Histone Lysine Methylation and Neurodevelopmental Disorders
Abstract
:1. Introduction
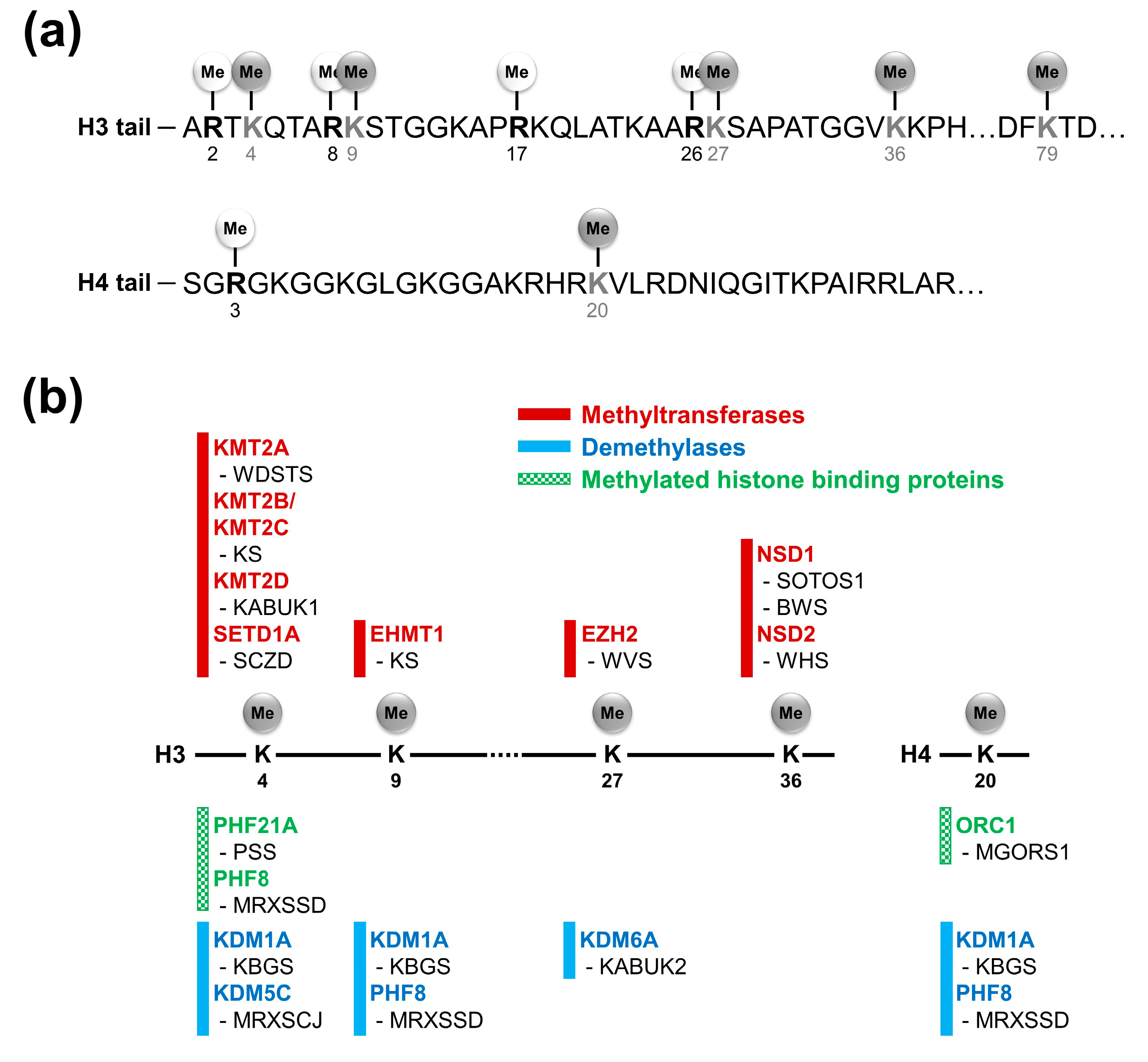
2. Histone Lysine Methylations and Related Factors
3. Neurodevelopmental Disorders Related with Histone Lysine Methylations
3.1. H3K4 Methylation
3.1.1. KMT2A and Wiedemann-Steiner Syndrome
3.1.2. KMT2D and Kabuki Syndrome 1
3.1.3. SETD1A and Schizophrenia
3.1.4. H3K4me Demethylases and Neurodevelopmental Disorders
3.1.5. PHF21A and Potocki-Shaffer Syndrome
3.2. H3K9 Methylation
3.2.1. EHMT1 and Kleefstra Syndrome
3.2.2. PHF8 and Siderius X-Linked Mental Retardation Syndrome
3.3. H3K27 Methylation
EZH2 and Weaver Syndrome
3.4. H3K36 Methylation
3.4.1. NSD1 Defects in Sotos Syndrome 1 and Beckwith-Wiedemann Syndrome
3.4.2. NSD2 and Wolf-Hirshhorn Syndrome
3.5. H4K20 Methylation
Siderius X-Linked Syndromic Mental Retardation and Meier-Gorlin Syndrome 1
4. Perspectives
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| APC2 | Adenomatosis polyposis coli 2 |
| ARX | Aristaless related homeobox |
| ASH1L | ASH1 like histone lysine methyltransferase |
| BHC complex | BRAF35/histone deacetylase complex |
| BWS | Beckwith-Wiedemann syndrome |
| DOT1L | DOT1 like histone lysine methyltransferase |
| EED | Embryonic ectoderm development |
| EHMT1 | Euchromatic histone lysine methyltransferase 1 |
| EHMT2 | Euchromatic histone lysine methyltransferase 2 |
| ERK | Extracellular signal-regulated kinase |
| ESC | Embryonic stem cells |
| EZH1 | Enhancer of zeste 1 polycomb repressive complex 2 subunit |
| EZH2 | Enhancer of zeste 2 polycomb repressive complex 2 subunit |
| FAD | Flavin adenosine dinucleotide |
| H3K4 | Histone H3 lysine 4 |
| H3K4me | Methylation on histone H3 lysine 4 |
| H3K9 | Histone H3 lysine 9 |
| H3K9me | Methylation on histone H3 lysine 9 |
| H3K27 | Histone H3 lysine 27 |
| H3K27me | Methylation on histone H3 lysine 27 |
| H3K36 | Histone H3 lysine 36 |
| H3K36me | Methylation on histone H3 lysine 36 |
| H3K79 | Histone H3 lysine 79 |
| H3K79me | Methylation on histone H3 lysine 79 |
| H3R2 | Histone H3 arginine 2 |
| H3R8 | Histone H3 arginine 8 |
| H3R17 | Histone H3 arginine 17 |
| H3R26 | Histone H3 arginine 26 |
| H4K20 | Histone H4 lysine 20 |
| H4K20me | Methylation on histone H4 lysine 20 |
| H4R3 | Histone H4 arginine 3 |
| HDAC | Histone deacetylase |
| HP1 | Heterochromatin protein 1 |
| JHDM | JmjC-domain containing histone demethylases |
| JmjC | Jumonji C |
| KABUK1 | Kabuki syndrome 1 |
| KABUK2 | Kabuki syndrome 2 |
| KBGS | KBG syndrome |
| KS | Kleefstra syndrome |
| LSD1n | Lysine-specific demethylase 1 variant |
| KDM | Lysine demethylase |
| KMT | Lysine methyl transferase |
| MAPK | Mitogen-activated protein kinase |
| MEIS2 | Myeloid ecotropic viral integration site 1 homolog 2 |
| MEK | Mitogen-activated protein kinase kinase |
| MGORS1 | Meier-Gorlin syndrome 1 |
| MRXSCJ | Mental retardation, X-linked, syndromic, Claes-Jensen type |
| MRXSSD | Siderius X-linked mental retardation syndrome |
| NFIX | Nuclear factor I X |
| NSD1 | Nuclear receptor-binding SET domain protein 1 |
| NSD2 | Nuclear receptor-binding SET domain protein 2 |
| NSD3 | Nuclear receptor-binding SET domain protein 3 |
| ORC1 | Origin recognition complex subunit 1 |
| PHF2 | PHD finger protein 2 |
| PHF21A | PHD finger protein 21A |
| PHF8 | PHD finger protein 8 |
| PRC1 | Polycomb repressive complex 1 |
| PRC2 | Polycomb repressive complex 2 |
| PRDM | PR/SET domain family |
| PSS | Potocki-Shaffer syndrome |
| RAP1A/B | RAS-related protein 1A/B |
| RIOX1 | Ribosomal oxygenase 1 |
| RUNX2 | Runt related transcription factor 2 |
| SCZD | Schizophrenia |
| SCN3A | Sodium voltage-gated channel alpha subunit 3 |
| SETD1A | SET domain containing 1A |
| SETD1B | SET domain containing 1B |
| SETD2 | SET domain containing 2 |
| SETD3 | SET domain containing 3 |
| SETDB1 | SET domain bifurcated 1 |
| SETMAR | SET domain and mariner transposase fusion gene |
| SMYD2 | SET and MYND domain containing 2 |
| SOTOS1 | Sotos syndrome 1 |
| SOTOS2 | Sotos syndrome 2 |
| SOTOS3 | Sotos syndrome 3 |
| SUV39H1 | Suppressor of variegation 3-9 homolog 1 |
| SUV39H2 | Suppressor of variegation 3-9 homolog 2 |
| UTY | Ubiquitously transcribed tetratricopeptide repeat containing, Y-linked |
| WDSTS | Wiedemann-Steiner syndrome |
| WHS | Wolf-Hirshhorn syndrome |
| WVS | Weaver syndrome |
References
- Jenuwein, T.; Allis, C.D. Translating the histone code. Science 2001, 293, 1074–1080. [Google Scholar] [CrossRef] [PubMed]
- Kouzarides, T. Chromatin modifications and their function. Cell 2007, 128, 693–705. [Google Scholar] [CrossRef] [PubMed]
- Martin, C.; Zhang, Y. The diverse functions of histone lysine methylation. Nat. Rev. Mol. Cell Biol. 2005, 6, 838–849. [Google Scholar] [CrossRef] [PubMed]
- Greer, E.L.; Shi, Y. Histone methylation: A dynamic mark in health, disease and inheritance. Nat. Rev. Genet. 2012, 13, 343–357. [Google Scholar] [CrossRef] [PubMed]
- Berger, S.L. The complex language of chromatin regulation during transcription. Nature 2007, 447, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Di Lorenzo, A.; Bedford, M.T. Histone arginine methylation. FEBS Lett. 2011, 585, 2024–2031. [Google Scholar] [CrossRef] [PubMed]
- Fuchs, J.; Demidov, D.; Houben, A.; Schubert, I. Chromosomal histone modification patterns–from conservation to diversity. Trends Plant Sci. 2006, 11, 199–208. [Google Scholar] [CrossRef] [PubMed]
- Hyun, K.; Jeon, J.; Park, K.; Kim, J. Writing, erasing and reading histone lysine methylations. Exp. Mol. Med. 2017, 49, e324. [Google Scholar] [CrossRef] [PubMed]
- Bale, T.L.; Baram, T.Z.; Brown, A.S.; Goldstein, J.M.; Insel, T.R.; McCarthy, M.M.; Nemeroff, C.B.; Reyes, T.M.; Simerly, R.B.; Susser, E.S. Early life programming and neurodevelopmental disorders. Biol. Psychiatry 2010, 68, 314–319. [Google Scholar] [CrossRef] [PubMed]
- Millan, M.J. An epigenetic framework for neurodevelopmental disorders: From pathogenesis to potential therapy. Neuropharmacology 2013, 68, 2–82. [Google Scholar] [CrossRef] [PubMed]
- Gilissen, C.; Hoischen, A.; Brunner, H.G.; Veltman, J.A. Unlocking Mendelian disease using exome sequencing. Genome Biol. 2011, 12, 228. [Google Scholar] [CrossRef] [PubMed]
- Ronan, J.L.; Wu, W.; Crabtree, G.R. From neural development to cognition: Unexpected roles for chromatin. Nat. Rev. Genet. 2013, 14, 347–359. [Google Scholar] [CrossRef] [PubMed]
- Vissers, L.E.; Gilissen, C.; Veltman, J.A. Genetic studies in intellectual disability and related disorders. Nat. Rev. Genet. 2016, 17, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Tatton-Brown, K.; Loveday, C.; Yost, S.; Clarke, M.; Ramsay, E.; Zachariou, A.; Elliott, A.; Wylie, H.; Ardissone, A.; Rittinger, O. Mutations in epigenetic regulation genes are a major cause of overgrowth with intellectual disability. Am. J. Hum. Genet. 2017, 100, 725–736. [Google Scholar] [CrossRef] [PubMed]
- Heintzman, N.D.; Stuart, R.K.; Hon, G.; Fu, Y.; Ching, C.W.; Hawkins, R.D.; Barrera, L.O.; Van Calcar, S.; Qu, C.; Ching, K.A. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 2007, 39, 311–318. [Google Scholar] [CrossRef] [PubMed]
- Heintzman, N.D.; Hon, G.C.; Hawkins, R.D.; Kheradpour, P.; Stark, A.; Harp, L.F.; Ye, Z.; Lee, L.K.; Stuart, R.K.; Ching, C.W. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 2009, 459, 108–112. [Google Scholar] [CrossRef] [PubMed]
- Allis, C.D.; Berger, S.L.; Cote, J.; Dent, S.; Jenuwien, T.; Kouzarides, T.; Pillus, L.; Reinberg, D.; Shi, Y.; Shiekhattar, R. New nomenclature for chromatin-modifying enzymes. Cell 2007, 131, 633–636. [Google Scholar] [CrossRef] [PubMed]
- Shilatifard, A. Chromatin modifications by methylation and ubiquitination: Implications in the regulation of gene expression. Annu. Rev. Biochem. 2006, 75, 243–269. [Google Scholar] [CrossRef] [PubMed]
- Rao, R.C.; Dou, Y. Hijacked in cancer: The KMT2 (MLL) family of methyltransferases. Nat. Rev. Cancer 2015, 15, 334–346. [Google Scholar] [CrossRef] [PubMed]
- Wagner, E.J.; Carpenter, P.B. Understanding the language of Lys36 methylation at histone H3. Nat. Rev. Mol. Cell Biol. 2012, 13, 115–126. [Google Scholar] [CrossRef] [PubMed]
- Bannister, A.J.; Zegerman, P.; Partridge, J.F.; Miska, E.A.; Thomas, J.O.; Allshire, R.C.; Kouzarides, T. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 2001, 410, 120–124. [Google Scholar] [CrossRef] [PubMed]
- Lachner, M.; O’carroll, D.; Rea, S.; Mechtler, K.; Jenuwein, T. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 2001, 410, 116–120. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, J.-I.; Rice, J.C.; Strahl, B.D.; Allis, C.D.; Grewal, S.I. Role of histone H3 lysine 9 methylation in epigenetic control of heterochromatin assembly. Science 2001, 292, 110–113. [Google Scholar] [CrossRef] [PubMed]
- Grewal, S.I.; Jia, S. Heterochromatin revisited. Nat. Rev. Genet. 2007, 8, 35–46. [Google Scholar] [CrossRef] [PubMed]
- Schultz, D.C.; Ayyanathan, K.; Negorev, D.; Maul, G.G.; Rauscher, F.J. SETDB1: A novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev. 2002, 16, 919–932. [Google Scholar] [CrossRef] [PubMed]
- Metzger, E.; Wissmann, M.; Yin, N.; Müller, J.M.; Schneider, R.; Peters, A.H.; Günther, T.; Buettner, R.; Schüle, R. LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature 2005, 437, 436–439. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Park, J.W.; Lee, K.H.; Yoon, H.; Shin, D.H.; Ju, U.I.; Seok, S.H.; Lim, S.H.; Lee, Z.H.; Kim, H.H. Plant homeodomain finger protein 2 promotes bone formation by demethylating and activating Runx2 for osteoblast differentiation. Cell Res. 2014, 24, 1231–1249. [Google Scholar] [CrossRef] [PubMed]
- Plath, K.; Fang, J.; Mlynarczyk-Evans, S.K.; Cao, R.; Worringer, K.A.; Wang, H.; Cecile, C.; Otte, A.P.; Panning, B.; Zhang, Y. Role of histone H3 lysine 27 methylation in X inactivation. Science 2003, 300, 131–135. [Google Scholar] [CrossRef] [PubMed]
- Wiles, E.T.; Selker, E.U. H3K27 methylation: A promiscuous repressive chromatin mark. Curr. Opin. Genet. Dev. 2017, 43, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Boyer, L.A.; Plath, K.; Zeitlinger, J.; Brambrink, T.; Medeiros, L.A.; Lee, T.I.; Levine, S.S.; Wernig, M.; Tajonar, A.; Ray, M.K. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 2006, 441, 349–353. [Google Scholar] [CrossRef] [PubMed]
- Lee, T.I.; Jenner, R.G.; Boyer, L.A.; Guenther, M.G.; Levine, S.S.; Kumar, R.M.; Chevalier, B.; Johnstone, S.E.; Cole, M.F.; Isono, K.-I. Control of developmental regulators by Polycomb in human embryonic stem cells. Cell 2006, 125, 301–313. [Google Scholar] [CrossRef] [PubMed]
- Harikumar, A.; Meshorer, E. Chromatin remodeling and bivalent histone modifications in embryonic stem cells. EMBO Rep. 2015, 16, 1609–1619. [Google Scholar] [CrossRef] [PubMed]
- Margueron, R.; Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 2011, 469, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Kizer, K.O.; Phatnani, H.P.; Shibata, Y.; Hall, H.; Greenleaf, A.L.; Strahl, B.D. A novel domain in Set2 mediates RNA polymerase II interaction and couples histone H3K36 methylation with transcript elongation. Mol. Cell. Biol. 2005, 25, 3305–3316. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Kim, H.; Fong, N.; Erickson, B.; Bentley, D.L. Pre-mRNA splicing is a determinant of histone H3K36 methylation. Proc. Natl. Acad. Sci. USA 2011, 108, 13564–13569. [Google Scholar] [CrossRef] [PubMed]
- Bell, O.; Conrad, T.; Kind, J.; Wirbelauer, C.; Akhtar, A.; Schübeler, D. Transcription-coupled methylation of histone H3 at lysine 36 regulates dosage compensation by enhancing recruitment of the MSL complex in Drosophila melanogaster. Mol. Cell. Biol. 2008, 28, 3401–3409. [Google Scholar] [CrossRef] [PubMed]
- Fnu, S.; Williamson, E.A.; De Haro, L.P.; Brenneman, M.; Wray, J.; Shaheen, M.; Radhakrishnan, K.; Lee, S.-H.; Nickoloff, J.A.; Hromas, R. Methylation of histone H3 lysine 36 enhances DNA repair by nonhomologous end-joining. Proc. Natl. Acad. Sci. USA 2011, 108, 540–545. [Google Scholar] [CrossRef] [PubMed]
- Strahl, B.D.; Grant, P.A.; Briggs, S.D.; Sun, Z.-W.; Bone, J.R.; Caldwell, J.A.; Mollah, S.; Cook, R.G.; Shabanowitz, J.; Hunt, D.F. Set2 is a nucleosomal histone H3-selective methyltransferase that mediates transcriptional repression. Mol. Cell. Biol. 2002, 22, 1298–1306. [Google Scholar] [CrossRef] [PubMed]
- Edmunds, J.W.; Mahadevan, L.C.; Clayton, A.L. Dynamic histone H3 methylation during gene induction: HYPB/Setd2 mediates all H3K36 trimethylation. EMBO J. 2008, 27, 406–420. [Google Scholar] [CrossRef] [PubMed]
- Cloos, P.A.; Christensen, J.; Agger, K.; Helin, K. Erasing the methyl mark: Histone demethylases at the center of cellular differentiation and disease. Genes Dev. 2008, 22, 1115–1140. [Google Scholar] [CrossRef] [PubMed]
- Li, G.-M. Decoding the histone code: Role of H3K36me3 in mismatch repair and implications for cancer susceptibility and therapy. Cancer Res. 2013, 73, 6379–6383. [Google Scholar] [CrossRef] [PubMed]
- Sinha, K.M.; Yasuda, H.; Coombes, M.M.; Dent, S.Y.; De Crombrugghe, B. Regulation of the osteoblast-specific transcription factor Osterix by NO66, a Jumonji family histone demethylase. EMBO J. 2010, 29, 68–79. [Google Scholar] [CrossRef] [PubMed]
- Farooq, Z.; Banday, S.; Pandita, T.K.; Altaf, M. The many faces of histone H3K79 methylation. Mutat. Res. Rev. Mutat. Res. 2016, 768, 46–52. [Google Scholar] [CrossRef] [PubMed]
- Van Leeuwen, F.; Gafken, P.R.; Gottschling, D.E. Dot1p modulates silencing in yeast by methylation of the nucleosome core. Cell 2002, 109, 745–756. [Google Scholar] [CrossRef]
- Wang, Z.; Zang, C.; Rosenfeld, J.A.; Schones, D.E.; Barski, A.; Cuddapah, S.; Cui, K.; Roh, T.-Y.; Peng, W.; Zhang, M.Q. Combinatorial patterns of histone acetylations and methylations in the human genome. Nat. Genet. 2008, 40, 897–903. [Google Scholar] [CrossRef] [PubMed]
- Kuo, A.J.; Song, J.; Cheung, P.; Ishibe-Murakami, S.; Yamazoe, S.; Chen, J.K.; Patel, D.J.; Gozani, O. The BAH domain of ORC1 links H4K20me2 to DNA replication licensing and Meier-Gorlin syndrome. Nature 2012, 484, 115–119. [Google Scholar] [CrossRef] [PubMed]
- Botuyan, M.V.; Lee, J.; Ward, I.M.; Kim, J.-E.; Thompson, J.R.; Chen, J.; Mer, G. Structural basis for the methylation state-specific recognition of histone H4K20 by 53BP1 and Crb2 in DNA repair. Cell 2006, 127, 1361–1373. [Google Scholar] [CrossRef] [PubMed]
- Jørgensen, S.; Schotta, G.; Sørensen, C.S. Histone H4 lysine 20 methylation: Key player in epigenetic regulation of genomic integrity. Nucleic Acids Res. 2013, 41, 2797–2806. [Google Scholar] [CrossRef] [PubMed]
- Nishioka, K.; Rice, J.C.; Sarma, K.; Erdjument-Bromage, H.; Werner, J.; Wang, Y.; Chuikov, S.; Valenzuela, P.; Tempst, P.; Steward, R. PR-Set7 is a nucleosome-specific methyltransferase that modifies lysine 20 of histone H4 and is associated with silent chromatin. Mol. Cell 2002, 9, 1201–1213. [Google Scholar] [CrossRef]
- Yang, H.; Pesavento, J.J.; Starnes, T.W.; Cryderman, D.E.; Wallrath, L.L.; Kelleher, N.L.; Mizzen, C.A. Preferential dimethylation of histone H4 lysine 20 by Suv4–20. J. Biol. Chem. 2008, 283, 12085–12092. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Tanasa, B.; Tyurina, O.V.; Zhou, T.Y.; Gassmann, R.; Liu, W.T.; Ohgi, K.A.; Benner, C.; Garcia-Bassets, I.; Aggarwal, A.K. PHF8 mediates histone H4 lysine 20 demethylation events involved in cell cycle progression. Nature 2010, 466, 508–512. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Telese, F.; Tan, Y.; Li, W.; Jin, C.; He, X.; Basnet, H.; Ma, Q.; Merkurjev, D.; Zhu, X. LSD1n is an H4K20 demethylase regulating memory formation via transcriptional elongation control. Nat. Neurosci. 2015, 18, 1256–1264. [Google Scholar] [CrossRef] [PubMed]
- Stender, J.D.; Pascual, G.; Liu, W.; Kaikkonen, M.U.; Do, K.; Spann, N.J.; Boutros, M.; Perrimon, N.; Rosenfeld, M.G.; Glass, C.K. Control of proinflammatory gene programs by regulated trimethylation and demethylation of histone H4K20. Mol. Cell 2012, 48, 28–38. [Google Scholar] [CrossRef] [PubMed]
- Strom, S.P.; Lozano, R.; Lee, H.; Dorrani, N.; Mann, J.; O’Lague, P.F.; Mans, N.; Deignan, J.L.; Vilain, E.; Nelson, S.F. De Novo variants in the KMT2A (MLL) gene causing atypical Wiedemann-Steiner syndrome in two unrelated individuals identified by clinical exome sequencing. BMC Med. Genet. 2014, 15, 49. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Kim, S.Y.; Artis, S.; Molfese, D.L.; Schumacher, A.; Sweatt, J.D.; Paylor, R.E.; Lubin, F.D. Histone methylation regulates memory formation. J. Neurosci. 2010, 30, 3589–3599. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.Y.; Levenson, J.M.; Korsmeyer, S.; Sweatt, J.D.; Schumacher, A. Developmental regulation of Eed complex composition governs a switch in global histone modification in brain. J. Biol. Chem. 2007, 282, 9962–9972. [Google Scholar] [CrossRef] [PubMed]
- Jakovcevski, M.; Ruan, H.; Shen, E.Y.; Dincer, A.; Javidfar, B.; Ma, Q.; Peter, C.J.; Cheung, I.; Mitchell, A.C.; Jiang, Y. Neuronal Kmt2a/Mll1 histone methyltransferase is essential for prefrontal synaptic plasticity and working memory. J. Neurosci. 2015, 35, 5097–5108. [Google Scholar] [CrossRef] [PubMed]
- Niikawa, N.; Matsuura, N.; Fukushima, Y.; Ohsawa, T.; Kajii, T. Kabuki make-up syndrome: A syndrome of mental retardation, unusual facies, large and protruding ears, and postnatal growth deficiency. J. Pediatr. 1981, 99, 565–569. [Google Scholar] [CrossRef]
- Ng, S.B.; Bigham, A.W.; Buckingham, K.J.; Hannibal, M.C.; McMillin, M.J.; Gildersleeve, H.I.; Beck, A.E.; Tabor, H.K.; Cooper, G.M.; Mefford, H.C. Exome sequencing identifies MLL2 mutations as a cause of Kabuki syndrome. Nat. Genet. 2010, 42, 790–793. [Google Scholar] [CrossRef] [PubMed]
- Paulussen, A.D.; Stegmann, A.; Blok, M.J.; Tserpelis, D.; Posma-Velter, C.; Detisch, Y.; Smeets, E.E.; Wagemans, A.; Schrander, J.J.; van den Boogaard, M.J.H. MLL2 mutation spectrum in 45 patients with Kabuki syndrome. Hum. Mutat. 2011, 32, E2018–E2025. [Google Scholar] [CrossRef] [PubMed]
- Bögershausen, N.; Wollnik, B. Unmasking Kabuki syndrome. Clin. Genet. 2013, 83, 201–211. [Google Scholar] [CrossRef] [PubMed]
- Lederer, D.; Grisart, B.; Digilio, M.C.; Benoit, V.; Crespin, M.; Ghariani, S.C.; Maystadt, I.; Dallapiccola, B.; Verellen-Dumoulin, C. Deletion of KDM6A, a histone demethylase interacting with MLL2, in three patients with Kabuki syndrome. Am. J. Hum. Genet. 2012, 90, 119–124. [Google Scholar] [CrossRef] [PubMed]
- Benjamin, J.S.; Pilarowski, G.O.; Carosso, G.A.; Zhang, L.; Huso, D.L.; Goff, L.A.; Vernon, H.J.; Hansen, K.D.; Bjornsson, H.T. A ketogenic diet rescues hippocampal memory defects in a mouse model of Kabuki syndrome. Proc. Natl. Acad. Sci. USA 2017, 114, 125–130. [Google Scholar] [CrossRef] [PubMed]
- Bjornsson, H.T.; Benjamin, J.S.; Zhang, L.; Weissman, J.; Gerber, E.E.; Chen, Y.-C.; Vaurio, R.G.; Potter, M.C.; Hansen, K.D.; Dietz, H.C. Histone deacetylase inhibition rescues structural and functional brain deficits in a mouse model of Kabuki syndrome. Sci. Transl. Med. 2014, 6, 256ra135. [Google Scholar] [CrossRef] [PubMed]
- Takata, A.; Xu, B.; Ionita-Laza, I.; Roos, J.L.; Gogos, J.A.; Karayiorgou, M. Loss-of-function variants in schizophrenia risk and SETD1A as a candidate susceptibility gene. Neuron 2014, 82, 773–780. [Google Scholar] [CrossRef] [PubMed]
- Singh, T.; Kurki, M.I.; Curtis, D.; Purcell, S.M.; Crooks, L.; McRae, J.; Suvisaari, J.; Chheda, H.; Blackwood, D.; Breen, G. Rare loss-of-function variants in SETD1A are associated with schizophrenia and developmental disorders. Nat. Neurosci. 2016, 19, 571–577. [Google Scholar] [CrossRef] [PubMed]
- Takata, A.; Ionita-Laza, I.; Gogos, J.A.; Xu, B.; Karayiorgou, M. De novo synonymous mutations in regulatory elements contribute to the genetic etiology of autism and schizophrenia. Neuron 2016, 89, 940–947. [Google Scholar] [CrossRef] [PubMed]
- Najmabadi, H.; Hu, H.; Garshasbi, M.; Zemojtel, T.; Abedini, S.S.; Chen, W.; Hosseini, M.; Behjati, F.; Haas, S.; Jamali, P. Deep sequencing reveals 50 novel genes for recessive cognitive disorders. Nature 2011, 478, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Jensen, L.R.; Amende, M.; Gurok, U.; Moser, B.; Gimmel, V.; Tzschach, A.; Janecke, A.R.; Tariverdian, G.; Chelly, J.; Fryns, J.-P. Mutations in the JARID1C gene, which is involved in transcriptional regulation and chromatin remodeling, cause X-linked mental retardation. Am. J. Hum. Genet. 2005, 76, 227–236. [Google Scholar] [CrossRef] [PubMed]
- Tahiliani, M.; Mei, P.; Fang, R.; Leonor, T.; Rutenberg, M.; Shimizu, F.; Li, J.; Rao, A.; Shi, Y. The histone H3K4 demethylase SMCX links REST target genes to X-linked mental retardation. Nature 2007, 447, 601–605. [Google Scholar] [CrossRef] [PubMed]
- Poeta, L.; Fusco, F.; Drongitis, D.; Shoubridge, C.; Manganelli, G.; Filosa, S.; Paciolla, M.; Courtney, M.; Collombat, P.; Lioi, M.B. A regulatory path associated with X-linked intellectual disability and epilepsy links KDM5C to the polyalanine expansions in ARX. Am. J. Hum. Genet. 2013, 92, 114–125. [Google Scholar] [CrossRef] [PubMed]
- Strømme, P.; Mangelsdorf, M.E.; Scheffer, I.E.; Gécz, J. Infantile spasms, dystonia, and other X-linked phenotypes caused by mutations in Aristaless related homeobox gene, ARX. Brain Dev. 2002, 24, 266–268. [Google Scholar] [CrossRef]
- Kitamura, K.; Yanazawa, M.; Sugiyama, N.; Miura, H.; Iizuka-Kogo, A.; Kusaka, M.; Omichi, K.; Suzuki, R.; Kato-Fukui, Y.; Kamiirisa, K. Mutation of ARX causes abnormal development of forebrain and testes in mice and X-linked lissencephaly with abnormal genitalia in humans. Nat. Genet. 2002, 32, 359–369. [Google Scholar] [CrossRef] [PubMed]
- Kato, M.; Das, S.; Petras, K.; Kitamura, K.; Morohashi, K.I.; Abuelo, D.N.; Barr, M.; Bonneau, D.; Brady, A.F.; Carpenter, N.J. Mutations of ARX are associated with striking pleiotropy and consistent genotype–phenotype correlation. Hum. Mutat. 2004, 23, 147–159. [Google Scholar] [CrossRef] [PubMed]
- Tunovic, S.; Barkovich, J.; Sherr, E.H.; Slavotinek, A.M. De novo ANKRD11 and KDM1A gene mutations in a male with features of KBG syndrome and Kabuki syndrome. Am. J. Med. Genet. 2014, 164, 1744–1749. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y. Histone lysine demethylases: Emerging roles in development, physiology and disease. Nat. Rev. Genet. 2007, 8, 829–833. [Google Scholar] [CrossRef] [PubMed]
- Lan, F.; Collins, R.E.; De Cegli, R.; Alpatov, R.; Horton, J.R.; Shi, X.; Gozani, O.; Cheng, X.; Shi, Y. Recognition of unmethylated histone H3 lysine 4 links BHC80 to LSD1-mediated gene repression. Nature 2007, 448, 718–722. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.-G.; Kim, H.-T.; Leach, N.T.; Lan, F.; Ullmann, R.; Silahtaroglu, A.; Kurth, I.; Nowka, A.; Seong, I.S.; Shen, Y. Translocations disrupting PHF21A in the Potocki-Shaffer-syndrome region are associated with intellectual disability and craniofacial anomalies. Am. J. Hum. Genet. 2012, 91, 56–72. [Google Scholar] [CrossRef] [PubMed]
- Labonne, J.D.; Vogt, J.; Reali, L.; Kong, I.K.; Layman, L.C.; Kim, H.G. A microdeletion encompassing PHF21A in an individual with global developmental delay and craniofacial anomalies. Am. J. Med. Genet. 2015, 167, 3011–3018. [Google Scholar] [CrossRef] [PubMed]
- Hakimi, M.-A.; Bochar, D.A.; Chenoweth, J.; Lane, W.S.; Mandel, G.; Shiekhattar, R. A core–BRAF35 complex containing histone deacetylase mediates repression of neuronal-specific genes. Proc. Natl. Acad. Sci. USA 2002, 99, 7420–7425. [Google Scholar] [CrossRef] [PubMed]
- Kleefstra, T.; Smidt, M.; Banning, M.; Oudakker, A.; Van Esch, H.; De Brouwer, A.; Nillesen, W.; Sistermans, E.; Hamel, B.; De Bruijn, D. Disruption of the gene Euchromatin Histone Methyl Transferase1 (Eu-HMTase1) is associated with the 9q34 subtelomeric deletion syndrome. J. Med. Genet. 2005, 42, 299–306. [Google Scholar] [CrossRef] [PubMed]
- Kleefstra, T.; Brunner, H.G.; Amiel, J.; Oudakker, A.R.; Nillesen, W.M.; Magee, A.; Geneviève, D.; Cormier-Daire, V.; Van Esch, H.; Fryns, J.-P. Loss-of-function mutations in euchromatin histone methyl transferase 1 (EHMT1) cause the 9q34 subtelomeric deletion syndrome. Am. J. Hum. Genet. 2006, 79, 370–377. [Google Scholar] [CrossRef] [PubMed]
- Harada, N.; Visser, R.; Dawson, A.; Fukamachi, M.; Iwakoshi, M.; Okamoto, N.; Kishino, T.; Niikawa, N.; Matsumoto, N. A 1-Mb critical region in six patients with 9q34. 3 terminal deletion syndrome. J. Hum. Genet. 2004, 49, 440–444. [Google Scholar] [CrossRef] [PubMed]
- Kramer, J.M.; Kochinke, K.; Oortveld, M.A.; Marks, H.; Kramer, D.; de Jong, E.K.; Asztalos, Z.; Westwood, J.T.; Stunnenberg, H.G.; Sokolowski, M.B. Epigenetic regulation of learning and memory by Drosophila EHMT/G9a. PLoS Biol. 2011, 9, e1000569. [Google Scholar] [CrossRef] [PubMed]
- Balemans, M.; Nadif Kasri, N.; Kopanitsa, M.V.; Afinowi, N.O.; Ramakers, G.; Peters, T.A.; Beynon, A.J.; Janssen, S.M.; van Summeren, R.C.; Eeftens, J.M. Hippocampal dysfunction in the Euchromatin histone methyltransferase 1 heterozygous knockout mouse model for Kleefstra syndrome. Hum. Mol. Genet. 2013, 22, 852–866. [Google Scholar] [CrossRef] [PubMed]
- Martens, M.B.; Frega, M.; Classen, J.; Epping, L.; Bijvank, E.; Benevento, M.; van Bokhoven, H.; Tiesinga, P.; Schubert, D.; Kasri, N.N. Euchromatin histone methyltransferase 1 regulates cortical neuronal network development. Sci. Rep. 2016, 6, 35756. [Google Scholar] [CrossRef] [PubMed]
- Benevento, M.; Iacono, G.; Selten, M.; Ba, W.; Oudakker, A.; Frega, M.; Keller, J.; Mancini, R.; Lewerissa, E.; Kleefstra, T. Histone methylation by the Kleefstra syndrome protein EHMT1 mediates homeostatic synaptic scaling. Neuron 2016, 91, 341–355. [Google Scholar] [CrossRef] [PubMed]
- Kleefstra, T.; Kramer, J.M.; Neveling, K.; Willemsen, M.H.; Koemans, T.S.; Vissers, L.E.; Wissink-Lindhout, W.; Fenckova, M.; van den Akker, W.M.; Kasri, N.N. Disruption of an EHMT1-associated chromatin-modification module causes intellectual disability. Am. J. Hum. Genet. 2012, 91, 73–82. [Google Scholar] [CrossRef] [PubMed]
- Agha, Z.; Iqbal, Z.; Azam, M.; Ayub, H.; Vissers, L.E.; Gilissen, C.; Ali, S.H.B.; Riaz, M.; Veltman, J.A.; Pfundt, R. Exome sequencing identifies three novel candidate genes implicated in intellectual disability. PLoS ONE 2014, 9, e112687. [Google Scholar] [CrossRef] [PubMed]
- Siderius, L.E.; Hamel, B.C.; van Bokhoven, H.; de Jager, F.; van den Helm, B.; Kremer, H.; Heineman-de Boer, J.A.; Ropers, H.H.; Mariman, E.C. X-linked mental retardation associated with cleft lip/palate maps to Xp11.3-q21.3. Am. J. Med. Genet. 1999, 85, 216–220. [Google Scholar] [CrossRef]
- Laumonnier, F.; Holbert, S.; Ronce, N.; Faravelli, F.; Lenzner, S.; Schwartz, C.; Lespinasse, J.; Van Esch, H.; Lacombe, D.; Goizet, C. Mutations in PHF8 are associated with X linked mental retardation and cleft lip/cleft palate. J. Med. Genet. 2005, 42, 780–786. [Google Scholar] [CrossRef] [PubMed]
- Abidi, F.; Miano, M.; Murray, J.; Schwartz, C. A novel mutation in the PHF8 gene is associated with X-linked mental retardation with cleft lip/cleft palate. Clin. Genet. 2007, 72, 19–22. [Google Scholar] [CrossRef] [PubMed]
- Koivisto, A.; Ala-Mello, S.; Lemmelä, S.; Komu, H.; Rautio, J.; Järvelä, I. Screening of mutations in the PHF8 gene and identification of a novel mutation in a Finnish family with XLMR and cleft lip/cleft palate. Clin. Genet. 2007, 72, 145–149. [Google Scholar] [CrossRef] [PubMed]
- Feng, W.; Yonezawa, M.; Ye, J.; Jenuwein, T.; Grummt, I. PHF8 activates transcription of rRNA genes through H3K4me3 binding and H3K9me1/2 demethylation. Nat. Struct. Mol. Biol. 2010, 17, 445–450. [Google Scholar] [CrossRef] [PubMed]
- Kleine-Kohlbrecher, D.; Christensen, J.; Vandamme, J.; Abarrategui, I.; Bak, M.; Tommerup, N.; Shi, X.; Gozani, O.; Rappsilber, J.; Salcini, A.E. A functional link between the histone demethylase PHF8 and the transcription factor ZNF711 in X-linked mental retardation. Mol. Cell 2010, 38, 165–178. [Google Scholar] [CrossRef] [PubMed]
- Qi, H.H.; Sarkissian, M.; Hu, G.-Q.; Wang, Z.; Bhattacharjee, A.; Gordon, D.B.; Gonzales, M.; Lan, F.; Ongusaha, P.P.; Huarte, M. Histone H4K20/H3K9 demethylase PHF8 regulates zebrafish brain and craniofacial development. Nature 2010, 466, 503–507. [Google Scholar] [CrossRef] [PubMed]
- Riveiro, A.R.; Mariani, L.; Malmberg, E.; Amendola, P.G.; Peltonen, J.; Wong, G.; Salcini, A.E. JMJD-1.2/PHF8 controls axon guidance by regulating Hedgehog-like signaling. Development 2017, 144, 856–865. [Google Scholar] [CrossRef] [PubMed]
- Walsh, R.M.; Shen, E.Y.; Bagot, R.C.; Anselmo, A.; Jiang, Y.; Javidfar, B.; Wojtkiewicz, G.J.; Cloutier, J.; Chen, J.W.; Sadreyev, R. Phf8 loss confers resistance to depression-like and anxiety-like behaviors in mice. Nat. Commun. 2017, 8, 15142. [Google Scholar] [CrossRef] [PubMed]
- Weaver, D.D.; Graham, C.B.; Thomas, I.; Smith, D.W. A new overgrowth syndrome with accelerated skeletal maturation, unusual facies, and camptodactyly. J. Pediatr. 1974, 84, 547–552. [Google Scholar] [CrossRef]
- Cohen, M.M. Mental deficiency, alterations in performance, and CNS abnormalities in overgrowth syndromes. Am. J. Med. Genet. 2003, 117C, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Tatton-Brown, K.; Murray, A.; Hanks, S.; Douglas, J.; Armstrong, R.; Banka, S.; Bird, L.M.; Clericuzio, C.L.; Cormier-Daire, V.; Cushing, T. Weaver syndrome and EZH2 mutations: Clarifying the clinical phenotype. Am. J. Med. Genet. 2013, 161, 2972–2980. [Google Scholar] [CrossRef] [PubMed]
- Tatton-Brown, K.; Hanks, S.; Ruark, E.; Zachariou, A.; Duarte, S.D.V.; Ramsay, E.; Snape, K.; Murray, A.; Perdeaux, E.R.; Seal, S. Germline mutations in the oncogene EZH2 cause Weaver syndrome and increased human height. Oncotarget 2011, 2, 1127–1133. [Google Scholar] [CrossRef] [PubMed]
- Gibson, W.T.; Hood, R.L.; Zhan, S.H.; Bulman, D.E.; Fejes, A.P.; Moore, R.; Mungall, A.J.; Eydoux, P.; Babul-Hirji, R.; An, J. Mutations in EZH2 cause Weaver syndrome. Am. J. Hum. Genet. 2012, 90, 110–118. [Google Scholar] [CrossRef] [PubMed]
- Cao, R.; Wang, L.; Wang, H.; Xia, L.; Erdjument-Bromage, H.; Tempst, P.; Jones, R.S.; Zhang, Y. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 2002, 298, 1039–1043. [Google Scholar] [CrossRef] [PubMed]
- Cohen, A.S.; Tuysuz, B.; Shen, Y.; Bhalla, S.K.; Jones, S.J.; Gibson, W.T. A novel mutation in EED associated with overgrowth. J. Hum. Genet. 2015, 60, 339–342. [Google Scholar] [CrossRef] [PubMed]
- Pereira, J.D.; Sansom, S.N.; Smith, J.; Dobenecker, M.-W.; Tarakhovsky, A.; Livesey, F.J. Ezh2, the histone methyltransferase of PRC2, regulates the balance between self-renewal and differentiation in the cerebral cortex. Proc. Natl. Acad. Sci. USA 2010, 107, 15957–15962. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Juan, A.H.; Wang, H.A.; Ko, K.D.; Zare, H.; Sartorelli, V. Polycomb Ezh2 controls the fate of GABAergic neurons in the embryonic cerebellum. Development 2016, 143, 1971–1980. [Google Scholar] [CrossRef] [PubMed]
- Akizu, N.; García, M.A.; Estarás, C.; Fueyo, R.; Badosa, C.; de la Cruz, X.; Martínez-Balbás, M.A. EZH2 regulates neuroepithelium structure and neuroblast proliferation by repressing p21. Open Biol. 2016, 6, 150227. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Ji, F.; Liu, Y.; Lei, X.; Li, H.; Ji, G.; Yuan, Z.; Jiao, J. Ezh2 regulates adult hippocampal neurogenesis and memory. J. Neurosci. 2014, 34, 5184–5199. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Taylor, R.J.; La Torre, A.; Wilken, M.S.; Cox, K.E.; Reh, T.A.; Vetter, M.L. Ezh2 maintains retinal progenitor proliferation, transcriptional integrity, and the timing of late differentiation. Dev. Biol. 2015, 403, 128–138. [Google Scholar] [CrossRef] [PubMed]
- Hirabayashi, Y.; Suzki, N.; Tsuboi, M.; Endo, T.A.; Toyoda, T.; Shinga, J.; Koseki, H.; Vidal, M.; Gotoh, Y. Polycomb limits the neurogenic competence of neural precursor cells to promote astrogenic fate transition. Neuron 2009, 63, 600–613. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.-L.; Chou, R.-H.; Chen, L.-T.; Shyu, W.-C.; Hsieh, S.-C.; Wu, C.-S.; Zeng, H.-J.; Yeh, S.-P.; Yang, D.-M.; Hung, S.-C. EZH2 regulates neuronal differentiation of mesenchymal stem cells through PIP5K1C-dependent calcium signaling. J. Biol. Chem. 2011, 286, 9657–9667. [Google Scholar] [CrossRef] [PubMed]
- Hwang, W.W.; Salinas, R.D.; Siu, J.J.; Kelley, K.W.; Delgado, R.N.; Paredes, M.F.; Alvarez-Buylla, A.; Oldham, M.C.; Lim, D.A. Distinct and separable roles for EZH2 in neurogenic astroglia. Elife 2014, 3, e02439. [Google Scholar] [CrossRef] [PubMed]
- Di Meglio, T.; Kratochwil, C.F.; Vilain, N.; Loche, A.; Vitobello, A.; Yonehara, K.; Hrycaj, S.M.; Roska, B.; Peters, A.H.; Eichmann, A. Ezh2 orchestrates topographic migration and connectivity of mouse precerebellar neurons. Science 2013, 339, 204–207. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Rank, G.; Tan, Y.T.; Li, H.; Moritz, R.L.; Simpson, R.J.; Cerruti, L.; Curtis, D.J.; Patel, D.J.; Allis, C.D. PRMT5-mediated methylation of histone H4R3 recruits DNMT3A, coupling histone and DNA methylation in gene silencing. Nat. Struct. Mol. Biol. 2009, 16, 304–311. [Google Scholar] [CrossRef] [PubMed]
- Tien, C.-L.; Jones, A.; Wang, H.; Gerigk, M.; Nozell, S.; Chang, C. Snail2/Slug cooperates with Polycomb repressive complex 2 (PRC2) to regulate neural crest development. Development 2015, 142, 722–731. [Google Scholar] [CrossRef] [PubMed]
- Kurotaki, N.; Imaizumi, K.; Harada, N.; Masuno, M.; Kondoh, T.; Nagai, T.; Ohashi, H.; Naritomi, K.; Tsukahara, M.; Makita, Y. Haploinsufficiency of NSD1 causes Sotos syndrome. Nat. Genet. 2002, 30, 365–366. [Google Scholar] [CrossRef] [PubMed]
- Sotos, J.F.; Dodge, P.R.; Muirhead, D.; Crawford, J.D.; Talbot, N.B. Cerebral gigantism in childhood: A syndrome of excessively rapid growth with acromegalic features and a nonprogressive neurologic disorder. N. Engl. J. Med. 1964, 271, 109–116. [Google Scholar] [CrossRef] [PubMed]
- Žilina, O.; Reimand, T.; Tammur, P.; Tillmann, V.; Kurg, A.; Õunap, K. Patient with dup (5) (q35.2-q35.3) reciprocal to the common Sotos syndrome deletion and review of the literature. Eur. J. Med. Genet. 2013, 56, 202–206. [Google Scholar] [CrossRef] [PubMed]
- Berdasco, M.; Ropero, S.; Setien, F.; Fraga, M.F.; Lapunzina, P.; Losson, R.; Alaminos, M.; Cheung, N.-K.; Rahman, N.; Esteller, M. Epigenetic inactivation of the Sotos overgrowth syndrome gene histone methyltransferase NSD1 in human neuroblastoma and glioma. Proc. Natl. Acad. Sci. USA 2009, 106, 21830–21835. [Google Scholar] [CrossRef] [PubMed]
- Visser, R.; Landman, E.B.; Goeman, J.; Wit, J.M.; Karperien, M. Sotos syndrome is associated with deregulation of the MAPK/ERK-signaling pathway. PLoS ONE 2012, 7, e49229. [Google Scholar] [CrossRef] [PubMed]
- Malan, V.; Rajan, D.; Thomas, S.; Shaw, A.C.; Louis Dit Picard, H.; Layet, V.; Till, M.; van Haeringen, A.; Mortier, G.; Nampoothiri, S.; et al. Distinct effects of allelic NFIX mutations on nonsense-mediated mRNA decay engender either a Sotos-like or a Marshall-Smith syndrome. Am. J. Hum. Genet. 2010, 87, 189–198. [Google Scholar] [CrossRef] [PubMed]
- Almuriekhi, M.; Shintani, T.; Fahiminiya, S.; Fujikawa, A.; Kuboyama, K.; Takeuchi, Y.; Nawaz, Z.; Nadaf, J.; Kamel, H.; Kitam, A.K. Loss-of-function mutation in APC2 causes Sotos syndrome features. Cell Rep. 2015, 10, 1585–1598. [Google Scholar] [CrossRef] [PubMed]
- Engström, W.; Lindham, S.; Schofield, P. Wiedemann-Beckwith syndrome. Eur. J. Pediatr. 1988, 147, 450–457. [Google Scholar] [CrossRef] [PubMed]
- Martínez-y-Martínez, R.; Martínez-Carboney, R.; Ocampo-Campos, R.; Rivera, H.; Cuevas, A.; Martín, M.M. Wiedemann-Beckwith syndrome: Clinical, cytogenetical and radiological observations in 39 new cases. Genet. Couns. 1992, 3, 67–76. [Google Scholar] [PubMed]
- Waziri, M.; Patil, S.R.; Hanson, J.W.; Bartley, J.A. Abnormality of chromosome 11 in patients with features of Beckwith-Wiedemann syndrome. J. Pediatr. 1983, 102, 873–876. [Google Scholar] [CrossRef]
- Baujat, G.; Rio, M.; Rossignol, S.; Sanlaville, D.; Lyonnet, S.; Le Merrer, M.; Munnich, A.; Gicquel, C.; Cormier-Daire, V.; Colleaux, L. Paradoxical NSD1 mutations in Beckwith-Wiedemann syndrome and 11p15 anomalies in Sotos syndrome. Am. J. Hum. Genet. 2004, 74, 715–720. [Google Scholar] [CrossRef] [PubMed]
- Battaglia, A.; Carey, J.C.; Cederholm, P.; Viskochil, D.H.; Brothman, A.R.; Galasso, C. Natural history of Wolf-Hirschhorn syndrome: Experience with 15 cases. Pediatrics 1999, 103, 830–836. [Google Scholar] [CrossRef] [PubMed]
- Näf, D.; Wilson, L.A.; Bergstrom, R.A.; Smith, R.S.; Goodwin, N.C.; Verkerk, A.; van Ommen, G.J.; Ackerman, S.L.; Frankel, W.N.; Schimenti, J.C. Mouse models for the Wolf–Hirschhorn deletion syndrome. Hum. Mol. Genet. 2001, 10, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.F.; Nimura, K.; Lo, W.N.; Saga, K.; Kaneda, Y. Histone H3 lysine 36 methyltransferase Whsc1 promotes the association of RUNX2 and p300 in the activation of bone-related genes. PLoS ONE 2014, 9, e106661. [Google Scholar] [CrossRef] [PubMed]
- Bicknell, L.S.; Walker, S.; Klingseisen, A.; Stiff, T.; Leitch, A.; Kerzendorfer, C.; Martin, C.-A.; Yeyati, P.; Al Sanna, N.; Bober, M. Mutations in ORC1, encoding the largest subunit of the origin recognition complex, cause microcephalic primordial dwarfism resembling Meier-Gorlin syndrome. Nat. Genet. 2011, 43, 350–355. [Google Scholar] [CrossRef] [PubMed]
- Shalev, S.A.; Hall, J.G. Another adult with Meier-Gorlin syndrome-insights into the natural history. Clin. Dysmorphol. 2003, 12, 167–169. [Google Scholar] [CrossRef] [PubMed]
- Bicknell, L.S.; Bongers, E.M.; Leitch, A.; Brown, S.; Schoots, J.; Harley, M.E.; Aftimos, S.; Al-Aama, J.Y.; Bober, M.; Brown, P.A. Mutations in the pre-replication complex cause Meier-Gorlin syndrome. Nat. Genet. 2011, 43, 356–359. [Google Scholar] [CrossRef] [PubMed]
- Peter, C.J.; Akbarian, S. Balancing histone methylation activities in psychiatric disorders. Trends Mol. Med. 2011, 17, 372–379. [Google Scholar] [CrossRef] [PubMed]
- Copeland, R.A.; Solomon, M.E.; Richon, V.M. Protein methyltransferases as a target class for drug discovery. Nat. Rev. Drug Discov. 2009, 8, 724–732. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liu, K.; Qin, S.; Xu, C.; Min, J. Epigenetic targets and drug discovery: Part 1: Histone methylation. Pharmacol. Ther. 2014, 143, 275–294. [Google Scholar] [CrossRef] [PubMed]
- Knutson, S.K.; Wigle, T.J.; Warholic, N.M.; Sneeringer, C.J.; Allain, C.J.; Klaus, C.R.; Sacks, J.D.; Raimondi, A.; Majer, C.R.; Song, J. A selective inhibitor of EZH2 blocks H3K27 methylation and kills mutant lymphoma cells. Nat. Chem. Biol. 2012, 8, 890–896. [Google Scholar] [CrossRef] [PubMed]
- McCabe, M.T.; Ott, H.M.; Ganji, G.; Korenchuk, S.; Thompson, C.; Van Aller, G.S.; Liu, Y.; Graves, A.P.; Diaz, E.; LaFrance, L.V. EZH2 inhibition as a therapeutic strategy for lymphoma with EZH2-activating mutations. Nature 2012, 492, 108–112. [Google Scholar] [CrossRef] [PubMed]
- Qi, W.; Chan, H.; Teng, L.; Li, L.; Chuai, S.; Zhang, R.; Zeng, J.; Li, M.; Fan, H.; Lin, Y. Selective inhibition of Ezh2 by a small molecule inhibitor blocks tumor cells proliferation. Proc. Natl. Acad. Sci. USA 2012, 109, 21360–21365. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.H.; Sætrom, P.; Snøve, O.; Rossi, J.J. MicroRNA-directed transcriptional gene silencing in mammalian cells. Proc. Natl. Acad. Sci. USA 2008, 105, 16230–16235. [Google Scholar] [CrossRef] [PubMed]
- Varambally, S.; Cao, Q.; Mani, R.-S.; Shankar, S.; Wang, X.; Ateeq, B.; Laxman, B.; Cao, X.; Jing, X.; Ramnarayanan, K. Genomic loss of microRNA-101 leads to overexpression of histone methyltransferase EZH2 in cancer. Science 2008, 322, 1695–1699. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, P.G.; Santoso, S.; Adams, C.; Anest, V.; Morris, K.V. Promoter targeted small RNAs induce long-term transcriptional gene silencing in human cells. Nucleic Acids Res. 2009, 37, 2984–2995. [Google Scholar] [CrossRef] [PubMed]
| Symbol | Previous Symbol | Synonym(s) | Residue | Function |
|---|---|---|---|---|
| ASH1L | ASH1L | ASH1, ASH1L1, huASH1, KMT2H | H3K36 | Methyltransferase |
| DOT1L | DOT1L | DOT1, KIAA1814, KMT4 | H3K79 | Methyltransferase |
| EHMT1 | EHMT1 | bA188C12.1, Eu-HMTase1, FLJ12879, KIAA1876, KMT1D | H3K9 | Methyltransferase |
| EHMT2 | BAT8, C6orf30 | Em:AF134726.3, G9A, KMT1C, NG36/G9a | H3K9 | Methyltransferase |
| EZH1 | EZH1 | KIAA0388, KMT6B | H3K27 | Methyltransferase |
| EZH2 | EZH2 | ENX-1, EZH1, KMT6, KMT6A | H3K27 | Methyltransferase |
| KDM1A | AOF2, KDM1 | BHC110, KIAA0601, LSD1 | H3K4, H3K9, H4K20 | Demethylase |
| KDM2A | FBXL11, KDM2A | CXXC8, DKFZP434M1735, FBL11, FBL7, FLJ00115, JHDM1A, KIAA1004, LILINA | H3K36 | Demethylase |
| KDM2B | FBXL10, KDM2B | CXXC2, Fbl10, JHDM1B, PCCX2 | H3K36 | Demethylase |
| KDM3A | JMJD1, JMJD1A, KDM3A | JHMD2A, KIAA0742, TSGA | H3K9 | Demethylase |
| KDM3B | C5orf7, JMJD1B, KDM3B | KIAA1082, NET22 | H3K9 | Demethylase |
| KDM4A | JMJD2, JMJD2A, KDM4A | JHDM3A, KIAA0677, TDRD14A | H3K9, H3K36 | Demethylase |
| KDM4B | JMJD2B, KDM4B | KIAA0876, TDRD14B | H3K9, H3K36 | Demethylase |
| KDM4C | JMJD2C, KDM4C | GASC1, KIAA0780, TDRD14C | H3K9, H3K36 | Demethylase |
| KDM5A | JARID1A, KDM5A, RBBP2 | - | H3K4 | Demethylase |
| KDM5C | JARID1C, KDM5C, MRX13, SMCX | DXS1272E, XE169 | H3K4 | Demethylase |
| KDM6A | KDM6A, UTX | - | H3K27 | Demethylase |
| KDM6B | JMJD3, KDM6B | KIAA0346 | H3K27 | Demethylase |
| KMT2A | KMT2A, MLL | ALL-1, CXXC7, HRX, HTRX1, MLL1A, TRX1 | H3K4 | Methyltransferase |
| KMT2B | KMT2B | CXXC10, HRX2, KIAA0304, MLL1B, MLL2, MLL4, TRX2, WBP7 | H3K4 | Methyltransferase |
| KMT2C | KMT2C, MLL3 | HALR, KIAA1506 | H3K4 | Methyltransferase |
| KMT2D | KMT2D, MLL2, TNRC21 | ALR, CAGL114, MLL4 | H3K4 | Methyltransferase |
| KMT5A | KMT5A, SETD8 | PR-Set7, SET07, SET8 | H4K20 | Methyltransferase |
| KMT5B | KMT5B, SUV420H1 | CGI-85 | H4K20 | Methyltransferase |
| KMT5C | KMT5C, SUV420H2 | MGC2705 | H4K20 | Methyltransferase |
| NSD1 | STO | ARA267, FLJ22263, KMT3B | H3K36 | Methyltransferase |
| NSD2 | WHSC1 | KMT3G, MMSET | H3K36 | Methyltransferase |
| NSD3 | WHSC1L1 | FLJ20353, KMT3F, WHISTLE | H3K36 | Methyltransferase |
| ORC1 | ORC1L | HSORC1, PARC1 | H4K20 | Recognition |
| PHF2 | - | CENP-35, JHDM1E, KDM7C, KIAA0662 | H3K9, H4K20 | Demethylase |
| PHF8 | - | JHDM1F, KDM7B, KIAA1111, ZNF422 | H3K9, H4K20/H3K4 | Demethylase/Recognition |
| PHF21A | - | BHC80, BM-006, KIAA1696 | H3K4 | Recognition |
| RIOX1 | C14orf169 | FLJ21802, JMJD9, MAPJD, NO66 | H3K4, H3K36 | Demethylase |
| SETD1A | - | KIAA0339, KMT2F, Set1 | H3K4 | Methyltransferase |
| SETD1B | - | KIAA1076, KMT2G, Set1B | H3K4 | Methyltransferase |
| SETD2 | - | FLJ23184, HIF-1, HYPB, KIAA1732, KMT3A | H3K36 | Methyltransferase |
| SETD3 | C14orf154 | FLJ23027 | H3K4, H3K36 | Methyltransferase |
| SETDB1 | SETDB1 | ESET, KG1T, KIAA0067, KMT1E, TDRD21 | H3K9 | Methyltransferase |
| SETMAR | - | Mentase | H3K4, H3K36 | Methyltransferase |
| SMYD2 | - | HSKM-B, KMT3C, ZMYND14 | H3K4, H3K36 | Methyltransferase |
| SUV39H1 | SUV39H | KMT1A | H3K9 | Methyltransferase |
| SUV39H2 | SUV39H2 | KMT1B FLJ23414 | H3K9 | Methyltransferase |
| UTY | UTY | KDM6AL, KDM6C | H3K27 | Demethylase |
| The names of the proteins are followed by HUGO Gene Nomenclature Committee (http://www.genenames.org/) | ||||
| Disorder | OMIM | Symptom | Gene | Residue | Function |
|---|---|---|---|---|---|
| Beckwith-Wiedemann syndrome (BWS) | 130650 | Pediatric overgrowth disorder involving a predisposition to tumor development | NSD1 | H3K36 | Methyltransferase |
| Kabuki syndrome 1 | 147920 | Congenital mental retardation, postnatal dwarfism, peculiar faces, broad and depressed nasal tip, large prominent earlobes, cleft or high-arched palate, scoliosis, short fifth finger, and persistence of finger pads | KMT2D KDM6A | H3K4 H3K27 | Methyltransferase Demethylase |
| Kabuki syndrome 2 (KABUK1/2) | 300867 | ||||
| KBG syndrome (KBGS) | 148050 | Macrodontia of the upper central incisors, distinctive craniofacial findings, short stature, skeletal anomalies, neurologic involvement that includes global developmental delay, seizures, and intellectual disability | KDM1A | H3K4 H3K9 H4K20 | Demethylase Demethylase Demethylase |
| Kleefstra syndrome (KS) | 610253 | Severe mental retardation, hypotonia, epileptic seizures, flat face with hypertelorism, synophrys, anteverted nares, everted lower lip, carp mouth with macroglossia, and heart defects | KMT2B, KMT2C EHMT1 | H3K4 H3K4 H3K9 | Methyltransferase Methyltransferase Methyltransferase |
| Meier-Gorlin syndrome 1 (MGORS1) | 224690 | Severe intrauterine and postnatal growth retardation, microcephaly, bilateral microtia, and aplasia or hypoplasia of the patellae | ORC1 | H4K20 | Recognition |
| Mental retardation, X-linked, syndromic, Claes-Jensen type (MRXSCJ) | 300534 | Severe mental retardation, slowly progressive spastic paraplegia, facial hypotonia, and maxillary hypoplasia | KDM5C | H3K4 | Demethylase |
| Potocki-Shaffer syndrome (PSS) | 601224 | Craniofacial abnormalities, developmental delay, intellectual disability, multiple exostoses, and biparietal foramina | PHF21A | H3K4 | Recognition |
| Schizophrenia (SCZD) | 181500 | Hallucinations and delusions, severely inappropriate emotional responses, disordered thinking and concentration, erratic behavior, as well as social and occupational deterioration | SETD1A | H3K4 | Methyltransferase |
| Siderius X-linked mental retardation syndrome (MRXSSD) | 300263 | Mental retardation, a repaired cleft lip, a long face with broad nasal tip, long hands with long thin fingers, and flat feet with long thin toes | PHF8 | H3K4 H3K9 H4K20 | Recognition Demethylase Demethylase |
| Sotos syndrome 1 (SOTOS1) | 117550 | Excessively rapid growth, acromegalic features, and non-progressive cerebral disorder with mental retardation | NSD1 | H3K36 | Methyltransferase |
| Weaver syndrome (WVS) | 277590 | Pre- and postnatal overgrowth, accelerated osseous maturation, characteristic craniofacial appearance, and developmental delay, broad forehead and face, ocular hypertelorism, prominent wide philtrum, micrognathia, deep horizontal chin groove, and deep-set nails | EZH2 | H3K27 | Methyltransferase |
| Wiedemann-Steiner syndrome (WDSTS) | 605130 | Hypertrichosis cubiti associated with short stature, consistent facial features, including long eyelashes, thick or arched eyebrows with a lateral flare, down slanting and vertically narrow palpebral fissures, mild to moderate intellectual disability, behavioral difficulties, and hypertrichosis on the back | KMT2A | H3K4 | Methyltransferase |
| Wolf-Hirschhorn syndrome (WHS) | 194190 | Pre- and postnatal growth deficiency, developmental disability of variable degree, characteristic craniofacial features, and a seizure disorder | NSD2 | H3K36 | Methyltransferase |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, J.-H.; Lee, J.H.; Lee, I.-S.; Lee, S.B.; Cho, K.S. Histone Lysine Methylation and Neurodevelopmental Disorders. Int. J. Mol. Sci. 2017, 18, 1404. https://doi.org/10.3390/ijms18071404
Kim J-H, Lee JH, Lee I-S, Lee SB, Cho KS. Histone Lysine Methylation and Neurodevelopmental Disorders. International Journal of Molecular Sciences. 2017; 18(7):1404. https://doi.org/10.3390/ijms18071404
Chicago/Turabian StyleKim, Jeong-Hoon, Jang Ho Lee, Im-Soon Lee, Sung Bae Lee, and Kyoung Sang Cho. 2017. "Histone Lysine Methylation and Neurodevelopmental Disorders" International Journal of Molecular Sciences 18, no. 7: 1404. https://doi.org/10.3390/ijms18071404
APA StyleKim, J.-H., Lee, J. H., Lee, I.-S., Lee, S. B., & Cho, K. S. (2017). Histone Lysine Methylation and Neurodevelopmental Disorders. International Journal of Molecular Sciences, 18(7), 1404. https://doi.org/10.3390/ijms18071404





