Anti-NMDA Receptor Encephalitis and Vaccination
Abstract
:1. Introduction
2. Results
2.1. Case Report
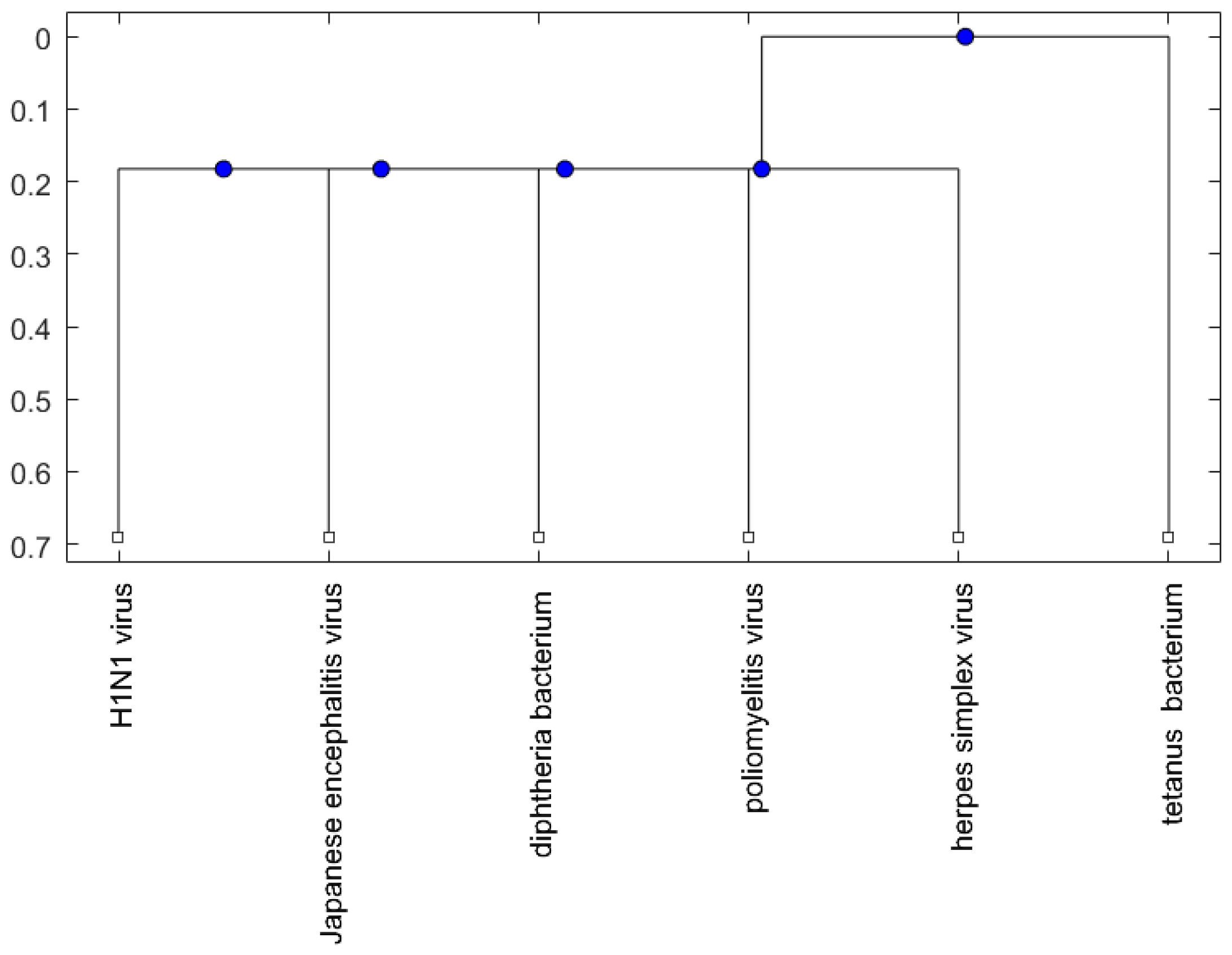
2.2. Analysis
3. Discussion
4. Methods and Materials
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Titulaer, M.J.; McCracken, L.; Gabilondo, I.; Armangué, T.; Glaser, C.; Iizuka, T.; Honig, L.S.; Benseler, S.M.; Kawachi, I.; Martinez-Hernandez, E.; et al. Treatment and prognostic factors for long-term outcome in patients with anti-NMDA receptor encephalitis: An observational cohort study. Lancet Neurol. 2013, 12, 157–165. [Google Scholar] [CrossRef]
- Wang, H. Efficacies of treatments for anti-NMDA receptor encephalitis. Front. Biosci. 2016, 21, 651–663. [Google Scholar] [CrossRef]
- Duan, B.C.; Weng, W.C.; Lin, K.L.; Wong, L.C.; Li, S.T.; Hsu, M.H.; Lin, J.J.; Fan, P.C.; Lin, M.I.; Chiu, N.C. Variations of movement disorders in anti-N-methyl-d-aspartate receptor encephalitis: A nationwide study in Taiwan. Medicine 2016, 95, e4365. [Google Scholar] [CrossRef] [PubMed]
- Offit, P.A.; Hackett, C.J. Addressing parents’ concerns: Do vaccines cause allergic or autoimmune diseases? Pediatrics 2003, 111, 653–659. [Google Scholar] [CrossRef] [PubMed]
- Shim, B.S.; Wu, W.; Kyriakis, C.S.; Bakre, A.; Jorquera, P.A.; Perwitasari, O.; Tripp, R.A. MicroRNA-555 has potent antiviral properties against poliovirus. J. Gen. Virol. 2016, 97, 659–668. [Google Scholar] [CrossRef] [PubMed]
- Steinman, L. Multiple sclerosis: A two-stage disease. Nat. Immunol. 2001, 2, 762–764. [Google Scholar] [CrossRef] [PubMed]
- Regner, M.; Lambert, P.H. Autoimmunity through infection or immunization? Nat. Immunol. 2001, 2, 185–188. [Google Scholar] [CrossRef] [PubMed]
- Albert, L.J.; Inman, R.D. Mechanisms of disease: Molecular mimicry and autoimmunity. N. Engl. J. Med. 1999, 341, 2068–2074. [Google Scholar] [PubMed]
- Hofmann, C.; Baur, M.O.; Schroten, H. Anti-NMDA receptor encephalitis after TdaP-IPV booster vaccination: Cause or coincidence? J. Neurol. 2011, 258, 500–501. [Google Scholar] [CrossRef] [PubMed]
- Dalmau, J.; Lancaster, E.; Martinez-Hernandez, E.; Rosenfeld, M.R.; Balice-Gordon, R. Clinical experience and laboratory investigations in patients with anti-NMDAR encephalitis. Lancet Neurol. 2011, 10, 63–74. [Google Scholar] [CrossRef]
- Mohammad, S.S.; Sinclair, K.; Pillai, S.; Merheb, V.; Aumann, T.D.; Gill, D.; Dale, R.C.; Brilot, F. Herpes simplex encephalitis relapse with chorea is associated with autoantibodies to N-methyl-d-aspartate receptor or dopamine-2 receptor. Mov. Disord. 2014, 29, 117–122. [Google Scholar] [CrossRef] [PubMed]
- DeSena, A.; Graves, D.; Warnack, W.; Greenberg, B.M. Herpes simplex encephalitis as a potential cause of anti-N-methyl-d-aspartate receptor antibody encephalitis report of 2 cases. JAMA Neurol. 2014, 71, 344–346. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, K.; Arita, T. Experimental double infection of Japanese encephalitis-virus and herpes-simplex virus in mouse-brain. Jpn. J. Exp. Med. 1977, 47, 9–13. [Google Scholar] [PubMed]
- Chuang, J.C.; Jones, P.A. Epigenetics and microRNAs. Pediatr. Res. 2007, 61, 24R–29R. [Google Scholar] [CrossRef] [PubMed]
- Wang, H. Predicting cancer-related miRNAs using expression profiles in tumor tissue. Curr. Pharm. Biotechnol. 2014, 15, 438–444. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, W.J.; Lin, F.; Huang, H.; Wang, H. Investigating microRNA-Target interaction-supported tissues in human cancer tissues based on miRNA and target gene expression profiling. PLoS ONE 2014, 9, e95697. [Google Scholar] [CrossRef] [PubMed]
- Taguchi, Y.H. Identification of more feasible microRNA–mRNA interactions within multiple cancers using principal component analysis based unsupervised feature extraction. Int. J. Mol. Sci. 2016, 17, 696. [Google Scholar] [CrossRef] [PubMed]
- Wang, H. Predicting microRNA biomarkers for cancer using phylogenetic tree and microarray analysis. Int. J. Mol. Sci. 2016, 17, 773. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Liu, H.; Gao, S.; Jiang, W.; Huang, W. Cellular microRNAs inhibit replication of the H1N1 influenza A virus in infected cells. J. Virol. 2010, 84, 8849–8860. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.J.; Yang, J.; Fan, X.L.; Zhao, H.B.; Hu, W.; Li, Z.P.; Yu, G.C.; Ding, X.R.; Wang, J.Z.; Bo, X.C.; et al. Cellular microRNA let-7c inhibits M1 protein expression of the H1N1 influenza A virus in infected human lung epithelial cells. J. Cell. Mol. Med. 2012, 16, 2539–2546. [Google Scholar] [CrossRef] [PubMed]
- Song, H.; Wang, Q.; Guo, Y.; Liu, S.; Song, R.; Gao, X.; Dai, L.; Li, B.; Zhang, D.; Cheng, J. Microarray analysis of microRNA expression in peripheral blood mononuclear cells of critically ill patients with influenza A (H1N1). BMC Infect. Dis. 2013, 13, 257. [Google Scholar] [CrossRef] [PubMed]
- Terrier, O.; Textoris, J.; Carron, C.; Marcel, V.; Bourdon, J.C.; Rosa-Calatrava, M. Host microRNA molecular signatures associated with human H1N1 and H3N2 influenza A viruses reveal an unanticipated antiviral activity for miR-146a. J. Gen. Virol. 2013, 94, 985–995. [Google Scholar] [CrossRef] [PubMed]
- Ge, Y.Y.; Zhao, K.; Qi, Y.; Min, X.; Shi, Z.; Qi, X.; Shan, Y.; Cui, L.; Zhou, M.; Wang, Y. Serum microRNA expression profile as a biomarker for the diagnosis of pertussis. Mol. Biol. Rep. 2013, 40, 1325–1332. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Diao, C.; Yang, X.; Yang, Z.; Liu, M.; Li, X.; Tang, H. ICP4-induced miR-101 attenuates HSV-1 replication. Sci. Rep. 2016, 6, 23205. [Google Scholar] [CrossRef] [PubMed]
- Ashraf, U.; Zhu, B.; Ye, J.; Wan, S.; Nie, Y.; Chen, Z.; Cui, M.; Wang, C.; Duan, X.; Zhang, H. MicroRNA-19b-3p modulates Japanese encephalitis virus-mediated inflammation via targeting RNF11. J. Virol. 2016, 90, 4780–4795. [Google Scholar] [CrossRef] [PubMed]
- Thounaojam, M.C.; Kundu, K.; Kaushik, D.K.; Swaroop, S.; Mahadevan, A.; Shankar, S.K.; Basu, A. MicroRNA 155 regulates Japanese encephalitis virus-induced inflammatory response by targeting Src homology 2-containing inositol phosphatase 1. J. Virol. 2014, 88, 4798–4810. [Google Scholar] [CrossRef] [PubMed]
- Sharma, N.; Verma, R.; Kumawat, K.L.; Basu, A.; Singh, S.K. miR-146a Suppresses cellular immune response during Japanese encephalitis virus JaOArS982 strain infection in human microglial cells. J. Neuroinflamm. 2015, 12, 30. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Xu, X.; Zhao, S.; Gong, Z.; Liu, P.; Guan, W.; He, X.; Wang, T.; Peng, T.; Teng, J.; et al. The expression and significance of the plasma let-7 family in anti-N-methyl-d-aspartate receptor encephalitis. J. Mol. Neurosci. 2015, 56, 531–539. [Google Scholar] [CrossRef] [PubMed]
- Li, J.M.; Kao, K.C.; Li, L.F.; Yang, T.M.; Wu, C.P.; Horng, Y.M.; Jia, W.W.; Yang, C.T. MicroRNA-145 regulates oncolytic herpes simplex virus-1 for selective killing of human non-small cell lung cancer cells. Virol. J. 2013, 10, 241. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Ye, J.; Ashraf, U.; Li, Y.; Wei, S.; Wan, S.; Zohaib, A.; Song, Y.; Chen, H.; Cao, S. MicroRNA-33a-5p modulates Japanese encephalitis virus replication by targeting eukaryotic translation elongation factor 1A1. J. Virol. 2016, 90, 3722–3734. [Google Scholar] [CrossRef] [PubMed]
- Bunai, Y.; Ishii, A.; Akaza, K.; Nagai, A.; Nishida, N.; Yamaguchi, S. A case of sudden death after Japanese encephalitis vaccination. Leg. Med. (Tokyo) 2015, 17, 279–282. [Google Scholar] [CrossRef] [PubMed]
- Plesner, A.M.; Arlien-Soborg, P.; Herning, M. Neurological complications to vaccination against Japanese encephalitis. Eur. J. Neurol. 1998, 5, 479–485. [Google Scholar] [CrossRef] [PubMed]
- Ohtaki, E.; Murakami, Y.; Komori, H.; Yamashita, Y.; Matsuishi, T. Acute disseminated encephalomyelitis after Japanese B encephalitis vaccination. Pediatr. Neurol. 1992, 8, 137–139. [Google Scholar] [CrossRef]
- Ohya, T.; Nagamitsu, S.; Yamashita, Y.; Matsuishi, T. Serial magnetic resonance imaging and single photon emission computed tomography study of acute disseminated encephalomyelitis patient after Japanese encephalitis vaccination. Kurume Med. J. 2007, 54, 95–99. [Google Scholar] [CrossRef] [PubMed]
- Nei, M.; Kumar, S.; Ebrary Inc. Molecular Evolution and Phylogenetics; Oxford University Press: Oxford, UK; New York, NY, USA, 2000. [Google Scholar]
- Kozomara, A.; Griffiths-Jones, S. miRBase: Annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014, 42, D68–D73. [Google Scholar] [CrossRef] [PubMed]
- MATLAB and Bioinformatics Toolbox Release 2012b; The MathWorks, Inc.: Natick, MA, USA, 2012.
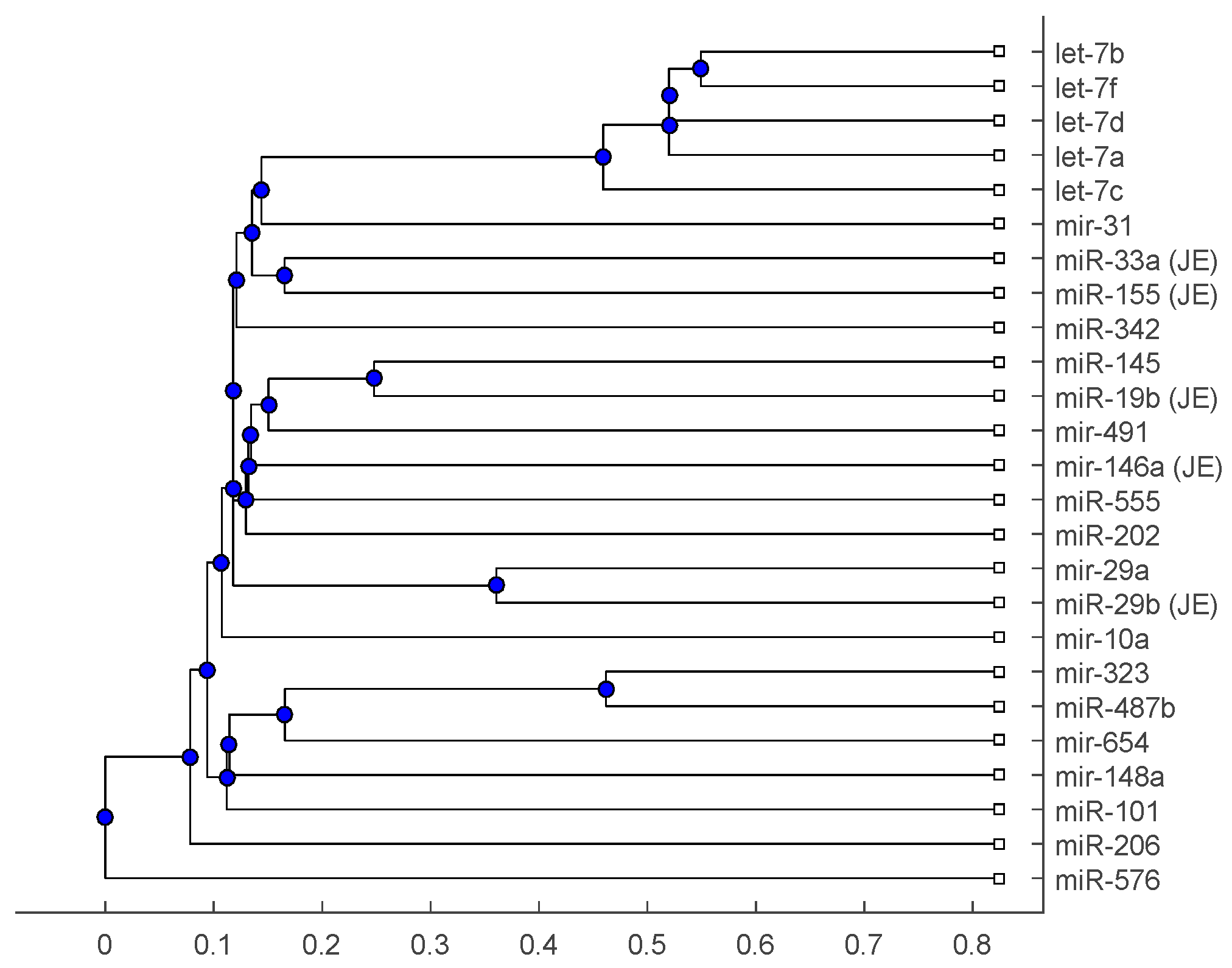
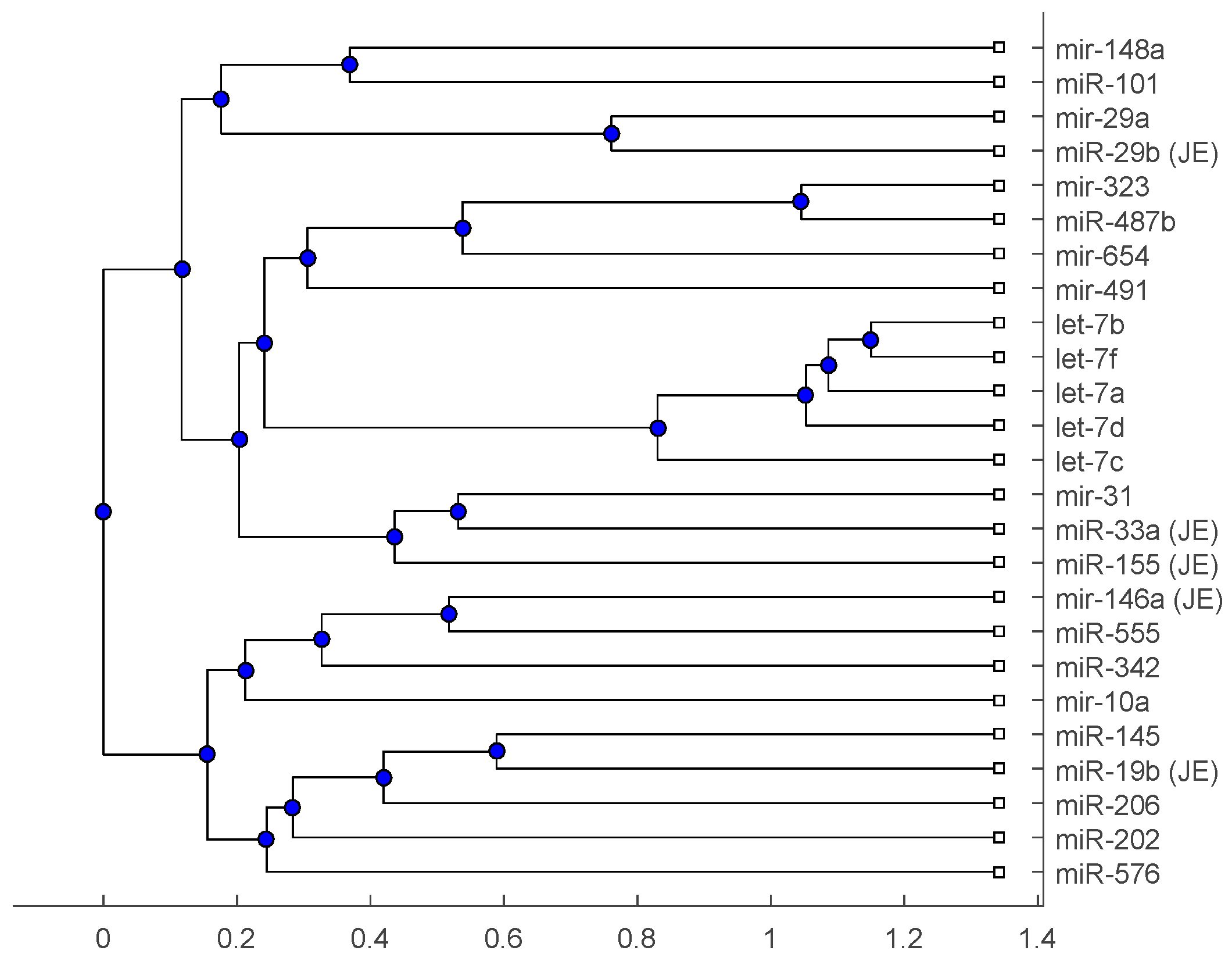
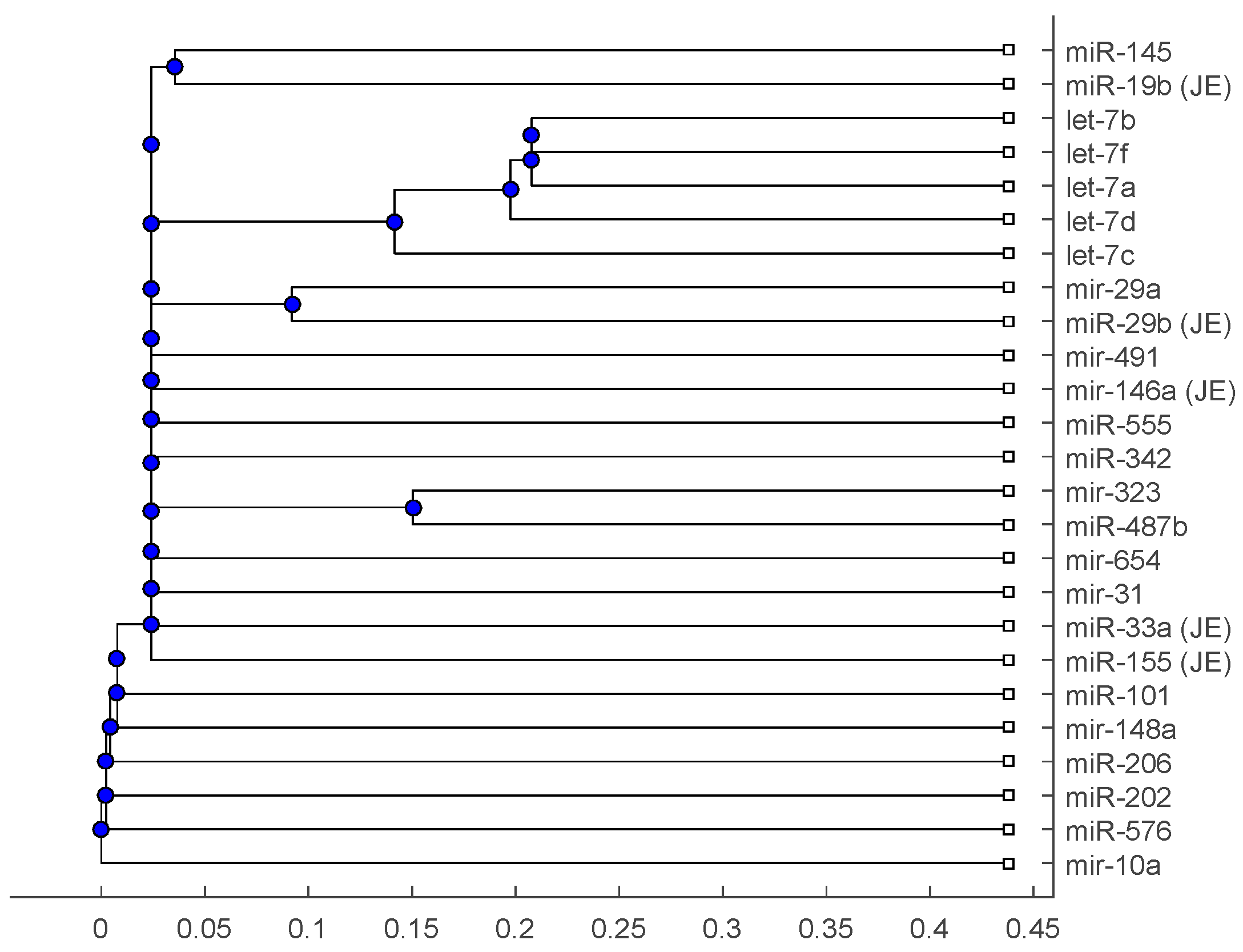

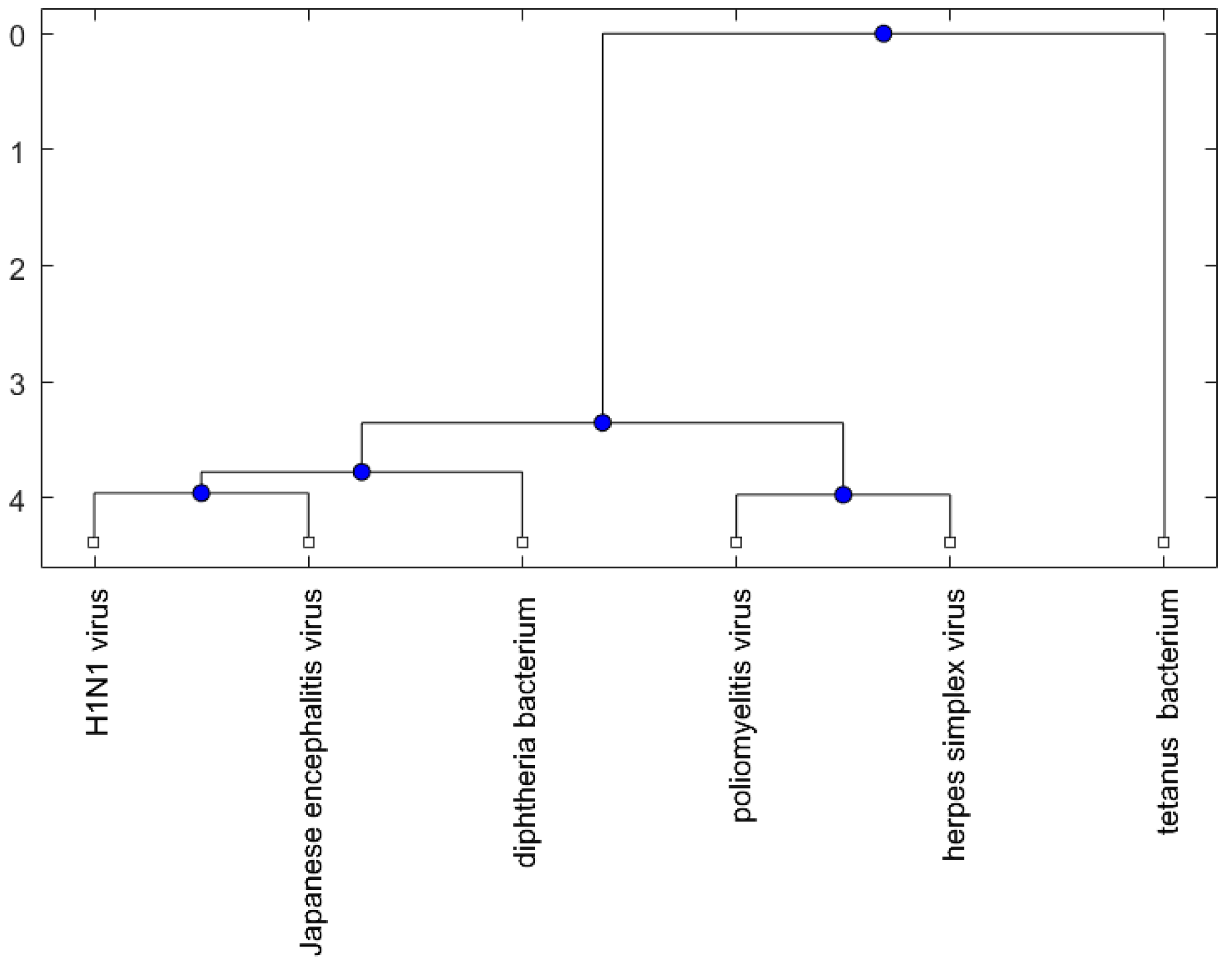
| Vaccine or Encephalitis | MircoRNA | References |
|---|---|---|
| H1N1 | miR-323, miR-491, miR-654, miR-10a, let-7c, let-7f, miR-31, miR-29a, miR-148a, miR-146a | [19,20,21,22] |
| pertussis | miR-202, miR-342, miR-206, miR-487b, miR-576 | [23] |
| poliomyelitis | miR-555 | [5] |
| herpes simplex virus | miR-145, miR-101 | [21,24] |
| Japanese encephalitis virus | miR-19b-3p, miR-33a-5p, miR-155, miR-29b, miR-146a | [25,26,27] |
| Anti-NMDA receptor encephalitis | let-7a, let-7b, let-7d, and let-7f | [28] |
| Vaccine | Virus (or Bacterium) Sequence Accession Number |
|---|---|
| H1N1 | AF250365 |
| Tetanus | X04436 |
| Diphtheria | K01722 |
| Poliomyelitis | KC880521 |
| Herpes simplex virus | M38699.1 |
| Japanese encephalitis virus | FJ938222 |
© 2017 by the author; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, H. Anti-NMDA Receptor Encephalitis and Vaccination. Int. J. Mol. Sci. 2017, 18, 193. https://doi.org/10.3390/ijms18010193
Wang H. Anti-NMDA Receptor Encephalitis and Vaccination. International Journal of Molecular Sciences. 2017; 18(1):193. https://doi.org/10.3390/ijms18010193
Chicago/Turabian StyleWang, Hsiuying. 2017. "Anti-NMDA Receptor Encephalitis and Vaccination" International Journal of Molecular Sciences 18, no. 1: 193. https://doi.org/10.3390/ijms18010193
APA StyleWang, H. (2017). Anti-NMDA Receptor Encephalitis and Vaccination. International Journal of Molecular Sciences, 18(1), 193. https://doi.org/10.3390/ijms18010193






