Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse
Abstract
:1. Introduction
2. Results
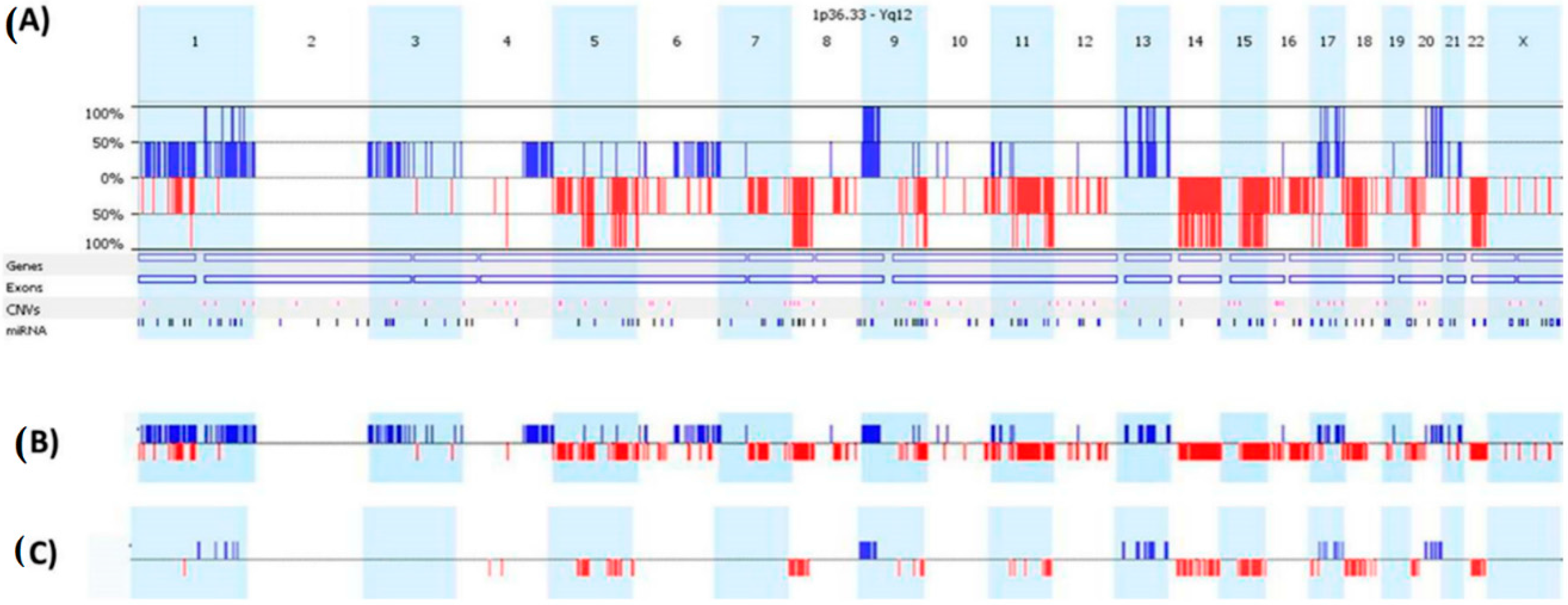
2.1. CRC aCGH Profile
2.2. Expression Analysis of Candidate Oncogenes
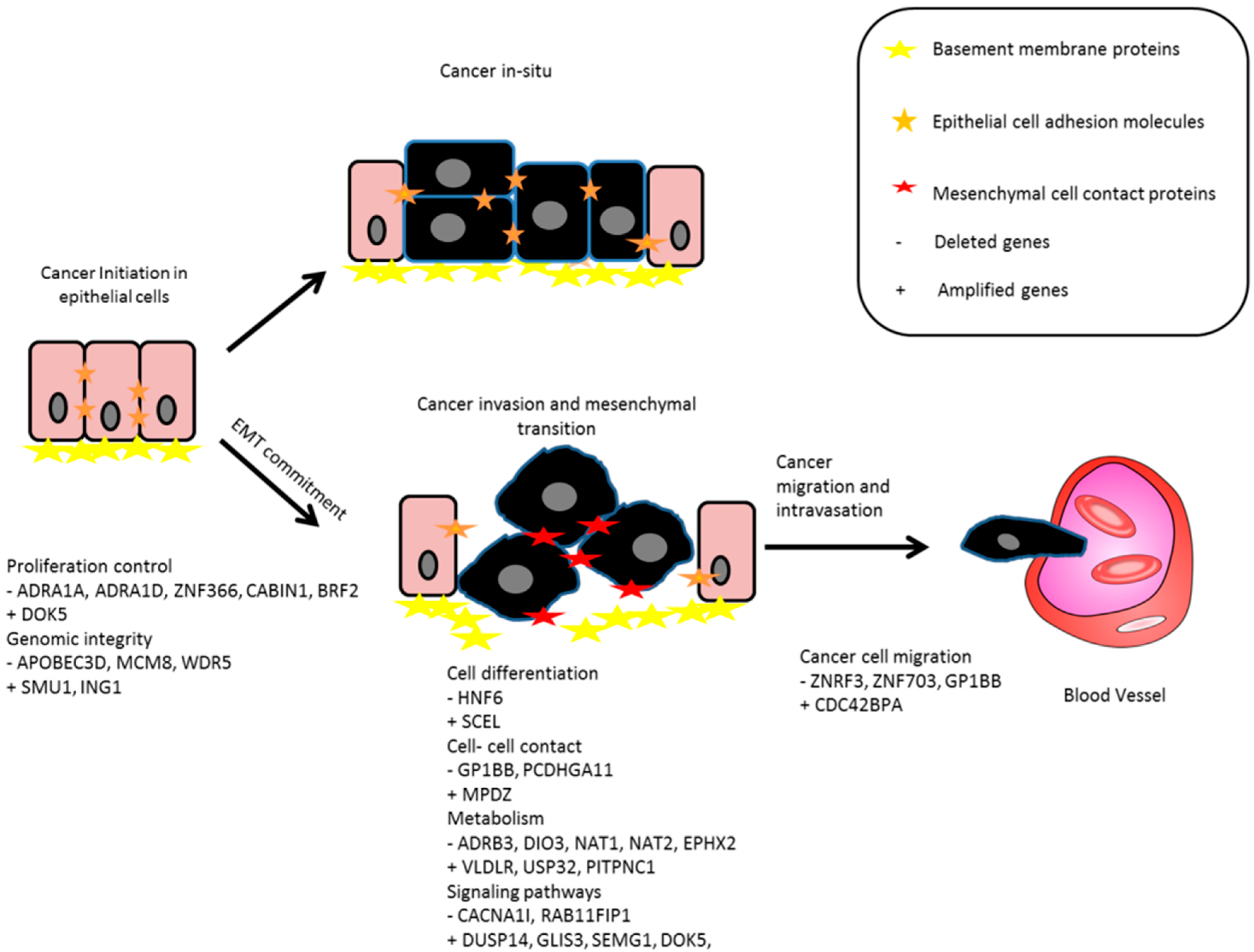
3. Discussion
4. Materials and Methods
4.1. CRC Samples
4.2. Microsatellite Instability Analysis
4.3. Genomic Landscaping Using aCGH
4.4. Data Clustering and Statistical Analysis
4.5. Data Preprocessing of Affymetrix Microarray Gene Expression
4.6. Estimation of Epithelial-Mesenchymal Transition Score
4.7. Statistical Analysis
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| aCGH | Array comparative genomic hybridization |
| CRC | Colorectal Cancer |
| CAN | Copy number aberration |
| OS | Overall survival |
| DFS | Disease free survival |
| EMT | Epithelial mesenchymal transition |
| MSI | Microsatellite instable |
| MSS | Microsatellite stable |
| MMR | Mismatch repair |
| STAC | Significance testing for aberrant copy number |
| DLRS | Derivative of log ratio spread |
| CNG | Copy number gain |
| CNL | Copy number loss |
| FFPE | Formalin-fixed paraffin embedded |
| FASST2 | Fast Adaptive State Segmentation Technique |
| SPPS | survival predictive power statistical |
| BinReg | Binary regression |
| SAM | Significance analysis of microarray |
| ROC | Receiver-operating characteristics curve |
References
- Dotan, E.; Cohen, S.J. Challenges in the management of stage II colon cancer. Semin. Oncol. 2011, 38, 511–520. [Google Scholar] [CrossRef] [PubMed]
- Ogino, S.; Chan, A.T.; Fuchs, C.S.; Giovannucci, E. Molecular pathological epidemiology of colorectal neoplasia: An emerging transdisciplinary and interdisciplinary field. Gut 2011, 60, 397–411. [Google Scholar] [CrossRef] [PubMed]
- Kahlenberg, M.S.; Sullivan, J.M.; Witmer, D.D.; Petrelli, N.J. Molecular prognostics in colorectal cancer. Surg. Oncol. 2003, 12, 173–186. [Google Scholar] [CrossRef]
- Compton, C.C. Colorectal carcinoma: Diagnostic, prognostic, and molecular features. Mod. Pathol. 2003, 16, 376–388. [Google Scholar] [CrossRef] [PubMed]
- Berg, M.; Nordgaard, O.; Korner, H.; Oltedal, S.; Smaaland, R.; Soreide, J.A.; Soreide, K. Molecular subtypes in stage II–III colon cancer defined by genomic instability: Early recurrence-risk associated with a high copy-number variation and loss of RUNX3 and CDKN2A. PLoS ONE 2015, 10, e0122391. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Carvalho, C.M.; Hastings, P.J.; Lupski, J.R. Mechanisms for recurrent and complex human genomic rearrangements. Curr. Opin. Genet. Dev. 2012, 22, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Sanz-Pamplona, R.; Berenguer, A.; Cordero, D.; Riccadonna, S.; Sole, X.; Crous-Bou, M.; Guino, E.; Sanjuan, X.; Biondo, S.; Soriano, A.; et al. Clinical value of prognosis gene expression signatures in colorectal cancer: A systematic review. PLoS ONE 2012, 7, e48877. [Google Scholar] [CrossRef] [PubMed]
- Shi, M.; Beauchamp, R.D.; Zhang, B. A network-based gene expression signature informs prognosis and treatment for colorectal cancer patients. PLoS ONE 2012, 7, e41292. [Google Scholar] [CrossRef] [PubMed]
- Wellcome Trust Sanger Institute. Available online: http://www.sanger.ac.uk/research/areas/humangenetics/CNA/ (accessed on 20 June 2014).
- Database of Genomic Variants. Available online: http://dgv.tcag.ca/dgv/app/chromosome?ref=NCBI36/hg18 (accessed on 11 July 2014).
- Markle, B.; May, E.J.; Majumdar, A.P. Do nutraceutics play a role in the prevention and treatment of colorectal cancer? Cancer Metastasis Rev. 2010, 29, 395–404. [Google Scholar] [CrossRef] [PubMed]
- Jones, S.; Chen, W.D.; Parmigiani, G.; Diehl, F.; Beerenwinkel, N.; Antal, T.; Traulsen, A.; Nowak, M.A.; Siegel, C.; Velculescu, V.E.; et al. Comparative lesion sequencing provides insights into tumor evolution. Proc. Natl. Acad. Sci. USA 2008, 105, 4283–4288. [Google Scholar] [CrossRef] [PubMed]
- Kim, S. New and emerging factors in tumorigenesis: An overview. Cancer Manag. Res. 2015, 7, 225–239. [Google Scholar] [CrossRef] [PubMed]
- Shull, A.Y.; Clendenning, M.L.; Ghoshal-Gupta, S.; Farrell, C.L.; Vangapandu, H.V.; Dudas, L.; Wilkerson, B.J.; Buckhaults, P.J. Somatic mutations, allele loss, and DNA methylation of the Cub and Sushi Multiple Domains 1 (CSMD1) gene reveals association with early age of diagnosis in colorectal cancer patients. PLoS ONE 2013, 8, e58731. [Google Scholar] [CrossRef] [PubMed]
- Udali, S.; Guarini, P.; Ruzzenente, A.; Ferrarini, A.; Guglielmi, A.; Lotto, V.; Tononi, P.; Pattini, P.; Moruzzi, S.; Campagnaro, T.; et al. DNA methylation and gene expression profiles show novel regulatory pathways in hepatocellular carcinoma. Clin. Epigenet. 2015, 7, 43. [Google Scholar] [CrossRef] [PubMed]
- Yi, J.K.; Kim, H.J.; Yu, D.H.; Park, S.J.; Shin, M.J.; Yuh, H.S.; Bae, K.B.; Ji, Y.R.; Kim, N.R.; Park, S.J.; et al. Regulation of inflammatory responses and fibroblast-like synoviocyte apoptosis by calcineurin-binding protein 1 in mice with collagen-induced arthritis. Arthritis Rheum. 2012, 64, 2191–2200. [Google Scholar] [CrossRef] [PubMed]
- Tsai, J.H.; Yang, J. Epithelial-mesenchymal plasticity in carcinoma metastasis. Genes Dev. 2013, 27, 2192–2206. [Google Scholar] [CrossRef] [PubMed]
- Collette, K.M.; Zhou, X.D.; Amoth, H.M.; Lyons, M.J.; Papay, R.S.; Sens, D.A.; Perez, D.M.; Doze, V.A. Long-term α1B-adrenergic receptor activation shortens lifespan, while α1A-adrenergic receptor stimulation prolongs lifespan in association with decreased cancer incidence. Age 2014, 36, 9675. [Google Scholar] [CrossRef] [PubMed]
- Ansems, M.; Hontelez, S.; Karthaus, N.; Span, P.N.; Adema, G.J. Crosstalk and DC-script: Expanding nuclear receptor modulation. Biochim. Biophys. Acta 2010, 1806, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Hulme, A.E.; Bogerd, H.P.; Cullen, B.R.; Moran, J.V. Selective inhibition of Alu retrotransposition by APOBEC3G. Gene 2007, 390, 199–205. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Long, D.T.; Lee, K.Y.; Abbas, T.; Shibata, E.; Negishi, M.; Luo, Y.; Schimenti, J.C.; Gambus, A.; Walter, J.C.; et al. The MCM8-MCM9 complex promotes RAD51 recruitment at DNA damage sites to facilitate homologous recombination. Mol. Cell. Biol. 2013, 33, 1632–1644. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.Z.; Tsai, Y.P.; Yang, M.H.; Huang, C.H.; Chang, S.Y.; Chang, C.C.; Teng, S.C.; Wu, K.J. Interplay between HDAC3 and WDR5 is essential for hypoxia-induced epithelial-mesenchymal transition. Mol. Cell. 2011, 43, 811–822. [Google Scholar] [CrossRef] [PubMed]
- Perrault, C.; Mangin, P.; Santer, M.; Baas, M.J.; Moog, S.; Cranmer, S.L.; Pikovski, I.; Williamson, D.; Jackson, S.P.; Cazenave, J.P.; et al. Role of the intracellular domains of GPIB in controlling the adhesive properties of the platelet GPIB/V/IX complex. Blood 2003, 101, 3477–3484. [Google Scholar] [CrossRef] [PubMed]
- Waha, A.; Guntner, S.; Huang, T.H.; Yan, P.S.; Arslan, B.; Pietsch, T.; Wiestler, O.D.; Waha, A. Epigenetic silencing of the protocadherin family member PCDH-γ-A11 in astrocytomas. Neoplasia 2005, 7, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Sibilio, A.; Ambrosio, R.; Bonelli, C.; de Stefano, M.A.; Torre, V.; Dentice, M.; Salvatore, D. Deiodination in cancer growth: The role of type III deiodinase. Minerv. Endocrinol. 2012, 37, 315–327. [Google Scholar]
- Zhou, Y.; Lan, J.; Wang, W.; Shi, Q.; Lan, Y.; Cheng, Z.; Guan, H. ZNRF3 acts as a tumour suppressor by the Wnt signalling pathway in human gastric adenocarcinoma. J. Mol. Histol. 2013, 44, 555–563. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Holterman, A.X. Pathophysiologic role of hepatocyte nuclear factor 6. Cell Signal. 2012, 24, 9–16. [Google Scholar] [CrossRef] [PubMed]
- Vega, S.; Morales, A.V.; Ocana, O.H.; Valdes, F.; Fabregat, I.; Nieto, M.A. Snail blocks the cell cycle and confers resistance to cell death. Genes Dev. 2004, 18, 1131–1143. [Google Scholar] [CrossRef] [PubMed]
- Tsai, J.H.; Donaher, J.L.; Murphy, D.A.; Chau, S.; Yang, J. Spatiotemporal regulation of epithelial-mesenchymal transition is essential for squamous cell carcinoma metastasis. Cancer Cell 2012, 22, 725–736. [Google Scholar] [CrossRef] [PubMed]
- Ren, L.; Liu, Y.; Guo, L.; Wang, H.; Ma, L.; Zeng, M.; Shao, X.; Yang, C.; Tang, Y.; Wang, L.; et al. Loss of SMU1 function de-represses DNA replication and over-activates ATR-dependent replication checkpoint. Biochem. Biophys. Res. Commun. 2013, 436, 192–198. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Chen, Y.; Yao, N.; Liu, H.; Wang, Z. MicroRNA LET-7B suppresses human gastric cancer malignancy by targeting ING1. Cancer Gene Ther. 2015, 22, 122–129. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Wu, F.; Chen, F.M.; Tian, J.; Qu, S. Variations of very low-density lipoprotein receptor subtype expression in gastrointestinal adenocarcinoma cells with various differentiations. World J. Gastroenterol. 2005, 11, 2817–2821. [Google Scholar] [CrossRef] [PubMed]
- Akhavantabasi, S.; Akman, H.B.; Sapmaz, A.; Keller, J.; Petty, E.M.; Erson, A.E. USP32 is an active, membrane-bound ubiquitin protease overexpressed in breast cancers. Mamm. Genome Off. J. Int. Mamm. Genome Soc. 2010, 21, 388–397. [Google Scholar] [CrossRef] [PubMed]
- Garner, K.; Hunt, A.N.; Koster, G.; Somerharju, P.; Groves, E.; Li, M.; Raghu, P.; Holic, R.; Cockcroft, S. Phosphatidylinositol transfer protein, cytoplasmic 1 (PITPNC1) binds and transfers phosphatidic acid. J. Biol. Chem. 2012, 287, 32263–32276. [Google Scholar] [CrossRef] [PubMed]
- Eckert, M.A.; Lwin, T.M.; Chang, A.T.; Kim, J.; Danis, E.; Ohno-Machado, L.; Yang, J. TWIST1-induced invadopodia formation promotes tumor metastasis. Cancer Cell. 2011, 19, 372–386. [Google Scholar] [CrossRef] [PubMed]
- Pignatelli, J.; Tumbarello, D.A.; Schmidt, R.P.; Turner, C.E. Hic-5 promotes invadopodia formation and invasion during TGF-β-induced epithelial-mesenchymal transition. J. Cell Biol. 2012, 197, 421–437. [Google Scholar] [CrossRef] [PubMed]
- Bosso, M.; Al-Mulla, F. Whole genome amplification of DNA extracted from ffpe tissues. Methods Mol. Biol. 2011, 724, 161–180. [Google Scholar] [PubMed]
- Al-Mulla, F. Microarray-based CGH and copy number analysis of FFPE samples. Methods Mol. Biol. 2011, 724, 131–145. [Google Scholar] [PubMed]
- Diskin, S.J.; Eck, T.; Greshock, J.; Mosse, Y.P.; Naylor, T.; Stoeckert, C.J., Jr.; Weber, B.L.; Maris, J.M.; Grant, G.R. STAC: A method for testing the significance of DNA copy number aberrations across multiple array-CGH experiments. Genome Res. 2006, 16, 1149–1158. [Google Scholar] [CrossRef] [PubMed]
- Johnson, W.E.; Li, C.; Rabinovic, A. Adjusting batch effects in microarray expression data using empirical bayes methods. Biostatistics 2007, 8, 118–127. [Google Scholar] [CrossRef] [PubMed]
- Tan, T.Z.; Miow, Q.H.; Miki, Y.; Noda, T.; Mori, S.; Huang, R.Y.; Thiery, J.P. Epithelial-mesenchymal transition spectrum quantification and its efficacy in deciphering survival and drug responses of cancer patients. EMBO Mol. Med. 2014, 6, 1279–1293. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.M.; Dedhar, S.; Kalluri, R.; Thompson, E.W. The epithelial-mesenchymal transition: New insights in signaling, development, and disease. J. Cell Biol. 2006, 172, 973–981. [Google Scholar] [CrossRef] [PubMed]
- Schweizer, J.; Bowden, P.E.; Coulombe, P.A.; Langbein, L.; Lane, E.B.; Magin, T.M.; Maltais, L.; Omary, M.B.; Parry, D.A.; Rogers, M.A.; et al. New consensus nomenclature for mammalian keratins. J. Cell Biol. 2006, 174, 169–174. [Google Scholar] [CrossRef] [PubMed]
- Moll, R.; Divo, M.; Langbein, L. The human keratins: Biology and pathology. Histochem. Cell Biol. 2008, 129, 705–733. [Google Scholar] [CrossRef] [PubMed]
- Gatza, M.L.; Lucas, J.E.; Barry, W.T.; Kim, J.W.; Wang, Q.; Crawford, M.D.; Datto, M.B.; Kelley, M.; Mathey-Prevot, B.; Potti, A.; et al. A pathway-based classification of human breast cancer. Proc. Natl. Acad. Sci. USA 2010, 107, 6994–6999. [Google Scholar] [CrossRef] [PubMed]
- Verhaak, R.G.; Tamayo, P.; Yang, J.Y.; Hubbard, D.; Zhang, H.; Creighton, C.J.; Fereday, S.; Lawrence, M.; Carter, S.L.; Mermel, C.H.; et al. Prognostically relevant gene signatures of high-grade serous ovarian carcinoma. J. Clin. Investig. 2013, 123, 517–525. [Google Scholar] [CrossRef] [PubMed]
- Ballou, L.M.; Cross, M.E.; Huang, S.; McReynolds, E.M.; Zhang, B.X.; Lin, R.Z. Differential regulation of the phosphatidylinositol 3-kinase/akt and p70 s6 kinase pathways by the α(1a)-adrenergic receptor in rat-1 fibroblasts. J. Biol. Chem. 2000, 275, 4803–4809. [Google Scholar] [CrossRef] [PubMed]
- Kassahun, W.T.; Gunl, B.; Jonas, S.; Ungemach, F.R.; Abraham, G. Altered liver α1-adrenoceptor density and phospholipase c activity in the human hepatocellular carcinoma. Eur. J. Pharmacol. 2011, 670, 92–95. [Google Scholar] [CrossRef] [PubMed]
- Keffel, S.; Alexandrov, A.; Goepel, M.; Michel, M.C. α(1)-adrenoceptor subtypes differentially couple to growth promotion and inhibition in chinese hamster ovary cells. Biochem. Biophys. Res. Commun. 2000, 272, 906–911. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Cabrera, P.J.; Shi, T.; Yun, J.; McCune, D.F.; Rorabaugh, B.R.; Perez, D.M. Differential regulation of the cell cycle by α1-adrenergic receptor subtypes. Endocrinology 2004, 145, 5157–5167. [Google Scholar] [CrossRef] [PubMed]
- Vinci, M.C.; Bellik, L.; Filippi, S.; Ledda, F.; Parenti, A. Trophic effects induced by α1d-adrenoceptors on endothelial cells are potentiated by hypoxia. Am. J. Physiol. Heart Circ. Physiol. 2007, 293, H2140–H2147. [Google Scholar] [CrossRef] [PubMed]
- Shibata, K.; Katsuma, S.; Koshimizu, T.; Shinoura, H.; Hirasawa, A.; Tanoue, A.; Tsujimoto, G. Α 1-adrenergic receptor subtypes differentially control the cell cycle of transfected cho cells through a camp-dependent mechanism involving p27kip1. J. Biol. Chem. 2003, 278, 672–678. [Google Scholar] [CrossRef] [PubMed]
- Babol, K.; Przybylowska, K.; Lukaszek, M.; Pertynski, T.; Blasiak, J. An association between the trp64arg polymorphism in the beta3-adrenergic receptor gene and endometrial cancer and obesity. J. Exp. Clin. Cancer Res. 2004, 23, 669–674. [Google Scholar] [PubMed]
- Guay, S.P.; Brisson, D.; Lamarche, B.; Biron, S.; Lescelleur, O.; Biertho, L.; Marceau, S.; Vohl, M.C.; Gaudet, D.; Bouchard, L. Adrb3 gene promoter DNA methylation in blood and visceral adipose tissue is associated with metabolic disturbances in men. Epigenomics 2014, 6, 33–43. [Google Scholar] [CrossRef] [PubMed]
- Lackey, L.; Law, E.K.; Brown, W.L.; Harris, R.S. Subcellular localization of the apobec3 proteins during mitosis and implications for genomic DNA deamination. Cell Cycle 2013, 12, 762–772. [Google Scholar] [CrossRef] [PubMed]
- Stenglein, M.D.; Harris, R.S. Apobec3b and apobec3f inhibit l1 retrotransposition by a DNA deamination-independent mechanism. J. Biol. Chem. 2006, 281, 16837–16841. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Tian, H.; Yue, W.; Li, L.; Li, S.; Qi, L.; Hu, W.; Gao, C.; Si, L. Overexpression of tfiib-related factor 2 is significantly correlated with tumor angiogenesis and poor survival in patients with esophageal squamous cell cancer. Med. Oncol. 2013, 30, 553. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Lu, M.; Yue, W.; Li, L.; Li, S.; Gao, C.; Si, L.; Qi, L.; Hu, W.; Tian, H. Tfiib-related factor 2 is associated with poor prognosis of nonsmall cell lung cancer patients through promoting tumor epithelial-mesenchymal transition. BioMed Res. Int. 2014, 2014, 530786. [Google Scholar] [CrossRef] [PubMed]
- Yu, D.H.; Yi, J.K.; Park, S.J.; Kim, M.O.; Kim, H.J.; Yuh, H.S.; Bae, K.B.; Ji, Y.R.; Lee, H.S.; Lee, S.G.; et al. Tissue-specific expression of human calcineurin-binding protein 1 in mouse synovial tissue can suppress inflammatory arthritis. J. Interferon Cytokine Res. 2012, 32, 6–11. [Google Scholar] [CrossRef] [PubMed]
- Talavera, K.; Nilius, B. Biophysics and structure-function relationship of t-type ca2+ channels. Cell Calcium 2006, 40, 97–114. [Google Scholar] [CrossRef] [PubMed]
- Farrell, C.; Crimm, H.; Meeh, P.; Croshaw, R.; Barbar, T.; Vandersteenhoven, J.J.; Butler, W.; Buckhaults, P. Somatic mutations to csmd1 in colorectal adenocarcinomas. Cancer Biol. Ther. 2008, 7, 609–613. [Google Scholar] [CrossRef] [PubMed]
- Dentice, M.; Ambrosio, R.; Salvatore, D. Role of type 3 deiodinase in cancer. Exp. Opin. Ther. Targets 2009, 13, 1363–1373. [Google Scholar] [CrossRef] [PubMed]
- Newman, J.W.; Morisseau, C.; Harris, T.R.; Hammock, B.D. The soluble epoxide hydrolase encoded by epxh2 is a bifunctional enzyme with novel lipid phosphate phosphatase activity. Proc. Natl. Acad. Sci. USA 2003, 100, 1558–1563. [Google Scholar] [CrossRef] [PubMed]
- Diede, S.J.; Yao, Z.; Keyes, C.C.; Tyler, A.E.; Dey, J.; Hackett, C.S.; Elsaesser, K.; Kemp, C.J.; Neiman, P.E.; Weiss, W.A.; et al. Fundamental differences in promoter cpg island DNA hypermethylation between human cancer and genetically engineered mouse models of cancer. Epigenetics 2013, 8, 1254–1260. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, K.; Ishiai, M.; Horikawa, K.; Fukagawa, T.; Takata, M.; Takisawa, H.; Kanemaki, M.T. Mcm8 and mcm9 form a complex that functions in homologous recombination repair induced by DNA interstrand crosslinks. Mol. Cell 2012, 47, 511–522. [Google Scholar] [CrossRef] [PubMed]
- Lilla, C.; Verla-Tebit, E.; Risch, A.; Jager, B.; Hoffmeister, M.; Brenner, H.; Chang-Claude, J. Effect of nat1 and nat2 genetic polymorphisms on colorectal cancer risk associated with exposure to tobacco smoke and meat consumption. Cancer Epidemiol. Biomark. Prev. 2006, 15, 99–107. [Google Scholar] [CrossRef] [PubMed]
- Johansson, I.; Nilsson, C.; Berglund, P.; Lauss, M.; Ringner, M.; Olsson, H.; Luts, L.; Sim, E.; Thorstensson, S.; Fjallskog, M.L.; et al. Gene expression profiling of primary male breast cancers reveals two unique subgroups and identifies N-acetyltransferase-1 (NAT1) as a novel prognostic biomarker. Breast Cancer Res. 2012, 14, R31. [Google Scholar] [CrossRef] [PubMed]
- Cai, J.; Zhao, Y.; Zhu, C.L.; Li, J.; Huang, Z.H. The association of nat1 polymorphisms and colorectal carcinoma risk: Evidence from 20,000 subjects. Mol. Biol. Rep. 2012, 39, 7497–7503. [Google Scholar] [CrossRef] [PubMed]
- Zhuo, W.; Zhang, L.; Qiu, Z.; Cai, L.; Zhu, B.; Chen, Z. Association of nat2 polymorphisms with risk of colorectal adenomas: Evidence from 3,197 cases and 4,681 controls. Exp. Ther. Med. 2012, 4, 895–900. [Google Scholar] [CrossRef] [PubMed]
- Jacquemin, P.; Durviaux, S.M.; Jensen, J.; Godfraind, C.; Gradwohl, G.; Guillemot, F.; Madsen, O.D.; Carmeliet, P.; Dewerchin, M.; Collen, D.; et al. Transcription factor hepatocyte nuclear factor 6 regulates pancreatic endocrine cell differentiation and controls expression of the proendocrine gene NGN3. Mol. Cell. Biol. 2000, 20, 4445–4454. [Google Scholar] [CrossRef] [PubMed]
- Prevot, P.P.; Simion, A.; Grimont, A.; Colletti, M.; Khalaileh, A.; Van den Steen, G.; Sempoux, C.; Xu, X.; Roelants, V.; Hald, J.; et al. Role of the ductal transcription factors HNF6 and Sox9 in pancreatic acinar-to-ductal metaplasia. Gut 2012, 61, 1723–1732. [Google Scholar] [CrossRef] [PubMed]
- Lehner, F.; Kulik, U.; Klempnauer, J.; Borlak, J. The hepatocyte nuclear factor 6 (hnf6) and foxa2 are key regulators in colorectal liver metastases. FASEB J. 2007, 21, 1445–1462. [Google Scholar] [CrossRef] [PubMed]
- Cullis, D.N.; Philip, B.; Baleja, J.D.; Feig, L.A. Rab11-Fip2, an adaptor protein connecting cellular components involved in internalization and recycling of epidermal growth factor receptors. J. Biol. Chem. 2002, 277, 49158–49166. [Google Scholar] [CrossRef] [PubMed]
- Carson, B.P.; Del Bas, J.M.; Moreno-Navarrete, J.M.; Fernandez-Real, J.M.; Mora, S. The rab11 effector protein fip1 regulates adiponectin trafficking and secretion. PLoS ONE 2013, 8, e74687. [Google Scholar] [CrossRef] [PubMed]
- Gori, F.; Friedman, L.; Demay, M.B. Wdr5, a novel wd repeat protein, regulates osteoblast and chondrocyte differentiation in vivo. J. Musculoskelet. Neuronal Interact. 2005, 5, 338–339. [Google Scholar] [PubMed]
- Ang, Y.S.; Tsai, S.Y.; Lee, D.F.; Monk, J.; Su, J.; Ratnakumar, K.; Ding, J.; Ge, Y.; Darr, H.; Chang, B.; et al. Wdr5 mediates self-renewal and reprogramming via the embryonic stem cell core transcriptional network. Cell 2011, 145, 183–197. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Garcia, J.; Periyasamy, M.; Thomas, R.S.; Christian, M.; Leao, M.; Jat, P.; Kindle, K.B.; Heery, D.M.; Parker, M.G.; Buluwela, L.; et al. Znf366 is an estrogen receptor corepressor that acts through ctbp and histone deacetylases. Nucleic Acids Res. 2006, 34, 6126–6136. [Google Scholar] [CrossRef] [PubMed]
- Ansems, M.; Karthaus, N.; Hontelez, S.; Aalders, T.; Looman, M.W.; Verhaegh, G.W.; Schalken, J.A.; Adema, G.J. Dc-script: Ar and vdr regulator lost upon transformation of prostate epithelial cells. Prostate 2012, 72, 1708–1717. [Google Scholar] [CrossRef] [PubMed]
- Sircoulomb, F.; Nicolas, N.; Ferrari, A.; Finetti, P.; Bekhouche, I.; Rousselet, E.; Lonigro, A.; Adelaide, J.; Baudelet, E.; Esteyries, S.; et al. Znf703 gene amplification at 8p12 specifies luminal b breast cancer. EMBO Mol. Med. 2011, 3, 153–166. [Google Scholar] [CrossRef] [PubMed]
- Ma, F.; Bi, L.; Yang, G.; Zhang, M.; Liu, C.; Zhao, Y.; Wang, Y.; Wang, J.; Bai, Y.; Zhang, Y. Znf703 promotes tumor cell proliferation and invasion and predicts poor prognosis in patients with colorectal cancer. Oncol. Rep. 2014, 32, 1071–1077. [Google Scholar] [CrossRef] [PubMed]
- Warnatz, H.J.; Schmidt, D.; Manke, T.; Piccini, I.; Sultan, M.; Borodina, T.; Balzereit, D.; Wruck, W.; Soldatov, A.; Vingron, M.; et al. The btb and cnc homology 1 (BACH1) target genes are involved in the oxidative stress response and in control of the cell cycle. J. Biol. Chem. 2011, 286, 23521–23532. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, S.; Paterson, H.F.; Marshall, C.J. Cdc42-mrck and rho-rock signalling cooperate in myosin phosphorylation and cell invasion. Nat. Cell Biol. 2005, 7, 255–261. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Yue, J.; You, Y.; Yin, B.; Gong, Y.; Xu, C.; Qiang, B.; Yuan, J.; Liu, Y.; Peng, X. Dok5 is substrate of trkb and trkc receptors and involved in neurotrophin induced mapk activation. Cell Signal. 2006, 18, 1995–2003. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Zhang, J.; Liu, W.; Shu, P.; Yin, B.; Yuan, J.; Qiang, B.; Peng, X. Dok5 is involved in the signaling pathway of neurotrophin-3 against trkc-induced apoptosis. Neuro Lett. 2013, 553, 46–51. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Li, Q.; Chen, R.; Zhang, J.; Ran, Y.; He, X.; Li, S.; Shu, H.B. The dual-specificity phosphatase dusp14 negatively regulates tumor necrosis factor- and interleukin-1-induced nuclear factor-kappab activation by dephosphorylating the protein kinase tak1. J. Biol. Chem. 2013, 288, 819–825. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.Y.; Li, J.P.; Chiu, L.L.; Lan, J.L.; Chen, D.Y.; Chuang, H.C.; Huang, C.Y.; Tan, T.H. Dual-specificity phosphatase 14 (DUSP14/MKP6) negatively regulates tcr signaling by inhibiting tab1 activation. J. Immunol. 2014, 192, 1547–1557. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Prasad, M.; Lemon, W.J.; Hampel, H.; Wright, F.A.; Kornacker, K.; LiVolsi, V.; Frankel, W.; Kloos, R.T.; Eng, C.; et al. Gene expression in papillary thyroid carcinoma reveals highly consistent profiles. Proc. Natl. Acad. Sci. USA 2001, 98, 15044–15049. [Google Scholar] [CrossRef] [PubMed]
- Lichti-Kaiser, K.; ZeRuth, G.; Kang, H.S.; Vasanth, S.; Jetten, A.M. Gli-similar proteins: Their mechanisms of action, physiological functions, and roles in disease. Vitam. Horm. 2012, 88, 141–171. [Google Scholar] [PubMed]
- Adachi, M.; Hamazaki, Y.; Kobayashi, Y.; Itoh, M.; Tsukita, S.; Furuse, M. Similar and distinct properties of mupp1 and patj, two homologous PDZ domain-containing tight-junction proteins. Mol. Cell. Biol. 2009, 29, 2372–2389. [Google Scholar] [CrossRef] [PubMed]
- Ernkvist, M.; Luna Persson, N.; Audebert, S.; Lecine, P.; Sinha, I.; Liu, M.; Schlueter, M.; Horowitz, A.; Aase, K.; Weide, T.; et al. The amot/patj/syx signaling complex spatially controls rhoa gtpase activity in migrating endothelial cells. Blood 2009, 113, 244–253. [Google Scholar] [CrossRef] [PubMed]
- Hamazaki, Y.; Itoh, M.; Sasaki, H.; Furuse, M.; Tsukita, S. Multi-pdz domain protein 1 (mupp1) is concentrated at tight junctions through its possible interaction with claudin-1 and junctional adhesion molecule. J. Biol. Chem. 2002, 277, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Png, K.J.; Halberg, N.; Yoshida, M.; Tavazoie, S.F. A microrna regulon that mediates endothelial recruitment and metastasis by cancer cells. Nature 2012, 481, 190–194. [Google Scholar] [CrossRef] [PubMed]
- Fonseca-Sanchez, M.A.; Perez-Plasencia, C.; Fernandez-Retana, J.; Arechaga-Ocampo, E.; Marchat, L.A.; Rodriguez-Cuevas, S.; Bautista-Pina, V.; Arellano-Anaya, Z.E.; Flores-Perez, A.; Diaz-Chavez, J.; et al. Microrna-18b is upregulated in breast cancer and modulates genes involved in cell migration. Oncol. Rep. 2013, 30, 2399–2410. [Google Scholar] [PubMed]
- Chen, L.S.; Wei, J.B.; Zhou, Y.C.; Zhang, S.; Liang, J.L.; Cao, Y.F.; Tang, Z.J.; Zhang, X.L.; Gao, F. Genetic alterations and expression of inhibitor of growth 1 in human sporadic colorectal cancer. World J. Gastroenterol. 2005, 11, 6120–6124. [Google Scholar] [CrossRef] [PubMed]

| Patients’ Characteristics | Stage II Who Stayed Disease Free | Stage II with Local Recurrences | Stage II Who Relapsed with Distant Metastasis | p Value a | |
|---|---|---|---|---|---|
| Mean age in years | 64.4 | 75.5 | 75.4 | 0.004 b | |
| Sex | Male | 41 | 2 | 5 | 0.43 |
| Female | 37 | 5 | 6 | ||
| Total | 78 | 7 | 11 | ||
| Site | Right | 19 | 2 | 1 | 0.16 |
| Left | 28 | 1 | 9 | ||
| Rectum | 16 | 1 | 1 | ||
| Unknown | 15 | 3 | 0 | ||
| T-stage | T3 | 44 | 3 | 9 | 0.27 |
| T4 | 19 | 4 | 2 | ||
| Unknown | 15 | 0 | 0 | ||
| Differentiation | Well | 10 | 1 | 1 | 0.8 |
| Moderate | 56 | 5 | 9 | ||
| Poor | 5 | 1 | 0 | ||
| Unknown | 7 | 0 | 1 | ||
| MMR status | MSI | 14 | 1 | 0 | 0.45 |
| MSS | 59 | 5 | 9 | ||
| Unknown | 5 | 1 | 2 | ||
| Follow-up | Mean DFS | 9.5 years | 3.9 years | 3.08 years |
| Gene | Gene ID | aCGH CNA Event | %CNA Overlap with Normal | Cox’s OS (p-Value) | Cox’s DFS (p-Value) |
|---|---|---|---|---|---|
| ADRA1A | 148 | Loss | 0.42 | −0.40185, (0.152322) | −1.67582, (0.001948) |
| ADRA1D | 146 | Loss | 1.15 | −0.5968, (0.020507) | 0.066504, (0.880385) |
| ADRB3 | 155 | Loss | 0 | −1.21809, (0.002366) | −1.12818, (0.08601) |
| APOBEC3D | 140564 | Loss | 0 | −0.45803, (0.24853) | −1.56085, (0.0224) |
| BRF2 | 55290 | Loss | 0 | −0.45526, (0.02013) | 0.061887, (0.8515) |
| C20orf202 | 400831 | Loss | 0.89 | −0.80141, (0.000481) | −0.19907, (0.628292) |
| CABIN1 | 23523 | Loss | 4.02 | −0.47782, (0.0210) | −0.51601, (0.14935) |
| CACNA1I | 8911 | Loss | 0 | −0.69524, (0.00404) | −0.73386, (0.082677) |
| CSMD1 | 64478 | Loss | 0 | −0.01578, (0.956357) | −1.30789, (0.01948) |
| DIO3 | 1735 | Loss | 1.76 | −0.32561, (0.041777) | −0.00829, (0.973719) |
| EPHX2 | 2053 | Loss | 0.42 | 0.036582, (0.61411) | −0.30515, (0.0099) |
| FAM83F | 113828 | Loss | 0 | −0.13746, (0.287004) | −0.72925, (0.00085) |
| GP1BB | 2812 | Loss | 0 | −0.56827, (0.000246) | −0.19289, (0.495577) |
| KIAA1656 | 85371 | Loss | 0 | −0.60241, (0.01922) | −1.02118, (0.032326) |
| LOC339593 | 339593 | Loss | 0.17 | 0.29021, (0.385267) | −1.12678, (0.047789) |
| MCM8 | 84515 | Loss | 0 | 0.043419, (0.614947) | −0.31948, (0.034203) |
| NAT1 | 9 | Loss | 0 | 0.028159, (0.713606) | −0.36337, (0.007197) |
| NAT2 | 10 | Loss | 0 | −0.09059, (0.160865) | −0.30169, (0.003804) |
| HNF6 | 3175 | Loss | 0 | −0.66924, (0.017955) | −0.51152, (0.31) |
| PCDHGA11 | 56105 | Loss | 0 | −0.85799, (0.001447) | −0.577, (0.33928) |
| RAB11FIP1 | 80223 | Loss | 0 | −0.38036, (5.55× 10−05) | −0.02196, (0.87) |
| SPAG11A | 653423 | Loss | 68.65 | −0.81665, (0.001447) | −0.13298, (0.57) |
| SIRPD | 128646 | Loss | 0.89 | −0.89166, (0.014166) | −0.13392, (0.82) |
| TEX43 | 389320 | Loss | 0 | −0.46611, (0.175471) | −0.19907, (0.01162) |
| TOP1P2 | 7152 | Loss | 0 | −1.13965, (0.005692) | 0.599602, (0.32) |
| WDR5 | 11091 | Loss | 0 | −0.0007, (0.995962) | −0.52505, (0.024307) |
| ZNF366 | 167465 | Loss | 0.07 | −0.76274, (0.01779) | −0.93429, (0.09208) |
| ZNF703 | 80139 | Loss | 0 | −0.32601, (0.000744) | −0.14954, (0.366064) |
| ZNRF3 | 84133 | Loss | 0 | −0.03445, (0.605797) | −0.33884, (0.002065) |
| ANXA2P2 | 304 | Gain | 0 | 0.478663, (0.003259) | 0.997702, (0.000504) |
| CDC42BPA | 8476 | Gain | 0 | 0.032966, (0.690294) | 0.401718, (0.005473) |
| DOK5 | 55816 | Gain | 0.57 | 0.105512, (0.1) | 0.577502, (0.002218) |
| DUSP14 | 11072 | Gain | 0.70 | 0.207902, (0.047604) | 0.709569, (0.000159) |
| GLIS3 | 169792 | Gain | 1.25 | 0.0916, (0.210004) | 0.397791, (0.000566) |
| ING1 | 3621 | Gain | 0 | 0.461934, (0.020587) | 0.127394, (0.710857) |
| MPDZ | 8777 | Gain | 0 | 0.079568, (0.406857) | 0.657405, (3.98 × 10−5) |
| PITPNC1 | 26207 | Gain | 0 | 0.257758, (0.029048) | 0.418362, (0.026674) |
| SCEL | 8796 | Gain | 0 | 0.081094, (0.202236) | 0.301113, (0.000357) |
| SEMG1 | 6406 | Gain | 0 | 0.098826, (0.043099) | 0.153704, (0.061048) |
| SMU1 | 55234 | Gain | 0.73 | 0.36857, (0.008896) | 0.016381, (0.942047) |
| USP32 | 84669 | Gain | 0 | 0.162429, (0.221638) | 0.586012, (0.01236) |
| VLDLR | 7436 | Gain | 0.14 | −0.00629, (0.92868) | 0.323607, (0.004972) |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Al-Temaimi, R.A.; Tan, T.Z.; Marafie, M.J.; Thiery, J.P.; Quirke, P.; Al-Mulla, F. Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse. Int. J. Mol. Sci. 2016, 17, 598. https://doi.org/10.3390/ijms17050598
Al-Temaimi RA, Tan TZ, Marafie MJ, Thiery JP, Quirke P, Al-Mulla F. Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse. International Journal of Molecular Sciences. 2016; 17(5):598. https://doi.org/10.3390/ijms17050598
Chicago/Turabian StyleAl-Temaimi, Rabeah A., Tuan Zea Tan, Makia J. Marafie, Jean Paul Thiery, Philip Quirke, and Fahd Al-Mulla. 2016. "Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse" International Journal of Molecular Sciences 17, no. 5: 598. https://doi.org/10.3390/ijms17050598
APA StyleAl-Temaimi, R. A., Tan, T. Z., Marafie, M. J., Thiery, J. P., Quirke, P., & Al-Mulla, F. (2016). Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse. International Journal of Molecular Sciences, 17(5), 598. https://doi.org/10.3390/ijms17050598







