Selection of Suitable Endogenous Reference Genes for Relative Copy Number Detection in Sugarcane
Abstract
:1. Introduction
2. Results
2.1. Assessment of Primer Within-Species-Specificity
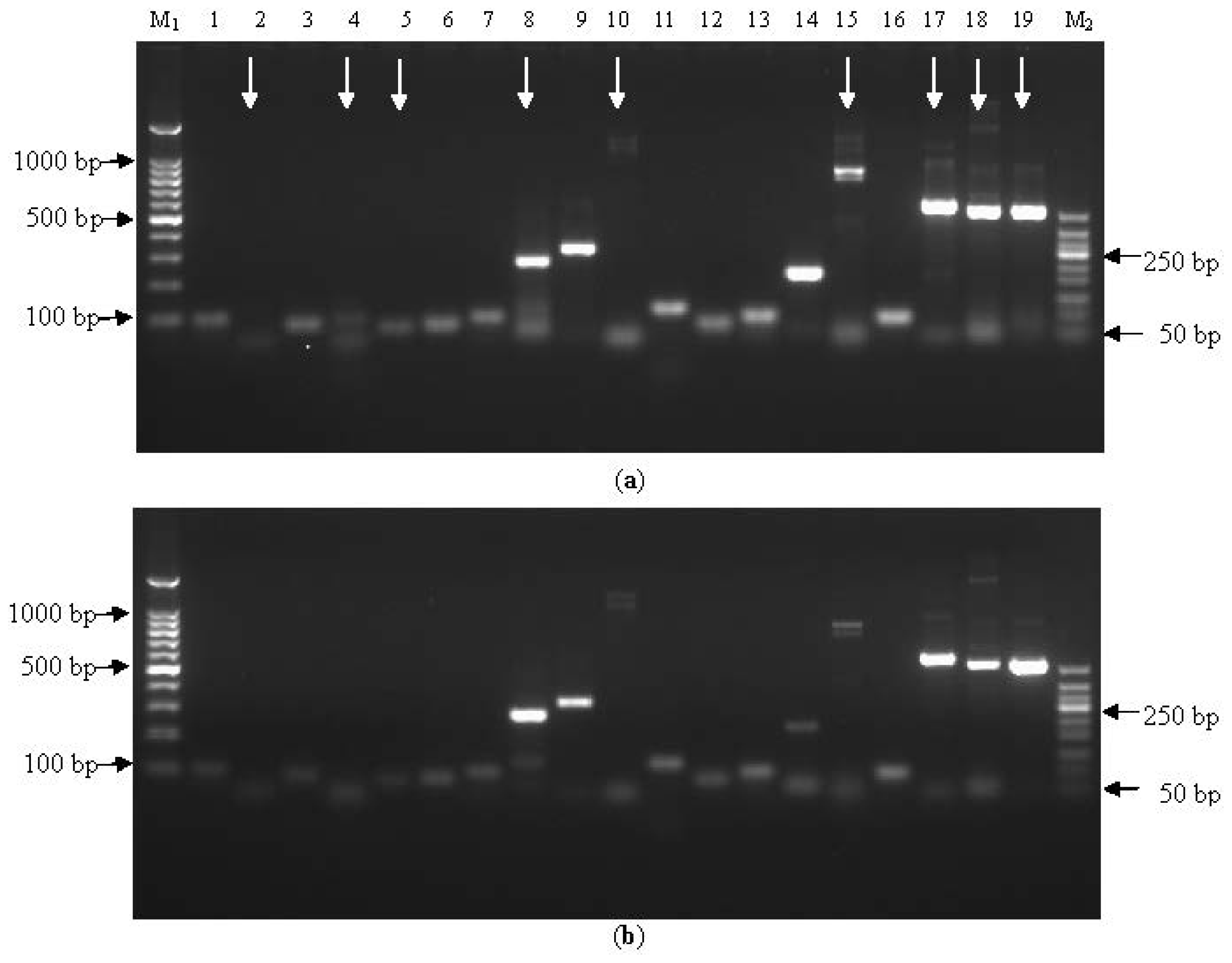
2.1.1. Assessment of Primer Within-Species-Specificity by PCR
2.1.2. Assessment of Primer Performance by qPCR
2.2. Screening for Low Copy Genes
2.2.1. Copy Number Calculation of the Candidate Reference Genes
2.2.2. Stability Analysis of the Selected Endogenous Reference Genes in Different Sugarcane Varieties
2.2.3. Evaluation of Three Candidate Genes in a Multi-Target Reference Plasmid
3. Discussion
4. Experimental Section
4.1. Tissue Samples
4.2. Reagents and Consumables
4.3. Extraction and Isolation of Genomic DNA
4.4. Primer Design for Candidate Endogenous Reference Genes
4.5. Polymerase Chain Reaction (PCR)
4.6. Preparation of Recombinant Plasmid
4.7. Calculation of Plasmid Copy Number
4.8. Real-Time Quantitative PCR
4.9. Establishment of Standard Curve
4.10. Copy Number Calculation of Endogenous Reference Genes in Sugarcane
4.11. Construction of Multi-Target Reference Plasmid for Screening Low Copy Genes
5. Conclusions
Supplementary Information
ijms-15-08846-s001.pdfAcknowledgments
Conflicts of Interest
- Author ContributionsConceived and designed the experiments: L.X., J.G., Y.Q.; Performed the experiments: B.X., J.G., Z.F.; Analyzed the data: B.X., J.G., L.X.; Wrote the paper: B.X., L.X., J.G., Y.Q.; Revised the paper: L.X., J.G., Y.Q., L.W.; Approved the final version of the paper: L.X.
References
- Casu, R.E.; Selivanova, A.; Perroux, J.M. High-throughput assessment of transgene copy number in sugarcane using real-time quantitative PCR. Plant Cell Rep 2012, 31, 167–177. [Google Scholar]
- Arencibia, A.; Vázquez, R.I.; Prieto, D.; Téllez, D.; Carmona, E.R.; Coego, A.; Hernández, L.; de la riva, G.A.; Selman-Housein, G. Transgenic sugarcane plants resistant to stem borer attack. Mol. Breed 1997, 3, 247–255. [Google Scholar]
- Li, W.; Bin, T.; Guo, W.W. Estimating transgene copy number in precocious trifoliate orange by TaqMan real-time PCR. Plant Cell Tissue Organ Cult 2012, 109, 363–371. [Google Scholar]
- Srikanth, J.; Subramonian, N.; Premachandran, M.N. Advances in transgenic research for insect resistance in sugarcane. Trop. Plant Biol 2011, 4, 52–61. [Google Scholar]
- Lakshmanan, P.; Geijskes, R.J.; Aitken, K.S.; Grof, C.L.P.; Bonnett, G.D.; Smith, G.R. Sugarcane biotechnology: The challenges and opportunities. In Vitro Cell. Dev. Biol.-Plant 2005, 41, 345–363. [Google Scholar]
- Vaucheret, H.; Fagard, M. Transcriptional gene silencing in plants: targets, inducers and regulators. Trends Genet 2001, 17, 29–35. [Google Scholar]
- Tang, W.; Newton, R.J.; Weidner, D.A. Genetic transformation and gene silencing mediated by multiple copies of a transgene in eastern white pine. J. Exp. Bot 2007, 58, 545–554. [Google Scholar]
- Dai, S.H.; Zheng, P.; Marmey, P.; Zhang, S.P.; Tian, W.Z.; Chen, S.Y.; Beachy, R.N.; Fauquet, C. Comparative analysis of transgenic rice plants obtained by Agrobacterium-mediated transformation and particle bombardment. Mol. Breed 2001, 7, 25–33. [Google Scholar]
- Travella, S.; Ross, S.M.; Harden, J.; Everett, C.; Snape, J.W.; Harwood, W.A. A comparison of transgenic barley lines produced by particle bombardment and Agrobacterium-mediated techniques. Plant Cell Rep 2005, 23, 780–789. [Google Scholar]
- Arencibia, A.D.; Carmona, E.R.; Tellez, P.; Chan, M.T.; Yu, S.M.; Trujillo, L.E.; Oramas, P. An efficient protocol for sugarcane (Saccharum spp. L.) transformation mediated by Agrobacterium tumefaciens. Transgenic Res 1998, 7, 213–222. [Google Scholar]
- Lucito, R.; West, J.; Reiner, A.; Alexander, J.; Esposito, D.; Mishra, B.; Powers, S.; Norton, L.; Wigler, M. Detecting gene copy number fluctuation in tumor cells by microarray analysis of genomic representations. Genome Res 2000, 10, 1726–1736. [Google Scholar]
- Song, P.; Cai, C.; Skokut, M.; Kosegi, B.; Petolino, J. Quantitative real-time PCR as a screening tool for estimating transgene copy number in WHISKERS™-derived transgenic maize. Plant Cell Rep 2002, 20, 948–954. [Google Scholar]
- Mason, G.; Provero, P.; Vaira, A.M.; Accotto, G.P. Estimating the number of integrations in transformed plants by quantitative real-time PCR. BMC Biotechnol 2002, 2, 20–30. [Google Scholar]
- D’Hont, A.; Grivet, L.; Feldmann, P.; Rao, S.; Berding, N.; Glaszmann, J.C. Characterisation of the double genome structure of modern sugarcane cultivars (Saccharum spp.) by molecular cytogenetics. Mol. Gen. Genet. MGG 1996, 250, 405–413. [Google Scholar]
- Edmé, S.J.; Comstock, J.C.; Miller, J.D.; Tai, P.Y.P. Determination of DNA content and genome size in sugarcane. Am. Soc. Sugar Cane Technol 2005, 25, 1–16. [Google Scholar]
- Yang, L.T.; Chen, J.X.; Huang, C.; Liu, Y.H.; Jia, S.R.; Pan, L.W.; Zhang, D.B. Validation of a cotton-specific gene, Sad1, used as an endogenous reference gene in qualitative and real-time quantitative PCR detection of transgenic cottons. Plant Cell Rep 2005, 24, 237–245. [Google Scholar]
- Schefe, J.H.; Lehmann, K.E.; Buschmann, I.R.; Unger, T.; Funke-Kaiser, H. Quantitative real-time RT-PCR data analysis: Current concepts and the novel “gene expression’s CT difference” formula. J. Mol. Med 2006, 84, 901–910. [Google Scholar]
- Li, H.; Lv, X.W.; Wang, J.; Li, J.G.; Yang, H.F.; Qin, Y.C. Quantitative determination of soybean meal content in compound feeds: Comparison of near-infrarded spectroscopy and real-time PCR. Anal. Bioanal. Chem 2007, 389, 2313–2322. [Google Scholar]
- Randhawa, G.J.; Singh, M.; Sharma, R. Validation of ST-LS1 as an endogenous reference gene for detection of AmA1 and cry1Ab genes in genetically modified potatoes using multiplex and real time PCR. Am. J. Potato Res 2009, 86, 398–405. [Google Scholar]
- Shou, H.X.; Frame, B.R.; Whitham, S.A.; Wang, K. Assessment of transgenic maize events produced by particle bombardment or Agrobacterium- mediated transformation. Mol. Breed 2004, 13, 201–208. [Google Scholar]
- Shen, H.F.; Qian, B.J.; Yang, L.T.; Liang, W.Q.; Chen, W.W.; Liu, Z.H.; Zhang, D.B. Estimation of the homoplasmy degree for transplastomic tobacco using quantitative real-time PCR. Eur. Food Res. Technol 2010, 231, 143–150. [Google Scholar]
- Sellars, M.J.; Vuocolo, T.; Leeton, L.A.; Coman, G.J.; Degnan, B.M.; Preston, N.P. Real-time RT-PCR quantification of Kuruma shrimp transcripts: A comparison of relative and absolute quantification procedures. J. Biotechnol 2007, 129, 391–399. [Google Scholar]
- Leong, D.T.; Gupta, A.; Bai, H.F.; Wan, G.Q.; Yoong, L.F.; Too, H.P.; Chew, F.T.; Hutmacher, D.W. Absolute quantification of gene expression in biomaterials research using real-time PCR. Biomaterials 2007, 28, 203–210. [Google Scholar]
- Bubner, B.; Gase, K.; Baldwin, I.T. Two fold differences are the detection limit for determining transgene copy numbers in plants by real-time PCR. BMC Biotechnol 2004, 4. [Google Scholar] [CrossRef] [Green Version]
- Weng, H.B.; Pan, A.H.; Yang, L.H.; Zhang, C.M.; Liu, Z.L.; Zhang, D.B. Estimating number of transgene copies in transgenic rapeseed by real-time PCR assay with HMG I/Y as an endogenous reference gene. Plant Mol. Biol. Rep 2004, 22, 289–300. [Google Scholar]
- Yang, L.T.; Ding, J.Y.; Zhang, C.M.; Jia, J.W.; Weng, H.B.; Liu, W.X.; Zhang, D.B. Estimating the copy number of transgenes in transformed rice by real-time quantitative PCR. Plant Cell Rep 2005, 23, 759–763. [Google Scholar]
- Yang, L.T.; Pan, A.H.; Zhang, K.W.; Yin, C.S.; Qian, B.J.; Chen, J.X.; Huang, C.; Zhang, D.B. Qualitative and quantitative PCR methods for event-specific detection of genetically modified cotton Mon1445 and Mon531. Transgenic Res 2005, 14, 817–831. [Google Scholar]
- Huang, H.L.; Cheng, F.; Wang, R.A.; Zhang, D.B.; Yang, L.T. Evaluation of four endogenous reference genes and their real-time PCR assay for common wheat quantification in GMOs detection. PLoS One 2013, 8, e75850. [Google Scholar]
- James, D.; Schmidt, A.M.; Wall, E.; Green, M.; Masri, S. Reliable detection and identification of genetically modified maize,soybean and canola by multiplex PCR analysis. J. Agric. Food Chem 2003, 51, 5829–5834. [Google Scholar]
- Hernández, M.; Duplan, M.N.; Berthier, G.; Vaïtilingom, M.; Hauser, W.; Freyer, R.; Pla, M.; Bertheau, Y. Development and comparison of four real-time polymerase chain reaction systems for specific detection and quantification of Zea mays L. J. Agric. Food Chem 2004, 52, 4632–4637. [Google Scholar]
- Yang, L.T.; Guo, J.C.; Zhang, H.B.; Liu, J.; Zhang, D.B. Qualitative and quantitative event-specific PCR detection methods for oxy235 canola based on the 3 integration flanking sequence. J. Agric. Food Chem 2008, 56, 1804–1809. [Google Scholar]
- Demeke, T.; Ratnayaka, I. Multiplex qualitative PCR assay for identification of genetically modified canola events and real-time event specific PCR assay for quantification of the GT73 canola event. Food Control 2008, 19, 893–897. [Google Scholar]
- Chaouachi, M.; El Malki, R.; Berard, A.; Romaniuk, M.; Laval, V.; Brunel, D.; Bertheau, Y. Development of a real-time PCR method for the diferential detection and quantification of four solanaceae in GMO analysis: potato (Solanum tuberosum), tomato (Solarium lycopersicum), eggplant (Solanum melongena) and pepper (Capsicum annuum). J. Agric. Food Chem 2008, 56, 1818–1828. [Google Scholar]
- Zhang, J.S.; Nagai, C.; Yu, Q.Y.; Pan, Y.B.; Ayala-Silva, T.; Schnell, R.J.; Comstock, J.C.; Arumuganathan, A.K.; Ming, R. Genome size variation in three Saccharum species. Euphytica 2012, 180, 511–519. [Google Scholar]
- Fraley, R.T.; Rogers, S.G.; Horsch, R.B.; Sanders, P.R.; Flick, J.S.; Adams, S.P.; Bittner, M.L.; Brand, L.A.; Fink, C.L.; Fry, J.S.; et al. Expression of bacterial genes in plant cells. Proc. Natl. Acad. Sci.USA 1983, 80, 4803–4807. [Google Scholar]
- Kok, E.J.; Kuiper, H.A. Comparative safety assessment for biotech crops. Trends Biotechnol 2003, 21, 439–444. [Google Scholar]
- Faclo, M.C.; Neto, A.T.; Ulian, E.C. Transformation and expression of a gene for herbicide resistance in a Brazilian sugarcane. Plant Cell Rep 2000, 19, 1188–1194. [Google Scholar]
- Yao, W.; Yu, A.L.; Xu, J.S.; Geng, G.L.; Zhang, M.Q.; Chen, R.C. Analysis and identification for transgenic sugarcane of ScMV-CP gene. Mol. Plant Breed 2004, 2, 13–18. [Google Scholar]
- Chen, R.K.; Chen, P.H.; Lin, M.J.; Xue, Z.P. A study on genetic transformation of GAN gene in sugarcane. Acta Agric. Univ. Jiangxiensis 2004, 26, 740–743. [Google Scholar]
- Chen, P.H.; Xu, L.P.; Chen, R.K. Construction of expression vector of nematode-resistant gene and transformation of sugarcane. Chin. J. Ecol.-Agric 2004, 12, 57–59. [Google Scholar]
- Wang, J.H.; Cao, G.; Zhang, M.Q. Physiological characteristics of Bt transgenic sugarcane. Chin. J. Trop. Crops 2011, 32, 1864–1867. [Google Scholar]
- Xu, W.T.; Bai, W.B.; Guo, F.; Luo, Y.B.; Yuan, Y.F.; Huang, K.L. A papaya-specific gene, papain, used as an endogenous reference gene in qualitative and real-time quantitative PCR detection of transgenic papayas. Eur. Food Res. Technol 2008, 228, 301–309. [Google Scholar]
- Beltrán, J.; Jaimes, H.; Echeverry, M.; Ladino, Y.; López, D.; Duque, M.C.; Chavarriaga, P.; Tohme, J. Quantitative analysis of transgenes in cassava plants using real-time PCR technology. In Vitro Cell. Dev. Biol.-Plant 2009, 45, 48–56. [Google Scholar]
- Fan, M.J.; Chen, S.; Kung, Y.J.; Cheng, Y.H.; Bau, H.J.; Su, T.T.; Yeh, S.D. Transgene-specific and event-specific molecular markers for characterization of transgenic papaya lines resistant to Papaya ringspot virus. Transgenic Res 2009, 18, 971–986. [Google Scholar]
- Paterson, A.H.; Brubaker, C.L.; Wendel, J.F. A rapid method for extraction of cotton (Gossypium spp.) genomic DNA suitable for RELP or PCR analsis. Plant Mol. Biol. Rep 1993, 11, 122–127. [Google Scholar]
- Li, H.F.; Li, L.; Zhang, L.J.; Xu, Y.L.; Li, W.F. Standard curve generation of PepT1 gene for absolute quantification using real-time PCR. J. Shanxi Agric. Univ. 2010, 30, 332–335. [Google Scholar]
- Li, L.; Zhao, C.P.; Li, H.L.; Wu, F.; Zhang, L.H.; Xu, D.M.; Wang, J.S.; Li, H.F. Establishment of the plasmid standard curve generation method for absolute quantification PCR. J. Agric. Biotechnol 2011, 19, 1157–1162. [Google Scholar]
- Lee, C.; Kim, J.; Shin, S.G.; Hwang, S. Absolute and relative qPCR quantification of plasmid copy number in Escherichia coli. J. Biotechnol 2006, 123, 273–280. [Google Scholar]
- Grivet, L.; Arruda, P. Sequence polymorphism from EST data in sugarcane: A fine analysis of 6-phosphogluconate dehydrogenase genes. Genet. Mol. Biol 2001, 24, 161–167. [Google Scholar]
- Li, F.W.; Shao, G.G.; Xing, Z.J.; Ling, C.C.; Xia, W.; Zhang, M. Constuction of the plasmid reference molecule for detection of transgenic Soybean MON89788. Agric. Sci. Technol 2010, 11, 55–58. [Google Scholar]
- Su, C.Q.; Xie, J.J.; Sun, Y.; Peng, Y.F. Construction and application of a reference plasmid suitable for determination of CpTI and cry1A gene dosages in genetically modified cottons. Acta Agron. Sin 2011, 37, 1533–1539. [Google Scholar]
| Gene | Primer pair # | Primer sequence | Product size/bp | Source sequence ID |
|---|---|---|---|---|
| P4H | P4H-1* | F: 5′-GCGACATCAGAACAGTGTGAA-3′ R: 5′-TTGTACTCTCCGCGGTTTCT-3′ | 100 | sb01g007280* |
| P4H-2 | F: 5′-GTCCGTAATCCCATACCAGATTTT-3′ R: 5′-CACACTGTTCTGATGTCGCAAA-3′ | 80 | CA107003, CA106927, CA209790, CA131236 | |
| P4H-3 | F: 5′-GTGAAAATATAGTAAAAACTGCGAAGGA-3′ R: 5′-TTGTACTCTCCGCGGTTTCTC-3′ | 84 | ||
| ENOL | ENOL-1* | F: 5′-TCCTTACAAAGGATGGGAGC-3′ R: 5′-TGTACAGATCACCCAGACGC-3′ | 96 | sb02g023480* |
| ENOL-2 | F: 5′-TTTTGATCAGGATGACTGGAGTTC-3′ R: 5′-AAATCATCACCCACAATTTGGAT-3′ | 75 | CA229198, CA131106 | |
| ENOL-3 | F: 5′-GGACCCTTTTGATCAGGATGAC-3′ R: 5′-AATCATCACCCACAATTTGGATATC-3′ | 80 | ||
| TST | TST-1* | F: 5′-ACATGCTGCCATCTGAAAAG-3′ R: 5′-CAGCCCCTTTCCATCATAAA-3′ | 95 | sb08g020860* |
| TST-2 | F: 5′-GCTGTTCAGTGCTGCTCGTGTT-3′ R: 5′-GGCAAACCTCCATCTAACACCC-3′ | 81 | ca182497, ca095999, ca251048, ca176811, ca076277, cf575388, ca085087, ca205081, ca074129, ca076192 | |
| TST-3 | F: 5′-TCAGTGCTGCTCGTGTTTGGT-3′ R: 5′-ATGGCATCACTGGAGGCACTT-3′ | 130 | ||
| TST-4 | F: 5′-AAGTGCCTCCAGTGATGCCA-3′ R: 5′-ACCTGAGGAAACGGAACGCA-3′ | 200 | ||
| APRT | APRT-1* | F: 5′-TGACACATTCCCAACCTCAA-3′ R: 5′-ATCTCTCTCCCTGAGTGGCA-3′ | 119 | sb02g033370* |
| APRT-2 | F: 5′-AGGGAAGTGGTTCGGTGATG-3′ R: 5′-TGATAAAGAGCACATGAACCAACA-3′ | 74 | CA089504, CA089592, CA146761, CA150154, DV640571 | |
| CYC | CYC-1* | F:5′-CTCATGGAAAACTTACCGGG-3′ R:5′-TGCATCCAGCAAGAAAGTTG-3′ | 95 | sb10g030790* |
| CYC-2 | F:5′-ACTGATGACATTCCCTTGCCTAT-3′ R:5′-CCGACAGCATAGGCAAGGGA-3′ | 226 | ca176931, ca235106, ca256302, ca101120, bu103284, ca289292, ca074749, ca253051, ca126707 | |
| CYC-3 | R:5′-ATAGTCAACCACAGCCAGGGA-3′ F:5′-AACCAACCTACGGTTGCCCA-3′ | 114 | ||
| PRR | PRR-1* | F: 5′-GCCAAATTCAGGCAGAAAAG-3′ R: 5′-CACCCTAGGCCTTGTTTCAG-3′ | 93 | sb04g026190* |
| PRR-2 | F: 5′-GCACCACCCTCCTCTCAGAC-3′ R: 5′-AATGAGCTGGTGGTTGGGGT-3′ | 261 | ca275375, ca297639, ca134882, ca246262, ca245313, ca083303 | |
| PRR-3 | F: 5′-ACCAATAGCACCACCCTCCTC-3′ R: 5′-GACGACCCAGCAACCCTCAG-3′ | 239 | ||
| PRR-4 | F: 5′-ACCAATAGCACCACCCTCCTC-3′ R: 5′-GACGACTCAGCAATCCTCAG-3′ | 239 |
| Gene | Primer pairs | Standard curve formula | R2 | E |
|---|---|---|---|---|
| ROC22 Badila | ROC22 Badila | ROC22 Badila | ||
| P4H | P4H-1 | y = −3.212 x + 35.610 y = −3.051 x + 35.328 | 0.989 0.996 | 1.048 1.127 |
| P4H-3 | y = −3.275 x + 37.509 y = −3.194 x + 36.323 | 0.998 0.993 | 1.020 1.056 | |
| APRT | APRT-1 | y = −3.174 x + 35.962 y = −2.918 x + 34.326 | 0.996 0.994 | 1.066 1.200 |
| APRT-2 | y = −3.328 x + 37.704 y = −3.202 x + 39.339 | 0.998 0.999 | 0.998 1.053 | |
| CYC | CYC-1 | y = −2.321 x + 32.405 y = −2.218 x + 37.622 | 0.955 0.816 | 1.697 1.824 |
| CYC-2 | y = −3.047 x + 34.961 y = −3.159 x + 37.622 | 0.992 0.998 | 1.129 1.073 | |
| TST | TST-1 | y = −3.272 x + 38.265 y = −3.402 x + 39.975 | 0.994 0.999 | 1.021 0.968 |
| TST-3 | y = −3.161 x + 37.308 y = −3.410 x + 38.854 | 0.999 0.999 | 1.072 0.965 | |
| ENOL | ENOL-3 | y = −3.214 x + 37.237 y = −3.403 x + 44.813 | 0.999 0.999 | 1.047 0.967 |
| PRR | PRR-1 | y = −3.185 x + 35.916 y = −3.514 x + 40.872 | 0.996 0.995 | 1.061 0.926 |
| Primer pair | Corresponding gene | Ct value | Copy number ± SE |
|---|---|---|---|
| ROC 22 Badila | ROC 22 Badila | ||
| P4H-3 | P4H | 23.259 22.725 | 10.47 ± 0.04 C 6.28 ± 0.36 B |
| ENOL-3 | ENOL | 21.050 24.220 | 47.68 ± 0.12 A 389.26 ± 1.09 A |
| TST-1 | TST | 21.689 24.558 | 51.14 ± 0.21 A 11.76 ± 0.17 B |
| APRT-2 | APRT | 22.952 24.874 | 8.39 ± 0.27 C 11.41 ± 0.55 B |
| CYC-2 | CYC | 21.574 23.678 | 11.07 ± 0.13 C 9.00 ± 0.53 B |
| PRR-1 | PRR | 21.143 23.920 | 19.07 ± 0.23 B 23.06 ± 0.29 B |
| Sugarcane genotype | Copy number ± SE | ||
|---|---|---|---|
| P4H–P4H-3 | APRT–APRT-2 | CYC–CYC-2 | |
| Black Cheribon (Yunnan) (Saccharum officinarum) | 3.58 ± 0.01 | 4.73 ± 0.47 | 4.97 ± 0.07 |
| Black Cheribon (Fujian) (S. officinarum) | 7.32 ± 0.78 | 9.69 ± 0.16 | 9.67 ± 0.17 |
| Loethers (S. officinarum) | 6.37 ± 0.13 | 9.24 ± 1.03 | 10.06 ± 0.24 |
| Crystalina (S. officinarum) | 3.18 ± 0.02 | 5.03 ± 0.53 | 4.28 ± 0.28 |
| SES208 (S. spontaneum) | 7.20 ± 0.66 | 11.82 ± 0.24 | 10.95 ± 0.16 |
| LA Purple (S. officinarum) | 4.44 ± 0.13 | 7.62 ± 0.95 | 6.87 ± 0.21 |
| Q117 (S. spp. hybrids) | 20.81 ± 0.53 | 22.14 ± 0.27 | 28.45 ± 0.10 |
| YCE05-179 (S. spp. hybrids) | 21.51 ± 0.17 | 13.18 ± 0.12 | 21.35 ± 0.72 |
| Gene | R2 | E | Copy number ± SE | Relative abundance value | ||
|---|---|---|---|---|---|---|
| ROC22 Badila | ROC22 Badila | |||||
| P4H | 0.999 | 1.040 | 2.91 ± 0.09 | 2.12 ± 0.06 | 1.5 | 1.0 |
| APRT | 0.999 | 1.049 | 2.45 ± 0.09 | 2.21 ± 0.13 | 1.0 | 1.0 |
| CYC | 0.998 | 1.047 | 4.27 ± 0.08 | 3.05 ± 0.18 | 2.0 | 1.5 |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Xue, B.; Guo, J.; Que, Y.; Fu, Z.; Wu, L.; Xu, L. Selection of Suitable Endogenous Reference Genes for Relative Copy Number Detection in Sugarcane. Int. J. Mol. Sci. 2014, 15, 8846-8862. https://doi.org/10.3390/ijms15058846
Xue B, Guo J, Que Y, Fu Z, Wu L, Xu L. Selection of Suitable Endogenous Reference Genes for Relative Copy Number Detection in Sugarcane. International Journal of Molecular Sciences. 2014; 15(5):8846-8862. https://doi.org/10.3390/ijms15058846
Chicago/Turabian StyleXue, Bantong, Jinlong Guo, Youxiong Que, Zhiwei Fu, Luguang Wu, and Liping Xu. 2014. "Selection of Suitable Endogenous Reference Genes for Relative Copy Number Detection in Sugarcane" International Journal of Molecular Sciences 15, no. 5: 8846-8862. https://doi.org/10.3390/ijms15058846
APA StyleXue, B., Guo, J., Que, Y., Fu, Z., Wu, L., & Xu, L. (2014). Selection of Suitable Endogenous Reference Genes for Relative Copy Number Detection in Sugarcane. International Journal of Molecular Sciences, 15(5), 8846-8862. https://doi.org/10.3390/ijms15058846





