Abstract
Ankylosing spondylitis (AS) is a systemic autoimmune disease mainly affecting the lumbar spine and sacroiliac joints, and exhibits peripheral inflammatory arthropathy. More than 25 loci have been identified as associated with AS. Because both AS and rheumatoid arthritis (RA) are autoimmune diseases that may share some common genetic factors, we therefore examined if the newly identified RA genetic polymorphisms were associated with AS in a Taiwanese population. In this study, we enrolled 475 AS patients and 11,301 healthy subjects from a Taiwanese biobank as controls. Although none of single-nucleotide polymorphisms (SNPs) were associated with the susceptibility to AS, the AS disease index Bath AS Global (BAS-G) clinical phenotype was observed as significantly correlated to the AA genotype of rs657075 (CSF2). The significance remains after gender/age/disease duration adjustment and after group categorization by human leukocyte antigen-B 27 (HLA-B27) genotype. We further investigated the possible functions of rs657075 through bioinformatics approaches. Results revealed that polymorphism of rs657075 is able to influence the expression of acyl-CoA synthetase long-chain family member 6 (ACSL6). In conclusion, our study indicated that rs657075 (CSF2) is strongly associated with the AS disease index Bath AS Global (BAS-G) clinical phenotype.
1. Introduction
Ankylosing spondylitis (AS) is a systemic autoimmune disease, which is characterized by inflammation of the lumbar spine and sacroiliac joints, peripheral inflammatory arthropathy, and the absence of rheumatoid factor [1,2]. The disease predominantly strikes men between the age of 20 and 40 years, in their peak productive years, leading to significant loss of work productivity and a decreased quality of life [3]. Family and twin studies indicated that genetic factors contribute to over 90% to the overall AS susceptibility [4,5]. Human leukocyte antigen (HLA)-B 27 has been known as the major AS-susceptibility gene for more than 40 years [6], but HLA-B27 accounts for only 16% of the genetic variability in AS [7]. The other HLA-B allele operative in AS susceptibility is HLA-B60, which was identified in studies of US and UK patients with AS [8,9]. In addition, HLA-B60 and HLA-B61 were associated with AS in HLA-B27-negative patients in Taiwan [10]. The susceptibility to AS can be enhanced when combining different patterns of HLA-B60 and HLA-B27 genotypes in Dutch and Taiwanese populations, implying genetic interaction mechanisms that may contribute to AS risk among two genes [11,12]. The interleukin (IL)-1 and IL-23R genes were also proven to be important in the pathogenesis of AS [13,14]. Genes involved in regulating peripheral tolerance were found have a combined effect on the development of AS, such as the PD-1 G-536A/PD-L1 A8923C/PD-L2 C47103T [15] or PTPN22 G-1123C/CTLA-4 A49G [16], and the CTLA-4 +49A/G genotype associated with circulatory C-reactive protein (CRP) level [17]. Our previous studies reported that genetic polymorphisms of ORAI1 (rs12313273 and rs7135617) and STIM1 (rs3750996) were associated with the pathogenesis of HLA-B27-positive AS patients [18,19].
Additionally, many non-major histocompatibility complex (MHC) regions were found to be significantly associated with AS in genome-wide association studies (GWASs) [20,21,22]. Twelve loci were previously confirmed to be associated with AS in Europeans [20,21,23], 2 loci were recently reported in Han Chinese [22], and an additional 13 new loci were identified in a recent global GWAS, bringing the total AS-associated loci to 43 [24]. Two studies confirmed the findings of previous AS studies that ERAP1 and rs10865331 are risk factors for AS susceptibility [25,26]. However, some susceptibility loci, such as EDIL3, HAPLN1, and ANO6, discovered in a Han Chinese GWAS were not associated in a Taiwanese AS population [27].
Rheumatoid arthritis (RA) is a type of autoimmune arthritis, triggered by a faulty immune system resulting in chronic synovitis and the destruction of localized cartilage and bone [28]. Previous twin and family-base studies indicate that the associations with major histocompatibility complex (MHC) regions explain around 12% of total heritability in the susceptibility of RA [29]. A recent GWAS meta-analysis indicated that nine new loci were associated with RA in a Japanese population [30]. Because AS and RA are autoimmune diseases that may share similar genetic factors as well as immune regulatory pathways, we therefore examined if the RA susceptibility polymorphisms from GWAS meta-analysis are also associated with the pathogenesis of AS. In this study, we selected eight single-nucleotide polymorphisms (SNPs) from previous GWAS meta-analysis study. The AS activity index (Bath AS Disease Activity Index, BASDAI), Bath AS Functional Index (BASFI), and Bath AS Global (BAS-G), as well as inflammatory biochemical parameters (immunoglobulin A, IgA, erythrocyte sedimentation rate, ESR, and CRP) were analyzed. We found that the AA genotype of rs657075 (CSF2) was significantly associated to the clinical phenotype Bath AS Global (BAS-G).
2. Results
2.1. Association Study between RA Polymorphisms and Susceptibility to AS
Eight selected RA polymorphisms from previous GWAS meta-analysis were selected for genotyping (Table 1). A total of 475 AS patients were recruited in this study (Table 2). The mean age was 39.1 ± 11.3 years; 68% were men, and 431 patients (90.7%) were HLA-B27 (+). Their mean BASDAI, BASFI and BAS-G scores were 4.3 ± 2.2, 2.1 ± 2.2, and 4.4 ± 2.8, respectively. The genotypes of 11,301 healthy subjects were used as controls to analyze the susceptibility to AS. All genotyped SNPs were in Hardy–Weinberg equilibrium (Table S1). As shown in Table 3, no significant association was found between SNPs and the susceptibility of AS. Two SNPs, rs11900673 (B3GNT2) and rs2847297 (PTPN2), showed an significant association with HLA-B27-positive AS patients in the recessive model (p = 0.0394) and dominant model (p = 0.0316), respectively (Table S2). However, the significance disappeared after the Bonferroni correction.

Table 1.
Eight previously reported single-nucleotide polymorphisms (SNP) associated with rheumatoid arthritis in the Japanese population.

Table 2.
Basal characteristics of patients with ankylosing spondylitis (AS) and of Taiwanese biobank controls.

Table 3.
Genotype and allelic frequencies in a Taiwanese biobank of controls and patients with ankylosing spondylitis.
2.2. rs657075 Is Associated with the Disease Activity Index
We further analyzed the correlation between genetic polymorphisms and clinical phenotypes including the BASDAI, BASFI and BAS-G in the AS patients (Table 4). The results indicated that a higher score of BAS-G in AS patients was associated with the AA genotypes of rs657075 (CSF2) (p = 0.011; q = 0.088). Subgroup analysis indicated a similar result (p = 0.021; q = 0.168) in HLA-B27-positive AS patients (Table S3). However, no association was found between these genetic polymorphisms and other two phenotypes.

Table 4.
Difference in the scores of the Bath Ankylosing Spondylosis (AS) Disease Activity Index (BASDAI), Bath AS Functional Index (BASFI), and Bath AS Global (BAS-G) among AS patients stratified by different genotypes.
2.3. Association Study between RA Polymorphisms and Inflammatory Biochemical Parameters of AS
The correlation of genetic polymorphisms with inflammatory biochemical parameters (IgA, ESR and CRP) in AS patients were also evaluated (Table 5). The rs657075 (CSF2) AA genotype was associated with increased level of IgA (p = 0.029), however this is insignificant after multiple testing correction (q = 0.194). Subgroup analysis indicated a weak correlation between rs11900673 (B3GNT2) and the ESR level in HLA-B27-positive patients, but the result was not still significant after multiple testing correction (p = 0.029; q = 0.232) (Table S4).

Table 5.
Differences in the values of immunoglobulin A (IgA), the erythrocyte sedimentation rate (ESR), and C-reactive protein (CRP) among ankylosing spondylitis (AS) patients stratified by different genotypes.
2.4. Studies for Tissue Expression Quantitative Trait Loci (eQTLs) of rs657075
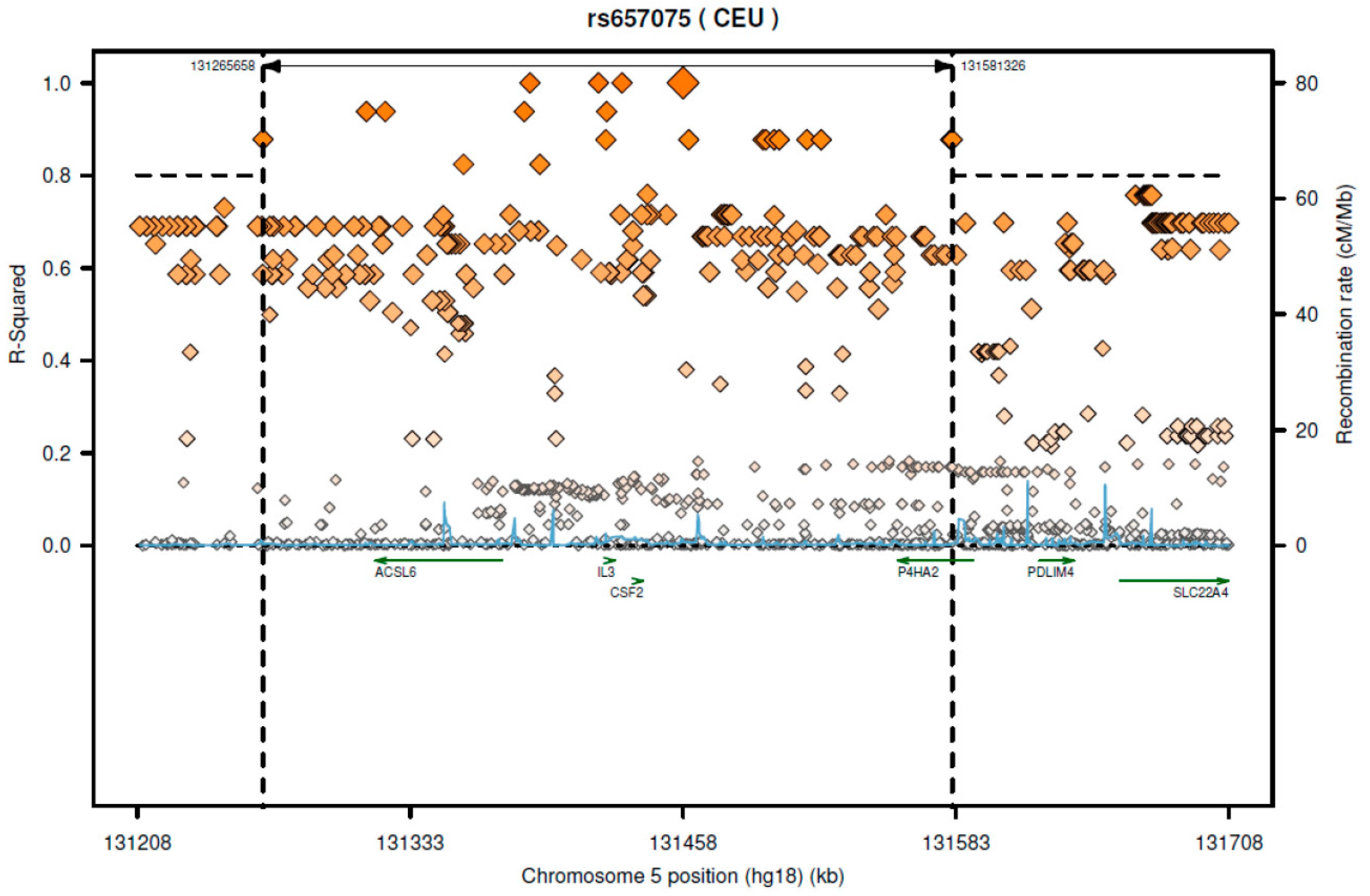
Based on 1000-genome CEU (Utah residents with Northern and Western European ancestry from the CEPH collection) population, this SNP is in great LD (linkage disequilibrium) (r2 > 0.8) with several SNPs which are located on different genes such as CSF2, acyl-CoA synthetase long-chain family member 6 (ACSL6), IL3, and P4HA2 (Figure 1). From data of the GTEx portal, we found that rs657075 can influence the expression of ACSL6 gene in colon-sigmoid cells (p = 4.7 × 10−7) (Table 6).
Figure 1.
The regional LD plot of rs657075 in CEU population.

Table 6.
Expression Quantitative Trait Loci (eQTL) results from Genotype-Tissue Expression (GTEx).
3. Discussion
AS and RA are autoimmune diseases with distinct phenotypes. RA is characterized as a chronic inflammatory joint disease with cartilage and bone damage, whereas spondyloarthropathies in AS are illustrated by entheses and subchondral bone marrow inflammation, particularly abnormal osteoproliferation at involved sites. Familial aggregations of AS and RA have been known for years [31,32]. In a 2008 Sweden study, Sundquist and coworkers analyzed 30 years of hospitalizations (1973–2004) for concordant and discordant associations among RA, AS, and systemic lupus erythematosus (SLE). They reported that significant concordant measures with standardized incidence ratios (SIRs) or sibling risks for RA and AS were 5.12 and 17.14, respectively. However, the discordant association between RA and AS was not significant, and AS was only associated with AS when discordance was taken into account [31]. A further study in 2009 indicated that offspring of parental probands with RA were significantly associated with AS with a SIR of 2.96 [32]. This laid the foundation for the identification of common genetic components for RA and AS. Recently, some autoimmune disease loci were identified as being shared among multiple autoimmune diseases [33,34]. Sirota and coworkers (2009) compared genetic variation profiles of six autoimmune disease and found that RA and AS displayed an autoimmune disease locus cluster that was distinct from the others. For instance, the G allele of rs2076530 in BTNL2 predisposed patients to RA, AS, and type 1 diabetes, but may play a protective role instead in multiple sclerosis and autoimmune thyroid disease.
To date, more than 43 AS risky loci [24] and 101 RA risky loci [35] have been identified. Most of the identified risk loci for autoimmune diseases are related to B-cell or T-cell activation pathways, differentiation, or innate immunity, or involved cytokine signaling or regulating peripheral tolerance. HLA-B27 has been known for years to be the major AS-susceptibility gene [6], but fewer than 5% of HLA-B27 carriers develop AS [24], suggesting that non-HLA-B27 alleles are important for AS susceptibility. Indeed, we recently provided evidence that genetic polymorphisms of ORAI1 (rs12313273 and rs7135617) and STIM1 (rs3750996) were associated with the pathogenesis of HLA-B27-positive AS patients [18,19]. There is evidence that three of the SNPs (rs4552569, rs17095830, and rs13210693) which correspond to two newly identified AS risky loci (EDIL3-HAPLN1 and ANO6) in Han Chinese [22] were not replicated in Caucasian populations [20,21,23] and were negatively associated with Taiwanese AS subjects in a previous study of ours [27], eliciting the question of whether these two newly-identified loci associated with AS in Han Chinese [22] may have been over-represented. Indeed the association of one SNP, rs13210693 at the 6q21 locus, did not reach a p value at the genome-wide significance level (p > 5 × 10−8) [22].
Bath ankylosing spondylitis global score (BAS-G) was a valuable quantitative measure which gives a global assessment of the well-being of the person with AS over a given time period as completed by the patient. It was first reported by Jones et al. in 1960 [36], and endorsed by Assessment of SpondyloArthritis international Society (ASAS) [37]. It uses two horizontal visual analogue scales (10 cm) to measure the effect of AS on the respondent’s well-being, where none = 0 and severe = 10. The first estimates over the last week, and the second over the last six months. In this study, we firstly reported that the genetic polymorphism rs657075 (CSF2) was associated with BAS-G.
CSF2 loci were previously reported to be associated with RA and exhibited high levels in joints of RA patients [30]. In this study, for the first time we showed that CSF2 loci were not associated with AS development, but may be weakly associated with AS clinical phenotype BAS-G levels in a Taiwanese population. CSF2, also known as granulocyte-macrophage colony-stimulating factor, functions as a cytokine and is secreted by macrophages, T cells, mast cells, natural killer (NK) cells, endothelial cells, and fibroblasts. It stimulates stem cells to produce granulocytes and is part of the immune/inflammatory cascade. On the other hand, the T helper Th17 cells require Csf2 to induce autoimmune encephalomyelitis or autoimmune neuroinflammation [38]. It is known that IL-23 and IL-23R are involved in pathogenic Th17 responses, and IL-23R is associated with many autoimmune diseases such as psoriasis, AS, and Crohn’s disease [24]. Perhaps levels of the weakly associated AS clinical phenotype BAS-G seen may have something to do with some of these gene–gene interactions.
According to SNAP website, this SNP has high LD with several genes including CSF2 and ACSL6 (Figure 1). Since the GTEx database includes limited cell types, rs657075 does not show significantly associated eQTL with the CSF2 gene in the current database. It is associated with expression of ACSL6 in sigmoid colon cells (Table 6). On searching HaploReg V4.1, rs657075 is involved in changing chromatin status in primary T cells. Therefore, further functional studies are needed to validate the impact of rs657075 on the expression of CSF2.
Using a high-density immune-related loci platform (Immunochip), Cortes and coworkers reported that some of the AS risky loci overlapped with those of other immune diseases. Eleven of these AS risky loci were positively associated with ulcerative colitis and 12 loci with Crohn’s disease. However, only two of the AS risky loci were marginally associated with RA; rs11065898 (SH2B3 or LNK) and rs2283790 (UBE2L3) displayed a concordant and discordant mode, respectively. As SH2B3 encoded an adaptor protein involving T cell signaling, it may be functionally related to the pathogenesis of AS. Interestingly, rs11065898 loci were indeed positively associated with CD4+ lymphocyte counts [24].
The modest correlation between RA risk SNPs and AS susceptibility that we observed in this study may due to several reasons.
Firstly, the SNPs were identified from a RA GWAS study conducted in a Japanese cohort. In this study, we assess their contribution to AS susceptibility and disease severity in a Taiwanese cohort. The nonsignificant results regarding susceptibility to Taiwanese AS patients may be attributed to the heterogeneity of the phenotype (diagnostic criteria of the disease). In considering of phenotypic heterogeneity, we have noted different criteria of clinical diagnosis between RA (in the GWAS study from a Japanese population) and AS (in this study based on a Taiwanese population).
Secondly, it is likely that different genetic backgrounds (variation in allele frequencies across different ethnic population admixture) in two populations may play a role in the observed inconsistent results. In particular, different genetic backgrounds between Japanese and Taiwanese populations may have an impact on the genetic influence of rs657075 in the two study populations, consequently, this may result in the inconsistent association that we observed in these two studies.
Thirdly, the prevalences of AS between Taiwanese and Japanese populations are different. The prevalence in Japanese population is lower than in Taiwanese population (0.0065% vs. 0.19%–0.54%) [39,40,41]. The distinct prevalence of AS may imply various genetic factors influencing different underlying regulations in AS across population, which may partially explain the nonsignificant association observed in our study. For example, a previous study reported that three SNPs (rs4552569, rs17095830, and rs13210693) were associated with AS in Han Chinese [22], but this was not replicated in Caucasian population [20,21,23], nor were these associated with Taiwanese AS subjects [27].
Fourthly, it has been known that the genetic effect size of SNP may be different across populations. Since some important confounding factors are unmeasured and can not be adjusted in the analyses, this may have a certain degree of influence when determining the risk effect of those SNPs on AS.
Although no correlation between RA susceptible SNPs and AS risk was observed, we have shown that these SNPs are good candidates for AS disease severity, i.e., BASDAI, BASFI, and BAS-G after adjusting for the effects of gender, age, and disease duration. It would still be interesting to consider those previously identified SNPs as good candidates for determining increased risk of AS. As such, further investigation would be merited to validate these SNPs identified from the Japanese AS GWAS study. It will be also important to the authors to further examine the association between these SNPs and increasing AS risk when sample size allows in the future.
The small sample size in this study may have limited the statistical power for detecting sizeable changes in AS risk assessments. Larger cohort studies and replication studies in different populations are required to validate our findings in this study. In summary, although the SNPs (rs657075 on CSF2 and rs11900673 on B3GNT2) are not the risk alleles for the susceptibility of AS, we provide supportive evidence that these SNPs are important generic markers for clinical manifestations of AS patients in a Taiwanese population. Further investigation would be needed to validate the findings in this study.
4. Materials and Methods
4.1. Study Subjects
Patients were sequentially recruited from the AS clinic at Chung Shan Medical University Hospital (Taichung, Taiwan). Our study conformed to the Declaration of Helsinki, and the design of the work and final report were performed with the approval of the institutional review board of Chung Shan Medical University Hospital (CSMUH No: CS12255). We received informed consent from all subjects before any data were collected. Patients with AS who met the selection criteria were asked to participate in the study. In total 475 AS patients were recruited from the Arthritis Clinic, and diagnosed by a qualified rheumatologist. The criteria included (i) patients being aged 17–82 years; (ii) an AS diagnosis having been made by modified New York criteria developed in 1984 [42]; (iii) being fluent Chinese language speakers; and (iv) having cognitive performance that was not influenced by other disease such as dementia. Patients with sacroiliitis were confirmed by a qualified radiologist. The detailed clinical history included age at initial symptoms, extra-spinal manifestations , and laboratory parameters of inflammation, i.e., the ESR, CRP, and IgA. The age of onset of AS symptom was defined as the time when the first symptom (axial symptom, peripheral arthritis, uveitis, or enthesitis) was noted. The BASDAI, BASFI, and BAS-G were applied to evaluate the disease activity, physical functions, and global wellbeing, respectively. A family history of AS was also recorded. Peripheral arthritis was defined as the presence of at least one swollen joint. These symptoms were determined by a rheumatologist, ophthalmologist, and gastroenterologist.
4.2. Candidate Single-Nucleotide Polymorphisms (SNPs)
We included eight SNPs which showed a significant association with RA in previous GWASs in a Japanese population (Table 1) [30]. rs2867461 (ANXA3) was excluded from the statistical analysis, because many subjects were unsuccessfully genotyped due to failure of polymerase chain reaction (PCR) amplification.
4.3. DNA Extraction
Peripheral venous blood was collected during medical surveillance, and processed in the same day. Blood was centrifuged to separate the serum and cells. DNA of blood cells was extracted by first treating them with 0.5% sodium dodecylsulfate lysis buffer and then protease K (1 mg/mL) to digest nuclear proteins for 4 h at 60 °C according to the manufacturer’s procedures. Total DNA was harvested using a Gentra (Qiagen, Valencia, CA, USA) extraction kit followed by 70% alcohol precipitation as recommended.
4.4. Genotyping
Genotyping for the eight SNPs was performed using the TaqMan Allelic Discrimination Assay (Applied Biosystems, Foster City, CA, USA). PCR was performed in the 96-well microplate and with the ABI9700 Thermal Cycler (Applied Biosystems). After the PCR, fluorescence was measured and analyzed using the StepOne software vers. 2.2.2 (Applied Biosystems, Foster City, CA, USA). HLA-B27 carriage was assessed by flow cytometry as previously described [43].
4.5. SNP Annotation Data Query
In order to understand to potential regulated genes by each SNP, we queried SNAP (https://www.broadinstitute.org/mpg/snap/index.php) to download the regional LD plot of interests for SNPs. The GTEx Portal (http://www.gtexportal.org/home/), which includes a variety of tissue expression quantitative trait loci (eQTLs), was used to investigate the association between gene expression profiles and the SNPs.
4.6. Statistical Analysis
SAS 9.3 (SAS Institute, Cary, NC, USA) for Windows was used for the statistical analysis. Hardy–Weinberg equilibrium (HWE) was evaluated to test the allelic frequencies of SNP’s using a chi-squared test (χ2 test). Statistical differences between the patient and control group in genotype frequencies were evaluated by a χ2 test. An analysis of variance (ANOVA) was used to compare the mean of continuous variables (BASDAI, BASFI, BAS-F, IgA, ESR, and CRP) among different genotypes in AS patients. An analysis of covariance (ANCOVA) was applied to adjust for gender, age, and disease duration. The false discovery rate (FDR) was applied in order to correct for multiple testing errors. FDR q values of <0.05 was considered statistically significant.
Supplementary Materials
Supplementary materials can be found at www.mdpi.com/1422-0067/18/1/83/s1.
Acknowledgments
This work was supported by funding from an Excellence for Cancer Research Center grant, Department of Health, Executive Yuan, Taiwan, ROC (DOH102-TD-C-111-002) and grants from the National Science Council, Taiwan, ROC (MOST 104-2320-B-038-016) and from Taipei Medical University (12310-0223) and from Chung Shan Medical University Hospital (CSH-2011-C-004).
Author Contributions
Experimental design: Wei-Chiao Chen, James Cheng-Chung Wei, and Jin-Ding Huang; Implementation of experiments: Wei-Chiao Chen, Peng Yeong Woon, and Yu-Wen Hsu; Data analysis: Wei-Chiao Chen, Hsing-Fang Lu, Wei-Chiao Chang, James Cheng-Chung Wei, and Jin-Ding Huang; Contribution of reagents/materials/analysis tools: James Cheng-Chung Wei, Wei-Chiao Chang, and Jin-Ding Huang; Clinical samples and data collection: James Cheng-Chung Wei; Writing of the paper: Wei-Chiao Chen, Hsing-Fang Lu, Henry Sung-Ching Wong, and Wei-Chiao Chang.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Braun, J.; Sieper, J. Ankylosing spondylitis. Lancet 2007, 369, 1379–1390. [Google Scholar] [CrossRef]
- Pal, B. Ankylosing spondylitis, a seronegative spondarthritis. Practitioner 1987, 231, 785–793. [Google Scholar] [PubMed]
- Calin, A.; Brophy, S.; Blake, D. Impact of sex on inheritance of ankylosing spondylitis: A cohort study. Lancet 1999, 354, 1687–1690. [Google Scholar] [CrossRef]
- Brown, M.A.; Laval, S.H.; Brophy, S.; Calin, A. Recurrence risk modelling of the genetic susceptibility to ankylosing spondylitis. Ann. Rheum. Dis. 2000, 59, 883–886. [Google Scholar] [CrossRef] [PubMed]
- Brown, M.A.; Kennedy, L.G.; MacGregor, A.J.; Darke, C.; Duncan, E.; Shatford, J.L.; Taylor, A.; Calin, A.; Wordsworth, P. Susceptibility to ankylosing spondylitis in twins: The role of genes, HLA, and the environment. Arthritis Rheumatol. 1997, 40, 1823–1828. [Google Scholar] [CrossRef]
- Brewerton, D.A.; Hart, F.D.; Nicholls, A.; Caffrey, M.; James, D.C.; Sturrock, R.D. Ankylosing spondylitis and HL-A 27. Lancet 1973, 1, 904–907. [Google Scholar] [CrossRef]
- Khan, M.A.; Ball, E.J. Genetic aspects of ankylosing spondylitis. Best Pract. Res. Clin. Rheumatol. 2002, 16, 675–690. [Google Scholar] [CrossRef]
- Robinson, W.P.; Van der Linden, S.M.; Khan, M.A.; Rentsch, H.U.; Cats, A.; Russell, A.; Thomson, G. HLA-Bw60 increases susceptibility to ankylosing spondylitis in HLA-B27+ patients. Arthritis Rheumatol. 1989, 32, 1135–1141. [Google Scholar] [CrossRef]
- Brown, M.A.; Pile, K.D.; Kennedy, L.G.; Calin, A.; Darke, C.; Bell, J.; Wordsworth, B.P.; Cornelis, F. HLA class I associations of ankylosing spondylitis in the white population in the United Kingdom. Ann. Rheum. Dis. 1996, 55, 268–270. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.C.; Tsai, W.C.; Lin, H.S.; Tsai, C.Y.; Chou, C.T. HLA-B60 and B61 are strongly associated with ankylosing spondylitis in HLA-B27-negative Taiwan Chinese patients. Rheumatology 2004, 43, 839–842. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.C.; Sung-Ching, H.W.; Hsu, Y.W.; Wen, Y.F.; Wang, W.C.; Wong, R.H.; Lu, H.F.; Gaalen, F.A.; Chang, W.C. Interaction between HLA-B60 and HLA-B27 as a Better Predictor of Ankylosing Spondylitis in a Taiwanese Population. PLoS ONE 2015, 10, e0137189. [Google Scholar] [CrossRef] [PubMed]
- Van Gaalen, F.A.; Verduijn, W.; Roelen, D.L.; Bohringer, S.; Huizinga, T.W.; Van der Heijde, D.M.; Toes, R.E. Epistasis between two HLA antigens defines a subset of individuals at a very high risk for ankylosing spondylitis. Ann. Rheum. Dis. 2013, 72, 974–978. [Google Scholar] [CrossRef] [PubMed]
- Guo, Z.S.; Li, C.; Lin, Z.M.; Huang, J.X.; Wei, Q.J.; Wang, X.W.; Xie, Y.Y.; Liao, Z.T.; Chao, S.Y.; Gu, J.R. Association of IL-1 gene complex members with ankylosing spondylitis in Chinese Han population. Int. J. Immunogenet. 2010, 37, 33–37. [Google Scholar] [CrossRef] [PubMed]
- Safrany, E.; Pazar, B.; Csongei, V.; Jaromi, L.; Polgar, N.; Sipeky, C.; Horvath, I.F.; Zeher, M.; Poor, G.; Melegh, B. Variants of the IL23R gene are associated with ankylosing spondylitis but not with Sjogren syndrome in Hungarian population samples. Scand. J. Immunol. 2009, 70, 68–74. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.H.; Wong, R.H.; Wei, J.C.; Tsay, M.D.; Chen, W.C.; Chen, H.Y.; Shih, W.T.; Chiou, S.P.; Tu, Y.C.; Lee, H.S. Effects of genetic polymorphisms of programmed cell death 1 and its ligands on the development of ankylosing spondylitis. Rheumatology 2011, 50, 1809–1813. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.H.; Wei, J.C.; Chen, C.C.; Chuang, C.S.; Chou, C.H.; Lin, Y.J.; Wang, M.F.; Wong, R.H. Associations of the PTPN22 and CTLA-4 genetic polymorphisms with Taiwanese ankylosing spondylitis. Rheumatol. Int. 2014, 34, 683–691. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.Y.; Chang, Y.H.; Lo, M.K.; Chang, C.P.; Yang, S.C.; Yang, T.P.; Ho, K.T.; Juan, C.W.; Shiau, M.Y. Polymorphisms of cytotoxic T lymphocyte-associated antigen-4 and cytokine genes in Taiwanese patients with ankylosing spondylitis. Tissue Antigens 2010, 75, 119–126. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.C.; Yen, J.H.; Juo, S.H.; Chen, W.C.; Wang, Y.S.; Chiu, Y.C.; Hsieh, T.J.; Guo, Y.C.; Huang, C.H.; Wong, R.H.; et al. Association of ORAI1 haplotypes with the risk of HLA-B27 positive ankylosing spondylitis. PLoS ONE 2011, 6, e20426. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.C.; Hung, K.S.; Hsu, Y.W.; Wong, R.H.; Huang, C.H.; Jan, M.S.; Wu, S.J.; Juan, Y.S.; Chang, W.C. Genetic polymorphisms of stromal interaction molecule 1 associated with the erythrocyte sedimentation rate and C-reactive protein in HLA-B27 positive ankylosing spondylitis patients. PLoS ONE 2012, 7, e49698. [Google Scholar] [CrossRef] [PubMed]
- Australo-Anglo-American Spondyloarthritis Consortium; Reveille, J.D.; Sims, A.M.; Danoy, P.; Evans, D.M.; Leo, P.; Pointon, J.J.; Jin, R.; Zhou, X.; Bradbury, L.A.; et al. Genome-wide association study of ankylosing spondylitis identifies non-MHC susceptibility loci. Nat. Genet. 2010, 42, 123–127. [Google Scholar]
- Evans, D.M.; Spencer, C.C.; Pointon, J.J.; Su, Z.; Harvey, D.; Kochan, G.; Oppermann, U.; Dilthey, A.; Pirinen, M.; Stone, M.A.; et al. Interaction between ERAP1 and HLA-B27 in ankylosing spondylitis implicates peptide handling in the mechanism for HLA-B27 in disease susceptibility. Nat. Genet. 2011, 43, 761–767. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Bei, J.X.; Shen, M.; Li, Q.; Liao, Z.; Zhang, Y.; Lv, Q.; Wei, Q.; Low, H.Q.; Guo, Y.M.; et al. A genome-wide association study in Han Chinese identifies new susceptibility loci for ankylosing spondylitis. Nat. Genet. 2012, 44, 73–77. [Google Scholar] [CrossRef] [PubMed]
- Wellcome Trust Case Control Consortium; Australo-Anglo-American Spondylitis Consortium; Burton, P.R.; Clayton, D.G.; Cardon, L.R.; Craddock, N.; Deloukas, P.; Duncanson, A.; Kwiatkowski, D.P.; McCarthy, M.I.; et al. Association scan of 14,500 nonsynonymous SNPs in four diseases identifies autoimmunity variants. Nat. Genet. 2007, 39, 1329–1337. [Google Scholar]
- International Genetics of Ankylosing Spondylitis Consortium; Cortes, A.; Hadler, J.; Pointon, J.P.; Robinson, P.C.; Karaderi, T.; Leo, P.; Cremin, K.; Pryce, K.; Harris, J.; et al. Identification of multiple risk variants for ankylosing spondylitis through high-density genotyping of immune-related loci. Nat. Genet. 2013, 45, 730–738. [Google Scholar]
- Wang, C.M.; Ho, H.H.; Chang, S.W.; Wu, Y.J.; Lin, J.C.; Chang, P.Y.; Wu, J.; Chen, J.Y. ERAP1 genetic variations associated with HLA-B27 interaction and disease severity of syndesmophytes formation in Taiwanese ankylosing spondylitis. Arthritis Res. Ther. 2012, 14, R125. [Google Scholar] [CrossRef] [PubMed]
- Wen, Y.F.; Wei, J.C.; Hsu, Y.W.; Chiou, H.Y.; Wong, H.S.; Wong, R.H.; Ikegawa, S.; Chang, W.C. rs10865331 associated with susceptibility and disease severity of ankylosing spondylitis in a Taiwanese population. PLoS ONE 2014, 9, e104525. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.C.; Hsu, Y.W.; Hung, K.S.; Wong, R.H.; Huang, C.H.; Liu, Y.T.; Guo, Y.C.; Ikegawa, S.; Chang, W.C. Association study of polymorphisms rs4552569 and rs17095830 and the risk of ankylosing spondylitis in a Taiwanese population. PLoS ONE 2013, 8, e52801. [Google Scholar] [CrossRef] [PubMed]
- Firestein, G.S. Evolving concepts of rheumatoid arthritis. Nature 2003, 423, 356–361. [Google Scholar] [CrossRef] [PubMed]
- Terao, C.; Raychaudhuri, S.; Gregersen, P.K. Recent Advances in Defining the Genetic Basis of Rheumatoid Arthritis. Annu. Rev. Genom. Hum. Genet. 2016, 17, 273–301. [Google Scholar] [CrossRef] [PubMed]
- Okada, Y.; Terao, C.; Ikari, K.; Kochi, Y.; Ohmura, K.; Suzuki, A.; Kawaguchi, T.; Stahl, E.A.; Kurreeman, F.A.; Nishida, N.; et al. Meta-analysis identifies nine new loci associated with rheumatoid arthritis in the Japanese population. Nat. Genet. 2012, 44, 511–516. [Google Scholar] [CrossRef] [PubMed]
- Sundquist, K.; Martineus, J.C.; Li, X.; Hemminki, K.; Sundquist, J. Concordant and discordant associations between rheumatoid arthritis, systemic lupus erythematosus and ankylosing spondylitis based on all hospitalizations in Sweden between 1973 and 2004. Rheumatology 2008, 47, 1199–1202. [Google Scholar] [CrossRef] [PubMed]
- Hemminki, K.; Li, X.; Sundquist, J.; Sundquist, K. Familial associations of rheumatoid arthritis with autoimmune diseases and related conditions. Arthritis Rheum. 2009, 60, 661–668. [Google Scholar] [CrossRef] [PubMed]
- Sirota, M.; Schaub, M.A.; Batzoglou, S.; Robinson, W.H.; Butte, A.J. Autoimmune disease classification by inverse association with SNP alleles. PLoS Genet. 2009, 5, e1000792. [Google Scholar] [CrossRef] [PubMed]
- Richard-Miceli, C.; Criswell, L.A. Emerging patterns of genetic overlap across autoimmune disorders. Genom. Med. 2012, 4, 6. [Google Scholar] [CrossRef] [PubMed]
- Okada, Y.; Wu, D.; Trynka, G.; Raj, T.; Terao, C.; Ikari, K.; Kochi, Y.; Ohmura, K.; Suzuki, A.; Yoshida, S.; et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 2014, 506, 376–381. [Google Scholar] [CrossRef] [PubMed]
- Jones, S.D.; Steiner, A.; Garrett, S.L.; Calin, A. The Bath Ankylosing Spondylitis Patient Global Score (BAS-G). Br. J. Rheumatol. 1996, 35, 66–71. [Google Scholar] [CrossRef] [PubMed]
- Zochling, J. Measures of symptoms and disease status in ankylosing spondylitis: Ankylosing Spondylitis Disease Activity Score (ASDAS), Ankylosing Spondylitis Quality of Life Scale (ASQoL), Bath Ankylosing Spondylitis Disease Activity Index (BASDAI), Bath Ankylosing Spondylitis Functional Index (BASFI), Bath Ankylosing Spondylitis Global Score (BAS-G), Bath Ankylosing Spondylitis Metrology Index (BASMI), Dougados Functional Index (DFI), and Health Assessment Questionnaire for the Spondylarthropathies (HAQ-S). Arthritis Care Res. 2011, 63, S47–S58. [Google Scholar]
- Codarri, L.; Gyulveszi, G.; Tosevski, V.; Hesske, L.; Fontana, A.; Magnenat, L.; Suter, T.; Becher, B. RORgammat drives production of the cytokine GM-CSF in helper T cells, which is essential for the effector phase of autoimmune neuroinflammation. Nat. Immunol. 2011, 12, 560–567. [Google Scholar] [CrossRef] [PubMed]
- Hukuda, S.; Minami, M.; Saito, T.; Mitsui, H.; Matsui, N.; Komatsubara, Y.; Makino, H.; Shibata, T.; Shingu, M.; Sakou, T.; et al. Spondyloarthropathies in Japan: Nationwide questionnaire survey performed by the Japan Ankylosing Spondylitis Society. J. Rheumatol. 2001, 28, 554–559. [Google Scholar] [PubMed]
- Chou, C.T.; Pei, L.; Chang, D.M.; Lee, C.F.; Schumacher, H.R.; Liang, M.H. Prevalence of rheumatic diseases in Taiwan: A population study of urban, suburban, rural differences. J. Rheumatol. 1994, 21, 302–306. [Google Scholar] [PubMed]
- Chou, C.T.; Schumacher, H.R. Human leucocyte antigens (class I and II) in central Taiwan aborigines: Can these explain the observed differences in rheumatic disease patterns compared with Han Chinese? Rheumatology 2000, 39, 1297–1299. [Google Scholar] [CrossRef] [PubMed]
- Van der Linden, S.; Valkenburg, H.A.; Cats, A. Evaluation of diagnostic criteria for ankylosing spondylitis. A proposal for modification of the New York criteria. Arthritis Rheum. 1984, 27, 361–368. [Google Scholar] [CrossRef] [PubMed]
- Chou, C.T.; Tsai, Y.F.; Liu, J.; Wei, J.C.; Liao, T.S.; Chen, M.L.; Liu, L.Y. The detection of the HLA-B27 antigen by immunomagnetic separation and enzyme-linked immunosorbent assay-comparison with a flow cytometric procedure. J. Immunol. Methods 2001, 255, 15–22. [Google Scholar] [CrossRef]
© 2017 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).