Polymorphisms of Leptin-b Gene Associated with Growth Traits in Orange-Spotted Grouper (Epinephelus coioides)
Abstract
:1. Introduction
2. Results and Discussion
2.1. Single Nucleotide Polymorphisms Identification and Genotyping
2.2. Association Analysis with Growth Traits
| SNP | Position | Mutation Type | Sample Size | Genotype Frequencies (%) | Allele Frequencies (%) | H o | H e | p-Value(χ2, HWE) | PIC a | |||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| c.14G>A | Exon1 | Missense | 200 | GG | GA | AA | G | A | 0.525 | 0.497 | 0.731 | 0.374 |
| Locus14 | R: CGG→Q: CAG | - | 27.5 | 52.5 | 20.0 | 53.75 | 46.25 | - | - | - | - | |
| c.93A>G | Exon1 | Synonymous | 200 | AA | AG | GG | A | G | 0.430 | 0.428 | 0.997 | 0.336 |
| Locus93 | T: ACA→ACG | - | 47.5 | 43.0 | 9.5 | 69.00 | 31.00 | - | - | - | - | |
| c.149G>A | Exon1 | Missense | 200 | GG | GA | AA | G | A | 0.080 | 0.077 | 0.841 | 0.074 |
| Locus149 | R: CGG→Q: CAG | - | 92.0 | 8.0 | 0.0 | 96.00 | 4.00 | - | - | - | - | |
| c.181A>G | Intron1 | Untranslated | 200 | AA | GA | GG | A | G | 0.060 | 0.068 | 0.287 | 0.065 |
| Locus181 | - | - | 93.5 | 6.0 | 0.5 | 96.50 | 3.50 | - | - | - | - | |
| c.193G>A | Intron1 | Untranslated | 200 | GG | GA | AA | G | A | 0.400 | 0.416 | 0.863 | 0.329 |
| Locus193 | - | - | 50.5 | 40.0 | 9.5 | 70.50 | 29.50 | - | - | - | - | |
| c.360C>T | Exon2 | Synonymous | 200 | CC | CT | TT | C | T | 0.495 | 0.467 | 0.707 | 0.358 |
| Locus360 | L: CTC→CTT | - | 38.0 | 49.5 | 12.5 | 62.75 | 37.25 | - | - | - | - | |
| SNP | Genotypes | N | BWT (g) | BWH (cm) | OL (cm) | BL (cm) | TW (cm) | HL (cm) | CPW (cm) | CPL (cm) | SL (cm) | ED (cm) | ID (cm) | K (%) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| c.14G>A | GG | 55 | 87.89 ± 37.17 | 4.66 ± 0.72 | 17.78 ± 2.65 | 14.58 ± 2.22 | 2.72 ± 0.49 | 5.80 ± 0.88 | 1.58 ± 0.23 | 2.42 ± 0.49 | 1.16 ± 0.22 | 0.94 ± 0.08 | 0.92 ± 0.16 | 2.70 ± 0.33 | ||||||||||
| GA | 105 | 92.86 ± 45.81 | 4.69 ± 0.72 | 18.07 ± 2.72 | 14.88 ± 2.38 | 2.76 ± 0.55 | 5.84 ± 0.92 | 1.62 ± 0.24 | 2.48 ± 0.50 | 1.20 ± 0.25 | 0.94 ± 0.09 | 0.93 ± 0.16 | 2.66 ± 0.31 | |||||||||||
| AA | 40 | 92.60 ± 45.66 | 4.69 ± 0.77 | 17.97 ± 2.87 | 14.75 ± 2.56 | 2.75 ± 0.55 | 5.78 ± 1.01 | 1.59 ± 0.26 | 2.43 ± 0.54 | 1.19 ± 0.20 | 0.95 ± 0.12 | 0.96 ± 0.16 | 2.70 ± 0.32 | |||||||||||
| p-value | - | 0.777 | 0.964 | 0.816 | 0.753 | 0.882 | 0.930 | 0.597 | 0.748 | 0.581 | 0.869 | 0.571 | 0.659 | |||||||||||
| GG/GA | - | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 0.895 | 1.000 | 1.000 | 1.000 | |||||||||||
| GA/AA | - | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | |||||||||||
| AA/GG | - | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 0.889 | 1.000 | |||||||||||
| c.93A>G | AA | 95 | 93.74 ± 47.82 | 4.70 ± 0.77 | 18.09 ± 2.87 | 14.93 ± 2.52 | 2.78 ± 0.56 | 5.84 ± 0.97 | 1.61 ± 0.25 | 2.47 ± 0.53 | 1.19 ± 0.24 | 0.95 ± 0.10 | 0.95 ± 0.16 | 2.64 ± 0.34 | ||||||||||
| AG | 86 | 88.11 ± 40.16 | 4.64 ± 0.72 | 17.74 ± 2.60 | 14.54 ± 2.24 | 2.69 ± 0.53 | 5.76 ± 0.92 | 1.60 ± 0.24 | 2.43 ± 0.49 | 1.18 ± 0.23 | 0.94 ± 0.09 | 0.92 ± 0.17 | 2.70 ± 0.30 | |||||||||||
| GG | 19 | 94.95 ± 34.51 | 4.74 ± 0.51 | 18.37 ± 2.53 | 15.03 ± 2.19 | 2.86 ± 0.36 | 5.92 ± 0.75 | 1.62 ± 0.19 | 2.57 ± 0.44 | 1.20 ± 0.18 | 0.94 ± 0.07 | 0.96 ± 0.12 | 2.74 ± 0.30 | |||||||||||
| p-value | - | 0.641 | 0.796 | 0.547 | 0.492 | 0.330 | 0.745 | 0.949 | 0.888 | 0.878 | 0.630 | 0.385 | 0.322 | |||||||||||
| AA/AG | - | 1.000 | 1.000 | 1.000 | 0.834 | 0.748 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 0.645 | 0.760 | |||||||||||
| AG/GG | - | 1.000 | 1.000 | 1.000 | 1.000 | 0.633 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | |||||||||||
| GG/AA | - | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 0.620 | |||||||||||
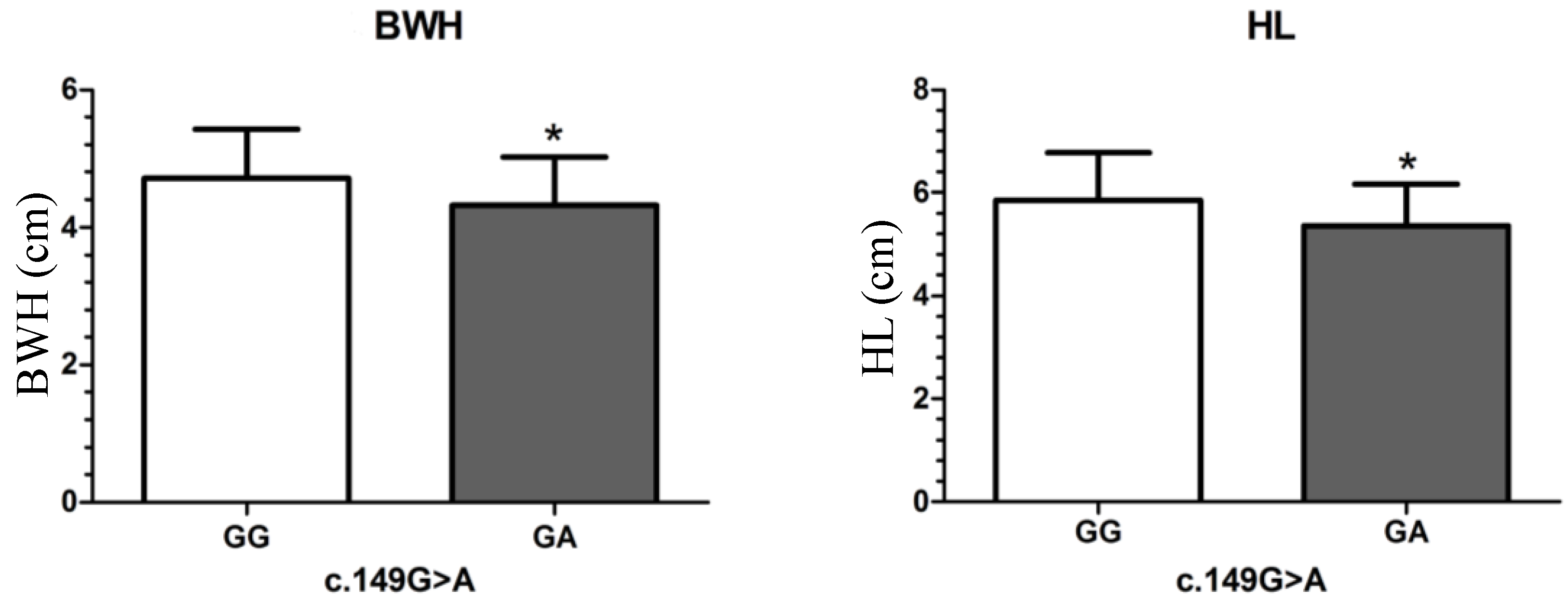
| c.149G>A | GG | 184 | 93.02 ± 44.09 | 4.71 ± 0.72 a | 18.06 ± 2.73 | 14.86 ± 2.37 | 2.76 ± 0.53 | 5.86 ± 0.93 a | 1.61 ± 0.24 | 2.47 ± 0.50 | 1.19 ± 0.23 | 0.95 ± 0.09 | 0.94 ± 0.16 | 2.68 ± 0.32 | ||||||||||
| GA | 16 | 73.25 ± 30.17 | 4.33 ± 0.70 b | 16.88 ± 2.42 | 13.78 ± 2.15 | 2.56 ± 0.51 | 5.36 ± 0.82 b | 1.50 ± 0.22 | 2.23 ± 0.50 | 1.14 ± 0.19 | 0.91 ± 0.09 | 0.89 ± 0.11 | 2.67 ± 0.30 | |||||||||||
| p-value | - | 0.081 | 0.041 * | 0.095 | 0.081 | 0.147 | 0.038 * | 0.070 | 0.073 | 0.445 | 0.139 | 0.269 | 0.987 | |||||||||||
| c.181A>G | AA | 187 | 91.04 ± 43.05 | 4.68 ± 0.72 | 17.93 ± 2.65 | 14.75 ± 2.29 | 2.75 ± 0.53 | 5.80 ± 0.91 | 1.60 ± 0.24 | 2.45 ± 0.50 | 1.18 ± 0.23 | 0.94 ± 0.09 | 0.93 ± 0.16 | 2.68 ± 0.32 | ||||||||||
| AG | 12 | 98.50 ± 51.91 | 4.72 ± 0.89 | 18.48 ± 3.92 | 15.18 ± 3.51 | 2.78 ± 0.69 | 6.09 ± 1.16 | 1.66 ± 0.26 | 2.60 ± 0.59 | 1.26 ± 0.25 | 0.96 ± 0.11 | 0.97 ± 0.21 | 2.63 ± 0.35 | |||||||||||
| GG | 1 | 82.00 ± 0.00 | 4.60 ± 0.00 | 18.00 ± 0.00 | 14.50 ± 0.00 | 2.50 ± 0.00 | 6.00 ± 0.00 | 1.60 ± 0.00 | 2.30 ± 0.00 | 1.10 ± 0.00 | 0.90 ± 0.00 | 0.90 ± 0.00 | 2.69 ± 0.00 | |||||||||||
| p-value | - | 0.828 | 0.979 | 0.802 | 0.829 | 0.875 | 0.557 | 0.741 | 0.570 | 0.502 | 0.790 | 0.783 | 0.870 | |||||||||||
| AA/AG | - | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 0.863 | 1.000 | 0.931 | 0.798 | 1.000 | 1.000 | 1.000 | |||||||||||
| AG/GG | - | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | |||||||||||
| c.193G>A | GG/AA | - | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | ||||||||||
| GG | 101 | 93.62 ± 47.51 | 4.71 ± 0.79 | 18.05 ± 2.89 | 14.88 ± 2.53 | 2.78 ± 0.58 | 5.83 ± 0.98 | 1.61 ± 0.25 | 2.47 ± 0.52 | 1.20 ± 0.24 | 0.95 ± 0.10 | 0.95 ± 0.17 | 2.66 ± 0.33 | |||||||||||
| GA | 80 | 89.40 ± 40.78 | 4.67 ± 0.68 | 17.85 ± 2.64 | 14.66 ± 2.26 | 2.71 ± 0.48 | 5.81 ± 0.91 | 1.60 ± 0.24 | 2.46 ± 0.50 | 1.18 ± 0.23 | 0.94 ± 0.08 | 0.93 ± 0.17 | 2.68 ± 0.31 | |||||||||||
| AA | 19 | 88.42 ± 30.70 | 4.59 ± 0.59 | 18.05 ± 2.23 | 14.68 ± 1.97 | 2.74 ± 0.47 | 5.80 ± 0.75 | 1.59 ± 0.21 | 2.34 ± 0.44 | 1.17 ± 0.17 | 0.93 ± 0.07 | 0.92 ± 0.11 | 2.72 ± 0.27 | |||||||||||
| p-value | - | 0.771 | 0.804 | 0.883 | 0.823 | 0.695 | 0.986 | 0.916 | 0.584 | 0.842 | 0.494 | 0.667 | 0.736 | |||||||||||
| GG/GA | - | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | |||||||||||
| GA/AA | - | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | |||||||||||
| AA/GG | - | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 0.906 | 1.000 | 0.815 | 1.000 | 1.000 | |||||||||||
| c.360C>T | CC | 76 | 86.60 ± 37.50 | 4.61 ± 0.70 | 17.74 ± 2.76 | 14.54 ± 2.37 | 2.68 ± 0.48 | 5.78 ± 0.89 | 1.58 ± 0.22 | 2.41 ± 0.50 | 1.16 ± 0.21 | 0.94 ± 0.08 | 0.92 ± 0.16 | 2.68 ± 0.31 | ||||||||||
| CT | 99 | 92.46 ± 46.06 | 4.70 ± 0.75 | 17.99 ± 2.64 | 14.81 ± 2.30 | 2.77 ± 0.57 | 5.80 ± 0.93 | 1.62 ± 0.26 | 2.46 ± 0.50 | 1.20 ± 0.25 | 0.95 ± 0.09 | 0.93 ± 0.16 | 2.67 ± 0.32 | |||||||||||
| TT | 25 | 102.08 ± 48.72 | 4.82 ± 0.71 | 18.56 ± 2.94 | 15.34 ± 2.64 | 2.86 ± 0.52 | 6.01 ± 1.01 | 1.62 ± 0.26 | 2.58 ± 0.55 | 1.22 ± 0.20 | 0.96 ± 0.13 | 1.00 ± 0.17 | 2.68 ± 0.36 | |||||||||||
| p-value | - | 0.288 | 0.410 | 0.428 | 0.340 | 0.312 | 0.537 | 0.552 | 0.326 | 0.428 | 0.522 | 0.119 | 0.944 | |||||||||||
| CC/CT | - | 1.000 | 1.000 | 1.000 | 1.000 | 0.899 | 1.000 | 0.937 | 1.000 | 0.901 | 0.890 | 1.000 | 1.000 | |||||||||||
| CT/TT | - | 0.969 | 1.000 | 1.000 | 0.966 | 1.000 | 0.938 | 1.000 | 0.795 | 1.000 | 0.785 | 0.261 | 1.000 | |||||||||||
| TT/CC | - | 0.370 | 0.629 | 0.586 | 0.440 | 0.461 | 0.839 | 1.000 | 0.405 | 0.791 | 0.659 | 0.120 | 1.000 | |||||||||||
2.3. Discussion
3. Experimental Section
3.1. Materials and Phenotypic Data Collection
3.2. PCR Amplification and SNP Identification
3.3. SNP Identification and Genotyping
3.4. Statistical Analysis
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Zhang, F.; Chen, Y.; Heiman, M.; DiMarchi, R. Leptin: Structure, function and biology. Vitam. Horm. 2005, 71, 345–372. [Google Scholar] [CrossRef]
- Denver, R.J.; Bonett, R.M.; Boorse, G.C. Evolution of leptin structure and function. Neuroendocrinology 2011, 94, 21–38. [Google Scholar] [CrossRef]
- Houseknecht, K.L.; Baile, C.A.; Matteri, R.L.; Spurlock, M.E. The biology of leptin: A review. J. Anim. Sci. 1998, 76, 1405–1420. [Google Scholar]
- Moschos, S.; Chan, J.L.; Mantzoros, C.S. Leptin and reproduction: A review. Fertil. Steril. 2002, 77, 433–444. [Google Scholar] [CrossRef]
- Máčajová, M.; Lamošová, D.; Zeman, M. Role of leptin in farm animals: A review. J. Vet. Med. A Physiol. Pathol. Clin. Med. 2004, 51, 157–166. [Google Scholar] [CrossRef]
- Kurokawa, T.; Uji, S.; Suzuki, T. Identification of cDNA coding for a homologue to mammalian leptin from pufferfish, Takifugu. rubripes. Peptides 2005, 26, 745–750. [Google Scholar]
- Huising, M.O.; Geven, E.J.; Kruiswijk, C.P.; Nabuurs, S.B.; Stolte, E.H.; Spanings, F.A.; Verburg-van, K.B.; Flik, G. Increased leptin expression in common Carp (Cyprinus. carpio) after food intake but not after fasting or feeding to satiation. Endocrinology 2006, 147, 5786–5797. [Google Scholar]
- Gorissen, M.; Bernier, N.J.; Nabuurs, S.B.; Flik, G.; Huising, M.O. Two divergent leptin paralogues in zebrafish (Danio. rerio) that originate early in teleostean evolution. J. Endocrinol. 2009, 201, 329–339. [Google Scholar] [CrossRef]
- Kurokawa, T.; Murashita, K. Genomic characterization of multiple leptin genes and a leptin receptor gene in the Japanese medaka, Oryzias. latipes. Gen. Comp. Endocrinol. 2009, 161, 229–237. [Google Scholar]
- Li, G.; Liang, X.; Xie, Q.; Li, G.; Yu, Y.; Lai, K. Gene structure, recombinant expression and functional characterization of grass carp leptin. Gen. Comp. Endocrinol. 2010, 166, 117–127. [Google Scholar] [CrossRef]
- Murashita, K.; Uji, S.; Yamamoto, T.; Rønnestad, I.; Kurokawa, T. Production of recombinant leptin and its effects on food intake in rainbow trout (Oncorhynchus. mykiss). Comp. Biochem. Physiol. B 2008, 150, 377–384. [Google Scholar] [CrossRef]
- Rønnestad, I.; Nilsen, T.O.; Murashita, K.; Angotzi, A.R.; Moen, A.G.; Stefansson, S.O.; Kling, P.; Björnsson, B.T.; Kurokawa, T. Leptin and leptin receptor genes in Atlantic salmon: Cloning, phylogeny, tissue distribution and expression correlated to long-term feeding status. Gen. Comp. Endocrinol. 2010, 168, 55–70. [Google Scholar]
- Won, E.T.; Baltzegar, D.A.; Picha, M.E.; Borski, R.J. Cloning and characterization of leptin in a Perciform fish, the striped bass (Morone. saxatilis): Control of feeding and regulation by nutritional state. Gen. Comp. Endocrinol. 2012, 178, 98–107. [Google Scholar] [CrossRef]
- Tinoco, A.B.; Nisembaum, L.G.; Isorna, E.; Delgado, M.J.; Pedro, N. Leptins and leptin receptor expression in the goldfish (Carassius. auratus). Regulation by food intake and fasting/overfeeding conditions. Peptides 2012, 34, 329–335. [Google Scholar]
- Zhang, H.; Chen, H.P.; Zhang, Y.; Li, S.S.; Lu, D.Q.; Zhang, H.F.; Meng, Z.N.; Liu, X.C.; Lin, H.R. Molecular cloning, characterization and expression profiles of multiple leptin genes and a leptin receptor gene in orange-spotted grouper (Epinephelus. coioides). Gen. Comp. Endocrinol. 2013, 181, 295–305. [Google Scholar]
- Ovilo, C.; Fernández, A.; Noguera, J.L.; Barragán, C.; Letón, R.; Rodríguez, C.; Mercadé, A.; Alves, E.; Folch, J.M.; Varona, L.; et al. Fine mapping of porcine chromosome 6 QTL and LEPR effects on body composition in multiple generations of an Iberian by Landrace intercross. Genet. Res. 2005, 85, 57–67. [Google Scholar] [CrossRef]
- Stachowiak, M.; Mackowski, M.; Madeja, Z.; Szydlowski, M.; Buszka, A.; Kaczmarek, P.; Rubis, B.; Mackowiak, P.; Nowak, K.W.; Switonski, M. Polymorphism of the porcine leptin gene promoter and analysis of its association with gene expression and fatness traits. Biochem. Genet. 2007, 45, 245–253. [Google Scholar] [CrossRef]
- Uemoto, Y.; Kikuchi, T.; Nakano, H.; Sato, S.; Shibata, T.; Kadowaki, H.; Katoh, K.; Kobayashi, E.; Suzuki, K. Effects of porcine leptin receptor gene polymorphisms on backfat thickness, fat area ratios by image analysis, and serum leptin concentrations in a Duroc purebred population. Anim. Sci. J. 2012, 83, 375–385. [Google Scholar]
- Buchanan, F.C.; Fitzsimmons, C.J.; van Kessel, A.G.; Thue, T.D.; Winkelman-Sim, D.C.; Schmutz, S.M. Association of a missense mutation in the bovine leptin gene with carcass fat content and leptin mRNA levels. Genet. Sel. Evol. 2002, 34, 105–116. [Google Scholar] [CrossRef]
- Guo, Y.; Chen, H.; Lan, X.; Zhang, B.; Pan, C.; Zhang, L.; Zhang, C.; Zhao, M. Novel SNPs of the bovine LEPR gene and their association with growth traits. Biochem. Genet. 2008, 46, 828–834. [Google Scholar] [CrossRef]
- Kulig, H.; Kmieć, M. Association between leptin gene polymorphisms and growth traits in Limousin cattle. Russ. J. Genet. 2009, 45, 738–741. [Google Scholar]
- Clempson, A.M.; Pollott, G.E.; Brickell, J.S.; Bourne, N.E.; Munce, N.; Wathes, D.C. Evidence that leptin genotype is associated with fertility, growth, and milk production in Holstein cows. J. Dairy Sci. 2011, 94, 3618–3628. [Google Scholar] [CrossRef]
- Melucci, L.M.; Panarace, M.; Feula, P.; Villarreal, E.L.; Grigioni, G.; Carduza, F.; Soria, L.A.; Mezzadra, C.A.; Arceo, M.E.; Papaleo Mazzucco, J.; et al. Genetic and management factors affecting beef quality in grazing Hereford steers. Meat Sci. 2012, 92, 768–774. [Google Scholar] [CrossRef]
- Da, S.R.; Ferraz, J.B.; Meirelles, F.V.; Eler, J.P.; Balieiro, J.C.; Cucco, D.C.; Mattos, E.C.; Rezende, F.M.; Silva, S.L. Association of single nucleotide polymorphisms in the bovine leptin and leptin receptor genes with growth and ultrasound carcass traits in Nellore cattle. Genet. Mol. Res. 2012, 11, 3721–3728. [Google Scholar]
- Wei, Y.; Huang, H.; Meng, Z.; Zhang, Y.; Luo, J.; Chen, G.; Lin, H. Single nucleotide polymorphisms in the leptin-a Gene and associations with growth traits in the orange-spotted grouper (Epinephelus. coioides). Int. J. Mol. Sci. 2013, 14, 8625–8637. [Google Scholar] [CrossRef]
- Von Heijne, G. The signal peptide. J. Membr. Biol. 1990, 115, 195–201. [Google Scholar]
- Copeland, D.L.; Duff, R.J.; Liu, Q.; Prokop, J.; Londraville, R.L. Leptin in teleost fishes: An argument for comparative study. Front. Physiol. 2011, 2, 1–11. [Google Scholar]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual, 3rd ed.; Cold Spring Harbor Laboratory Press: Woodbury, NY, USA, 2001. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Huang, H.; Wei, Y.; Meng, Z.; Zhang, Y.; Liu, X.; Guo, L.; Luo, J.; Chen, G.; Lin, H. Polymorphisms of Leptin-b Gene Associated with Growth Traits in Orange-Spotted Grouper (Epinephelus coioides). Int. J. Mol. Sci. 2014, 15, 11996-12006. https://doi.org/10.3390/ijms150711996
Huang H, Wei Y, Meng Z, Zhang Y, Liu X, Guo L, Luo J, Chen G, Lin H. Polymorphisms of Leptin-b Gene Associated with Growth Traits in Orange-Spotted Grouper (Epinephelus coioides). International Journal of Molecular Sciences. 2014; 15(7):11996-12006. https://doi.org/10.3390/ijms150711996
Chicago/Turabian StyleHuang, Hai, Yun Wei, Zining Meng, Yong Zhang, Xiaochun Liu, Liang Guo, Jian Luo, Guohua Chen, and Haoran Lin. 2014. "Polymorphisms of Leptin-b Gene Associated with Growth Traits in Orange-Spotted Grouper (Epinephelus coioides)" International Journal of Molecular Sciences 15, no. 7: 11996-12006. https://doi.org/10.3390/ijms150711996
APA StyleHuang, H., Wei, Y., Meng, Z., Zhang, Y., Liu, X., Guo, L., Luo, J., Chen, G., & Lin, H. (2014). Polymorphisms of Leptin-b Gene Associated with Growth Traits in Orange-Spotted Grouper (Epinephelus coioides). International Journal of Molecular Sciences, 15(7), 11996-12006. https://doi.org/10.3390/ijms150711996




