Mapping Quantitative Trait Loci (QTL) for Resistance to Late Blight in Tomato
Abstract
1. Introduction
2. Results
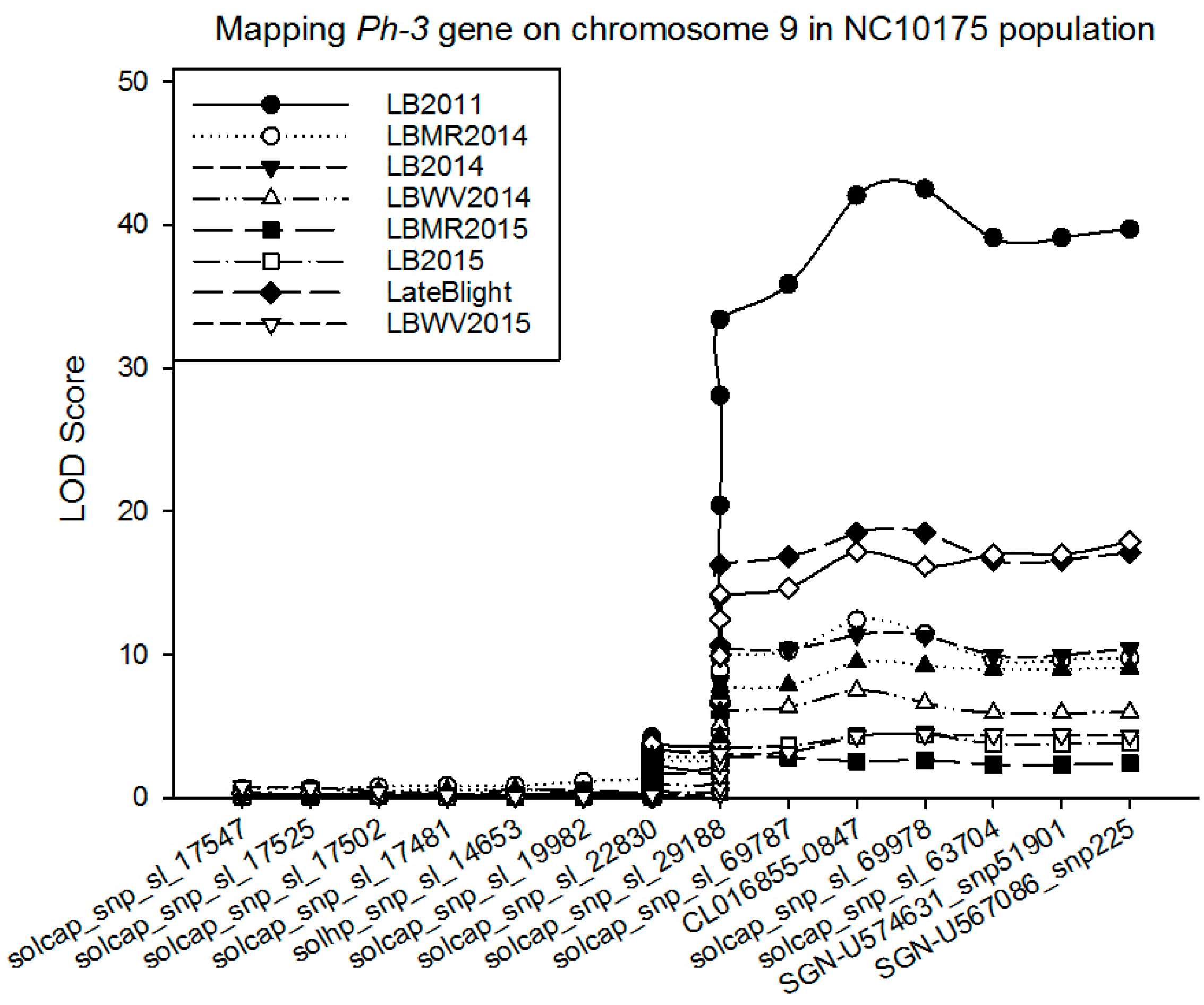
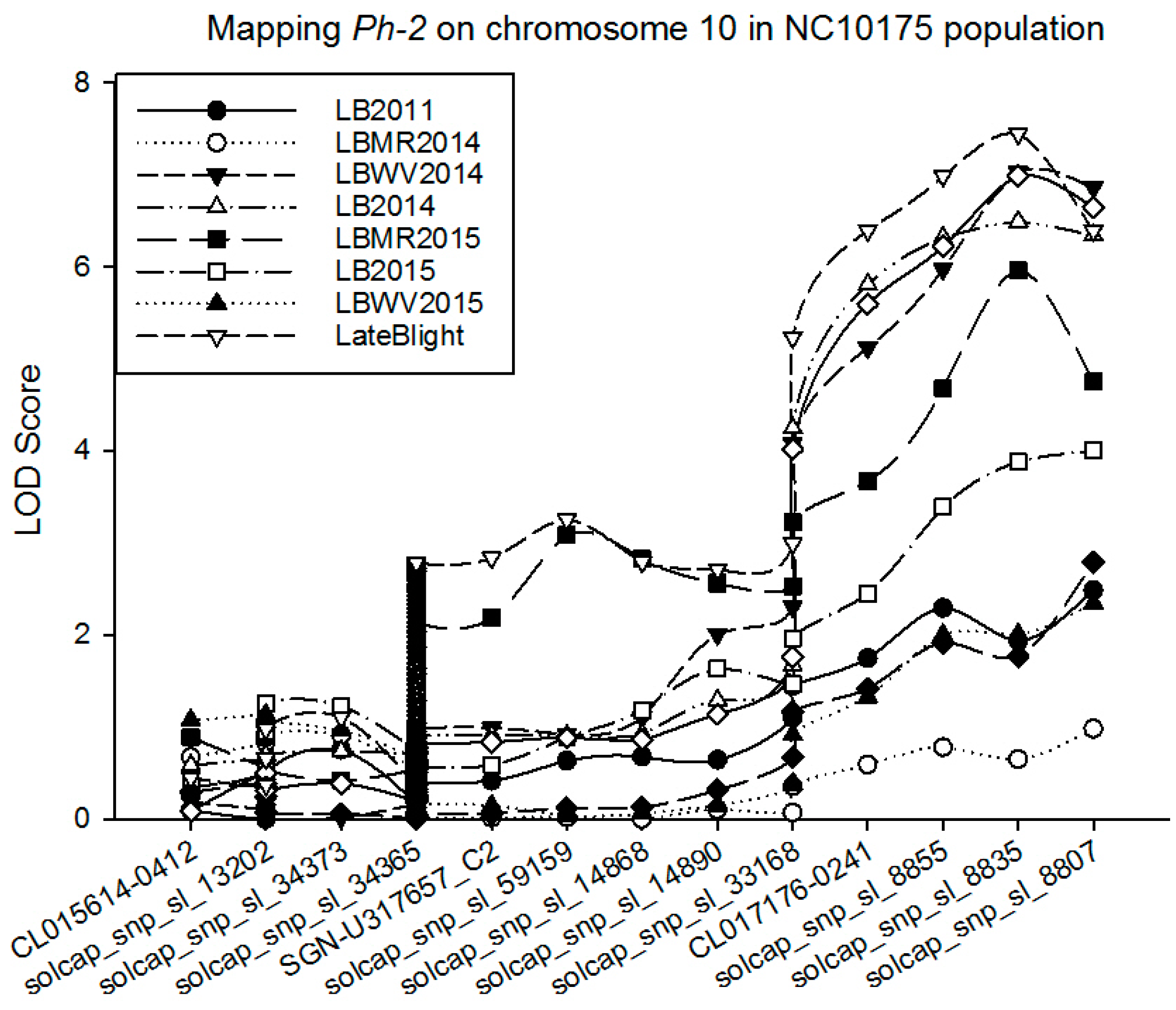
Quantitative Trait Loci (QTL) Analysis
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Phenotyping for Disease Resistance in Field in 2011
4.3. Phenotyping for Disease Resistance in the Field in 2014 and 2015
4.4. DNA Isolation and SNP Genotyping
4.5. Phenotypic Data Analysis
4.6. QTL Analysis
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Peirce, L.C. Linkage tests with Ph. conditioning resistance to race 0, Phytophthora infestans. Rep. Tomato Genet. Coop. 1971, 21, 30. [Google Scholar]
- AVRDC. AVRDC Report 1998; Asian Vegetable Research and Development Center: Tainan, Taiwan, 1999; pp. 9–13. [Google Scholar]
- Merk, H.L.; Ashrafi, H.; Foolad, M.R. Selective genotyping to identify late blight resistance genes in an accession of the tomato wild species Solanum pimpinellifolium. Euphytica 2012, 187, 63–75. [Google Scholar] [CrossRef]
- Moreau, P.; Thoquet, P.; Olivier, J.; Laterrot, H.; Grimsley, N. Genetic mapping of Ph-2, a single locus controlling partial resistance to Phytophtora infestans in tomato. Mol. Plant-Microbe Interact. 1998, 11, 259–269. [Google Scholar] [CrossRef]
- Gallegly, M.E. Resistance to the Late Blight Fungus in Tomato; Campbell Soup Co.: Camden, NJ, USA, 1960; pp. 113–135. [Google Scholar]
- Black, L.L.; Wang, T.C.; Hanson, P.M.; Chen, J.T. Late blight resistance in four wild tomato accessions: Effectiveness in diverse locations and inheritance of resistance. Phytopathology 1996, 86, S24. [Google Scholar]
- Goodwin, S.B.; Schneider, R.E.; Fry, W.E. Use of cellulose-acetate electrophoresis for rapid identification of allozyme genotypes of Phytophthora infestans. Plant Dis. 1995, 79, 1181–1185. [Google Scholar] [CrossRef]
- Chunwongse, J.; Chunwongse, C.; Black, L.; Hanson, P. Molecular mapping of the Ph-3 gene for late blight resistance in tomato. J. Hortic. Sci. Biotechnol. 2002, 77, 281–286. [Google Scholar] [CrossRef]
- Zhang, C.Z.; Liu, L.; Zheng, Z.; Sun, Y.Y.; Zhou, L.X.; Yang, Y.H.; Cheng, F.; Zhang, Z.H.; Wang, X.W.; Huang, S.W.; et al. Fine mapping of the Ph-3 gene conferring resistance to late blight (Phytophthora infestans) in tomato. Theor. Appl. Genet. 2013, 126, 2643–2653. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.Z.; Liu, L.; Wang, X.X.; Vossen, J.; Li, G.C.; Li, T.; Zheng, Z.; Gao, J.C.; Guo, Y.M.; Visser, R.G.F.; et al. The Ph-3 gene from Solanum pimpinellifolium encodes CC-NBS-LRR protein conferring resistance to Phytophthora infestans. Theor. Appl. Genet. 2014, 127, 1353–1364. [Google Scholar] [CrossRef] [PubMed]
- Park, Y.; Hwang, J.; Kim, K.; Kang, J.; Kim, B.; Xu, S.; Ahn, Y. Development of the gene-based SCARs for the Ph-3 locus, which confers late blight resistance in tomato. Sci. Hortic. 2013, 164, 9–16. [Google Scholar] [CrossRef]
- Robbins, M.D.; Masud, M.A.T.; Panthee, D.R.; Gardner, C.O.; Francis, D.; Stevens, M.A. Marker-assisted selection for coupling phase resistance to tomato spotted wilt virus and Phytophthora infestans (late blight) in tomato. HortScience 2010, 45, 1424–1428. [Google Scholar]
- Kim, M.J.; Mutschler, M.A. Transfer to processing tomato and characterization of late blight resistance derived from Solanum pimpinellifolium L. L3708. J. Am. Soc. Hortic. Sci. 2005, 130, 877–884. [Google Scholar]
- Lee, S.J.; Kelley, B.S.; Damasceno, C.M.B.; John, B.S.; Kim, B.S.; Kim, B.D.; Rose, J.K.C. A functional screen to characterize the secretomes of eukaryotic pathogens and their hosts in planta. Mol. Plant-Microbe Interact. 2006, 1, 1368–1377. [Google Scholar] [CrossRef] [PubMed]
- Gardner, R.G.; Panthee, D.R. “Plum Regal” Fresh-market plum tomato hybrid and its parents, NC 25P and NC 30P. HortScience 2010, 45, 824–825. [Google Scholar]
- Panthee, D.R.; Gardner, R.G. “Mountain Merit”: A late blight-resistant large-fruited tomato hybrid. HortScience 2010, 45, 1547–1548. [Google Scholar]
- Panthee, D.R.; Gardner, R.G. “Mountain Rouge”: A Pink-fruited, Heirloom-type hybrid tomato and its parent line NC 161L. Hortscience 2014, 49, 1463–1464. [Google Scholar]
- Gardner, R.G.; Panthee, D.R. NC 1 CELBR and NC 2 CELBR: Early blight and late blight resistant fresh market tomato breeding lines. HortScience 2010, 45, 975–976. [Google Scholar]
- Broman, K.W.; Wu, H.; Sen, S.; Churchill, G.A. R/qtl: QTL mapping in experimental crosses. Bioinformatics 2003, 19, 889–890. [Google Scholar] [CrossRef] [PubMed]
- Chunwongse, J.; Chunwongse, C.; Black, L.; Hanson, P. Mapping of the Ph-3 gene for late blight from L. pimpinellifolium L 3708. TGC Rep. 1998, 48, 13–16. [Google Scholar]
- Kim, M.J.; Mutschler, M.A. Characterization of late blight resistance derived from Solanum pimpinellifolium L3708 against multiple isolates of the pathogen Phytophthora infestans. J. Am. Soc. Hortic. Sci. 2006, 131, 637–645. [Google Scholar]
- Chen, C.H.; Shen, Z.M.; Wang, T.C. Host specificity and tomato-related race composition of Phytophthora infestans isolates in Taiwan during 2004 and 2005. Plant Dis. 2008, 92, 751–755. [Google Scholar] [CrossRef]
- Chen, A.L.; Liu, C.Y.; Chen, C.H.; Wang, J.F.; Liao, Y.C.; Chang, C.H.; Tsai, M.H.; Hwu, K.K.; Chen, K.Y. Reassessment of QTLs for late blight resistance in the tomato accession L3708 using a restriction site associated DNA (RAD) linkage map and highly aggressive isolates of phytophthora infestans. PLoS ONE 2014, 9, e96417. [Google Scholar] [CrossRef] [PubMed]
- Brouwer, D.J.; St Clair, D.A. Fine mapping of three quantitative trait loci for late blight resistance in tomato using near isogenic lines (NILs) and sub-NILs. Theor. Appl. Genet. 2004, 108, 628–638. [Google Scholar] [CrossRef] [PubMed]
- Brouwer, D.J.; Jones, E.S.; St Clair, D.A. QTL analysis of quantitative resistance to Phytophthora infestans (late blight) in tomato and comparisons with potato. Genome 2004, 47, 475–492. [Google Scholar] [CrossRef] [PubMed]
- Haggard, J.E.; Johnson, E.B.; St Clair, D.A. Linkage relationships among multiple QTL for horticultural traits and late blight (P. infestans) resistance on chromosome 5 introgressed from wild tomato solanum habrochaites. G3-Genes Genomes Genet. 2013, 3, 2131–2146. [Google Scholar] [CrossRef] [PubMed]
- Haggard, J.E.; Johnson, E.B.; St Clair, D.A. Multiple QTL for horticultural traits and quantitative resistance to phytophthora infestans linked on solanum habrochaites chromosome 11. G3-Genes Genomes Genet. 2015, 5, 219–233. [Google Scholar] [CrossRef] [PubMed]
- Li, J.M.; Liu, L.; Bai, Y.L.; Finkers, R.; Wang, F.; Du, Y.C.; Yang, Y.H.; Xie, B.Y.; Visser, R.G.F.; van Heusden, A.W. Identification and mapping of quantitative resistance to late blight (Phytophthora infestans) in Solanum habrochaites LA1777. Euphytica 2011, 179, 427–438. [Google Scholar] [CrossRef]
- Smart, C.D.; Tanksley, S.D.; Mayton, H.; Fry, W.E. Resistance to phytophthora infestans in lycopersicon pennellii. Plant Dis. 2007, 91, 1045–1049. [Google Scholar] [CrossRef]
- Shandil, R.K.; Chakrabarti, S.K.; Singh, B.P.; Sharma, S.; Sundaresha, S.; Kaushik, S.K.; Bhatt, A.K.; Sharma, N.N. Genotypic background of the recipient plant is crucial for conferring RB gene mediated late blight resistance in potato. BMC Genet. 2017, 18, 22. [Google Scholar] [CrossRef] [PubMed]
- Jo, K.R.; Arens, M.; Kim, T.Y.; Jongsma, M.A.; Visser, R.G.F.; Jacobsen, E.; Vossen, J.H. Mapping of the S. demissum late blight resistance gene R8 to a new locus on chromosome IX. Theor. Appl. Genet. 2011, 123, 1331–1340. [Google Scholar] [CrossRef] [PubMed]
- Rietman, H.; Bijsterbosch, G.; Cano, L.M.; Lee, H.R.; Vossen, J.H.; Jacobsen, E.; Visser, R.G.F.; Kamoun, S.; Vleeshouwers, V. Qualitative and quantitative late blight resistance in the potato cultivar sarpo mira is determined by the perception of five distinct RXLR effectors. Mol. Plant-Microbe Interact. 2012, 25, 910–919. [Google Scholar] [CrossRef] [PubMed]
- Barbary, A.; Palloix, A.; Fazari, A.; Marteu, N.; Castagnone-Sereno, P.; Djian-Caporalino, C. The plant genetic background affects the efficiency of the pepper major nematode resistance genes Me1 and Me3. Theor. Appl. Genet. 2014, 127, 499–507. [Google Scholar] [CrossRef] [PubMed]
- Barbary, A.; Djian-Caporalino, C.; Marteu, N.; Fazari, A.; Caromel, B.; Castagnone-Sereno, P.; Palloix, A. Plant genetic background increasing the efficiency and durability of major resistance genes to root-knot nematodes can be resolved into a few resistance QTLs. Front. Plant Sci. 2016, 7, 632. [Google Scholar] [CrossRef] [PubMed]
- Irzhansky, I.; Cohen, Y. Inheritance of resistance against Phytophthora infestans in Lycopersicon pimpenellifolium L3707. Euphytica 2006, 149, 309–316. [Google Scholar] [CrossRef]
- Sim, S.C.; van Deynze, A.; Stoffel, K.; Douches, D.S.; Zarka, D.; Ganal, M.W.; Chetelat, R.T.; Hutton, S.F.; Scott, J.W.; Gardner, R.G.; et al. High-density SNP genotyping of tomato (Solanum lycopersicum L.) reveals patterns of genetic variation due to breeding. PLoS ONE 2012, 7, e45520. [Google Scholar] [CrossRef] [PubMed]
- Sim, S.-C.; Durstewitz, G.; Plieske, J.; Wieseke, R.; Ganal, M.W.; van Deynze, A.; Hamilton, J.P.; Buell, C.R.; Causse, M.; Wijeratne, S. Development of a large SNP genotyping array and generation of high-density genetic maps in tomato. PLoS ONE 2012, 7, e40563. [Google Scholar] [CrossRef] [PubMed]
- SAS Institute Inc. The SAS System, Version 9.3 for Windows, 9th ed.; SAS Institute: Cary, NC, USA, 2011. [Google Scholar]
- Meng, L.; Li, H.H.; Zhang, L.Y.; Wang, J.K. QTL IciMapping: Integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop. J. 2015, 3, 269–283. [Google Scholar] [CrossRef]
- Kosambi, D.D. The estimation of map distances from recombination values. Ann. Eugen. 1943, 12, 172–175. [Google Scholar] [CrossRef]
- Wang, S.C.; Basten, J.; Gaffney, P.; Zeng, Z.B. Windows QTL Cartographer V2.5; Department of Statistics, North Carolina State University: Raleigh, NC, USA, 2010. [Google Scholar]

| Variables | LB2011 | LBMR2014 | LBWV2014 | LB2014 | LBMR2015 | LBWV2015 | LB2015 | Average Late Blight |
|---|---|---|---|---|---|---|---|---|
| Mean | 2.5 | 3.0 | 1.9 | 2.5 | 1.8 | 0.4 | 1.1 | 1.9 |
| Minimum | 0.0 | 1.0 | 0.0 | 0.8 | 0.0 | 0.0 | 0.0 | 0.3 |
| Maximum | 5.0 | 5.0 | 5.0 | 5.0 | 5.0 | 2.0 | 3.0 | 4.0 |
| Standard Deviation | 1.5 | 1.0 | 2.0 | 1.2 | 1.3 | 0.7 | 0.8 | 0.9 |
| Variable | LBMR2014 | LBWV2014 | LB2014 | LBMR2015 | LBWV2015 | LB2015 | Average Late Blight |
|---|---|---|---|---|---|---|---|
| LB2011 | 0.34 | 0.50 | 0.57 | 0.29 | 0.34 | 0.35 | 0.79 |
| LBMR2014 | 0.10 | 0.54 | 0.21 | 0.28 | 0.27 | 0.46 | |
| LBWV2014 | 0.91 | 0.28 | 0.31 | 0.34 | 0.75 | ||
| LB2014 | 0.35 | 0.39 | 0.42 | 0.83 | |||
| LBMR2015 | 0.30 | 0.94 | 0.58 | ||||
| LBWV2015 | 0.64 | 0.52 | |||||
| LB2015 | 0.65 |
| Environment | Chromosome | Markers | Position (cM) | LOD Score | Additive Effect | Dominance Effect | R2-Value |
|---|---|---|---|---|---|---|---|
| LB2011 | 9 | solcap_snp_sl_22830 | 0.32 | 9.18 | −0.33 | −2.93 | 0.81 |
| LBMR2014 | 9 | solcap_snp_sl_22830 | 0.27 | 2.75 | −1.40 | 1.48 | 0.69 |
| 9 | solcap_snp_sl_29188 | 0.66 | 33.36 | −1.66 | 0.52 | 0.60 | |
| 9 | solcap_snp_sl_69787 | 0.66 | 35.81 | −1.68 | 0.58 | 0.64 | |
| 9 | CL016855-0847 | 0.67 | 41.99 | −1.69 | 0.58 | 0.66 | |
| 9 | solcap_snp_sl_69978 | 0.67 | 42.44 | −1.72 | 0.50 | 0.67 | |
| 9 | solcap_snp_sl_63704 | 0.67 | 39.05 | −1.68 | 0.52 | 0.63 | |
| 9 | SGN-U574631_snp51901 | 0.67 | 39.05 | −1.68 | 0.52 | 0.63 | |
| 9 | SGN-U567086_snp225 | 0.67 | 39.65 | −1.71 | 0.53 | 0.65 | |
| LBWV2014 | 9 | solcap_snp_sl_22830 | 0.40 | 3.25 | −0.59 | −0.38 | 0.19 |
| 9 | solcap_snp_sl_29188 | 0.66 | 10.01 | −0.72 | −0.30 | 0.23 | |
| 9 | solcap_snp_sl_69787 | 0.66 | 10.27 | −0.71 | −0.30 | 0.22 | |
| 9 | CL016855-0847 | 0.67 | 12.40 | −0.73 | −0.42 | 0.26 | |
| 9 | solcap_snp_sl_69978 | 0.67 | 11.40 | −0.71 | −0.36 | 0.24 | |
| 9 | solcap_snp_sl_63704 | 0.67 | 9.62 | −0.67 | −0.28 | 0.21 | |
| 9 | SGN-U574631_snp51901 | 0.67 | 9.62 | −0.67 | −0.28 | 0.21 | |
| 9 | SGN-U567086_snp225 | 0.67 | 9.74 | −0.68 | −0.28 | 0.21 | |
| LB2014 | 9 | solcap_snp_sl_29188 | 0.66 | 6.09 | −1.00 | 0.32 | 0.14 |
| 9 | solcap_snp_sl_69787 | 0.66 | 6.31 | −0.98 | 0.35 | 0.13 | |
| 9 | CL016855-0847 | 0.67 | 7.49 | −1.06 | 0.33 | 0.16 | |
| 9 | solcap_snp_sl_69978 | 0.67 | 6.59 | −1.02 | 0.22 | 0.14 | |
| 9 | solcap_snp_sl_63704 | 0.67 | 5.92 | −0.94 | 0.36 | 0.13 | |
| 9 | SGN-U574631_snp51901 | 0.67 | 5.92 | −0.94 | 0.36 | 0.13 | |
| 9 | SGN-U567086_snp225 | 0.67 | 5.96 | −0.95 | 0.36 | 0.13 | |
| LBMR2015 | 9 | solcap_snp_sl_22830 | 0.61 | 3.30 | −0.42 | −0.24 | 0.07 |
| 9 | solcap_snp_sl_29188 | 0.66 | 10.44 | −0.81 | −0.30 | 0.21 | |
| 9 | solcap_snp_sl_69787 | 0.66 | 10.37 | −0.78 | −0.25 | 0.20 | |
| 9 | CL016855-0847 | 0.67 | 11.36 | −0.80 | −0.24 | 0.22 | |
| 9 | solcap_snp_sl_69978 | 0.67 | 11.27 | −0.78 | −0.30 | 0.21 | |
| 9 | solcap_snp_sl_63704 | 0.67 | 9.95 | −0.75 | −0.23 | 0.19 | |
| 9 | SGN-U574631_snp51901 | 0.67 | 9.95 | −0.75 | −0.23 | 0.19 | |
| 9 | SGN-U567086_snp225 | 0.67 | 10.41 | −0.78 | −0.26 | 0.21 | |
| LBWV2015 | 9 | solcap_snp_sl_22830 | 0.61 | 2.91 | −0.21 | −0.13 | 0.07 |
| 9 | solcap_snp_sl_29188 | 0.66 | 7.74 | −0.37 | 0.07 | 0.17 | |
| 9 | CL016855-0847 | 0.67 | 9.46 | −0.38 | 0.12 | 0.19 | |
| 9 | solcap_snp_sl_69978 | 0.67 | 9.22 | −0.38 | 0.08 | 0.19 | |
| 9 | solcap_snp_sl_63704 | 0.67 | 8.97 | −0.37 | 0.09 | 0.18 | |
| 9 | SGN-U574631_snp51901 | 0.67 | 8.97 | −0.37 | 0.09 | 0.18 | |
| 9 | SGN-U567086_snp225 | 0.67 | 9.05 | −0.38 | 0.09 | 0.19 | |
| Late Blight | 9 | solcap_snp_sl_29188 | 0.66 | 3.46 | −0.39 | 0.07 | 0.09 |
| 9 | solcap_snp_sl_69787 | 0.66 | 3.66 | −0.38 | 0.09 | 0.09 | |
| 9 | CL016855-0847 | 0.67 | 4.31 | −0.42 | 0.05 | 0.10 | |
| 9 | solcap_snp_sl_69978 | 0.67 | 4.44 | −0.43 | 0.02 | 0.11 | |
| 9 | solcap_snp_sl_63704 | 0.67 | 3.77 | −0.39 | 0.07 | 0.09 | |
| 9 | SGN-U574631_snp51901 | 0.67 | 3.77 | −0.39 | 0.07 | 0.09 | |
| 9 | SGN-U567086_snp225 | 0.67 | 3.80 | −0.40 | 0.08 | 0.09 | |
| LBMR2014 | 10 | solcap_snp_sl_8807 | 0.64 | 2.48 | −0.34 | −0.03 | 0.02 |
| LB2014 | 10 | solcap_snp_sl_14890 | 0.60 | 2.01 | −0.58 | 0.24 | 0.04 |
| 10 | solcap_snp_sl_33168 | 0.62 | 4.07 | −0.86 | 0.28 | 0.09 | |
| 10 | CL017176-0241 | 0.63 | 5.12 | −0.95 | 0.23 | 0.11 | |
| 10 | solcap_snp_sl_8835 | 0.63 | 7.01 | −1.08 | 0.52 | 0.14 | |
| 10 | solcap_snp_sl_8807 | 0.64 | 6.85 | −1.10 | 0.41 | 0.14 | |
| LBMR2015 | 10 | solcap_snp_sl_33168 | 0.62 | 4.24 | −0.49 | −0.08 | 0.08 |
| 10 | CL017176-0241 | 0.63 | 5.80 | −0.55 | −0.08 | 0.09 | |
| 10 | solcap_snp_sl_8835 | 0.63 | 6.48 | −0.59 | −0.01 | 0.10 | |
| 10 | solcap_snp_sl_8807 | 0.64 | 6.33 | −0.61 | 0.03 | 0.10 | |
| 10 | solcap_snp_sl_33168 | 0.62 | 4.01 | −0.28 | −0.02 | 0.09 | |
| 10 | CL017176-0241 | 0.63 | 5.59 | −0.31 | −0.04 | 0.11 | |
| 10 | solcap_snp_sl_8835 | 0.63 | 6.98 | −0.36 | 0.05 | 0.14 | |
| 10 | solcap_snp_sl_8807 | 0.64 | 6.64 | −0.36 | 0.02 | 0.13 | |
| LB2015 | 10 | solcap_snp_sl_34365 | 0.54 | 2.00 | −0.49 | 0.19 | 0.07 |
| 10 | SGN-U317657_C2 | 0.60 | 2.18 | −0.47 | 0.18 | 0.06 | |
| 10 | solcap_snp_sl_59159 | 0.60 | 3.08 | −0.54 | 0.36 | 0.09 | |
| 10 | solcap_snp_sl_33168 | 0.62 | 3.22 | −0.61 | 0.06 | 0.10 | |
| 10 | CL017176-0241 | 0.63 | 3.67 | −0.65 | −0.03 | 0.10 | |
| 10 | solcap_snp_sl_8835 | 0.63 | 5.96 | −0.79 | −0.20 | 0.16 | |
| 10 | solcap_snp_sl_8807 | 0.64 | 4.74 | −0.74 | 0.02 | 0.13 | |
| Late Blight | 10 | CL017176-0241 | 0.63 | 2.45 | −0.33 | −0.01 | 0.06 |
| 10 | solcap_snp_sl_8835 | 0.63 | 3.88 | −0.42 | 0.02 | 0.09 | |
| 10 | solcap_snp_sl_8807 | 0.64 | 4.00 | −0.44 | 0.06 | 0.10 | |
| LBWV2014 | 12 | solcap_snp_sl_12664 | 0.00 | 2.21 | −0.30 | 0.10 | 0.04 |
| 12 | solcap_snp_sl_1490 | 0.01 | 3.10 | −0.37 | 0.07 | 0.06 | |
| 12 | solcap_snp_sl_1504 | 0.01 | 3.13 | −0.37 | 0.07 | 0.06 | |
| 12 | CL015195-0336 | 0.01 | 3.13 | −0.37 | 0.08 | 0.06 | |
| 12 | solcap_snp_sl_1525 | 0.01 | 2.91 | −0.34 | 0.11 | 0.06 | |
| LB2014 | 12 | solcap_snp_sl_14415 | 0.60 | 1.86 | −0.27 | 0.62 | 0.04 |
| LBMR2015 | 12 | solcap_snp_sl_1525 | 0.01 | 2.16 | −0.32 | 0.01 | 0.03 |
| Late Blight | 12 | solcap_snp_sl_14415 | 0.60 | 2.58 | −0.06 | 0.41 | 0.06 |
| Source | Degree of Freedom | Sum of Square | Mean Sum of Square | Likelihood of Odd (LOD) | %var | p-Value (χ2) | p-Value (F) |
|---|---|---|---|---|---|---|---|
| Model | 4 | 28.03788 | 7.009469 | 2.790598 | 7.080374 | 0.012026 | 0.013657 |
| Error | 170 | 367.9564 | 2.164449 | ||||
| Total | 174 | 395.9943 | |||||
| Estimated effects: | |||||||
| Parameters | Estimates | Standard Error | t-Value | ||||
| Intercept | 2.50944 | 0.11355 | 22.101 | ||||
| 9@48.0a | 0.68239 | 0.20929 | 3.261 | ||||
| 9@48.0d | −0.29844 | 0.37945 | −0.787 | ||||
| 12@12.3a | 0.01011 | 0.16344 | 0.062 | ||||
| 12@12.3d | 0.16303 | 0.22532 | 0.724 | ||||
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Panthee, D.R.; Piotrowski, A.; Ibrahem, R. Mapping Quantitative Trait Loci (QTL) for Resistance to Late Blight in Tomato. Int. J. Mol. Sci. 2017, 18, 1589. https://doi.org/10.3390/ijms18071589
Panthee DR, Piotrowski A, Ibrahem R. Mapping Quantitative Trait Loci (QTL) for Resistance to Late Blight in Tomato. International Journal of Molecular Sciences. 2017; 18(7):1589. https://doi.org/10.3390/ijms18071589
Chicago/Turabian StylePanthee, Dilip R., Ann Piotrowski, and Ragy Ibrahem. 2017. "Mapping Quantitative Trait Loci (QTL) for Resistance to Late Blight in Tomato" International Journal of Molecular Sciences 18, no. 7: 1589. https://doi.org/10.3390/ijms18071589
APA StylePanthee, D. R., Piotrowski, A., & Ibrahem, R. (2017). Mapping Quantitative Trait Loci (QTL) for Resistance to Late Blight in Tomato. International Journal of Molecular Sciences, 18(7), 1589. https://doi.org/10.3390/ijms18071589





