Starting to Gel: How Arabidopsis Seed Coat Epidermal Cells Produce Specialized Secondary Cell Walls
Abstract
:1. Introduction
2. The Development of Seed Coat Epidermal Cells
2.1. Specialized Wall Production
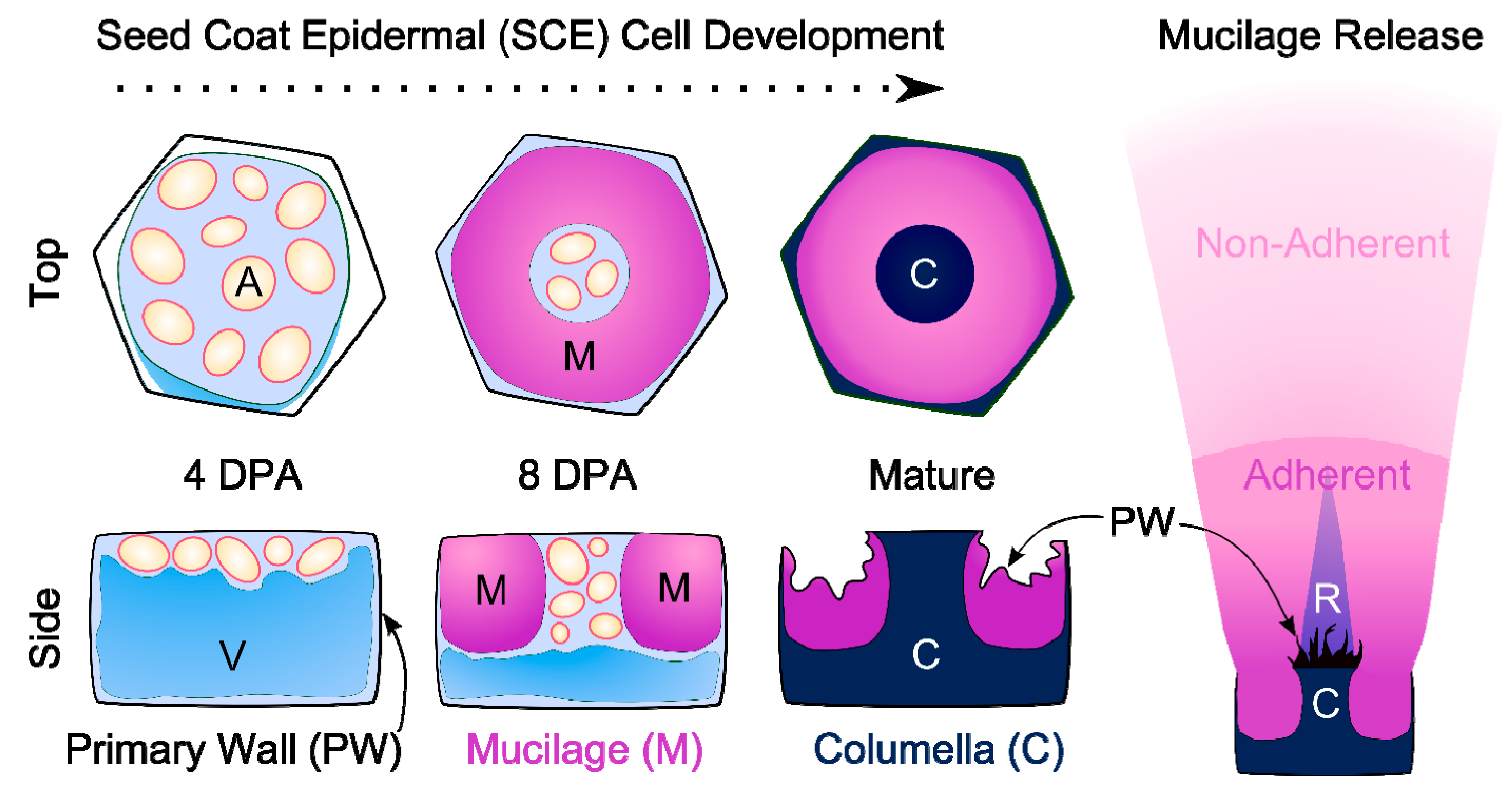
2.2. Covered by a Cuticle?
3. Mucilage Composition: More than Just Pectin
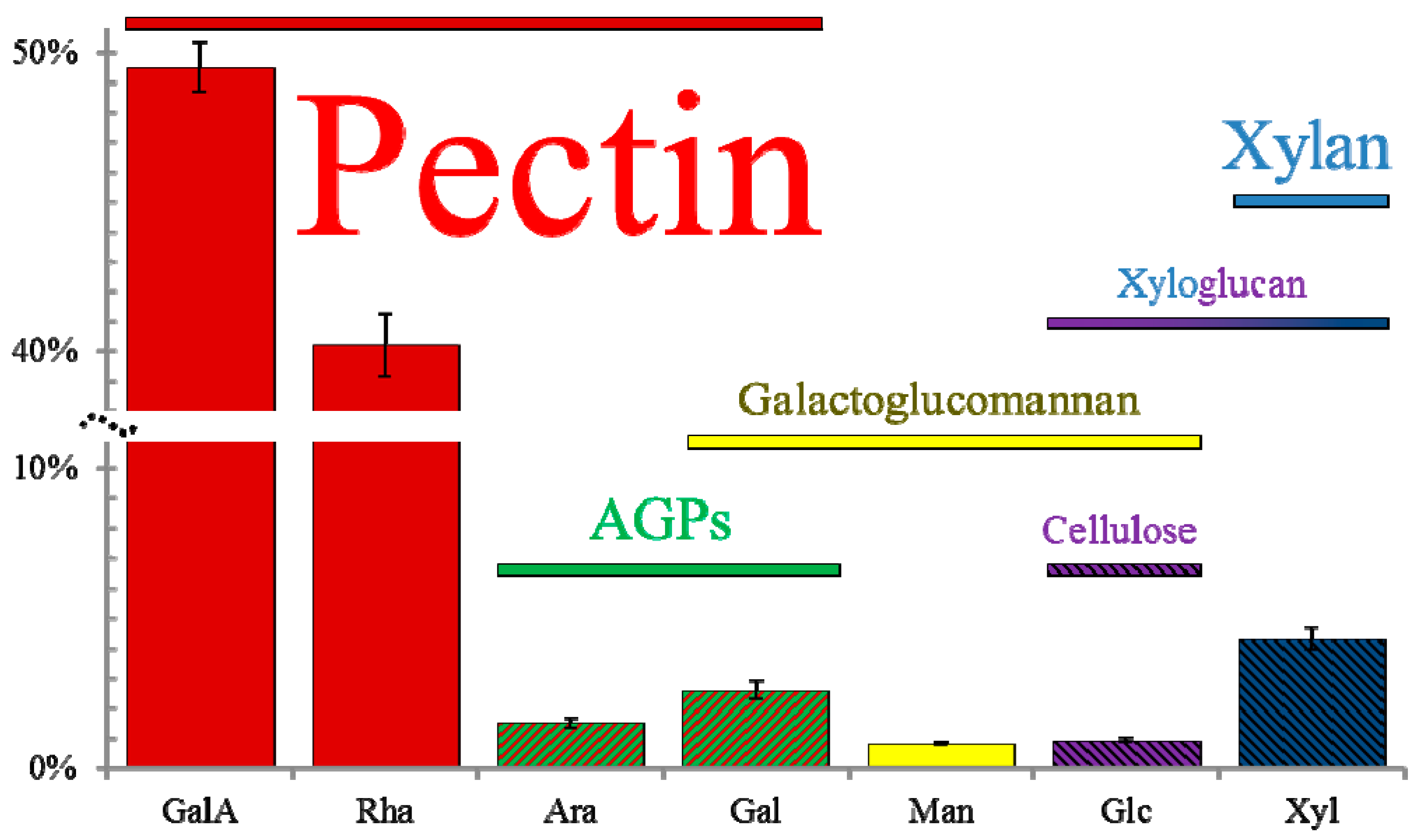
3.1. Pectins
3.1.1. Rhamnogalacturonan I
3.1.2. Homogalacturonan
3.2. Hemicelluloses
3.2.1. Galactoglucomannan
3.2.2. Xylans
3.2.3. Xyloglucan
3.3. Cellulose
3.4. Proteins
4. Regulation of SCE Development
4.1. Outer Integument Establishment
4.2. Control of SCE Differentiation
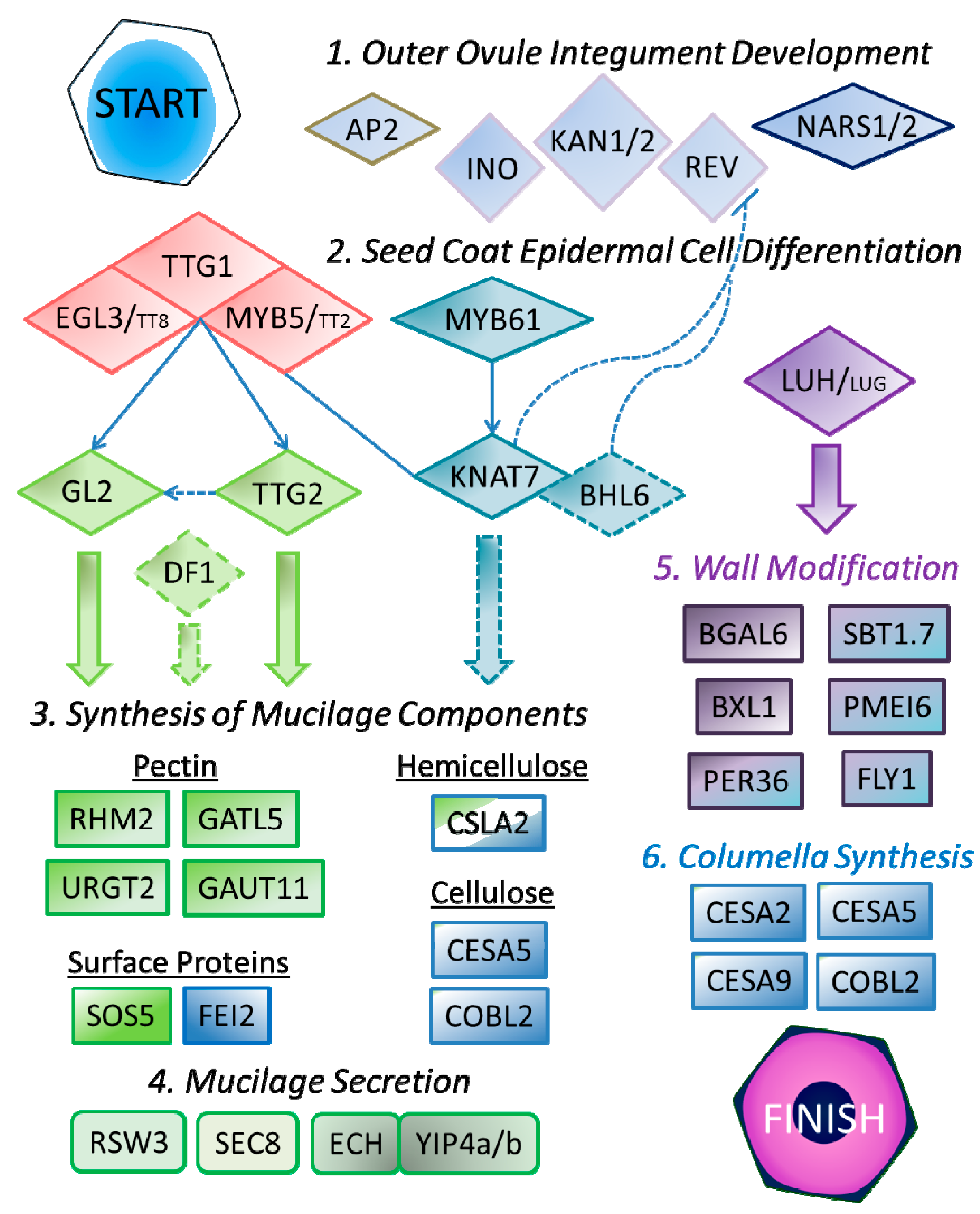
5. Genes Involved in Mucilage Production
5.1. Mucilage Synthesis
5.2. Polar Secretion of Mucilage
6. Genes Involved in Mucilage Modification
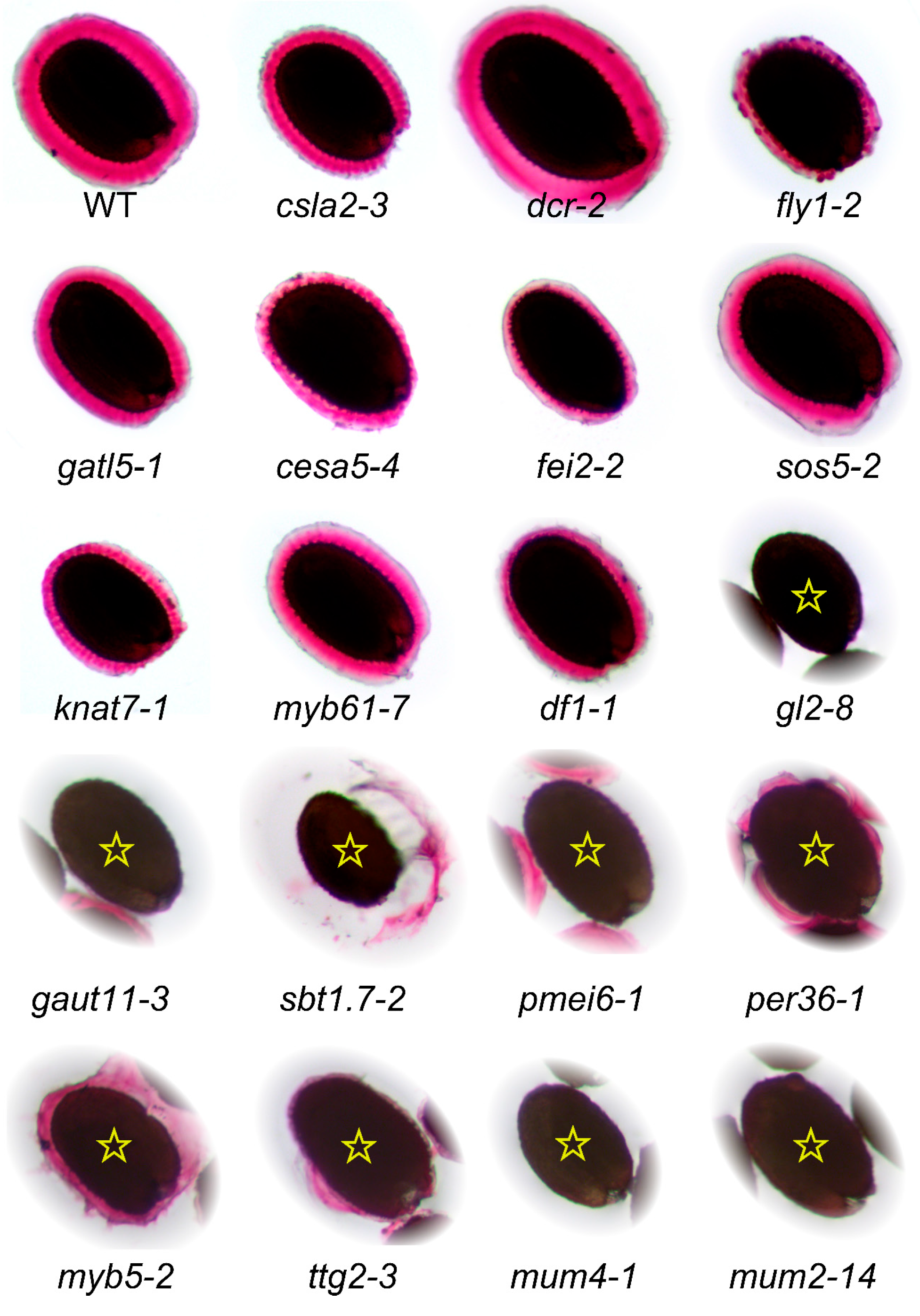
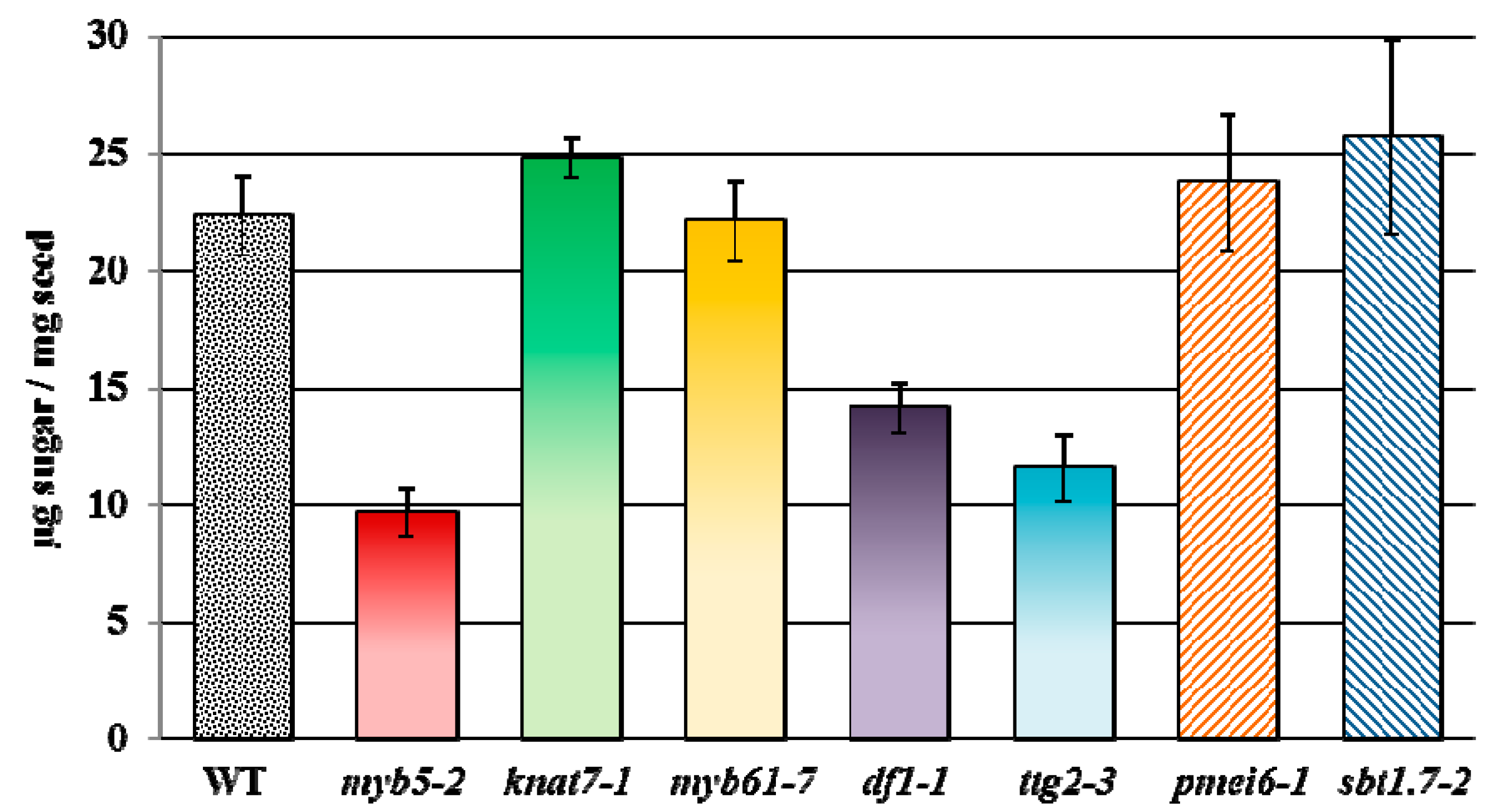
7. Re-Assessment of Mucilage Defects
| Mutant | Locus | Polymorphism | References | Mucilage RR Staining |
|---|---|---|---|---|
| gatl5-1 | At1g02720 | SALK_106615C | [35] | As previously described |
| myb61-7 | At1g09540 | SALK_106556C | [97] | Similar to [78], unlike [28] |
| gaut11-3 | At1g18580 | SAIL_567_H05 | New allele | Similar to gaut11-2 [86] |
| mum4-1 | At1g53500 | EMS mutation | [40] | As previously described |
| knat7-1 | At1g62990 | SALK_002098C | [80] | As previously described |
| df1-1 | At1g76880 | SALK_106258C | [34] | Very small capsule |
| gl2-8 | At1g79840 | SALK_130213C | [98] | As previously described |
| fei2-2 | At2g35620 | SALK_044226 | [24,99] | As previously described |
| ttg2-3 | At2g37260 | SALK_148838 | [77] | As previously described |
| pmei6-1 | At2g47670 | SM_3.19557 | [36] | As previously described |
| myb5-2 | At3g13540 | SALK_105723C | [100] | Similar to myb5-1 mucilage |
| sos5-2 | At3g46550 | SALK_125874 | [24,55] | As previously described |
| per36-1 | At3g50990 | SAIL_194_G03 | [37] | As previously described |
| fly1-2 | At4g28370 | SALK_067290 | [20] | As previously described |
| cesa5-4 | At5g09870 | SALK_207154C | New allele | Similar defects to [22,23,24] |
| csla2-3 | At5g22740 | SALK_149092C | [26] | As previously described |
| dcr-2 | At5g23940 | SALK_128228C | [19] | Similar to wild-type |
| mum2-14 | At5g63800 | SALK_060221C | New allele | Similar defects to [30,31,97] |
| sbt1.7-2 | At5g67360 | GK-544E06.01 | [43] | As previously described |
8. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Somerville, C.; Bauer, S.; Brininstool, G.; Facette, M.; Hamann, T.; Milne, J.; Osborne, E.; Paredez, A.; Persson, S.; Raab, T.; et al. Toward a systems approach to understanding plant cell walls. Science 2004, 306, 2206–2211. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Growth of the plant cell wall. Nat. Rev. Mol. Cell Biol. 2005, 6, 850–861. [Google Scholar] [CrossRef] [PubMed]
- Pauly, M.; Keegstra, K. Plant cell wall polymers as precursors for biofuels. Curr. Opin. Plant Biol. 2010, 13, 305–312. [Google Scholar] [CrossRef] [PubMed]
- Liepman, A.H.; Wightman, R.; Geshi, N.; Turner, S.R.; Scheller, H.V. Arabidopsis—A powerful model system for plant cell wall research. Plant J. 2010, 61, 1107–1121. [Google Scholar] [CrossRef] [PubMed]
- Turner, S.R.; Somerville, C.R. Collapsed xylem phenotype of Arabidopsis identifies mutants deficient in cellulose deposition in the secondary cell wall. Plant Cell 1997, 9, 689–701. [Google Scholar] [CrossRef] [PubMed]
- Brown, D.M.; Zeef, L.A.H.; Ellis, J.; Goodacre, R.; Turner, S.R. Identification of novel genes in Arabidopsis involved in secondary cell wall formation using expression profiling and reverse genetics. Plant Cell 2005, 17, 2281–2295. [Google Scholar] [CrossRef] [PubMed]
- Arsovski, A.A.; Haughn, G.W.; Western, T.L. Seed coat mucilage cells of Arabidopsis thaliana as a model for plant cell wall research. Plant Signal. Behav. 2010, 5, 796–801. [Google Scholar] [CrossRef] [PubMed]
- Haughn, G.W.; Western, T.L. Arabidopsis seed coat mucilage is a specialized cell wall that can be used as a model for genetic analysis of plant cell wall structure and function. Front. Plant Sci. 2012, 3, 64. [Google Scholar] [CrossRef] [PubMed]
- North, H.M.; Berger, A.; Saez-Aguayo, S.; Ralet, M.-C. Understanding polysaccharide production and properties using seed coat mutants: Future perspectives for the exploitation of natural variants. Ann. Bot. 2014, 114, 1251–1263. [Google Scholar] [CrossRef] [PubMed]
- Western, T.L. Changing spaces: The Arabidopsis mucilage secretory cells as a novel system to dissect cell wall production in differentiating cells. Can. J. Bot. 2006, 84, 622–630. [Google Scholar] [CrossRef]
- Western, T.L. The sticky tale of seed coat mucilages: Production, genetics, and role in seed germination and dispersal. Seed Sci. Res. 2011, 22, 1–25. [Google Scholar] [CrossRef]
- Beeckman, T.; de Rycke, R.; Viane, R.; Inzé, D. Histological study of seed coat development in Arabidopsis thaliana. J. Plant Res. 2000, 113, 139–148. [Google Scholar] [CrossRef]
- Western, T.L.; Skinner, D.J.; Haughn, G.W. Differentiation of mucilage secretory cells of the Arabidopsis seed coat. Plant Physiol. 2000, 122, 345–356. [Google Scholar] [CrossRef] [PubMed]
- Windsor, J.B.; Symonds, V.V.; Mendenhall, J.; Lloyd, A.M. Arabidopsis seed coat development: Morphological differentiation of the outer integument. Plant J. 2000, 22, 483–493. [Google Scholar] [CrossRef] [PubMed]
- Young, R.E.; McFarlane, H.E.; Hahn, M.G.; Western, T.L.; Haughn, G.W.; Samuels, L. Analysis of the Golgi apparatus in Arabidopsis seed coat cells during polarized secretion of pectin-rich mucilage. Plant Cell 2008, 20, 1623–1638. [Google Scholar] [CrossRef] [PubMed]
- Western, T.; Burn, J.; Tan, W.; Skinner, D.J.; Martin-McCaffrey, L.; Moffatt, B.A.; Haughn, G.W. Isolation and characterization of mutants defective in seed coat mucilage secretory cell development in Arabidopsis. Plant Physiol. 2001, 127, 998–1011. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Baskin, J.M.; Baskin, C.C.; Huang, Z. More than just a coating: Ecological importance, taxonomic occurrence and phylogenetic relationships of seed coat mucilage. Perspect. Plant Ecol. Evol. Syst. 2012, 14, 434–442. [Google Scholar] [CrossRef]
- Watanabe, M.; Tanaka, H.; Watanabe, D.; Machida, C.; Machida, Y. The ACR4 receptor-like kinase is required for surface formation of epidermis-related tissues in Arabidopsis thaliana. Plant J. 2004, 39, 298–308. [Google Scholar] [CrossRef] [PubMed]
- Panikashvili, D.; Shi, J.; Schreiber, L.; Aharoni, A. The Arabidopsis DCR encoding a soluble BAHD acyltransferase is required for cutin polyester formation and seed hydration properties. Plant Physiol. 2009, 151, 1773–1789. [Google Scholar] [CrossRef] [PubMed]
- Voiniciuc, C.; Dean, G.H.; Griffiths, J.S.; Kirchsteiger, K.; Hwang, Y.T.; Gillett, A.; Dow, G.; Western, T.L.; Estelle, M.; Haughn, G.W.; et al. Flying saucer1 is a transmembrane RING E3 ubiquitin ligase that regulates the degree of pectin methylesterification in Arabidopsis seed mucilage. Plant Cell 2013, 25, 944–959. [Google Scholar] [CrossRef] [PubMed]
- Molina, I.; Ohlrogge, J.B.; Pollard, M. Deposition and localization of lipid polyester in developing seeds of Brassica napus and Arabidopsis thaliana. Plant J. 2008, 53, 437–449. [Google Scholar] [CrossRef] [PubMed]
- Mendu, V.; Griffiths, J.S.; Persson, S.; Stork, J.; Downie, B.; Voiniciuc, C.; Haughn, G.W.; DeBolt, S. Subfunctionalization of cellulose synthases in seed coat epidermal cells mediates secondary radial wall synthesis and mucilage attachment. Plant Physiol. 2011, 157, 441–453. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, S.; Ralet, M.-C.; Berger, A.; Diatloff, E.; Bischoff, V.; Gonneau, M.; Marion-Poll, A.; North, H.M. CESA5 is required for the synthesis of cellulose with a role in structuring the adherent mucilage of Arabidopsis seeds. Plant Physiol. 2011, 156, 1725–1739. [Google Scholar] [CrossRef] [PubMed]
- Harpaz-Saad, S.; McFarlane, H.E.; Xu, S.; Divi, U.K.; Forward, B.; Western, T.L.; Kieber, J.J. Cellulose synthesis via the FEI2 RLK/SOS5 pathway and cellulose synthase 5 is required for the structure of seed coat mucilage in Arabidopsis. Plant J. 2011, 68, 941–953. [Google Scholar] [CrossRef] [PubMed]
- Macquet, A.; Ralet, M.-C.; Kronenberger, J.; Marion-Poll, A.; North, H.M. In situ, chemical and macromolecular study of the composition of Arabidopsis thaliana seed coat mucilage. Plant Cell Physiol. 2007, 48, 984–999. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Shi, D.; Li, J.; Kong, Y.; Yu, Y.; Chai, G.; Hu, R.; Wang, J.; Hahn, M.G.; Zhou, G.; et al. CSLA2, a glucomannan synthase, is involved in maintaining adherent mucilage structure in Arabidopsis seed. Plant Physiol. 2014, 164, 1842–1856. [Google Scholar] [CrossRef] [PubMed]
- Kulich, I.; Cole, R.; Drdová, E.; Cvrcková, F.; Soukup, A.; Fowler, J.; Zárský, V. Arabidopsis exocyst subunits SEC8 and EXO70A1 and exocyst interactor ROH1 are involved in the localized deposition of seed coat pectin. New Phytol. 2010, 188, 615–625. [Google Scholar] [CrossRef] [PubMed]
- Penfield, S.; Meissner, R.C.; Shoue, D.A.; Carpita, N.C.; Bevan, M.W. MYB61 is required for mucilage deposition and extrusion in the Arabidopsis seed coat. Plant Cell 2001, 13, 2777–2791. [Google Scholar] [CrossRef] [PubMed]
- Usadel, B.; Kuschinsky, A.; Rosso, M.G.; Eckermann, N.; Pauly, M. RHM2 is involved in mucilage pectin synthesis and is required for the development of the seed coat in Arabidopsis. Plant Physiol. 2004, 134, 286–295. [Google Scholar] [CrossRef] [PubMed]
- Dean, G.H.; Zheng, H.; Tewari, J.; Huang, J.; Young, D.S.; Hwang, Y.T.; Western, T.L.; Carpita, N.C.; McCann, M.C.; Mansfield, S.D.; et al. The Arabidopsis MUM2 gene encodes a β-galactosidase required for the production of seed coat mucilage with correct hydration properties. Plant Cell 2007, 19, 4007–4021. [Google Scholar] [CrossRef] [PubMed]
- Macquet, A.; Ralet, M.-C.; Loudet, O.; Kronenberger, J.; Mouille, G.; Marion-Poll, A.; North, H.M. A naturally occurring mutation in an Arabidopsis accession affects a β-d-galactosidase that increases the hydrophilic potential of rhamnogalacturonan I in seed mucilage. Plant Cell 2007, 19, 3990–4006. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; DeBowles, D.; Esfandiari, E.; Dean, G.; Carpita, N.C.; Haughn, G.W. The Arabidopsis transcription factor LUH/MUM1 is required for extrusion of seed coat mucilage. Plant Physiol. 2011, 156, 491–502. [Google Scholar] [CrossRef] [PubMed]
- Walker, M.; Tehseen, M.; Doblin, M.S.; Pettolino, F.A; Wilson, S.M.; Bacic, A.; Golz, J.F. The transcriptional regulator LEUNIG_HOMOLOG regulates mucilage release from the Arabidopsis testa. Plant Physiol. 2011, 156, 46–60. [Google Scholar] [CrossRef]
- Vasilevski, A.; Giorgi, F.M.; Bertinetti, L.; Usadel, B. LASSO modeling of the Arabidopsis thaliana seed/seedling transcriptome: A model case for detection of novel mucilage and pectin metabolism genes. Mol. Biosyst. 2012, 8, 2566–2574. [Google Scholar] [CrossRef] [PubMed]
- Kong, Y.; Zhou, G.; Abdeen, A.; Schafhauser, J.; Richardson, B.; Atmodjo, M.; Jung, J.; Wicker, L.; Mohnen, D.; Western, T.; et al. GALACTURONOSYLTRANSFERASE-LIKE5 is involved in the production of Arabidopsis seed coat mucilage. Plant Physiol. 2013, 163, 1203–1217. [Google Scholar] [CrossRef] [PubMed]
- Saez-Aguayo, S.; Ralet, M.-C.; Berger, A.; Botran, L.; Ropartz, D.; Marion-Poll, A.; North, H.M. PECTIN METHYLESTERASE INHIBITOR6 promotes Arabidopsis mucilage release by limiting methylesterification of homogalacturonan in seed coat epidermal cells. Plant Cell 2013, 25, 308–323. [Google Scholar] [CrossRef] [PubMed]
- Kunieda, T.; Shimada, T.; Kondo, M.; Nishimura, M.; Nishitani, K.; Hara-Nishimura, I. Spatiotemporal secretion of PEROXIDASE36 is required for seed coat mucilage extrusion in Arabidopsis. Plant Cell 2013, 25, 1355–1367. [Google Scholar] [CrossRef] [PubMed]
- Arsovski, A.; Popma, T.M.; Haughn, G.W.; Carpita, N.C.; Mccann, M.C.; Western, T.L. AtBXL1 encodes a bifunctional β-d-xylosidase/alpha-l-arabinofuranosidase required for pectic arabinan modification in Arabidopsis mucilage secretory cells. Plant Physiol. 2009, 150, 1219–1234. [Google Scholar] [CrossRef] [PubMed]
- Hanke, D.E.; Northcote, D.H. Molecular visualization of pectin and DNA by ruthenium red. Biopolymers 1975, 14, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Western, T.L.; Young, D.S.; Dean, G.H.; Tan, W.L.; Samuels, A.L.; Haughn, G.W. MUCILAGE-MODIFIED4 encodes a putative pectin biosynthetic enzyme developmentally regulated by APETALA2, TRANSPARENT TESTA GLABRA1, and GLABRA2 in the arabidopsis seed coat. Plant Physiol. 2004, 134, 296–306. [Google Scholar] [CrossRef] [PubMed]
- Ralet, M.-C.; Tranquet, O.; Poulain, D.; Moïse, A.; Guillon, F. Monoclonal antibodies to rhamnogalacturonan I backbone. Planta 2010, 231, 1373–1383. [Google Scholar] [CrossRef] [PubMed]
- Willats, W.G.T.; McCartney, L.; Knox, J.P. In-situ analysis of pectic polysaccharides in seed mucilage and at the root surface of Arabidopsis thaliana. Planta 2001, 213, 37–44. [Google Scholar] [CrossRef] [PubMed]
- Rautengarten, C.; Usadel, B.; Neumetzler, L.; Hartmann, J.; Büssis, D.; Altmann, T. A subtilisin-like serine protease essential for mucilage release from Arabidopsis seed coats. Plant J. 2008, 54, 466–480. [Google Scholar] [CrossRef] [PubMed]
- Naran, R.; Chen, G.; Carpita, N.C. Novel rhamnogalacturonan I and arabinoxylan polysaccharides of flax seed mucilage. Plant Physiol. 2008, 148, 132–141. [Google Scholar] [CrossRef] [PubMed]
- Ralet, M.-C.; Crépeau, M.-J.; Buchholt, H.-C.; Thibault, J.-F. Polyelectrolyte behaviour and calcium binding properties of sugar beet pectins differing in their degrees of methylation and acetylation. Biochem. Eng. J. 2003, 16, 191–201. [Google Scholar] [CrossRef]
- Harholt, J.; Suttangkakul, A.; Vibe Scheller, H. Biosynthesis of pectin. Plant Physiol. 2010, 153, 384–395. [Google Scholar] [CrossRef] [PubMed]
- Gille, S.; Pauly, M. O-acetylation of plant cell wall polysaccharides. Front. Plant Sci. 2012, 3, 12. [Google Scholar] [CrossRef] [PubMed]
- Levesque-Tremblay, G.; Müller, K.; Mansfield, S.D.; Haughn, G.W. HIGHLY METHYL ESTERIFIED SEEDS is a pectin methyl esterase involved in embryo development. Plant Physiol. 2015. [Google Scholar] [CrossRef]
- Liners, F.; van Cutsem, P. Distribution of pectic polysaccharides throughout walls of suspension-cultured carrot cells: An immunocytochemical study. Protoplasma 1992, 170, 10–21. [Google Scholar] [CrossRef]
- Marcus, S.E.; Blake, A.W.; Benians, T.A.S.; Lee, K.J.D.; Poyser, C.; Donaldson, L.; Leroux, O.; Rogowski, A.; Petersen, H.L.; Boraston, A.; et al. Restricted access of proteins to mannan polysaccharides in intact plant cell walls. Plant J. 2010, 64, 191–203. [Google Scholar] [CrossRef] [PubMed]
- Liepman, A.H.; Wilkerson, C.G.; Keegstra, K. Expression of cellulose synthase-like (Csl) genes in insect cells reveals that CslA family members encode mannan synthases. Proc. Natl. Acad. Sci. USA 2005, 102, 2221–2226. [Google Scholar] [CrossRef] [PubMed]
- Pauly, M.; Gille, S.; Liu, L.; Mansoori, N.; de Souza, A.; Schultink, A.; Xiong, G. Hemicellulose biosynthesis. Planta 2013, 238, 627–642. [Google Scholar] [CrossRef] [PubMed]
- McCartney, L.; Marcus, S.E.; Knox, J.P. Monoclonal antibodies to plant cell wall xylans and arabinoxylans. J. Histochem. Cytochem. 2005, 53, 543–546. [Google Scholar] [CrossRef] [PubMed]
- Rennie, E.A.; Scheller, H.V. Xylan biosynthesis. Curr. Opin. Biotechnol. 2014, 26, 100–107. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, J.S.; Tsai, A.Y.-L.; Xue, H.; Voiniciuc, C.; Sola, K.; Seifert, G.J.; Mansfield, S.D.; Haughn, G.W. SALT-OVERLY SENSITIVE5 mediates arabidopsis seed coat mucilage adherence and organization through pectins. Plant Physiol. 2014, 165, 991–1004. [Google Scholar] [CrossRef] [PubMed]
- Zabotina, O. Xyloglucan and its biosynthesis. Front. Plant Sci. 2012, 3, 134. [Google Scholar] [CrossRef] [PubMed]
- Moore, P.J.; Darvill, A.G.; Albersheim, P.; Andrew, L.; Al, M.E.T. Immunogold localization of xyloglucan and rhamnogalacturonan I in the cell walls of suspension-cultured sycamore cells. Plant Physiol. 1986, 82, 787–794. [Google Scholar] [CrossRef] [PubMed]
- Updegraff, D.M. Semimicro determination of cellulose in biological materials. Anal. Biochem. 1969, 32, 420–424. [Google Scholar] [CrossRef] [PubMed]
- Ben-Tov, D.; Abraham, Y.; Stav, S.; Thompson, K.; Loraine, A.; Elbaum, R.; de Souza, A.; Pauly, M.; Kieber, J.J.; Harpaz-Saad, S.; et al. COBRA-LIKE 2, a member of the GPI-anchored COBRA-LIKE family, plays a role in cellulose deposition in Arabidopsis seed coat mucilage secretory cells. Plant Physiol. 2015. [Google Scholar] [CrossRef]
- Blake, A.W.; McCartney, L.; Flint, J.E.; Bolam, D.N.; Boraston, A.B.; Gilbert, H.J.; Knox, J.P. Understanding the biological rationale for the diversity of cellulose-directed carbohydrate-binding modules in prokaryotic enzymes. J. Biol. Chem. 2006, 281, 29321–29329. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.T.; Carroll, A.; Akhmetova, L.; Somerville, C. Real-time imaging of cellulose reorientation during cell wall expansion in Arabidopsis roots. Plant Physiol. 2010, 152, 787–796. [Google Scholar] [CrossRef] [PubMed]
- Ellis, M.; Egelund, J.; Schultz, C.J.; Bacic, A. Arabinogalactan-proteins: Key regulators at the cell surface? Plant Physiol. 2010, 153, 403–419. [Google Scholar] [CrossRef] [PubMed]
- Dean, G.; Cao, Y.; Xiang, D.; Provart, N.J.; Ramsay, L.; Ahad, A.; White, R.; Selvaraj, G.; Datla, R.; Haughn, G.; et al. Analysis of gene expression patterns during seed coat development in Arabidopsis. Mol. Plant 2011, 4, 1074–1091. [Google Scholar] [CrossRef] [PubMed]
- Winter, D.; Vinegar, B.; Nahal, H.; Ammar, R.; Wilson, G.V.; Provart, N.J. An “electronic fluorescent pictograph” Browser for exploring and analyzing large-scale biological data sets. PLoS One 2007, 2, e718. [Google Scholar] [CrossRef] [PubMed]
- Le, B.H.; Cheng, C.; Bui, A.Q.; Wagmaister, J.A; Henry, K.F.; Pelletier, J.; Kwong, L.; Belmonte, M.; Kirkbride, R.; Horvath, S.; et al. Global analysis of gene activity during Arabidopsis seed development and identification of seed-specific transcription factors. Proc. Natl. Acad. Sci. USA 2010, 107, 8063–8070. [Google Scholar] [CrossRef] [PubMed]
- Jofuku, K.; Den Boer, B.G. Control of Arabidopsis flower and seed development by the homeotic gene APETALA2. Plant Cell 1994, 6, 1211–1225. [Google Scholar] [CrossRef] [PubMed]
- Kunieda, T.; Mitsuda, N.; Ohme-Takagi, M.; Takeda, S.; Aida, M.; Tasaka, M.; Kondo, M.; Nishimura, M.; Hara-Nishimura, I. NAC family proteins NARS1/NAC2 and NARS2/NAM in the outer integument regulate embryogenesis in Arabidopsis. Plant Cell 2008, 20, 2631–2642. [Google Scholar] [CrossRef] [PubMed]
- Kelley, D.R.; Skinner, D.J.; Gasser, C.S. Roles of polarity determinants in ovule development. Plant J. 2009, 57, 1054–1064. [Google Scholar] [CrossRef] [PubMed]
- McAbee, J.M.; Hill, T.A.; Skinner, D.J.; Izhaki, A.; Hauser, B.A.; Meister, R.J.; Venugopala Reddy, G.; Meyerowitz, E.M.; Bowman, J.L.; Gasser, C.S.; et al. ABERRANT TESTA SHAPE encodes a KANADI family member, linking polarity determination to separation and growth of Arabidopsis ovule integuments. Plant J. 2006, 46, 522–531. [Google Scholar] [CrossRef] [PubMed]
- Leon-Kloosterziel, K.M.; Keijzer, C.J.; Koornneef, M. A seed shape mutant of Arabidopsis that is affected in integument development. Plant Cell 1994, 6, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Gonzalez, A.; Zhao, M.; Payne, C.T.; Lloyd, A. A network of redundant bHLH proteins functions in all TTG1-dependent pathways of Arabidopsis. Development 2003, 130, 4859–4869. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.R.; Davison, P.A.; Bolognesi-Winfield, A.C.; James, C.M.; Srinivasan, N.; Blundell, T.L.; Esch, J.J.; Marks, M.D.; Gray, J.C.; Graya, J.C.; et al. The TRANSPARENT TESTA GLABRA1 locus, which regulates trichome differentiation and anthocyanin biosynthesis in Arabidopsis, encodes a WD40 repeat protein. Plant Cell 1999, 11, 1337–1349. [Google Scholar] [CrossRef] [PubMed]
- Baudry, A.; Heim, M.A; Dubreucq, B.; Caboche, M.; Weisshaar, B.; Lepiniec, L. TT2, TT8, and TTG1 synergistically specify the expression of BANYULS and proanthocyanidin biosynthesis in Arabidopsis thaliana. Plant J. 2004, 39, 366–380. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, A.; Mendenhall, J.; Huo, Y.; Lloyd, A. TTG1 complex MYBs, MYB5 and TT2, control outer seed coat differentiation. Dev. Biol. 2009, 325, 412–421. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, A.; Zhao, M.; Leavitt, J.M.; Lloyd, A.M. Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J. 2008, 53, 814–827. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.S.; Kolevski, B.; Smyth, D.R. TRANSPARENT TESTA GLABRA2, a trichome and seed coat development gene of Arabidopsis, encodes a WRKY transcription factor. Plant Cell 2002, 14, 1359–1375. [Google Scholar] [CrossRef] [PubMed]
- Ishida, T.; Hattori, S.; Sano, R.; Inoue, K.; Shirano, Y.; Hayashi, H.; Shibata, D.; Sato, S.; Kato, T.; Tabata, S.; et al. Arabidopsis TRANSPARENT TESTA GLABRA2 is directly regulated by R2R3 MYB transcription factors and is involved in regulation of GLABRA2 transcription in epidermal differentiation. Plant Cell 2007, 19, 2531–2543. [Google Scholar] [CrossRef] [PubMed]
- Romano, J.M.; Dubos, C.; Prouse, M.B.; Wilkins, O.; Hong, H.; Poole, M.; Kang, K.-Y.; Li, E.; Douglas, C.J.; Western, T.L.; et al. AtMYB61, an R2R3-MYB transcription factor, functions as a pleiotropic regulator via a small gene network. New Phytol. 2012, 195, 774–786. [Google Scholar] [CrossRef] [PubMed]
- Li, E.; Bhargava, A.; Qiang, W.; Friedmann, M.C.; Forneris, N.; Savidge, R.A.; Johnson, L.A.; Mansfield, S.D.; Ellis, B.E.; Douglas, C.J.; et al. The Class II KNOX gene KNAT7 negatively regulates secondary wall formation in Arabidopsis and is functionally conserved in Populus. New Phytol. 2012, 194, 102–115. [Google Scholar] [CrossRef] [PubMed]
- Bhargava, A.; Ahad, A.; Wang, S.; Mansfield, S.D.; Haughn, G.W.; Douglas, C.J.; Ellis, B.E. The interacting MYB75 and KNAT7 transcription factors modulate secondary cell wall deposition both in stems and seed coat in Arabidopsis. Planta 2013, 237, 1199–1211. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; You, S.; Taylor-Teeples, M.; Li, W.L.; Schuetz, M.; Brady, S.M.; Douglas, C.J. BEL1-LIKE HOMEODOMAIN6 and KNOTTED ARABIDOPSIS THALIANA7 Interact and Regulate Secondary Cell Wall Formation via Repression of REVOLUTA. Plant Cell 2014. [Google Scholar] [CrossRef]
- Stahle, M.I.; Kuehlich, J.; Staron, L.; von Arnim, A.G.; Golz, J.F. YABBYs and the transcriptional corepressors LEUNIG and LEUNIG_HOMOLOG maintain leaf polarity and meristem activity in Arabidopsis. Plant Cell 2009, 21, 3105–3118. [Google Scholar] [CrossRef] [PubMed]
- Bui, M.; Lim, N.; Sijacic, P.; Liu, Z. LEUNIG_HOMOLOG and LEUNIG regulate seed mucilage extrusion in Arabidopsis. J. Integr. Plant Biol. 2011, 53, 399–408. [Google Scholar] [CrossRef] [PubMed]
- Oka, T.; Nemoto, T.; Jigami, Y. Functional analysis of Arabidopsis thaliana RHM2/MUM4, a multidomain protein involved in UDP-d-glucose to UDP-l-rhamnose conversion. J. Biol. Chem. 2007, 282, 5389–5403. [Google Scholar] [CrossRef] [PubMed]
- Rautengarten, C.; Ebert, B.; Moreno, I.; Temple, H.; Herter, T.; Link, B.; Doñas-Cofré, D.; Moreno, A.; Saéz-Aguayo, S.; Blanco, F.; et al. The Golgi localized bifunctional UDP-rhamnose/UDP-galactose transporter family of Arabidopsis. Proc. Natl. Acad. Sci. USA 2014, 111, 11563–11568. [Google Scholar] [CrossRef] [PubMed]
- Caffall, K.H.; Pattathil, S.; Phillips, S.E.; Hahn, M.G.; Mohnen, D. Arabidopsis thaliana T-DNA mutants implicate GAUT genes in the biosynthesis of pectin and xylan in cell walls and seed testa. Mol. Plant 2009, 2, 1000–1014. [Google Scholar] [CrossRef] [PubMed]
- Stork, J.; Harris, D.; Griffiths, J.; Williams, B.; Beisson, F.; Li-Beisson, Y.; Mendu, V.; Haughn, G.; Debolt, S. CELLULOSE SYNTHASE9 serves a nonredundant role in secondary cell wall synthesis in Arabidopsis epidermal testa cells. Plant Physiol. 2010, 153, 580–589. [Google Scholar] [CrossRef] [PubMed]
- Harpaz-Saad, S.; Western, T.L.; Kieber, J.J. The FEI2-SOS5 pathway and CELLULOSE SYNTHASE 5 are required for cellulose biosynthesis in the Arabidopsis seed coat and affect pectin mucilage structure. Plant Signal. Behav. 2012, 7, 285–288. [Google Scholar] [CrossRef] [PubMed]
- Burn, J.E.; Hurley, U.A.; Birch, R.J.; Arioli, T.; Cork, A.; Williamson, R.E. The cellulose-deficient Arabidopsis mutant rsw3 is defective in a gene encoding a putative glucosidase II, an enzyme processing N-glycans during ER quality control. Plant J. 2002, 32, 949–960. [Google Scholar] [CrossRef] [PubMed]
- Gendre, D.; McFarlane, H.E.; Johnson, E.; Mouille, G.; Sjödin, A.; Oh, J.; Levesque-Tremblay, G.; Watanabe, Y.; Samuels, L.; Bhalerao, R.P.; et al. Trans-Golgi network localized ECHIDNA/Ypt interacting protein complex is required for the secretion of cell wall polysaccharides in Arabidopsis. Plant Cell 2013, 25, 2633–2646. [Google Scholar] [CrossRef] [PubMed]
- McFarlane, H.E.; Watanabe, Y.; Gendre, D.; Carruthers, K.; Levesque-Tremblay, G.; Haughn, G.W.; Bhalerao, R.P.; Samuels, L. Cell wall polysaccharides are mislocalized to the Vacuole in echidna mutants. Plant Cell Physiol. 2013, 54, 1867–1880. [Google Scholar] [CrossRef] [PubMed]
- Wolf, S.; Mouille, G.; Pelloux, J. Homogalacturonan methyl-esterification and plant development. Mol. Plant 2009, 2, 851–860. [Google Scholar] [CrossRef] [PubMed]
- Müller, K.; Levesque-Tremblay, G.; Bartels, S.; Weitbrecht, K.; Wormit, A.; Usadel, B.; Haughn, G.; Kermode, A.R. Demethylesterification of cell wall pectins in Arabidopsis plays a role in seed germination. Plant Physiol. 2013, 161, 305–316. [Google Scholar] [CrossRef] [PubMed]
- Scheler, C.; Weitbrecht, K.; Pearce, S.P.; Hampstead, A.; Büttner-Mainik, A.; Lee, K.; Voegele, A.; Oracz, K.; Dekkers, B.; Wang, X.; et al. Promotion of testa rupture during lepidium sativum germination involves seed compartment-specific expression and activity of pectin methylesterases. Plant Physiol. 2015, 167, 200–215. [Google Scholar] [CrossRef] [PubMed]
- Sénéchal, F.; Graff, L.; Surcouf, O.; Marcelo, P.; Rayon, C.; Bouton, S.; Mareck, A.; Mouille, G.; Stintzi, A.; Höfte, H.; et al. Arabidopsis PECTIN METHYLESTERASE17 is co-expressed with and processed by SBT3.5, a subtilisin-like serine protease. Ann. Bot. 2014, 114, 1161–1175. [Google Scholar] [CrossRef] [PubMed]
- Berendzen, K.; Searle, I.; Ravenscroft, D.; Koncz, C.; Batschauer, A.; Coupland, G.; Somssich, I.E.; Ulker, B. A rapid and versatile combined DNA/RNA extraction protocol and its application to the analysis of a novel DNA marker set polymorphic between Arabidopsis thaliana ecotypes Col-0 and Landsberg erecta. Plant Methods 2005, 1. [Google Scholar] [CrossRef] [PubMed]
- Saez-Aguayo, S.; Rondeau-Mouro, C.; Macquet, A.; Kronholm, I.; Ralet, M.-C.; Berger, A.; Sallé, C.; Poulain, D.; Granier, F.; Botran, L.; et al. Local evolution of seed flotation in Arabidopsis. PLoS Genet. 2014, 10, e1004221. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Katavic, V.; Yu, Y.; Kunst, L.; Haughn, G. Arabidopsis GLABRA2 mutant seeds deficient in mucilage biosynthesis produce more oil. Plant J. 2012, 69, 37–46. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.-L.; Rahman, A.; Baskin, T.I.; Kieber, J.J. Two leucine-rich repeat receptor kinases mediate signaling, linking cell wall biosynthesis and ACC synthase in Arabidopsis. Plant Cell 2008, 20, 3065–3079. [Google Scholar] [CrossRef] [PubMed]
- Li, S.F.; Milliken, O.N.; Pham, H.; Seyit, R.; Napoli, R.; Preston, J.; Koltunow, A.M.; Parish, R.W. The Arabidopsis MYB5 transcription factor regulates mucilage synthesis, seed coat development, and trichome morphogenesis. Plant Cell 2009, 21, 72–89. [Google Scholar] [CrossRef] [PubMed]
- Arsovski, A.A.; Villota, M.M.; Rowland, O.; Subramaniam, R.; Western, T.L. MUM ENHANCERS are important for seed coat mucilage production and mucilage secretory cell differentiation in Arabidopsis thaliana. J. Exp. Bot. 2009, 60, 2601–2612. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Voiniciuc, C.; Yang, B.; Schmidt, M.H.-W.; Günl, M.; Usadel, B. Starting to Gel: How Arabidopsis Seed Coat Epidermal Cells Produce Specialized Secondary Cell Walls. Int. J. Mol. Sci. 2015, 16, 3452-3473. https://doi.org/10.3390/ijms16023452
Voiniciuc C, Yang B, Schmidt MH-W, Günl M, Usadel B. Starting to Gel: How Arabidopsis Seed Coat Epidermal Cells Produce Specialized Secondary Cell Walls. International Journal of Molecular Sciences. 2015; 16(2):3452-3473. https://doi.org/10.3390/ijms16023452
Chicago/Turabian StyleVoiniciuc, Cătălin, Bo Yang, Maximilian Heinrich-Wilhelm Schmidt, Markus Günl, and Björn Usadel. 2015. "Starting to Gel: How Arabidopsis Seed Coat Epidermal Cells Produce Specialized Secondary Cell Walls" International Journal of Molecular Sciences 16, no. 2: 3452-3473. https://doi.org/10.3390/ijms16023452
APA StyleVoiniciuc, C., Yang, B., Schmidt, M. H.-W., Günl, M., & Usadel, B. (2015). Starting to Gel: How Arabidopsis Seed Coat Epidermal Cells Produce Specialized Secondary Cell Walls. International Journal of Molecular Sciences, 16(2), 3452-3473. https://doi.org/10.3390/ijms16023452






