Comparative Analysis of Whole-Genome Gene Expression Changes in Cultured Human Embryonic Stem Cells in Response to Low, Clinical Diagnostic Relevant, and High Doses of Ionizing Radiation Exposure
Abstract
:1. Introduction
2. Results and Discussion
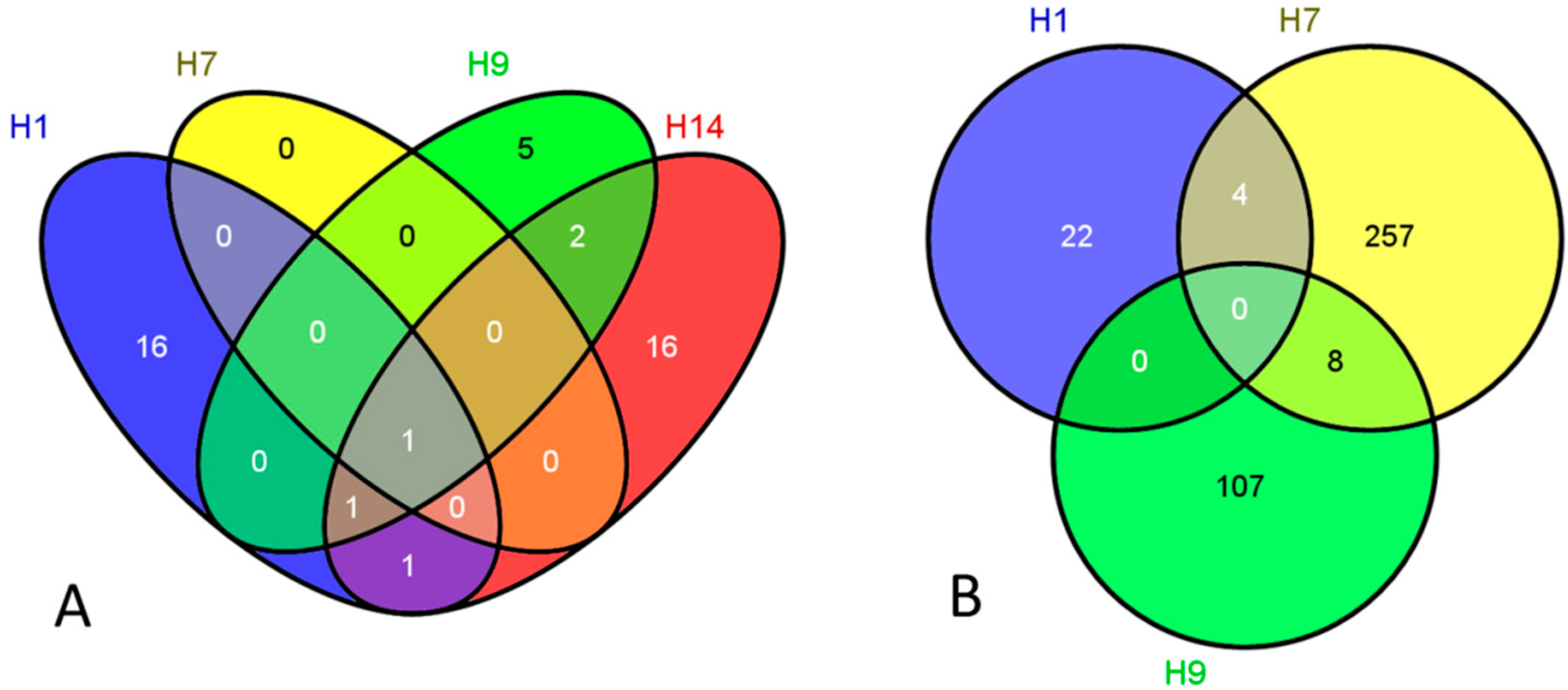
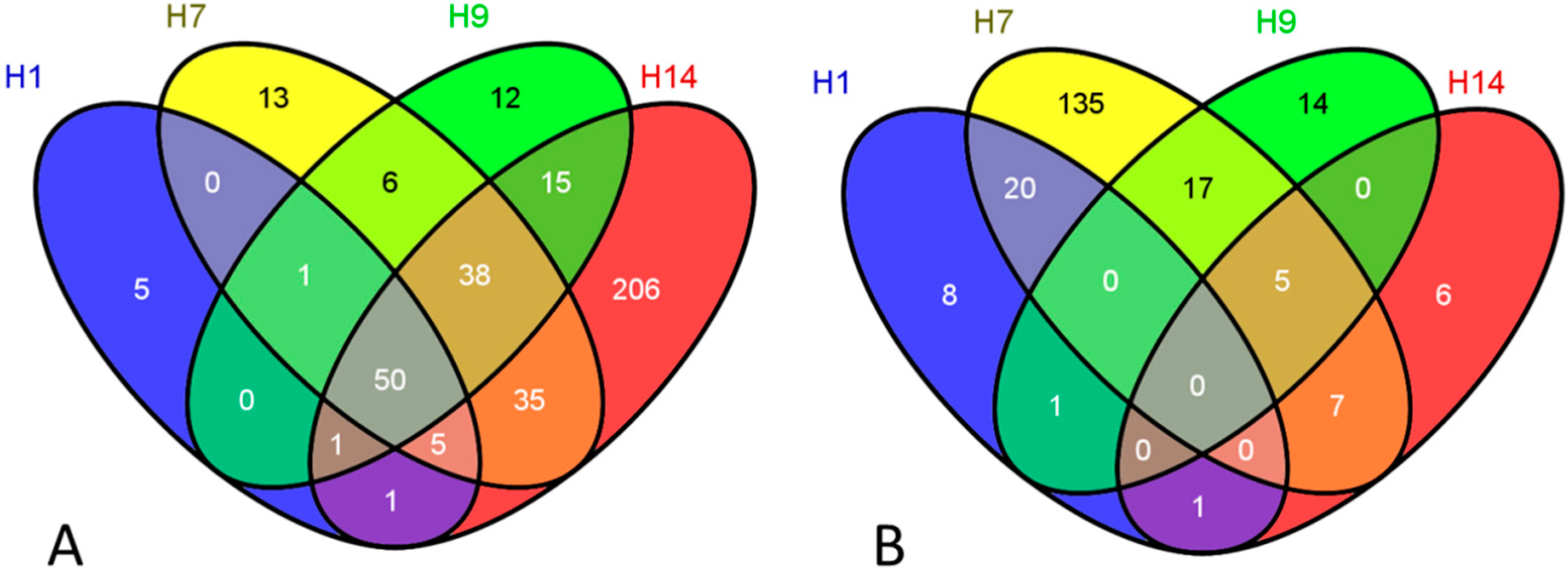
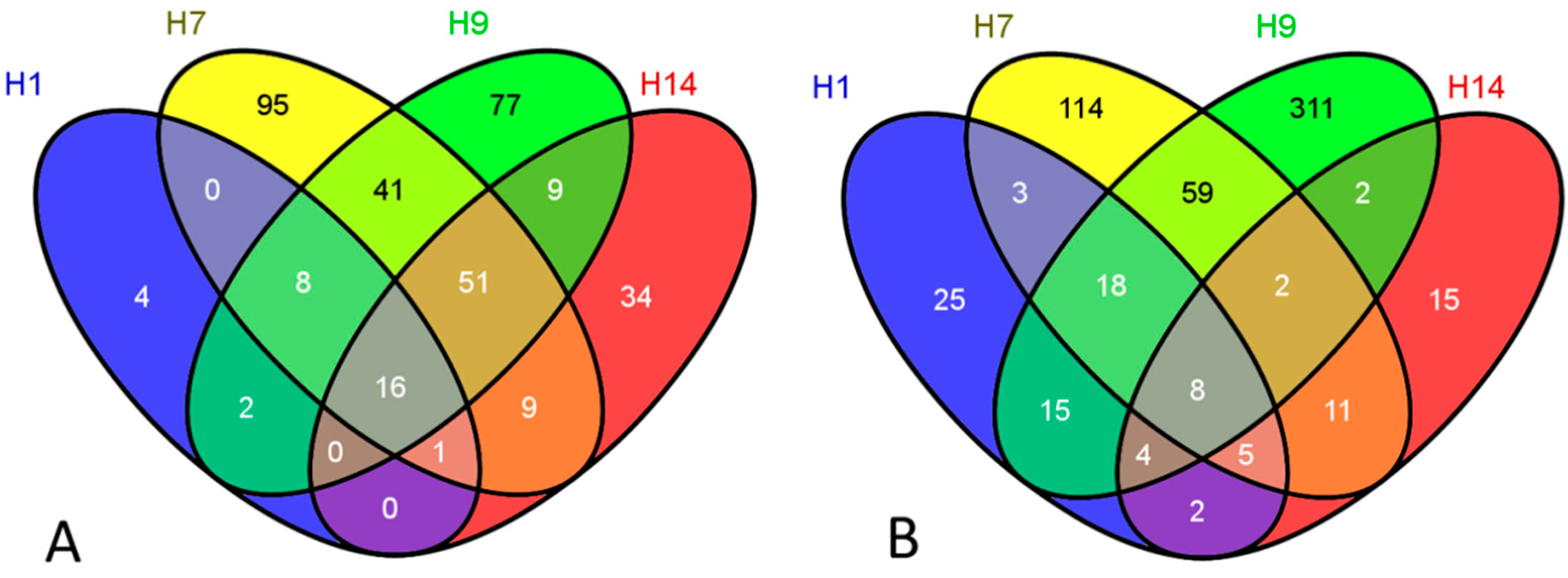
| Exposures | Over-Represented Categories (Upregulation) | EASE Score |
|---|---|---|
| 1 Gy, 2 h | P53 signaling pathway | 7.9 × 10−12 |
| Response to DNA damage stimulus | 1.5 × 10−7 | |
| Positive regulation of apoptosis | 5.6 × 10−7 | |
| Cellular response to stress | 6.7 × 10−6 | |
| Negative regulation of cell proliferation | 1.1 × 10−4 | |
| Cell cycle arrest | 2.8 × 10−4 | |
| Response to radiation | 3.7 × 10−4 | |
| Regulation of protein kinase activity | 4.1 × 10−3 | |
| Regulation of transferase activity | 5.6 × 10−3 | |
| 1 Gy, 16 h | Positive regulation of anti-apoptosis | 4.3 × 10−4 |
| Response to DNA damage stimulus | 6.0 × 10−3 | |
| Cellular response to stress | 0.019 | |
| Exposures | Over-Represented Categories (Downregulation) | EASE Score |
| 1 Gy, 16 h | Cytoplasmic vesicle | 0.033 |
3. Experimental Section
3.1. Cell Culture
3.2. Cell Culture Treatments
3.3. RNA Sample Preparation, Probe Labeling and DNA Microarray Procedure
3.4. Data Statistical Analysis
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Tubiana, M.; Feinendegen, L.E.; Yang, C.; Kaminski, J.M. The linear no-threshold relationship is inconsistent with radiation biologic and experimental data. Radiology 2009, 251, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Dauer, L.T.; Brooks, A.L.; Hoel, D.G.; Morgan, W.F.; Stram, D.; Tran, P. Review and evaluation of updated research on the health effects associated with low-dose ionising radiation. Radiat. Prot. Dosim. 2010, 140, 103–136. [Google Scholar] [CrossRef] [PubMed]
- Feinendegen, L.E.; Pollycove, M.; Neumann, R.D. Low-dose cancer risk modeling must recognize upregulation of protection. Dose Response 2010, 8, 227–252. [Google Scholar] [CrossRef] [PubMed]
- Feinendegen, L.; Neumann, R.D.; Pollycove, M. Systems-related facts and consequences in assessing risk from low-level irradiation. Health Phys. 2011, 100, 274–276. [Google Scholar] [CrossRef] [PubMed]
- Feinendegen, L.E.; Brooks, A.L.; Morgan, W.F. Biological consequences and health risks of low-level exposure to ionizing radiation: Commentary on the workshop. Health Phys. 2011, 100, 247–259. [Google Scholar] [CrossRef] [PubMed]
- Morgan, W.F.; Bair, W.J. Issues in low dose radiation biology: The controversy continues. A perspective. Radiat. Res. 2013, 179, 501–510. [Google Scholar] [CrossRef] [PubMed]
- Brenner, D.J.; Mossman, K.L. Do radiation doses below 1 cGy increase cancer risks? Radiat. Res. 2005, 163, 692–693. [Google Scholar] [PubMed]
- Pearce, M.S.; Salotti, J.A.; Little, M.P.; McHugh, K.; Lee, C.; Kim, K.P.; Howe, N.L.; Ronckers, C.M.; Rajaraman, P.; Sir Craft, A.W.; et al. Radiation exposure from CT scans in childhood and subsequent risk of leukaemia and brain tumours: A retrospective cohort study. Lancet 2012, 380, 499–505. [Google Scholar] [CrossRef]
- Pearce, M.S. Patterns in paediatric CT use: An international and epidemiological perspective. J. Med. Imaging Radiat. Oncol. 2011, 55, 107–109. [Google Scholar] [CrossRef] [PubMed]
- Amundson, S.A.; Do, K.T.; Fornace, A.J., Jr. Induction of stress genes by low doses of γ rays. Radiat. Res. 1999, 152, 225–231. [Google Scholar] [CrossRef] [PubMed]
- Sokolov, M.; Neumann, R. Effects of low doses of ionizing radiation exposures on stress-responsive gene expression in human embryonic stem cells. Int. J. Mol. Sci. 2014, 15, 588–604. [Google Scholar] [CrossRef] [PubMed]
- Sokolov, M.V.; Smirnova, N.A.; Camerini-Otero, R.D.; Neumann, R.D.; Panyutin, I.G. Microarray analysis of differentially expressed genes after exposure of normal human fibroblasts to ionizing radiation from an external source and from DNA-incorporated iodine-125 radionuclide. Gene 2006, 382, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Amundson, S.A.; Lee, R.A.; Koch-Paiz, C.A.; Bittner, M.L.; Meltzer, P.; Trent, J.M.; Fornace, A.J., Jr. Differential responses of stress genes to low dose-rate γ irradiation. Mol. Cancer Res. 2003, 1, 445–452. [Google Scholar] [PubMed]
- Ding, L.H.; Shingyoji, M.; Chen, F.; Hwang, J.J.; Burma, S.; Lee, C.; Cheng, J.F.; Chen, D.J. Gene expression profiles of normal human fibroblasts after exposure to ionizing radiation: A comparative study of low and high doses. Radiat. Res. 2005, 164, 17–26. [Google Scholar] [CrossRef] [PubMed]
- Franco, N.; Lamartine, J.; Frouin, V.; Le Minter, P.; Petat, C.; Leplat, J.J.; Libert, F.; Gidrol, X.; Martin, M.T. Low-dose exposure to γ rays induces specific gene regulations in normal human keratinocytes. Radiat. Res. 2005, 163, 623–635. [Google Scholar] [CrossRef] [PubMed]
- Goldberg, Z.; Rocke, D.M.; Schwietert, C.; Berglund, S.R.; Santana, A.; Jones, A.; Lehmann, J.; Stern, R.; Lu, R.; Hartmann Siantar, C. Human in vivo dose-response to controlled, low-dose low linear energy transfer ionizing radiation exposure. Clin. Cancer Res. 2006, 12, 3723–3729. [Google Scholar] [CrossRef] [PubMed]
- Albrecht, H.; Durbin-Johnson, B.; Yunis, R.; Kalanetra, K.M.; Wu, S.; Chen, R.; Stevenson, T.R.; Rocke, D.M. Transcriptional response of ex vivo human skin to ionizing radiation: Comparison between low- and high-dose effects. Radiat. Res. 2012, 177, 69–83. [Google Scholar] [CrossRef] [PubMed]
- Knops, K.; Boldt, S.; Wolkenhauer, O.; Kriehuber, R. Gene expression in low- and high-dose-irradiated human peripheral blood lymphocytes: Possible applications for biodosimetry. Radiat. Res. 2012, 178, 304–312. [Google Scholar] [CrossRef] [PubMed]
- Wilson, K.D.; Sun, N.; Huang, M.; Zhang, W.Y.; Lee, A.S.; Li, Z.; Wang, S.X.; Wu, J.C. Effects of ionizing radiation on self-renewal and pluripotency of human embryonic stem cells. Cancer Res. 2010, 70, 5539–5548. [Google Scholar] [CrossRef] [PubMed]
- Sokolov, M.V.; Panyutin, I.V.; Panyutin, I.G.; Neumann, R.D. Dynamics of the transcriptome response of cultured human embryonic stem cells to ionizing radiation exposure. Mutat. Res. 2011, 709–710, 40–48. [Google Scholar] [CrossRef]
- Sokolov, M.V.; Panyutin, I.V.; Neumann, R.D. Unraveling the global microRNAome responses to ionizing radiation in human embryonic stem cells. PLoS ONE 2012, 7, e31028. [Google Scholar] [CrossRef] [PubMed]
- Paul, S.; Amundson, S.A. Development of gene expression signatures for practical radiation biodosimetry. Int. J. Radiat. Oncol. Biol. Phys. 2008, 71, 1236–1244. [Google Scholar] [CrossRef] [PubMed]
- El-Saghire, H.; Thierens, H.; Monsieurs, P.; Michaux, A.; Vandevoorde, C.; Baatout, S. Gene set enrichment analysis highlights different gene expression profiles in whole blood samples X-irradiated with low and high doses. Int. J. Radiat. Biol. 2013, 89, 628–638. [Google Scholar] [CrossRef] [PubMed]
- Di Leonardo, A.; Linke, S.P.; Clarkin, K.; Wahl, G.M. DNA damage triggers a prolonged p53-dependent G1 arrest and long-term induction of Cip1 in normal human fibroblasts. Genes Dev. 1994, 8, 2540–2551. [Google Scholar] [CrossRef] [PubMed]
- Becker, K.A.; Ghule, P.N.; Therrien, J.A.; Lian, J.B.; Stein, J.L.; van Wijnen, A.J.; Stein, G.S. Self-renewal of human embryonic stem cells is supported by a shortened G1 cell cycle phase. J. Cell. Physiol. 2006, 209, 883–893. [Google Scholar] [CrossRef] [PubMed]
- Filion, T.M.; Qiao, M.; Ghule, P.N.; Mandeville, M.; van Wijnen, A.J.; Stein, J.L.; Lian, J.B.; Altieri, D.C.; Stein, G.S. Survival responses of human embryonic stem cells to DNA damage. J. Cell. Physiol. 2009, 220, 586–592. [Google Scholar] [CrossRef] [PubMed]
- Rouault, J.P.; Falette, N.; Guehenneux, F.; Guillot, C.; Rimokh, R.; Wang, Q.; Berthet, C.; Moyret-Lalle, C.; Savatier, P.; Pain, B.; et al. Identification of BTG2, an antiproliferative p53-dependent component of the DNA damage cellular response pathway. Nat. Genet. 1996, 14, 482–486. [Google Scholar] [CrossRef] [PubMed]
- Okazaki, R.; Moon, Y.; Norimura, T.; Eling, T. Ionizing radiation enhances the expression of the nonsteroidal anti-inflammatory drug-activated gene (NAG1) by increasing the expression of TP53 in human colon cancer cells. Radiat. Res. 2006, 165, 125–130. [Google Scholar] [CrossRef] [PubMed]
- Kabacik, S.; Mackay, A.; Tamber, N.; Manning, G.; Finnon, P.; Paillier, F.; Ashworth, A.; Bouffler, S.; Badie, C. Gene expression following ionising radiation: Identification of biomarkers for dose estimation and prediction of individual response. Int. J. Radiat. Biol. 2011, 87, 115–129. [Google Scholar] [CrossRef] [PubMed]
- Kis, E.; Szatmari, T.; Keszei, M.; Farkas, R.; Esik, O.; Lumniczky, K.; Falus, A.; Safrany, G. Microarray analysis of radiation response genes in primary human fibroblasts. Int. J. Radiat. Oncol. Biol. Phys. 2006, 66, 1506–1514. [Google Scholar] [CrossRef] [PubMed]
- Kelly, J.A.; Lucia, M.S.; Lambert, J.R. p53 controls prostate-derived factor/macrophage inhibitory cytokine/NSAID-activated gene expression in response to cell density, DNA damage and hypoxia through diverse mechanisms. Cancer Lett. 2009, 277, 38–47. [Google Scholar] [CrossRef] [PubMed]
- Qian, Y.; Jung, Y.S.; Chen, X. Differentiated embryo-chondrocyte expressed gene 1 regulates p53-dependent cell survival versus cell death through macrophage inhibitory cytokine-1. Proc. Natl. Acad. Sci. USA 2012, 109, 11300–11305. [Google Scholar] [CrossRef] [PubMed]
- Tucker, J.D.; Joiner, M.C.; Thomas, R.A.; Grever, W.E.; Bakhmutsky, M.V.; Chinkhota, C.N.; Smolinski, J.M.; Divine, G.W.; Auner, G.W. Accurate gene expression-based biodosimetry using a minimal set of human gene transcripts. Int. J. Radiat. Oncol. Biol. Phys. 2014, 88, 933–939. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.S.; Kim, J.Y.; Lim, S.K.; Choi, Y.W.; Kim, Y.H.; Kang, S.Y.; Park, T.J.; Lim, I.K. TIS21(/BTG2/PC3) accelerates the repair of DNA double strand breaks by enhancing Mre11 methylation and blocking damage signal transfer to the Chk2(T68)-p53(S20) pathway. DNA Repair 2012, 11, 965–975. [Google Scholar] [CrossRef] [PubMed]
- Sokolov, M.V.; Neumann, R.D.; Panyutin, I.G. Effects of DNA-targeted ionizing radiation produced by 5-(125I)iodo-2′-deoxyuridine on global gene expression in primary human cells. BMC Genomics 2007, 8, 192. [Google Scholar] [CrossRef] [PubMed]
- Li, J.J.; Biggin, M.D. Gene expression. Statistics requantitates the central dogma. Science 2015, 347, 1066–1067. [Google Scholar] [CrossRef]
- Sokolov, M.V.; Panyutin, I.V.; Onyshchenko, M.I.; Panyutin, I.G.; Neumann, R.D. Expression of pluripotency-associated genes in the surviving fraction of cultured human embryonic stem cells is not significantly affected by ionizing radiation. Gene 2010, 455, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Sokolov, M.; Panyutin, I.G.; Neumann, R. Genome-wide gene expression changes in normal human fibroblasts in response to low-LET γ-radiation and high-LET-like 125IUdR exposures. Radiat. Prot. Dosim. 2006, 122, 195–201. [Google Scholar] [CrossRef] [PubMed]
- Sokolov, M.V.; Panyutin, I.G.; Neumann, R.D. Whole-genome gene expression profiling reveals the major role of nitric oxide in mediating the cellular transcriptional response to ionizing radiation in normal human fibroblasts. Genomics 2012, 100, 277–281. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sokolov, M.; Nguyen, V.; Neumann, R. Comparative Analysis of Whole-Genome Gene Expression Changes in Cultured Human Embryonic Stem Cells in Response to Low, Clinical Diagnostic Relevant, and High Doses of Ionizing Radiation Exposure. Int. J. Mol. Sci. 2015, 16, 14737-14748. https://doi.org/10.3390/ijms160714737
Sokolov M, Nguyen V, Neumann R. Comparative Analysis of Whole-Genome Gene Expression Changes in Cultured Human Embryonic Stem Cells in Response to Low, Clinical Diagnostic Relevant, and High Doses of Ionizing Radiation Exposure. International Journal of Molecular Sciences. 2015; 16(7):14737-14748. https://doi.org/10.3390/ijms160714737
Chicago/Turabian StyleSokolov, Mykyta, Van Nguyen, and Ronald Neumann. 2015. "Comparative Analysis of Whole-Genome Gene Expression Changes in Cultured Human Embryonic Stem Cells in Response to Low, Clinical Diagnostic Relevant, and High Doses of Ionizing Radiation Exposure" International Journal of Molecular Sciences 16, no. 7: 14737-14748. https://doi.org/10.3390/ijms160714737
APA StyleSokolov, M., Nguyen, V., & Neumann, R. (2015). Comparative Analysis of Whole-Genome Gene Expression Changes in Cultured Human Embryonic Stem Cells in Response to Low, Clinical Diagnostic Relevant, and High Doses of Ionizing Radiation Exposure. International Journal of Molecular Sciences, 16(7), 14737-14748. https://doi.org/10.3390/ijms160714737





