High Concentration of Melatonin Regulates Leaf Development by Suppressing Cell Proliferation and Endoreduplication in Arabidopsis
Abstract
:1. Introduction
2. Results
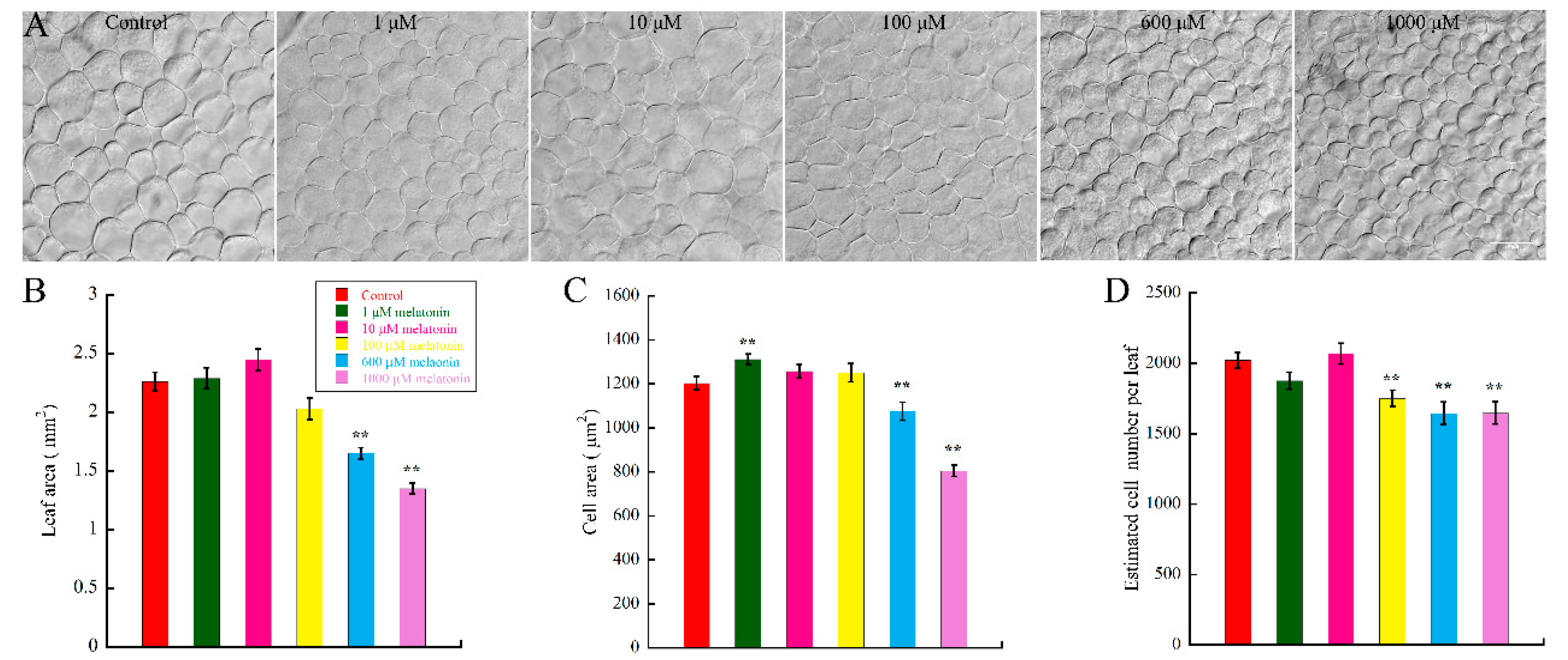
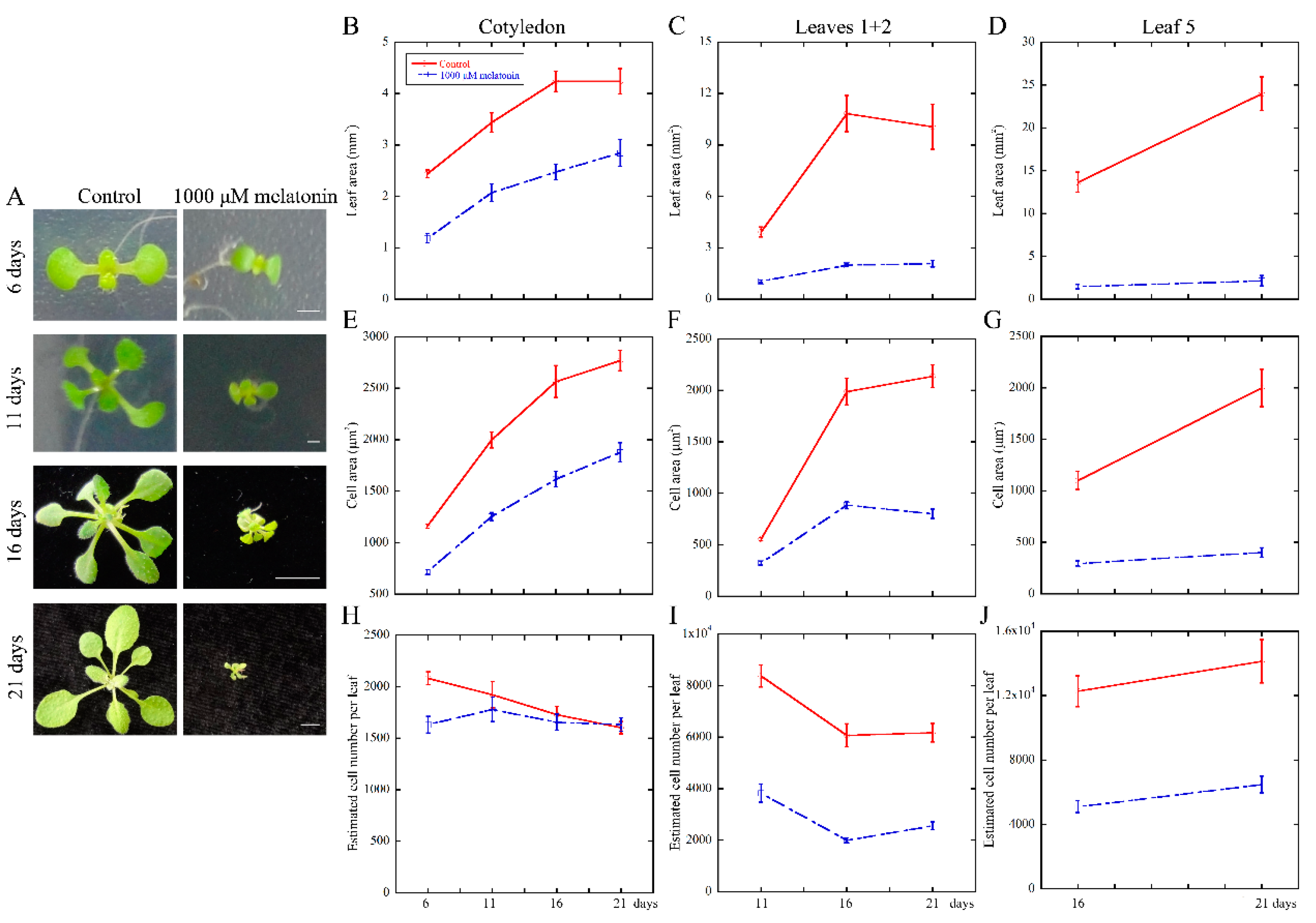
2.1. High Concentration of Melatonin Suppresses the Leaf Growth in Arabidopsis by Reduced Cell Size and Cell Number
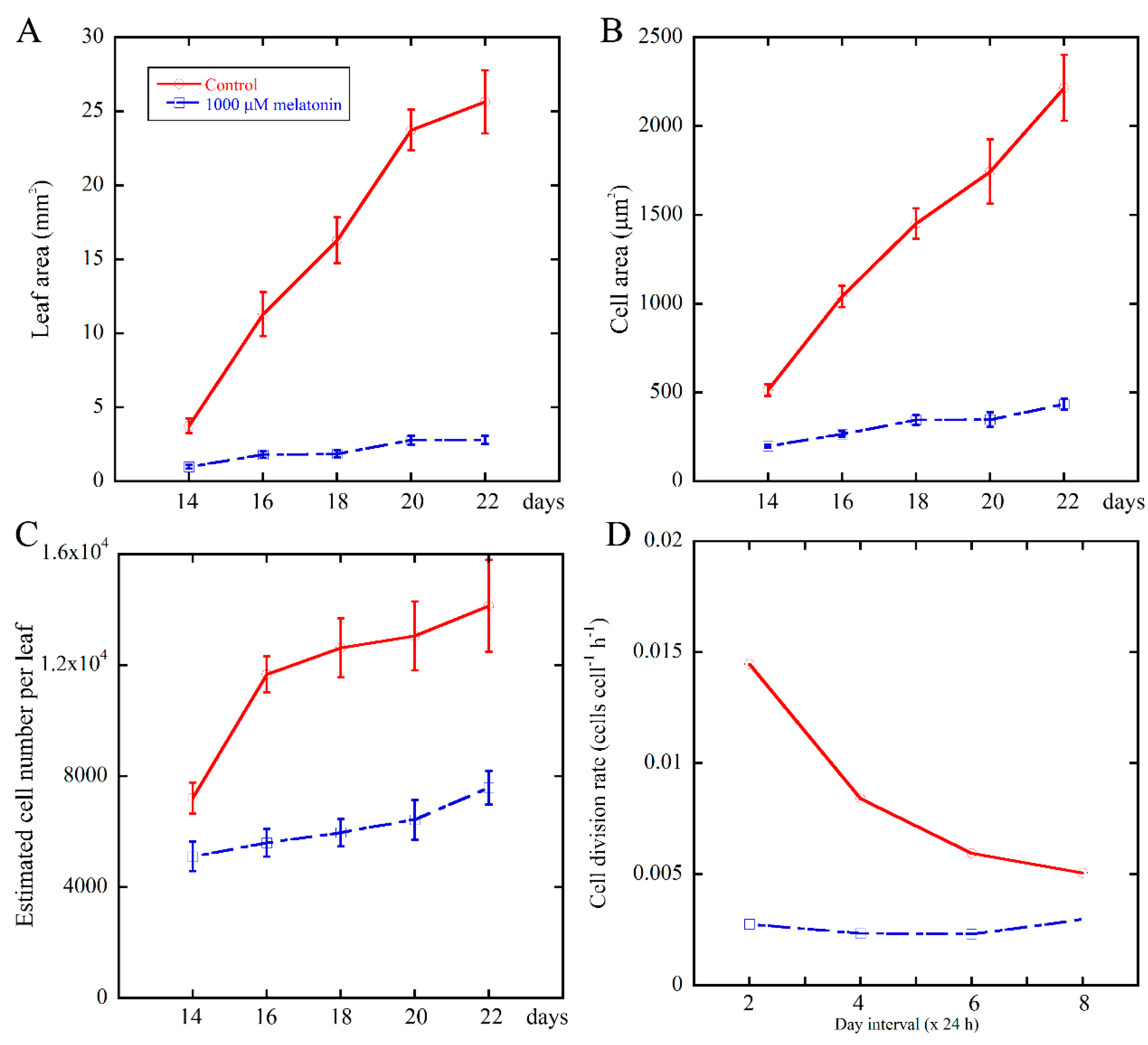
2.2. Melatonin Negatively Affects the Cell Division Rate of Leaves During Leaf Growth
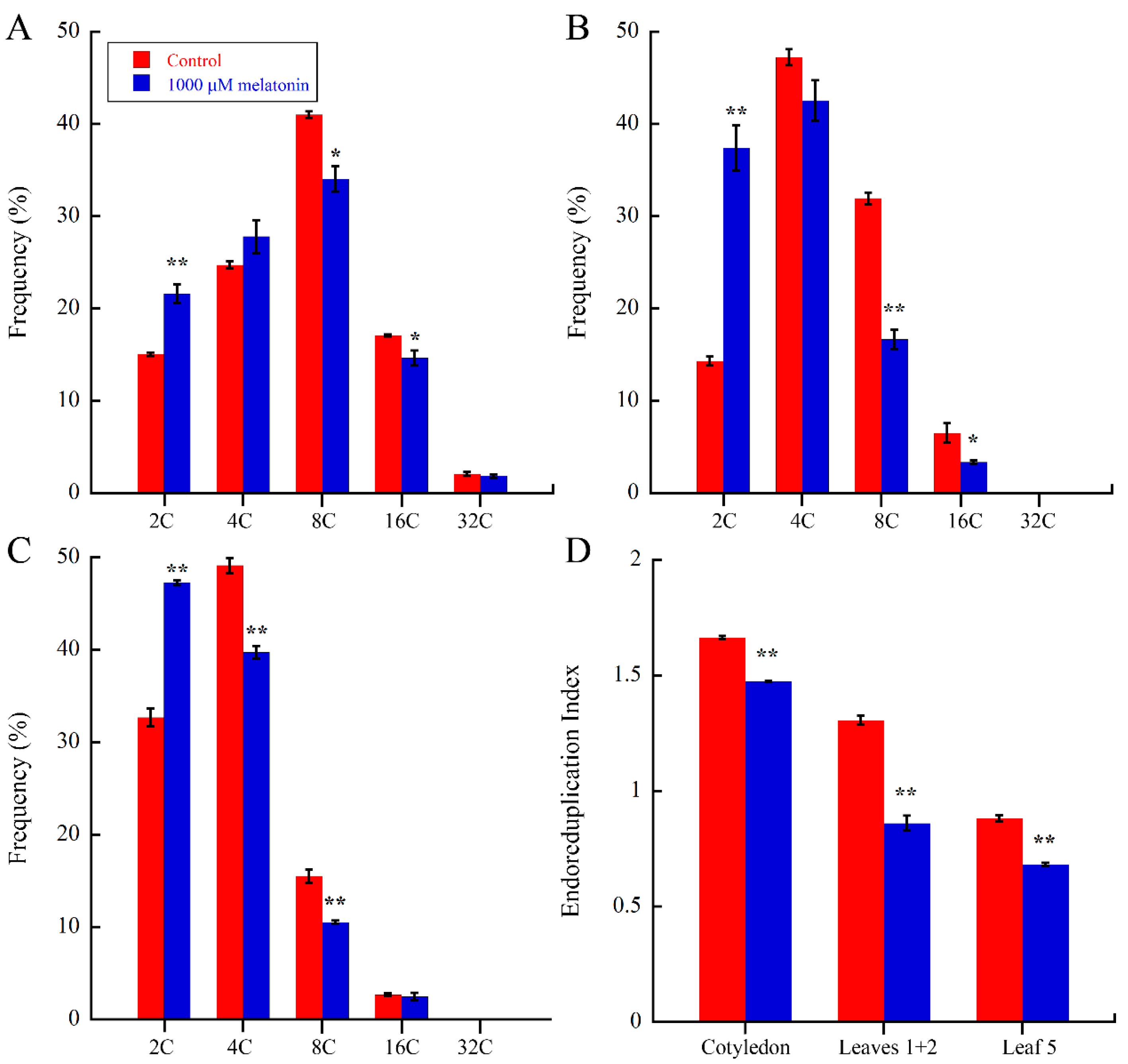
2.3. Melatonin Regulate Endoreduplication during Leaf Development
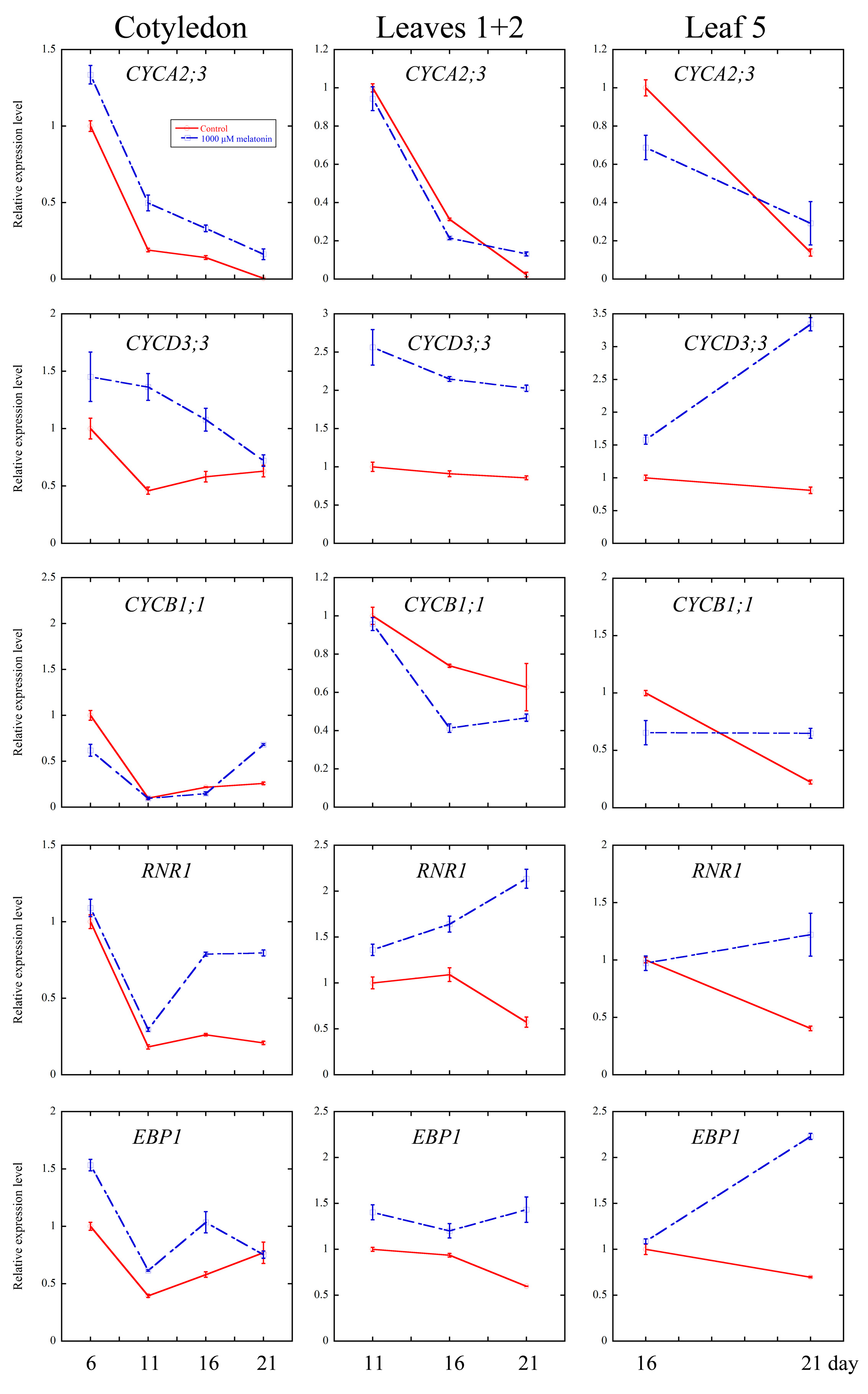
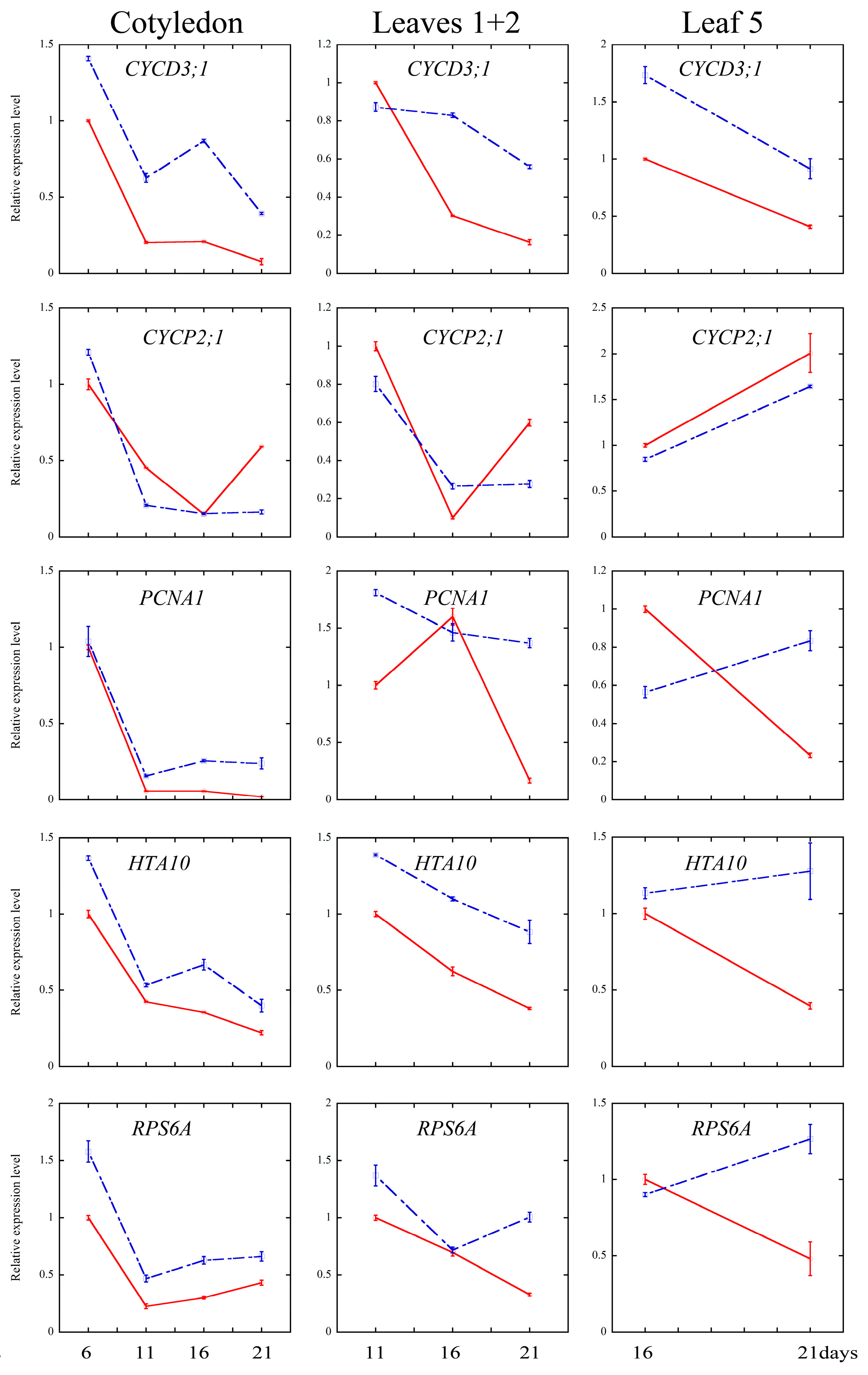
2.4. Melatonin Regulated Several Cell Cycle and Ribosomal Related Genes
3. Discussion
3.1. Melatonin Represses Cell Proliferation and Cell Expansion in Arabidopsis Leaf Growth
3.2. Melatonin Inhibites Endoreduplication during Leaf Growth
3.3. Melatonin Regulates Cell Cycle Progression
4. Experimental Section
4.1. Plant Materials and Growth Conditions
4.2. Leaf Measurements
4.3. Flow Cytometry Experiments and Ploidy Measurement
4.4 Quantitative Real-Time Pcr Analysis
5. Conclusions
Supplementary Materials
Acknowledgements
Author Contributions
Conflicts of Interest
References
- Lerner, A.B.; Case, J.D.; Takahashi, Y.; Lee, T.H.; Mori, W. Isolation of melatonin, a pineal factor that lightens melanocytes. J. Am. Chem. Soc. 1958, 80, 2587. [Google Scholar] [CrossRef]
- Poeggeler, B.; Balzer, I.; Hardeland, R.; Lerchl, A. Pineal hormone melatonin oscillates also in the dinoflagellate Gonyaulax polyedra. Naturwissenschaften 1991, 78, 268–269. [Google Scholar] [CrossRef]
- Dubbels, R.; Reiter, R.J.; Klenke, E.; Goebel, A.; Schnakenberg, E.; Ehlers, C.; Schiwara, H.W.; Schloot, W. Melatonin in edible plants identified by radioimmunoassay and by high performance liquid chromatography-mass spectrometry. J. Pineal Res. 1995, 18, 28–31. [Google Scholar] [CrossRef] [PubMed]
- Hattori, A.; Migitaka, H.; Iigo, M.; Itoh, M.; Yamamoto, K.; Ohtani-Kaneko, R.; Hara, M.; Suzuki, T.; Reiter, R.J. Identification of melatonin in plants and its effects on plasma melatonin levels and binding to melatonin receptors in vertebrates. Biochem. Mol. Biol. Int. 1995, 35, 627–634. [Google Scholar] [PubMed]
- Reiter, R.J.; Tan, D.X.; Rosales-Corral, S.; Manchester, L.C. The universal nature, unequal distribution and antioxidant functions of melatonin and its derivatives. Mini. Rev. Med. Chem. 2013, 13, 373–384. [Google Scholar] [CrossRef] [PubMed]
- Hardeland, R. Melatonin in plants-diversity of levels and multiplicity of functions. Front. Plant Sci. 2016, 7, 198. [Google Scholar] [CrossRef] [PubMed]
- Jan, J.E.; Reiter, R.J.; Wasdell, M.B.; Bax, M. The role of the thalamus in sleep, pineal melatonin production, and circadian rhythm sleep disorders. J. Pineal Res. 2009, 46, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Hardeland, R.; Madrid, J.A.; Tan, D.X.; Reiter, R.J. Melatonin, the circadian multioscillator system and health: The need for detailed analyses of peripheral melatonin signaling. J. Pineal Res. 2012, 52, 139–166. [Google Scholar] [CrossRef] [PubMed]
- Carrillo-Vico, A.; Lardone, P.J.; Álvarez-Sánchez, N.; Rodríguez-Rodríguez, A.; Guerrero, J.M. Melatonin: Buffering the immune system. Int. J. Mol. Sci. 2013, 14, 8638–8683. [Google Scholar] [CrossRef] [PubMed]
- Reiter, R.J.; Tan, D.X.; Galano, A. Melatonin: Exceeding expectations. Physiology 2014, 29, 325–333. [Google Scholar] [CrossRef] [PubMed]
- Kang, K.; Lee, K.; Park, S.; Kim, Y.S.; Back, K. Enhanced production of melatonin by ectopic overexpression of human serotonin N-acetyltransferase plays a role in cold resistance in transgenic rice seedlings. J. Pineal Res. 2010, 49, 176–182. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.; Wang, P.; Li, M.; Ke, X.; Li, C.; Liang, D.; Wu, S.; Ma, X.; Li, C.; Zou, Y.; et al. Exogenous melatonin improves Malus resistance to marssonina apple blotch. J. Pineal Res. 2013, 54, 426–434. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Zhao, B.; Zhang, H.J.; Weeda, S.; Yang, C.; Yang, Z.C.; Ren, S.; Guo, Y.D. Melatonin promotes water-stress tolerance, lateral root formation, and seed germination in cucumber (Cucumis sativus L.). J. Pineal Res. 2013, 54, 15–23. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Chan, Z. The Cysteine2/Histidine2-type transcription factor ZINC FINGER OF ARABIDOPSIS thaliana 6- activated C-REPEAT-BINDING FACTOR pathway is essential for melatonin-mediated freezing stress resistance in Arabidopsis. J. Pineal Res. 2014, 57, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Weeda, S.; Zhang, N.; Zhao, X.; Ndip, G.; Guo, Y.; Buck, G.A.; Fu, C.; Ren, S. Arabidopsis transcriptome analysis reveals key roles of melatonin in plant defense systems. PLoS ONE 2014, 9, e93462. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Chen, Y.; Tan, D.X.; Reiter, R.J.; Chan, Z.; He, C. Melatonin induces nitric oxide and the potential mechanisms relate to innate immunity against bacterial pathogen infection in Arabidopsis. J. Pineal Res. 2015, 59, 102–108. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Jiang, C.; Ye, T.; Tan, D.X.; Reiter, R.J.; Zhang, H.; Liu, R.; Chan, Z. Comparative physiological, metabolomic, and transcriptomic analyses reveal mechanisms of improved abiotic stress resistance in bermudagrass (Cynodon dactylon (L). Pers.) by exogenous melatonin. J. Exp. Bot. 2015, 66, 681–694. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Reiter, R.J.; Tan, D.X.; Chan, Z. INDOLE-3-ACETIC ACID INDUCIBLE 17 positively modulates natural leaf senescence through melatonin-mediated pathway in Arabidopsis. J. Pineal Res. 2015, 58, 26–33. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Tan, D.X.; Reiter, R.J.; Ye, T.; Yang, F.; Chan, Z. Melatonin induces class A1 heat shock factors (HSFA1s) and their possible involvement of thermotolerance in Arabidopsis. J. Pineal Res. 2015, 58, 335–342. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Wang, X.; Tan, D.X.; Reiter, R.J.; Chan, Z. Comparative physiological and proteomic analyses reveal the actions of melatonin in the reduction of oxidative stress in bermudagrass (Cynodon dactylon (L). Pers.). J. Pineal Res. 2015, 59, 120–131. [Google Scholar] [CrossRef] [PubMed]
- Cai, S.Y.; Zhang, Y.; Xu, Y.P.; Qi, Z.Y.; Li, M.Q.; Ahammed, G.J.; Xia, X.J.; Shi, K.; Zhou, Y.H.; Reiter, R.J.; et al. HsfA1a upregulates melatonin biosynthesis to confer cadmium tolerance in tomato plants. J. Pineal Res. 2017, 62, e12387. [Google Scholar] [CrossRef] [PubMed]
- Reiter, R.J.; Tan, D.X.; Zhou, Z.; Cruz, M.H.; Fuentes-Broto, L.; Galano, A. Phytomelatonin: Assisting plants to survive and thrive. Molecules 2015, 20, 7396–7437. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; An, B.; Wei, Y.; Reiter, R.J.; Shi, H.; Luo, H.; He, C. Melatonin regulates root meristem by repressing auxin synthesis and polar auxin transport in Arabidopsis. Front. Plant Sci. 2016, 15, 1882. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Wei, Y.; Wang, Q.; Reiter, R.J.; He, C. Melatonin mediates the stabilization of DELLA proteins to repress the floral transition in Arabidopsis. J. Pineal Res. 2016, 60, 373–379. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Zhang, N.; Wang, J.; Zhang, H.; Li, D.; Shi, J.; Li, R.; Weeda, S.; Zhao, B.; Ren, S.; et al. Melatonin promotes ripening and improves quality of tomato fruit during postharvest life. J. Exp. Bot. 2014, 66, 657–658. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Zhang, N.; Wang, J.; Cao, Y.; Li, X.; Zhang, H.; Zhang, L.; Tan, D.X.; Guo, Y.D. A label-free differential proteomics analysis reveals the effect of melatonin on promoting fruit ripening and anthocyanin accumulation upon postharvest in tomato. J. Pineal Res. 2016, 61, 138–153. [Google Scholar] [CrossRef] [PubMed]
- Wen, D.; Gong, B.; Sun, S.; Liu, S.; Wang, X.; Wei, M.; Yang, F.; Li, Y.; Shi, Q. Promoting roles of melatonin in adventitious root development of Solanum lycopersicum L. by regulating auxin and nitric oxide signaling. Front. Plant Sci. 2016, 7, 718. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Zhang, H.J.; Sun, Q.Q.; Cao, Y.Y.; Li, X.; Zhao, B.; Wu, P.; Guo, Y.D. Proteomic analysis reveals a role of melatonin in promoting cucumber seed germination under high salinity by regulating energy production. Sci. Rep. 2017, 7, 503. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Sun, Y.; Liu, Z.; Sun, Y. Effects of melatonin on seedling growth, mineral nutrition, and nitrogen metabolism in cucumber under nitrate stress. J. Pineal Res. 2017, 62. [Google Scholar] [CrossRef] [PubMed]
- Arnao, M.B.; Hernández-Ruiz, J. Melatonin: Plant growth regulator and/or biostimulator during stress? Trends Plant Sci. 2014, 19, 789–797. [Google Scholar] [CrossRef] [PubMed]
- Arnao, M.B.; Hernández-Ruiz, J. Functions of melatonin plants: A review. J. Pineal Res. 2015, 59, 133–150. [Google Scholar] [CrossRef] [PubMed]
- Paredes, S.D.; Korkmaz, A.; Manchester, L.C.; Tan, D.X.; Reiter, R.J. Phytomelatonin: A review. J. Exp. Bot. 2009, 60, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Tan, D.X.; Hardeland, R.; Manchester, L.C.; Korkmaz, A.; Ma, S.; Rosales-Corral, S.; Reiter, R.J. Functional roles of melatonin in plants and perspectives in nutritional and agricultural science. J. Exp. Bot. 2012, 63, 577–597. [Google Scholar] [CrossRef] [PubMed]
- Arnao, M.B. Phytomelatonin: Discovery, content, and role in plants. Adv. Bot. 2014, 2014, 815769. [Google Scholar] [CrossRef]
- Hardeland, R. Melatonin in plants and other phototrophs—Advances and gaps concerning the diversity of functions. J. Exp. Bot. 2015, 66, 627–646. [Google Scholar] [CrossRef] [PubMed]
- Pelagio-Flores, R.; Muñoz-Parra, E.; Ortiz-Castro, R.; López-Bucio, J. Melatonin regulates Arabidopsis root system architecture likely acting independently of auxin signaling. J. Pineal Res. 2012, 53, 279–288. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Seo, H.; Park, C.; Park, W.J. Examination of the auxin hypothesis of phytomelatonin action in classical auxin assay systems in maize. J. Plant Physiol. 2016, 190, 67–71. [Google Scholar] [CrossRef] [PubMed]
- Tsukaya, H. Organ shape and size: A lesson from studies of leaf morphogenesis. Curr. Opin. Plant Biol. 2003, 6, 57–62. [Google Scholar] [CrossRef] [PubMed]
- Mizukami, Y. A matter of size: Developmental control of organ size in plants. Curr. Opin. Plant Biol. 2001, 4, 533–539. [Google Scholar] [CrossRef]
- Walch-Liu, P.; Neumann, G.; Bangerth, F.; Engels, C. Rapid effects of nitrogen form on leaf morphogenesis in tobacco. J. Exp. Bot. 2000, 51, 227–237. [Google Scholar] [CrossRef] [PubMed]
- Thingnaes, E.; Torre, S.; Ernstsen, A.; Moe, R. Day and night temperature responses in Arabidopsis: Effects on gibberellin and auxin content, cell size, morphology and flowering time. Ann. Bot. 2003, 92, 601–612. [Google Scholar] [CrossRef] [PubMed]
- Kozuka, T.; Horiguchi, G.; Kim, G.T.; Ohgishi, M.; Sakai, T.; Tsukaya, H. The different growth responses of the Arabidopsis thaliana leaf blade and the petiole during shade avoidance are regulated by photoreceptors and sugar. Plant Cell Physiol. 2005, 46, 213–223. [Google Scholar] [CrossRef] [PubMed]
- Smith, L.G.; Hake, S. The initiation and determination of leaves. Plant Cell 1992, 4, 1017–1027. [Google Scholar] [CrossRef] [PubMed]
- Anastasiou, E.; Lenhard, M. Growing up to one’s standard. Curr. Opin. Plant Biol. 2007, 10, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, N.; Vanhaeren, H.; Inzé, D. Leaf size control: Complex coordination of cell division and expansion. Trends Plant Sci. 2012, 17, 332–340. [Google Scholar] [CrossRef] [PubMed]
- Donnelly, P.M.; Bonetta, D.; Tsukaya, H.; Dengler, R.E.; Dengler, N.G. Cell cycling and cell enlargement in developing leaves of Arabidopsis. Dev. Biol. 1999, 215, 407–419. [Google Scholar] [CrossRef] [PubMed]
- Nath, U.; Crawford, B.C.; Carpenter, R.; Coen, E. Genetic control of surface curvature. Science 2003, 299, 1404–1407. [Google Scholar] [CrossRef] [PubMed]
- Tsukaya, H. Controlling size in multicellular organs: Focus on the leaf. PLoS Biol. 2008, 6, e174. [Google Scholar] [CrossRef] [PubMed]
- Beemster, G.T.; Fiorani, F.; Inzé, D. Cell cycle: The key to plant growth control? Trends Plant Sci. 2003, 8, 154–158. [Google Scholar] [CrossRef]
- Horiguchi, G.; Ferjani, A.; Fujikura, U.; Tsukaya, H. Coordination of cell proliferation and cell expansion in the control of leaf size in Arabidopsis thaliana. J. Plant Res. 2006, 119, 37–42. [Google Scholar] [CrossRef] [PubMed]
- Fiorani, F.; Beemster, G.T. Quantitative analyses of cell division in plants. Plant Mol. Boil. 2006, 60, 963–979. [Google Scholar] [CrossRef] [PubMed]
- Beemster, G.T.; de Veylder, L.; Vercruysse, S.; West, G.; Rombaut, D.; van Hummelen, P.; Galichet, A.; Gruissem, W.; Inzé, D.; Vuylesteke, M. Genome-wide analysis of gene expression profiles associated with cell cycle transitions in growing organs of Arabidopsis. Plant Physiol. 2005, 138, 734–743. [Google Scholar] [CrossRef] [PubMed]
- Inzé, D.; de Veylder, L. Cell cycle regulation in plant development. Annu. Rev. Genet. 2006, 40, 77–105. [Google Scholar] [CrossRef] [PubMed]
- Breuer, C.; Ishida, T.; Sugimoto, K. Developmental control of endocycles and cell growth in plants. Curr. Opin. Plant Biol. 2010, 13, 654–660. [Google Scholar] [CrossRef] [PubMed]
- Cookson, S.J.; Radziejwoski, A.; Granier, C. Cell and leaf size plasticity in Arabidopsis: What is the role of endoreduplication? Plant Cell Environ. 2006, 29, 1273–1283. [Google Scholar] [CrossRef] [PubMed]
- Burssens, S.; Montagu, M.V.; Inzé, D. The cell cycle in Arabidopsis. Plant Physiol. Biochem. 1998, 36, 9–19. [Google Scholar] [CrossRef]
- Vandepoele, K.; Raes, J.; de Veylder, L.; Rouzé, P.; Rombauts, S.; Inzé, D. Genome-wide analysis of core cell cycle genes in Arabidopsis. Plant Cell 2002, 14, 903–916. [Google Scholar] [CrossRef] [PubMed]
- Menges, M.; de Jager, S.M.; Gruissem, W.; Murray, J.A. Global analysis of the core cell cycle regulators of Arabidopsis identifies novel genes, reveals multiple and highly specific profiles of expression and provides a coherent model for plant cell cycle control. Plant J. 2005, 41, 546–566. [Google Scholar] [CrossRef] [PubMed]
- López-Juez, E.; Dillon, E.; Magyar, Z.; Khan, S.; Hazeldine, S.; de Jager, S.M.; Murray, J.A.; Beemster, G.T.; Bögre, L.; Shanahan, H. Distinct light-initiated gene expression and cell cycle programs in the shoot apex and cotyledons of Arabidopsis. Plant Cell 2008, 20, 947–968. [Google Scholar] [CrossRef] [PubMed]
- Noir, S.; Bömer, M.; Takahashi, N.; Ishida, T.; Tsui, T.L.; Balbi, V.; Shanahan, H.; Sugimoto, K.; Devoto, A. Jasmonate controls leaf growth by repressing cell proliferation and the onset of endoreduplication while maintaining a potential stand-by mode. Plant Physiol. 2013, 161, 1930–1951. [Google Scholar] [CrossRef] [PubMed]
- Dewitte, W.; Riou-Khamlichi, C.; Scofield, S.; Healy, J.S.; Jacqmard, A.; Kilby, N.J.; Murray, J.A. Altered cell cycle distribution, hyperplasia, and inhibited differentiation in Arabidopsis caused by the D-type cyclin CYCD3. Plant Cell 2003, 15, 79–92. [Google Scholar] [CrossRef] [PubMed]
- Imai, K.K.; Ohashi, Y.; Tsuge, T.; Yoshizumi, T.; Matsui, M.; Oka, A.; Aoyama, T. The A-type cyclin CYCA2;3 is a key regulator of ploidy levels in Arabidopsis endoreduplication. Plant Cell 2006, 18, 382–396. [Google Scholar] [CrossRef] [PubMed]
- González-García, M.P.; Vilarrasa-Blasi, J.; Zhiponova, M.; Divol, F.; Mora-García, S.; Russinova, E.; Caño-Delgado, A.I. Brassinosteroids control meristem size by promoting cell cycle progression in Arabidopsis roots. Development 2011, 138, 849–859. [Google Scholar] [CrossRef] [PubMed]
- Peng, L.; Skylar, A.; Chang, P.L.; Bisova, K.; Wu, X. CYCP2;1 integrates genetic and nutritional information to promote meristem cell division in Arabidopsis. Dev. Boil. 2014, 393, 160–170. [Google Scholar] [CrossRef] [PubMed]
- Francis, D. The plant cell cycle—15 Years on. New Phytol. 2007, 174, 261–278. [Google Scholar] [CrossRef] [PubMed]
- Ohnishi, N.; Kodama, H.; Ando, S.; Komamine, A. Synthesis of protein and mRNA is necessary for the transition of suspension-cultured Catharanthus roseus cells from G1 to the S phase of the cell cycle. Physiol. Plant. 1990, 80, 95–101. [Google Scholar] [CrossRef]
- Ito, T.; Kim, G.T.; Shinozaki, K. Disruption of an Arabidopsis S-13 homologous gene by transposon-mediated mutagenesis causes aberrant growth and development. Plant J. 2000, 22, 257–264. [Google Scholar] [CrossRef] [PubMed]
- Weijers, D.; van Dijk, M.F.; Venken, R.J.; Quint, A.; Hooykaas, P.; Offringa, R. An Arabidopsis minute-like phenotype caused by a semi-dominant mutation in a RIBOSOMAL PROTEIN S5 gene. Development 2001, 128, 4289–4299. [Google Scholar] [PubMed]
- Shi, H.; Chen, K.; Wei, Y.; He, C. Fundamental issues of melatonin-mediated stress signaling in plants. Front. Plant Sci. 2016, 7, 1124. [Google Scholar] [CrossRef] [PubMed]
- Mansfield, S.G.; Briarty, L.G. Cotyledon cell development in Arabidopsis thaliana during reserve deposition. Can. J. Bot. 1992, 70, 151–164. [Google Scholar] [CrossRef]
- Tsukaya, H.; Tsuge, T.; Uchimiya, H. The cotyledon: A superior system for studies of leaf development. Planta 1994, 195, 309–312. [Google Scholar] [CrossRef]
- Ferjani, A.; Segami, S.; Horiguchi, G.; Muto, Y.; Maeshima, M.; Tsukaya, H. Keep an eye on PPi: The vacuolar-type H+-pyrophosphatase regulates postgerminative development in Arabidopsis. Plant Cell 2011, 23, 2895–2908. [Google Scholar] [CrossRef] [PubMed]
- Sabelli, P.A.; Larkins, B.A. The endoreduplication cell cycle: Regulation and function. Plant Cell Monogr. 2008, 9, 75–100. [Google Scholar]
- Yoshizumi, T.; Breuer, C.; Matsui, M.; Sugimoto-Shirasu, K. Plant cell growth signaling and its link to ploidy. Plant Cell Monogr. 2008, 10, 107–125. [Google Scholar]
- Edgar, B.A.; Orr-Weaver, T.L. Endoreplication cell cycles: More for less. Cell 2001, 105, 297–306. [Google Scholar] [CrossRef]
- Sugimoto-Shirasu, K.; Roberts, K. “Big it up”: Endoreduplication and cell-size control in plants. Curr. Opin. Plant Biol. 2003, 6, 544–553. [Google Scholar] [CrossRef] [PubMed]
- Bramsiepe, J.; Wester, K.; Weinl, C.; Roodbarkelari, F.; Kasili, R.; Larkin, J.C.; Hülskamp, M.; Schnittger, A. Endoreplication controls cell fate maintenance. PLoS Genet. 2010, 6, e1000996. [Google Scholar] [CrossRef] [PubMed]
- Orr-Weaver, T.L. When bigger is better: The role of polyploidy in organogenesis. Trends Genet. 2015, 31, 307–315. [Google Scholar] [CrossRef] [PubMed]
- Katagiri, Y.; Hasegawa, J.; Fujikura, U.; Hoshino, R.; Matsunaga, S.; Tsukaya, H. The coordination of ploidy and cell size differs between cell layers in leaves. Development 2016, 143, 1120–1125. [Google Scholar] [CrossRef] [PubMed]
- Breuer, C.; Stacey, N.J.; West, C.E.; Zhao, Y.; Chory, J.; Tsukaya, H.; Azumi, Y.; Maxwell, A.; Roberts, K.; Sugimoto-Shirasu, K. BIN4, a novel component of the plant DNA topoisomerase VI complex, is required for endoreduplication in Arabidopsis. Plant Cell 2007, 19, 3655–3668. [Google Scholar] [CrossRef] [PubMed]
- Hur, Y.S.; Um, J.H.; Kim, S.; Kim, K.; Park, H.J.; Lim, J.S.; Kim, W.Y.; Jun, S.E.; Yoon, E.K.; Lim, J.; et al. Arabidopsis thaliana homeobox 12 (ATHB12), a homeodomain-leucine zipper protein, regulates leaf growth by promoting cell expansion and endoreduplication. New Phytol. 2015, 205, 316–328. [Google Scholar] [CrossRef] [PubMed]
- Posmyk, M.M.; Kuran, H.; Marciniak, K.; Janas, K.M. Presowing seed treatment with melatonin protects red cabbage seedlings against toxic copper ion concentrations. J Pineal Res. 2008, 45, 24–31. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Gusti, A.; Cheminant, S.; Alioua, M.; Dhondt, S.; Coppens, F.; Beemster, G.T.; Genschik, P. Gibberellin signaling controls cell proliferation rate in Arabidopsis. Curr. Boil. 2009, 19, 1188–1193. [Google Scholar] [CrossRef] [PubMed]
- Andersen, S.U.; Buechel, S.; Zhao, Z.; Ljung, K.; Novák, O.; Busch, W.; Schuster, C.; Lohmann, J.U. Requirement of B2-type cyclin-dependent kinases for meristem integrity in Arabidopsis thaliana. Plant Cell 2008, 20, 88–100. [Google Scholar] [CrossRef] [PubMed]
- Takatsuka, H.; Ohno, R.; Umeda, M. The Arabidopsis cyclin-dependent kinase-activating kinase CDKF;1 is a major regulator of cell proliferation and cell expansion but is dispensable for CDKA activation. Plant J. 2009, 59, 475–487. [Google Scholar] [CrossRef] [PubMed]
- Menges, M.; Samland, A.K.; Planchais, S.; Murray, J.A. The D-type cyclin CYCD3;1 is limiting for the G1-to-S-phase transition in Arabidopsis. Plant Cell 2006, 18, 893–906. [Google Scholar] [CrossRef] [PubMed]
- Dewitte, W.; Scofield, S.; Alcasabas, A.A.; Maughan, S.C.; Menges, M.; Braun, N.; Collins, C.; Nieuwland, J.; Prinsen, E.; Sundaresan, V.; et al. Arabidopsis CYCD3 D-type cyclins link cell proliferation and endocycles and are rate-limiting for cytokinin responses. Proc. Natl. Acad. Sci. USA 2007, 104, 14537–14542. [Google Scholar] [CrossRef] [PubMed]
- Torres Acosta, J.A.; de Almeida Engler, J.; Raes, J.; Magyar, Z.; de Groodt, R.; Inzé, D.; de Veylder, L. Molecular characterization of Arabidopsis PHO80-like proteins, a novel class of CDKA;1-interacting cyclins. Cell Mol. Life Sci. 2004, 61, 1485–1497. [Google Scholar] [CrossRef] [PubMed]
- Doerner, P.; Jørgensen, J.E.; You, R.; Steppuhn, J.; Lamb, C. Control of root growth and development by cyclin expression. Nature 1996, 380, 520–523. [Google Scholar] [CrossRef] [PubMed]
- Shaul, O.; Mironov, V.; Burssens, S.; van Montagu, M.; Inze’, D. Two Arabidopsis cyclin promoters mediate distinctive transcriptional oscillation in synchronized tobacco BY-2 cells. Proc. Natl. Acad. Sci. USA 1996, 93, 4868–4872. [Google Scholar] [CrossRef] [PubMed]
- Morris, G.F.; Mathews, M.B. Regulation of proliferating cell nuclear antigen during the cell cycle. J. Biol. Chem. 1989, 264, 13856–13864. [Google Scholar] [PubMed]
- Strzalka, W.; Ziemienowicz, A. Proliferating cell nuclear antigen (PCNA): A key factor in DNA replication and cell cycle regulation. Ann. Bot. 2011, 107, 1127–1140. [Google Scholar] [CrossRef] [PubMed]
- Amoroso, A.; Concia, L.; Maggio, C.; Raynaud, C.; Bergounioux, C.; Crespan, E.; Cella, R.; Maga, G. Oxidative DNA damage bypass in Arabidopsis thaliana requires DNA polymerase λ and proliferating cell nuclear antigen 2. Plant Cell 2011, 23, 806–822. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.; Zhou, Z.; Elledge, S.J. The DNA replication and damage checkpoint pathways induce transcription by inhibition of the Crt1 repressor. Cell 1998, 94, 595–605. [Google Scholar] [CrossRef]
- Wang, C.; Liu, Z. Arabidopsis ribonucleotide reductases are critical for cell cycle progression, DNA damage repair, and plant development. Plant Cell 2006, 18, 350–365. [Google Scholar] [CrossRef] [PubMed]
- Yi, H.; Sardesai, N.; Fujinuma, T.; Chan, C.W.; Gelvin, S.B. Constitutive expression exposes functional redundancy between the Arabidopsis histone H2A gene HTA1 and other H2A gene family members. Plant Cell 2006, 18, 1575–1589. [Google Scholar] [CrossRef] [PubMed]
- Sundas, A.; Tandre, K.; Kvarnheden, A.; Engstrom, P. cDNA sequence and expression of an intron-containing histone H2A gene from Norway spruce, Picea abies. Plant Mol. Biol. 1993, 21, 595–605. [Google Scholar] [CrossRef] [PubMed]
- Reichheld, J.P.; Gigot, C.; Gigot, N.C. Multilevel regulation of histone gene expression during the cell cycle in tobacco cells. Nucleic Acids Res. 1998, 26, 3255–3262. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Kennedy, B.K.; Lawrence, B.D.; Barbie, D.A.; Matera, G.; Fletcher, J.A.; Harlow, E. NPAT links cyclin E-Cdk2 to the regulation of replication-dependent histone gene transcription. Genes Dev. 2000, 14, 2283–2297. [Google Scholar] [CrossRef] [PubMed]
- Nelson, D.M.; Ye, X.; Hall, C.; Santos, H.; Ma, T.; Kao, G.D.; Yen, T.J.; Harper, J.W.; Adams, P.D. Coupling of DNA synthesis and histone synthesis in S phase independent of cyclin/cdk2 activity. Mol. Cell. Biol. 2002, 22, 7459–7472. [Google Scholar] [CrossRef] [PubMed]
- Feng, G.; Qin, Z.; Yan, J.; Zhang, X.; Hu, Y. Arabidopsis ORGAN SIZE RELATED1 regulates organ growth and final organ size in orchestration with ARGOS and ARL. New Phytol. 2011, 191, 635–646. [Google Scholar] [CrossRef] [PubMed]
- Galbraith, D.W.; Harkins, K.R.; Maddox, J.M.; Ayres, N.M.; Sharma, D.P.; Firoozabady, E. Rapid flow cytometric analysis of the cell cycle in intact plant tissues. Science 1983, 220, 1049–1051. [Google Scholar] [CrossRef] [PubMed]
- Czechowski, T.; Stitt, M.; Altmann, T.; Udvardi, M.K.; Scheible, W.R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 2005, 139, 5–17. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Q.; An, B.; Shi, H.; Luo, H.; He, C. High Concentration of Melatonin Regulates Leaf Development by Suppressing Cell Proliferation and Endoreduplication in Arabidopsis. Int. J. Mol. Sci. 2017, 18, 991. https://doi.org/10.3390/ijms18050991
Wang Q, An B, Shi H, Luo H, He C. High Concentration of Melatonin Regulates Leaf Development by Suppressing Cell Proliferation and Endoreduplication in Arabidopsis. International Journal of Molecular Sciences. 2017; 18(5):991. https://doi.org/10.3390/ijms18050991
Chicago/Turabian StyleWang, Qiannan, Bang An, Haitao Shi, Hongli Luo, and Chaozu He. 2017. "High Concentration of Melatonin Regulates Leaf Development by Suppressing Cell Proliferation and Endoreduplication in Arabidopsis" International Journal of Molecular Sciences 18, no. 5: 991. https://doi.org/10.3390/ijms18050991
APA StyleWang, Q., An, B., Shi, H., Luo, H., & He, C. (2017). High Concentration of Melatonin Regulates Leaf Development by Suppressing Cell Proliferation and Endoreduplication in Arabidopsis. International Journal of Molecular Sciences, 18(5), 991. https://doi.org/10.3390/ijms18050991




