Transcriptome Profiling Analysis of Wolf Spider Pardosa pseudoannulata (Araneae: Lycosidae) after Cadmium Exposure
Abstract
:1. Introduction
2. Results and Discussion
2.1. Sequence Analysis and De Novo Assembly
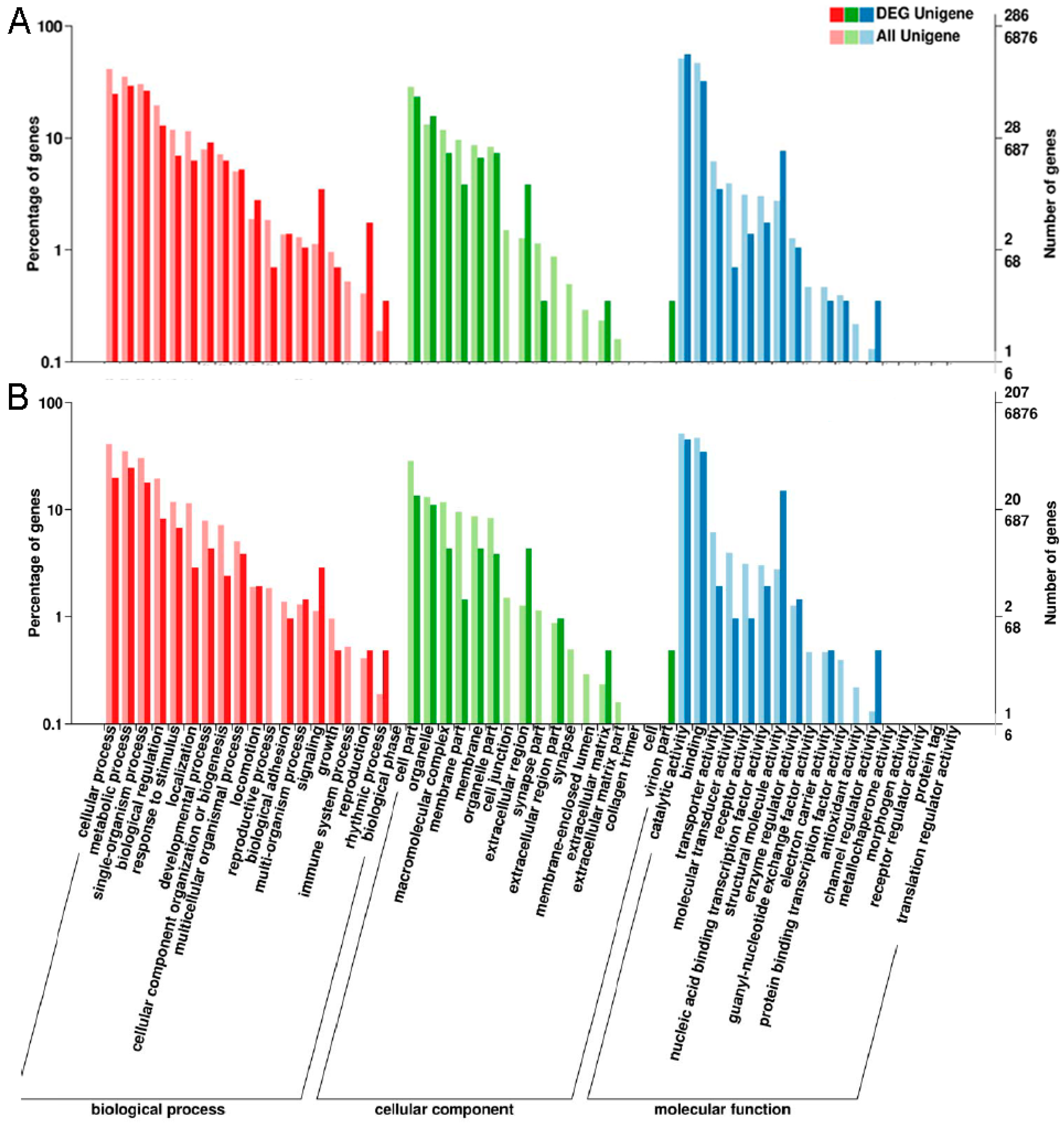
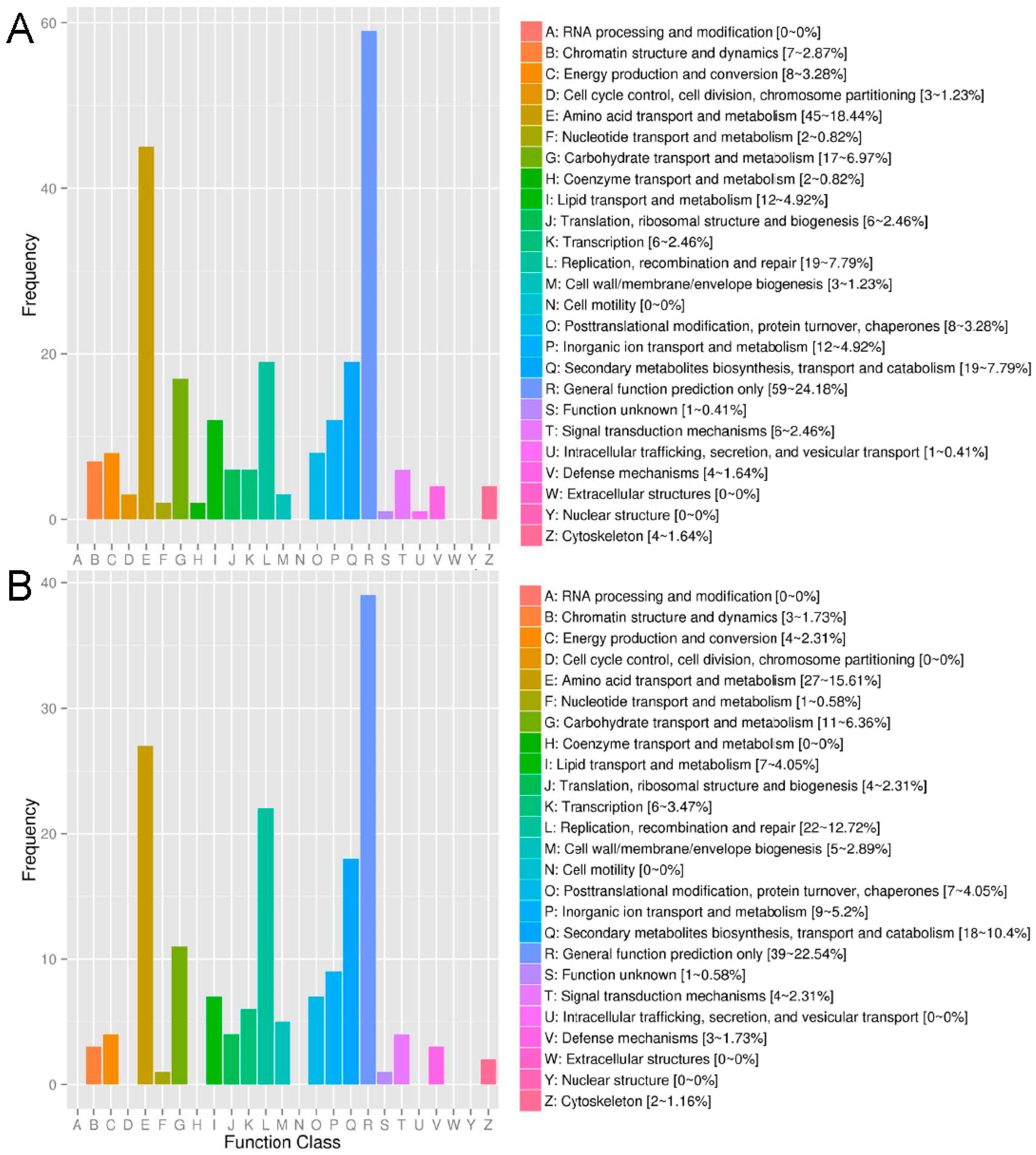
2.2. Annotation of Unigenes
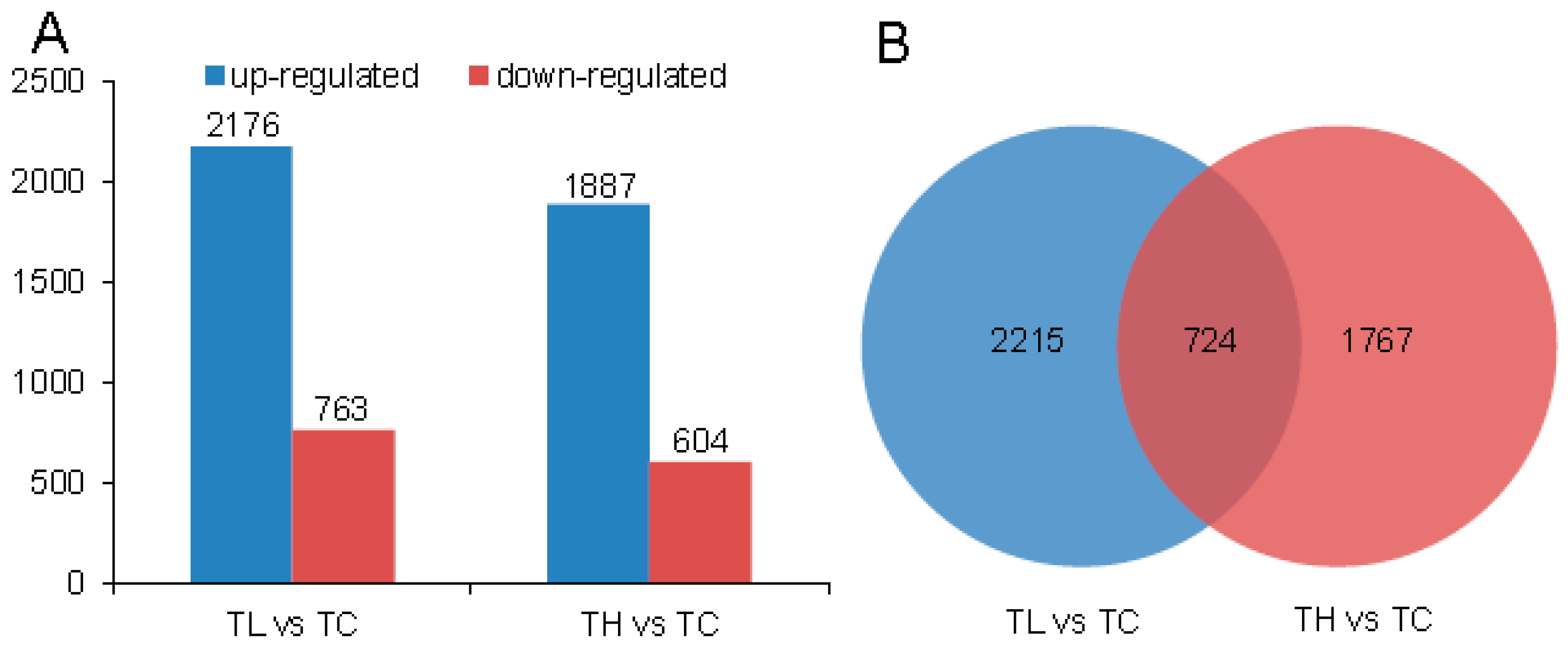
2.3. Differential Expression Analysis
2.4. Functional Enrichment Analysis of DEGs
2.5. Analysis of Genes Related to Cd Detoxification
2.6. Validation of mRNA-Seq Data by RT-qPCR
3. Materials and Methods
3.1. Animal Materials and RNA Extraction
3.2. cDNA Library Construction, Sequencing and De Novo Assembly
3.3. Unigene Functional Annotation
3.4. Differential Dene Expression and Functional Annotation Analyses
3.5. Validation of mRNA-Seq Data
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Nemmiche, S.; Chabane-Sari, D.; Kadri, M.; Guiraud, P. Cadmium chloride-induced oxidative stress and DNA damage in the human Jurkat T cell line is not linked to intracellular trace elements depletion. Toxicol. In Vitro 2011, 25, 191–198. [Google Scholar] [CrossRef] [PubMed]
- He, B.; Yun, Z.J.; Shi, J.B.; Jiang, G.B. Research progress of heavy metal pollution in China: Sources, analytical methods, status, and toxicity. Chin. Sci. Bull. 2013, 58, 134–140. [Google Scholar] [CrossRef]
- Li, Z.Y.; Ma, Z.W.; van der Kuijp, T.J.; Yuan, Z.W.; Huang, L. A review of soil heavy metal pollution from mines in China: Pollution and health risk assessment. Sci. Total Environ. 2014, 468–469, 843–853. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, S.A.; Kristiansen, E.; Andersen, R.A.; Zachariassen, K.E. Cadmium is deposited in the gut content of larvae of the beetle Tenebrio molitor and involves a Cd-binding protein of the low cysteine type. Comp. Biochem. Physiol. C 2008, 148, 217–222. [Google Scholar] [CrossRef] [PubMed]
- Nacke, H.; Gonçalves, A.C., Jr.; Schwantes, D.; Nava, I.A.; Strey, L.; Coelho, G.F. Availability of heavy metals (Cd, Pb, and Cr) in agriculture from commercial fertilizers. Arch. Environ. Contam. Toxicol. 2013, 64, 537–544. [Google Scholar] [CrossRef] [PubMed]
- Jung, C.S.; Lee, S.B.; Jung, M.P.; Lee, J.H.; Lee, S.; Lee, S.H. Accumulated heavy metal content in wolf spider, Pardosa astrigera (Araneae: Lycosidae), as a bioindicator of exposure. J. Asia-Pac. Entomol. 2005, 8, 185–192. [Google Scholar] [CrossRef]
- Heikens, A.; Peijnenburg, W.J.; Hendriks, A.J. Bioaccumulation of heavy metals in terrestrial invertebrates. Environ. Pollut. 2001, 113, 385–393. [Google Scholar] [CrossRef]
- Eraly, D.; Hendrickx, F.; Backeljau, T.; Bervoets, L.; Lens, L. Direct and indirect effects of metal stress on physiology and life history variation in field populations of a lycosid spider. Ecotoxicol. Environ. Saf. 2011, 74, 1489–1497. [Google Scholar] [CrossRef] [PubMed]
- Wilczek, G.; Babczyńska, A.; Wilczek, P. Antioxidative responses in females and males of the spider Xerolycosa nemoralis (Lycosidae) exposed to natural and anthropogenic stressors. Comp. Biochem. Physiol. C 2013, 157, 119–131. [Google Scholar] [CrossRef] [PubMed]
- Li, C.C.; Li, G.Y.; Yun, Y.L.; Chen, J.; Zhang, Z.T.; Peng, Y. The effect of cadmium exposure on fitness-related traits and antioxidant responses in the wolf spider, Pardosa pseudoannulata. Bull. Environ. Contam. Toxicol. 2016, 97, 31–36. [Google Scholar] [CrossRef]
- Rani, A.; Kumar, A.; Lal, A.; Pant, M. Cellular mechanisms of cadmium-induced toxicity: A review. Int. J. Environ. Health Res. 2014, 24, 378–399. [Google Scholar] [CrossRef] [PubMed]
- Stalmach, M.; Wilczek, G.; Wilczek, P.; Skowronek, M.; Mędrzak, M. DNA damage in haemocytes and midgut gland cells of Steatoda grossa (Theridiidae) spiders exposed to food contaminated with cadmium. Ecotoxicol. Environ. Saf. 2015, 113, 353–361. [Google Scholar] [CrossRef] [PubMed]
- Shao, X.L.; Peng, Y.; Hose, G.C.; Chen, J.; Liu, F.X. Spider webs as indicators of heavy metal pollution in air. Bull. Environ. Contam. Toxicol. 2006, 76, 271–277. [Google Scholar]
- Chen, X.Q.; Zhang, Z.T.; Liu, R.; Zhang, X.L.; Chen, J.; Peng, Y. Effects of the metals lead and zinc on the growth, development, and reproduction of Pardosa astrigera (Araneae: Lycosidae). Bull. Environ. Contam. Toxicol. 2011, 86, 203–207. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.K.; Zhang, Y.X.; Bao, H.B.; Liu, Z.W. Sequence analysis of insecticide action and detoxification-related genes in the insect pest natural enemy Pardosa pseudoannulata. PLoS ONE 2015, 10, e0125242. [Google Scholar] [CrossRef] [PubMed]
- Clarke, T.H.; Garb, J.E.; Hayashi, C.Y.; Haney, R.A.; Lancaster, A.K.; Corbett, S.; Ayoub, N.A. Multi-tissue transcriptomics of the black widow spider reveals expansions, co-options, and functional processes of the silk gland gene toolkit. BMC Genom. 2014, 15, 365. [Google Scholar] [CrossRef] [PubMed]
- Haney, R.A.; Ayoub, N.A.; Clarke, T.H.; Hayashi, C.Y.; Garb, J.E. Dramatic expansion of the black widow toxin arsenal uncovered by multi-tissue transcriptomics and venom proteomics. BMC Genom. 2014, 15, 366. [Google Scholar] [CrossRef] [PubMed]
- Xiao, R.; Wang, L.; Cao, Y.S.; Zhang, G. Transcriptome response to temperature stress in the wolf spider Pardosa pseudoannulata (Araneae: Lycosidae). Ecol. Evol. 2016, 6, 3540–3554. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.L.; Liu, M.; Jiang, K.Y.; Wang, B.J.; Tian, X.; Sun, S.J.; Luo, Z.Y.; Qiu, C.W.; Wang, L. De novo characterization of Japanese scallop Mizuhopecten yessoensis transcriptome and analysis of its gene expression following cadmium exposure. PLoS ONE 2013, 8, e64485. [Google Scholar] [CrossRef] [PubMed]
- Jeppe, K.J.; Carew, M.E.; Long, S.M.; Lee, S.F.; Pettigrove, V.; Hoffmann, A.A. Genes involved in cysteine metabolism of Chironomus tepperiare regulated differently by copper and by cadmium. Comp. Biochem. Physiol. C 2014, 162, 1–6. [Google Scholar]
- Mehinto, A.C.; Melinda, S.; Prucha, M.S.; Colli-Dulac, R.C.; Kroll, K.J.; Lavelle, C.M.; Barber, D.S.; Vulpe, C.D.; Denslow, N.D. Gene networks and toxicity pathways induced by acute cadmium exposure in adult largemouth bass (Micropterus salmoides). Aquat. Toxicol. 2014, 152, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Lou, Y.G.; Zhang, G.R.; Zhang, W.Q.; Hu, Y.; Zhang, J. Biological control of rice insect pests in China. Biol. Control 2013, 67, 8–20. [Google Scholar] [CrossRef]
- Bu, C.Y.; Li, J.L.; Wang, X.Q.; Shi, G.L.; Peng, B.; Han, J.Y.; Gao, P.; Wang, Y.N. Transcriptome analysis of the carmine spider mite, Tetranychus cinnabarinus (Boisduval, 1867) (Acari: Tetranychidae), and its response to β-Sitosterol. BioMed Res. Int. 2015. [Google Scholar] [CrossRef] [PubMed]
- Chen, E.H.; Wei, D.D.; Shen, G.M.; Yuan, G.R.; Bai, P.P.; Wang, J.J. De novo characterization of the Dialeurodes citri transcriptome: Mining genes involved in stress resistance and simple sequence repeats (SSRs) discovery. Insect Mol. Biol. 2014, 23, 52–66. [Google Scholar] [CrossRef] [PubMed]
- Karray, S.; Marchand, J.; Moreau, B.; Tastard, E.; Thiriet-Rupert, S.; Geffard, A.; Delahaut, L.; Denis, F.; Hamza-Chaffai, A.; Chénais, B. Transcriptional response of stress-regulated genes to cadmium exposure in the cockle Cerastoderma glaucum from the gulf of Gabès area (Tunisia). Environ. Sci. Pollut. Res. 2015, 22, 17290–17302. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.H.; Shao, Y.N.; Li, C.H.; Zhang, W.W.; Duan, X.M.; Zhao, X.L.; Qiu, Q.F.; Jin, C.H. RNA-seq analysis revealed ROS-mediated related genes involved in cadmium detoxification in the razor clam Sinonovacula constricta. Fish Shellfish Immun. 2016, 57, 350–361. [Google Scholar] [CrossRef] [PubMed]
- Jo, P.G.; Choi, Y.K.; Choi, C.Y. Cloning and mRNA expression of antioxidant enzymes in the Pacific oyster, Crassostrea gigas in response to cadmium exposure. Comp. Biochem. Physiol. C 2008, 147, 460–469. [Google Scholar] [CrossRef] [PubMed]
- Hao, C.F.; Hao, W.D.; Wei, X.T.; Xing, L.N.; Jiang, J.J.; Shang, L.Q. The role of MAPK in the biphasic dose-response phenomenon induced by cadmium and mercury in HEK293 cells. Toxicol. In Vitro 2009, 23, 660–666. [Google Scholar] [CrossRef] [PubMed]
- Sun, M.; Li, Y.T.; Liu, Y.; Lee, S.C.; Wang, L. Transcriptome assembly and expression profiling of molecular responses to cadmium toxicity in hepatopancreas of the freshwater crab Sinopotamon henanense. Sci. Rep. 2016. [Google Scholar] [CrossRef] [PubMed]
- Messner, B.; Türkcan, A.; Ploner, C.; Laufer, G.; Bernhard, D. Cadmium overkill: Autophagy, apoptosis and necrosis signalling in endothelial cells exposed to cadmium. Cell. Mol. Life Sci. 2016. [Google Scholar] [CrossRef] [PubMed]
- Hendrickx, F.; Maelfait, J.P.; Lens, L. Effect of metal stress on life history divergence and quantitative genetic architecture in a wolf spider. J. Evol. Biol. 2008, 21, 183–193. [Google Scholar] [CrossRef] [PubMed]
- Babczyńska, A.; Wilczek, G.; Szulińska, E.; Kędziorski, A.; Franiel, I.; Migula, P. The reproductive potential of the spiders Agelena labyrinthica and Xerolycosa nemoralis from areas contaminated with metals. Sci. Total Environ. 2012, 435–436, 374–379. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Gao, J.; Yun, Y.L.; Hu, Z.Y.; Peng, Y. Bioaccumulation of mercury and its effects on survival, development and web-weaving in the funnel-web spider Agelena labyrinthica (Araneae: Agelenidae). Bull. Environ. Contam. Toxicol. 2013, 90, 558–562. [Google Scholar] [CrossRef] [PubMed]
- Wilczek, G. Apoptosis and biochemical biomarkers of stress in spiders from industrially polluted areas exposed to high temperature and dimethoate. Comp. Biochem. Physiol. C 2005, 141, 194–206. [Google Scholar] [CrossRef] [PubMed]
- Wilczek, G.; Babczyńska, A.; Wilczek, P.; Doleżych, B.; Migula, P.; Młyńska, H. Cellular stress reactions in female and male spiders from areas variously polluted with heavy metals. Ecotoxicol. Environ. Saf. 2008, 70, 127–137. [Google Scholar] [CrossRef] [PubMed]
- Aimola, P.; Carmignani, M.; Volpe, A.R.; Benedetto, A.D.; Claudio, L.; Waalkes, M.P.; Bokhoven, A.; Tokar, E.J.; Claudio, P.C. Cadmium induces p53-dependent apoptosis in human prostate epithelial cells. PLoS ONE 2012, 7, e33647. [Google Scholar] [CrossRef] [PubMed]
- Hall, J. Cellular mechanisms for heavy metal detoxification and tolerance. J. Exp. Bot. 2002, 53, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Kingtong, S.; Chitramvong, Y.; Janvilisri, T. ATP-binding cassette multidrug transporters in Indian-rock oyster Saccostrea forskali and their role in the export of an environmental organic pollutant tributyltin. Aquat. Toxicol. 2007, 85, 124–132. [Google Scholar] [CrossRef] [PubMed]
- Tian, S.Q.; Gu, C.S.; Liu, L.Q.; Zhu, X.D.; Zhao, Y.H.; Huang, S.Z. Transcriptome profiling of Louisiana iris root and identification of genes involved in lead-stress response. Int. J. Mol. Sci. 2015, 16, 28087–28097. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Xu, L.; Chen, Y.L.; Shen, H.; Gong, Y.Q.; Limera, C.; Liu, L.W. Transcriptome profiling of radish (Raphanus sativus L.) root and identification of genes involved in response to lead (Pb) stress with next generation sequencing. PLoS ONE 2013, 8, e66539. [Google Scholar] [CrossRef] [PubMed]
- Kazemi-Dinan, A.; Thomaschky, S.; Stein, R.J.; Krämer, U.; Müller, C. Zinc and cadmium hyperaccumulation act as deterrents towards specialist herbivores and impede the performance of a generalist herbivore. New Phytol. 2014, 202, 628–639. [Google Scholar] [CrossRef] [PubMed]
- Shi, B.; Huang, Z.K.; Xiang, X.; Huang, M.Q.; Wang, W.X.; Ke, C.H. Transcriptome analysis of the key role of GAT2 gene in the hyper-accumulation of copper in the oyster Crassostrea angulata. Sci. Rep. 2015. [Google Scholar] [CrossRef] [PubMed]
- Yepiskoposyan, H.; Egli, D.; Fergestad, T.; Selvaraj, A.; Treiber, C.; Multhaup, G.; Georgiev, O.; Schaffner, W. Transcriptome response to heavy metal stress in Drosophila reveals a new zinc transporter that confers resistance to zinc. Nucleic Acids Res. 2006, 34, 4866–4877. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.X.; McBride, S.J.; Boyd, W.A.; Alper, S.; Freedman, J.H. Toxicogenomic analysis of Caenorhabditis elegans reveals novel genes and pathways involved in the resistance to cadmium toxicity. Genome Biol. 2007, 8, R122. [Google Scholar] [CrossRef] [PubMed]
- Misra, U.K.; Gawdi, G.; Akabani, G.; Pizzo, S.V. Cadmium-induced DNA synthesis and cell proliferation in macrophages: The role of intracellular calcium and signal transduction mechanisms. Cell Signal. 2002, 14, 327–340. [Google Scholar] [CrossRef]
- Liu, J.; Kadiiska, M.B.; Corton, J.C.; Qu, W.; Waalkes, M.P.; Mason, R.P.; Liu, Y.; Klaassen, C.D. Acute cadmium exposure induces stress-related gene expression in wild-type and metallothionein-I/II null mice. Free Radic. Biol. Med. 2002, 32, 525–535. [Google Scholar] [CrossRef]
- Chen, L.; Liu, L.; Huang, S. Cadmium activates the mitogen-activated protein kinase (MAPK) pathway via induction of reactive oxygen species and inhibition of protein phosphatases 2A and 5. Free Radic. Biol. Med. 2008, 45, 1035–1044. [Google Scholar] [CrossRef] [PubMed]
- Hofmann, G.E.; Buckley, B.A.; Place, S.P.; Zippay, M.L. Molecular chaperones in ectothermic marine animals: Biochemical function and gene expression. Integr. Comp. Biol. 2002, 42, 808–814. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.C.; Sharma, A.; Mishra, M.; Mishra, R.K.; Chowdhuri, D.K. Heat shock proteins in toxicology: How close and how far? Life Sci. 2010, 86, 377–384. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.M.; Rhee, J.S.; Jeong, C.B.; Seo, J.S.; Park, G.S.; Lee, Y.M.; Lee, J.S. Heavy metals induce oxidative stress and trigger oxidative stress-mediated heat shock protein (hsp) modulation in the intertidal copepod Tigriopus japonicus. Comp. Biochem. Physiol. C 2014, 166, 65–74. [Google Scholar] [CrossRef] [PubMed]
- King, A.M.; Macrae, T.H. Insect heat shock proteins during stress and diapause. Annu. Rev. Entomol. 2015, 60, 59–75. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.Y.; Guan, X.T.; Yao, L.L.; Zhang, H.; Jin, X.; Han, Y. Effects of single and joint subacute exposure of copper and cadmium on heat shock proteins in common carp (Cyprinus carpio). Biol. Trace Elem. Res. 2016, 169, 374. [Google Scholar] [CrossRef] [PubMed]
- Stohs, S.J.; Bagchi, D.; Hassoun, E.; Bagchi, M. Oxidative mechanisms in the toxicity of chromium and cadmium ions. J. Environ. Pathol. Toxicol. 2001, 20, 77–88. [Google Scholar] [CrossRef]
- Itziou, A.; Kaloyianni, M.; Dimitriadis, V.K. In vivo and in vitro effects of metals in reactive oxygen species production, protein carbonylation, and DNA damage in land snails Eobania vermiculata. Arch. Environ. Contam. Toxicol. 2011, 60, 697–707. [Google Scholar] [CrossRef] [PubMed]
- Blagojević, D.P.; Grubor-Lajšić, G. Multifunctionality of antioxidant system in insects. Arch. Biol. Sci. 2000, 52, 185–194. [Google Scholar]
- Wilczek, G.; Kramarz, P.; Babczyńska, A. Activity of carboxylesterase and glutathione S-transferase in different life-stages of carabid beetle (Poecilus cupreus) exposed to toxic metal concentrations. Comp. Biochem. Physiol. C 2003, 134, 501–512. [Google Scholar] [CrossRef]
- Wang, Y.; Fang, J.; Leonard, S.S.; Rao, K.M. Cadmium inhibits the electron transfer chain and induces reactive oxygen species. Free Radic. Biol. Med. 2004, 36, 1434–1443. [Google Scholar] [CrossRef] [PubMed]
- Pickett, C.B.; Lu, A.Y.H. Glutathione-S-transferases: Gene structure, regulation and biological function. Annu. Rev. Biochem. 1989, 58, 743–764. [Google Scholar] [CrossRef] [PubMed]
- Mirčić, D.; Blagojević, D.; Mataruga, V.P.; Ilijin, L.; Mrdaković, M.; Vlahović, M.; Lazarević, J. Cadmium effects on the fitness-related traits and antioxidative defense of Lymantria dispar L. larvae. Environ. Sci. Pollut. Res. 2013, 20, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Hui, K.M.; Hao, F.Y.; Li, W.; Zhang, C.Y.; Wang, W.; Ren, Q. Cloning and identification of four Mu-type glutathione S-transferases from the giant freshwater prawn Macrobrachium rosenbergii. Fish Shellfish Immun. 2013, 35, 546–552. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Quackenbush, J.; Cho, J.; Lee, D.; Liang, F.; Holt, I.; Karamycheva, S.; Parvizi, B.; Pertea, G.; Sultana, R.; White, J. The TIGR gene indices: Analysis of gene transcript sequences in highly sampled eukaryotic species. Nucleic Acids Res. 2001, 29, 159–164. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; Baren, M.J.; Salzberg, S.L.; Pachter, L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [PubMed]
- Leng, N.; Dawson, J.A.; Thomson, J.A.; Ruotti, V.; Rissman, A.I.; Smits, B.M.G.; Haag, J.D.; Gould, M.N.; Stewart, R.M.; Kendziorski, C. EBSeq: An empirical Bayes hierarchical model for inference in RNA-seq experiments. Bioinformatics 2013, 29, 1035–1043. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 1995, 57, 289–300. [Google Scholar]
- Alexa, A.; Rahnenführer, J.; Lengauer, T. Improved scoring of functional groups from gene expression data by decorrelating GO graph structure. Bioinformatics 2006, 22, 1600–1607. [Google Scholar] [CrossRef] [PubMed]
| Length Range (bp) | Transcript | Unigene |
|---|---|---|
| 0–300 | 11,907 (10.02%) | 0 (0%) |
| 300–500 | 40,345 (33.95%) | 27,245 (45.04%) |
| 500–1000 | 31,410 (26.43%) | 16,985 (28.08%) |
| 1000–2000 | 20,515 (17.26%) | 9867 (16.31%) |
| 2000+ | 14,674 (12.35%) | 6390 (10.56% ) |
| Total number | 118,853 | 60,489 |
| Total length | 119,999,996 | 57,521,612 |
| N50 length | 1637 | 1433 |
| Mean length | 1009.65 | 950.94 |
| Databases Used for Annotation | Annotated Number | 300 ≤ Length < 1000 | Length ≥ 1000 |
|---|---|---|---|
| COG | 5131 (8.48%) | 1312 | 3819 |
| GO | 6876 (11.37%) | 2310 | 4566 |
| KEGG | 8297 (13.72%) | 2484 | 5813 |
| KOG | 11,804 (19.51%) | 3694 | 8110 |
| Pfam | 13,550 (22.40%) | 4204 | 9346 |
| Swissprot | 10,216 (16.89%) | 3164 | 7052 |
| nr | 18,477 (30.55%) | 7477 | 11,000 |
| All | 18,773 (31.04%) | 7709 | 11,064 |
| TL vs. TC | TH vs. TC |
|---|---|
| Glycine, serine and threonine metabolism (11 DEGs) | Lysosome (7 DEGs) |
| Lysosome (10 DEGs) | Fat digestion and absorption (7 DEGs) |
| Fat digestion and absorption (10 DEGs) | Fatty acid elongation (5 DEGs) |
| Protein processing in endoplasmic reticulum (9 DEGs) | Protein processing in endoplasmic reticulum (5 DEGs) |
| Phagosome (8 DEGs) | Phagosome (5 DEGs) |
| Carbon metabolism (8 DEGs) | Notch signalling pathway (4 DEGs) |
| Pyruvate metabolism (7 DEGs) | Peroxisome (4 DEGs) |
| Drug metabolism-cytochrome P450 (7 DEGs) | Glycine, serine and threonine metabolism (4 DEGs) |
| Oxidative phosphorylation (6 DEGs) | Valine, leucine and isoleucine degradation (4 DEGs) |
| Notch signalling pathway (6 DEGs) | Glutathione metabolism (3 DEGs) |
| Valine, leucine and isoleucine degradation (6 DEGs) | Drug metabolism-cytochrome P450 (3 DEGs) |
| Transcript ID | Brief Description | DEG Library | Fold by RNA-Seq | Fold by qPCR |
|---|---|---|---|---|
| c59601.graph_c0 | Cytochrome P450 4C1 | TL vs. TC | 7.78 | 21.55 |
| c82407.graph_c1 | Heat shock protein 70 | TL vs. TC | −2.53 | 0.26 |
| c91015.graph_c0 | Myb-related transcription factor | TH vs. TC | 7.39 | 4.65 |
| c92582.graph_c0 | Glutathione S-transferase | TL vs. TC | 2.80 | 4.41 |
| c98395.graph_c0 | Cytochrome P450 4c3 | TL vs. TC | 9.06 | 42.27 |
| TH vs. TC | 8.47 | 15.51 | ||
| c98430.graph_c0 | gamma-Glutamyltransferase | TL vs. TC | 2.56 | 2.93 |
| TH vs. TC | 2.27 | 2.24 | ||
| c101656.graph_c1 | ABC transporter | TL vs. TC | 2.24 | 4.96 |
| c91520.graph_c0 | Glutathione S-transferase | TL vs. TC | 2.62 | 10.44 |
| TH vs. TC | 3.24 | 21.13 | ||
| c87886.graph_c0 | Dimethylaniline monooxygenase | TL vs. TC | 7.60 | 315.25 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, C.-C.; Wang, Y.; Li, G.-Y.; Yun, Y.-L.; Dai, Y.-J.; Chen, J.; Peng, Y. Transcriptome Profiling Analysis of Wolf Spider Pardosa pseudoannulata (Araneae: Lycosidae) after Cadmium Exposure. Int. J. Mol. Sci. 2016, 17, 2033. https://doi.org/10.3390/ijms17122033
Li C-C, Wang Y, Li G-Y, Yun Y-L, Dai Y-J, Chen J, Peng Y. Transcriptome Profiling Analysis of Wolf Spider Pardosa pseudoannulata (Araneae: Lycosidae) after Cadmium Exposure. International Journal of Molecular Sciences. 2016; 17(12):2033. https://doi.org/10.3390/ijms17122033
Chicago/Turabian StyleLi, Chang-Chun, Yong Wang, Guo-Yuan Li, Yue-Li Yun, Yu-Jun Dai, Jian Chen, and Yu Peng. 2016. "Transcriptome Profiling Analysis of Wolf Spider Pardosa pseudoannulata (Araneae: Lycosidae) after Cadmium Exposure" International Journal of Molecular Sciences 17, no. 12: 2033. https://doi.org/10.3390/ijms17122033
APA StyleLi, C.-C., Wang, Y., Li, G.-Y., Yun, Y.-L., Dai, Y.-J., Chen, J., & Peng, Y. (2016). Transcriptome Profiling Analysis of Wolf Spider Pardosa pseudoannulata (Araneae: Lycosidae) after Cadmium Exposure. International Journal of Molecular Sciences, 17(12), 2033. https://doi.org/10.3390/ijms17122033






