MicroRNA-24 Regulates Osteogenic Differentiation via Targeting T-Cell Factor-1
Abstract
:1. Introduction
2. Results
2.1. miR-24 Is Involved in Osteogenic Differentiation
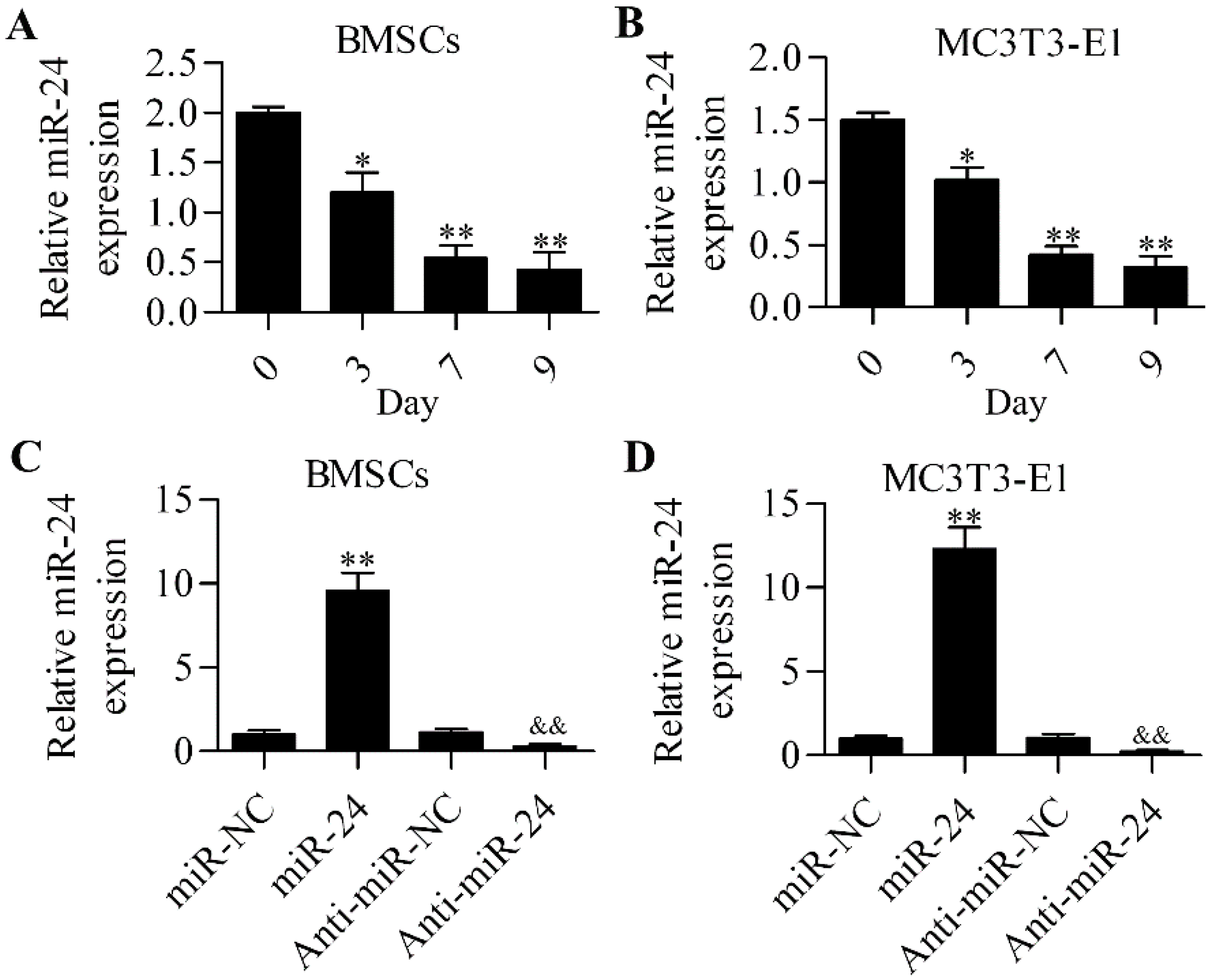
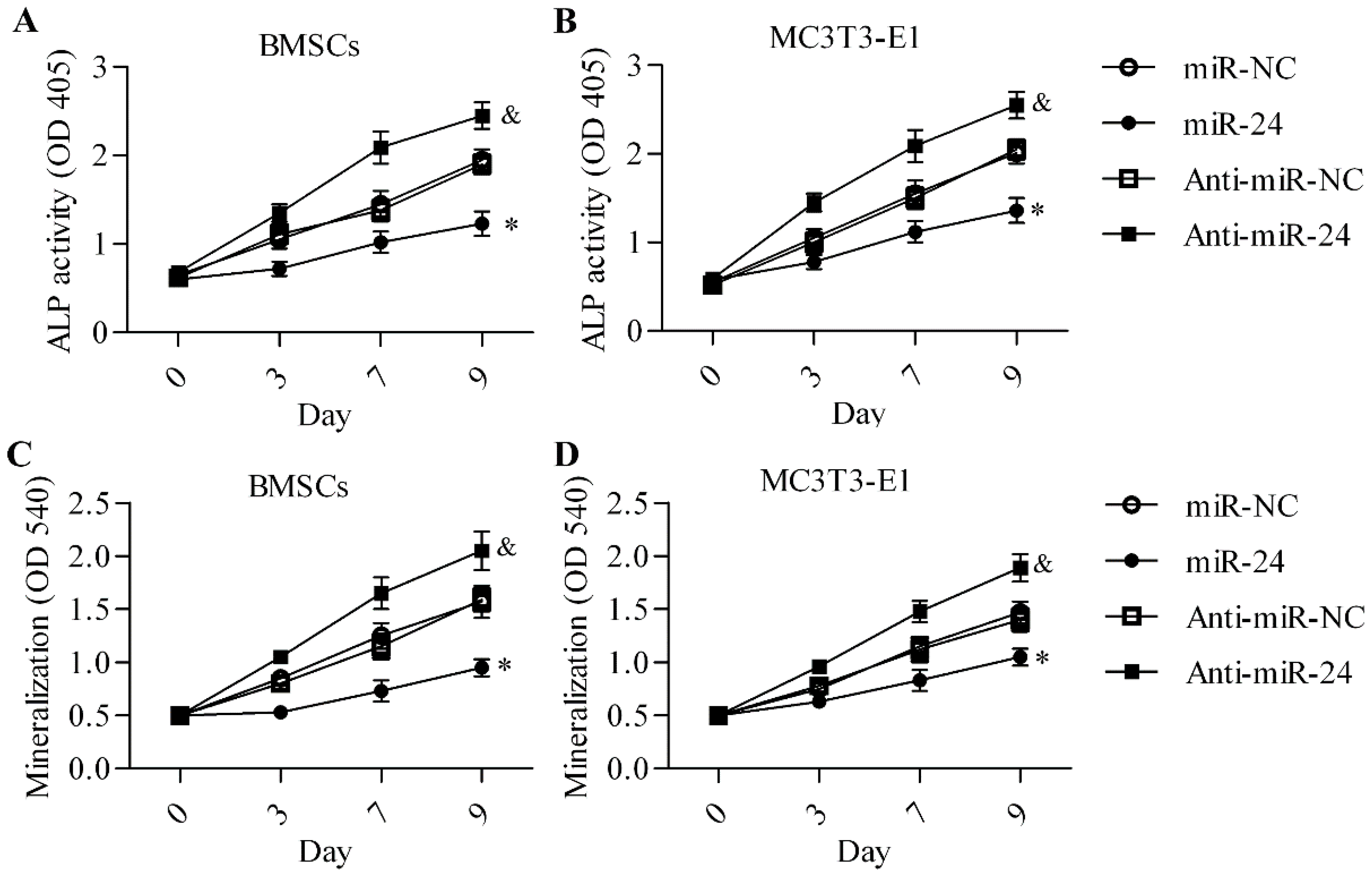
2.2. Inhibition of miR-24 Elevates the Expression of Osteogenic Differentiation Markers
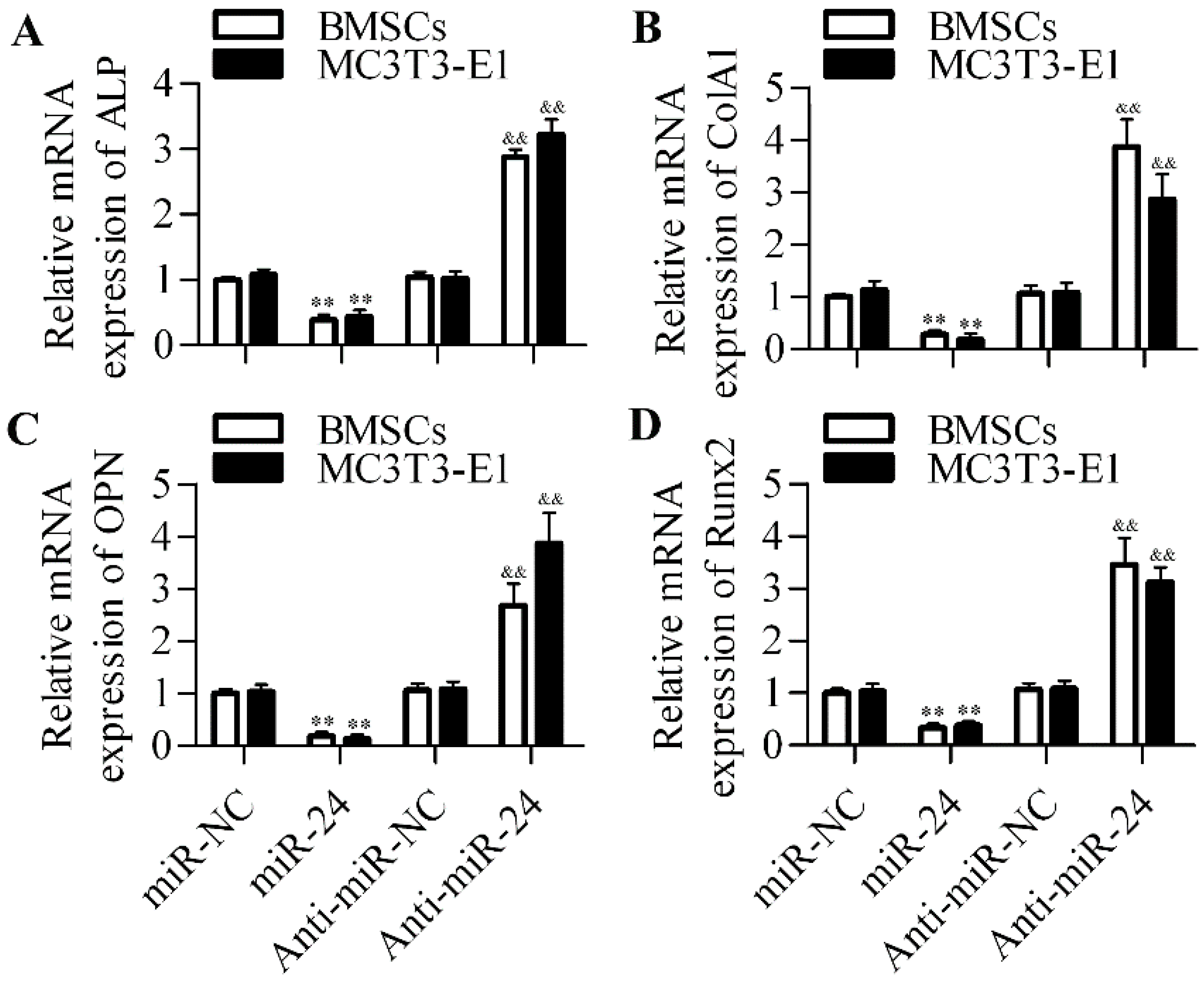
2.3. miR-24 Directly Targets the 3′-UTR of Tcf-1
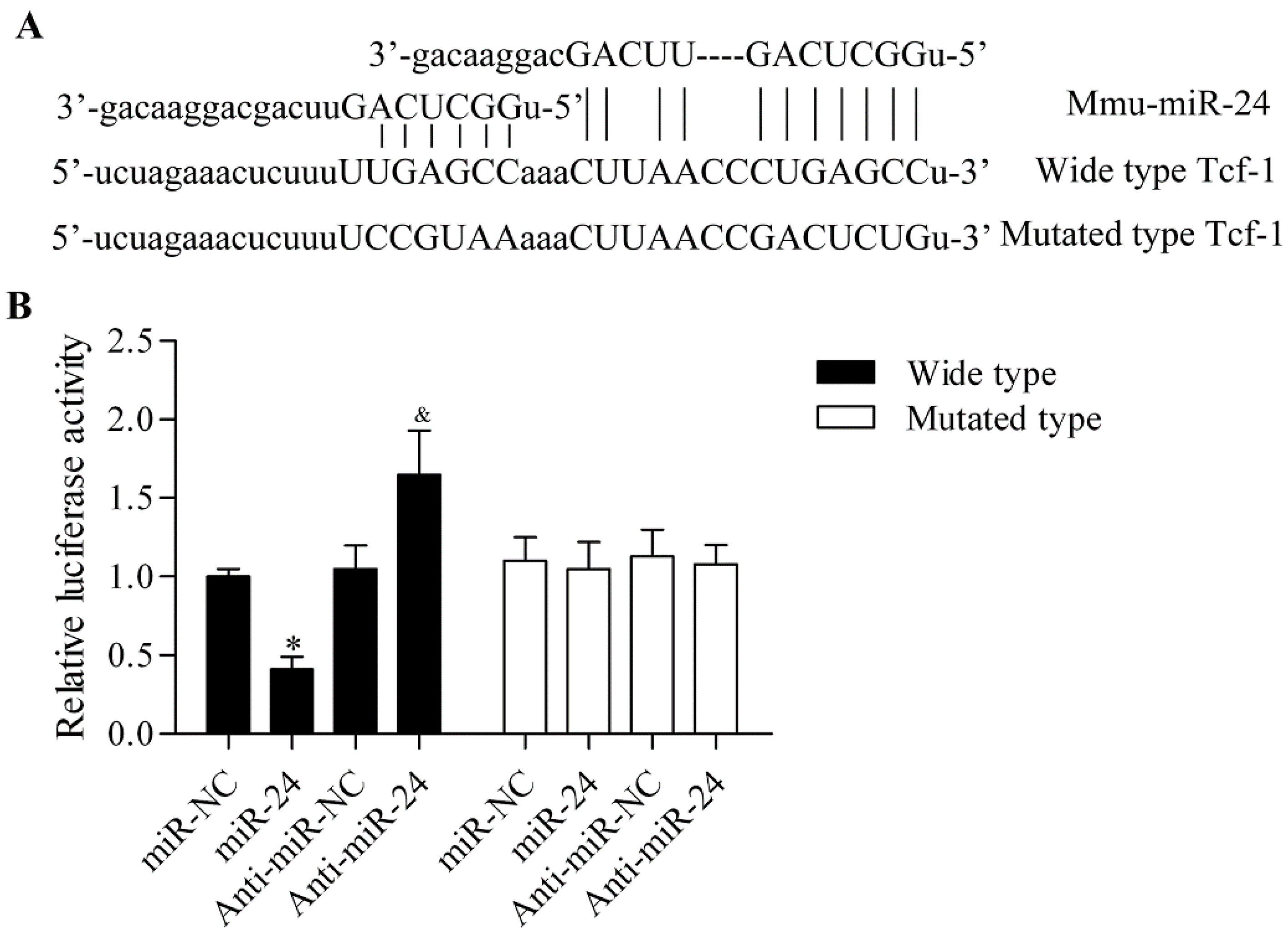
2.4. miR-24 Regulates Tcf-1 Expression in BMSCs and MC3T3-E1 Cells
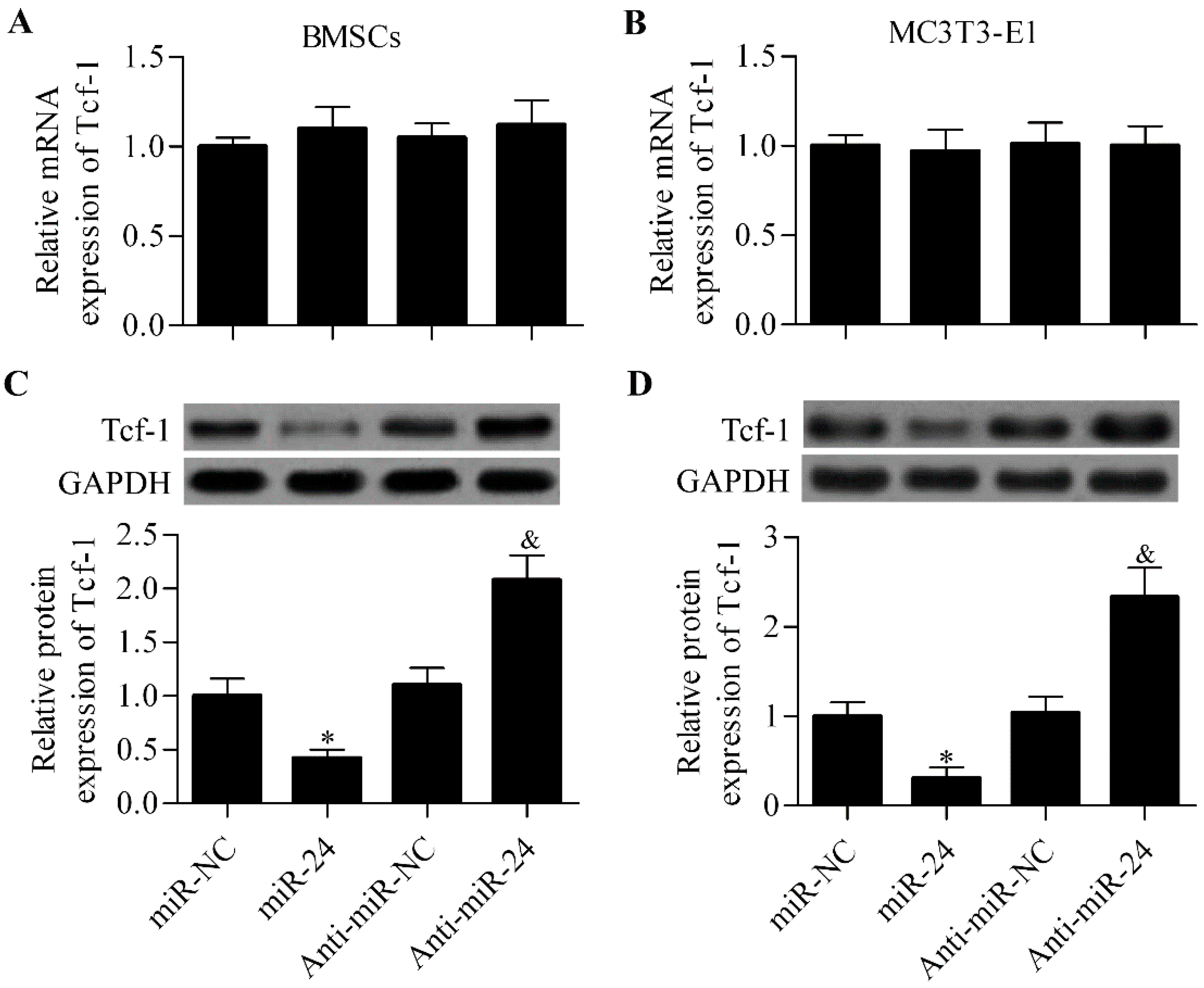
2.5. Silencing of Tcf-1 Abolishes the Positive Effect of miR-24 Inhibition on Osteoblast Differentiation
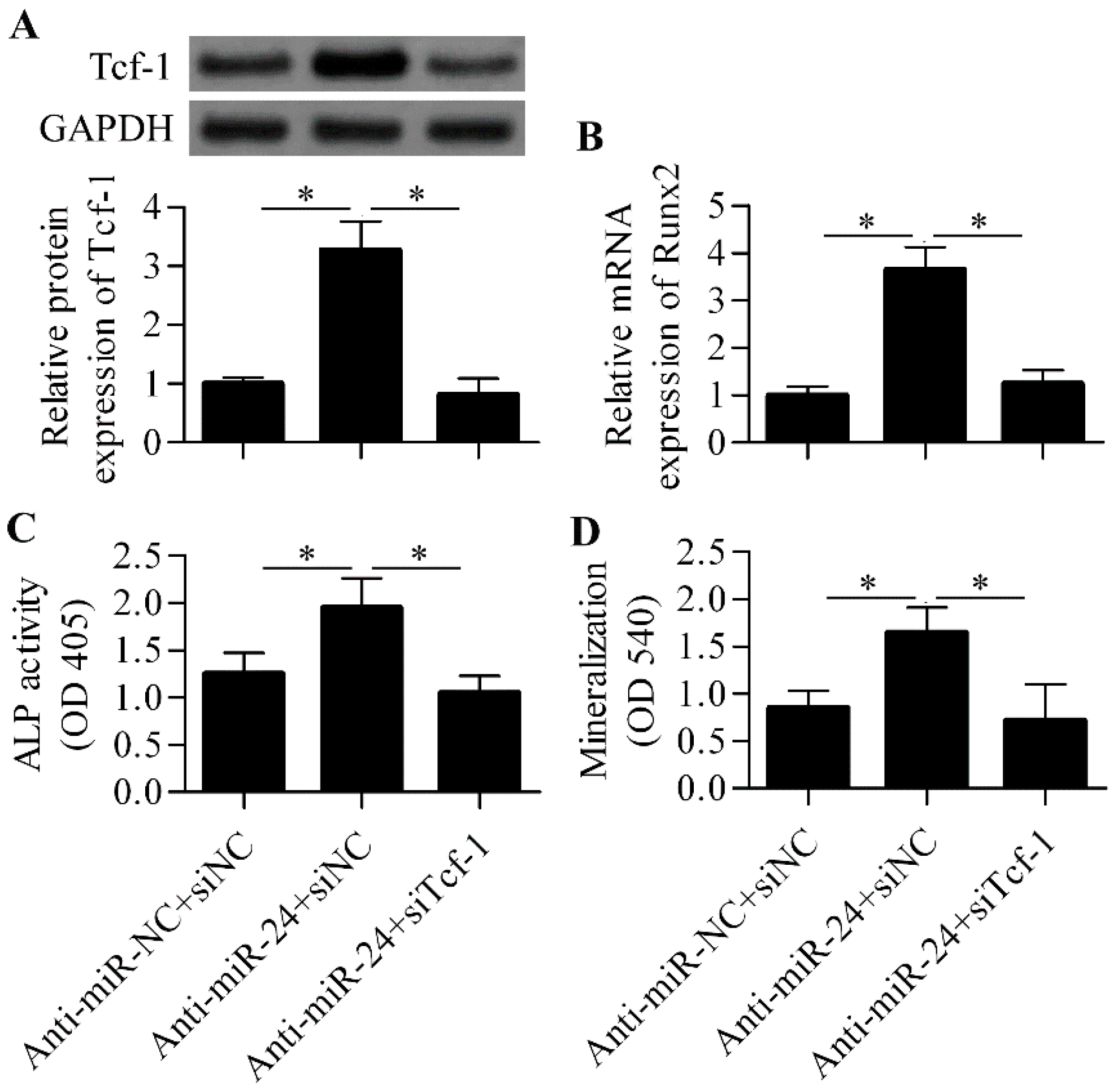
3. Discussion
4. Experimental Section
4.1. Cell Cultures
4.2. Real-Time Quantitative Polymerase Chain Reaction (RT-qPCR)
4.3. Alkaline Phosphatase (ALP) Activity
4.4. Measurement of Matrix Mineralization
4.5. Western Blot Analysis
4.6. Dual Luciferase Reporter Assay
4.7. Small Interfering RNA (siRNA) Transfection
4.8. Data Analysis
5. Conclusions
Author Contributions
Conflicts of Interest
References
- Franceschi, R.T. The developmental control of osteoblast-specific gene expression: Role of specific transcription factors and the extracellular matrix environment. Crit. Rev. Oral Biol. Med. 1999, 10, 40–57. [Google Scholar] [CrossRef] [PubMed]
- Rodan, G.A.; Martin, T.J. Therapeutic approaches to bone diseases. Science 2000, 289, 1508–1514. [Google Scholar] [CrossRef] [PubMed]
- Lo, Y.C.; Chang, Y.H.; Wei, B.L.; Huang, Y.L.; Chiou, W.F. Betulinic acid stimulates the differentiation and mineralization of osteoblastic mc3t3-e1 cells: Involvement of bmp/runx2 and beta-catenin signals. J. Agric. Food Chem. 2010, 58, 6643–6649. [Google Scholar] [CrossRef] [PubMed]
- Canalis, E.; Economides, A.N.; Gazzerro, E. Bone morphogenetic proteins, their antagonists, and the skeleton. Endocr. Rev. 2003, 24, 218–235. [Google Scholar] [CrossRef] [PubMed]
- Hughes, F.J.; Turner, W.; Belibasakis, G.; Martuscelli, G. Effects of growth factors and cytokines on osteoblast differentiation. Periodontology 2000 2006, 41, 48–72. [Google Scholar] [CrossRef] [PubMed]
- Mendell, J.T.; Olson, E.N. Micrornas in stress signaling and human disease. Cell 2012, 148, 1172–1187. [Google Scholar] [CrossRef] [PubMed]
- Ranganathan, K.; Sivasankar, V. Micrornas—Biology and clinical applications. J. Oral Maxillofac. Pathol. 2014, 18, 229–234. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. Micrornas: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [PubMed]
- Winter, J.; Jung, S.; Keller, S.; Gregory, R.I.; Diederichs, S. Many roads to maturity: Microrna biogenesis pathways and their regulation. Nat. Cell Biol. 2009, 11, 228–234. [Google Scholar] [CrossRef] [PubMed]
- Vimalraj, S.; Selvamurugan, N. Micrornas: Synthesis, gene regulation and osteoblast differentiation. Curr. Issues Mol. Biol. 2012, 15, 7–18. [Google Scholar] [PubMed]
- Hassan, M.Q.; Gordon, J.A.; Beloti, M.M.; Croce, C.M.; van Wijnen, A.J.; Stein, J.L.; Stein, G.S.; Lian, J.B. A network connecting Runx2, SATB2, and the miR-23a~27a~24-2 cluster regulates the osteoblast differentiation program. Proc. Natl. Acad. Sci. USA 2010, 107, 19879–19884. [Google Scholar] [CrossRef] [PubMed]
- Hu, R.; Li, H.; Liu, W.; Yang, L.; Tan, Y.F.; Luo, X.H. Targeting mirnas in osteoblast differentiation and bone formation. Expert Opin. Ther. Targets 2010, 14, 1109–1120. [Google Scholar] [CrossRef] [PubMed]
- Arfat, Y.; Xiao, W.Z.; Ahmad, M.; Zhao, F.; Li, D.J.; Sun, Y.L.; Hu, L.; Zhihao, C.; Zhang, G.; Iftikhar, S.; et al. Role of micrornas in osteoblasts differentiation and bone disorders. Curr. Med. Chem. 2015, 22, 748–758. [Google Scholar] [CrossRef] [PubMed]
- Mizuno, Y.; Tokuzawa, Y.; Ninomiya, Y.; Yagi, K.; Yatsuka-Kanesaki, Y.; Suda, T.; Fukuda, T.; Katagiri, T.; Kondoh, Y.; Amemiya, T.; et al. Mir-210 promotes osteoblastic differentiation through inhibition of AcvR1b. FEBS Lett. 2009, 583, 2263–2268. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Pan, Y.; Zhou, Y. Mir-96 promotes osteogenic differentiation by suppressing HBEGF-EGFR signaling in osteoblastic cells. FEBS Lett. 2014, 588, 4761–4768. [Google Scholar] [CrossRef] [PubMed]
- Kang, I.H.; Jeong, B.C.; Hur, S.W.; Choi, H.; Choi, S.H.; Ryu, J.H.; Hwang, Y.C.; Koh, J.T. MicroRNA-302a stimulates osteoblastic differentiation by repressing COUP-TFII expression. J. Cell Physiol. 2015, 230, 911–921. [Google Scholar] [CrossRef] [PubMed]
- Logan, C.Y.; Nusse, R. The wnt signaling pathway in development and disease. Annu. Rev. Cell Dev. Biol. 2004, 20, 781–810. [Google Scholar] [CrossRef] [PubMed]
- Travis, A.; Amsterdam, A.; Belanger, C.; Grosschedl, R. LEF-1, a gene encoding a lymphoid-specific protein with an HMG domain, regulates T-cell receptor alpha enhancer function [corrected]. Genes Dev. 1991, 5, 880–894. [Google Scholar] [CrossRef] [PubMed]
- Van de Wetering, M.; Oosterwegel, M.; Dooijes, D.; Clevers, H. Identification and cloning of TCF-1, a T lymphocyte-specific transcription factor containing a sequence-specific HMG box. EMBO J. 1991, 10, 123–132. [Google Scholar]
- Leclerc, N.; Noh, T.; Cogan, J.; Samarawickrama, D.B.; Smith, E.; Frenkel, B. Opposing effects of glucocorticoids and Wnt signaling on Krox20 and mineral deposition in osteoblast cultures. J. Cell. Biochem. 2008, 103, 1938–1951. [Google Scholar] [CrossRef] [PubMed]
- Armstrong, V.J.; Muzylak, M.; Sunters, A.; Zaman, G.; Saxon, L.K.; Price, J.S.; Lanyon, L.E. Wnt/β-catenin signaling is a component of osteoblastic bone cell early responses to load-bearing and requires estrogen receptor alpha. J. Biol. Chem. 2007, 282, 20715–20727. [Google Scholar] [CrossRef] [PubMed]
- Gaur, T.; Lengner, C.J.; Hovhannisyan, H.; Bhat, R.A.; Bodine, P.V.; Komm, B.S.; Javed, A.; van Wijnen, A.J.; Stein, J.L.; Stein, G.S.; et al. Canonical wnt signaling promotes osteogenesis by directly stimulating Runx2 gene expression. J. Biol. Chem. 2005, 280, 33132–33140. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Zhao, M.; Mundy, G.R. Bone morphogenetic proteins. Growth Factors 2004, 22, 233–241. [Google Scholar] [CrossRef] [PubMed]
- Amelio, I.; Lena, A.M.; Viticchie, G.; Shalom-Feuerstein, R.; Terrinoni, A.; Dinsdale, D.; Russo, G.; Fortunato, C.; Bonanno, E.; Spagnoli, L.G.; et al. Mir-24 triggers epidermal differentiation by controlling actin adhesion and cell migration. J. Cell Biol. 2012, 199, 347–363. [Google Scholar] [CrossRef] [PubMed]
- Lal, A.; Pan, Y.; Navarro, F.; Dykxhoorn, D.M.; Moreau, L.; Meire, E.; Bentwich, Z.; Lieberman, J.; Chowdhury, D. Mir-24-mediated downregulation of H2AX suppresses DNA repair in terminally differentiated blood cells. Nat. Struct. Mol. Biol. 2009, 16, 492–498. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Zhang, Y.; Yang, G.; Chen, X.; Cao, G.; Wang, J.; Sun, Y.; Zhang, P.; Fan, M.; Shao, N.; et al. Transforming growth factor-β-regulated mir-24 promotes skeletal muscle differentiation. Nucleic Acids Res. 2008, 36, 2690–2699. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.S.; Jung, E.Y.; Bae, S.H.; Kwon, K.H.; Kim, J.M.; Suh, H.J. Stimulation of osteoblastic differentiation and mineralization in MC3T3-E1 cells by yeast hydrolysate. Phytother. Res. 2011, 25, 716–723. [Google Scholar] [CrossRef] [PubMed]
- Roberto, V.P.; Tiago, D.M.; Silva, I.A.; Cancela, M.L. Mir-29a is an enhancer of mineral deposition in bone-derived systems. Arch. Biochem. Biophys. 2014, 564, 173–183. [Google Scholar] [CrossRef] [PubMed]
- Hwang, S.; Park, S.K.; Lee, H.Y.; Kim, S.W.; Lee, J.S.; Choi, E.K.; You, D.; Kim, C.S.; Suh, N. Mir-140-5p suppresses BMP2-mediated osteogenesis in undifferentiated human mesenchymal stem cells. FEBS Lett. 2014, 588, 2957–2963. [Google Scholar] [CrossRef] [PubMed]
- Zuo, B.; Zhu, J.; Li, J.; Wang, C.; Zhao, X.; Cai, G.; Li, Z.; Peng, J.; Wang, P.; Shen, C.; et al. MicroRNA-103a functions as a mechanosensitive microrna to inhibit bone formation through targeting Runx2. J. Bone Miner. Res. 2015, 30, 330–345. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; She, X.G.; Ming, Y.Z.; Wan, Q.Q. Mir-24 promotes the proliferation and invasion of HCC cells by targeting SOX7. Tumour Biol. 2014, 35, 10731–10736. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.; Hu, L.; Liu, B.; Yu, B.; Li, J.; Yan, M.; Yu, Y.; Li, C.; Su, L.; Zhu, Z.; et al. Tumor suppressor mir-24 restrains gastric cancer progression by downregulating regiv. Mol. Cancer 2014, 13, 127. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Qian, L. Mir-24 regulates intrinsic apoptosis pathway in mouse cardiomyocytes. PLoS ONE 2014, 9, e85389. [Google Scholar] [CrossRef] [PubMed]
- Fiedler, J.; Stohr, A.; Gupta, S.K.; Hartmann, D.; Holzmann, A.; Just, A.; Hansen, A.; Hilfiker-Kleiner, D.; Eschenhagen, T.; Thum, T. Functional microrna library screening identifies the hypoxamir mir-24 as a potent regulator of smooth muscle cell proliferation and vascularization. Antioxid. Redox Signal. 2014, 21, 1167–1176. [Google Scholar] [CrossRef] [PubMed]
- Naqvi, A.R.; Fordham, J.B.; Nares, S. Mir-24, mir-30b, and mir-142-3p regulate phagocytosis in myeloid inflammatory cells. J. Immunol. 2015, 194, 1916–1927. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Yao, N.; Liu, G.; Dong, L.; Liu, Y.; Zhang, M.; Wang, F.; Wang, B.; Wei, X.; Dong, H.; et al. Functional screen reveals essential roles of mir-27a/24 in differentiation of embryonic stem cells. EMBO J. 2015, 34, 361–378. [Google Scholar] [CrossRef] [PubMed]
- Philipot, D.; Guerit, D.; Platano, D.; Chuchana, P.; Olivotto, E.; Espinoza, F.; Dorandeu, A.; Pers, Y.M.; Piette, J.; Borzi, R.M.; et al. P16INK4a and its regulator mir-24 link senescence and chondrocyte terminal differentiation-associated matrix remodeling in osteoarthritis. Arthritis Res. Ther. 2014, 16, R58. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Huang, W.; Xu, R.; Nie, Y.; Cao, X.; Meng, J.; Xu, X.; Hu, S.; Zheng, Z. Microrna-24 regulates cardiac fibrosis after myocardial infarction. J. Cell Mol. Med. 2012, 16, 2150–2160. [Google Scholar] [CrossRef] [PubMed]
- Kang, M.; Yan, L.M.; Li, Y.M.; Zhang, W.Y.; Wang, H.; Tang, A.Z.; Ou, H.S. Inhibitory effect of microRNA-24 on fatty acid-binding protein expression on 3T3-L1 adipocyte differentiation. Genet. Mol. Res. 2013, 12, 5267–5277. [Google Scholar] [CrossRef] [PubMed]
- Vimalraj, S.; Selvamurugan, N. Micrornas expression and their regulatory networks during mesenchymal stem cells differentiation toward osteoblasts. Int. J. Biol. Macromol. 2014, 66, 194–202. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Cai, J.; Cai, X.H.; Chen, L. Mir-346 regulates osteogenic differentiation of human bone marrow-derived mesenchymal stem cells by targeting the wnt/β-catenin pathway. PLoS ONE 2013, 8, e72266. [Google Scholar] [CrossRef] [PubMed]
- Hassan, M.Q.; Maeda, Y.; Taipaleenmaki, H.; Zhang, W.; Jafferji, M.; Gordon, J.A.; Li, Z.; Croce, C.M.; van Wijnen, A.J.; Stein, J.L.; et al. Mir-218 directs a wnt signaling circuit to promote differentiation of osteoblasts and osteomimicry of metastatic cancer cells. J. Biol. Chem. 2012, 287, 42084–42092. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Tu, Q.; Bonewald, L.F.; He, X.; Stein, G.; Lian, J.; Chen, J. Effects of mir-335-5p in modulating osteogenic differentiation by specifically downregulating wnt antagonist DKK1. J. Bone Miner. Res. 2011, 26, 1953–1963. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Xu, Z. Mir-27 promotes osteoblast differentiation by modulating wnt signaling. Biochem. Biophys. Res. Commun. 2010, 402, 186–189. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Guan, X.; Guo, F.; Zhou, J.; Chang, A.; Sun, B.; Cai, Y.; Ma, Z.; Dai, C.; Li, X.; et al. Mir-30e reciprocally regulates the differentiation of adipocytes and osteoblasts by directly targeting low-density lipoprotein receptor-related protein 6. Cell Death Dis. 2013, 4, e845. [Google Scholar] [CrossRef] [PubMed]
- Staal, F.J.; Meeldijk, J.; Moerer, P.; Jay, P.; van de Weerdt, B.C.; Vainio, S.; Nolan, G.P.; Clevers, H. Wnt signaling is required for thymocyte development and activates TCF-1 mediated transcription. Eur. J. Immunol. 2001, 31, 285–293. [Google Scholar] [CrossRef] [PubMed]
- Glass, D.A., 2nd; Bialek, P.; Ahn, J.D.; Starbuck, M.; Patel, M.S.; Clevers, H.; Taketo, M.M.; Long, F.; McMahon, A.P.; Lang, R.A.; et al. Canonical wnt signaling in differentiated osteoblasts controls osteoclast differentiation. Dev. Cell 2005, 8, 751–764. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, W.; Wu, C.; Dong, Y.; Ma, Y.; Jin, Y.; Ji, Y. MicroRNA-24 Regulates Osteogenic Differentiation via Targeting T-Cell Factor-1. Int. J. Mol. Sci. 2015, 16, 11699-11712. https://doi.org/10.3390/ijms160511699
Zhao W, Wu C, Dong Y, Ma Y, Jin Y, Ji Y. MicroRNA-24 Regulates Osteogenic Differentiation via Targeting T-Cell Factor-1. International Journal of Molecular Sciences. 2015; 16(5):11699-11712. https://doi.org/10.3390/ijms160511699
Chicago/Turabian StyleZhao, Weigong, Caijun Wu, Yanying Dong, Yunfeng Ma, Yaofeng Jin, and Yanhong Ji. 2015. "MicroRNA-24 Regulates Osteogenic Differentiation via Targeting T-Cell Factor-1" International Journal of Molecular Sciences 16, no. 5: 11699-11712. https://doi.org/10.3390/ijms160511699
APA StyleZhao, W., Wu, C., Dong, Y., Ma, Y., Jin, Y., & Ji, Y. (2015). MicroRNA-24 Regulates Osteogenic Differentiation via Targeting T-Cell Factor-1. International Journal of Molecular Sciences, 16(5), 11699-11712. https://doi.org/10.3390/ijms160511699




