Expression of Human Endogenous Retrovirus env Genes in the Blood of Breast Cancer Patients
Abstract
:1. Introduction
2. Results and Discussion
| Patient No. | Gender | Age | Status | Stage | CTx | Rx | Expression (2−ΔΔCt) | |||
|---|---|---|---|---|---|---|---|---|---|---|
| HERV-R | HERV-H | HERV-K | HERV-P | |||||||
| 1 | Female | 25 | Normal | NA | − | − | 1.54806924 | 1.46936213 | 0.9275391 | 0.99967427 |
| 2 | Female | 31 | Normal | NA | − | − | 1.471890146 | 2.46791037 | 1.37724001 | 1.14023406 |
| 3 | Male | 25 | Normal | NA | − | − | 1.538228683 | 4.95879643 | 2.59526166 | 1.73282985 |
| 4 | Male | 38 | Normal | NA | − | − | 1.512746241 | 8.0583809 | 6.39294198 | 2.66203585 |
| 5 | Female | 25 | Normal | NA | − | − | 1.071157036 | 13.4086306 | 6.87891009 | 3.32217944 |
| 6 | Male | 31 | Normal | NA | − | − | 1.414870488 | 1.85316328 | 1.16342644 | 1.00558149 |
| 7 | Female | 25 | Normal | NA | − | − | 0.633655233 | 0.28319938 | 0.13652906 | 0.4591158 |
| 8 | Male | 26 | Normal | NA | − | − | 1.061327569 | 0.51662523 | 0.23777682 | 0.54624876 |
| 9 | Male | 25 | Normal | NA | − | − | 0.155951572 | 0.60970784 | 0.11957436 | 0.27466215 |
| 10 | Male | 26 | Normal | NA | − | − | 0.898300474 | 2.82527245 | 1.488831 | 0.80934684 |
| 11 | Female | 65 | Benign | NA | − | − | 0.969426438 | 6.60804418 | 1.16919319 | 2.01306906 |
| 12 | Female | 54 | Benign | NA | − | − | 1.316648275 | 10.5628214 | 2.48689685 | 3.75981932 |
| 13 | Female | 47 | Benign | NA | − | − | 1.131003453 | 7.46317751 | 2.59460214 | 1.52509241 |
| 14 | Female | 62 | Benign | NA | − | − | 1.235136082 | 11.1308356 | 2.20403429 | 1.42355345 |
| 15 | Female | 23 | Benign | NA | − | − | 3.067726784 | 5.04920645 | 0.92465046 | 2.9775588 |
| 16 | Female | 72 | Benign | NA | − | − | 1.958437833 | 3.11879775 | 0.72987096 | 0.95265781 |
| 17 | Female | 53 | Benign | NA | − | − | 8.088114008 | 0.20038179 | 0.03046887 | 9.30321907 |
| 18 | Female | 45 | Primary | 3c | − | − | 5.319792596 | 18.2529364 | 6.3774514 | 8.69446145 |
| 19 | Female | 64 | Primary | 2a | − | − | 16.14262545 | 39.6795383 | 10.1711671 | 15.4610172 |
| 20 | Female | 69 | Primary | 2a | − | − | 9.608447428 | 8.99396124 | 4.93020169 | 8.97988312 |
| 21 | Female | 48 | Primary | 1 | − | − | 8.913363856 | 87.2262221 | 66.7816577 | 20.8662098 |
| 22 | Female | 57 | Primary | 1 | − | − | 3.813819767 | 18.7491939 | 9.5227615 | 7.94857735 |
| 23 | Female | 58 | Primary | 4 | − | − | 4.12807734 | 3.28552366 | 1.48206974 | 3.14537058 |
| 24 | Female | 75 | Primary | 2a | − | − | 3.549551305 | 3.70504226 | 0.99168007 | 0.61249218 |
| 25 | Female | 49 | Primary | 0 | − | − | 6.627064679 | 33.2770737 | 63.1836773 | 16.2480724 |
| 26 | Female | 41 | Primary | 1 | − | − | 7.63980951 | 24.3444893 | 18.1808364 | 6.60914356 |
| 27 | Female | 64 | Primary | 2a | − | − | 5.867326733 | 2.70049398 | 0.86049974 | 5.10306258 |
| 28 | Female | 44 | Primary chemo | 2b | FEC | + | 0.583229932 | 3.42132858 | 1.35070567 | 0.70656621 |
| 29 | Female | 59 | Primary chemo | 1 | FEC | − | 1.922348628 | 19.9588497 | 13.4846546 | 2.12642504 |
| 30 | Female | 89 | Primary chemo | 2b | FEC | − | 2.655413743 | 1.21267252 | 0.32439272 | 1.91732734 |
| 31 | Female | 78 | Primary chemo | 2a | CMF | + | 6.824878361 | 9.60170093 | 4.13870667 | 4.4970014 |
| 32 | Female | 51 | Primary chemo | 1 | FEC | + | 1.27006588 | 6.65339183 | 1.08423689 | 7.41885563 |
| 33 | Female | 46 | Primary chemo | 1 | FAC | − | 1.703344207 | 28.0725848 | 5.5797997 | 7.00473271 |
| 34 | Female | 46 | Primary chemo | 3c | AC-taxol | + | 0.730760217 | 2.65103674 | 0.43893524 | 1.69693347 |
| 35 | Female | 52 | Primary chemo | 2a | AC | + | 3.813907886 | 22.6455656 | 5.9942531 | 6.20184155 |
| 36 | Female | 50 | Primary chemo | 1 | FEC | + | 2.83884353 | 14.931874 | 6.69126972 | 6.64835153 |
| 37 | Female | 61 | Primary chemo | 1 | FEC | + | 8.470829981 | 63.937481 | 24.7538325 | 21.6310231 |
| 38 | Female | 53 | Primary chemo | 1 | CMF | + | 2.113845596 | 12.3478675 | 3.84667871 | 3.80772688 |
| 39 | Female | 46 | Primary chemo | 2a | FEC | + | 2.873687797 | 11.3739122 | 8.0083961 | 5.84659504 |
| 40 | Female | 48 | Primary chemo | 2a | FEC | + | 2.776120133 | 25.2437908 | 17.7988649 | 10.6208697 |
| 41 | Female | 64 | Primary chemo | 2a | TA | + | 0.544990631 | 4.74241966 | 1.46403157 | 1.24584967 |
| 42 | Female | 56 | Primary chemo | 1 | FEC | + | 3.620787059 | 16.8844506 | 13.754324 | 4.36202085 |
| 43 | Female | 53 | Primary chemo | 2a | CMF | − | 1.035985157 | 2.41299234 | 1.01158204 | 1.64714017 |
| 44 | Female | 49 | Primary chemo | 2b | FEC | + | 1.265233213 | 45.0438009 | 6.23611126 | 1.15212453 |
| 45 | Female | 36 | Primary chemo | 3b | TA | + | 1.342234427 | 30.3275419 | 8.30499903 | 1.51382314 |
| 46 | Female | 50 | Primary chemo | 2b | CMF | + | 3.154118071 | 95.4643386 | 29.5362207 | 2.34984131 |
| 47 | Female | 64 | Primary chemo | 1 | FEC | + | 2.795558755 | 66.8485023 | 27.5099193 | 4.24480025 |
| 48 | Female | 33 | Meta no chemo | 4 | FEC | + | 0.746425408 | 0.45963376 | 0.07435361 | 0.68133206 |
| 49 | Female | 47 | Meta no chemo | 4 | CMF | + | 3.49050715 | 24.6256517 | 3.02817098 | 2.55660628 |
| 50 | Female | 43 | Meta no chemo | 4 | CMF | + | 0.397355807 | 1.72938211 | 0.38039759 | 0.10406547 |
| 51 | Female | 49 | Meta no chemo | 4 | CMF | + | 9.724135523 | 10.9923149 | 2.68548004 | 6.7503548 |
| 52 | Female | 56 | Meta no chemo | 4 | CMF | + | 6.626911563 | 13.408011 | 7.89577019 | 3.87500096 |
| 53 | Female | 49 | Meta chemo | 4 | CMF | + | 1.383737227 | 0.53792987 | 0.12805599 | 1.64584674 |
| 54 | Female | 53 | Meta chemo | 4 | FEC | + | 2.838056544 | 0.50303244 | 0.26561588 | 0.84934936 |
| 55 | Female | 47 | Meta chemo | 4 | AC | − | 0.973399092 | 1.9376706 | 0.39927279 | 3.47905785 |
| 56 | Female | 59 | Meta chemo | 4 | CMF | + | 1.757514591 | 8.90854611 | 5.13375104 | 2.23178286 |
| 57 | Female | 60 | Meta chemo | 4 | FEC | + | 0.976259554 | 11.4398006 | 4.19425139 | 1.56341054 |
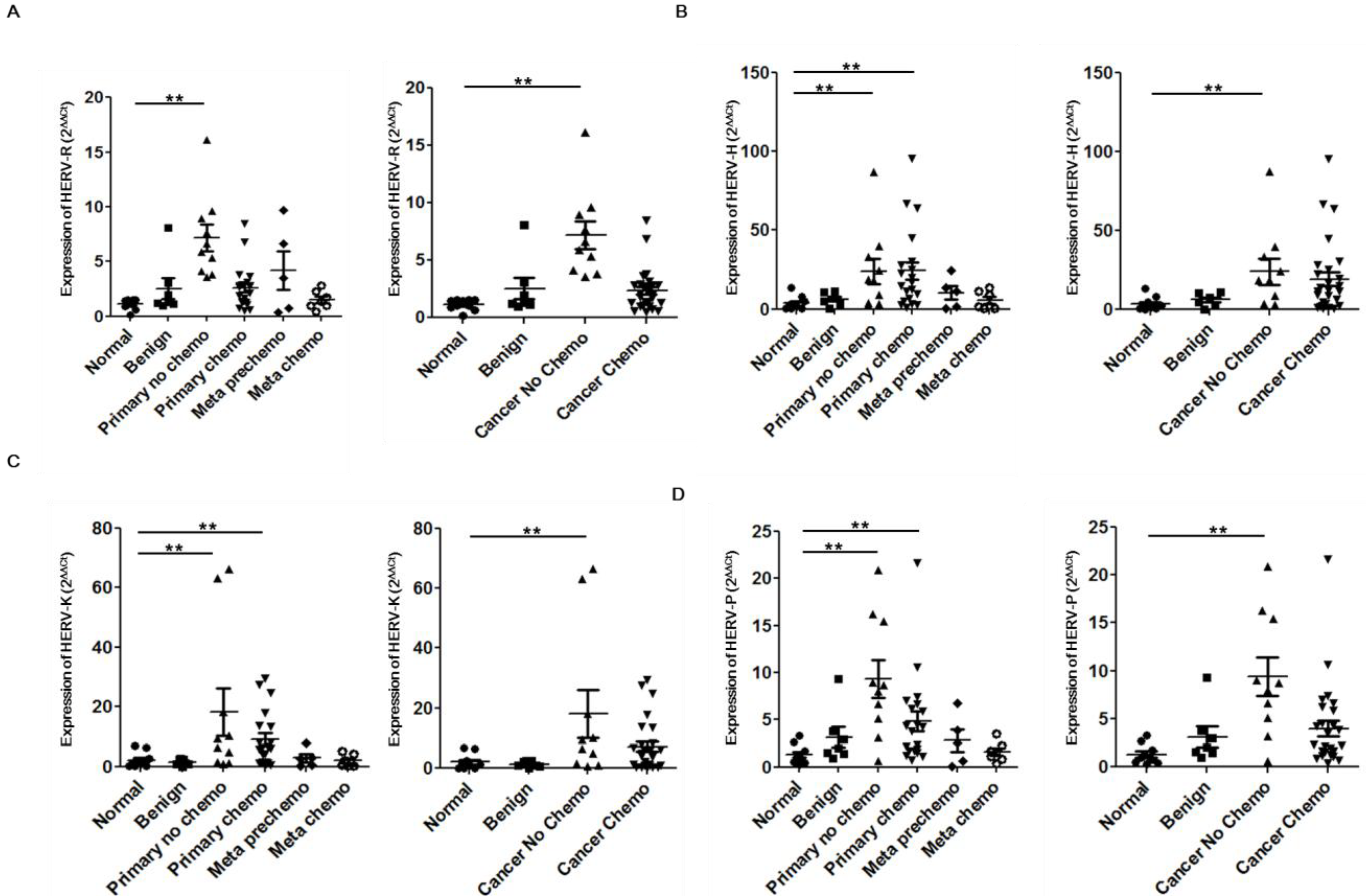
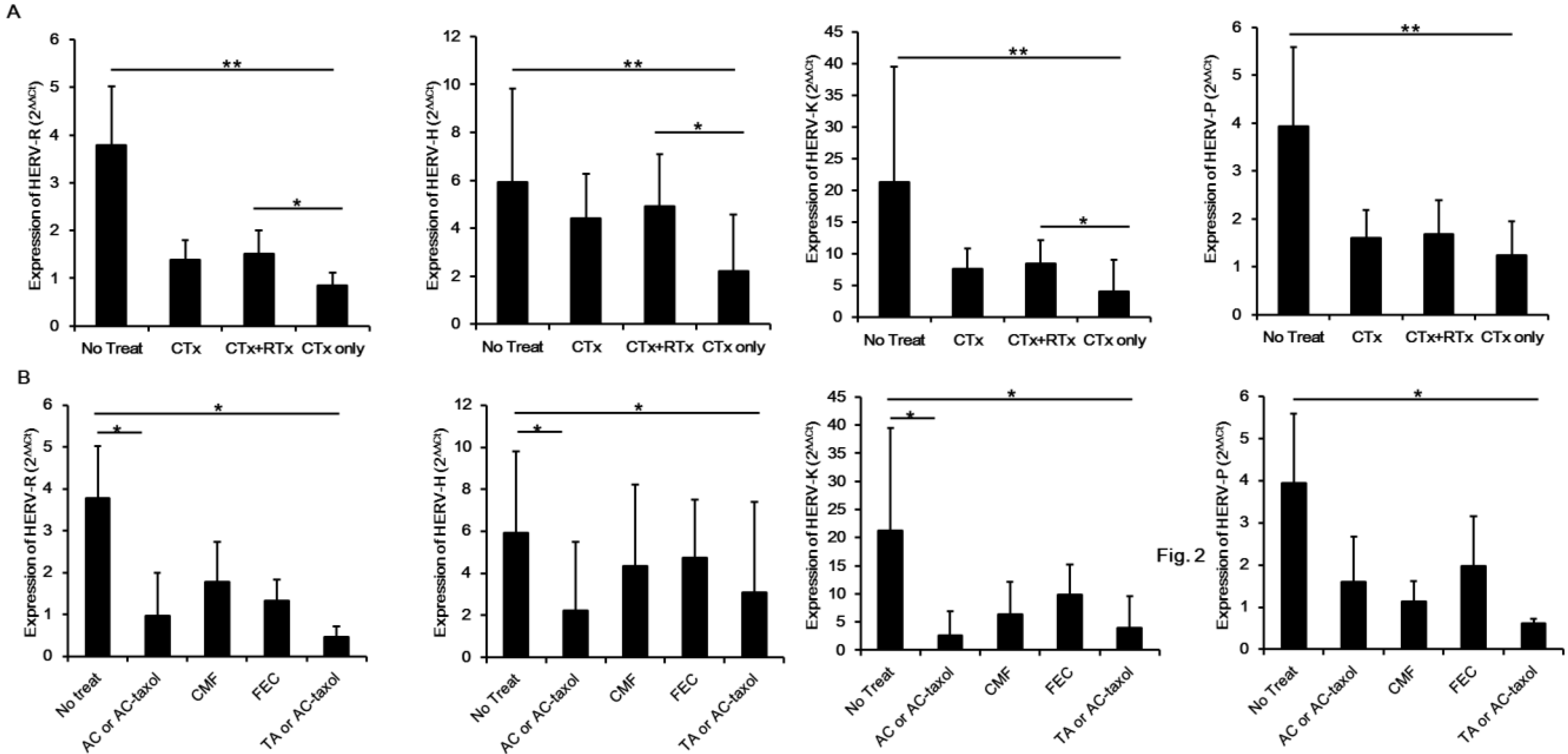
3. Experimental Section
3.1. Blood Samples of Patients
3.2. Real-Time PCR Analysis
| Target Gene | Primers | GenBank Accession No. | |
|---|---|---|---|
| Forward | Reverse | ||
| HERV-R env | 5'-CATGGGAAGCAAGGGAACT-3' | 5'-CTTTCCCCAGCGAGCAATAC-3' | AC073210 from Chr.7q11.21 |
| HERV-H env | 5'-TTCACTCCATCCTTGGCTAT-3' | 5'-CGTCGAGTATCTACGAGCAAT-3' | AJ289711 from Chr.2q24.3 |
| HERV-K env | 5'-CACAACTAAAGAAGCTGACG-3' | 5'-CATAGGCCCAGTTGGTATAG-3' | AC074261 from Chr12q14.1 |
| HERV-P env | 5'-CAAGATTGGGTCCCCTCAC-3' | 5'-CCTATGGGGTCTTTCCCTC-3' | DQ247958 from Chr.14q32.12 |
4. Conclusions
Supplementary Files
Acknowledgments
Conflicts of Interest
References
- Hayes, M.; Whitesell, M.; Brown, M.A. Pathological and evolutionary implications of retroviruses as mobile genetic elements. Genes 2013, 4, 573–582. [Google Scholar] [CrossRef]
- Belshaw, R.; Pereira, V.; Katzourakis, A.; Talbot, G.; Paces, J.; Burt, A.; Tristem, M. Long-term reinfection of the human genome by endogenous retroviruses. Proc. Natl. Acad. Sci. USA 2004, 101, 4894–4899. [Google Scholar]
- Belshaw, R.; Dawson, A.L.; Woolven-Allen, J.; Redding, J.; Burt, A.; Tristem, M. Genomewide screening reveals high levels of insertional polymorphism in the human endogenous retrovirus family HERV-K(HML2): Implications for present-day activity. J. Virol. 2005, 79, 12507–12514. [Google Scholar] [CrossRef]
- Bannert, N.; Kurth, R. Retroelements and the human genome: New perspectives on an old relation. Proc. Natl. Acad. Sci. USA 2004, 101, 14572–14579. [Google Scholar] [CrossRef]
- Nelson, P.N.; Carnegie, P.R.; Martin, J.; Davari Ejtehadi, H.; Hooley, P.; Roden, D.; Rowland-Jones, S.; Warren, P.; Astley, J.; Murray, P.G. Demystified. Human endogenous retroviruses. Mol. Pathol. 2003, 56, 11–18. [Google Scholar]
- Singh, S.K. Endogenous retroviruses: Suspects in the disease world. Future Microbiol. 2007, 2, 269–275. [Google Scholar]
- Mameli, G.; Astone, V.; Arru, G.; Marconi, S.; Lovato, L.; Serra, C.; Sotgiu, S.; Bonetti, B.; Dolei, A. Brains and peripheral blood mononuclear cells of multiple sclerosis (MS) patients hyperexpress MS-associated retrovirus/HERV-W endogenous retrovirus, but not Human herpesvirus 6. J. Gen. Virol. 2007, 88, 264–274. [Google Scholar] [CrossRef]
- Serra, C.; Mameli, G.; Arru, G.; Sotgiu, S.; Rosati, G.; Dolei, A. In vitro modulation of the multiple sclerosis (MS)-associated retrovirus by cytokines: Implications for MS pathogenesis. J. Neurovirol. 2003, 9, 637–643. [Google Scholar]
- Yolken, R. Viruses and schizophrenia: A focus on herpes simplex virus. Herpes 2004, 11, 83A–88A. [Google Scholar]
- Garrison, K.E.; Jones, R.B.; Meiklejohn, D.A.; Anwar, N.; Ndhlovu, L.C.; Chapman, J.M.; Erickson, A.L.; Agrawal, A.; Spotts, G.; Hecht, F.M.; et al. T cell responses to human endogenous retroviruses in HIV-1 infection. PLoS Pathog. 2007, 3, e165. [Google Scholar] [CrossRef]
- Boyd, M.T.; Bax, C.M.; Bax, B.E.; Bloxam, D.L.; Weiss, R.A. The human endogenous retrovirus ERV-3 is upregulated in differentiating placental trophoblast cells. Virology 1993, 196, 905–909. [Google Scholar] [CrossRef]
- Takeuchi, K.; Katsumata, K.; Ikeda, H.; Minami, M.; Wakisaka, A.; Yoshiki, T. Expression of endogenous retroviruses, ERV3 and lambda 4–1, in synovial tissues from patients with rheumatoid arthritis. Clin. Exp. Immunol. 1995, 99, 338–344. [Google Scholar]
- Larsson, E.; Venables, P.J.; Andersson, A.C.; Fan, W.; Rigby, S.; Botling, J.; Oberg, F.; Cohen, M.; Nilsson, K. Expression of the endogenous retrovirus ERV3 (HERV-R) during induced monocytic differentiation in the U-937 cell line. Int. J. Cancer 1996, 67, 451–456. [Google Scholar]
- Andersson, A.C.; Venables, P.J.; Tonjes, R.R.; Scherer, J.; Eriksson, L.; Larsson, E. Developmental expression of HERV-R (ERV3) and HERV-K in human tissue. Virology 2002, 297, 220–225. [Google Scholar] [CrossRef]
- Andersson, A.C.; Yun, Z.; Sperber, G.O.; Larsson, E.; Blomberg, J. ERV3 and related sequences in humans: Structure and RNA expression. J. Virol. 2005, 79, 9270–9284. [Google Scholar] [CrossRef]
- Ahn, K.; Kim, H.S. Structural and quantitative expression analyses of HERV gene family in human tissues. Mol. Cells 2009, 28, 99–103. [Google Scholar] [CrossRef]
- Wang-Johanning, F.; Liu, J.; Rycaj, K.; Huang, M.; Tsai, K.; Rosen, D.G.; Chen, D.T.; Lu, D.W.; Barnhart, K.F.; Johanning, G.L. Expression of multiple human endogenous retrovirus surface envelope proteins in ovarian cancer. Int. J. Cancer 2007, 120, 81–90. [Google Scholar] [CrossRef]
- Contreras-Galindo, R.; Kaplan, M.H.; Leissner, P.; Verjat, T.; Ferlenghi, I.; Bagnoli, F.; Giusti, F.; Dosik, M.H.; Hayes, D.F.; Gitlin, S.D.; et al. Human endogenous retrovirus K (HML-2) elements in the plasma of people with lymphoma and breast cancer. J. Virol. 2008, 82, 9329–9336. [Google Scholar] [CrossRef]
- Zhao, J.; Rycaj, K.; Geng, S.; Li, M.; Plummer, J.B.; Yin, B.; Liu, H.; Xu, X.; Zhang, Y.; Yan, Y.; et al. Expression of human endogenous retrovirus type K envelope protein is a novel candidate prognostic marker for human breast cancer. Genes Cancer 2011, 2, 914–922. [Google Scholar] [CrossRef]
- Wang-Johanning, F.; Rycaj, K.; Plummer, J.B.; Li, M.; Yin, B.; Frerich, K.; Garza, J.G.; Shen, J.; Lin, K.; Yan, P.; et al. Immunotherapeutic potential of anti-human endogenous retrovirus-K envelope protein antibodies in targeting breast tumors. J. Natl. Cancer Inst. 2012, 104, 189–210. [Google Scholar] [CrossRef]
- Wang-Johanning, F.; Li, M.; Esteva, F.J.; Hess, K.R.; Yin, B.; Rycaj, K.; Plummer, J.B.; Garza, J.G.; Ambs, S.; Johanning, G.L. Human endogenous retrovirus type K antibodies and mRNA as serum biomarkers of early-stage breast cancer. Int. J. Cancer 2014, 134, 587–595. [Google Scholar] [CrossRef]
- Cegolon, L.; Salata, C.; Weiderpass, E.; Vineis, P.; Palu, G.; Mastrangelo, G. Human endogenous retroviruses and cancer prevention: Evidence and prospects. BMC Cancer 2013, 13, 4. [Google Scholar] [CrossRef]
- Schiavetti, F.; Thonnard, J.; Colau, D.; Boon, T.; Coulie, P.G. A human endogenous retroviral sequence encoding an antigen recognized on melanoma by cytolytic T lymphocytes. Cancer Res. 2002, 62, 5510–5516. [Google Scholar]
- Kang, Y.J.; Jo, J.O.; Ock, M.S.; Chang, H.K.; Baek, K.W.; Lee, J.R.; Choi, Y.H.; Kim, W.J.; Leem, S.H.; Kim, H.S.; et al. Human ERV3–1 env protein expression in various human tissues and tumours. J. Clin. Pathol. 2014, 67, 86–90. [Google Scholar] [CrossRef]
- Golan, M.; Hizi, A.; Resau, J.H.; Yaal-Hahoshen, N.; Reichman, H.; Keydar, I.; Tsarfaty, I. Human endogenous retrovirus (HERV-K) reverse transcriptase as a breast cancer prognostic marker. Neoplasia 2008, 10, 521–533. [Google Scholar]
- Yi, J.M.; Kim, H.M.; Kim, H.S. Human endogenous retrovirus HERV-H family in human tissues and cancer cells: Expression, identification, and phylogeny. Cancer Lett. 2006, 231, 228–239. [Google Scholar] [CrossRef]
- Yi, J.M.; Schuebel, K.; Kim, H.S. Molecular genetic analyses of human endogenous retroviral elements belonging to the HERV-P family in primates, human tissues, and cancer cells. Genomics 2007, 89, 1–9. [Google Scholar] [CrossRef]
- Lee, J.R.; Ahn, K.; Kim, Y.J.; Jung, Y.D.; Kim, H.S. Radiation-induced human endogenous retrovirus (HERV)-R env gene expression by epigenetic control. Radiat. Res. 2012, 178, 379–384. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Rhyu, D.-W.; Kang, Y.-J.; Ock, M.-S.; Eo, J.-W.; Choi, Y.-H.; Kim, W.-J.; Leem, S.-H.; Yi, J.-M.; Kim, H.-S.; Cha, H.-J. Expression of Human Endogenous Retrovirus env Genes in the Blood of Breast Cancer Patients. Int. J. Mol. Sci. 2014, 15, 9173-9183. https://doi.org/10.3390/ijms15069173
Rhyu D-W, Kang Y-J, Ock M-S, Eo J-W, Choi Y-H, Kim W-J, Leem S-H, Yi J-M, Kim H-S, Cha H-J. Expression of Human Endogenous Retrovirus env Genes in the Blood of Breast Cancer Patients. International Journal of Molecular Sciences. 2014; 15(6):9173-9183. https://doi.org/10.3390/ijms15069173
Chicago/Turabian StyleRhyu, Dong-Won, Yun-Jeong Kang, Mee-Sun Ock, Jung-Woo Eo, Yung-Hyun Choi, Wun-Jae Kim, Sun-Hee Leem, Joo-Mi Yi, Heui-Soo Kim, and Hee-Jae Cha. 2014. "Expression of Human Endogenous Retrovirus env Genes in the Blood of Breast Cancer Patients" International Journal of Molecular Sciences 15, no. 6: 9173-9183. https://doi.org/10.3390/ijms15069173
APA StyleRhyu, D.-W., Kang, Y.-J., Ock, M.-S., Eo, J.-W., Choi, Y.-H., Kim, W.-J., Leem, S.-H., Yi, J.-M., Kim, H.-S., & Cha, H.-J. (2014). Expression of Human Endogenous Retrovirus env Genes in the Blood of Breast Cancer Patients. International Journal of Molecular Sciences, 15(6), 9173-9183. https://doi.org/10.3390/ijms15069173









