Genetic Variant rs10757278 on Chromosome 9p21 Contributes to Myocardial Infarction Susceptibility
Abstract
:1. Introduction
2. Results
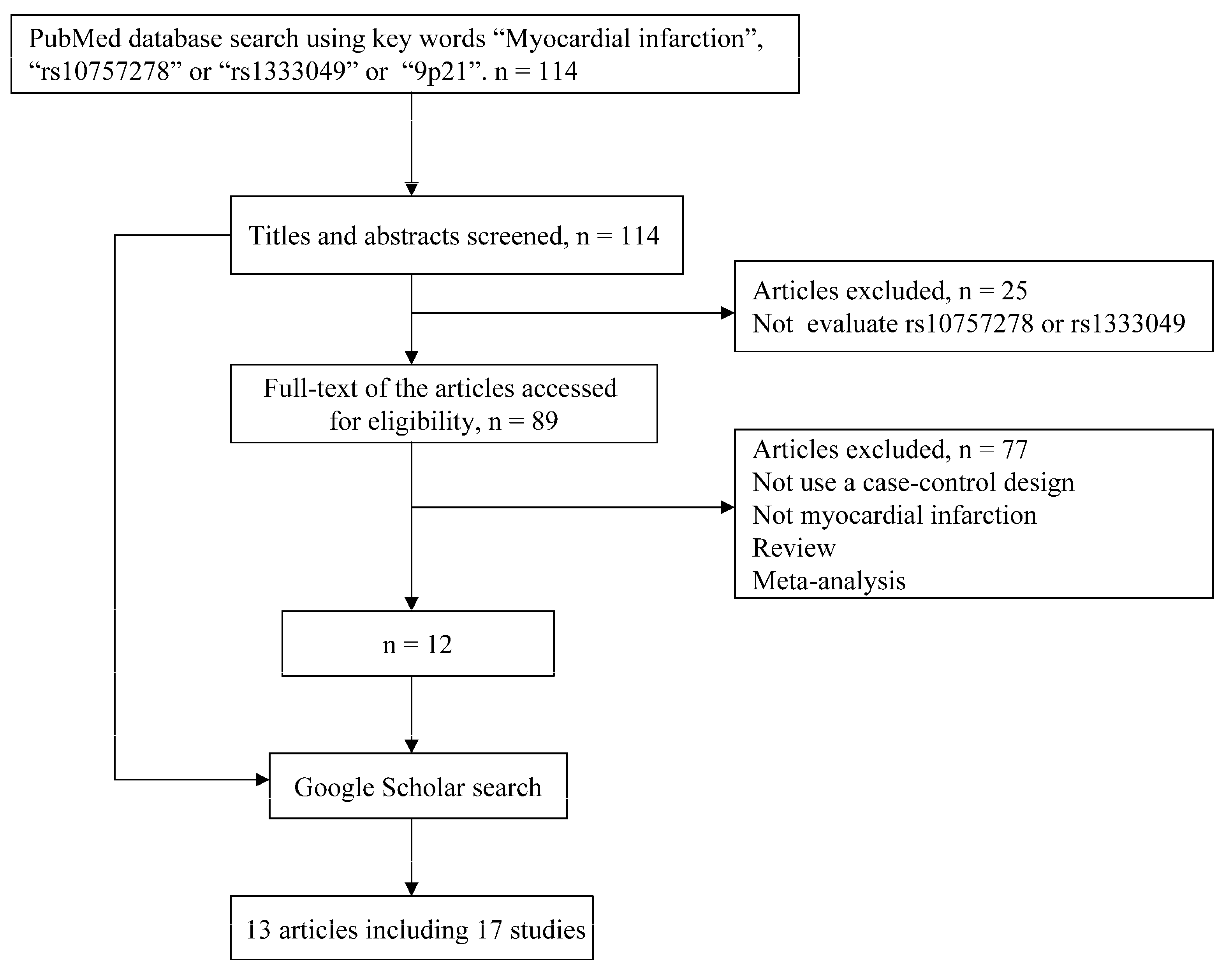
2.1. Literature Search
| Study | SNP/Risk Allele | Country | Ethnicity | Case # | Control # | Quality Score | Genotyping Platform |
|---|---|---|---|---|---|---|---|
| [4] | rs10757278/A | China | Asian | 432 | 430 | 8 | GenomeLab SNPstream |
| [9] | rs1333049/C | China | Asian | 425 | 1377 | 8 | TaqMan |
| [16] | rs1333049/C | China | Asian | 142 | 192 | 8 | PCR |
| [17] | rs1333049/C | China | Asian | 520 | 560 | 8 | NA |
| [18] | rs10757278/C | China | Asian | 1515 | 5019 | 8 | NA |
| [5] | rs1333049/C | Japan | Asian | 589 | 2475 | 9 | MALDI-TOF MS |
| [10] | rs10757278/A | India | Asian | 87 | 150 | 8 | PCR |
| [6] | rs1333049/C | Pakistan | Asian | 2587 | 2573 | 8 | NA |
| [11] | rs10757278/A | Russia | Siberian | 197 | 417 | 8 | NA |
| [12] | rs10757278/G | Italy | Caucasian | 416 | 308 | 8 | ABI PRISM 7900HT |
| [7] | rs10757278/G | Germany | Caucasian | 3657 | 1211 | 9 | TaqMan |
| [3] | rs10757278/G | Iceland (discovery) | Caucasian | 1067 | 6728 | 9 | IlluminaHap300 |
| [3] | rs10757278/G | Iceland (replication) | Caucasian | 665 | 3533 | 9 | IlluminaHap300 |
| [3] | rs10757278/G | United States (Atlanta) | Caucasian | 596 | 1284 | 9 | IlluminaHap300 |
| [3] | rs10757278/G | United States (Philadelphia) | Caucasian | 582 | 504 | 9 | IlluminaHap300 |
| [3] | rs10757278/G | United States (Durham) | Caucasian | 1137 | 718 | 9 | IlluminaHap300 |
| [8] | rs10757278/G | United States | Caucasian | 310 | 560 | 9 | TaqMan |
| n = 14,924 | n = 28,039 |
2.2. Heterogeneity Test and Meta-Analysis
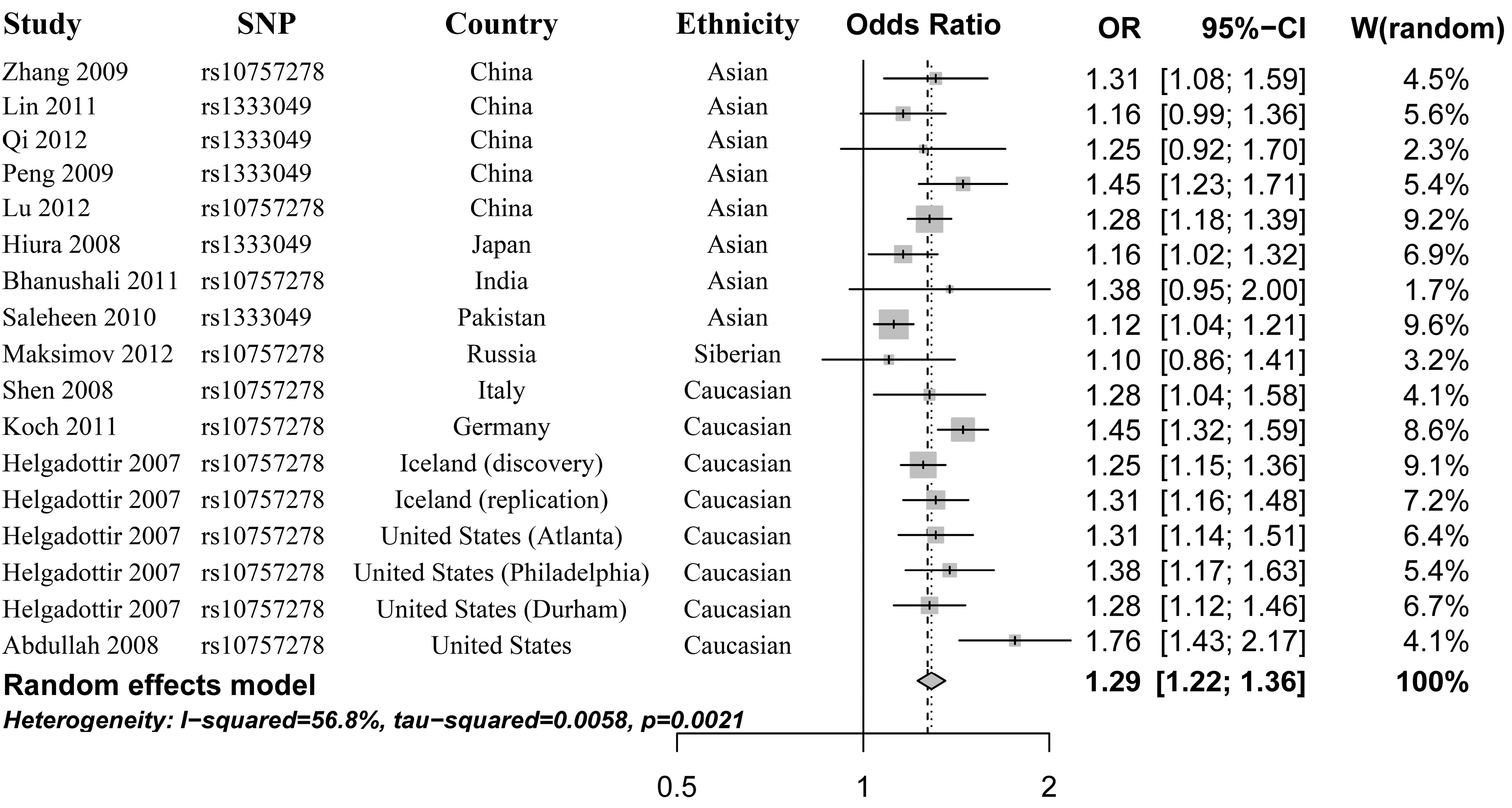
2.3. Heterogeneity Test and Subgroup Analysis
2.4. Sensitivity Analysis
2.5. Publication Bias Analysis
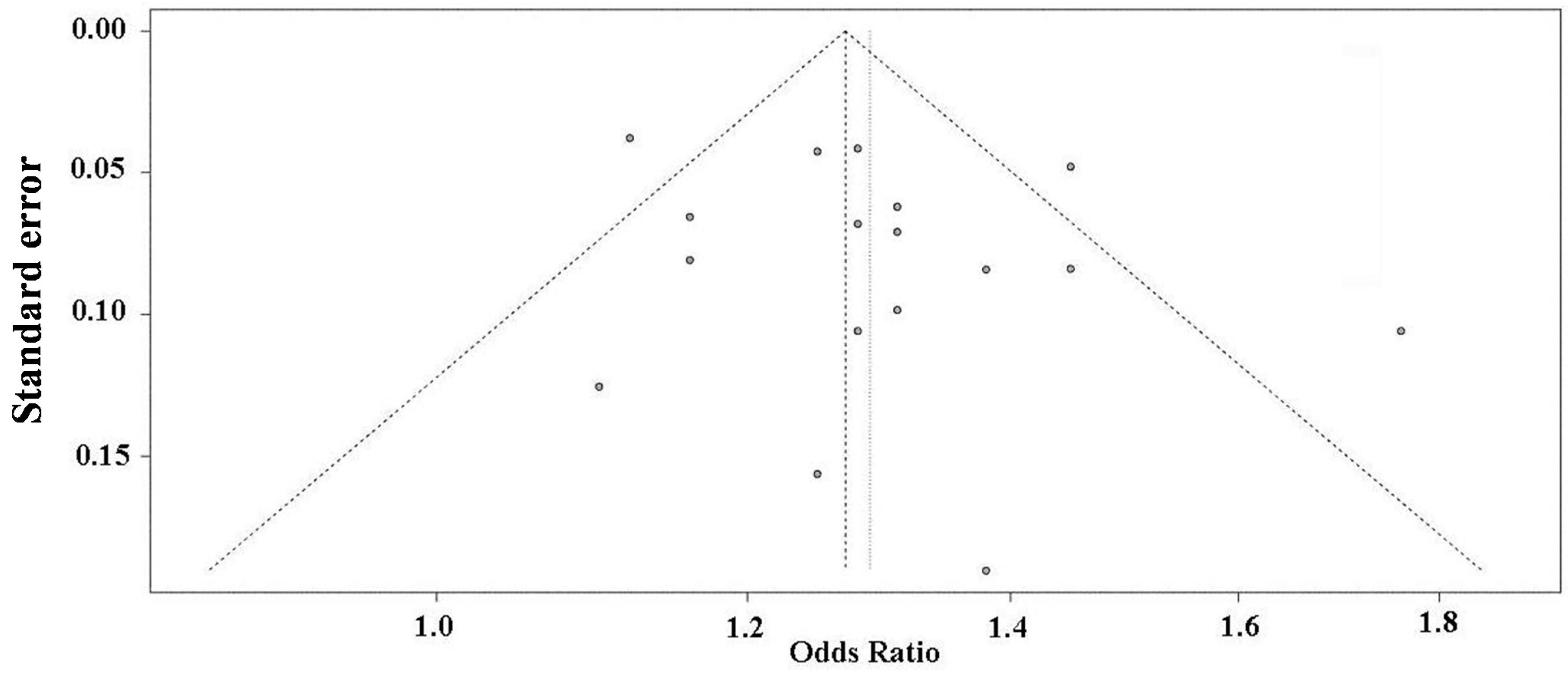
3. Discussion
4. Methods and Materials
4.1. Literature Search
4.2. Inclusion Criteria
4.3. Quality Evaluation
4.4. Data Extraction
4.5. Genetic Model
| Chromosome | Position (hg19) | Variant | Reference Allele | Altered Allele | AMR Freq. | ASN Freq. | EUR Freq. |
|---|---|---|---|---|---|---|---|
| 9 | 22124477 | rs10757278 | A | G | 0.5 | 0.51 | 0.48 |
| 9 | 22125503 | rs1333049 | G | C | 0.5 | 0.5 | 0.47 |
4.6. Heterogeneity Test
4.7. Meta-Analysis
4.8. Sensitivity Analysis
4.9. Publication Bias Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Shiffman, D.; Ellis, S.G.; Rowland, C.M.; Malloy, M.J.; Luke, M.M.; Iakoubova, O.A.; Pullinger, C.R.; Cassano, J.; Aouizerat, B.E.; Fenwick, R.G.; et al. Identification of four gene variants associated with myocardial infarction. Am. J. Hum. Genet. 2005, 77, 596–605. [Google Scholar] [CrossRef]
- Kathiresan, S.; Voight, B.F.; Purcell, S.; Musunuru, K.; Ardissino, D.; Mannucci, P.M.; Anand, S.; Engert, J.C.; Samani, N.J.; Schunkert, H.; et al. Genome-wide association of early-onset myocardial infarction with single nucleotide polymorphisms and copy number variants. Nat. Genet. 2009, 41, 334–341. [Google Scholar] [CrossRef]
- Helgadottir, A.; Thorleifsson, G.; Manolescu, A.; Gretarsdottir, S.; Blondal, T.; Jonasdottir, A.; Sigurdsson, A.; Baker, A.; Palsson, A.; Masson, G.; et al. A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science 2007, 316, 1491–1493. [Google Scholar] [CrossRef]
- Zhang, Q.; Wang, X.F.; Cheng, S.S.; Wan, X.H.; Cao, F.F.; Li, L.; Chen, X.D.; Liu, W.J.; Yang, X.C.; Jin, L. Three SNPs on chromosome 9p21 confer increased risk of myocardial infarction in Chinese subjects. Atherosclerosis 2009, 207, 26–28. [Google Scholar] [CrossRef] [PubMed]
- Hiura, Y.; Fukushima, Y.; Yuno, M.; Sawamura, H.; Kokubo, Y.; Okamura, T.; Tomoike, H.; Goto, Y.; Nonogi, H.; Takahashi, R.; et al. Validation of the association of genetic variants on chromosome 9p21 and 1q41 with myocardial infarction in a Japanese population. Circ. J. 2008, 72, 1213–1217. [Google Scholar] [CrossRef]
- Saleheen, D.; Alexander, M.; Rasheed, A.; Wormser, D.; Soranzo, N.; Hammond, N.; Butterworth, A.; Zaidi, M.; Haycock, P.; Bumpstead, S.; et al. Association of the 9p21.3 locus with risk of first-ever myocardial infarction in Pakistanis: Case-control study in South Asia and updated meta-analysis of Europeans. Arterioscler. Thromb. Vasc. Biol. 2010, 30, 1467–1473. [Google Scholar] [CrossRef]
- Koch, W.; Turk, S.; Erl, A.; Hoppmann, P.; Pfeufer, A.; King, L.; Schomig, A.; Kastrati, A. The chromosome 9p21 region and myocardial infarction in a European population. Atherosclerosis 2011, 217, 220–226. [Google Scholar] [CrossRef] [PubMed]
- Abdullah, K.G.; Li, L.; Shen, G.Q.; Hu, Y.; Yang, Y.; MacKinlay, K.G.; Topol, E.J.; Wang, Q.K. Four SNPS on chromosome 9p21 confer risk to premature, familial CAD and MI in an American Caucasian population (GeneQuest). Ann. Hum. Genet. 2008, 72, 654–657. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.F.; Tsai, P.C.; Liao, Y.C.; Lin, T.H.; Tai, C.T.; Juo, S.H.; Lin, R.T. Chromosome 9p21 genetic variants are associated with myocardial infarction but not with ischemic stroke in a Taiwanese population. J. Investig. Med. 2011, 59, 926–930. [Google Scholar] [PubMed]
- Bhanushali, A.A.; Parmar, N.; Contractor, A.; Shah, V.T.; Das, B.R. Variant on 9p21 is strongly associated with coronary artery disease but lacks association with myocardial infarction and disease severity in a population in Western India. Arch. Med. Res. 2011, 42, 469–474. [Google Scholar] [CrossRef] [PubMed]
- Maksimov, V.N.; Kulikov, I.V.; Orlov, P.S.; Gafarov, V.V.; Maliutina, S.K.; Romashchenko, A.G.; Voevoda, M.I. Evaluation of association between 9 genetic polymorphism and myocardial infarction in the Siberian population. Vestn. Ross. Akad. Med. Nauk 2012, 5, 24–29. [Google Scholar] [PubMed]
- Shen, G.Q.; Rao, S.; Martinelli, N.; Li, L.; Olivieri, O.; Corrocher, R.; Abdullah, K.G.; Hazen, S.L.; Smith, J.; Barnard, J.; et al. Association between four SNPs on chromosome 9p21 and myocardial infarction is replicated in an Italian population. J. Hum. Genet. 2008, 53, 144–150. [Google Scholar] [CrossRef]
- Dehghan, A.; van Hoek, M.; Sijbrands, E.J.; Oostra, B.A.; Hofman, A.; van Duijn, C.M.; Witteman, J.C. Lack of association of two common polymorphisms on 9p21 with risk of coronary heart disease and myocardial infarction; Results from a prospective cohort study. BMC Med. 2008, 6, 30. [Google Scholar] [CrossRef] [PubMed]
- Virani, S.S.; Brautbar, A.; Lee, V.V.; MacArthur, E.; Morrison, A.C.; Grove, M.L.; Nambi, V.; Frazier, L.; Wilson, J.M.; Willerson, J.T.; et al. Chromosome 9p21 single nucleotide polymorphisms are not associated with recurrent myocardial infarction in patients with established coronary artery disease. Circ. J. 2012, 76, 950–956. [Google Scholar] [CrossRef]
- Riley, R.D.; Lambert, P.C.; Abo-Zaid, G. Meta-analysis of individual participant data: Rationale, conduct, and reporting. BMJ 2010, 340, c221. [Google Scholar] [CrossRef] [PubMed]
- Qi, L.; Li, J.M.; Sun, H.; Huang, X.Q.; Lin, K.Q.; Chu, J.Y.; Yang, Z.Q. Association between gene polymorphisms and myocardial infarction in Han Chinese of Yunnan province. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 2012, 29, 413–419. [Google Scholar] [PubMed]
- Peng, W.H.; Lu, L.; Zhang, Q.; Zhang, R.Y.; Wang, L.J.; Yan, X.X.; Chen, Q.J.; Shen, W.F. Chromosome 9p21 polymorphism is associated with myocardial infarction but not with clinical outcome in Han Chinese. Clin. Chem. Lab. Med. 2009, 47, 917–922. [Google Scholar] [PubMed]
- Lu, X.; Wang, L.; Chen, S.; He, L.; Yang, X.; Shi, Y.; Cheng, J.; Zhang, L.; Gu, C.C.; Huang, J.; et al. Genome-wide association study in Han Chinese identifies four new susceptibility loci for coronary artery disease. Nat. Genet. 2012, 44, 890–894. [Google Scholar] [CrossRef]
- Clark, M.F.; Baudouin, S.V. A systematic review of the quality of genetic association studies in human sepsis. Intensive Care Med. 2006, 32, 1706–1712. [Google Scholar] [CrossRef] [PubMed]
- Szpakowicz, A.; Kiliszek, M.; Pepinski, W.; Waszkiewicz, E.; Franaszczyk, M.; Skawronska, M.; Ploski, R.; Niemcunowicz-Janica, A.; Dobrzycki, S.; Opolski, G.; et al. Polymorphism of 9p21.3 locus is associated with 5-year survival in high-risk patients with myocardial infarction. PLoS ONE 2014, 9, e104635. [Google Scholar] [CrossRef]
- Zeng, Q.; Yuan, Y.; Wang, S.; Sun, J.; Zhang, T.; Qi, M. Polymorphisms on chromosome 9p21 confer a risk for acute coronary syndrome in a Chinese Han population. Can. J. Cardiol. 2013, 29, 940–944. [Google Scholar] [CrossRef] [PubMed]
- Patel, R.S.; Asselbergs, F.W.; Quyyumi, A.A.; Palmer, T.M.; Finan, C.I.; Tragante, V.; Deanfield, J.; Hemingway, H.; Hingorani, A.D.; Holmes, M.V. Genetic variants at chromosome 9p21 and risk of first versus subsequent coronary heart disease events: A systematic review and meta-analysis. J. Am. Coll. Cardiol. 2014, 63, 2234–2245. [Google Scholar] [CrossRef]
- Gogele, M.; Minelli, C.; Thakkinstian, A.; Yurkiewich, A.; Pattaro, C.; Pramstaller, P.P.; Little, J.; Attia, J.; Thompson, J.R. Methods for meta-analyses of genome-wide association studies: Critical assessment of empirical evidence. Am. J. Epidemiol. 2012, 175, 739–749. [Google Scholar] [CrossRef] [PubMed]
- Lewis, C.M.; Knight, J. Introduction to genetic association studies. Cold Spring Harb. Protoc. 2012, 2012, 297–306. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Li, F.; Zhang, S.; Jiang, Y.; Ma, G.; Shang, H.; Liu, J.; Feng, R.; Zhang, L.; Liao, M.; et al. Analyzing large-scale samples confirms the association between the ABCA7 rs3764650 polymorphism and Alzheimer’s disease susceptibility. Mol. Neurobiol. 2014, 50, 757–764. [Google Scholar] [CrossRef]
- Ward, L.D.; Kellis, M. HaploReg: A resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 2012, 40, D930–D934. [Google Scholar] [CrossRef] [PubMed]
- Higgins, J.P.; Thompson, S.G.; Deeks, J.J.; Altman, D.G. Measuring inconsistency in meta-analyses. BMJ 2003, 327, 557–560. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Zhang, S.; Cai, Z.; Ma, G.; Zhang, L.; Jiang, Y.; Feng, R.; Liao, M.; Chen, Z.; Zhao, B.; et al. PICALM gene rs3851179 polymorphism contributes to Alzheimer’s disease in an Asian population. Neuromol. Med. 2013, 15, 384–388. [Google Scholar] [CrossRef]
- Egger, M.; Davey Smith, G.; Schneider, M.; Minder, C. Bias in meta-analysis detected by a simple, graphical test. BMJ 1997, 315, 629–634. [Google Scholar] [CrossRef] [PubMed]
- Sterne, J.A.; Egger, M. Funnel plots for detecting bias in meta-analysis: Guidelines on choice of axis. J. Clin. Epidemiol. 2001, 54, 1046–1055. [Google Scholar] [CrossRef] [PubMed]
- Song, F.; Khan, K.S.; Dinnes, J.; Sutton, A.J. Asymmetric funnel plots and publication bias in meta-analyses of diagnostic accuracy. Int. J. Epidemiol. 2002, 31, 88–95. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, G.; Fu, X.; Wang, G.; Liu, G.; Bai, X. Genetic Variant rs10757278 on Chromosome 9p21 Contributes to Myocardial Infarction Susceptibility. Int. J. Mol. Sci. 2015, 16, 11678-11688. https://doi.org/10.3390/ijms160511678
Chen G, Fu X, Wang G, Liu G, Bai X. Genetic Variant rs10757278 on Chromosome 9p21 Contributes to Myocardial Infarction Susceptibility. International Journal of Molecular Sciences. 2015; 16(5):11678-11688. https://doi.org/10.3390/ijms160511678
Chicago/Turabian StyleChen, Guangyuan, Xiuhua Fu, Guangyu Wang, Guiyou Liu, and Xiuping Bai. 2015. "Genetic Variant rs10757278 on Chromosome 9p21 Contributes to Myocardial Infarction Susceptibility" International Journal of Molecular Sciences 16, no. 5: 11678-11688. https://doi.org/10.3390/ijms160511678
APA StyleChen, G., Fu, X., Wang, G., Liu, G., & Bai, X. (2015). Genetic Variant rs10757278 on Chromosome 9p21 Contributes to Myocardial Infarction Susceptibility. International Journal of Molecular Sciences, 16(5), 11678-11688. https://doi.org/10.3390/ijms160511678





