Supervised Clustering of Adipokines and Hormonal Receptors Predict Prognosis in a Population of Obese Women with Type 1 Endometrial Cancer
Abstract
:1. Introduction
2. Results
2.1. Characteristics of General and Obese Populations
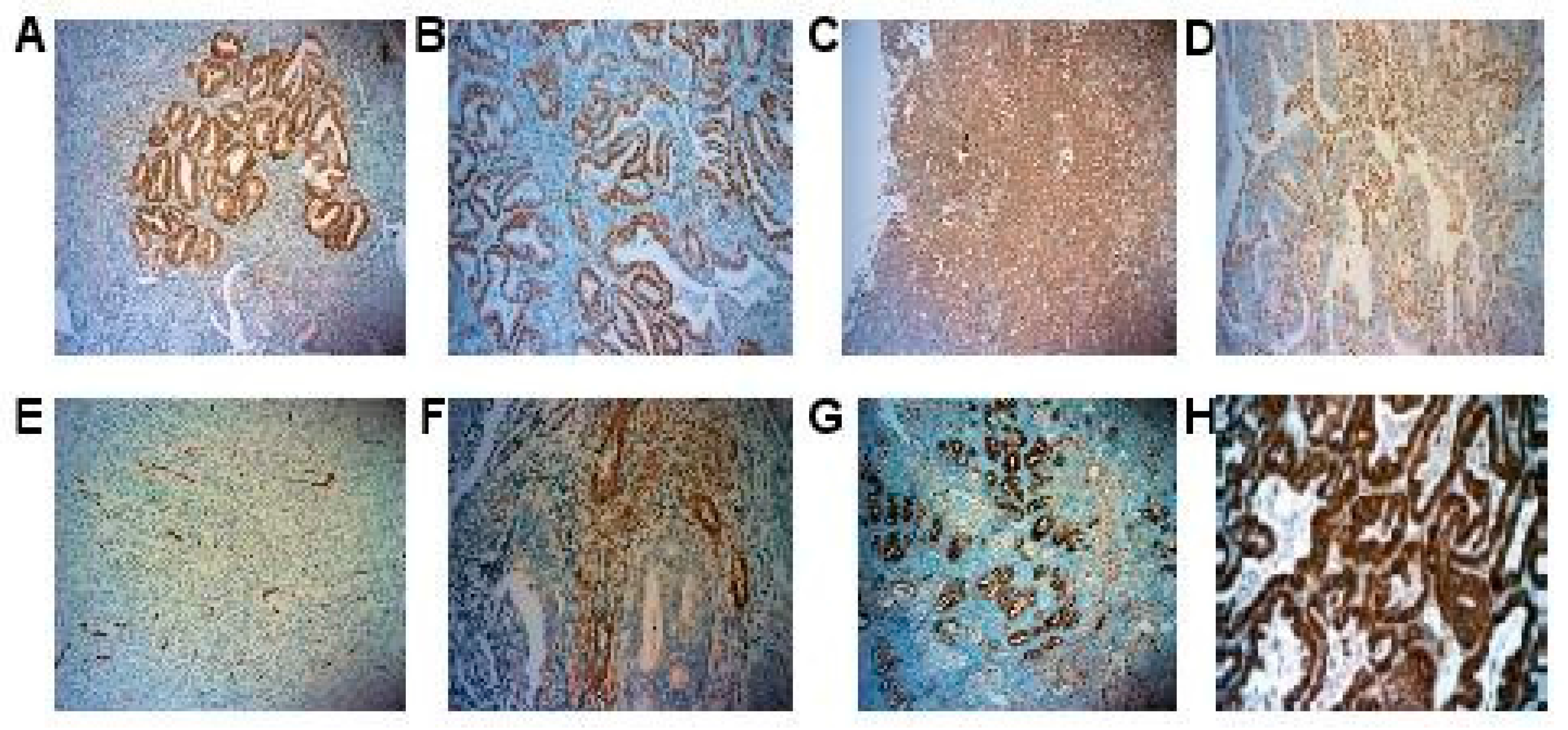
2.2. IHC Expression of Adipokines and Hormonal Receptors
2.3. Adipokines and Hormonal Receptors Expression Were Similar in Obese and Non-Obese Population
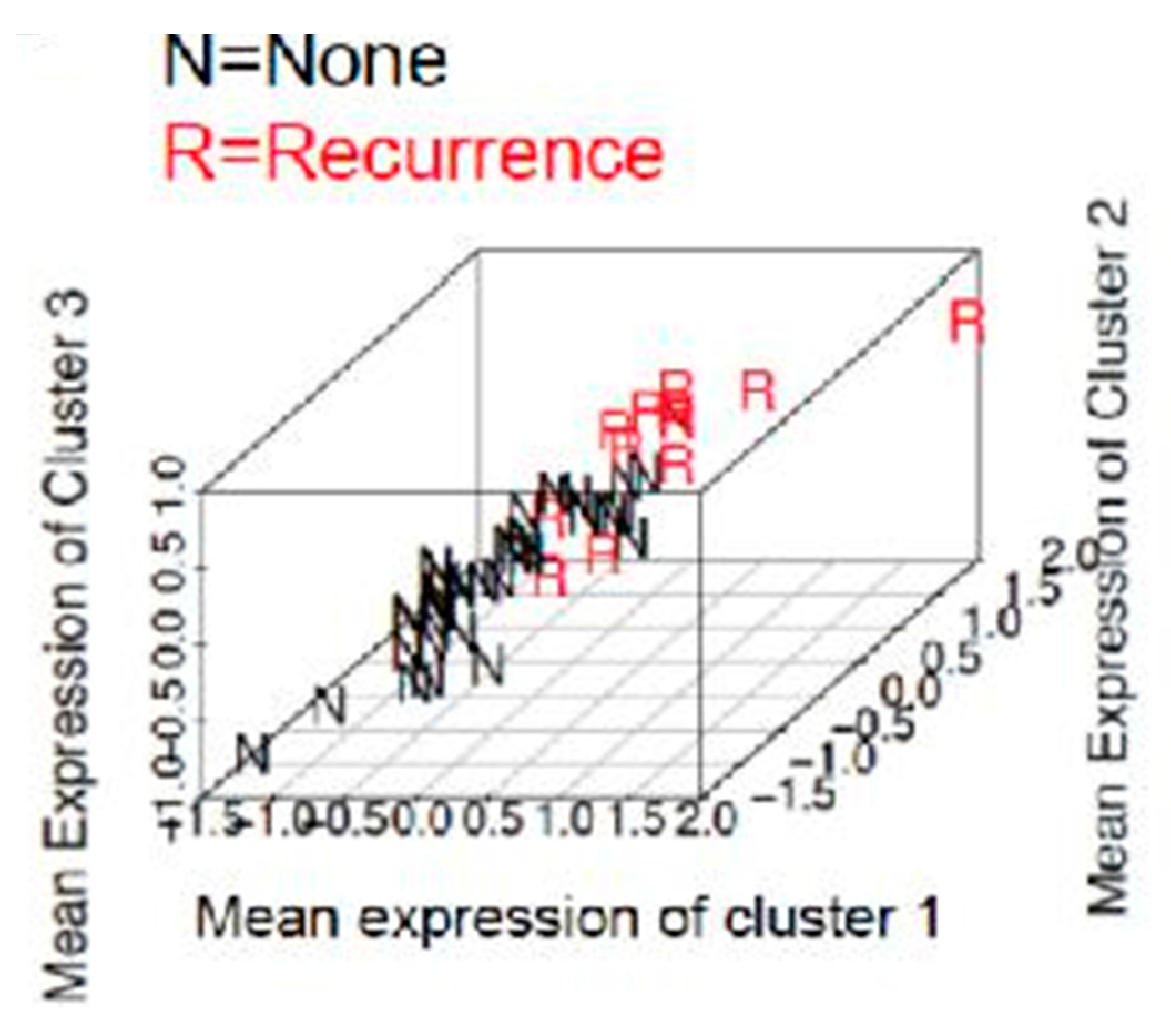
2.4. Prediction of Recurrence in Obese Population
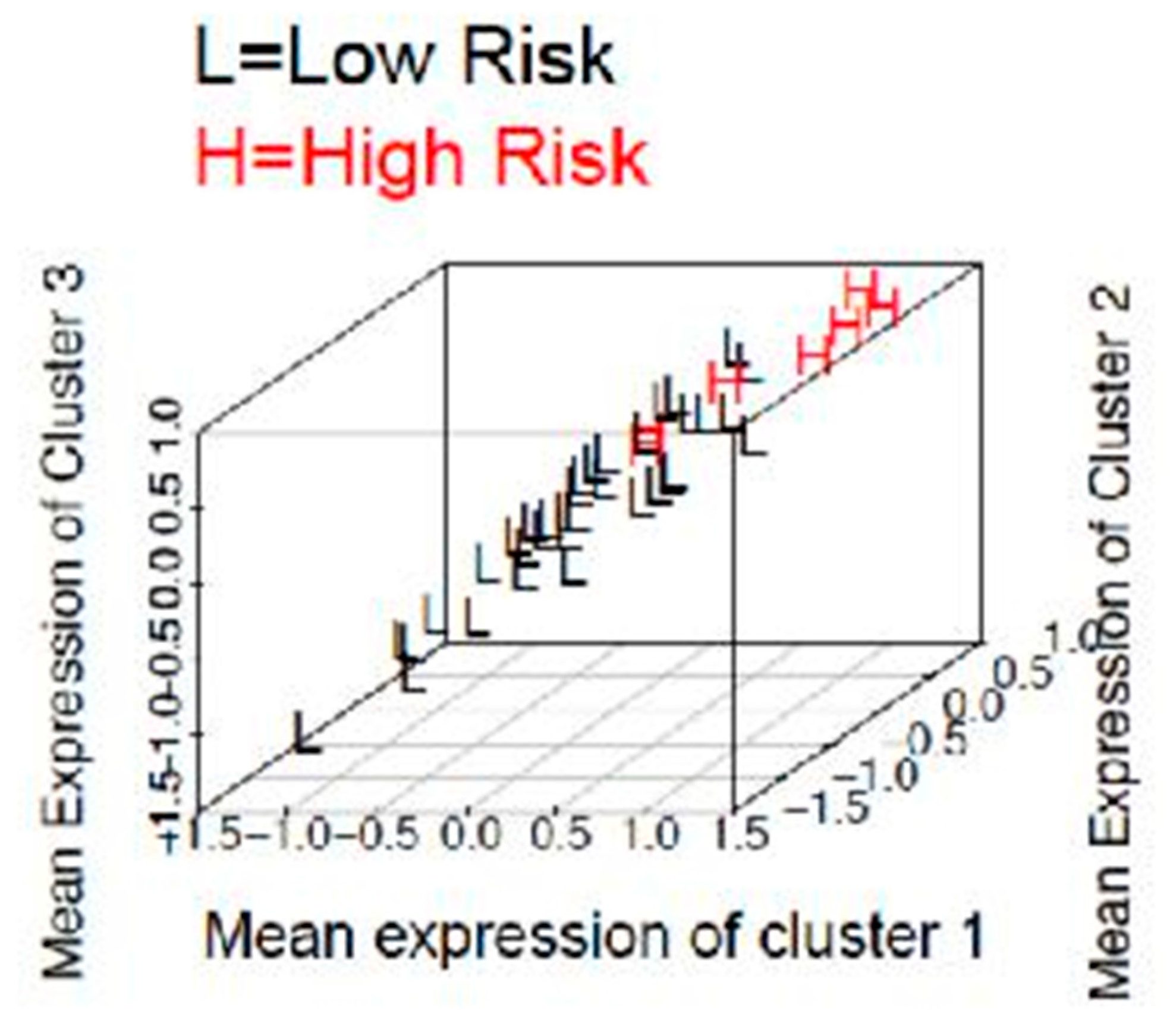
2.5. Prediction of A High Risk Stage I Group in Obese Population
2.6. Adipokines and Hormonal Receptors Expression Could Not Predict Prognostic Groups in Non Obese Population
3. Discussion
4. Material and Methods
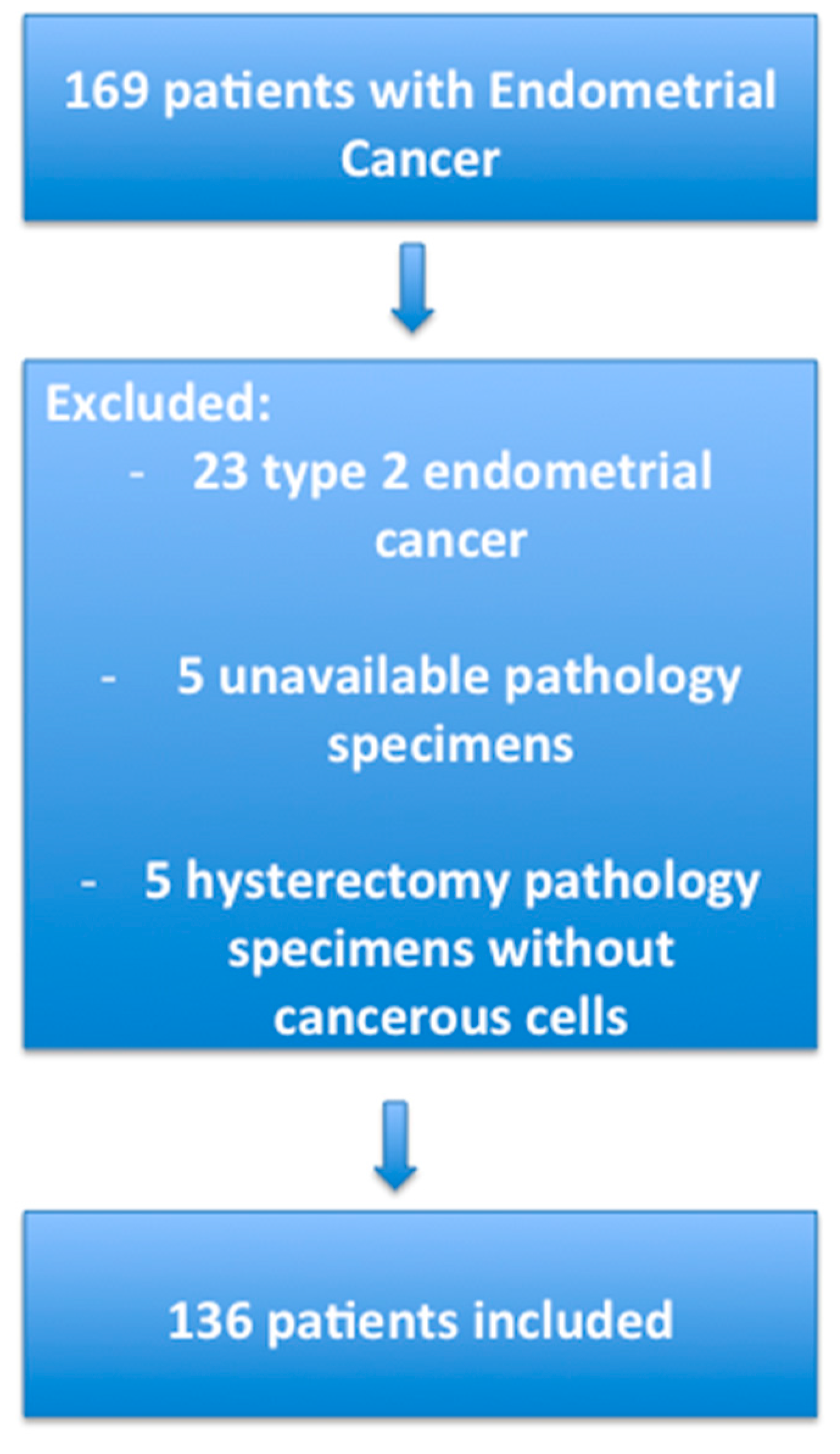
4.1. Patients
4.2. Immunohistochemistry
4.3. Adipokines and Hormonal Receptors Immunostaining
4.4. Analysis of the Immunohistochemical Results
4.5. Selection of the Groups
4.6. Statistical Analysis
4.7. Supervised Clustering
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lindemann, K.; Vatten, L.J.; Ellstrøm-Engh, M.; Eskild, A. Body mass, diabetes and smoking, and endometrial cancer risk: A follow-up study. Br. J. Cancer 2008, 98, 1582–1585. [Google Scholar] [CrossRef] [PubMed]
- Reeves, K.W.; Carter, G.C.; Rodabough, R.J.; Lane, D.; McNeeley, S.G.; Stefanick, M.L.; Paskett, E.D. Obesity in relation to endometrial cancer risk and disease characteristics in the Women’s Health Initiative. Gynecol. Oncol. 2011, 121, 376–382. [Google Scholar] [CrossRef] [PubMed]
- Fader, A.N.; Arriba, L.N.; Frasure, H.E.; von Gruenigen, V.E. Endometrial cancer and obesity: Epidemiology, biomarkers, prevention and survivorship. Gynecol. Oncol. 2009, 114, 121–127. [Google Scholar] [CrossRef] [PubMed]
- Calle, E.E.; Rodriguez, C.; Walker-Thurmond, K.; Thun, M.J. Overweight, obesity, and mortality from cancer in a prospectively studied cohort of U.S. adults. N. Engl. J. Med. 2003, 348, 1625–1638. [Google Scholar] [CrossRef] [PubMed]
- Pavelka, J.C.; Ben-Shachar, I.; Fowler, J.M.; Ramirez, N.C.; Copeland, L.J.; Eaton, L.A.; Manolitsas, T.P.; Cohn, D.E. Morbid obesity and endometrial cancer: surgical, clinical, and pathologic outcomes in surgically managed patients. Gynecol. Oncol. 2004, 95, 588–592. [Google Scholar] [CrossRef] [PubMed]
- Everett, E.; Tamimi, H.; Greer, B.; Swisher, E.; Paley, P.; Mandel, L.; Goff, B. The effect of body mass index on clinical/pathologic features, surgical morbidity, and outcome in patients with endometrial cancer. Gynecol. Oncol. 2003, 90, 150–157. [Google Scholar] [CrossRef]
- Akbayır, O.; Corbacıoglu Esmer, A.; Numanoglu, C.; Cılesız Goksedef, B.P.; Akca, A.; Bakır, L.V.; Kuru, O. Influence of body mass index on clinicopathologic features, surgical morbidity and outcome in patients with endometrial cancer. Arch. Gynecol. Obstet. 2012, 286, 1269–1276. [Google Scholar] [CrossRef] [PubMed]
- Modesitt, S.C.; Tian, C.; Kryscio, R.; Thigpen, J.T.; Randall, M.E.; Gallion, H.H.; Fleming, G.F.; Gynecologic Oncology Group. Impact of body mass index on treatment outcomes in endometrial cancer patients receiving doxorubicin and cisplatin: A Gynecologic Oncology Group study. Gynecol. Oncol. 2007, 105, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.L.; Hertan, L.; Rengan, R.; Teo, B.-K.K. Effect of body mass index on magnitude of setup errors in patients treated with adjuvant radiotherapy for endometrial cancer with daily image guidance. Int. J. Radiat. Oncol. Biol. Phys. 2012, 83, 670–675. [Google Scholar] [CrossRef] [PubMed]
- Khandwala, H.M.; McCutcheon, I.E.; Flyvbjerg, A.; Friend, K.E. The effects of insulin-like growth factors on tumorigenesis and neoplastic growth. Endocr. Rev. 2000, 21, 215–244. [Google Scholar] [CrossRef] [PubMed]
- Calle, E.E.; Kaaks, R. Overweight, obesity and cancer: Epidemiological evidence and proposed mechanisms. Nat. Rev. Cancer 2004, 4, 579–591. [Google Scholar] [CrossRef] [PubMed]
- Key, T.J.; Pike, M.C. The dose-effect relationship between “unopposed” oestrogens and endometrial mitotic rate: Its central role in explaining and predicting endometrial cancer risk. Br. J. Cancer 1988, 57, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Sherman, M.E. Theories of endometrial carcinogenesis: A multidisciplinary approach. Mod. Pathol. Off. J. U. S. Can. Acad. Pathol. 2000, 13, 295–308. [Google Scholar] [CrossRef] [PubMed]
- Zannoni, G.F.; Monterossi, G.; de Stefano, I.; Gargini, A.; Salerno, M.G.; Farulla, I.; Travaglia, D.; Vellone, V.G.; Scambia, G.; Gallo, D. The expression ratios of estrogen receptor α (ERα) to estrogen receptor β1 (ERβ1) and ERα to ERβ2 identify poor clinical outcome in endometrioid endometrial cancer. Hum. Pathol. 2013, 44, 1047–1054. [Google Scholar] [CrossRef] [PubMed]
- Huvila, J.; Talve, L.; Carpén, O.; Edqvist, P.-H.; Pontén, F.; Grénman, S.; Auranen, A. Progesterone receptor negativity is an independent risk factor for relapse in patients with early stage endometrioid endometrial adenocarcinoma. Gynecol. Oncol. 2013, 130, 463–469. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, K.; Mori, M.; Uchiyama, M.; Iwai, K.; Iwasaka, T.; Sugimori, H. Prognostic significance of progesterone receptor immunohistochemistry in endometrial carcinoma. Gynecol. Oncol. 1998, 69, 220–225. [Google Scholar] [CrossRef] [PubMed]
- Jongen, V.; Briët, J.; de Jong, R.; ten Hoor, K.; Boezen, M.; van der Zee, A.; Nijman, H.; Hollema, H. Expression of estrogen receptor-α and -β and progesterone receptor-A and -B in a large cohort of patients with endometrioid endometrial cancer. Gynecol. Oncol. 2009, 112, 537–542. [Google Scholar] [CrossRef] [PubMed]
- Maachi, M.; Piéroni, L.; Bruckert, E.; Jardel, C.; Fellahi, S.; Hainque, B.; Capeau, J.; Bastard, J.-P. Systemic low-grade inflammation is related to both circulating and adipose tissue TNFα, leptin and IL-6 levels in obese women. Int. J. Obes. 2004, 28, 993–997. [Google Scholar] [CrossRef] [PubMed]
- Modugno, F.; Ness, R.B.; Chen, C.; Weiss, N.S. Inflammation and endometrial cancer: A hypothesis. Cancer Epidemiol. Biomark. 2005, 14, 2840–2847. [Google Scholar] [CrossRef] [PubMed]
- Kos, K.; Wilding, J.P. H. SPARC: A key player in the pathologies associated with obesity and diabetes. Nat. Rev. Endocrinol. 2010, 6, 225–235. [Google Scholar] [CrossRef] [PubMed]
- Graham, T.E.; Yang, Q.; Blüher, M.; Hammarstedt, A.; Ciaraldi, T.P.; Henry, R.R.; Wason, C.J.; Oberbach, A.; Jansson, P.-A.; Smith, U.; et al. Retinol-binding protein 4 and insulin resistance in lean, obese, and diabetic subjects. N. Engl. J. Med. 2006, 354, 2552–2563. [Google Scholar] [CrossRef] [PubMed]
- Antuna-Puente, B.; Feve, B.; Fellahi, S.; Bastard, J.-P. Obesity, inflammation and insulin resistance: Which role for adipokines. Thérapie 2007, 62, 285–292. [Google Scholar] [CrossRef] [PubMed]
- Che, Q.; Liu, B.-Y.; Liao, Y.; Zhang, H.-J.; Yang, T.-T.; He, Y.-Y.; Xia, Y.-H.; Lu, W.; He, X.-Y.; Chen, Z.; et al. Activation of a positive feedback loop involving IL-6 and aromatase promotes intratumoral 17β-estradiol biosynthesis in endometrial carcinoma microenvironment. Int. J. Cancer J. Int. Cancer 2014, 135, 282–294. [Google Scholar] [CrossRef] [PubMed]
- Choi, D.S.; Kim, H.-J.; Yoon, J.-H.; Yoo, S.-C.; Jo, H.; Lee, S.Y.; Min, C.K.; Ryu, H.-S. Endometrial cancer invasion depends on cancer-derived tumor necrosis factor-α and stromal derived hepatocyte growth factor. Int. J. Cancer J. Int. Cancer 2009, 124, 2528–2538. [Google Scholar] [CrossRef] [PubMed]
- Cong, L.; Gasser, J.; Zhao, J.; Yang, B.; Li, F.; Zhao, A.Z. Human adiponectin inhibits cell growth and induces apoptosis in human endometrial carcinoma cells, HEC-1-A and RL95 2. Endocr. Relat. Cancer 2007, 14, 713–720. [Google Scholar] [CrossRef] [PubMed]
- Moon, H.-S.; Chamberland, J.P.; Aronis, K.; Tseleni-Balafouta, S.; Mantzoros, C.S. Direct role of adiponectin and adiponectin receptors in endometrial cancer: in vitro and ex vivo studies in humans. Mol. Cancer Ther. 2011, 10, 2234–2243. [Google Scholar] [CrossRef] [PubMed]
- Yamauchi, N.; Takazawa, Y.; Maeda, D.; Hibiya, T.; Tanaka, M.; Iwabu, M.; Okada-Iwabu, M.; Yamauchi, T.; Kadowaki, T.; Fukayama, M. Expression levels of adiponectin receptors are decreased in human endometrial adenocarcinoma tissues. Int. J. Gynecol. Pathol. 2012, 31, 352–357. [Google Scholar] [CrossRef] [PubMed]
- Futyma, K.; Kubiatowski, T.; Rózyńska, K.; Zdunek, M.; Kotarski, J.; Rechberger, T.; Wojcierowski, J. Decreased osteonectin and fibronectin gene expression in endometrial cancer cancer as a prognostic marker. Ginekol. Pol. 2009, 80, 907–913. [Google Scholar] [PubMed]
- Kinsel, L.B.; Szabo, E.; Greene, G.L.; Konrath, J.; Leight, G.S.; McCarty, K.S. Immunocytochemical analysis of estrogen receptors as a predictor of prognosis in breast cancer patients: Comparison with quantitative biochemical methods. Cancer Res. 1989, 49, 1052–1056. [Google Scholar] [PubMed]
- Smith, H.O.; Stephens, N.D.; Qualls, C.R.; Fligelman, T.; Wang, T.; Lin, C.-Y.; Burton, E.; Griffith, J.K.; Pollard, J.W. The clinical significance of inflammatory cytokines in primary cell culture in endometrial carcinoma. Mol. Oncol. 2013, 7, 41–54. [Google Scholar] [CrossRef] [PubMed]
- Ballester, M.; Dubernard, G.; Lécuru, F.; Heitz, D.; Mathevet, P.; Marret, H.; Querleu, D.; Golfier, F.; Leblanc, E.; Rouzier, R.; et al. Detection rate and diagnostic accuracy of sentinel-node biopsy in early stage endometrial cancer: A prospective multicentre study (SENTI-ENDO). Lancet Oncol. 2011, 12, 469–476. [Google Scholar] [CrossRef]
- Jacob, K.; Webber, M.; Benayahu, D.; Kleinman, H.K. Osteonectin promotes prostate cancer cell migration and invasion: A possible mechanism for metastasis to bone. Cancer Res. 1999, 59, 4453–4457. [Google Scholar] [PubMed]
- Liang, J.; Wang, H.; Xiao, H.; Li, N.; Cheng, C.; Zhao, Y.; Ma, Y.; Gao, J.; Bai, R.; Zheng, H. Relationship and prognostic significance of SPARC and VEGF protein expression in colon cancer. J. Exp. Clin. Cancer Res. CR 2010, 29, 71. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Jiménez, F.-J.; Caldés, T.; Iniesta, P.; Vidart, J.A.; Garcia-Asenjo, J.L.; Benito, M. Overexpression of SPARC protein contrasts with its transcriptional silencing by aberrant hypermethylation of SPARC CpG-rich region in endometrial carcinoma. Oncol. Rep. 2007, 17, 1301–1307. [Google Scholar] [CrossRef] [PubMed]
- Colombo, N.; Preti, E.; Landoni, F.; Carinelli, S.; Colombo, A.; Marini, C.; Sessa, C.; ESMO Guidelines Working Group. Endometrial cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2013, 24 (Suppl 6), vi33–vi38. [Google Scholar] [CrossRef] [PubMed]
- Dettling, M. Finding predictive genes groups from microarray data. J. Multivar. Anal. 2004, 90, 106–131. [Google Scholar] [CrossRef]
- Laas, E.; Ballester, M.; Cortez, A.; Gonin, J.; Daraï, E.; Graesslin, O. Supervised clustering of immunohistochemical markers to distinguish atypical endometrial hyperplasia from grade 1 endometrial cancer. Gynecol. Oncol. 2014, 133, 205–210. [Google Scholar] [CrossRef] [PubMed]
- Laas, E.; Ballester, M.; Cortez, A.; Gonin, J.; Canlorbe, G.; Daraï, E.; Graesslin, O. Supervised clustering of immunohistochemical markers to distinguish atypical and non-atypical endometrial hyperplasia. Gynecol. Endocrinol. 2015, 31, 282–285. [Google Scholar] [CrossRef] [PubMed]

| Characteristics | General Population | Obese Population | p |
|---|---|---|---|
| Total | 136 | 55 | - |
| Age: mean (SD) | 67.8 (11.1) | 66.1 (16.2) | 0.34 |
| Menopausal status | 128 (94.1%) | 52 (94.5%) | 1 |
| Hysterectomy with bilateral SO * | 129 (94.8%) | 52 (94.5%) | 1 |
| Hysterectomy with unilateral SO | 5 (3.6%) | 2 (3.6%) | |
| Hysterectomy without SO | 2 (1.6%) | 1 (1.9%) | |
| Lymphadenectomy | 0.43 | ||
| Pelvic | 110 (80.8%) | 41 (74.5%) | |
| Pelvic and para aortic | 8 (5.8%) | 3 (5.4%) | |
| Histological grade | 0.86 | ||
| 1 | 76 (55.8%) | 29 (52.7%) | |
| 2 | 38 (25.7%) | 17 (30.9%) | |
| 3 | 22 (18.5%) | 9 (16.4%) | |
| Lymphovascular space involvement (LVSI) | 0.49 | ||
| Yes | 33 (24.2%) | 17 (30.9%) | |
| No | 80 (58.8%) | 30 (54.5%) | |
| Data not available | 23 (17%) | 8 (14.6%) | |
| FIGO ** stage | 0.95 | ||
| I | 107 (78.7%) | 43 (78.1%) | |
| IA | 61 (44.8%) | 20 (36.3%) | |
| IB | 46 (33.8%) | 23 (41.8%) | |
| II | 8 (5.8%) | 3 (5.4%) | |
| III | 17 (12.7%) | 8 (14.5%) | |
| IV | 4 (2.8%) | 1 (2.0%) | |
| Recurrence | 25 (18.3%) | 13 (23.6%) | 0.61 |
| Positive lymph node | 11 (8.0%) | 5 (9.1%) | 0.90 |
| High risk stage I *** | 11 (8.0%) | 7 (12.7%) | 0.89 |
| Cluster | Proteins Overexpressed | Proteins Underexpressed |
|---|---|---|
| Cluster 1 | Stromal IL-6 | PR |
| RBP4 | ||
| ER | ||
| Cluster 2 | Stromal IL-6 | PR |
| RBP4 | ||
| Cluster 3 | Stromal IL-6 | PR |
| RBP4 | ||
| TNF α | ER | |
| Stromal adiponectin |
| Cluster | Proteins Overexpressed | Proteins Underexpressed |
|---|---|---|
| Cluster 1 | Stromal adiponectin | PR |
| ER | ||
| Stromal SPARC | ||
| RBP4 | ||
| Cluster 2 | Stromal adiponectin | Stromal SPARC |
| PR | ||
| RBP4 | ||
| Cluster 3 | Stromal adiponectin | Stromal SPARC |
| RBP4 | ||
| PR | ||
| Cluster 4 | Stromal adiponectin Stromal IL-6 | Stromal SPARC |
| RBP4 | ||
| PR | ||
| ER | ||
| SPARC |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Uzan, J.; Laas, E.; Alsamad, I.A.; Skalli, D.; Mansouri, D.; Haddad, B.; Touboul, C. Supervised Clustering of Adipokines and Hormonal Receptors Predict Prognosis in a Population of Obese Women with Type 1 Endometrial Cancer. Int. J. Mol. Sci. 2017, 18, 1055. https://doi.org/10.3390/ijms18051055
Uzan J, Laas E, Alsamad IA, Skalli D, Mansouri D, Haddad B, Touboul C. Supervised Clustering of Adipokines and Hormonal Receptors Predict Prognosis in a Population of Obese Women with Type 1 Endometrial Cancer. International Journal of Molecular Sciences. 2017; 18(5):1055. https://doi.org/10.3390/ijms18051055
Chicago/Turabian StyleUzan, Jennifer, Enora Laas, Issam Abd Alsamad, Dounia Skalli, Dhouha Mansouri, Bassam Haddad, and Cyril Touboul. 2017. "Supervised Clustering of Adipokines and Hormonal Receptors Predict Prognosis in a Population of Obese Women with Type 1 Endometrial Cancer" International Journal of Molecular Sciences 18, no. 5: 1055. https://doi.org/10.3390/ijms18051055
APA StyleUzan, J., Laas, E., Alsamad, I. A., Skalli, D., Mansouri, D., Haddad, B., & Touboul, C. (2017). Supervised Clustering of Adipokines and Hormonal Receptors Predict Prognosis in a Population of Obese Women with Type 1 Endometrial Cancer. International Journal of Molecular Sciences, 18(5), 1055. https://doi.org/10.3390/ijms18051055






