van Berkum, N.L.; Meisig, J.; Sedat, J.; et al. Spatial partitioning of the regulatory landscape of the x-inactivation centre. Nature 2012, 485, 381–385. [Google Scholar] [CrossRef] [PubMed]
- Rao, S.S.; Huntley, M.H.; Durand, N.C.; Stamenova, E.K.; Bochkov, I.D.; Robinson, J.T.; Sanborn, A.L.; Machol, I.; Omer, A.D.; Lander, E.S.; et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 2014, 159, 1665–1680. [Google Scholar] [CrossRef] [PubMed]
- Cremer, T.; Cremer, C.; Schneider, T.; Baumann, H.; Hens, L.; Kirsch-Volders, M. Analysis of chromosome positions in the interphase nucleus of chinese hamster cells by laser-UV-microirradiation experiments. Hum. Genet. 1982, 62, 201–209. [Google Scholar] [CrossRef] [PubMed]
- Postberg, J.; Lipps, H.J.; Cremer, T. Evolutionary origin of the cell nucleus and its functional architecture. Essays Biochem. 2010, 48, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Heride, C.; Ricoul, M.; Kieu, K.; von Hase, J.; Guillemot, V.; Cremer, C.; Dubrana, K.; Sabatier, L. Distance between homologous chromosomes results from chromosome positioning constraints. J. Cell Sci. 2010, 123, 4063–4075. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Su, J.H.; Beliveau, B.J.; Bintu, B.; Moffitt, J.R.; Wu, C.T.; Zhuang, X. Spatial organization of chromatin domains and compartments in single chromosomes. Science 2016, 353, 598–602. [Google Scholar] [CrossRef] [PubMed]
- Rowley, M.J.; Nichols, M.H.; Lyu, X.; Ando-Kuri, M.; Rivera, I.S.M.; Hermetz, K.; Wang, P.; Ruan, Y.; Corces, V.G. Evolutionarily conserved principles predict 3d chromatin organization. Mol. Cell 2017, 67, 837–852. [Google Scholar] [CrossRef] [PubMed]
- Bonev, B.; Mendelson Cohen, N.; Szabo, Q.; Fritsch, L.; Papadopoulos, G.L.; Lubling, Y.; Xu, X.; Lv, X.; Hugnot, J.P.; Tanay, A.; et al. Multiscale 3d genome rewiring during mouse neural development. Cell 2017, 171, 557–572. [Google Scholar] [CrossRef] [PubMed]
- Stadhouders, R.; Vidal, E.; Serra, F.; Di Stefano, B.; Le Dily, F.; Quilez, J.; Gomez, A.; Collombet, S.; Berenguer, C.; Cuartero, Y.; et al. Transcription factors orchestrate dynamic interplay between genome topology and gene regulation during cell reprogramming. Nat. Genet. 2018. [Google Scholar] [CrossRef] [PubMed]
- Hou, C.; Li, L.; Qin, Z.S.; Corces, V.G. Gene density, transcription, and insulators contribute to the partition of the drosophila genome into physical domains. Mol. Cell 2012, 48, 471–484. [Google Scholar] [CrossRef] [PubMed]
- Van Bortle, K.; Corces, V.G. The role of chromatin insulators in nuclear architecture and genome function. Curr. Opin. Genet. Dev. 2013, 23, 212–218. [Google Scholar] [CrossRef] [PubMed]
- Zhan, Y.; Mariani, L.; Barozzi, I.; Schulz, E.G.; Bluthgen, N.; Stadler, M.; Tiana, G.; Giorgetti, L. Reciprocal insulation analysis of Hi-C data shows that tads represent a functionally but not structurally privileged scale in the hierarchical folding of chromosomes. Genome Res. 2017, 27, 479–490. [Google Scholar] [CrossRef] [PubMed]
- Neems, D.S.; Garza-Gongora, A.G.; Smith, E.D.; Kosak, S.T. Topologically associated domains enriched for lineage-specific genes reveal expression-dependent nuclear topologies during myogenesis. Proc. Natl. Acad. Sci. USA 2016, 113, E1691–E1700. [Google Scholar] [CrossRef] [PubMed]
- Phillips-Cremins, J.E.; Sauria, M.E.; Sanyal, A.; Gerasimova, T.I.; Lajoie, B.R.; Bell, J.S.; Ong, C.T.; Hookway, T.A.; Guo, C.; Sun, Y.; et al. Architectural protein subclasses shape 3d organization of genomes during lineage commitment. Cell 2013, 153, 1281–1295. [Google Scholar] [CrossRef] [PubMed]
- Jin, F.; Li, Y.; Dixon, J.R.; Selvaraj, S.; Ye, Z.; Lee, A.Y.; Yen, C.A.; Schmitt, A.D.; Espinoza, C.A.; Ren, B. A high-resolution map of the three-dimensional chromatin interactome in human cells. Nature 2013, 503, 290–294. [Google Scholar] [CrossRef] [PubMed]
- Dowen, J.M.; Fan, Z.P.; Hnisz, D.; Ren, G.; Abraham, B.J.; Zhang, L.N.; Weintraub, A.S.; Schujiers, J.; Lee, T.I.; Zhao, K.; et al. Control of cell identity genes occurs in insulated neighborhoods in mammalian chromosomes. Cell 2014, 159, 374–387. [Google Scholar] [CrossRef] [PubMed]
- Weintraub, A.S.; Li, C.H.; Zamudio, A.V.; Sigova, A.A.; Hannett, N.M.; Day, D.S.; Abraham, B.J.; Cohen, M.A.; Nabet, B.; Buckley, D.L.; et al. Yy1 is a structural regulator of enhancer-promoter loops. Cell 2017, 171, 1573–1588. [Google Scholar] [CrossRef] [PubMed]
- Nagano, T.; Lubling, Y.; Yaffe, E.; Wingett, S.W.; Dean, W.; Tanay, A.; Fraser, P. Single-cell Hi-C for genome-wide detection of chromatin interactions that occur simultaneously in a single cell. Nat. Protoc. 2015, 10, 1986–2003. [Google Scholar] [CrossRef] [PubMed]
- Williamson, I.; Berlivet, S.; Eskeland, R.; Boyle, S.; Illingworth, R.S.; Paquette, D.; Dostie, J.; Bickmore, W.A. Spatial genome organization: Contrasting views from chromosome conformation capture and fluorescence in situ hybridization. Genes Dev. 2014, 28, 2778–2791. [Google Scholar] [CrossRef] [PubMed]
- Schwarzer, W.; Abdennur, N.; Goloborodko, A.; Pekowska, A.; Fudenberg, G.; Loe-Mie, Y.; Fonseca, N.A.; Huber, W.; Haering, C.H.; Mirny, L.; et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 2017, 551, 51–56. [Google Scholar] [CrossRef] [PubMed]
- Nora, E.P.; Goloborodko, A.; Valton, A.L.; Gibcus, J.H.; Uebersohn, A.; Abdennur, N.; Dekker, J.; Mirny, L.A.; Bruneau, B.G. Targeted degradation of ctcf decouples local insulation of chromosome domains from genomic compartmentalization. Cell 2017, 169, 930–944. [Google Scholar] [CrossRef] [PubMed]
- Rao, S.S.P.; Huang, S.C.; Glenn St Hilaire, B.; Engreitz, J.M.; Perez, E.M.; Kieffer-Kwon, K.R.; Sanborn, A.L.; Johnstone, S.E.; Bascom, G.D.; Bochkov, I.D.; et al. Cohesin loss eliminates all loop domains. Cell 2017, 171, 305–320. [Google Scholar] [CrossRef] [PubMed]
- Ross-Innes, C.S.; Stark, R.; Teschendorff, A.E.; Holmes, K.A.; Ali, H.R.; Dunning, M.J.; Brown, G.D.; Gojis, O.; Ellis, I.O.; Green, A.R.; et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 2012, 481, 389–393. [Google Scholar] [CrossRef] [PubMed]
- Creighton, C.J.; Cordero, K.E.; Larios, J.M.; Miller, R.S.; Johnson, M.D.; Chinnaiyan, A.M.; Lippman, M.E.; Rae, J.M. Genes regulated by estrogen in breast tumor cells in vitro are similarly regulated in vivo in tumor xenografts and human breast tumors. Genome Biol. 2006, 7, R28. [Google Scholar] [CrossRef] [PubMed]
- Rangel, N.; Villegas, V.E.; Rondon-Lagos, M. Profiling of gene expression regulated by 17β-estradiol and tamoxifen in estrogen receptor-positive and estrogen receptor-negative human breast cancer cell lines. Breast Cancer 2017, 9, 537–550. [Google Scholar] [PubMed]
- Carroll, J.S.; Meyer, C.A.; Song, J.; Li, W.; Geistlinger, T.R.; Eeckhoute, J.; Brodsky, A.S.; Keeton, E.K.; Fertuck, K.C.; Hall, G.F.; et al. Genome-wide analysis of estrogen receptor binding sites. Nat. Genet. 2006, 38, 1289–1297. [Google Scholar] [CrossRef] [PubMed]
- Joseph, R.; Orlov, Y.L.; Huss, M.; Sun, W.; Kong, S.L.; Ukil, L.; Pan, Y.F.; Li, G.; Lim, M.; Thomsen, J.S.; et al. Integrative model of genomic factors for determining binding site selection by estrogen receptor-α. Mol. Syst. Biol. 2010, 6, 456. [Google Scholar] [CrossRef] [PubMed]
- Zaret, K.S.; Carroll, J.S. Pioneer transcription factors: Establishing competence for gene expression. Genes Dev. 2011, 25, 2227–2241. [Google Scholar] [CrossRef] [PubMed]
- Hurtado, A.; Holmes, K.A.; Ross-Innes, C.S.; Schmidt, D.; Carroll, J.S. Foxa1 is a key determinant of estrogen receptor function and endocrine response. Nat. Genet. 2011, 43, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Verde, G.; De Llobet, L.I.; Wright, R.H.G.; Quilez, J.; Peiro, S.; Le Dily, F.; Beato, M. Progesterone receptor maintains estrogen receptor gene expression by regulating DNA methylation in hormone-free breast cancer cells. bioRxiv 2017. [Google Scholar] [CrossRef]
- Slattery, M.; Zhou, T.; Yang, L.; Dantas Machado, A.C.; Gordan, R.; Rohs, R. Absence of a simple code: How transcription factors read the genome. Trends Biochem. Sci. 2014, 39, 381–399. [Google Scholar] [CrossRef] [PubMed]
- Berg, O.G.; Winter, R.B.; von Hippel, P.H. Diffusion-driven mechanisms of protein translocation on nucleic acids. 1. Models and theory. Biochemistry 1981, 20, 6929–6948. [Google Scholar] [CrossRef] [PubMed]
- Cortini, R.; Filion, G. Principles of transcription factor traffic on folded chromatin. bioRxiv 2017. [Google Scholar] [CrossRef]
- Li, G.; Cai, L.; Chang, H.; Hong, P.; Zhou, Q.; Kulakova, E.V.; Kolchanov, N.A.; Ruan, Y. Chromatin interaction analysis with paired-end tag (chia-pet) sequencing technology and application. BMC Genom. 2014, 15 (Suppl. 12), S11. [Google Scholar] [CrossRef] [PubMed]
- Sanyal, A.; Lajoie, B.R.; Jain, G.; Dekker, J. The long-range interaction landscape of gene promoters. Nature 2012, 489, 109–113. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Yue, F.; McCleary, D.F.; Ye, Z.; Edsall, L.; Kuan, S.; Wagner, U.; Dixon, J.; Lee, L.; Lobanenkov, V.V.; et al. A map of the cis-regulatory sequences in the mouse genome. Nature 2012, 488, 116–120. [Google Scholar] [CrossRef] [PubMed]
- Lupianez, D.G.; Kraft, K.; Heinrich, V.; Krawitz, P.; Brancati, F.; Klopocki, E.; Horn, D.; Kayserili, H.; Opitz, J.M.; Laxova, R.; et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 2015, 161, 1012–1025. [Google Scholar] [CrossRef] [PubMed]
- Hon, G.C.; Hawkins, R.D.; Caballero, O.L.; Lo, C.; Lister, R.; Pelizzola, M.; Valsesia, A.; Ye, Z.; Kuan, S.; Edsall, L.E.; et al. Global DNA hypomethylation coupled to repressive chromatin domain formation and gene silencing in breast cancer. Genome Res. 2012, 22, 246–258. [Google Scholar] [CrossRef] [PubMed]
- Bert, S.A.; Robinson, M.D.; Strbenac, D.; Statham, A.L.; Song, J.Z.; Hulf, T.; Sutherland, R.L.; Coolen, M.W.; Stirzaker, C.; Clark, S.J. Regional activation of the cancer genome by long-range epigenetic remodeling. Cancer Cell 2013, 23, 9–22. [Google Scholar] [CrossRef] [PubMed]
- McDonald, O.G.; Wu, H.; Timp, W.; Doi, A.; Feinberg, A.P. Genome-scale epigenetic reprogramming during epithelial-to-mesenchymal transition. Nat. Struct. Mol. Biol. 2011, 18, 867–874. [Google Scholar] [CrossRef] [PubMed]
- Antony, J.; Dasgupta, T.; Rhodes, J.M.; McEwan, M.V.; Print, C.G.; O’Sullivan, J.M.; Horsfield, J.A. Cohesin modulates transcription of estrogen-responsive genes. Biochim. Biophys. Acta 2015, 1849, 257–269. [Google Scholar] [CrossRef] [PubMed]
- Ross-Innes, C.S.; Brown, G.D.; Carroll, J.S. A co-ordinated interaction between ctcf and er in breast cancer cells. BMC Genom. 2011, 12, 593. [Google Scholar] [CrossRef] [PubMed]
- Quintin, J.; Le Peron, C.; Palierne, G.; Bizot, M.; Cunha, S.; Serandour, A.A.; Avner, S.; Henry, C.; Percevault, F.; Belaud-Rotureau, M.A.; et al. Dynamic estrogen receptor interactomes control estrogen-responsive trefoil factor (TFF) locus cell-specific activities. Mol. Cell. Biol. 2014, 34, 2418–2436. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, D.; Schwalie, P.C.; Ross-Innes, C.S.; Hurtado, A.; Brown, G.D.; Carroll, J.S.; Flicek, P.; Odom, D.T. A ctcf-independent role for cohesin in tissue-specific transcription. Genome Res. 2010, 20, 578–588. [Google Scholar] [CrossRef] [PubMed]
- Chan, C.S.; Song, J.S. Ccctc-binding factor confines the distal action of estrogen receptor. Cancer Res. 2008, 68, 9041–9049. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.I.; Lessard, J.; Crabtree, G.R. Understanding the words of chromatin regulation. Cell 2009, 136, 200–206. [Google Scholar] [CrossRef] [PubMed]
- Johnson, T.A.; Elbi, C.; Parekh, B.S.; Hager, G.L.; John, S. Chromatin remodeling complexes interact dynamically with a glucocorticoid receptor-regulated promoter. Mol. Biol. Cell 2008, 19, 3308–3322. [Google Scholar] [CrossRef] [PubMed]
- Vicent, G.P.; Nacht, A.S.; Font-Mateu, J.; Castellano, G.; Gaveglia, L.; Ballare, C.; Beato, M. Four enzymes cooperate to displace histone h1 during the first minute of hormonal gene activation. Genes Dev. 2011, 25, 845–862. [Google Scholar] [CrossRef] [PubMed]
- Bretschneider, N.; Kangaspeska, S.; Seifert, M.; Reid, G.; Gannon, F.; Denger, S. E2-mediated cathepsin d (ctsd) activation involves looping of distal enhancer elements. Mol. Oncol. 2008, 2, 182–190. [Google Scholar] [CrossRef] [PubMed]
- Hsu, P.Y.; Hsu, H.K.; Singer, G.A.; Yan, P.S.; Rodriguez, B.A.; Liu, J.C.; Weng, Y.I.; Deatherage, D.E.; Chen, Z.; Pereira, J.S.; et al. Estrogen-mediated epigenetic repression of large chromosomal regions through DNA looping. Genome Res. 2010, 20, 733–744. [Google Scholar] [CrossRef] [PubMed]
- Belakavadi, M.; Fondell, J.D. Role of the mediator complex in nuclear hormone receptor signaling. Rev. Physiol. Biochem. Pharmacol. 2006, 156, 23–43. [Google Scholar] [PubMed]
- Malik, S.; Roeder, R.G. The metazoan mediator co-activator complex as an integrative hub for transcriptional regulation. Nat. Rev. Genet. 2010, 11, 761–772. [Google Scholar] [CrossRef] [PubMed]
- Bojcsuk, D.; Nagy, G.; Balint, B.L. Inducible super-enhancers are organized based on canonical signal-specific transcription factor binding elements. Nucleic Acids Res. 2017, 45, 3693–3706. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Notani, D.; Ma, Q.; Tanasa, B.; Nunez, E.; Chen, A.Y.; Merkurjev, D.; Zhang, J.; Ohgi, K.; Song, X.; et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature 2013, 498, 516–520. [Google Scholar] [CrossRef] [PubMed]
- Hah, N.; Murakami, S.; Nagari, A.; Danko, C.G.; Kraus, W.L. Enhancer transcripts mark active estrogen receptor binding sites. Genome Res. 2013, 23, 1210–1223. [Google Scholar] [CrossRef] [PubMed]
- Lai, F.; Orom, U.A.; Cesaroni, M.; Beringer, M.; Taatjes, D.J.; Blobel, G.A.; Shiekhattar, R. Activating RNAs associate with mediator to enhance chromatin architecture and transcription. Nature 2013, 494, 497–501. [Google Scholar] [CrossRef] [PubMed]
- Le Dily, F.; Vidal, E.; Cuartero, Y.; Quilez, J.; Nacht, S.; Vicent, G.P.; Sharma, P.; Verde, G.; Beato, M. Hormone control regions mediate opposing steroid receptor-dependent genome organizations. bioRxiv 2017. [Google Scholar] [CrossRef]
- Hakim, O.; Sung, M.H.; Voss, T.C.; Splinter, E.; John, S.; Sabo, P.J.; Thurman, R.E.; Stamatoyannopoulos, J.A.; de Laat, W.; Hager, G.L. Diverse gene reprogramming events occur in the same spatial clusters of distal regulatory elements. Genome Res. 2011, 21, 697–706. [Google Scholar] [CrossRef] [PubMed]
- Rafique, S.; Thomas, J.S.; Sproul, D.; Bickmore, W.A. Estrogen-induced chromatin decondensation and nuclear re-organization linked to regional epigenetic regulation in breast cancer. Genome Biol. 2015, 16, 145. [Google Scholar] [CrossRef] [PubMed]
- Jubb, A.W.; Boyle, S.; Hume, D.A.; Bickmore, W.A. Glucocorticoid receptor binding induces rapid and prolonged large-scale chromatin decompaction at multiple target loci. Cell Rep. 2017, 21, 3022–3031. [Google Scholar] [CrossRef] [PubMed]
- Nye, A.C.; Rajendran, R.R.; Stenoien, D.L.; Mancini, M.A.; Katzenellenbogen, B.S.; Belmont, A.S. Alteration of large-scale chromatin structure by estrogen receptor. Mol. Cell. Biol. 2002, 22, 3437–3449. [Google Scholar] [CrossRef] [PubMed]
- Mourad, R.; Hsu, P.Y.; Juan, L.; Shen, C.; Koneru, P.; Lin, H.; Liu, Y.; Nephew, K.; Huang, T.H.; Li, L. Estrogen induces global reorganization of chromatin structure in human breast cancer cells. PLoS ONE 2014, 9, e113354. [Google Scholar] [CrossRef] [PubMed]
- Hu, Q.; Kwon, Y.S.; Nunez, E.; Cardamone, M.D.; Hutt, K.R.; Ohgi, K.A.; Garcia-Bassets, I.; Rose, D.W.; Glass, C.K.; Rosenfeld, M.G.; et al. Enhancing nuclear receptor-induced transcription requires nuclear motor and lsd1-dependent gene networking in interchromatin granules. Proc. Natl. Acad. Sci. USA 2008, 105, 19199–19204. [Google Scholar] [CrossRef] [PubMed]
- Kocanova, S.; Kerr, E.A.; Rafique, S.; Boyle, S.; Katz, E.; Caze-Subra, S.; Bickmore, W.A.; Bystricky, K. Activation of estrogen-responsive genes does not require their nuclear co-localization. PLoS Genet. 2010, 6, e1000922. [Google Scholar] [CrossRef] [PubMed]
- Levin, E.R. Integration of the extranuclear and nuclear actions of estrogen. Mol. Endocrinol. 2005, 19, 1951–1959. [Google Scholar] [CrossRef] [PubMed]
- Vicent, G.P.; Ballare, C.; Nacht, A.S.; Clausell, J.; Subtil-Rodriguez, A.; Quiles, I.; Jordan, A.; Beato, M. Induction of progesterone target genes requires activation of erk and msk kinases and phosphorylation of histone h3. Mol. Cell 2006, 24, 367–381. [Google Scholar] [CrossRef] [PubMed]
- Migliaccio, A.; Piccolo, D.; Castoria, G.; Di Domenico, M.; Bilancio, A.; Lombardi, M.; Gong, W.; Beato, M.; Auricchio, F. Activation of the src/p21ras/erk pathway by progesterone receptor via cross-talk with estrogen receptor. EMBO J. 1998, 17, 2008–2018. [Google Scholar] [CrossRef] [PubMed]
- Revankar, C.M.; Cimino, D.F.; Sklar, L.A.; Arterburn, J.B.; Prossnitz, E.R. A transmembrane intracellular estrogen receptor mediates rapid cell signaling. Science 2005, 307, 1625–1630. [Google Scholar] [CrossRef] [PubMed]
- Wright, R.H.; Castellano, G.; Bonet, J.; Le Dily, F.; Font-Mateu, J.; Ballare, C.; Nacht, A.S.; Soronellas, D.; Oliva, B.; Beato, M. CDK2-dependent activation of PARP-1 is required for hormonal gene regulation in breast cancer cells. Genes Dev. 2012, 26, 1972–1983. [Google Scholar] [CrossRef] [PubMed]
- Wright, R.H.; Lioutas, A.; Le Dily, F.; Soronellas, D.; Pohl, A.; Bonet, J.; Nacht, A.S.; Samino, S.; Font-Mateu, J.; Vicent, G.P.; et al. ADP-ribose-derived nuclear ATP synthesis by NUDIX5 is required for chromatin remodeling. Science 2016, 352, 1221–1225. [Google Scholar] [CrossRef] [PubMed]
- Altmeyer, M.; Neelsen, K.J.; Teloni, F.; Pozdnyakova, I.; Pellegrino, S.; Grofte, M.; Rask, M.B.; Streicher, W.; Jungmichel, S.; Nielsen, M.L.; et al. Liquid demixing of intrinsically disordered proteins is seeded by poly(adp-ribose). Nat. Commun. 2015, 6, 8088. [Google Scholar] [CrossRef] [PubMed]
- Patel, A.; Malinovska, L.; Saha, S.; Wang, J.; Alberti, S.; Krishnan, Y.; Hyman, A.A. ATP as a biological hydrotrope. Science 2017, 356, 753–756. [Google Scholar] [CrossRef] [PubMed]
- Franke, M.; Ibrahim, D.M.; Andrey, G.; Schwarzer, W.; Heinrich, V.; Schopflin, R.; Kraft, K.; Kempfer, R.; Jerkovic, I.; Chan, W.L.; et al. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature 2016, 538, 265–269. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Gong, M.; Yuan, H.; Park, H.G.; Frierson, H.F.; Li, H. Chimeric transcript generated by cis-splicing of adjacent genes regulates prostate cancer cell proliferation. Cancer Discov. 2012, 2, 598–607. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.; Yang, L.; Tanasa, B.; Hutt, K.; Ju, B.G.; Ohgi, K.; Zhang, J.; Rose, D.W.; Fu, X.D.; Glass, C.K.; et al. Nuclear receptor-induced chromosomal proximity and DNA breaks underlie specific translocations in cancer. Cell 2009, 139, 1069–1083. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Le Dily, F.; Beato, M. Signaling by Steroid Hormones in the 3D Nuclear Space. Int. J. Mol. Sci. 2018, 19, 306. https://doi.org/10.3390/ijms19020306
Le Dily F, Beato M. Signaling by Steroid Hormones in the 3D Nuclear Space. International Journal of Molecular Sciences. 2018; 19(2):306. https://doi.org/10.3390/ijms19020306
Chicago/Turabian StyleLe Dily, François, and Miguel Beato. 2018. "Signaling by Steroid Hormones in the 3D Nuclear Space" International Journal of Molecular Sciences 19, no. 2: 306. https://doi.org/10.3390/ijms19020306
APA StyleLe Dily, F., & Beato, M. (2018). Signaling by Steroid Hormones in the 3D Nuclear Space. International Journal of Molecular Sciences, 19(2), 306. https://doi.org/10.3390/ijms19020306




