Non-Coding RNAs in Hodgkin Lymphoma
Abstract
:1. Introduction
2. miRNA Signatures in HL Tissue and Cell Lines
3. miRNAs Impacting Prognosis and Treatment Response in HL
4. miRNAs Involved in the Pathogenesis of HL
4.1. miR-155
4.2. miR-9 and let-7a
4.3. miR-17/106b Family
4.4. miR-21 and miR-30d
4.5. miR-135a
4.6. miR-96, miR-182 and miR-183
5. miRNAs Regulated by Methylation in HL
6. Genomic Changes Affecting miRNA Expression in HL
7. lncRNAs in HL
8. PiwiRNAs in HL
9. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| HL | Hodgkin lymphoma |
| HRS | Hodgkin and Reed-Sternberg cells |
| GC | Germinal center |
| EBV | Epstein Barr virus |
| cHL | Classical Hodgkin lymphoma |
| NS | Nodular sclerosis |
| MC | Mixed cellularity |
| ncRNAs | Non-coding RNAs |
| lncRNAs | Long non-coding RNAs |
| miRNAs | microRNAs |
| XPO5 | Exportin-5 |
| RLNs | Reactive lymph nodes |
| FFPE | Formalin-fixed paraffin-embedded |
| DFS | Disease-free survival |
| OS | Overall survival |
| IPS | International Prognostic Score |
| CR | Complete remission |
| MSP | Methylation specific PCR |
| CNVs | Copy number variations |
| SNPs | Single nucleotide polymorhisms |
| piRNAs | piwiRNAs |
References
- Hodgkin, T. On some morbid appearances of the absorbent glands and spleen. Med. Chir. Trans. 1832, 17, 68–114. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2016. CA cancer J. Clin. 2016, 66, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, R.; Stanelle, J.; Hansmann, M.-L.; Küppers, R. Pathogenesis of classical and lymphocyte-predominant hodgkin lymphoma. Annu. Rev. Pathol. Mech. Dis. 2009, 4, 151–174. [Google Scholar] [CrossRef] [PubMed]
- Campo, E.; Swerdlow, S.H.; Harris, N.L.; Pileri, S.; Stein, H.; Jaffe, E.S. The 2008 who classification of lymphoid neoplasms and beyond: Evolving concepts and practical applications. Blood 2011, 117, 5019–5032. [Google Scholar] [CrossRef] [PubMed]
- Harris, N.L.; Jaffe, E.S.; Stein, H.; Banks, P.M.; Chan, J.K.; Cleary, M.L.; Delsol, G.; De Wolf-Peeters, C.; Falini, B.; Gatter, K.C. A revised european-american classification of lymphoid neoplasms. Blood 1994, 84, 1361–1392. [Google Scholar] [PubMed]
- Basso, K.; Dalla-Favera, R. Germinal centres and B cell lymphomagenesis. Nat. Rev. Immunol. 2015, 15, 172–184. [Google Scholar] [CrossRef] [PubMed]
- Kanzler, H.; Küppers, R.; Hansmann, M.-L.; Rajewsky, K. Hodgkin and reed-sternberg cells in Hodgkin’s disease represent the outgrowth of a dominant tumor clone derived from (crippled) germinal center B cells. J. Exp. Med. 1996, 184, 1495–1505. [Google Scholar] [CrossRef] [PubMed]
- Pileri, S.; Ascani, S.; Leoncini, L.; Sabattini, E.; Zinzani, P.; Piccaluga, P.; Pileri, A.; Giunti, M.; Falini, B.; Bolis, G. Hodgkin’s lymphoma: The pathologist’s viewpoint. J. Clin. Pathol. 2002, 55, 162–176. [Google Scholar] [CrossRef] [PubMed]
- Vockerodt, M.; Cader, F.Z.; Shannon-Lowe, C.; Murray, P. Epstein-barr virus and the origin of hodgkin lymphoma. Cell Differ. 2014, 57, 58. [Google Scholar] [CrossRef] [PubMed]
- Esteller, M. Non-coding RNAs in human disease. Nat. Rev. Genet. 2011, 12, 861–874. [Google Scholar] [CrossRef] [PubMed]
- Nana-Sinkam, S.P.; Croce, C.M. Non-coding RNAs in cancer initiation and progression and as novel biomarkers. Mol. Oncol. 2011, 5, 483–491. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Wightman, B.; Ha, I.; Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 1993, 75, 855–862. [Google Scholar] [CrossRef]
- Lelandais-Brière, C.; Sorin, C.; Declerck, M.; Benslimane, A.; Crespi, M.; Hartmann, C. Small RNA diversity in plants and its impact in development. Curr. Genom. 2010, 11, 14–23. [Google Scholar] [CrossRef] [PubMed]
- Pfeffer, S.; Zavolan, M.; Grässer, F.A.; Chien, M.; Russo, J.J.; Ju, J.; John, B.; Enright, A.J.; Marks, D.; Sander, C. Identification of virus-encoded microRNAs. Science 2004, 304, 734–736. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Lim, L.P.; Glasner, M.E.; Yekta, S.; Burge, C.B.; Bartel, D.P. Vertebrate microRNA genes. Science 2003, 299, 1540. [Google Scholar] [CrossRef] [PubMed]
- Pillai, R.S. MicroRNA function: Multiple mechanisms for a tiny RNA? RNA 2005, 11, 1753–1761. [Google Scholar] [CrossRef] [PubMed]
- Kluiver, J.; Poppema, S.; de Jong, D.; Blokzijl, T.; Harms, G.; Jacobs, S.; Kroesen, B.J.; van den Berg, A. Bic and miR-155 are highly expressed in hodgkin, primary mediastinal and diffuse large B cell lymphomas. J. Pathol. 2005, 207, 243–249. [Google Scholar] [CrossRef] [PubMed]
- Van den Berg, A.; Kroesen, B.J.; Kooistra, K.; de Jong, D.; Briggs, J.; Blokzijl, T.; Jacobs, S.; Kluiver, J.; Diepstra, A.; Maggio, E. High expression of B-cell receptor inducible gene bic in all subtypes of hodgkin lymphoma. Genes Chromosomes Cancer 2003, 37, 20–28. [Google Scholar] [CrossRef] [PubMed]
- Eis, P.S.; Tam, W.; Sun, L.; Chadburn, A.; Li, Z.; Gomez, M.F.; Lund, E.; Dahlberg, J.E. Accumulation of mir-155 and BIC RNA in human B cell lymphomas. Proc. Natl. Acad. Sci. USA 2005, 102, 3627–3632. [Google Scholar] [CrossRef] [PubMed]
- Navarro, A.; Gaya, A.; Martinez, A.; Urbano-Ispizua, A.; Pons, A.; Balagué, O.; Gel, B.; Abrisqueta, P.; Lopez-Guillermo, A.; Artells, R. MicroRNA expression profiling in classic hodgkin lymphoma. Blood 2008, 111, 2825–2832. [Google Scholar] [CrossRef] [PubMed]
- Jones, K.; Nourse, J.P.; Keane, C.; Bhatnagar, A.; Gandhi, M.K. Plasma microRNA are disease response biomarkers in classical hodgkin lymphoma. Clin. Cancer Res. 2013. [CrossRef] [PubMed]
- Van Vlierberghe, P.; de Weer, A.; Mestdagh, P.; Feys, T.; de Preter, K.; de Paepe, P.; Lambein, K.; Vandesompele, J.; van Roy, N.; Verhasselt, B. Comparison of miRNA profiles of microdissected hodgkin/reed-sternberg cells and hodgkin cell lines versus cd77+b-cells reveals a distinct subset of differentially expressed miRNAs. Br. J Haematol. 2009, 147, 686–690. [Google Scholar] [CrossRef] [PubMed]
- Gibcus, J.H.; Tan, L.P.; Harms, G.; Schakel, R.N.; de Jong, D.; Blokzijl, T.; Möller, P.; Poppema, S.; Kroesen, B.-J.; van den Berg, A. Hodgkin lymphoma cell lines are characterized by a specific miRNA expression profile. Neoplasia 2009, 11, 167IN166–176IN169. [Google Scholar] [CrossRef]
- Sánchez-Espiridión, B.; Martín-Moreno, A.M.; Montalbán, C.; Figueroa, V.; Vega, F.; Younes, A.; Medeiros, L.J.; Alvés, F.J.; Canales, M.; Estévez, M. MICRORNA signatures and treatment response in patients with advanced classical hodgkin lymphoma. Br. J. Haematol. 2013, 162, 336–347. [Google Scholar] [CrossRef] [PubMed]
- Paydas, S.; Acikalin, A.; Ergin, M.; Celik, H.; Yavuz, B.; Tanriverdi, K. Micro-RNA (miRNA) profile in hodgkin lymphoma: Association between clinical and pathological variables. Med. Oncol. 2016, 33, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Volinia, S.; Calin, G.A.; Liu, C.-G.; Ambs, S.; Cimmino, A.; Petrocca, F.; Visone, R.; Iorio, M.; Roldo, C.; Ferracin, M. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc. Natl. Acad. Sci. USA 2006, 103, 2257–2261. [Google Scholar] [CrossRef] [PubMed]
- Medina, P.P.; Nolde, M.; Slack, F.J. Oncomir addiction in an in vivo model of microRNA-21-induced pre-b-cell lymphoma. Nature 2010, 467, 86–90. [Google Scholar] [CrossRef] [PubMed]
- Tiacci, E.; Döring, C.; Brune, V.; van Noesel, C.J.; Klapper, W.; Mechtersheimer, G.; Falini, B.; Küppers, R.; Hansmann, M.-L. Analyzing primary hodgkin and reed-sternberg cells to capture the molecular and cellular pathogenesis of classical hodgkin lymphoma. Blood 2012, 120, 4609–4620. [Google Scholar] [CrossRef] [PubMed]
- Navarro, A.; Diaz, T.; Martinez, A.; Gaya, A.; Pons, A.; Gel, B.; Codony, C.; Ferrer, G.; Martinez, C.; Montserrat, E. Regulation of jak2 by miR-135a: Prognostic impact in classic hodgkin lymphoma. Blood 2009, 114, 2945–2951. [Google Scholar] [CrossRef] [PubMed]
- Van Den Berg, A.V.; Vidal, F.; Cruz, A.M.P.; Ribeiro-dos-Santos, Â. MicroRNAs as biomarkers of the response to treatment with abvd scheme in hodgkin lymphoma. J. Leukemia 2015. [CrossRef]
- Van Eijndhoven, M.A.; Zijlstra, J.M.; Groenewegen, N.J.; Drees, E.E.; van Niele, S.; Baglio, S.R.; Koppers-Lalic, D.; van der Voorn, H.; Libregts, S.F.; Wauben, M.H. Plasma vesicle miRNAs for therapy response monitoring in hodgkin lymphoma patients. JCI Insight 2016. [CrossRef] [PubMed]
- Cordeiro, A.; Navarro, A.; Gaya, A.; Díaz-Beyá, M.; Gonzalez-Farré, B.; Castellano, J.J.; Fuster, D.; Martínez, C.; Martínez, A.; Monzó, M. PiwiRNA-651 as marker of treatment response and survival in classical hodgkin lymphoma. Oncotarget 2016, 7, 46002–46013. [Google Scholar] [CrossRef] [PubMed]
- Slezak-Prochazka, I.; Kluiver, J.; de Jong, D.; Smigielska-Czepiel, K.; Kortman, G.; Winkle, M.; Rutgers, B.; Koerts, J.; Visser, L.; Diepstra, A. Inhibition of the miR-155 target niam phenocopies the growth promoting effect of miR-155 in B-cell lymphoma. Oncotarget 2016, 7, 2391–2400. [Google Scholar] [PubMed]
- Costinean, S.; Sandhu, S.K.; Pedersen, I.M.; Tili, E.; Trotta, R.; Perrotti, D.; Ciarlariello, D.; Neviani, P.; Harb, J.; Kauffman, L.R. Src homology 2 domain–containing inositol-5-phosphatase and CCAAT enhancer-binding protein β are targeted by mir-155 in B cells of eμ-miR-155 transgenic mice. Blood 2009, 114, 1374–1382. [Google Scholar] [CrossRef] [PubMed]
- Costinean, S.; Zanesi, N.; Pekarsky, Y.; Tili, E.; Volinia, S.; Heerema, N.; Croce, C.M. Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in eμ-miR155 transgenic mice. Proc. Natl. Acad. Sci. USA 2006, 103, 7024–7029. [Google Scholar] [CrossRef] [PubMed]
- Babar, I.A.; Cheng, C.J.; Booth, C.J.; Liang, X.; Weidhaas, J.B.; Saltzman, W.M.; Slack, F.J. Nanoparticle-based therapy in an in vivo microRNA-155 (miR-155)-dependent mouse model of lymphoma. Proc. Natl. Acad. Sci. USA 2012, 109, E1695–E1704. [Google Scholar] [CrossRef] [PubMed]
- Nie, K.; Gomez, M.; Landgraf, P.; Garcia, J.-F.; Liu, Y.; Tan, L.H.; Chadburn, A.; Tuschl, T.; Knowles, D.M.; Tam, W. MicroRNA-mediated down-regulation of prdm1/blimp-1 in hodgkin/reed-sternberg cells: A potential pathogenetic lesion in hodgkin lymphomas. Am. J. Pathol. 2008, 173, 242–252. [Google Scholar] [CrossRef] [PubMed]
- Shaffer, A.; Lin, K.-I.; Kuo, T.C.; Yu, X.; Hurt, E.M.; Rosenwald, A.; Giltnane, J.M.; Yang, L.; Zhao, H.; Calame, K. Blimp-1 orchestrates plasma cell differentiation by extinguishing the mature B cell gene expression program. Immunity 2002, 17, 51–62. [Google Scholar] [CrossRef]
- Huang, X.; Zhou, X.; Wang, Z.; Li, F.; Liu, F.; Zhong, L.; Li, X.; Han, X.; Wu, Z.; Chen, S. Cd99 triggers upregulation of miR-9-modulated prdm1/blimp1 in hodgkin/reed–sternberg cells and induces redifferentiation. Int. J. Cancer 2012, 131, E382–E394. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Choi, E.Y.; Shin, Y.K.; Kim, T.J.; Chung, D.H.; Chang, S.I.; Kim, N.K.; Park, S.H. Generation of cells with Hodgkin’s and reed-sternberg phenotype through downregulation of cd99 (mic2). Blood 1998, 92, 4287–4295. [Google Scholar] [PubMed]
- Kim, S.H.; Shin, Y.K.; Lee, I.-S.; Bae, Y.M.; Sohn, H.W.; Suh, Y.H.; Ree, H.J.; Rowe, M.; Park, S.H. Viral latent membrane protein 1 (lmp-1)–induced cd99 down-regulation in b cells leads to the generation of cells with Hodgkin’s and reed-sternberg phenotype. Blood 2000, 95, 294–300. [Google Scholar] [PubMed]
- Leucci, E.; Zriwil, A.; Gregersen, L.; Jensen, K.; Obad, S.; Bellan, C.; Leoncini, L.; Kauppinen, S.; Lund, A. Inhibition of miR-9 de-represses HuR and DICER1 and impairs Hodgkin lymphoma tumour outgrowth in vivo. Oncogene 2012, 31, 5081–5089. [Google Scholar] [CrossRef] [PubMed]
- Kuhlen, M.; Hönscheid, A.; Schemme, J.; Merz, H.; Mauz-Körholz, C.; Borkhardt, A.; Troeger, A. Hodgkin lymphoma as a novel presentation of familial DICER1 syndrome. Eur. J. Pediatr. 2016, 175, 593–597. [Google Scholar] [CrossRef] [PubMed]
- Ivanovska, I.; Ball, A.S.; Diaz, R.L.; Magnus, J.F.; Kibukawa, M.; Schelter, J.M.; Kobayashi, S.V.; Lim, L.; Burchard, J.; Jackson, A.L. MicroRNAs in the miR-106b family regulate p21/CDKN1A and promote cell cycle progression. Mol. Cell. Biol. 2008, 28, 2167–2174. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.P.; Seinen, E.; Duns, G.; de Jong, D.; Sibon, O.C.; Poppema, S.; Kroesen, B.-J.; Kok, K.; van den Berg, A. A high throughput experimental approach to identify miRNA targets in human cells. Nucleic Acids Res. 2009, 37, e137. [Google Scholar] [CrossRef] [PubMed]
- Gibcus, J.H.; Kroesen, B.J.; Koster, R.; Halsema, N.; de Jong, D.; de Jong, S.; Poppema, S.; Kluiver, J.; Diepstra, A.; van den Berg, A. MiR-17/106b seed family regulates p21 in Hodgkin’s lymphoma. J. Pathol. 2011, 225, 609–617. [Google Scholar] [CrossRef] [PubMed]
- Navarro, A.; Gaya, A.; Cordeiro, A.; Gonzalez-Farre, B.; Díaz-Beyá, M.; Fuster, D.; Martinez, C.; Esteve, J.; Martinez, A.; Monzó, M. Prognostic significance of a 4-microRNA signature targeting jak2 in classical hodgkin lymphoma. Blood 2013, 122, 629. [Google Scholar]
- Xie, L.; Ushmorov, A.; Leithäuser, F.; Guan, H.; Steidl, C.; Färbinger, J.; Pelzer, C.; Vogel, M.J.; Maier, H.J.; Gascoyne, R.D. Foxo1 is a tumor suppressor in classical hodgkin lymphoma. Blood 2012, 119, 3503–3511. [Google Scholar] [CrossRef] [PubMed]
- Vogel, M.J.; Xie, L.; Guan, H.; Tooze, R.M.; Maier, T.; Kostezka, U.; Maier, H.J.; Holzmann, K.; Chan, F.C.; Steidl, C. Foxo1 repression contributes to block of plasma cell differentiation in classical hodgkin lymphoma. Blood 2014, 124, 3118–3129. [Google Scholar] [CrossRef] [PubMed]
- Ben Dhiab, M.; Ziadi, S.; Louhichi, T.; Ben Gacem, R.; Ksiaa, F.; Trimeche, M. Investigation of miR9-1, miR9-2 and miR9-3 methylation in hodgkin lymphoma. Pathobiology 2015, 82, 195–202. [Google Scholar] [CrossRef] [PubMed]
- Navarro, A.; Díaz, T.; Cordeiro, A.; Beyá, M.D.; Ferrer, G.; Fuster, D.; Martinez, A.; Monzó, M. Epigenetic regulation of microRNA expression in hodgkin lymphoma. Leuk. Lymphoma 2015, 56, 2683–2689. [Google Scholar] [CrossRef] [PubMed]
- Lujambio, A.; Ropero, S.; Ballestar, E.; Fraga, M.F.; Cerrato, C.; Setién, F.; Casado, S.; Suarez-Gauthier, A.; Sanchez-Cespedes, M.; Gitt, A. Genetic unmasking of an epigenetically silenced microRNA in human cancer cells. Cancer Res. 2007, 67, 1424–1429. [Google Scholar] [CrossRef] [PubMed]
- Schouten, H.C.; Sanger, W.G.; Duggan, M.; Weisenburger, D.D.; MacLennan, K.A.; Armitage, J. Chromosomal abnormalities in Hodgkin’s disease. Blood 1989, 73, 2149–2154. [Google Scholar] [PubMed]
- Hartmann, S.; Martin-Subero, J.I.; Gesk, S.; Hüsken, J.; Giefing, M.; Nagel, I.; Riemke, J.; Chott, A.; Klapper, W.; Parrens, M. Detection of genomic imbalances in microdissected hodgkin and reed-sternberg cells of classical Hodgkin’s lymphoma by array-based comparative genomic hybridization. Haematologica 2008, 93, 1318–1326. [Google Scholar] [CrossRef] [PubMed]
- Reichel, J.; Chadburn, A.; Rubinstein, P.G.; Giulino-Roth, L.; Tam, W.; Liu, Y.; Gaiolla, R.; Eng, K.; Brody, J.; Inghirami, G. Flow sorting and exome sequencing reveal the oncogenome of primary hodgkin and reed-sternberg cells. Blood 2015, 125, 1061–1072. [Google Scholar] [CrossRef] [PubMed]
- Hudnall, S.D.; Meng, H.; Lozovatsky, L.; Li, P.; Strout, M.; Kleinstein, S.H. Recurrent genetic defects in classical hodgkin lymphoma cell lines. Leuk. Lymphoma 2016, 57, 2890–2900. [Google Scholar] [CrossRef] [PubMed]
- Mishra, P.J.; Mishra, P.J.; Banerjee, D.; Bertino, J.R. Mirsnps or miR-polymorphisms, new players in microRNA mediated regulation of the cell: Introducing microRNA pharmacogenomics. Cell Cycle 2008, 7, 853–858. [Google Scholar] [CrossRef] [PubMed]
- Navarro, A.; Munoz, C.; Gaya, A.; Díaz-Beyá, M.; Gel, B.; Tejero, R.; Díaz, T.; Martinez, A.; Monzo, M. MiR-SNPs as markers of toxicity and clinical outcome in hodgkin lymphoma patients. PLoS ONE 2013, 8, e64716. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef] [PubMed]
- Díaz-Beyá, M.; Brunet, S.; Nomdedéu, J.; Pratcorona, M.; Cordeiro, A.; Gallardo, D.; Escoda, L.; Tormo, M.; Heras, I.; Ribera, J.M. The lincRNA hotairm1, located in the hoxa genomic region, is expressed in acute myeloid leukemia, impacts prognosis in patients in the intermediate-risk cytogenetic category, and is associated with a distinctive microRNA signature. Oncotarget 2015, 6, 31613. [Google Scholar] [PubMed]
- Leucci, E.; Patella, F.; Waage, J.; Holmstrøm, K.; Lindow, M.; Porse, B.; Kauppinen, S.; Lund, A.H. MicroRNA-9 targets the long non-coding RNA MALAT1 for degradation in the nucleus. Sci. Rep. 2013, 3, 2535. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Huang, C.; Meng, X.; Li, J. Long noncoding RNA MALAT1: Insights into its biogenesis and implications in human disease. Curr. Pharm. Des. 2015, 21, 5017–5028. [Google Scholar] [CrossRef] [PubMed]
- Tayari, M.M.; Winkle, M.; Kortman, G.; Sietzema, J.; de Jong, D.; Terpstra, M.; Mestdagh, P.; Kroese, F.G.; Visser, L.; Diepstra, A. Long noncoding RNA expression profiling in normal B-cell subsets and hodgkin lymphoma reveals hodgkin and reed-sternberg cell-specific long noncoding RNAs. Am. J. Pathol. 2016, 186, 2462–2472. [Google Scholar] [CrossRef] [PubMed]
- Iwasaki, Y.W.; Siomi, M.C.; Siomi, H. Piwi-interacting RNA: Its biogenesis and functions. Annu. Rev. Biochem. 2015, 84, 405–433. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.; Guo, J.-M.; Xiao, B.-X.; Miao, Y.; Jiang, Z.; Zhou, H.; Li, Q.-N. PiRNA, the new non-coding RNA, is aberrantly expressed in human cancer cells. Clin. Chim. Acta 2011, 412, 1621–1625. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.-L.; Hu, H.; Xue, X.; Shen, S.; Gao, E.-L.; Guo, G.; Shen, X.; Zhang, X. Altered expression of piRNAs and their relation with clinicopathologic features of breast cancer. Clin. Transl. Oncol. 2013, 15, 563–568. [Google Scholar] [CrossRef] [PubMed]
- Law, P.T.-Y.; Qin, H.; Ching, A.K.-K.; Lai, K.P.; Co, N.N.; He, M.; Lung, R.W.-M.; Chan, A.W.-H.; Chan, T.-F.; Wong, N. Deep sequencing of small RNA transcriptome reveals novel non-coding RNAs in hepatocellular carcinoma. J. Hepatol. 2013, 58, 1165–1173. [Google Scholar] [CrossRef] [PubMed]
- Yan, H.; Wu, Q.; Sun, C.; Ai, L.; Deng, J.; Zhang, L.; Chen, L.; Chu, Z.; Tang, B.; Wang, K. PiRNA-823 contributes to tumorigenesis by regulating de novo DNA methylation and angiogenesis in multiple myeloma. Leukemia 2015, 29, 196–206. [Google Scholar] [CrossRef] [PubMed]
- Cui, L.; Lou, Y.; Zhang, X.; Zhou, H.; Deng, H.; Song, H.; Yu, X.; Xiao, B.; Wang, W.; Guo, J. Detection of circulating tumor cells in peripheral blood from patients with gastric cancer using piRNAs as markers. Clin. Biochem. 2011, 44, 1050–1057. [Google Scholar] [CrossRef] [PubMed]
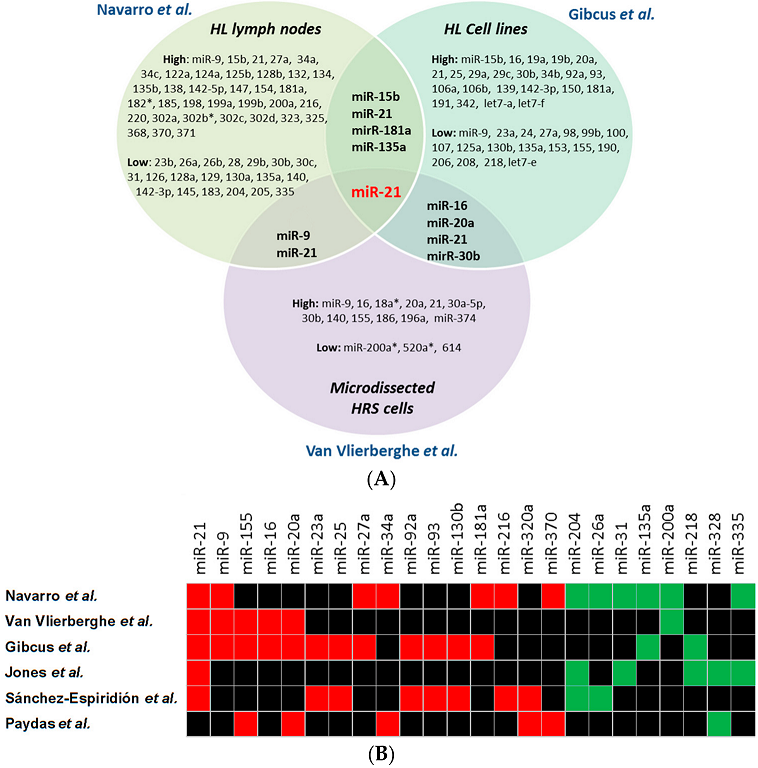

| miRNAs | Navarro | Van Vlierberghe | Gibcus | Jones | Sánchez-Espiridion | Paydas |
|---|---|---|---|---|---|---|
| miR-9 | ↑ cHL LN-RLN | ↑ HRS-CD77+ | ↑ HL cl-BL cl $ | |||
| miR-15b | ↓ cHL LN-RLN | ↑ HL cl-BL/LCL cl | ||||
| miR-16 | ↑ HRS-CD77+ | ↑ HL cl-BL/LCL cl | ||||
| miR-20a | ↑ HRS-CD77+ | ↑ HL cl-LCL/CLL cl | ↑ cHL LN-RLN | |||
| miR-21 | ↑ cHL LN-RLN | ↑ HRS-CD77+ | ↑ HL cl-BL cl | ↑ cHL-RLN | ↑ cHL LN-HL cl | |
| miR-23a | ↑ HL cl-DLBCL/BL cl | ↑ cHL LN-HL cl | ||||
| miR-25 | ↑ HL cl-BL/LCL cl | ↑ cHL LN-HL cl | ||||
| miR-26a | ↓ cHL LN-RLN | ↓ cHL LN-HL cl | ||||
| miR-27a | ↑ cHL LN-RLN | ↑ HL cl-DLBCL/BL cl | ||||
| miR-29a | ↑ HL cl-BL/CLL cl | ↓ cHL-RLN | ||||
| miR-30b | ↓ cHL LN-RLN | ↑ HRS-CD77+ | ↑ HL cl-BL cl | |||
| miR-31 | ↓ cHL LN-RLN | ↓ cHL-RLN | ||||
| miR-34a | ↑ cHL LN-RLN | ↑ cHL LN-RLN | ||||
| miR-92a | ↑ HL-BL/LCL/CLL cl | ↑ cHL LN-HL cl | ||||
| miR-93 | ↑ HL cl-BL/CLL cl | ↑ cHL LN-HL cl | ||||
| miR-140 | ↓ cHL LN-RLN | ↑ HRS-CD77+ | ||||
| miR-125a | ↓ cHL LN-RLN | ↑ HRS-CD77+ | ||||
| miR-128a | ↑ cHL LN-RLN | ↓ cHL LN-RLN | ||||
| miR-130b | ↑ HL cl-BL cl $ | ↑ cHL LN-HL cl | ||||
| miR-132 | ↑ cHL LN-RLN | ↓ cHL-RLN | ↓ cHL LN-HL cl | |||
| miR-135a | ↓ cHL LN-RLN | ↓ HL cl-PMBL cl | ||||
| miR-139 | ↑ HL cl-LCL cl | ↓ cHL-RLN | ||||
| miR-142-3p | ↓ cHL LN-RLN | ↑ HL cl-BL/CLL cl | ||||
| miR-145 | ↑ cHL LN-RLN | ↓ cHL LN-RLN | ||||
| miR-155 | ↑ HRS-CD77+ | #Non-sig. | #Non-sig. | ↑ cHL LN-RLN | ||
| miR-181a | ↑ cHL LN-RLN | |||||
| miR-196a | ↑ HRS-CD77+ | ↓ cHL-RLN | ||||
| miR-200a | ↓ cHL LN-RLN | ↓ HRS-CD77+ | ||||
| miR-204 | ↓ cHL LN-RLN | ↓ cHL-RLN | ↓ cHL LN-HL cl | |||
| miR-216 | ↑ cHL LN-RLN | ↑ cHL LN-HL cl | ||||
| miR-218 | ↓ HL cl-BL cl | ↓ cHL-RLN | ||||
| miR-320a | ↑ cHL LN-HL cl | ↑ cHL LN-RLN | ||||
| miR-328 | ↓ cHL-RLN | ↓ cHL LN-RLN | ||||
| miR-335 | ↓ cHL LN-RLN | ↓ cHL-RLN | ||||
| miR-370 | ↑ cHL LN-RLN | ↑ cHL LN-RLN |
| ncRNAs | Levels Associated with Bad Prognosis | Samples | Patients | Endpoint | Multivariate Analysis Performed | Ref. |
|---|---|---|---|---|---|---|
| miR-135a | Low | Lymph nodes | 89 cHL | DFS | Yes | [32] |
| miR-21 miR-92b-5p miR-30d miR-30e | High Low High Low | Lymph nodes | Discovery set with 29 and training set with 168 HIV-advanced cHL | Failure-free survival | Yes | [27] |
| miR-494 miR-21 miR-1973 | High High High | Plasma | 42 HIV-,HCV- and HBV-cHL | Treatment response | No | [24] |
| miR-9 miR-21 miR-155 | High High Low | Peripheral blood | 4 before treatment and 7 after treatment cHL | Treatment response | No | [33] |
| miR-21-5p miR-127-3-p miR-155-5p let-7a-5p | High High High High | Plasma (extracellular vesicles and protein bound associated miRNAs) | 20 cHL before treatment (13 primary and 7 relapsed) and 7 after treatment | Complete metabolic response (FDG-PET) | No | [34] |
| piR-651 | Low | Lymph nodes | 94 HIV-cHL | Treatment response, DFS and OS | Yes | [35] |
| piR-651 | Low | Serum | 11 cHL before treatment and 9 after treatment | Treatment response | No | [35] |
| miRNA | Location | HL Cell Lines | HL Patients | ||
|---|---|---|---|---|---|
| Gain+ | Loss+ | Gain+ | Loss+ | ||
| miR-9 | chr1: 156420341-156420429 [−] chr5: 88666853-88666939 [−] chr15: 89368017-89368106 [+] | 1/2 − 2/5 | − − 1/6 | 1/10 − − | − − 1/10 |
| miR-15 | chr3: 160404588-160404685 [+] | − | − | − | 1/10 |
| miR-16 | chr13: 50048973-50049061 [−] chr3: 160404745-160404825 [+] | − − | 2/2 0/2 | − − | 3/10 1/10 |
| miR-20a | chr13: 91351065-91351135 [+] | 2/2 | 2/10 | ||
| miR-21 | chr17: 59841266-59841337 [+] | 4/5 | 1/5 | − | − |
| miR-23a | chr19: 13836587-13836659 [−] | 0/2 | − | 8/22 | − |
| miR-25 | chr7: 100093560-100093643 [−] | − | − | − | − |
| miR-26a | chr3: 37969404-37969480 [+] chr12: 57824609-57824692 [−] | − 1/2 | 1/2 1/2 | − 1/10 | 0/10 0/10 |
| miR-27a | chr19: 13836440-13836517 [−] | 0/2 | − | 8/22 | 0/12 |
| miR-29a | chr7: 130876747-130876810 [−] | 2/5 | 0/6 | − | 2/10 |
| miR-30b | chr8: 134800520-134800607 [−] | 3/6 | 0/6 | 0/10 | 3/10 |
| miR-31 | chr9: 21512115-21512185 [−] | 0/2 | 1/2 | 3/10 | 1/10 |
| miR-34a | chr1: 9151668-9151777 [−] | 4/5 | 0/6 | − | 2/10 |
| miR-92a | chr13: 91351314-91351391 [+] chrX: 134169538-134169612 [−] | − 2/2 | 2/2 − | − 6/22 | 2/10 − |
| miR-93 | chr7: 100093768-100093847 [−] | − | − | − | − |
| miR-125a | chr19: 51693254-51693339 [+] | − | − | − | − |
| miR-128a | chr2: 135665397-135665478 [+] | 0/2 | − | 1/10 | − |
| miR-130b | chr22: 21653304-21653385 [+] | 1/5 | 1/6 | 4/12 | 4/22 |
| miR-132 | chr17: 2049908-2050008 [−] | 2/5 | 1/6 | 4/12 | 0/10 |
| miR-135a | chr3: 52294219-52294308 [−] chr12: 97563812-97563911 [+] | − 3/6 | 1/2 1/6* | − 1/10 | 0/10 0/10 |
| miR-139 | chr11: 72615063-72615130 [−] | − | 1/2 | 4/12 | 0/10 |
| miR-140 | chr16: 69933081-69933180 [+] | − | 0/2 | − | 1/10 |
| miR-142-3p | chr:17 58331232-58331318 [−] | − | − | − | − |
| miR-145 | chr5: 149430646-149430733 [+] | − | − | − | − |
| miR-155 | chr21: 25573980-25574044 [+] | 3/5 | 0/6 | − | 1/10 |
| miR-181a | chr1: 198859044-198859153 [−] chr9: 124692442-124692551 [+] | 1/2 2/6* | − 0/6 | 0/10 2/10 | − 1/10 |
| miR-196a | chr17: 48632490-48632559 [−] chr12: 53991738-53991847 [+] | 5/5 5/6* | 0/5 0/5 | − 6/22 | − − |
| miR-200a | chr1: 1167863-1167952 [+] | 2/5 | 1/6* | − | 2/10 |
| miR-204 | chr9: 70809975-70810084 [−] | 0/2 | 0/2 | 1/10 | 1/10 |
| miR-216 | chr2: 55988950-55989059 [−] | 6/6* | 0/5 | 3/10 | − |
| miR-218 | chr4: 20528275-20528384 [+] chr5: 168768146-168768255 [−] | − 0/2 | 1/2 − | − 2/10 | 0/10 − |
| miR−320a | chr8: 22244962−22245043 [−] | − | 1/2 | − | 7/22 |
| miR-328 | chr16: 67202321-67202395 [−] | − | 0/2 | − | 1/10 |
| miR-335 | chr7: 130496111-130496204 [−] | − | 0/2 | − | 2/10 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cordeiro, A.; Monzó, M.; Navarro, A. Non-Coding RNAs in Hodgkin Lymphoma. Int. J. Mol. Sci. 2017, 18, 1154. https://doi.org/10.3390/ijms18061154
Cordeiro A, Monzó M, Navarro A. Non-Coding RNAs in Hodgkin Lymphoma. International Journal of Molecular Sciences. 2017; 18(6):1154. https://doi.org/10.3390/ijms18061154
Chicago/Turabian StyleCordeiro, Anna, Mariano Monzó, and Alfons Navarro. 2017. "Non-Coding RNAs in Hodgkin Lymphoma" International Journal of Molecular Sciences 18, no. 6: 1154. https://doi.org/10.3390/ijms18061154
APA StyleCordeiro, A., Monzó, M., & Navarro, A. (2017). Non-Coding RNAs in Hodgkin Lymphoma. International Journal of Molecular Sciences, 18(6), 1154. https://doi.org/10.3390/ijms18061154







