Prognostic Value of MicroRNAs in Preoperative Treated Rectal Cancer
Abstract
:1. Introduction
2. Results
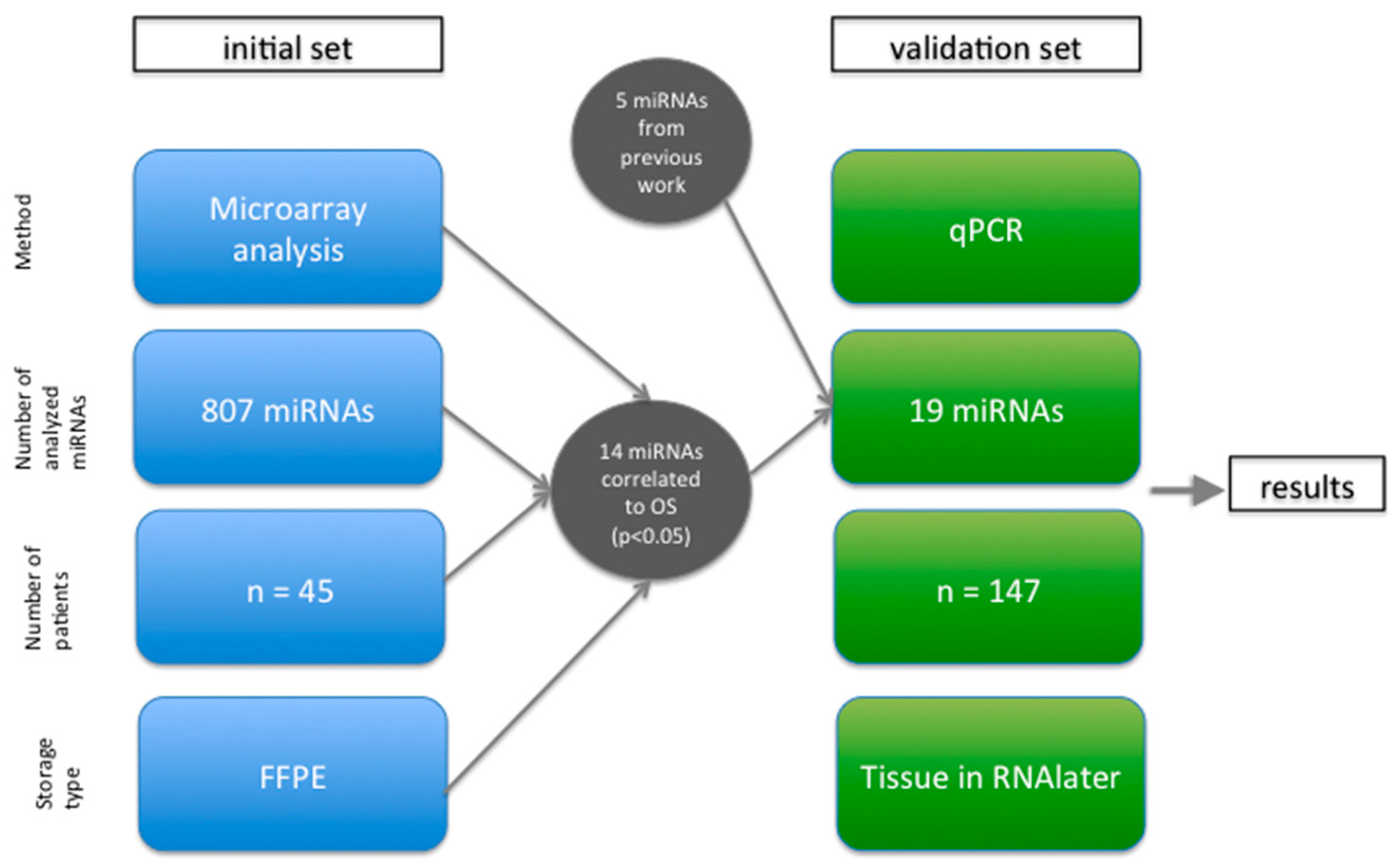
2.1. Microarray Analysis Identified 14 miRNAs Significantly Associated with Overall Survival
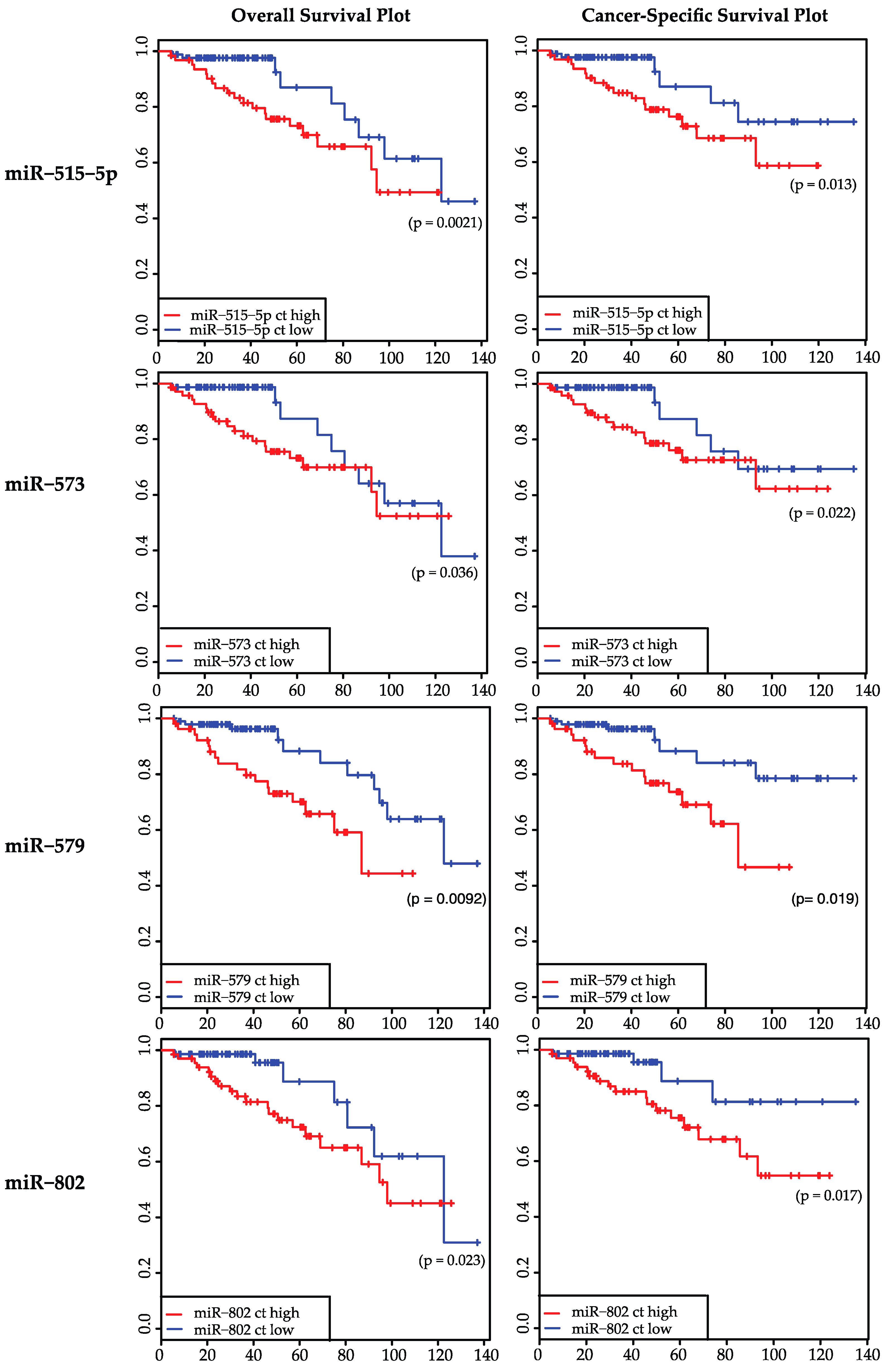
2.2. Expression Levels of 19 miRNAs Were Analyzed in 147 Samples: miR-515-5p, miR-573, miR-579, and miR-802 Were Significantly Correlated to Overall Survival and Cancer-Specific Survival
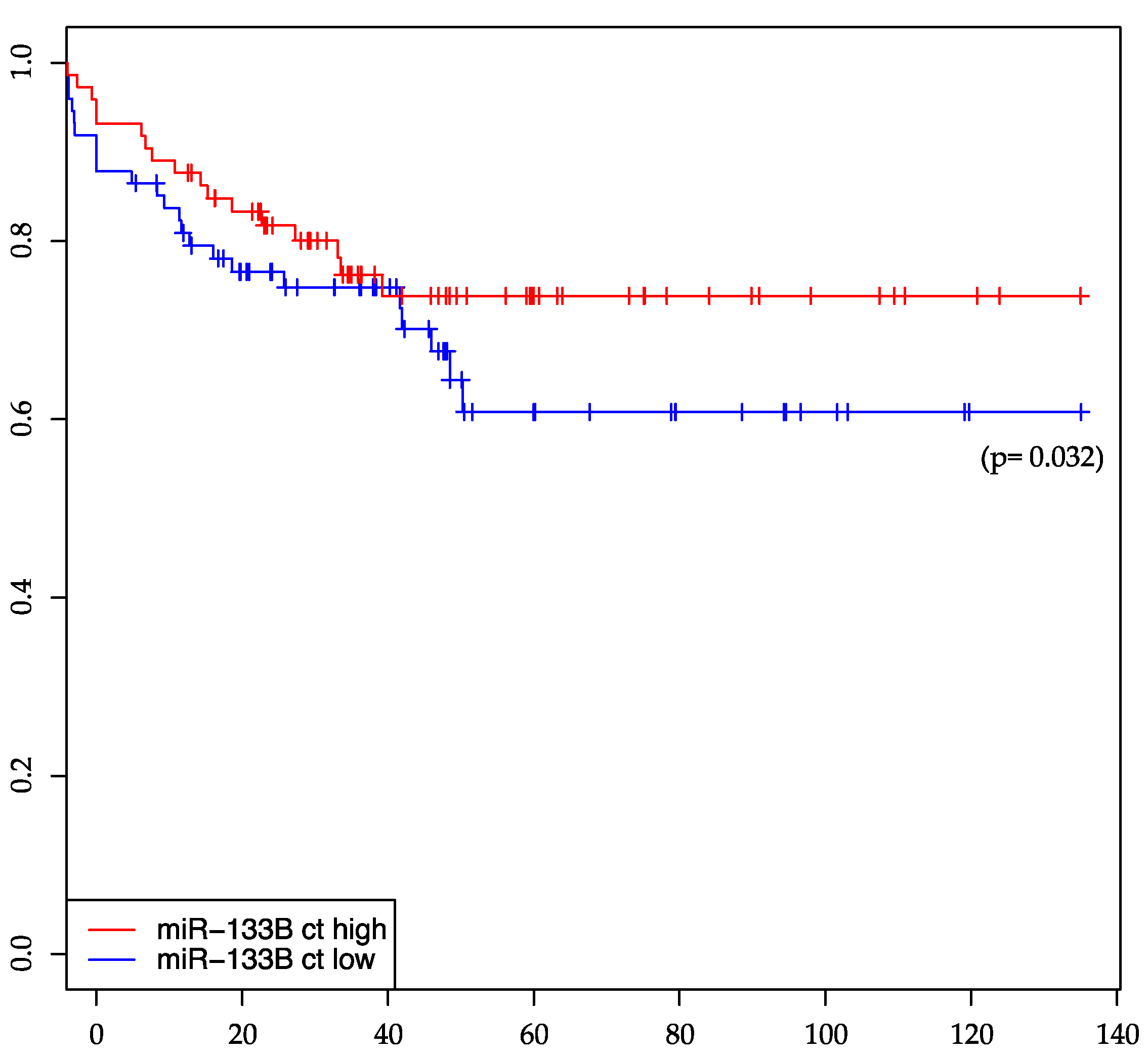
2.3. miR-133b Is Significantly Associated with Distant-Metastasis-Free Survival
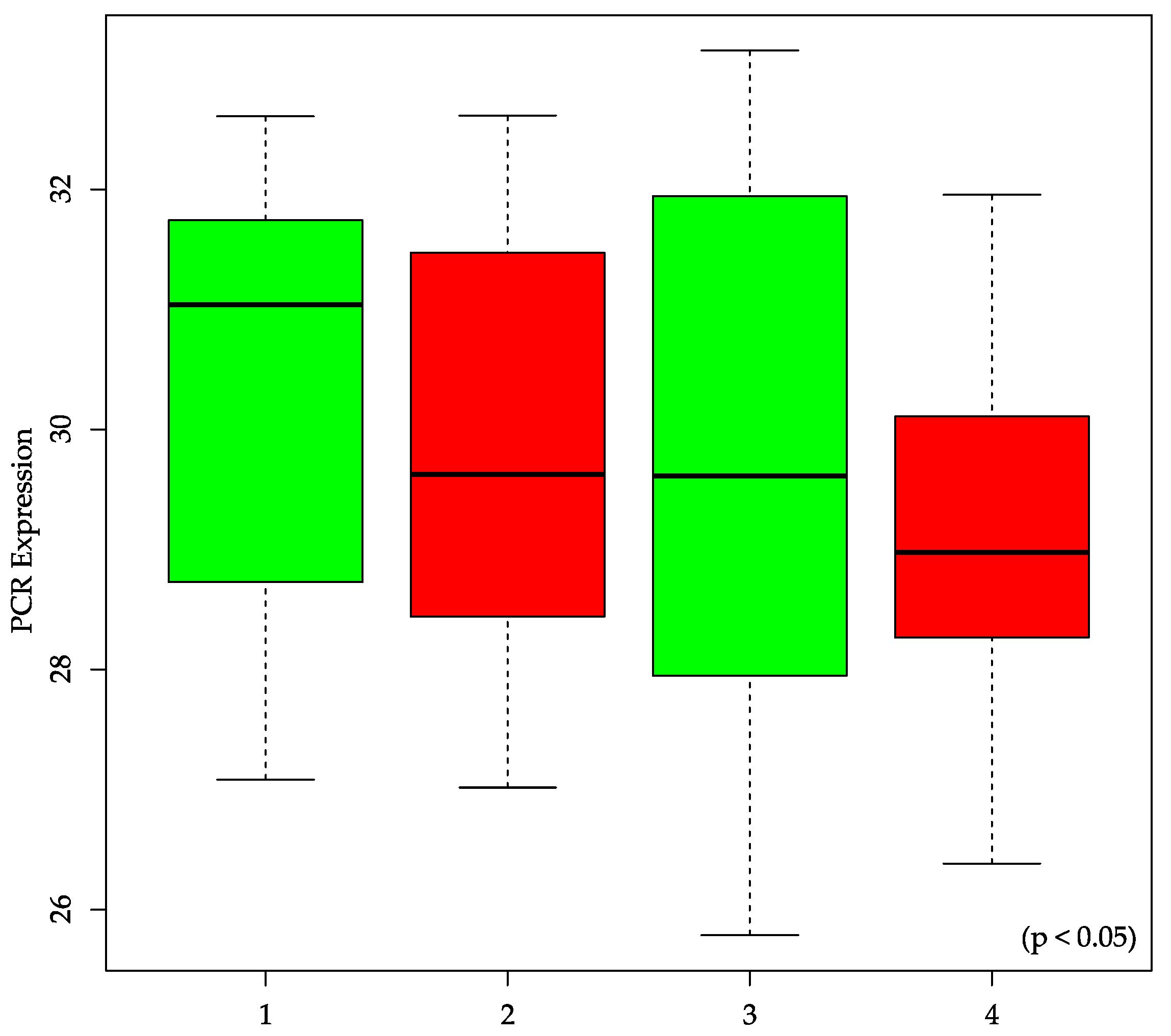
2.4. miR-146b Expression Levels Show a Significant Association with a Negative Post-Therapeutic Nodal Stage (ypN0)
3. Discussion
4. Experimental Section
4.1. Patients and Samples
4.2. Tumor Biopsies, RNA Isolation and Microarray Analysis
4.3. Semi-Quantitative Real-Time PCR
4.4. Statistical Analysis
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| CRT | chemoradiotherapy |
| TME | total mesorectal excision |
| TRG | tumor regression grade |
| miRNA | microRNA |
| OS | overall-survival |
| CSS | cancer-specific-survival |
| LR | local recurrence |
| DM | distant metastasis |
| DMS | distant-metastasis-free survival |
| DF | disease-free-survival |
| ypN | post-chemoradiotherapy nodal status |
| f | female |
| m | male |
References
- Ferlay, J.; Steliarova-Foucher, E.; Lortet-Tieulent, J.; Rosso, S.; Coebergh, J.W.W.; Comber, H.; Forman, D.; Bray, F. Cancer incidence and mortality patterns in Europe: Estimates for 40 countries in 2012. Eur. J. Cancer 2013, 49, 1374–1403. [Google Scholar] [CrossRef] [PubMed]
- Ambros, V. MicroRNA pathways in flies and worms: Growth, death, fat, stress, and timing. Cell 2003, 113, 673–676. [Google Scholar] [CrossRef]
- Li, Y.; Jiang, Z.; Xu, L.; Yao, H.; Guo, J.; Ding, X. Stability analysis of liver cancer-related microRNAs. Acta Biochim. Biophys. Sin. 2011, 43, 69–78. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Sevignani, C.; Dumitru, C.D.; Hyslop, T.; Noch, E.; Yendamuri, S.; Shimizu, M.; Rattan, S.; Bullrich, F.; Negrini, M.; et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc. Natl. Acad. Sci. USA 2004, 101, 2999–3004. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Getz, G.; Miska, E.A.; Alvarez-Saavedra, E.; Lamb, J.; Peck, D.; Sweet-Cordero, A.; Ebert, B.L.; Mak, R.H.; Ferrando, A.A.; et al. MicroRNA expression profiles classify human cancers. Nature 2005, 435, 834–838. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Lai, E.C. Micro RNAs are complementary to 3′ UTR sequence motifs that mediate negative post-transcriptional regulation. Nat. Genet. 2002, 30, 363–364. [Google Scholar] [CrossRef] [PubMed]
- Michael, M.Z.; O’ Connor, S.M.; van Holst Pellekaan, N.G.; Young, G.P.; James, R.J. Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol. Cancer Res. 2003, 1, 882–891. [Google Scholar] [PubMed]
- Xu, X.-H.; Wu, X.-B.; Wu, S.-B.; Liu, H.-B.; Chen, R.; Li, Y. Identification of miRNAs differentially expressed in clinical stages of human colorectal carcinoma—An investigation in Guangzhou, China. PLoS ONE 2014, 9, e94060. [Google Scholar] [CrossRef] [PubMed]
- Slattery, M.L.; Herrick, J.S.; Mullany, L.E.; Valeri, N.; Stevens, J.; Caan, B.J.; Samowitz, W.; Wolff, R.K. An evaluation and replication of miRNAs with disease stage and colorectal cancer-specific mortality. Int. J. Cancer 2015, 137, 428–438. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, B.S.; Jørgensen, S.; Fog, J.U.; Søkilde, R.; Christensen, I.J.; Hansen, U.; Brünner, N.; Baker, A.; Møller, S.; Nielsen, H.J. High levels of microRNA-21 in the stroma of colorectal cancers predict short disease-free survival in stage II colon cancer patients. Clin. Exp. Metastasis 2011, 28, 27–38. [Google Scholar] [CrossRef] [PubMed]
- Gaedcke, J.; Grade, M.; Camps, J.; Søkilde, R.; Kaczkowski, B.; Schetter, A.J.; Difilippantonio, M.J.; Harris, C.C.; Ghadimi, B.M.; Møller, S.; et al. The rectal cancer microRNAome--microRNA expression in rectal cancer and matched normal mucosa. Clin. Cancer Res. 2012, 18, 4919–4930. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhang, G.; Luo, F.; Ruan, J.; Huang, D.; Feng, D.; Xiao, D.; Zeng, Z.; Chen, X.; Wu, W. Identification of aberrantly expressed miRNAs in rectal cancer. Oncol. Rep. 2012, 28, 77–84. [Google Scholar] [PubMed]
- Azizian, A.; Kramer, F.; Jo, P.; Wolff, H.A.; Beissbarth, T.; Skarupke, R.; Bernhardt, M.; Grade, M.; Ghadimi, B.M.; Gaedcke, J. Preoperative prediction of lymph node status by circulating miR-18b and MiR-20a during chemoradiotherapy in patients with rectal cancer. World J. Surg. 2015, 39, 2329–2335. [Google Scholar] [CrossRef] [PubMed]
- Svoboda, M.; Izakovicova Holla, L.; Sefr, R.; Vrtkova, I.; Kocakova, I.; Tichy, B.; Dvorak, J. Micro-RNAs miR125b and miR137 are frequently upregulated in response to capecitabine chemoradiotherapy of rectal cancer. Int. J. Oncol. 2008, 33, 541–547. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Svoboda, M.; Sana, J.; Fabian, P.; Kocakova, I.; Gombosova, J.; Nekvindova, J.; Radova, L.; Vyzula, R.; Slaby, O. MicroRNA expression profile associated with response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer patients. Radiat. Oncol. 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Lopes-Ramos, C.M.; Habr-Gama, A.; Quevedo, B.S.; Felício, N.M.; Bettoni, F.; Koyama, F.C.; Asprino, P.F.; Galante, P.A.; Gama-Rodrigues, J.; Camargo, A.A.; et al. Overexpression of miR-21–5p as a predictive marker for complete tumor regression to neoadjuvant chemoradiotherapy in rectal cancer patients. BMC Med. Genom. 2014, 7. [Google Scholar] [CrossRef] [PubMed]
- Sauer, R.; Becker, H.; Hohenberger, W.; Rödel, C.; Wittekind, C.; Fietkau, R.; Martus, P.; Tschmelitsch, J.; Hager, E.; Hess, C.F.; et al. German Rectal Cancer Study Group Preoperative versus postoperative chemoradiotherapy for rectal cancer. N. Engl. J. Med. 2004, 351, 1731–1740. [Google Scholar] [CrossRef] [PubMed]
- Sauer, R.; Liersch, T.; Merkel, S.; Fietkau, R.; Hohenberger, W.; Hess, C.; Becker, H.; Raab, H.-R.; Villanueva, M.-T.; Witzigmann, H.; et al. Preoperative versus postoperative chemoradiotherapy for locally advanced rectal cancer: Results of the German CAO/ARO/AIO-94 randomized phase III trial after a median follow-up of 11 years. J. Clin. Oncol. 2012, 30, 1926–1933. [Google Scholar] [CrossRef] [PubMed]
- Fokas, E.; Liersch, T.; Fietkau, R.; Hohenberger, W.; Beissbarth, T.; Hess, C.; Becker, H.; Ghadimi, M.; Mrak, K.; Merkel, S.; et al. Tumor regression grading after preoperative chemoradiotherapy for locally advanced rectal carcinoma revisited: Updated results of the CAO/ARO/AIO-94 trial. J. Clin. Oncol. 2014, 32, 1554–1562. [Google Scholar] [CrossRef] [PubMed]
- Rödel, C.; Liersch, T.; Becker, H.; Fietkau, R.; Hohenberger, W.; Hothorn, T.; Graeven, U.; Arnold, D.; Lang-Welzenbach, M.; Raab, H.-R.; et al. German Rectal Cancer Study Group Preoperative chemoradiotherapy and postoperative chemotherapy with fluorouracil and oxaliplatin versus fluorouracil alone in locally advanced rectal cancer: Initial results of the German CAO/ARO/AIO-04 randomised phase 3 trial. Lancet Oncol. 2012, 13, 679–687. [Google Scholar] [PubMed]
- Rödel, C.; Graeven, U.; Fietkau, R.; Hohenberger, W.; Hothorn, T.; Arnold, D.; Hofheinz, R.-D.; Ghadimi, M.; Wolff, H.A.; Lang-Welzenbach, M.; et al. German Rectal Cancer Study Group Oxaliplatin added to fluorouracil-based preoperative chemoradiotherapy and postoperative chemotherapy of locally advanced rectal cancer (the German CAO/ARO/AIO-04 study): Final results of the multicentre, open-label, randomised, phase 3 trial. Lancet Oncol. 2015, 16, 979–989. [Google Scholar] [PubMed]
- Hotchi, M.; Shimada, M.; Kurita, N.; Iwata, T.; Sato, H.; Morimoto, S.; Yoshikawa, K.; Higashijima, J.; Miyatani, T. microRNA expression is able to predict response to chemoradiotherapy in rectal cancer. Mol. Clin. Oncol. 2013, 1, 137–142. [Google Scholar] [CrossRef]
- Toyota, M.; Suzuki, H.; Sasaki, Y.; Maruyama, R.; Imai, K.; Shinomura, Y.; Tokino, T. Epigenetic silencing of microRNA-34b/c and B-cell translocation gene 4 is associated with CpG island methylation in colorectal cancer. Cancer Res. 2008, 68, 4123–4132. [Google Scholar] [CrossRef] [PubMed]
- Quah, H.-M.; Chou, J.F.; Gonen, M.; Shia, J.; Schrag, D.; Saltz, L.B.; Goodman, K.A.; Minsky, B.D.; Wong, W.D.; Weiser, M.R. Pathologic stage is most prognostic of disease-free survival in locally advanced rectal cancer patients after preoperative chemoradiation. Cancer 2008, 113, 57–64. [Google Scholar] [CrossRef] [PubMed]
- Li, C.-F.; He, H.-L.; Wang, J.-Y.; Huang, H.-Y.; Wu, T.-F.; Hsing, C.-H.; Lee, S.-W.; Lee, H.-H.; Fang, J.-L.; Huang, W.-T.; et al. Fibroblast growth factor receptor 2 overexpression is predictive of poor prognosis in rectal cancer patients receiving neoadjuvant chemoradiotherapy. J. Clin. Pathol. 2014, 67, 1056–1061. [Google Scholar] [CrossRef] [PubMed]
- García-Flórez, L.J.; Gómez-Álvarez, G.; Frunza, A.M.; Barneo-Serra, L.; Martínez-Alonso, C.; Fresno-Forcelledo, M.F. Predictive markers of response to neoadjuvant therapy in rectal cancer. J. Surg. Res. 2015, 194, 120–126. [Google Scholar] [CrossRef] [PubMed]
- He, H.-L.; Lee, Y.-E.; Chen, H.-P.; Hsing, C.-H.; Chang, I.-W.; Shiue, Y.-L.; Lee, S.-W.; Hsu, C.-T.; Lin, L.-C.; Wu, T.-F.; et al. Overexpression of DNAJC12 predicts poor response to neoadjuvant concurrent chemoradiotherapy in patients with rectal cancer. Exp. Mol. Pathol. 2015, 98, 338–345. [Google Scholar] [CrossRef] [PubMed]
- Tsang, J.S.; Vencken, S.; Sharaf, O.; Leen, E.; Kay, E.W.; McNamara, D.A.; Deasy, J.; Mulligan, E.D. Global DNA methylation is altered by neoadjuvant chemoradiotherapy in rectal cancer and may predict response to treatment—A pilot study. Eur. J. Surg. Oncol. 2014, 40, 1459–1466. [Google Scholar] [CrossRef] [PubMed]
- Kim, I.Y.; You, S.H.; Kim, Y.W. Neutrophil-lymphocyte ratio predicts pathologic tumor response and survival after preoperative chemoradiation for rectal cancer. BMC Surg. 2014, 14. [Google Scholar] [CrossRef] [PubMed]
- Memon, S.; Lynch, A.C.; Akhurst, T.; Ngan, S.Y.; Warrier, S.K.; Michael, M.; Heriot, A.G. Systematic review of FDG-PET prediction of complete pathological response and survival in rectal cancer. Ann. Surg. Oncol. 2014, 21, 3598–3607. [Google Scholar] [CrossRef] [PubMed]
- Yiu, R.; Wong, S.K.; Cromwell, J.; Madoff, R.D.; Rothenberger, D.A.; Garcia-Aguilar, J. Pelvic wall involvement denotes a poor prognosis in T4 rectal cancer. Dis. Colon Rectum 2001, 44, 1676–1681. [Google Scholar] [CrossRef] [PubMed]
- Hwang, K.; Park, I.J.; Yu, C.S.; Lim, S.-B.; Lee, J.L.; Yoon, Y.S.; Kim, C.W.; Kim, J.C. Impression of prognosis regarding pathologic stage after preoperative chemoradiotherapy in rectal cancer. World J. Gastroenterol. 2015, 21, 563–570. [Google Scholar] [CrossRef] [PubMed]
- Slattery, M.L.; Herrick, J.S.; Bondurant, K.L.; Wolff, R.K. Toll-like receptor genes and their association with colon and rectal cancer development and prognosis. Int. J. Cancer 2012, 130, 2974–2980. [Google Scholar] [CrossRef] [PubMed]
- Huh, J.W.; Lee, W.Y.; Kim, S.H.; Park, Y.A.; Cho, Y.B.; Yun, S.H.; Kim, H.C.; Park, H.C.; Choi, D.H.; Park, J.O.; et al. Immunohistochemical detection of p53 expression in patients with preoperative chemoradiation for rectal cancer: Association with prognosis. Yonsei Med. J. 2015, 56, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Gaedcke, J.; Leha, A.; Claus, R.; Weichenhan, D.; Jung, K.; Kitz, J.; Grade, M.; Wolff, H.A.; Jo, P.; Doyen, J.; et al. Identification of a DNA methylation signature to predict disease-free survival in locally advanced rectal cancer. Oncotarget 2014, 5, 8123–8135. [Google Scholar] [CrossRef] [PubMed]
- Jo, P.; Jung, K.; Grade, M.; Conradi, L.-C.; Wolff, H.A.; Kitz, J.; Becker, H.; Rüschoff, J.; Hartmann, A.; Beissbarth, T.; et al. CpG island methylator phenotype infers a poor disease-free survival in locally advanced rectal cancer. Surgery 2012, 151, 564–570. [Google Scholar] [CrossRef] [PubMed]
- Ghadimi, B.M.; Grade, M.; Difilippantonio, M.J.; Varma, S.; Simon, R.; Montagna, C.; Füzesi, L.; Langer, C.; Becker, H.; Liersch, T.; et al. Effectiveness of gene expression profiling for response prediction of rectal adenocarcinomas to preoperative chemoradiotherapy. J. Clin. Oncol. 2005, 23, 1826–1838. [Google Scholar] [CrossRef] [PubMed]
- Scapoli, L.; Palmieri, A.; Lo Muzio, L.; Pezzetti, F.; Rubini, C.; Girardi, A.; Farinella, F.; Mazzotta, M.; Carinci, F. MicroRNA expression profiling of oral carcinoma identifies new markers of tumor progression. Int. J. Immunopathol. Pharmacol. 2010, 23, 1229–1234. [Google Scholar] [PubMed]
- Mancikova, V.; Castelblanco, E.; Pineiro-Yanez, E.; Perales-Paton, J.; de Cubas, A.A.; Inglada-Perez, L.; Matias-Guiu, X.; Capel, I.; Bella, M.; Lerma, E.; et al. MicroRNA deep-sequencing reveals master regulators of follicular and papillary thyroid tumors. Mod. Pathol. 2015, 28, 748–757. [Google Scholar] [CrossRef] [PubMed]
- Pardo, O.E.; Castellano, L.; Munro, C.E.; Hu, Y.; Mauri, F.; Krell, J.; Lara, R.; Pinho, F.G.; Choudhury, T.; Frampton, A.E.; et al. miR-515–5p controls cancer cell migration through MARK4 regulation. EMBO Rep. 2016, 17, 570–584. [Google Scholar] [CrossRef] [PubMed]
- Gilam, A.; Edry, L.; Mamluk-Morag, E.; Bar-Ilan, D.; Avivi, C.; Golan, D.; Laitman, Y.; Barshack, I.; Friedman, E.; Shomron, N. Involvement of IGF-1R regulation by miR-515–5p modifies breast cancer risk among BRCA1 carriers. Breast Cancer Res. Treat. 2013, 138, 753–760. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.; Luo, T.; Nie, M.; Pang, T.; Zhang, X.; Shen, X.; Ma, L.; Bi, J.; Wei, G.; Fang, G.; et al. TSPAN1 functions as an oncogene in gastric cancer and is downregulated by miR-573. FEBS Lett. 2015, 589, 1988–1994. [Google Scholar] [CrossRef] [PubMed]
- Danza, K.; de Summa, S.; Pinto, R.; Pilato, B.; Palumbo, O.; Merla, G.; Simone, G.; Tommasi, S. MiR-578 and miR-573 as potential players in BRCA-related breast cancer angiogenesis. Oncotarget 2015, 6, 471–483. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.-F.; Chen, H.; Ma, M.-W.; Wang, J.-A.; Tang, T.-T.; Ni, L.-S.; Yu, J.-L.; Li, Y.-Z.; Bai, B.-X. miR-573 regulates melanoma progression by targeting the melanoma cell adhesion molecule. Oncol. Rep. 2013, 30, 520–526. [Google Scholar] [PubMed]
- Quinn, E.M.; Wang, J.; Redmond, H.P. The emerging role of microRNA in regulation of endotoxin tolerance. J. Leukoc. Biol. 2012, 91, 721–727. [Google Scholar] [CrossRef] [PubMed]
- El Gazzar, M.; McCall, C.E. MicroRNAs distinguish translational from transcriptional silencing during endotoxin tolerance. J. Biol. Chem. 2010, 285, 20940–20951. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, D.E.; Nuovo, G.J.; Martin, M.M.; Malana, G.E.; Pleister, A.P.; Jiang, J.; Schmittgen, T.D.; Terry, A.V.; Gardiner, K.; Head, E.; et al. Human chromosome 21-derived miRNAs are overexpressed in down syndrome brains and hearts. Biochem. Biophys. Res. Commun. 2008, 370, 473–477. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, D.E.; Nuovo, G.J.; Terry, A.V.; Martin, M.M.; Malana, G.E.; Sansom, S.E.; Pleister, A.P.; Beck, W.D.; Head, E.; Feldman, D.S.; et al. Chromosome 21-derived microRNAs provide an etiological basis for aberrant protein expression in human Down syndrome brains. J. Biol. Chem. 2010, 285, 1529–1543. [Google Scholar] [CrossRef] [PubMed]
- Bofill-De Ros, X.; Santos, M.; Vila-Casadesús, M.; Villanueva, E.; Andreu, N.; Dierssen, M.; Fillat, C. Genome-wide miR-155 and miR-802 target gene identification in the hippocampus of Ts65Dn Down syndrome mouse model by miRNA sponges. BMC Genom. 2015, 16. [Google Scholar] [CrossRef]
- Kornfeld, J.W.; Baitzel, C.; Könner, A.C.; Nicholls, H.T.; Vogt, M.C.; Herrmanns, K.; Scheja, L.; Haumaitre, C.; Wolf, A.M.; Knippschild, U.; et al. Obesity-induced overexpression of miR-802 impairs glucose metabolism through silencing of Hnf1b. Nature 2013, 494, 111–115. [Google Scholar] [CrossRef] [PubMed]
- Yuan, F.; Wang, W. MicroRNA-802 suppresses breast cancer proliferation through downregulation of FoxM1. Mol. Med. Rep. 2015, 12, 4647–4651. [Google Scholar] [CrossRef] [PubMed]
- Tao, J.; Ji, J.; Li, X.; Ding, N.; Wu, H.; Liu, Y.; Wang, X.W.; Calvisi, D.F.; Song, G.; Chen, X. Distinct anti-oncogenic effect of various microRNAs in different mouse models of liver cancer. Oncotarget 2015, 6, 6977–6988. [Google Scholar] [CrossRef] [PubMed]
- Cao, Z.-Q.; Shen, Z.; Huang, W.-Y. MicroRNA-802 promotes osteosarcoma cell proliferation by targeting p27. Asian Pac. J. Cancer Prev. 2013, 14, 7081–7084. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.Q.; Chen, G.; Liu, X.Y.; Liu, F.Y.; Jiang, S.Y.; Wang, Z. MicroRNA‑802 promotes lung carcinoma proliferation by targeting the tumor suppressor menin. Mol. Med. Rep. 2014, 10, 1537–1542. [Google Scholar] [CrossRef] [PubMed]
- Xiang, K.-M.; Li, X.-R. MiR-133b acts as a tumor suppressor and negatively regulates TBPL1 in colorectal cancer cells. Asian Pac. J. Cancer Prev. 2014, 15, 3767–3772. [Google Scholar] [CrossRef] [PubMed]
- Kano, M.; Seki, N.; Kikkawa, N.; Fujimura, L.; Hoshino, I.; Akutsu, Y.; Chiyomaru, T.; Enokida, H.; Nakagawa, M.; Matsubara, H. miR-145, miR-133a and miR-133b: Tumor-suppressive miRNAs target FSCN1 in esophageal squamous cell carcinoma. Int. J. Cancer 2010, 127, 2804–2814. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharjya, S.; Roy, K.S.; Ganguly, A.; Sarkar, S.; Panda, C.K.; Bhattacharyya, D.; Bhattacharyya, N.P.; Roychoudhury, S. Inhibition of nucleoporin member Nup214 expression by miR-133b perturbs mitotic timing and leads to cell death. Mol. Cancer 2015, 14. [Google Scholar] [CrossRef] [PubMed]
- Qiu, T.; Zhou, X.; Wang, J.; Du, Y.; Xu, J.; Huang, Z.; Zhu, W.; Shu, Y.; Liu, P. miR-145, miR-133a and miR-133b inhibit proliferation, migration, invasion and cell cycle progression via targeting transcription factor Sp1 in gastric cancer. FEBS Lett. 2014, 588, 1168–1177. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Huang, J.; Zhang, L.; Qu, Y.; Li, J.; Yu, B.; Yan, M.; Yu, Y.; Liu, B.; Zhu, Z. MiR-133b is frequently decreased in gastric cancer and its overexpression reduces the metastatic potential of gastric cancer cells. BMC Cancer 2014, 14. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.W.; Li, X.R.; Zhang, Y.; Hu, G.; Guo, Y.H.; Zhou, J.Y.; Du, J.; Lv, L.; Gao, K.; Deng, H. TAp63 suppress metastasis via miR-133b in colon cancer cells. Br. J. Cancer 2014, 110, 2310–2320. [Google Scholar] [CrossRef] [PubMed]
- Boštjančič, E.; Jerše, M.; Glavač, D.; Zidar, N. miR-1, miR-133a/b, and miR-208a in human fetal hearts correlate to the apoptotic and proliferation markers. Exp. Biol. Med. 2015, 240, 211–219. [Google Scholar]
- Ferreira, L.R.P.; Frade, A.F.; Santos, R.H.B.; Teixeira, P.C.; Baron, M.A.; Navarro, I.C.; Benvenuti, L.A.; Fiorelli, A.I.; Bocchi, E.A.; Stolf, N.A.; et al. MicroRNAs miR-1, miR-133a, miR-133b, miR-208a and miR-208b are dysregulated in Chronic Chagas disease Cardiomyopathy. Int. J. Cardiol. 2014, 175, 409–417. [Google Scholar] [CrossRef] [PubMed]
- Koutsoulidou, A.; Mastroyiannopoulos, N.P.; Furling, D.; Uney, J.B.; Phylactou, L.A. Expression of miR-1, miR-133a, miR-133b and miR-206 increases during development of human skeletal muscle. BMC Dev. Biol. 2011, 11. [Google Scholar] [CrossRef] [PubMed]
- Boettger, T.; Wüst, S.; Nolte, H.; Braun, T. The miR-206/133b cluster is dispensable for development, survival and regeneration of skeletal muscle. Skelet. Muscle 2014, 4. [Google Scholar] [CrossRef] [PubMed]
- Chou, C.-K.; Yang, K.D.; Chou, F.-F.; Huang, C.-C.; Lan, Y.-W.; Lee, Y.-F.; Kang, H.-Y.; Liu, R.-T. Prognostic implications of miR-146b expression and its functional role in papillary thyroid carcinoma. J. Clin. Endocrinol. Metab. 2013, 98, E196–E205. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.S.; Lim, Y.S.; Lee, J.-C.; Wang, S.-G.; Park, H.-Y.; Kim, S.Y.; Lee, B.-J. Differential expression levels of plasma-derived miR-146b and miR-155 in papillary thyroid cancer. Oral Oncol. 2015, 51, 77–83. [Google Scholar] [CrossRef] [PubMed]
- Sun, M.; Fang, S.; Li, W.; Li, C.; Wang, L.; Wang, F.; Wang, Y. Associations of miR-146a and miR-146b expression and clinical characteristics in papillary thyroid carcinoma. Cancer Biomark. 2015, 15, 33–40. [Google Scholar] [PubMed]
- Deng, X.; Wu, B.; Xiao, K.; Kang, J.; Xie, J.; Zhang, X.; Fan, Y. MiR-146b-5p promotes metastasis and induces epithelial-mesenchymal transition in thyroid cancer by targeting ZNRF3. Cell. Physiol. Biochem. 2015, 35, 71–82. [Google Scholar] [CrossRef] [PubMed]
- Geraldo, M.V.; Yamashita, A.S.; Kimura, E.T. MicroRNA miR-146b-5p regulates signal transduction of TGF-β by repressing SMAD4 in thyroid cancer. Oncogene 2012, 31, 1910–1922. [Google Scholar] [CrossRef] [PubMed]
- Hardin, H.; Guo, Z.; Shan, W.; Montemayor-Garcia, C.; Asioli, S.; Yu, X.-M.; Harrison, A.D.; Chen, H.; Lloyd, R.V. The roles of the epithelial-mesenchymal transition marker PRRX1 and miR-146b-5p in papillary thyroid carcinoma progression. Am. J. Pathol. 2014, 184, 2342–2354. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Shan, F.; Xiong, G.; Wang, J.-M.; Wang, W.-L.; Xu, X.; Bai, Y. Transcriptional regulation of miR-146b by C/EBPβ LAP2 in esophageal cancer cells. Biochem. Biophys. Res. Commun. 2014, 446, 267–271. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wang, Y.; Yu, L.; Sun, C.; Cheng, D.; Yu, S.; Wang, Q.; Yan, Y.; Kang, C.; Jin, S.; et al. miR-146b-5p inhibits glioma migration and invasion by targeting MMP16. Cancer Lett. 2013, 339, 260–269. [Google Scholar] [CrossRef] [PubMed]
- Katakowski, M.; Buller, B.; Zheng, X.; Lu, Y.; Rogers, T.; Osobamiro, O.; Shu, W.; Jiang, F.; Chopp, M. Exosomes from marrow stromal cells expressing miR-146b inhibit glioma growth. Cancer Lett. 2013, 335, 201–204. [Google Scholar] [CrossRef] [PubMed]
- Drebber, U.; Lay, M.; Wedemeyer, I.; Vallböhmer, D.; Bollschweiler, E.; Brabender, J.; Mönig, S.P.; Hölscher, A.H.; Dienes, H.P.; Odenthal, M. Altered levels of the onco-microRNA 21 and the tumor-supressor microRNAs 143 and 145 in advanced rectal cancer indicate successful neoadjuvant chemoradiotherapy. Int. J. Oncol. 2011, 39, 409–415. [Google Scholar] [CrossRef] [PubMed]
- Kheirelseid, E.A.H.; Miller, N.; Chang, K.H.; Curran, C.; Hennessey, E.; Sheehan, M.; Newell, J.; Lemetre, C.; Balls, G.; Kerin, M.J. miRNA expressions in rectal cancer as predictors of response to neoadjuvant chemoradiation therapy. Int. J. Colorectal Dis. 2013, 28, 247–260. [Google Scholar] [CrossRef] [PubMed]
- Della Vittoria Scarpati, G.; Falcetta, F.; Carlomagno, C.; Ubezio, P.; Marchini, S.; de Stefano, A.; Singh, V.K.; D'Incalci, M.; de Placido, S.; Pepe, S. A specific miRNA signature correlates with complete pathological response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer. Int. J. Radiat. Oncol. Biol. Phys. 2012, 83, 1113–1119. [Google Scholar] [CrossRef] [PubMed]
- Mlcochova, J.; Faltejskova-Vychytilova, P.; Ferracin, M.; Zagatti, B.; Radova, L.; Svoboda, M.; Nemecek, R.; John, S.; Kiss, I.; Vyzula, R.; et al. MicroRNA expression profiling identifies miR-31–5p/3p as associated with time to progression in wild-type RAS metastatic colorectal cancer treated with cetuximab. Oncotarget 2015, 6, 38695–38704. [Google Scholar] [PubMed]
- Pichler, M.; Winter, E.; Ress, A.L.; Bauernhofer, T.; Gerger, A.; Kiesslich, T.; Lax, S.; Samonigg, H.; Hoefler, G. miR-181a is associated with poor clinical outcome in patients with colorectal cancer treated with EGFR inhibitor. J. Clin. Pathol. 2014, 67, 198–203. [Google Scholar] [CrossRef] [PubMed]
- Saridaki, Z.; Weidhaas, J.B.; Lenz, H.J.; Laurent-Puig, P.; Jacobs, B.; de Schutter, J.; de Roock, W.; Salzman, D.W.; Zhang, W.; Yang, D.; et al. A let-7 microRNA-binding site polymorphism in KRAS predicts improved outcome in patients with metastatic colorectal cancer treated with salvage cetuximab/panitumumab monotherapy. Clin. Cancer Res. 2014, 20, 4499–4510. [Google Scholar] [CrossRef] [PubMed]
- Sebio, A.; Paré, L.; Páez, D.; Salazar, J.; González, A.; Sala, N.; del Río, E.; Martín-Richard, M.; Tobeña, M.; Barnadas, A.; et al. The LCS6 polymorphism in the binding site of let-7 microRNA to the KRAS 3′-untranslated region: Its role in the efficacy of anti-EGFR-based therapy in metastatic colorectal cancer patients. Pharmacogenet. Genom. 2013, 23, 142–147. [Google Scholar] [CrossRef] [PubMed]
- Salendo, J.; Spitzner, M.; Kramer, F.; Zhang, X.; Jo, P.; Wolff, H.A.; Kitz, J.; Kaulfuß, S.; Beissbarth, T.; Dobbelstein, M.; et al. Identification of a microRNA expression signature for chemoradiosensitivity of colorectal cancer cells, involving miRNAs-320a, -224, -132 and let7g. Radiother. Oncol. 2013, 108, 451–457. [Google Scholar] [CrossRef] [PubMed]
- Bolstad, B.M.; Irizarry, R.A.; Astrand, M.; Speed, T.P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 2003, 19, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 2004, 3. [Google Scholar] [CrossRef] [PubMed]
- Cox, D.R.; Oakes, D. Analysis of Survival Data; CRC Press: London, UK, 1984. [Google Scholar]
- The R Project for Statistical Computing. Available online: http://www.r-project.org (accessed on 13 April 2016).
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar]
| MicroRNA | Overall Survival | Disease-Free Survival | Cancer-Specific Survival | Distant-Metastasis-Free Survival | ypN 1 | TRG 2 |
|---|---|---|---|---|---|---|
| miR-133b | 0.1981 | 0.1797 | 0.1054 | 0.032 * | 0.8869 | 0.5895 |
| miR-146b | 0.5912 | 0.4065 | 0.7999 | 0.4282 | 0.0465 * | 0.9552 |
| miR-198 | 0.5148 | 0.8378 | 0.4601 | 0.7771 | 0.3038 | 0.8873 |
| miR-223 | 0.8491 | 0.8887 | 0.2834 | 0.9801 | 0.1327 | 0.8693 |
| miR-224 * | 0.1097 | 0.8256 | 0.2389 | 0.664 | 0.4927 | 0.736 |
| miR-23c | 0.6434 | 0.8418 | 0.445 | 0.8975 | 0.2371 | 0.8907 |
| miR-3133 | 0.7581 | 0.6015 | 0.5951 | 0.4516 | 0.146 | 0.2699 |
| miR-320a | 0.2016 | 0.4967 | 0.3011 | 0.2985 | 0.2024 | 0.8778 |
| miR-34b | 0.9669 | 0.7904 | 0.9092 | 0.5506 | 0.6819 | 0.4109 |
| miR-3941 | 0.0637 | 0.5656 | 0.0518 | 0.9055 | 0.7289 | 0.2858 |
| miR-4263 | 0.3752 | 0.8541 | 0.3079 | 0.3768 | 0.9154 | 0.5357 |
| miR-450b-3p | 0.321 | 0.9334 | 0.3794 | 0.8234 | 0.4722 | 0.0878 |
| miR-497 | 0.2883 | 0.8704 | 0.4852 | 0.6942 | 0.8824 | 0.4389 |
| miR-515-5p | 0.0021 * | 0.6666 | 0.0132 * | 0.9754 | 0.4235 | 0.867 |
| miR-518f * | 0.3482 | 0.8358 | 0.5293 | 0.5179 | 0.1116 | 0.0849 |
| miR-573 | 0.0364 * | 0.8672 | 0.0223 * | 0.4698 | 0.0937 | 0.0416 * |
| miR-579 | 0.0092 * | 0.695 | 0.0186 * | 0.896 | 0.0984 | 0.1266 |
| miR-612 | 0.12 | 0.6907 | 0.0734 | 0.979 | 0.5931 | 0.4797 |
| miR-802 | 0.0231 * | 0.1745 | 0.0168 * | 0.411 | 0.152 | 0.1402 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Azizian, A.; Epping, I.; Kramer, F.; Jo, P.; Bernhardt, M.; Kitz, J.; Salinas, G.; Wolff, H.A.; Grade, M.; Beißbarth, T.; et al. Prognostic Value of MicroRNAs in Preoperative Treated Rectal Cancer. Int. J. Mol. Sci. 2016, 17, 568. https://doi.org/10.3390/ijms17040568
Azizian A, Epping I, Kramer F, Jo P, Bernhardt M, Kitz J, Salinas G, Wolff HA, Grade M, Beißbarth T, et al. Prognostic Value of MicroRNAs in Preoperative Treated Rectal Cancer. International Journal of Molecular Sciences. 2016; 17(4):568. https://doi.org/10.3390/ijms17040568
Chicago/Turabian StyleAzizian, Azadeh, Ingo Epping, Frank Kramer, Peter Jo, Markus Bernhardt, Julia Kitz, Gabriela Salinas, Hendrik A. Wolff, Marian Grade, Tim Beißbarth, and et al. 2016. "Prognostic Value of MicroRNAs in Preoperative Treated Rectal Cancer" International Journal of Molecular Sciences 17, no. 4: 568. https://doi.org/10.3390/ijms17040568
APA StyleAzizian, A., Epping, I., Kramer, F., Jo, P., Bernhardt, M., Kitz, J., Salinas, G., Wolff, H. A., Grade, M., Beißbarth, T., Ghadimi, B. M., & Gaedcke, J. (2016). Prognostic Value of MicroRNAs in Preoperative Treated Rectal Cancer. International Journal of Molecular Sciences, 17(4), 568. https://doi.org/10.3390/ijms17040568





