Bordetella holmesii: Lipid A Structures and Corresponding Genomic Sequences Comparison in Three Clinical Isolates and the Reference Strain ATCC 51541
Abstract
:1. Introduction
2. Results
2.1. Fatty Acids Composition
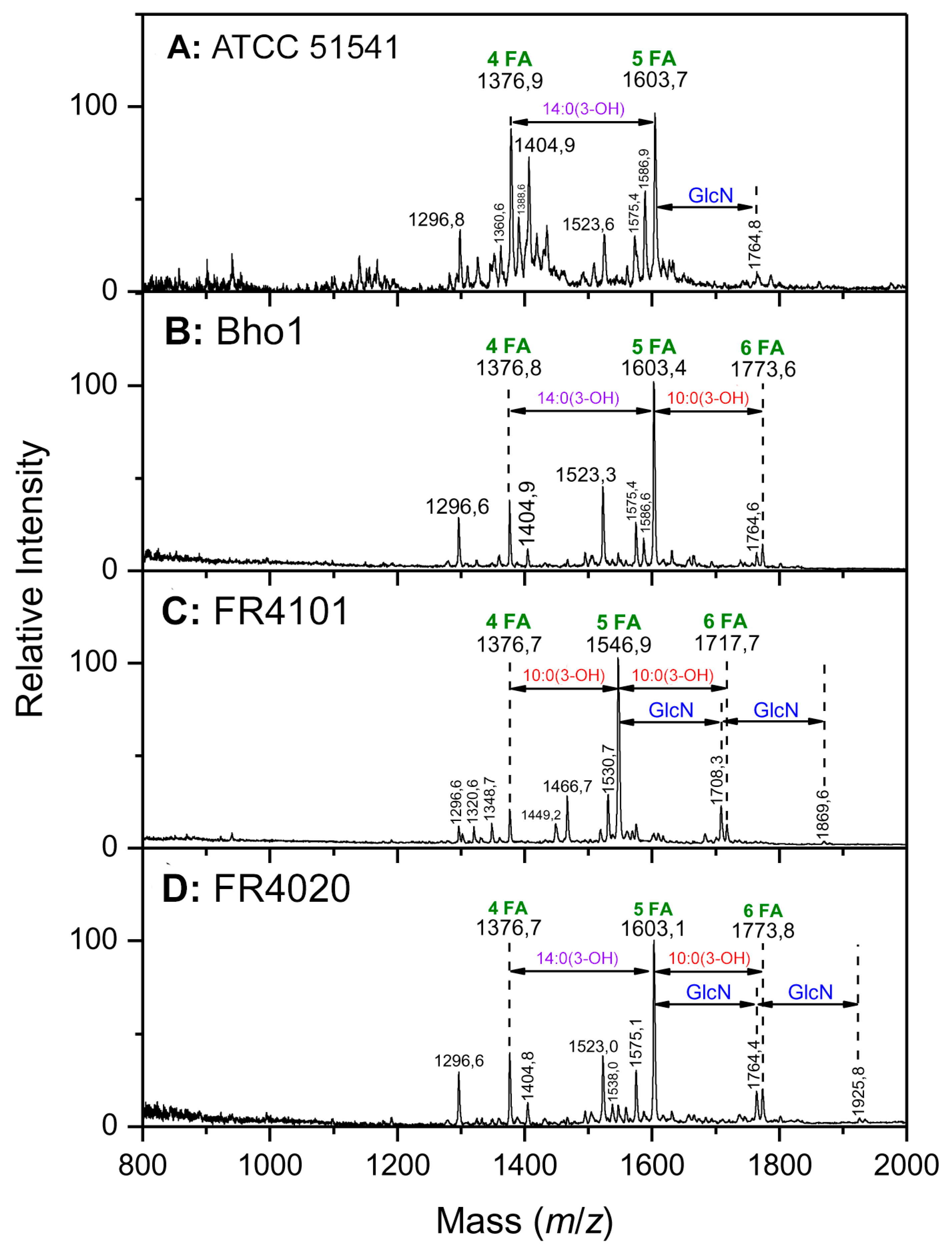
2.2. Matrix-Assisted Laser Desorption Ionization–Mass Spectrometry Structural Analyses
2.2.1. Interpretation of the Main Molecular Species in the Different Lipid a Spectra
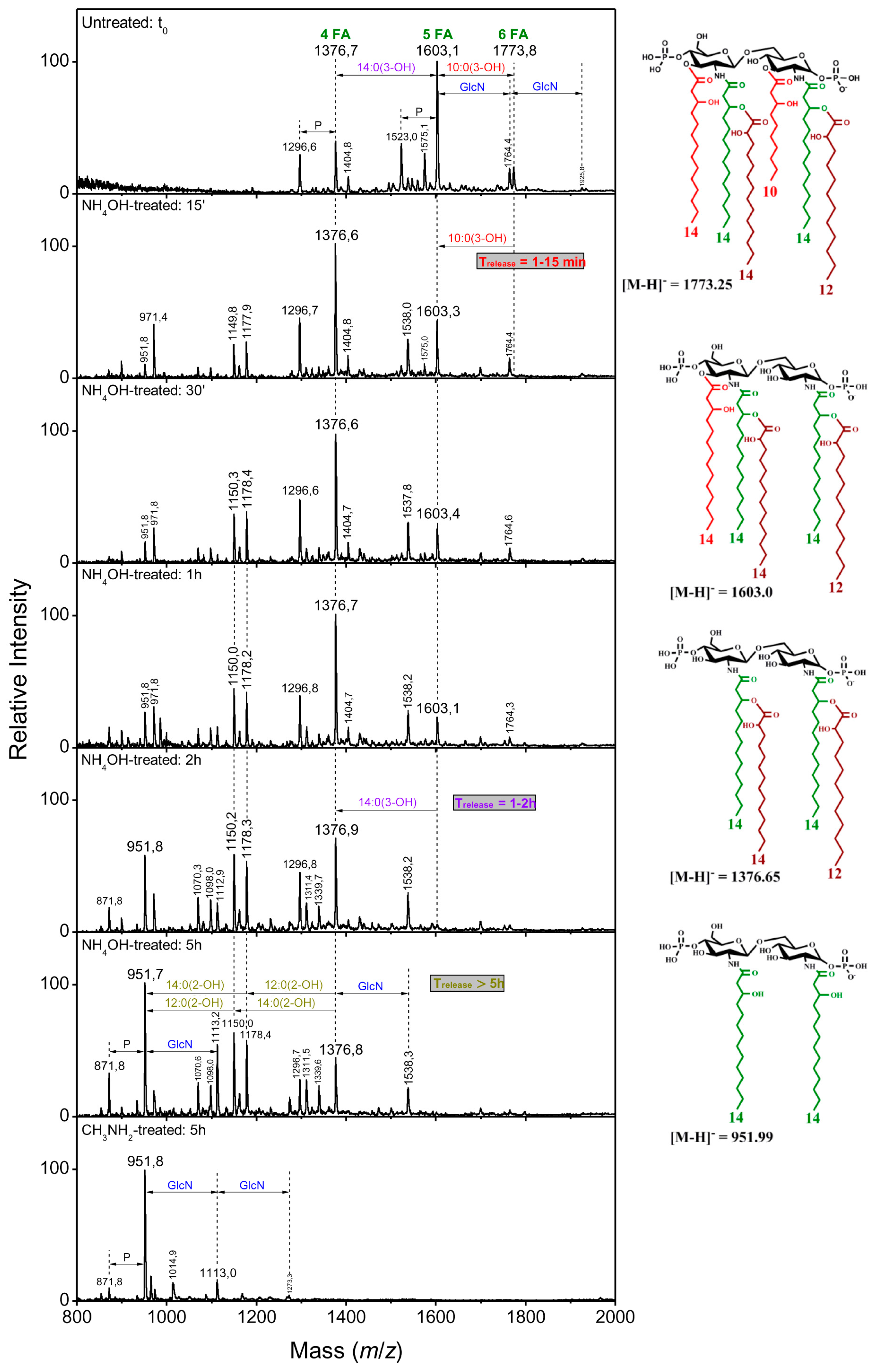
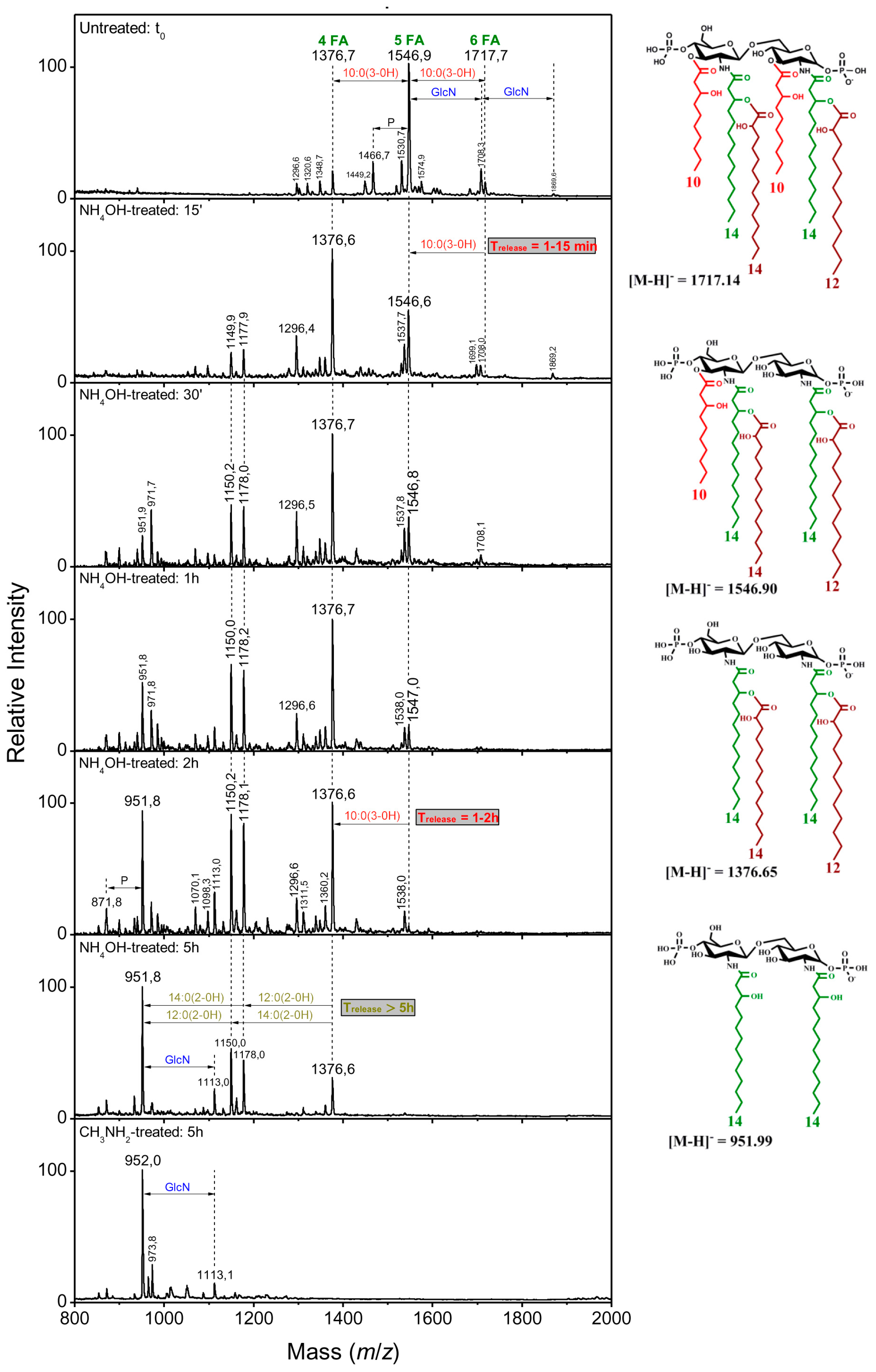
2.2.2. Distribution of the Fatty Acids on the d-Glucosamine Residues of the Clinical Isolates
2.2.3. Distribution of the Fatty Acids on the d-Glucosamine Residues of the Isolates and the Reference Strain
2.2.4. Linkage of Fatty Acids in B. holmesii Lipids A
2.3. Genomic Analyses of Genes Involved in Lipid A Biosynthesis
- (i)
- An orthologous sequence corresponding to the lgm locus was identified in the genome of the three clinical isolates as in the ATCC 51541 reference strain (position 1966909..1970398). We observe no sequence difference for this locus comprising ArnT (position 1967952..1969541) between ATCC type strain and the three clinical isolates.
- (ii)
- Bho1 and FR4020 lpxA sequence is identical to that of ATCC 51541 reference strain (position 3231726..3232520) but FR4101 lpxA sequence displays a non-synonymous SNP in position 508 leading to an amino acid modification (S instead of G in position 170 of LpxA protein). We also sequenced lpxA gene from 16 additional B. holmesii isolates collected from blood or from respiratory samples and found that this SNP was not linked to the origin of the isolates (unpublished data).
- (iii)
- pagL sequence is identical for the three clinical isolates and the ATCC 51541 reference strain (position 2144078..2144623). We did not observe any sequence difference between the three tested isolates for pagP but a difference was observed in ATCC 51541 reference strain pagL sequence (position 630144..630686) corresponding to an additional G leading to a frameshift.
- (iv)
- We found two homologs of lpxO in the genome of the three isolates of B. holmesii as in the ATCC 51541 reference strain (position 2899145..2900041 for the first one and 2402394..2403293 for the second). They respectively displayed 84% nucleotidic identity with the KF214918 lpxO1 of B. avium ATCC 35086 and 84% nucleotidic identity with the KF214919 LpxO2 of B. avium ATCC 35086. We then identified a lpxL1 homolog in the three isolates as in ATCC 51541 reference strain (position 1194726..1195580) with no sequence difference between them.
3. Discussion
- (i)
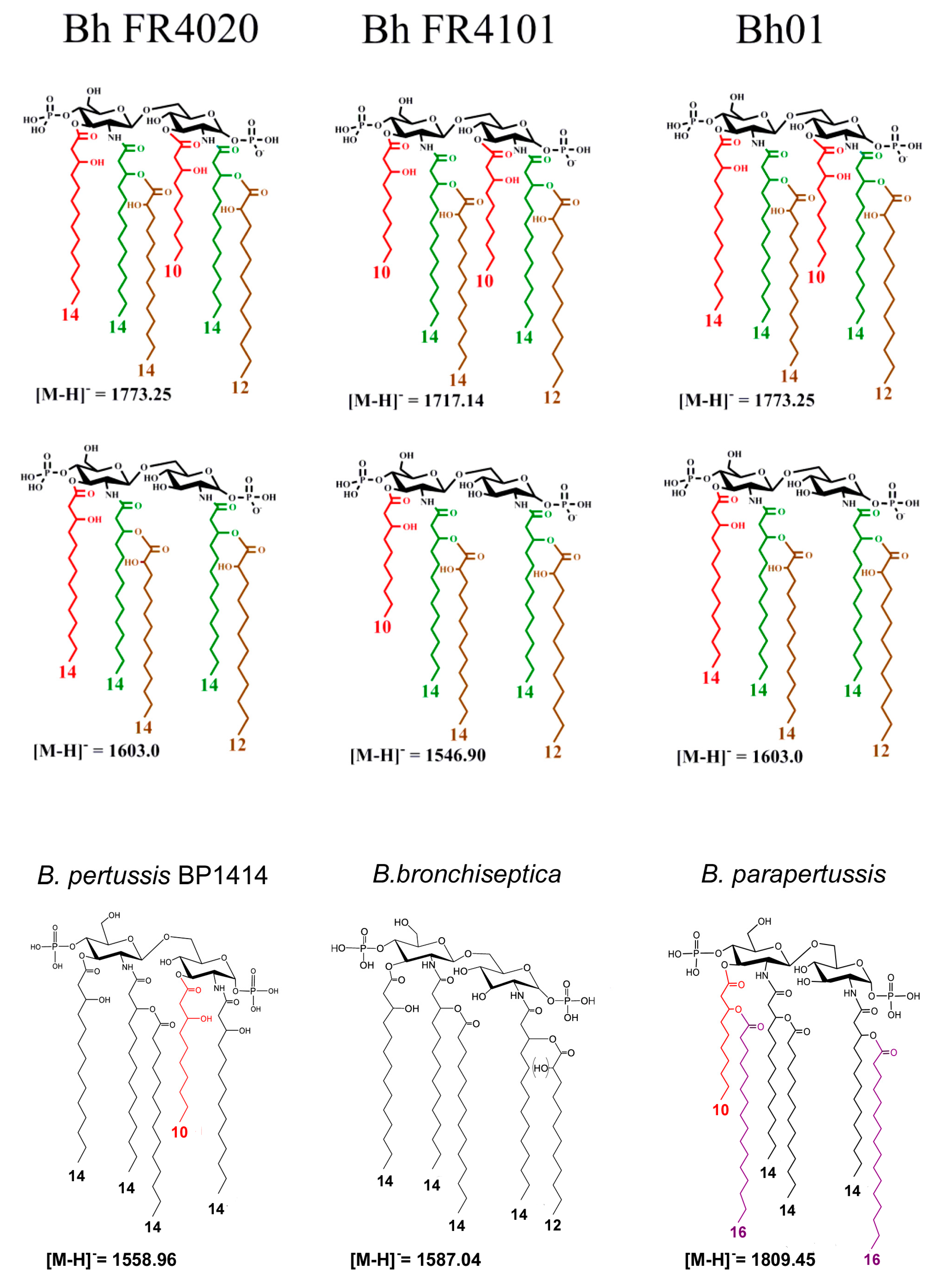
- The differences in length of the carbon FA at C3’, 14:0(3-OH) for Bho1, FR4020 and ATCC 51541 or 10:0(3-OH) for FR4101. LpxA is the first enzyme of the lipid A biosynthesis pathway catalyzing the addition of an acyl chain onto the C3’ carbon [40]. We identified lpxA B. holmesii gene and observed a non-synonymous SNP in FR4101, leading to an amino acid change, as compared to ATCC 51541, FR4020 and Bho1 lpxA sequences. Thus, we concluded that the difference observed in the length of the carbon FA at C3’ between FR4101 and the other isolates could be due to this difference within lpxA gene. Analysis of additional French B. holmesii isolates also led to the conclusion that this was not related to the blood or respiratory origin of the sample which was also confirmed by the lpxA sequence analysis of isolates with available genomes on NCBI.
- (ii)
- The presence of 10:0(3-OH) at C-3, like in B. pertussis [1] and B. petrii [41] is also a common trait between the three clinical isolates. In B. pertussis, this is the consequence of the lack of activity of the C-3 de-O-acylase PagL, which has been shown to result in a lower cytokine induction capacity of the Bordetella human pathogens [29,42]. The presence of this fatty acid was interpreted as resulting in the facility of such bacteria, with short-chain fatty acid structures, and low acylation pattern, to escape the human host defense through the MD2–TLR4 complex [42,43]. We found that the pagL gene sequence is the same for the three isolates and the ATCC 51541 strain. Both H and S residues in the C-terminal part of PagL protein, described as essential for catalytic activity are present in B. holmesii [44]. In B. pertussis, pagL is a pseudogene because of a frameshift due to a deletion of “CA” bases. Such a deletion is not found in the B. holmesii sequence. Further investigations are necessary to understand why PagL is inactive in these isolates.
- (iii)
- The presence of a secondary palmitoyl chain at two positions of lipid A (3′ and 2). We previously described the presence of such palmitoyl chains at C-3′ in B. avium, and in both C3′ and C2 in B. parapertussis [27,45] as a result of PagP action. In the present study, we observed no pagP sequence difference between the three tested isolates. They nevertheless display an additional G insertion as compared to ATCC 51541 reference strain, leading to a frameshift that probably explains why no palmitoyl acid is observed in these isolates.
- (iv)
- The hydroxylation of 12:0 and 14:0 at position 2 of the two secondary acylated residues was observed in the lipid A of the three isolates (Figure 5). In Novikov et al. [27], such hydroxylations have been observed in B. avium, B. hinzii and B. trematum and were attributed to two homologs of the LpxO enzyme. We found two homologs of lpxO in the genome of the three isolates of B. holmesii as in the ATCC 51541 type strain that support the degree of hydroxylation observed.
- (v)
- The presence of a 12:0(2-OH) residue in a secondary position at C-2 on the amide-linked 14:0(3-OH) is a common trait between all the B. holmesii structures. LpxL is a lipid A lauroyl acyltransferase. Geurtsen et al. [46] showed that LpxL1, a homolog of the classical LpxL2 lauroyl acytransferase, is present but usually poorly expressed in B. pertussis, compared to LpxL2 [40,46]. However, through overexpressing it in B. pertussis B213 strain, they showed that this enzyme led to the presence of an extra secondary 12:0(2-OH) chain at the C-2 position, like we describe here in the B. holmesii isolates, and they reported that this type of acylation was required for efficient infection in human macrophages and could help host infection. We found a lpxL1 homolog in the three isolates as in ATCC 51541 type strain supporting this substitution.
4. Materials and Methods
4.1. Strains and Isolates
4.2. LPS Preparation
4.3. Lipid A Isolation from Whole Cells
4.4. Lipid A Isolation from LPS as Performed by the Triethylamine Citrate (TEA) Method
4.5. Thin-Layer Chromatography
4.6. Fatty Acid
4.7. O-Deacylation
4.8. Mass Spectrometry
4.9. Genomic Analysis
5. Conclusions
- (i)
- Presence of free amino GlcN on the phosphate groups of lipid A structure: in Bho1 as in ATCC 51541, the failure of BvgA to function leads to a weaker substitution than in the two other isolates.
- (ii)
- Presence of 10:0(3-OH) at C-3, which we already observed in other Bordetella species such as B. pertussis [31], is thought to be the consequence of the lack of activity of the 3 O-deacylase PagL, even if genomic explanation could not be found to support this hypothesis.
- (iii)
- The 14:0(2-OH) and 12:0(2-OH) fatty acids found in all the presented structures were previously shown to be important structural features for the infection of human macrophages [42].
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| LPS | Lipopolysaccharide |
| MALDI | Matrix-ssisted laser desorption |
| MS | Mass spectrometry |
| FA | Fatty acid |
References and Notes
- Basheer, S.M.; Guiso, N.; Tirsoaga, A.; Caroff, M.; Novikov, A. Structural modifications occurring in lipid A of Bordetella bronchiseptica clinical isolates as demonstrated by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Rapid Commun. Mass Spectrom. 2011, 25, 1075–1081. [Google Scholar] [CrossRef] [PubMed]
- Flak, T.A.; Goldman, W.E. Signalling and cellular specificity of airway nitric oxide production in pertussis. Cell microbiol. 1999, 1, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Preston, A.; Maxim, E.; Toland, E.; Pishko, E.J.; Harvill, E.T.; Caroff, M.; Maskell, D.J. Bordetella bronchiseptica Pagp is a Bvg-regulated lipid A palmitoyl transferase that is required for persistent colonization of the mouse respiratory tract. Mol. Microbiol. 2003, 48, 725–736. [Google Scholar] [CrossRef] [PubMed]
- Schaeffer, L.M.; McCormack, F.X.; Wu, H.; Weiss, A.A. Interactions of pulmonary collectins with Bordetella bronchiseptica and Bordetella pertussis lipopolysaccharide elucidate the structural basis of their antimicrobial activities. Infect. Immun. 2004, 72, 7124–7130. [Google Scholar] [CrossRef] [PubMed]
- Rietschel, E.T.; Schade, U.; Jensen, M.; Wollenweber, H.W.; Luderitz, O.; Greisman, S.G. Bacterial endotoxins: Chemical structure, biological activity and role in septicaemia. Scand. J. Infect. Dis. Suppl. 1982, 31, 8–21. [Google Scholar] [CrossRef] [PubMed]
- Weyant, R.S.; Hollis, D.G.; Weaver, R.E.; Amin, M.F.; Steigerwalt, A.G.; O'Connor, S.P.; Whitney, A.M.; Daneshvar, M.I.; Moss, C.W.; Brenner, D.J. Bordetella holmesii sp. Nov., a new gram-negative species associated with septicemia. J. Clin. Microbiol. 1995, 33, 1–7. [Google Scholar] [PubMed]
- Bottero, D.; Griffith, M.M.; Lara, C.; Flores, D.; Pianciola, L.; Gaillard, M.E.; Mazzeo, M.; Zamboni, M.I.; Spoleti, M.J.; Anchart, E.; et al. Bordetella holmesii in children suspected of pertussis in Argentina. Epidemiol. Infect. 2013, 141, 714–717. [Google Scholar] [CrossRef] [PubMed]
- Rodgers, L.; Martin, S.W.; Cohn, A.; Budd, J.; Marcon, M.; Terranella, A.; Mandal, S.; Salamon, D.; Leber, A.; Tondella, M.L.; et al. Epidemiologic and laboratory features of a large outbreak of pertussis-like illnesses associated with cocirculating Bordetella holmesii and Bordetella pertussis–Ohio, 2010–2011. Clin. Infect. Dis. 2013, 56, 322–331. [Google Scholar] [CrossRef] [PubMed]
- Kamiya, H.; Otsuka, N.; Ando, Y.; Odaira, F.; Yoshino, S.; Kawano, K.; Takahashi, H.; Nishida, T.; Hidaka, Y.; Toyoizumi-Ajisaka, H.; et al. Transmission of Bordetella holmesii during pertussis outbreak, Japan. Emerg. Infect. Dis. 2012, 18, 1166–1169. [Google Scholar] [CrossRef] [PubMed]
- Miranda, C.; Porte, L.; Garcia, P. Bordetella holmesii in nasopharyngeal samples from chilean patients with suspected Bordetella pertussis infection. J. Clin. Microbiol. 2012, 50, 1505; Author reply 1506. [Google Scholar] [CrossRef] [PubMed]
- Mooi, F.R.; Bruisten, S.; Linde, I.; Reubsaet, F.; Heuvelman, K.; van der Lee, S.; King, A.J. Characterization of Bordetella holmesii isolates from patients with pertussis-like illness in the netherlands. FEMS Immunol. Med. Microbiol. 2012, 64, 289–291. [Google Scholar] [CrossRef] [PubMed]
- Njamkepo, E.; Bonacorsi, S.; Debruyne, M.; Gibaud, S.A.; Guillot, S.; Guiso, N. Significant finding of Bordetella holmesii DNA in nasopharyngeal samples from french patients with suspected pertussis. J. Clin. Microbiol. 2011, 49, 4347–4348. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.B.; Epelboin, L.; Gabarre, J.; Lecso, M.; Guillot, S.; Bricaire, F.; Caumes, E.; Guiso, N. Recurrent Bordetella holmesii bacteremia and nasal carriage in a patient receiving rituximab. Emerg. Infect. Dis. 2013, 19, 1703–1705. [Google Scholar] [CrossRef] [PubMed]
- Guiso, N.; Hegerle, N. Other bordetellas, lessons for and from pertussis vaccines. Expert Rev. Vaccines 2014, 13, 1125–1133. [Google Scholar] [CrossRef] [PubMed]
- Pittet, L.F.; Posfay-Barbe, K.M. Bordetella holmesii infection: Current knowledge and a vision for future research. Expert. Rev. Anti. Infect. Ther. 2015, 13, 965–971. [Google Scholar] [CrossRef] [PubMed]
- Pittet, L.F.; Posfay-Barbe, K.M. Bordetella holmesii: Still emerging and elusive 20 years on. Microbiol. spectr. 2016, 4. [Google Scholar] [CrossRef]
- Planet, P.J.; Narechania, A.; Hymes, S.R.; Gagliardo, C.; Huard, R.C.; Whittier, S.; Della-Latta, P.; Ratner, A.J. Bordetella holmesii: Initial genomic analysis of an emerging opportunist. Pathog. Dis. 2013, 67, 132–135. [Google Scholar] [CrossRef] [PubMed]
- Tatti, K.M.; Loparev, V.N.; Ranganathanganakammal, S.; Changayil, S.; Frace, M.; Weil, M.R.; Sammons, S.; Maccannell, D.; Mayer, L.W.; Tondella, M.L. Draft genome sequences of Bordetella holmesii strains from blood (F627) and nasopharynx (H558). Genome Announc. 2013, 1, e0005613. [Google Scholar] [CrossRef] [PubMed]
- Bouchez, V.; Guiso, N. Bordetella holmesii: Comparison of two isolates from blood and a respiratory sample. Adv. Infect. Dis. 2013, 3, 123–133. [Google Scholar] [CrossRef]
- NCBI, Bordetella pertussis genome and annotation report.
- Linz, B.; Ivanov, Y.V.; Preston, A.; Brinkac, L.; Parkhill, J.; Kim, M.; Harris, S.R.; Goodfield, L.L.; Fry, N.K.; Gorringe, A.R.; et al. Acquisition and loss of virulence-associated factors during genome evolution and speciation in three clades of Bordetella species. BMC Genom. 2016, 17, 767. [Google Scholar] [CrossRef] [PubMed]
- Gerlach, G.; Janzen, S.; Beier, D.; Gross, R. Functional characterization of the BvgAS two-component system of Bordetella holmesii. Microbiology 2004, 150, 3715–3729. [Google Scholar] [CrossRef] [PubMed]
- Diavatopoulos, D.A.; Cummings, C.A.; van der Heide, H.G.; van Gent, M.; Liew, S.; Relman, D.A.; Mooi, F.R. Characterization of a highly conserved island in the otherwise divergent Bordetella holmesii and Bordetella pertussis genomes. J. Bacteriol. 2006, 188, 8385–8394. [Google Scholar] [CrossRef] [PubMed]
- Hiramatsu, Y.; Saito, M.; Otsuka, N.; Suzuki, E.; Watanabe, M.; Shibayama, K.; Kamachi, K. BipA is associated with preventing autoagglutination and promoting biofilm formation in Bordetella holmesii. PLoS ONE 2016, 11, e0159999. [Google Scholar] [CrossRef] [PubMed]
- Link, S.; Schmitt, K.; Beier, D.; Gross, R. Identification and regulation of expression of a gene encoding a filamentous hemagglutinin-related protein in Bordetella holmesii. BMC Microbiol. 2007, 7, 100. [Google Scholar] [CrossRef] [PubMed]
- Van den Akker, W.M. Lipopolysaccharide expression within the genus Bordetella: Influence of temperature and phase variation. Microbiology 1998, 144 Pt 6, 1527–1535. [Google Scholar] [CrossRef] [PubMed]
- Novikov, A.; Shah, N.R.; Albitar-Nehme, S.; Basheer, S.M.; Trento, I.; Tirsoaga, A.; Moksa, M.; Hirst, M.; Perry, M.B.; Hamidi, A.E.; et al. Complete Bordetella avium, Bordetella hinzii and Bordetella trematum lipid A structures and genomic sequence analyses of the loci involved in their modifications. Innate Immun. 2013. [Google Scholar] [CrossRef] [PubMed]
- Caroff, M.; Aussel, L.; Zarrouk, H.; Martin, A.; Richards, J.C.; Therisod, H.; Perry, M.B.; Karibian, D. Structural variability and originality of the Bordetella endotoxins. J. Endotoxin Res. 2001, 7, 63–68. [Google Scholar] [CrossRef] [PubMed]
- MacArthur, I.; Jones, J.W.; Goodlett, D.R.; Ernst, R.K.; Preston, A. Role of pagl and lpxo in Bordetella bronchiseptica lipid A biosynthesis. J. Bact. 2011, 193, 4726–4735. [Google Scholar] [CrossRef] [PubMed]
- Kawasaki, K. Complexity of lipopolysaccharide modifications in Salmonella enterica: Its effects on endotoxin activity, membrane permeability, and resistance to antimicrobial peptides. Food Res. Int. 2012, 45, 493–501. [Google Scholar] [CrossRef]
- Albitar-Nehme, S.; Basheer, S.M.; Njamkepo, E.; Brisson, J.R.; Guiso, N.; Caroff, M. Comparison of lipopolysaccharide structures of Bordetella pertussis clinical isolates from pre- and post-vaccine era. Carbohyd. Res. 2013. [Google Scholar] [CrossRef] [PubMed]
- Tirsoaga, A.; El Hamidi, A.; Perry, M.B.; Caroff, M.; Novikov, A. A rapid, small-scale procedure for the structural characterization of lipid A applied to Citrobacter and Bordetella strains: Discovery of a new structural element. J. Lipid Res. 2007, 48, 2419–2427. [Google Scholar] [CrossRef] [PubMed]
- Marr, N.; Tirsoaga, A.; Blanot, D.; Fernandez, R.; Caroff, M. Glucosamine found as a substituent of both phosphate groups in Bordetella lipid A backbones: Role of a BvgAS-activated Arnt ortholog. J. Bact. 2008, 190, 4281–4290. [Google Scholar] [CrossRef] [PubMed]
- Geurtsen, J.; Dzieciatkowska, M.; Steeghs, L.; Hamstra, H.J.; Boleij, J.; Broen, K.; Akkerman, G.; El Hassan, H.; Li, J.; Richards, J.C.; et al. Identification of a novel lipopolysaccharide core biosynthesis gene cluster in Bordetella pertussis, and influence of core structure and lipid A glucosamine substitution on endotoxic activity. Infect. Immun. 2009, 77, 2602–2611. [Google Scholar] [CrossRef] [PubMed]
- Karibian, D.; Brunelle, A.; Aussel, L.; Caroff, M. 252Cf-plasma desorption mass spectrometry of unmodified lipid A: Fragmentation patterns and localization of fatty acids. Rapid Commun. Mass Spectrom. 1999, 13, 2252–2259. [Google Scholar] [CrossRef]
- Albitar-Nehme, S. Endotoxins of the Bordetella Genus: Structure, Evolution and Impact on Bacterial Virulence. Ph.D. Thesis, Université de Paris-Sud, Orsay, France, 13 June 2014. [Google Scholar]
- Caroff, M.; Deprun, C.; Richards, J.C.; Karibian, D. Structural characterization of the lipid A of Bordetella pertussis 1414 endotoxin. J. Bacteriol. 1994, 176, 5156–5159. [Google Scholar] [CrossRef] [PubMed]
- El Hamidi, A.; Novikov, A.; Karibian, D.; Perry, M.B.; Caroff, M. Structural characterization of Bordetella parapertussis lipid A. J. Lipid Res. 2009, 50, 854–859. [Google Scholar] [CrossRef] [PubMed]
- Therisod, H.; Labas, V.; Caroff, M. Direct microextraction and analysis of rough-type lipopolysaccharides by combined thin-layer chromatography and MALDI mass spectrometry. Anal. Chem. 2001, 73, 3804–3807. [Google Scholar] [CrossRef] [PubMed]
- Shah, N.R.; Albitar-Nehme, S.; Kim, E.; Marr, N.; Novikov, A.; Caroff, M.; Fernandez, R.C. Minor modifications to the phosphate groups and the C3’ acyl chain length of lipid a in two Bordetella pertussis strains, BP338 and 18–323, independently affect Toll-like receptor 4 protein activation. J. Biol. Chem. 2013, 288, 11751–11760. [Google Scholar] [CrossRef] [PubMed]
- Basheer, S.M.; Bouchez, V.; Novikov, A.; Augusto, L.A.; Guiso, N.; Caroff, M. Structure activity characterization of Bordetella petrii lipid A, from environment to human isolates. Biochimie 2016, 120, 87–95. [Google Scholar] [CrossRef] [PubMed]
- Marr, N.; Hajjar, A.M.; Shah, N.R.; Novikov, A.; Yam, C.S.; Caroff, M.; Fernandez, R.C. Substitution of the Bordetella pertussis lipid A phosphate groups with glucosamine is required for robust NF-kB activation and release of proinflammatory cytokines in cells expressing human but not murine Toll-like receptor 4-MD-2-CD14. Infect. Immun. 2010, 78, 2060–2069. [Google Scholar] [CrossRef] [PubMed]
- Marr, N.; Novikov, A.; Hajjar, A.M.; Caroff, M.; Fernandez, R.C. Variability in the lipooligosaccharide structure and endotoxicity among Bordetella pertussis strains. J. Infect. Dis. 2010, 202, 1897–1906. [Google Scholar] [CrossRef] [PubMed]
- Geurtsen, J.; Steeghs, L.; Hove, J.T.; van der Ley, P.; Tommassen, J. Dissemination of lipid A deacylases (Pagl) among gram-negative bacteria: Identification of active-site histidine and serine residues. J. Biol. Chem. 2005, 280, 8248–8259. [Google Scholar] [CrossRef] [PubMed]
- Hittle, L.E.; Jones, J.W.; Hajjar, A.M.; Ernst, R.K.; Preston, A. Bordetella parapertussis Pagp mediates the addition of two palmitates to the lipopolysaccharide lipid A. J. Bacteriol. 2015, 197, 572–580. [Google Scholar] [CrossRef] [PubMed]
- Geurtsen, J.; Angevaare, E.; Janssen, M.; Hamstra, H.J.; ten Hove, J.; de Haan, A.; Kuipers, B.; Tommassen, J.; van der Ley, P. A novel secondary acyl chain in the lipopolysaccharide of Bordetella pertussis required for efficient infection of human macrophages. J. Biol. Chem. 2007, 282, 37875–37884. [Google Scholar] [CrossRef] [PubMed]
- Di Fabio, J.L.; Caroff, M.; Karibian, D.; Richards, J.C.; Perry, M.B. Characterization of the common antigenic lipopolysaccharide O-chains produced by Bordetella bronchiseptica and Bordetella parapertussis. FEMS Microbiol. Lett. 1992, 76, 275–281. [Google Scholar] [CrossRef] [PubMed]
- Njamkepo, E.; Delisle, F.; Hagege, I.; Gerbaud, G.; Guiso, N. Bordetella holmesii isolated from a patient with sickle cell anemia: Analysis and comparison with other Bordetella holmesii isolates. Clin. Microbiol. Infect. 2000, 6, 131–136. [Google Scholar] [CrossRef] [PubMed]
- Tizolova, A.; Guiso, N.; Guillot, S. Insertion sequences shared by Bordetella species and implications for the biological diagnosis of pertussis syndrome. Eur. J. Clin. Microbiol. Infect. Dis. 2013, 32, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Stainer, D.W.; Scholte, M.J. A simple chemically defined medium for the production of phase I Bordetella pertussis. J. Gen. Microbiol. 1970, 63, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Caroff, M. Brevet Français. Brevet International Patent wo2004/062690a1, 9 December 2002. [Google Scholar]
- El Hamidi, A.; Tirsoaga, A.; Novikov, A.; Hussein, A.; Caroff, M. Microextraction of bacterial lipid A: Easy and rapid method for mass spectrometric characterization. J. Lipid Res. 2005, 46, 1773–1778. [Google Scholar] [CrossRef] [PubMed]
- Chafchaouni-Moussaoui, I.; Novikov, A.; Bhrada, F.; Perry, M.B.; Filali-Maltouf, A.; Caroff, M. A new rapid and micro-scale hydrolysis, using triethylamine citrate, for lipopolysaccharide characterization by mass spectrometry. Rapid Commun. Mass Spectrom. 2011, 25, 2043–2048. [Google Scholar] [CrossRef] [PubMed]
- Caroff, M.; Tacken, A.; Szabo, L. Detergent-accelerated hydrolysis of bacterial endotoxins and determination of the anomeric configuration of the glycosyl phosphate present in the “isolated lipid A” fragment of the bordetella pertussis endotoxin. Carbohydr. Res. 1988, 175, 273–282. [Google Scholar] [CrossRef]
- Haeffner, N.; Chaby, R.; Szabo, L. Identification of 2-methyl-3-hydroxydecanoic and 2-methyl-3-hydroxytetradecanoic acids in the ‘Lipid X’ fraction of the Bordetella pertussis endotoxin. Eur. J. Biochem. 1977, 77, 535–544. [Google Scholar] [CrossRef] [PubMed]
- Wollenweber, H.W.; Rietschel, E.T. Analysis of lipopolysaccharide (lipid A) fatty acids. J. Microbiol. Methods 1990, 11, 195–211. [Google Scholar] [CrossRef]

| Treatment and Time Point | [M-H]− Ions m/z | Fatty Acid Released | |||
|---|---|---|---|---|---|
| Type | Position | ||||
| A | |||||
| t0 | 1376 | 1603 | 1773 | ||
| NH4OH, 15 min | 1376 | 1603 | 1603 | 10:0(3-OH) | C3 |
| NH4OH, 2 h | 1376 | 1376 | 1376 | 14:0(3-OH) | C3′ |
| NH4OH, 5 h | 1178 | 1178 | 1178 | 12:0(2-OH) | secondary C2 partial release |
| 1150 | 1150 | 1150 | 14:0(2-OH) | secondary C2′ partial release | |
| CH3NH2, 5 h | 952 | 952 | 952 | 14:0(2-OH) | secondary C2′ |
| 12:0(2-OH) | secondary C2 | ||||
| B | |||||
| t0 | 1376.6 | 1547 | 1717 | ||
| NH4OH, 15 min | 1376.6 | 1547 | 1547 | 10:0(3-OH) | C3 |
| NH4OH, 2 h | 1376.6 | 1376.6 | 1376.6 | 10:0(3-OH) | C3′ |
| NH4OH, 5 h | 1178 | 1178 | 1178 | 12:0(2-OH) | secondary C2 partial release |
| 1150 | 1150 | 1150 | 14:0(2-OH) | secondary C2′ partial release | |
| CH3NH2, 5 h | 952 | 952 | 952 | 14:0(2-OH) | secondary C2′ |
| 12:0(2-OH) | secondary C2 | ||||
| Genes of Interest | Position within ATCC 51541 Reference Strain (Accession Number: CP007494.1) | Sequences Differences Observed Between Isolates |
|---|---|---|
| Lgm locus | 1966909..1970398 | None |
| ArnT | 1967952..1969541 | None |
| lpxA | 3231726..3232520 | No sequence differences between Bho1, FR4020 and ATCC 51541 |
| Non-synonymous SNP in position 508 for FR4101 (*) | ||
| pagL | 2144078..2144623 | None |
| pagP | 630144..630686 | No sequence differences between Bho1, FR4020 and FR4101 |
| Additional G compared to ATCC51541 leading to a frameshift | ||
| lpxO1 (**) | 2899145..2900041 | None |
| lpxO2 (***) | 2402394..2403293 | None |
| lpxL1 | 1194726..1195580 | None |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bouchez, V.; AlBitar-Nehmé, S.; Novikov, A.; Guiso, N.; Caroff, M. Bordetella holmesii: Lipid A Structures and Corresponding Genomic Sequences Comparison in Three Clinical Isolates and the Reference Strain ATCC 51541. Int. J. Mol. Sci. 2017, 18, 1080. https://doi.org/10.3390/ijms18051080
Bouchez V, AlBitar-Nehmé S, Novikov A, Guiso N, Caroff M. Bordetella holmesii: Lipid A Structures and Corresponding Genomic Sequences Comparison in Three Clinical Isolates and the Reference Strain ATCC 51541. International Journal of Molecular Sciences. 2017; 18(5):1080. https://doi.org/10.3390/ijms18051080
Chicago/Turabian StyleBouchez, Valérie, Sami AlBitar-Nehmé, Alexey Novikov, Nicole Guiso, and Martine Caroff. 2017. "Bordetella holmesii: Lipid A Structures and Corresponding Genomic Sequences Comparison in Three Clinical Isolates and the Reference Strain ATCC 51541" International Journal of Molecular Sciences 18, no. 5: 1080. https://doi.org/10.3390/ijms18051080
APA StyleBouchez, V., AlBitar-Nehmé, S., Novikov, A., Guiso, N., & Caroff, M. (2017). Bordetella holmesii: Lipid A Structures and Corresponding Genomic Sequences Comparison in Three Clinical Isolates and the Reference Strain ATCC 51541. International Journal of Molecular Sciences, 18(5), 1080. https://doi.org/10.3390/ijms18051080






